Deacetylation of Histone H4 Accompanying Cardiomyogenesis is Weakened in HDAC1-Depleted ES Cells
Abstract
:1. Introduction
2. Results
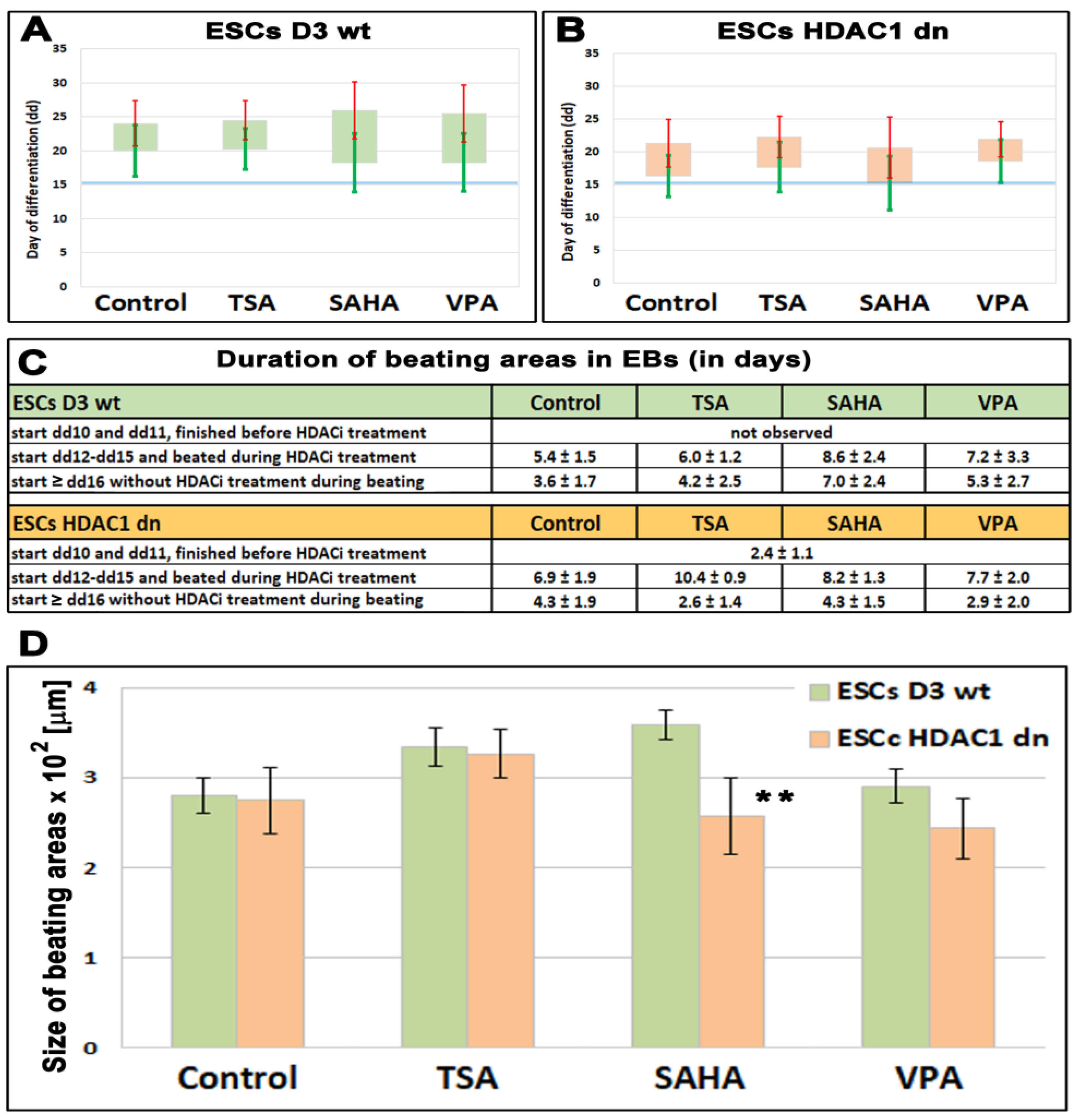
2.1. The Beating of Cardiomyocytes Studied in wt and HDAC1 dn Cells
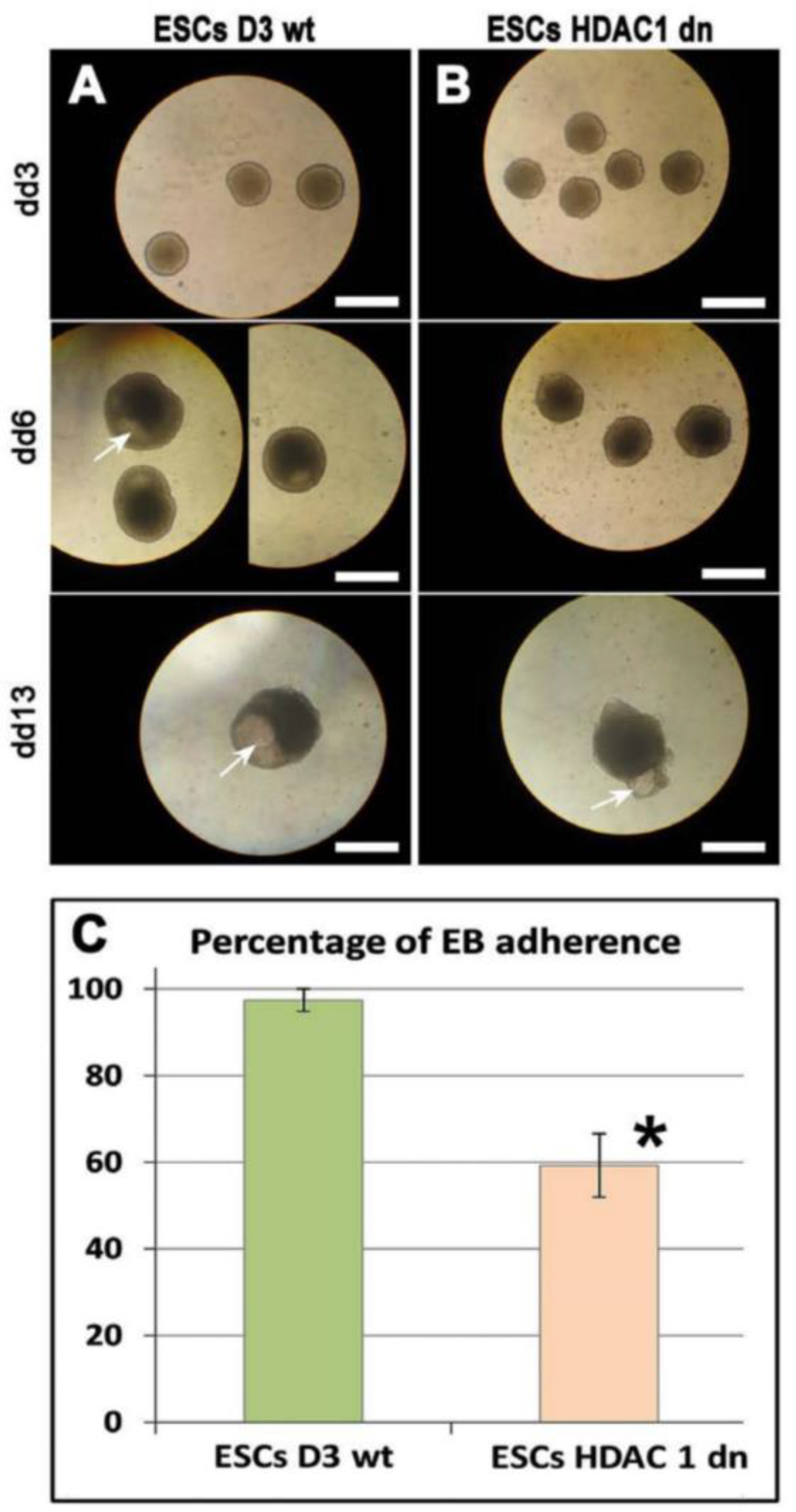
2.2. Adherence of Embryonic Bodies Is Affected by HDAC1 Depletion
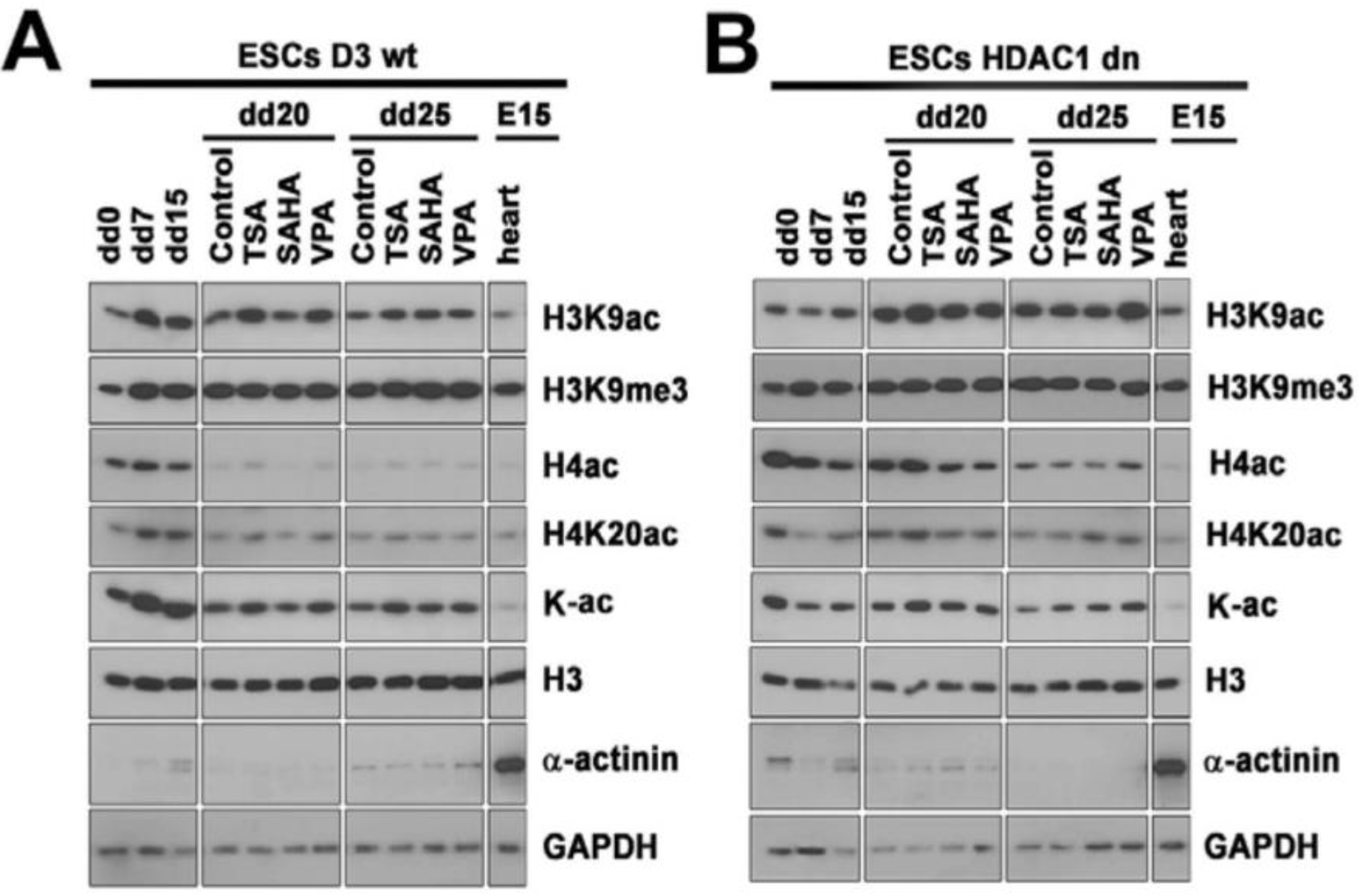
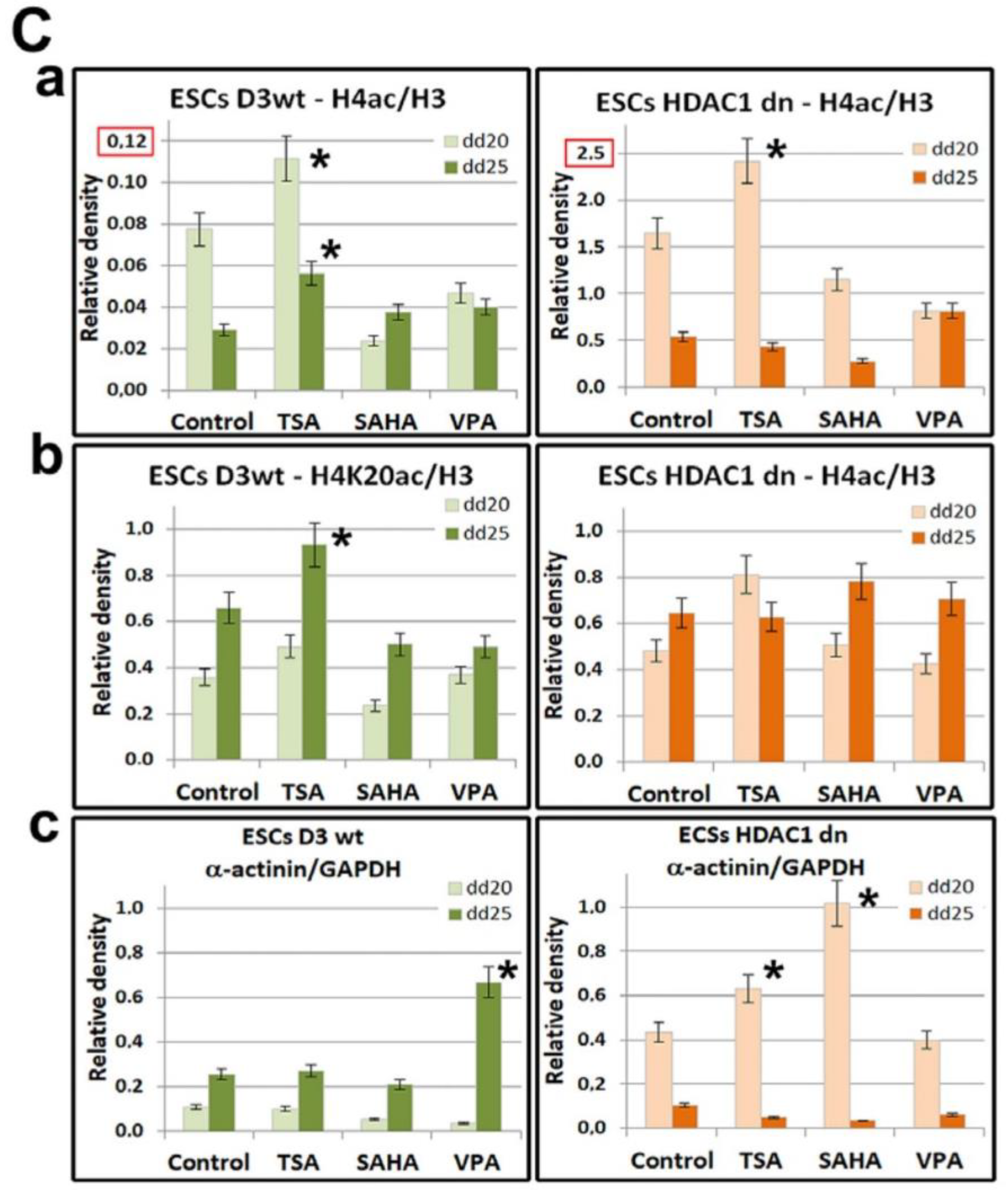
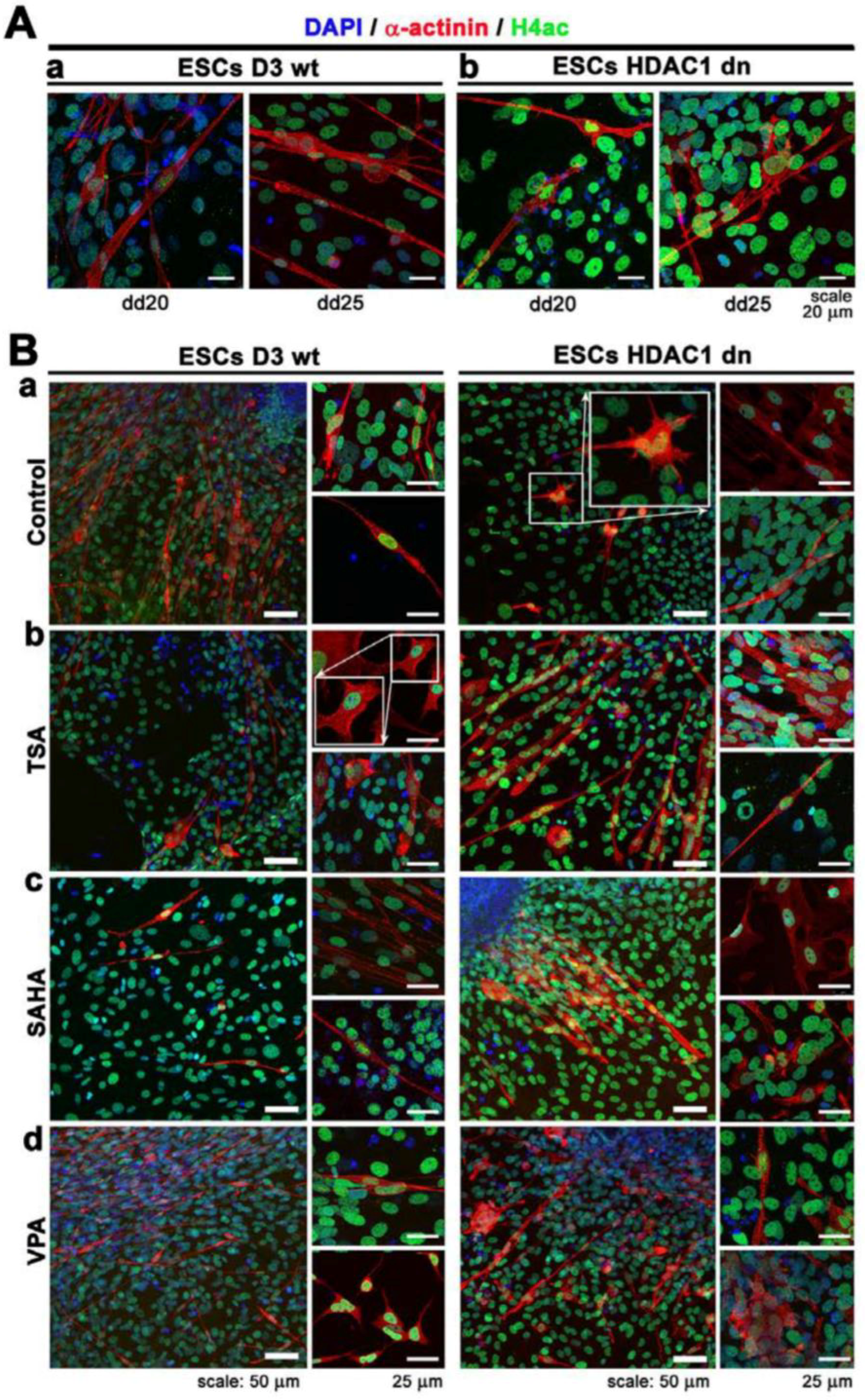
2.3. Differentiation-Specific Deacetylation of Histone H4 was Weakened in HDAC1-Depleted Cells
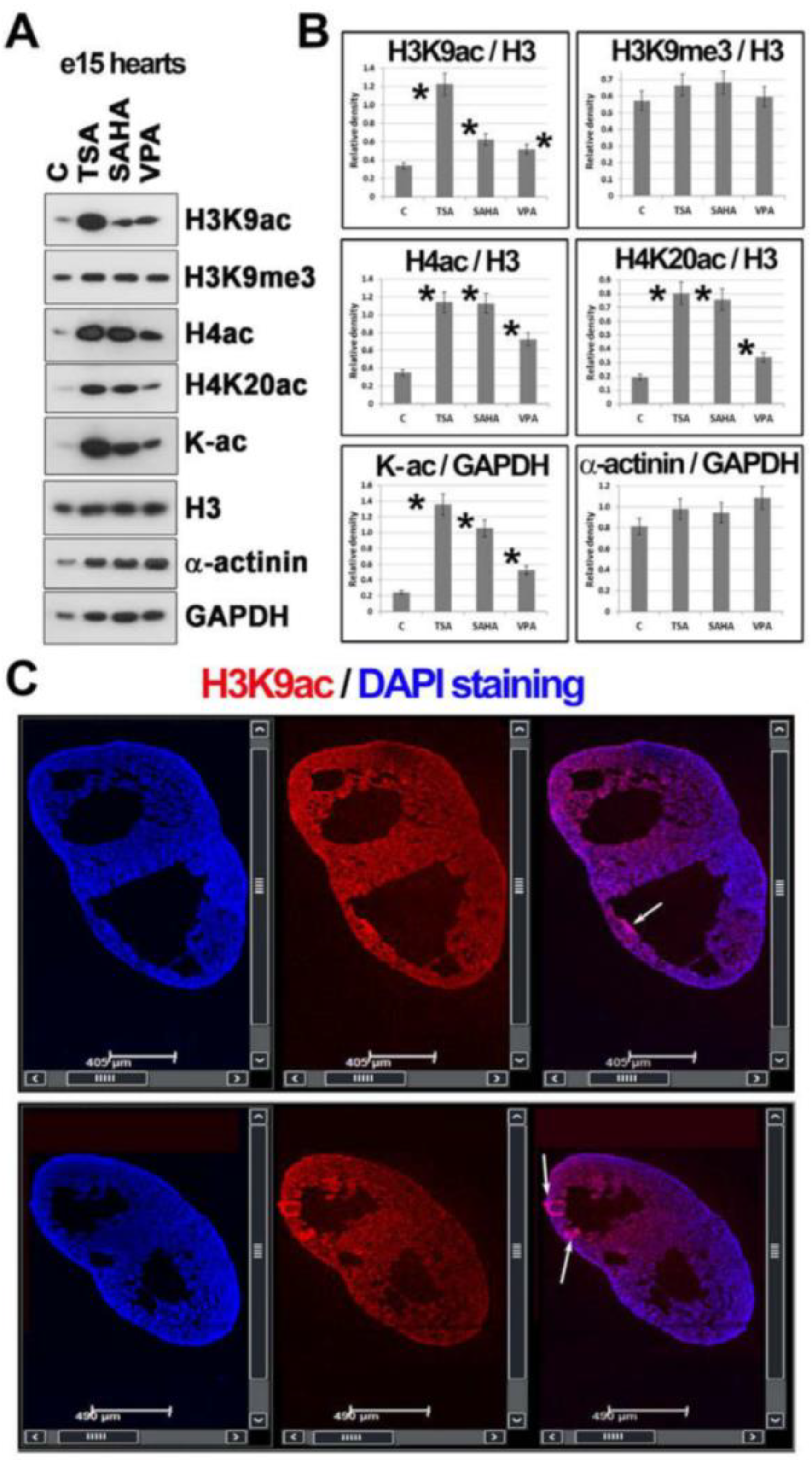
2.4. Histone Hyperacetylation Can Be Induced by HDACs Inhibitors in Hearts Explanted from Mouse Embryos at Stage e15
3. Discussion
4. Materials and Methods
4.1. Mouse ESCs Cultivation and Differentiation
4.2. Experimental Animals
4.3. Immunostaining
4.4. Western Blot Analysis
4.5. Confocal Microscopy
4.6. The Tile-Scanning
4.7. Statistical Analysis, Image Analysis, and Image Processing
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| H3 | histone H3 |
| H4 | histone H4 |
| me1 | mono-methylated histones |
| me2 | di-methylated histones |
| me3 | tri-methylated histones |
| H3K9me1/me2/me3 | histone H3 methylated on lysine 9 (mono-; di-; tri-methylation) |
| H3K9ac | histone H3 acetylated on lysine 9 |
| HDAC | histone deacetylases; HDAC1: histone deacetylase 1 |
| HDACi | histone deacetylase inhibitors |
| Kac | lysine acetylation |
| PTM | post-translational modification |
References
- Kimes, B.W.; Brandt, B.L. Properties of a clonal muscle cell line from rat heart. Exp. Cell Res. 1976, 98, 367–381. [Google Scholar] [CrossRef]
- Jaffredo, T.; Chestier, A.; Bachnou, N.; Dieterlen-Lièvre, F. MC29-immortalized clonal avian heart cell lines can partially differentiate in vitro. Exp. Cell Res. 1991, 192, 481–491. [Google Scholar] [CrossRef]
- Field, L.J. Transgenic mice in cardiovascular research. Annu. Rev. Physiol. 1993, 55, 97–114. [Google Scholar] [CrossRef] [PubMed]
- Sen, A.; Dunnmon, P.; Henderson, S.A.; Gerard, R.D.; Chien, K.R. Terminally differentiated neonatal rat myocardial cells proliferate and maintain specific differentiated functions following expression of SV40 large T. antigen. J. Biol. Chem. 1988, 263, 19132–19136. [Google Scholar] [PubMed]
- White, S.M.; Claycomb, W.C. Embryonic stem cells form an organized, functional cardiac conduction system in vitro. Am. J. Physiol. Heart Circ. Physiol. 2005, 288, H670–H679. [Google Scholar] [CrossRef] [PubMed]
- Wobus, A.M.; Kaomei, G.; Shan, J.; Wellner, M.C.; Rohwedel, J.; Guanju, J.; Fleischmann, B.; Katus, H.A.; Hescheler, J.; Franz, W.M. Retinoic acid accelerates embryonic stem cell-derived cardiac differentiation and enhances development of ventricular cardiomyocytes. J. Mol. Cell. Cardiol. 1997, 29, 1525–1539. [Google Scholar] [CrossRef] [PubMed]
- Wobus, A.M. Potential of embryonic stem cells. Mol. Aspects Med. 2001, 22, 149–164. [Google Scholar] [CrossRef]
- Rohwedel, J.; Maltsev, V.; Bober, E.; Arnold, H.H.; Hescheler, J.; Wobus, A.M. Muscle cell differentiation of embryonic stem cells reflects myogenesis in vivo: Developmentally regulated expression of myogenic determination genes and functional expression of ionic currents. Dev. Biol. 1994, 164, 87–101. [Google Scholar] [CrossRef] [PubMed]
- Strübing, C.; Ahnert-Hilger, G.; Shan, J.; Wiedenmann, B.; Hescheler, J.; Wobus, A.M. Differentiation of pluripotent embryonic stem cells into the neuronal lineage in vitro gives rise to mature inhibitory and excitatory neurons. Mech. Dev. 1995, 53, 275–287. [Google Scholar] [CrossRef]
- Risau, W.; Sariola, H.; Zerwes, H.-G.; Sasse, J.; Ekblom, P.; Kemler, R.; Doetschman, T. Vasculogenesis and angiogenesis in embryonic-stem-cell-derived embryoid bodies. Development 1988, 102, 471–478. [Google Scholar] [PubMed]
- Bagutti, C.; Wobus, A.M.; Fässler, R.; Watt, F.M. Differentiation of embryonal stem cells into keratinocytes: Comparison of wild-type and β1integrin-deficient cells. Dev. Biol. 1996, 179, 184–196. [Google Scholar] [CrossRef] [PubMed]
- Fassler, R.; Rohwedel, J.; Maltsev, V.; Bloch, W.; Lentini, S.; Guan, K.; Gullberg, D.; Hescheler, J.; Addicks, K.; Wobus, A.M. Differentiation and integrity of cardiac muscle cells are impaired in the absence of beta1 integrin. J. Cell Sci. 1996, 109, 2989–2999. [Google Scholar] [PubMed]
- Fässler, R.; Sasaki, T.; Timpl, R.; Chu, M.L.; Werner, S. Differential regulation of fibulin, tenascin-C., and nidogen expression during wound healing of normal and glucocorticoid-treated mice. Exp. Cell Res. 1996, 222, 111–116. [Google Scholar] [CrossRef] [PubMed]
- Hescheler, J.; Fleischmann, B.K.; Lentini, S.; Maltsev, V.A.; Rohwedel, J.; Wobus, A.M.; Addicks, K. Embryonic stem cells: A model to study structural and functional properties in cardiomyogenesis. Cardiovasc. Res. 1997, 36, 149–162. [Google Scholar] [CrossRef]
- Davies, M.P.; An, R.H.; Doevendans, P.; Kubalak, S.; Chien, K.R.; Kass, R.S. Developmental changes in ionic channel activity in the embryonic murine heart. Circ. Res. 1996, 78, 15–25. [Google Scholar] [CrossRef] [PubMed]
- Schaub, M.C.; Hefti, M.A.; Zuellig, R.A.; Morano, I. Modulation of contractility in human cardiac hypertrophy by myosin essential light chain isoforms. Cardiovasc. Res. 1998, 37, 381–404. [Google Scholar] [CrossRef] [Green Version]
- Rodenhiser, D.; Mann, M. Epigenetics and human disease: Translating basic biology into clinical applications. Can. Med. Assoc. J. 2006, 174, 341–348. [Google Scholar] [CrossRef] [PubMed]
- Yanazume, T.; Hasegawa, K.; Morimoto, T.; Kawamura, T.; Wada, H.; Matsumori, A.; Kawase, Y.; Hirai, M.; Kita, T. Cardiac p300 is involved in myocyte growth with decompensated heart failure. Mol. Cell. Biol. 2003, 23, 3593–3606. [Google Scholar] [CrossRef] [PubMed]
- Hohl, M.; Wagner, M.; Reil, J.C.; Müller, S.A.; Tauchnitz, M.; Zimmer, A.M.; Lehmann, L.H.; Thiel, G.; Böhm, M.; Backs, J.; Maack, C. HDAC4 controls histone methylation in response to elevated cardiac load. J. Clin. Investig. 2013, 123, 1359–1370. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Banerjee, A. A review of family history of cardiovascular disease: Risk factor and research tool. Int. J. Clin. Pract. 2012, 66, 536–543. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization; Mendis, S.; Puska, P.; Norrving, B. Global Atlas on Cardiovascular Disease Prevention and Control; World Health Organization in collaboration with the World Heart Federation and the World Stroke Organization, 2011; ISBN 9789240687905. [Google Scholar]
- Ordovás, J.M.; Smith, C.E. Epigenetics and cardiovascular disease. Nat. Rev. Cardiol. 2010, 7, 510–519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gusterson, R.J.; Jazrawi, E.; Adcock, I.M.; Latchman, D.S. The transcriptional co-activators CREB-binding protein (CBP) and p300 play a critical role in cardiac hypertrophy that is dependent on their histone acetyltransferase activity. J. Biol. Chem. 2003, 278, 6838–6847. [Google Scholar] [CrossRef] [PubMed]
- Antos, C.L.; McKinsey, T.A.; Dreitz, M.; Hollingsworth, L.M.; Zhang, C.L.; Schreiber, K.; Rindt, H.; Gorczynski, R.J.; Olson, E.N. Dose-dependent blockade to cardiomyocyte hypertrophy by histone deacetylase inhibitors. J. Biol. Chem. 2003, 278, 28930–28937. [Google Scholar] [CrossRef] [PubMed]
- Cao, D.J.; Wang, Z.V.; Battiprolu, P.K.; Jiang, N.; Morales, C.R.; Kong, Y.; Rothermel, B.A.; Gillette, T.G.; Hill, J.A. Histone deacetylase (HDAC) inhibitors attenuate cardiac hypertrophy by suppressing autophagy. Proc. Natl. Acad. Sci. USA 2011, 108, 4123–4128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, F.; Levin, M.D.; Petrenko, N.B.; Lu, M.M.; Wang, T.; Yuan, L.J.; Stout, A.L.; Epstein, J.A.; Patel, V.V. Histone-deacetylase inhibition reverses atrial arrhythmia inducibility and fibrosis in cardiac hypertrophy independent of angiotensin. J. Mol. Cell. Cardiol. 2008, 45, 715–723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montgomery, R.L.; Davis, C.A.; Potthoff, M.J.; Haberland, M.; Fielitz, J.; Qi, X.; Hill, J.A.; Richardson, J.A.; Olson, E.N. Histone deacetylases 1 and 2 redundantly regulate cardiac morphogenesis, growth, and contractility. Genes Dev. 2007, 21, 1790–1802. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Montgomery, R.L.; Potthoff, M.J.; Haberland, M.; Qi, X.; Matsuzaki, S.; Humphries, K.M.; Richardson, J.A.; Bassel-Duby, R.; Olson, E.N. Maintenance of cardiac energy metabolism by histone deacetylase 3 in mice. J. Clin. Investig. 2008, 118, 3588–3597. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McKinsey, T.A. Therapeutic Potential for HDAC Inhibitors in the Heart. Annu. Rev. Pharmacol. Toxicol. 2012, 52, 303–319. [Google Scholar] [CrossRef] [PubMed]
- Kawamura, T.; Ono, K.; Morimoto, T.; Wada, H.; Hirai, M.; Hidaka, K.; Morisaki, T.; Heike, T.; Nakahata, T.; Kita, T.; et al. Acetylation of GATA-4 is involved in the differentiation of embryonic stem cells into cardiac myocytes. J. Biol. Chem. 2005, 280, 19682–19688. [Google Scholar] [CrossRef] [PubMed]
- Iezzi, S.; Di Padova, M.; Serra, C.; Caretti, G.; Simone, C.; Maklan, E.; Minetti, G.; Zhao, P.; Hoffman, E.P.; Puri, P.L.; et al. Deacetylase inhibitors increase muscle cell size by promoting myoblast recruitment and fusion through induction of follistatin. Dev. Cell 2004, 6, 673–684. [Google Scholar] [CrossRef]
- Eom, G.H.; Nam, Y.S.; Oh, J.G.; Choe, N.; Min, H.K.; Yoo, E.K.; Kang, G.; Nguyen, V.H.; Min, J.J.; Kim, J.K.; et al. Regulation of acetylation of histone deacetylase 2 by p300/CBP-associated factor/histone deacetylase 5 in the development of cardiac hypertrophy. Circ. Res. 2014, 114, 1133–1143. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Wu, C.T.; Qi, X.Y.; Meijering, R.A.M.; Hoogstra-Berends, F.; Tadevosyan, A.; Deniz, G.C.; Durdu, S.; Akar, A.R.; Sibon, O.C.M.; et al. Activation of histone deacetylase-6 induces contractile dysfunction through derailment of α-tubulin proteostasis in experimental and human atrial fibrillation. Circulation 2014, 129, 346–358. [Google Scholar] [CrossRef] [PubMed]
- Glozak, M.A.; Sengupta, N.; Zhang, X.; Seto, E. Acetylation and deacetylation of non-histone proteins. Gene 2005, 363, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Tao, R.; De Zoeten, E.F.; Özkaynak, E.; Chen, C.; Wang, L.; Porrett, P.M.; Li, B.; Turka, L.A.; Olson, E.N.; Greene, M.I.; et al. Deacetylase inhibition promotes the generation and function of regulatory T. cells. Nat. Med. 2007, 13, 1299–1307. [Google Scholar] [CrossRef] [PubMed]
- Doetschman, T.C.; Eistetter, H.; Katz, M.; Schmidt, W.; Kemler, R. The in vitro development of blastocyst-derived embryonic stem cell lines: Formation of visceral yolk sac, blood islands and myocardium. J. Embryol. Exp. Morphol. 1985, 87, 27–45. [Google Scholar] [CrossRef] [PubMed]
- Zupkovitz, G.; Grausenburger, R.; Brunmeir, R.; Senese, S.; Tischler, J.; Jurkin, J.; Rembold, M.; Meunier, D.; Egger, G.; Lagger, S.; et al. The Cyclin-Dependent Kinase Inhibitor p21 Is a Crucial Target for Histone Deacetylase 1 as a Regulator of Cellular Proliferation. Mol. Cell. Biol. 2010, 30, 1171–1181. [Google Scholar] [CrossRef] [PubMed]
- Kudová, J.; Procházková, J.; Vašiček, O.; Perečko, T.; Sedláčková, M.; Pešl, M.; Pacherník, J.; Kubala, L. HIF-1alpha deficiency attenuates the cardiomyogenesis of mouse embryonic stem cells. PLoS ONE 2016, 11, e0158358. [Google Scholar] [CrossRef] [PubMed]
- Večeřa, J.; Bártová, E.; Krejčí, J.; Legartová, S.; Komůrková, D.; Rudá-Kučerová, J.; Štark, T.; Dražanová, E.; Kašpárek, T.; Šulcová, A.; et al. HDAC1 and HDAC3 underlie dynamic H3K9 acetylation during embryonic neurogenesis and in schizophrenia-like animals. J. Cell. Physiol. 2018, 233, 530–548. [Google Scholar] [CrossRef] [PubMed]
- Krejčí, J.; Harničarová, A.; Kůrová, J.; Uhlířová, R.; Kozubek, S.; Legartová, S.; Hájek, R.; Bártová, E. Nuclear organization of PML bodies in leukaemic and multiple myeloma cells. Leuk. Res. 2008, 32, 1866–1877. [Google Scholar] [CrossRef] [PubMed]
- Krejčí, J.; Legartová, S.; Bártová, E. Neural Differentiation in HDAC1-Depleted Cells Is Accompanied by Coilin Downregulation and the Accumulation of Cajal Bodies in Nucleoli. Stem Cells Int. 2017, 2017, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Legartová, S.; Suchánková, J.; Krejčí, J.; Kovaříková, A.; Bártová, E. Advanced Confocal Microscopy Techniques to Study Protein-protein Interactions and Kinetics at DNA Lesions. J. Vis. Exp. 2017. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arcidiacono, O.A.; Krejčí, J.; Suchánková, J.; Bártová, E. Deacetylation of Histone H4 Accompanying Cardiomyogenesis is Weakened in HDAC1-Depleted ES Cells. Int. J. Mol. Sci. 2018, 19, 2425. https://doi.org/10.3390/ijms19082425
Arcidiacono OA, Krejčí J, Suchánková J, Bártová E. Deacetylation of Histone H4 Accompanying Cardiomyogenesis is Weakened in HDAC1-Depleted ES Cells. International Journal of Molecular Sciences. 2018; 19(8):2425. https://doi.org/10.3390/ijms19082425
Chicago/Turabian StyleArcidiacono, Orazio Angelo, Jana Krejčí, Jana Suchánková, and Eva Bártová. 2018. "Deacetylation of Histone H4 Accompanying Cardiomyogenesis is Weakened in HDAC1-Depleted ES Cells" International Journal of Molecular Sciences 19, no. 8: 2425. https://doi.org/10.3390/ijms19082425
APA StyleArcidiacono, O. A., Krejčí, J., Suchánková, J., & Bártová, E. (2018). Deacetylation of Histone H4 Accompanying Cardiomyogenesis is Weakened in HDAC1-Depleted ES Cells. International Journal of Molecular Sciences, 19(8), 2425. https://doi.org/10.3390/ijms19082425






