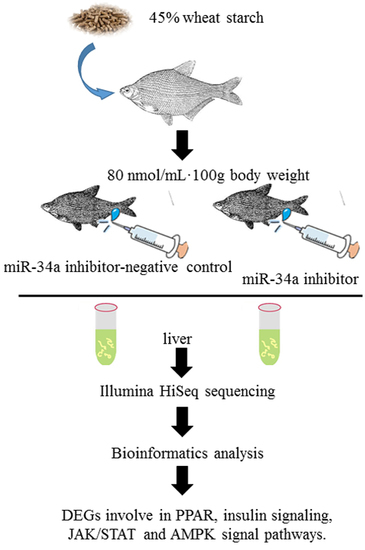
In Vivo Analysis of miR-34a Regulated Glucose Metabolism Related Genes in Megalobrama amblycephala
Abstract
1. Introduction
2. Results
2.1. Transcriptome Sequencing in M. amblycephala
2.2. De Novo Assembly of the M. amblycephala Transcriptome
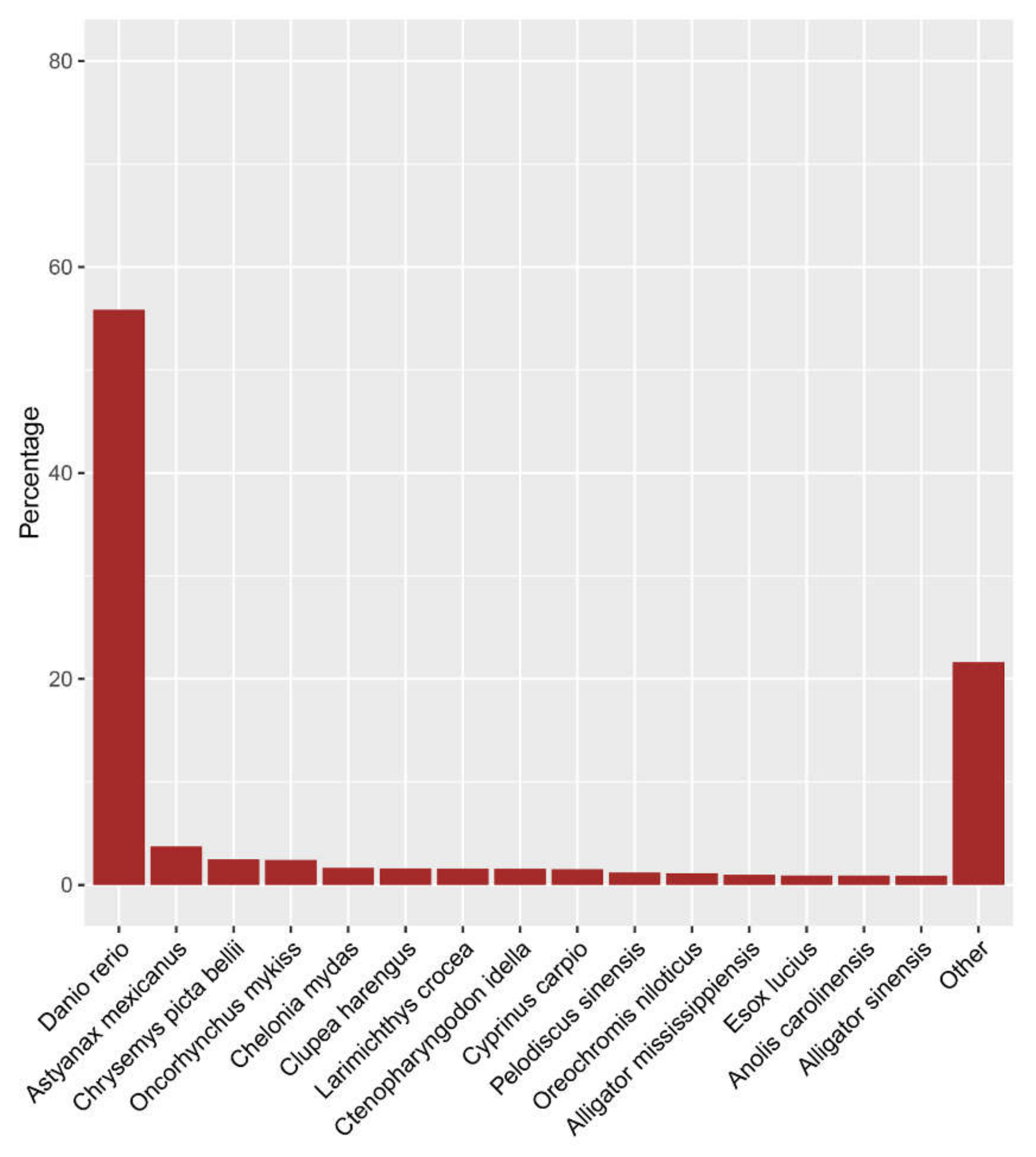
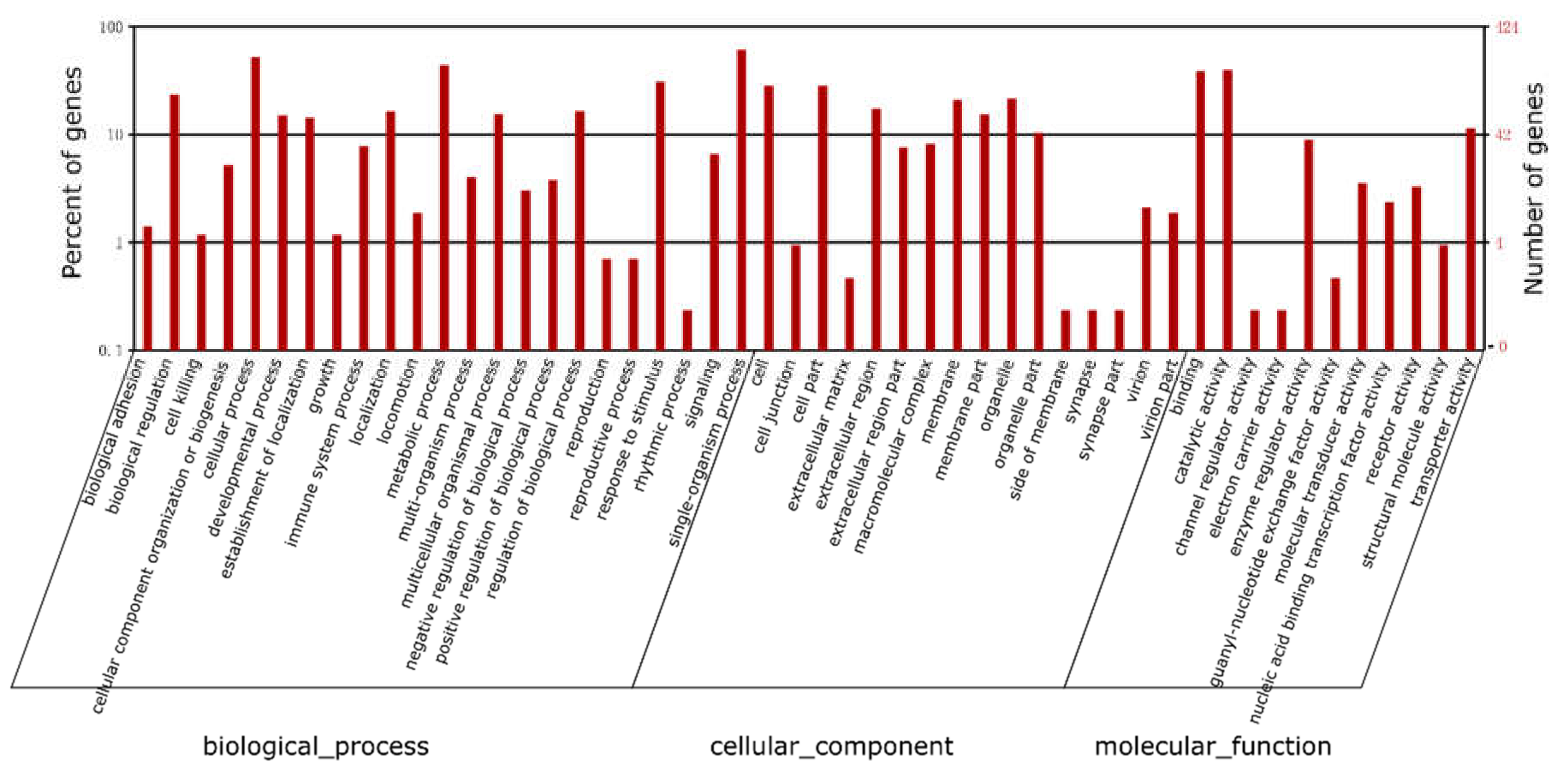
2.3. Functional Annotation
2.4. Identification of Simple Sequence Repeat
2.5. Open Reading Frame Identification and Prediction
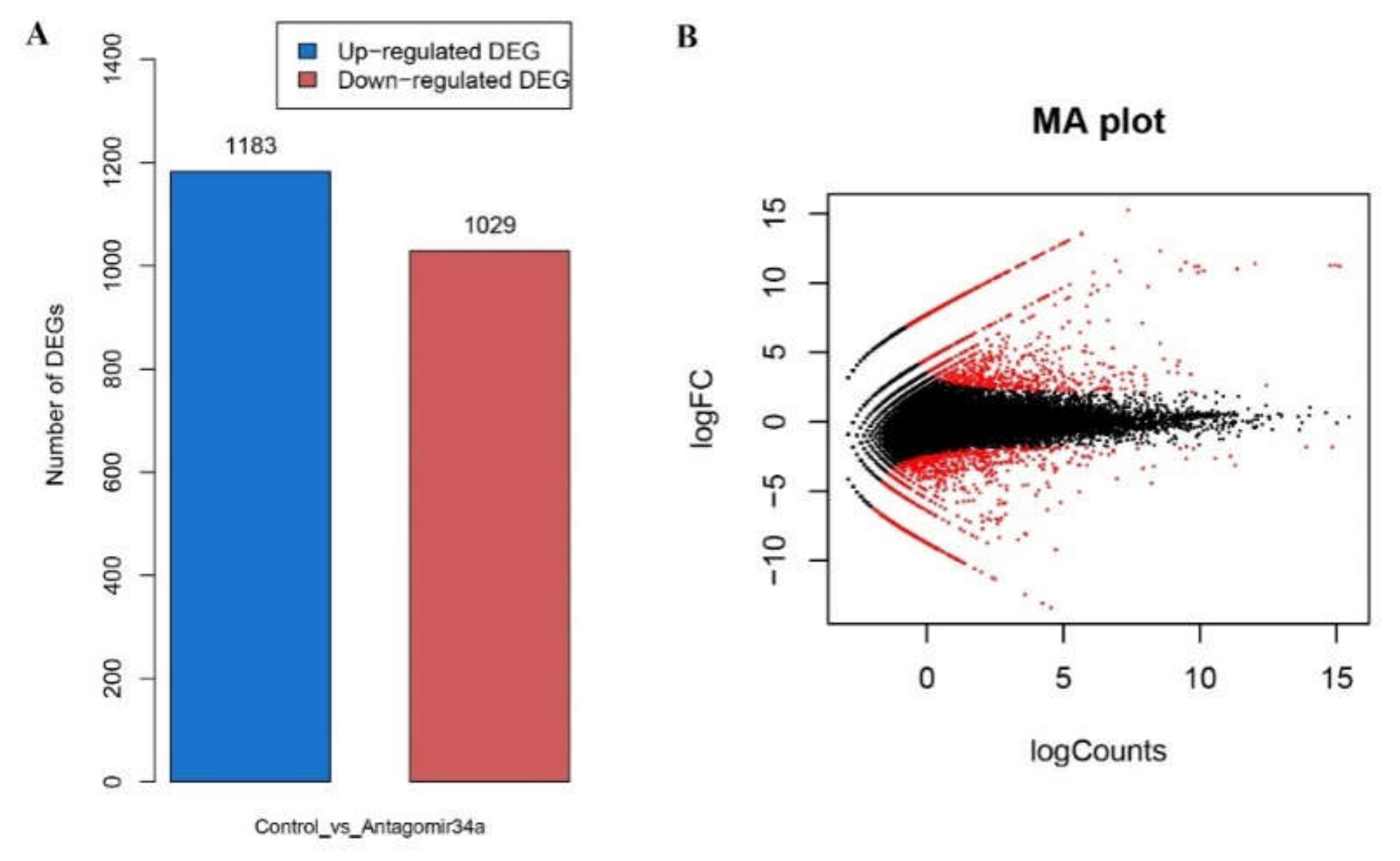
2.6. DEG Predication and Clustering Analysis
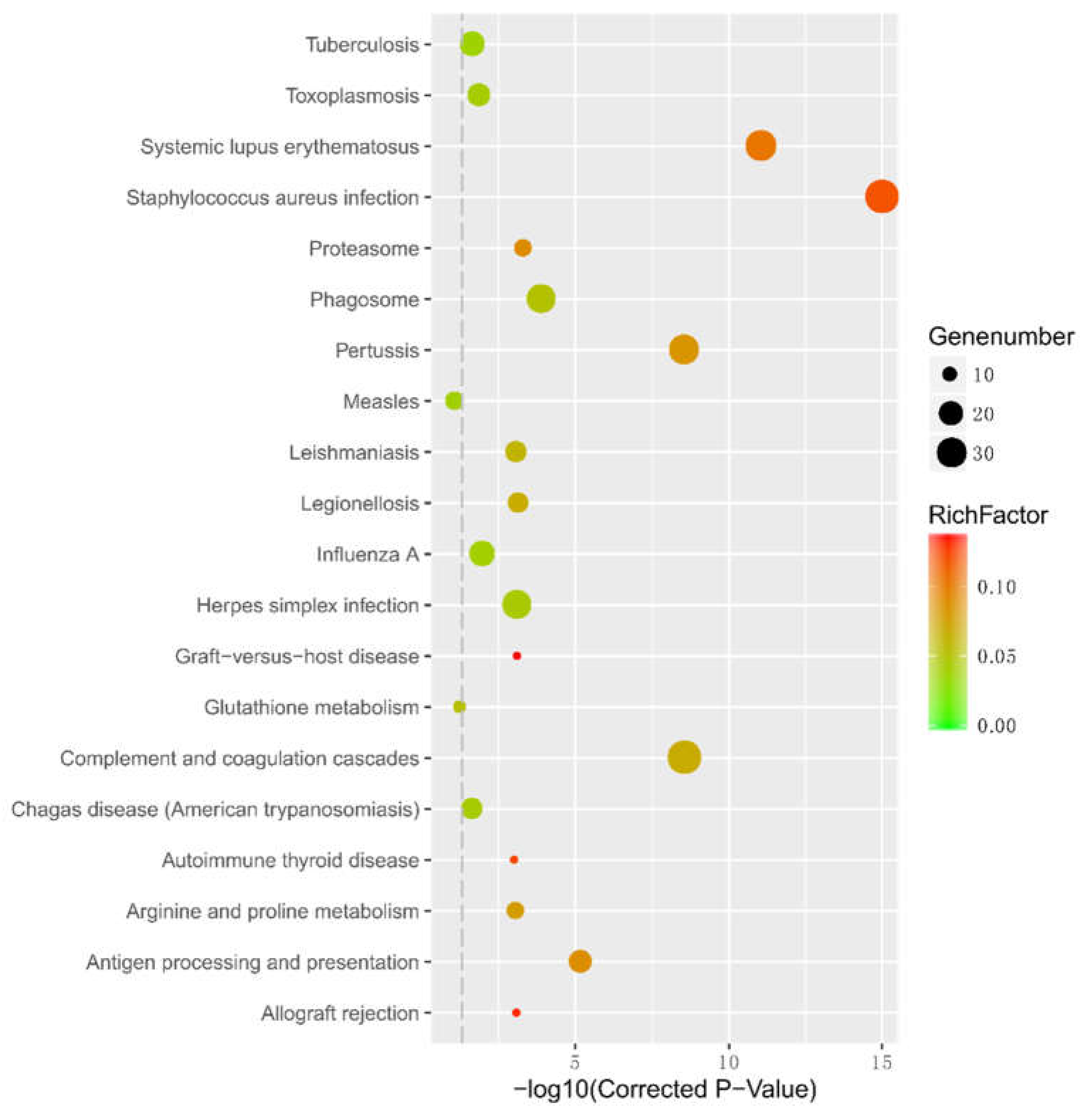
2.7. Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) Pathways Enrichment Analysis of differentially expressed genes (DEGs)
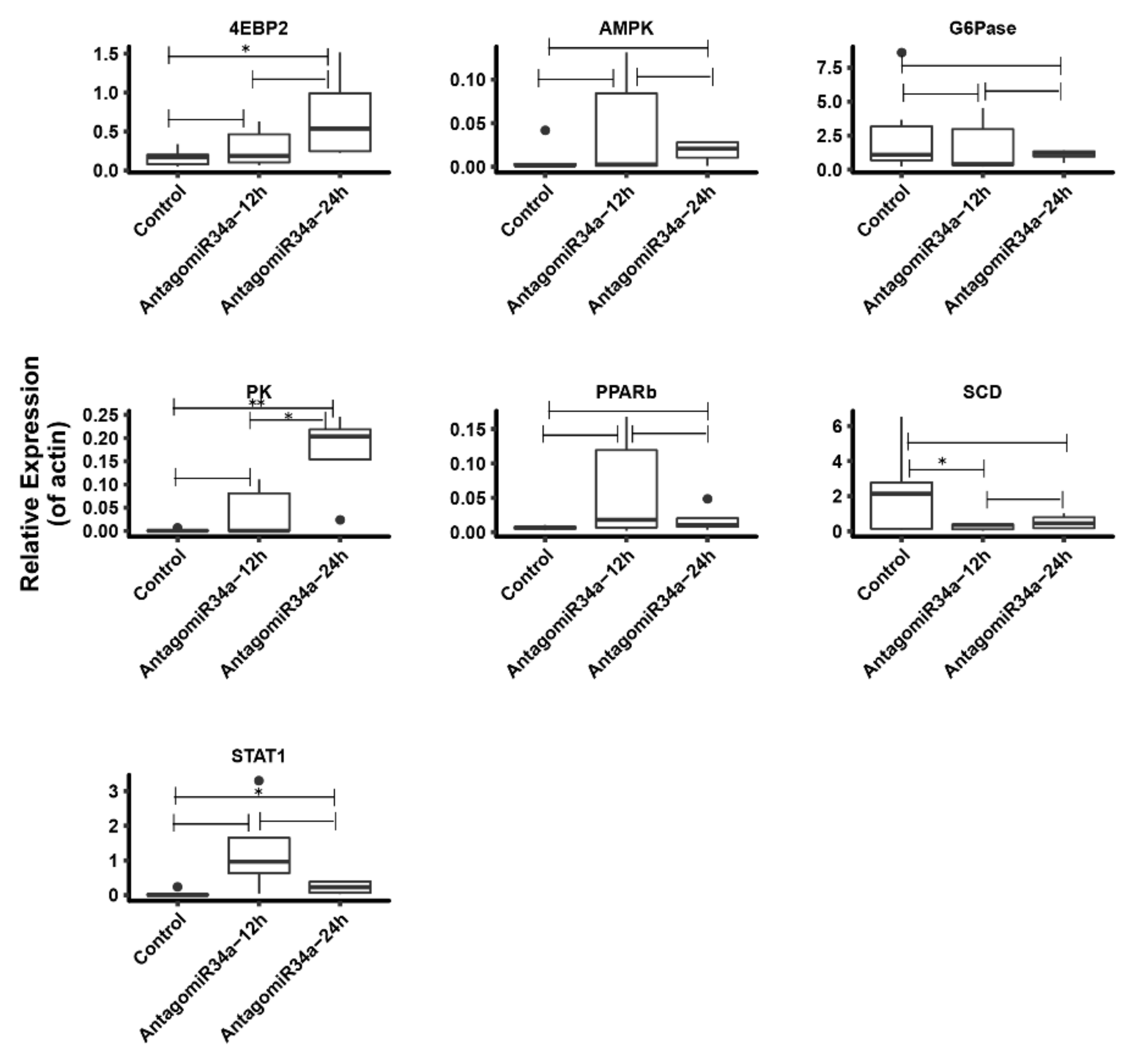
2.8. Change of Some Other Glucose Metabolism Related Genes after Inhibition of miR-34a
3. Discussion and Conclusions
4. Material and Methods
4.1. Ethics Statement
4.2. Sample Collection
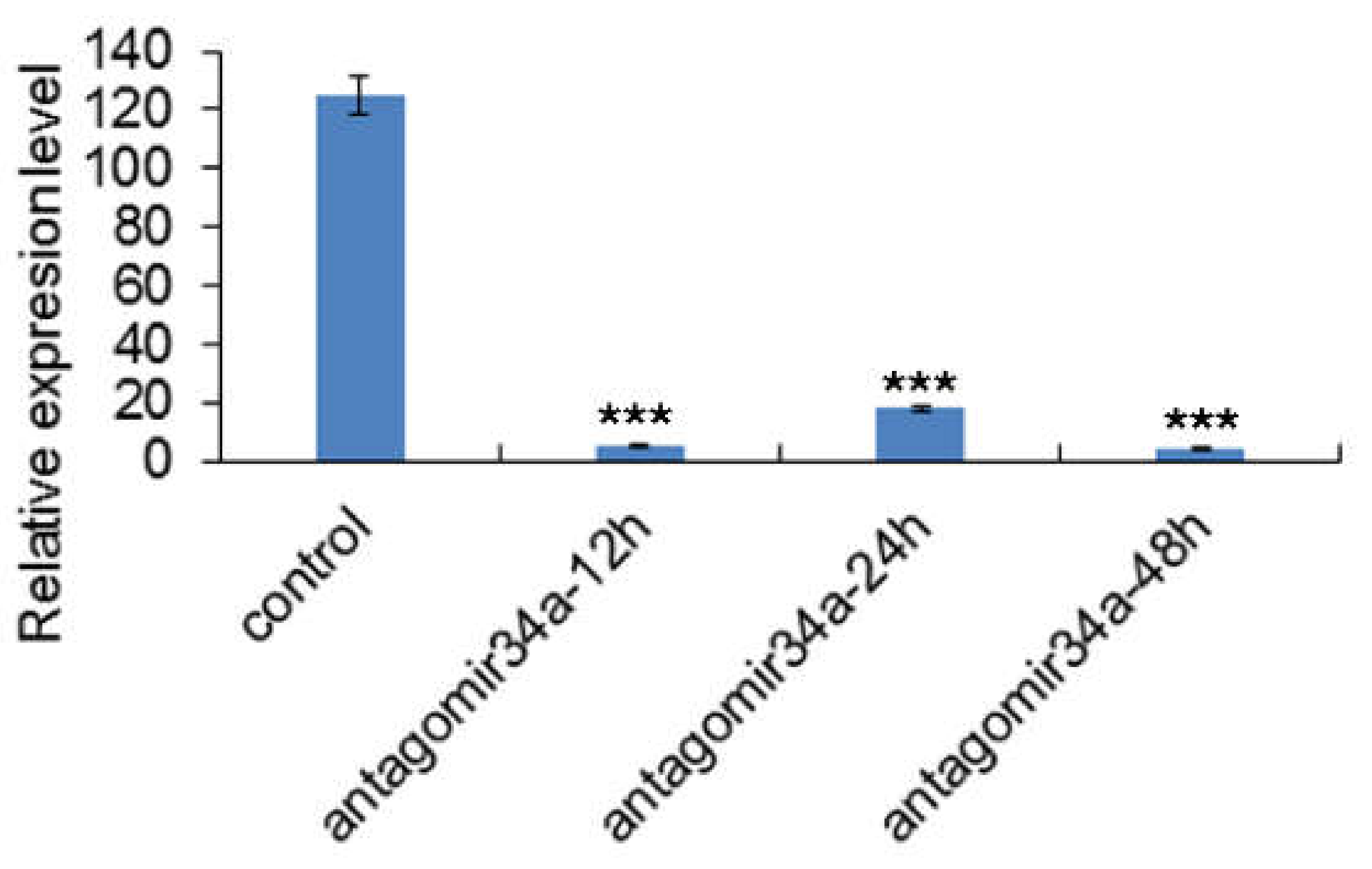
4.3. qRT-PCR Analysis for the miR-34a
4.4. cDNA Library Construction and Sequencing
4.5. De Novo Assembly of Reference Sequences
4.6. Assembled Sequence Annotation and Classification
4.7. SSRs Identification
4.8. ORF Identification and Prediction
4.9. Identification of DEGs
4.10. QRT-PCR Analysis for DEGs
4.11. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| FAO | Food and Agriculture Organization |
| DEM | Differentially expressed miRNA |
| T2D | Type 2 diabetes |
| NAFLD | Non-alcoholic fatty liver disease |
| ACSL1 | Acyl-CoA synthetase long-chain family member 1 |
| FOXO1 | Forkhead box protein O1 |
| PPARs | Peroxisome proliferator-activated receptors |
| PGC-1α | Peroxisome proliferator-activated receptor γ coactivator 1-α |
| COG | Cluster of Orthologous Groups of protein |
| MISA | MIcroSAtellite |
| SSR | Simple sequence repeat |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| ORF | Open reading frame |
| FPKM | Fragments per kilobase of exon per million reads mapped |
| FDR | False Discovery Rate |
| HSD | High starch diet |
References
- Englyst, H.N.; Hudson, G.J. The classification and measurement of dietary carbohydrates. Food Chem. 1996, 57, 15–21. [Google Scholar] [CrossRef]
- Asp, N.G. Dietary carbohydrates: Classification by chemistry and physiology. Food Chem. 1996, 57, 9–14. [Google Scholar] [CrossRef]
- Hemre, G.I.; Mommsen, T.P.; Krogdahl, A. Carbohydrates in fish nutrition: Effects on growth, glucose metabolism and hepatic enzymes. Aquac. Nutr. 2015, 8, 175–194. [Google Scholar] [CrossRef]
- Li, X.F.; Wang, Y.; Liu, W.B.; Jiang, G.Z.; Zhu, J. Effects of dietary carbohydrate/lipid ratios on growth performance, body composition and glucose metabolism of fingerling blunt snout bream Megalobrama amblycephala. Aquac. Nutr. 2013, 19, 701–708. [Google Scholar] [CrossRef]
- Hung, L.T.; Lazard, J.; Mariojouls, C.; Moreau, Y. Comparison of starch utilization in fingerlings of two Asian catfishes from the Mekong River (Pangasius bocourti Sauvage, 1880, Pangasius Hypophthalmus Sauvage, 1878). Aquac. Nutr. 2015, 9, 215–222. [Google Scholar] [CrossRef]
- Watanabe, T. Strategies for further development of aquatic feeds. Fish. Sci. 2002, 68, 242–252. [Google Scholar] [CrossRef]
- Yi, S.; Gao, Z.X.; Zhao, H.; Zeng, C.; Luo, W.; Chen, B.; Wang, W.M. Identification and characterization of microRNAs involved in growth of blunt snout bream (Megalobrama amblycephala) by Solexa sequencing. BMC Genom. 2013, 14, 754. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Xu, L.; Liu, W.; Liu, Y.; Ringø, E.; Du, Z.; Zhou, Z. Protein replacement in practical diets altered gut allochthonous bacteria of cultured cyprinid species with different food habits. Aquac. Int. 2015, 23, 913–928. [Google Scholar] [CrossRef]
- Miao, L.H.; Lin, Y.; Pan, W.J.; Huang, X.; Ge, X.P.; Ren, M.C.; Zhou, Q.L.; Liu, B. Identification of differentially expressed microRNAs associate with glucose metabolism in different organs of blunt snout bream (Megalobrama amblycephala). Int. J. Mol. Sci. 2017, 18, 1161. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Liu, B.; Xie, J.; Ge, X. Effect of High Dietary Carbohydrate on Growth, Serum Physiological Response, and Hepatic Heat Shock Protein 70 Expression of Wuchang Bream (Megalobrama amblycephala) at Two Temperatures. Turk. J. Fish. Aquat. Sci. 2013, 13, 211–220. [Google Scholar] [CrossRef]
- Zhou, C.; Liu, B.; Ge, X.; Xie, J.; Xu, P. Effect of dietary carbohydrate on the growth performance, immune response, hepatic antioxidant abilities and heat shock protein 70 expression of Wuchang bream, Megalobrama amblycephala. J. Appl. Ichthyol. 2013, 29, 1348–1356. [Google Scholar] [CrossRef]
- Zhou, C.; Ge, X.P.; Liu, B.; Xie, J.; Miao, L.H. Effect of Dietary Carbohydrate Level on Growth Performance, Blood Chemistry, Hepatic Enzyme Activity, and Growth Hormone Gene Expression in Wuchang Bream (Megalobrama amblycephala). Isr. J. Aquac.-Bamidgeh 2013, 65, 1–8. [Google Scholar]
- Ghawanmeh, T.; Thunberg, U.; Castro, J.; Murray, F.; Laytragoon-Lewin, N. miR-34a expression, cell cycle arrest and cell death of malignant mesothelioma cells upon treatment with radiation, docetaxel or combination treatment. Oncology 2011, 81, 330–335. [Google Scholar] [CrossRef] [PubMed]
- Li, X.J.; Ren, Z.J.; Tang, J.H. MicroRNA-34a: A potential therapeutic target in human cancer. Cell Death Dis. 2014, 5, e1327. [Google Scholar] [CrossRef] [PubMed]
- Lovis, P.; Roggli, E.; Laybutt, D.R.; Gattesco, S.; Yang, J.Y.; Widmann, C.; Abderrahmani, A.; Regazzi, R. Alterations in microRNA expression contribute to fatty acid-induced pancreatic β-cell dysfunction. Diabetes 2008, 57, 2728–2736. [Google Scholar] [CrossRef] [PubMed]
- Roggli, E.; Britan, A.; Gattesco, S.; Lin-Marq, N.; Abderrahmani, A.; Meda, P.; Regazzi, R. Involvement of microRNAs in the cytotoxic effects exerted by proinflammatory cytokines on pancreatic β-cells. Diabetes 2010, 59, 978–986. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Chen, X.; Zhang, H.; Liang, X.; Xiang, Y.; Yu, C.; Zen, K.; Li, Y.; Zhang, C.Y. Differential expression of microRNAs in mouse liver under aberrant energy metabolic status. J. Lipid Res. 2009, 50, 1756–1765. [Google Scholar] [CrossRef] [PubMed]
- Trajkovski, M.; Hausser, J.; Soutschek, J.; Bhat, B.; Akin, A.; Zavolan, M.; Heim, M.H.; Stoffel, M. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 2011, 474, 649–653. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.; Bing, L.; Lodish, H.F. MicroRNAs induced during adipogenesis that accelerate fat cell development are downregulated in obesity. Diabetes 2009, 58, 1050–1057. [Google Scholar] [CrossRef] [PubMed]
- Cermelli, S.; Ruggieri, A.; Marrero, J.A.; Ioannou, G.N.; Beretta, L. Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS ONE 2011, 6, e23937. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhu, J.; Han, W.; Jiang, X.; Xu, M.; Zhao, Y.; Dong, Q.; Pang, Z.; Guan, Q.; Gao, L.; et al. Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2diabetes: A clinical study. Acta Diabetol. 2011, 48, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Yamakuchi, M.; Ferlito, M.; Lowenstein, C.J. miR-34a repression of sirt1 regulates apoptosis. Proc. Natl. Acad. Sci. USA 2008, 105, 13421–13426. [Google Scholar] [CrossRef] [PubMed]
- Li, W.Q.; Chen, C.; Xu, M.D.; Guo, J.; Li, Y.M.; Xia, Q.M.; Liu, H.M.; He, J.; Yu, H.Y.; Zhu, L. The rno-miR-34 family is upregulated and targets ACSL1 in dimethylnitrosamine-induced hepatic fibrosis in rats. FEBS J. 2011, 278, 1522–1532. [Google Scholar] [CrossRef] [PubMed]
- Rottiers, V.; Näär, A.M. MicroRNAs in metabolism and metabolic disorders. Nat. Rev. Mol. Cell Biol. 2012, 13, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Meng, Y.; Tian, S.; Chen, J.; Liu, M.; Zhuo, M.; Zhang, Y.; Du, H.; Wang, X. Comparative microRNA expression profiles of cynomolgus monkeys, rat, and human reveal that miR-182 is involved in T2D pathogenic processes. J. Diabetes Res. 2014, 2014, 760397. [Google Scholar] [CrossRef] [PubMed]
- Tryndyak, V.P.; Ross, S.A.; Beland, F.A.; Pogribny, I.P. Down-regulation of the microRNAs miR-34a, miR-127, and miR-200b in rat liver during hepatocarcinogenesis induced by a methyl-deficient diet. Mol. Carcinog. 2009, 48, 479–487. [Google Scholar] [CrossRef] [PubMed]
- Castro, R.E.; Ferreira, D.M.; Afonso, M.B.; Borralho, P.M.; Machado, M.V.; Cortez-Pinto, H.; Rodrigues, C.M. miR-34a/SIRT1/p53 is suppressed by ursodeoxycholic acid in the rat liver and activated by disease severity in human non-alcoholic fatty liver disease. J. Hepatol. 2013, 58, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Lam, M. Registered report: The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. J. Urol. 2011, 186, 211–215. [Google Scholar]
- Gurzov, E.N.; Stanley, W.J.; Pappas, E.G.; Thomas, H.E.; Gough, D.J. The JAK-STAT pathway in obesity and diabetes. FEBS J. 2016, 283, 3002–3015. [Google Scholar] [CrossRef] [PubMed]
- Farooqi, I.S.; O’Rahilly, S. 20 years of leptin:human disorders of leptin action. J. Endocrinol. 2014, 223, T63–T70. [Google Scholar] [CrossRef] [PubMed]
- Hamilton-Williams, E.E.; Palmer, S.E.; Charlton, B.; Slattery, R.M. Beta cell MHC class I is a late requirement for diabetes. Proc. Natl. Acad. Sci. USA 2003, 100, 6688–6693. [Google Scholar] [CrossRef] [PubMed]
- Habte-Tsion, H.M.; Ren, M.; Liu, B.; Xie, J.; Ge, X.; Chen, R.; Zhou, Q.; Pan, L. Threonine affects digestion capacity and hepatopancreatic gene expression of juvenile blunt snout bream (Megalobrama amblycephala). Br. J. Nutr. 2015, 114, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Kemper, J.K. Controlling SIRT1 expression by microRNAs in health and metabolic disease. Aging 2010, 2, 527–534. [Google Scholar] [CrossRef] [PubMed]
- Kemper, J.K.; Xiao, Z.; Ponugoti, B.; Miao, J.; Fang, S.; Kanamaluru, D.; Tsang, S.; Wu, S.Y.; Chiang, C.M.; Veenstra, T.D. FXR acetylation is normally dynamically regulated by p300 and SIRT1 but constitutively elevated in metabolic disease states. Cell Metab. 2009, 10, 392–404. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, S.; Blander, G.; Tse, J.G.; Krieger, M.; Guarente, L. SIRT1 deacetylates and positively regulates the nuclear receptor LXR. Mol. Cell 2007, 28, 91–106. [Google Scholar] [CrossRef] [PubMed]
- Motta, M.C.; Divecha, N.; Lemieux, M.; Kamel, C.; Chen, D.; Gu, W.; Bultsma, Y.; McBurney, M.; Guarente, L. Mammalian SIRT1 represses forkhead transcription factors. Cell 2004, 116, 551–563. [Google Scholar] [CrossRef]
- Ponugoti, B.; Kim, D.H.; Xiao, Z.; Smith, Z.; Miao, J.; Zang, M.; Wu, S.Y.; Chiang, C.M.; Veenstra, T.D.; Kemper, J.K. SIRT1 deacetylates and inhibits SREBP-1C activity in regulation of hepatic lipid metabolism. J. Biol. Chem. 2010, 285, 33959–33970. [Google Scholar] [CrossRef] [PubMed]
- Purushotham, A.; Schug, T.T.; Xu, Q.; Surapureddi, S.; Guo, X.; Li, X. Hepatocyte-specific deletion of SIRT1 alters fatty acid metabolism and results in hepatic steatosis and inflammation. Cell Metab. 2009, 9, 327–338. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, H.; Dessain, S.K.; Ng Eaton, E.; Imai, S.I.; Frye, R.A.; Pandita, T.K.; Guarente, L.; Weinberg, R.A. hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 2001, 107, 149–159. [Google Scholar] [CrossRef]
- Walker, A.K.; Yang, F.; Jiang, K.; Ji, J.Y.; Watts, J.L.; Purushotham, A.; Boss, O.; Hirsch, M.L.; Ribich, S.; Smith, J.J.; et al. Conserved role of SIRT1 orthologs in fasting-dependent inhibition of the lipid/cholesterol regulator SREBP. Genes Dev. 2010, 24, 1403–1417. [Google Scholar] [CrossRef] [PubMed]
- Fan, Y.; Hoberg, J.E.; Ramsey, C.S.; Keller, M.D.; Jones, D.R.; Frye, R.A.; Mayo, M.W. Modulation of NF-κB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J. 2004, 23, 2369–2380. [Google Scholar]
- And, J.B.; Moller, D.E. The mechanisms of action of PPARs. Annu. Rev. Med. 2002, 53, 409–435. [Google Scholar]
- Feige, J.N.; Gelman, L.; Michalik, L.; Desvergne, B.; Wahli, W. From molecular action to physiological outputs: Peroxisome proliferator-activated receptors are nuclear receptors at the crossroads of key cellular functions. Prog. Lipid Res. 2002, 45, 120–159. [Google Scholar] [CrossRef] [PubMed]
- Sanderson, L.M.; Boekschoten, M.V.; Desvergne, B.; Müller, M.; Kersten, S. Transcriptional profiling reveals divergent roles of PPARα and PPARβ/δ in regulation of gene expression in mouse liver. Physiol. Genom. 2010, 41, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Miao, L.H.; Pan, W.J.; Ge, X.P.; Liu, B.; Ren, M.C.; Zhou, Q.L. Bioinformatic prediction and analysis of glucolipid metabolic regulation by miR-34a in Megalobrama amblycephala. Genes Genom. 2017, 39, 1407–1417. [Google Scholar] [CrossRef]
- Chen, K.; Zhao, L.; He, H.; Wan, X.; Wang, F.; Mo, Z. Silibinin protects β cells from glucotoxicity through regulation of the Insig-1/SREBP-1c pathway. Int. J. Mol. Med. 2014, 34, 1073–1080. [Google Scholar] [CrossRef] [PubMed]
- Zhan, T.; Poppelreuther, M.; Ehehalt, R.; Füllekrug, J. Overexpressed FATP1, ACSVL4/FATP4 and ACSL1 increase the cellular fatty acid uptake of 3T3-L1 adipocytes but are localized on intracellular membranes. PLoS ONE 2012, 7, e45087. [Google Scholar] [CrossRef] [PubMed]
- Phosat, C.; Panprathip, P.; Chumpathat, N.; Prangthip, P.; Chantratita, N.; Soonthornworasiri, N.; Kwanbunjan, K. Elevated, C-reactive protein, interleukin 6, tumor necrosis factor α and glycemic load associated with type 2 diabetes mellitus in rural Thais: A cross-sectional study. BMC Endocr. Disord. 2017, 17, 44. [Google Scholar] [CrossRef] [PubMed]
- Long, Y.C.; Zierath, J.R. AMP-activated protein kinase signaling in metabolic regulation. J. Clin. Investig. 2006, 116, 1776–1783. [Google Scholar] [CrossRef] [PubMed]
- Saltiel, A.R.; Kahn, C.R. Insulin signaling and the regulation of glucose and lipid metabolism. Nature 2002, 414, 799–806. [Google Scholar] [CrossRef] [PubMed]
- Lochhead, P.A.; Salt, I.P.; Walker, K.S.; Hardie, D.G.; Sutherland, C. 5-aminoimidazole-4-carboxamide riboside mimics the effects of insulin on the expression of the 2 key gluconeogenic genes PEPCK and glucose-6-phosphatase. Diabetes 2000, 49, 896–903. [Google Scholar] [CrossRef] [PubMed]
- Andreelli, F.; Foretz, M.; Knauf, C.; Cani, P.D.; Perrin, C.; Iglesias, M.A.; Pillot, B.; Bado, A.; Tronche, F.; Mithieux, G.; et al. Liver adenosine monophosphate-activated kinase-α2 catalytic subunit is a key target for the control of hepatic glucose production by adiponectin and leptin but not insulin. Endocrinology 2006, 147, 2432–2441. [Google Scholar] [CrossRef] [PubMed]
- Gan, Z.; Burkart-Hartman, E.M.; Han, D.H.; Finck, B.; Leone, T.C.; Smith, E.Y.; Ayala, J.E.; Holloszy, J.; Kelly, D.P. The nuclear receptor PPARβ/δ programs muscle glucose metabolism in cooperation with AMPK and MEF2. Genes Dev. 2011, 25, 2619–2630. [Google Scholar] [CrossRef] [PubMed]
- Ren, M.; Liao, Y.; Xie, J.; Liu, B.; Zhou, Q.; Ge, X.; Cui, H.; Pan, L.; Chen, R. Dietary arginine requirement of juvenile blunt snout bream, Megalobrama amblycephala. Aquaculture 2013, 414, 229–234. [Google Scholar] [CrossRef]
- Grabherr, M.G.; Haas, B.J.; Yassour, M.; Levin, J.Z.; Thompson, D.A.; Amit, I.; Adiconis, X.; Fan, L.; Raychowdhury, R.; Zeng, Q.; et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Li, J.; Wu, S.; Zhu, Y.; Chen, Y.; He, F. Integrated NR database in protein annotation system and its localization. Comput. Eng. 2006, 32, 71–74. [Google Scholar]
- Apweiler, R.; Bairoch, A.; Wu, C.H.; Barker, W.C.; Boeckmann, B.; Ferro, S.; Gasteiger, E.; Huang, H.; Lopez, R.; Magrane, M.; et al. UniProt: The Universal Protein knowledgebase. Nucleic Acids Res. 2004, 32, D115–D119. [Google Scholar] [CrossRef] [PubMed]
- Tatusov, R.L.; Galperin, M.Y.; Natale, D.A.; Koonin, E.V. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28, 33–36. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, 222–230. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Dcwcy, C.N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; Mccarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
- Boyle, E.I.; Weng, S.; Gollub, J.; Jin, H.; Botstein, D.; Cherry, J.M.; Sherlock, G. GO::TermFinder—Open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics 2004, 20, 3710–3715. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, W316–W322. [Google Scholar] [CrossRef] [PubMed]



| Sample | Raw Reads | Clean Reads | Clean Q20 (%) | Clean Q30 (%) | Clean GC (%) |
|---|---|---|---|---|---|
| AntagomiR34a-12h | 40,929,414 | 40,359,036 | 97.33 | 92.81 | 46.96 |
| Control-1 | 51,674,516 | 50,991,682 | 97.39 | 92.97 | 46.35 |
| Control-2 | 52,931,508 | 52,218,818 | 97.47 | 93.14 | 47.20 |
| Item | Value |
|---|---|
| Total number of sequences | 222,411 |
| Total number of genes | 197,425 |
| Total length of sequences (bp) | 151,079,306 |
| Maximum sequence length (bp) | 16,839 |
| Minimum sequence length (bp) | 200 |
| N50 | 1041 |
| Annotated Database | Annotated Number | Transcript Ratio (%) |
|---|---|---|
| All_Assembly_Contig | 222,411 | 100 |
| Nr | 82,223 | 36.97 |
| SwissProt | 57,581 | 25.89 |
| KEGG | 71,045 | 31.94 |
| KOG | 45,994 | 20.68 |
| Pfam | 27,677 | 12.44 |
| GO | 25,433 | 11.44 |
| All_anno_contig | 83,587 | 37.58 |
| Repeat Numbers | SSR Type | Total | |||||
|---|---|---|---|---|---|---|---|
| Mono- | Di- | Tri- | Tetra- | Penta- | Hexa- | ||
| 5 | 0 | 0 | 3436 | 996 | 154 | 21 | 4607 |
| 6 | 0 | 4847 | 1545 | 710 | 38 | 11 | 7151 |
| 7 | 0 | 2673 | 948 | 75 | 18 | 1 | 3715 |
| 8 | 0 | 1674 | 873 | 93 | 24 | 1 | 2665 |
| 9 | 0 | 1266 | 134 | 60 | 35 | 0 | 1495 |
| 10 | 11,710 | 1138 | 198 | 60 | 22 | 1 | 13,129 |
| 11 | 7843 | 2036 | 129 | 38 | 24 | 0 | 10,070 |
| 12 | 5217 | 1165 | 84 | 27 | 27 | 0 | 6520 |
| ≥13 | 37,415 | 5640 | 172 | 190 | 64 | 1 | 43,482 |
| Total | 62,185 | 20,439 | 7519 | 2249 | 406 | 36 | 92,834 |
| % | 66.99% | 22.02% | 8.10% | 2.42% | 0.44% | 0.04% | |
| Ingredient | Quantity (g) | Proximate Composition | (% of Dry Matter) |
|---|---|---|---|
| Casein 1 | 20 | Crude protein | 33.32 |
| Gelatin | 5 | Crude lipid | 8.58 |
| Fish meal 2 | 16 | Digestible carbohydrate | 45.25 |
| Wheat starch 3 | 45 | Energy (kJ·g−1) | 18.55 |
| Soybean oil 4 | 7 | Calcium | 1.06 |
| Vitamin additive 5 | 1 | Total phosphorus | 1.32 |
| Mineral additives 5 | 1 | ||
| Carboxymethycellulose | 4 | ||
| Monocalcium phosphate 5 | 1 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Miao, L.-H.; Lin, Y.; Huang, X.; Pan, W.-J.; Zhou, Q.-L.; Liu, B.; Ren, M.-C.; Ge, X.-P.; Pan, L.-K. In Vivo Analysis of miR-34a Regulated Glucose Metabolism Related Genes in Megalobrama amblycephala. Int. J. Mol. Sci. 2018, 19, 2417. https://doi.org/10.3390/ijms19082417
Miao L-H, Lin Y, Huang X, Pan W-J, Zhou Q-L, Liu B, Ren M-C, Ge X-P, Pan L-K. In Vivo Analysis of miR-34a Regulated Glucose Metabolism Related Genes in Megalobrama amblycephala. International Journal of Molecular Sciences. 2018; 19(8):2417. https://doi.org/10.3390/ijms19082417
Chicago/Turabian StyleMiao, Ling-Hong, Yan Lin, Xin Huang, Wen-Jing Pan, Qun-Lan Zhou, Bo Liu, Ming-Chun Ren, Xian-Ping Ge, and Liang-Kun Pan. 2018. "In Vivo Analysis of miR-34a Regulated Glucose Metabolism Related Genes in Megalobrama amblycephala" International Journal of Molecular Sciences 19, no. 8: 2417. https://doi.org/10.3390/ijms19082417
APA StyleMiao, L.-H., Lin, Y., Huang, X., Pan, W.-J., Zhou, Q.-L., Liu, B., Ren, M.-C., Ge, X.-P., & Pan, L.-K. (2018). In Vivo Analysis of miR-34a Regulated Glucose Metabolism Related Genes in Megalobrama amblycephala. International Journal of Molecular Sciences, 19(8), 2417. https://doi.org/10.3390/ijms19082417