Protein Expression in Tonsillar and Base of Tongue Cancer and in Relation to Human Papillomavirus (HPV) and Clinical Outcome
Abstract
:1. Introduction
2. Results
2.1. Comparison between Tumor and Normal Samples
2.2. Predictive Models for HPV-Positive and HPV-Negative Tumors
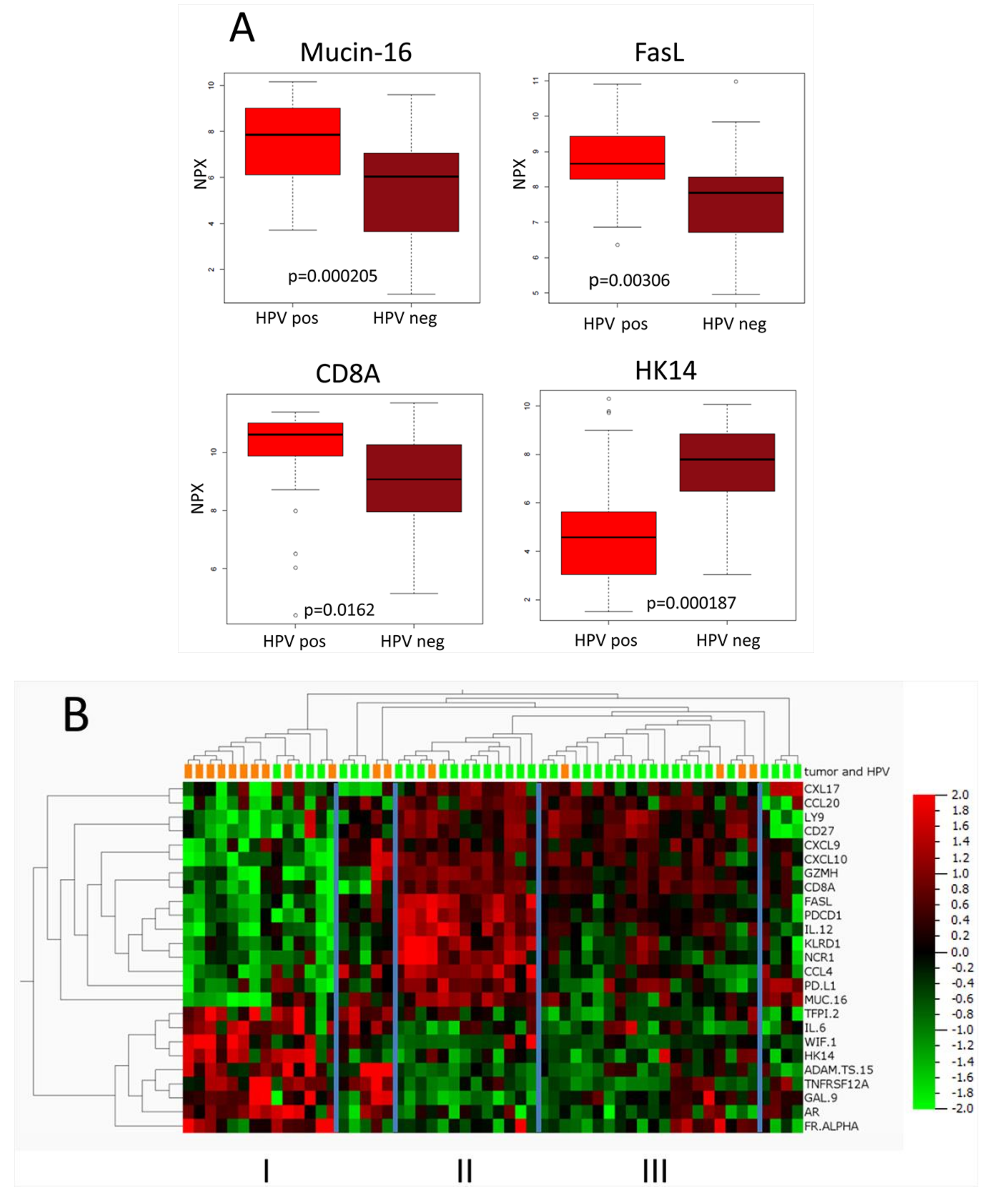
2.3. Comparison between HPV-Positive and HPV-Negative Tumors
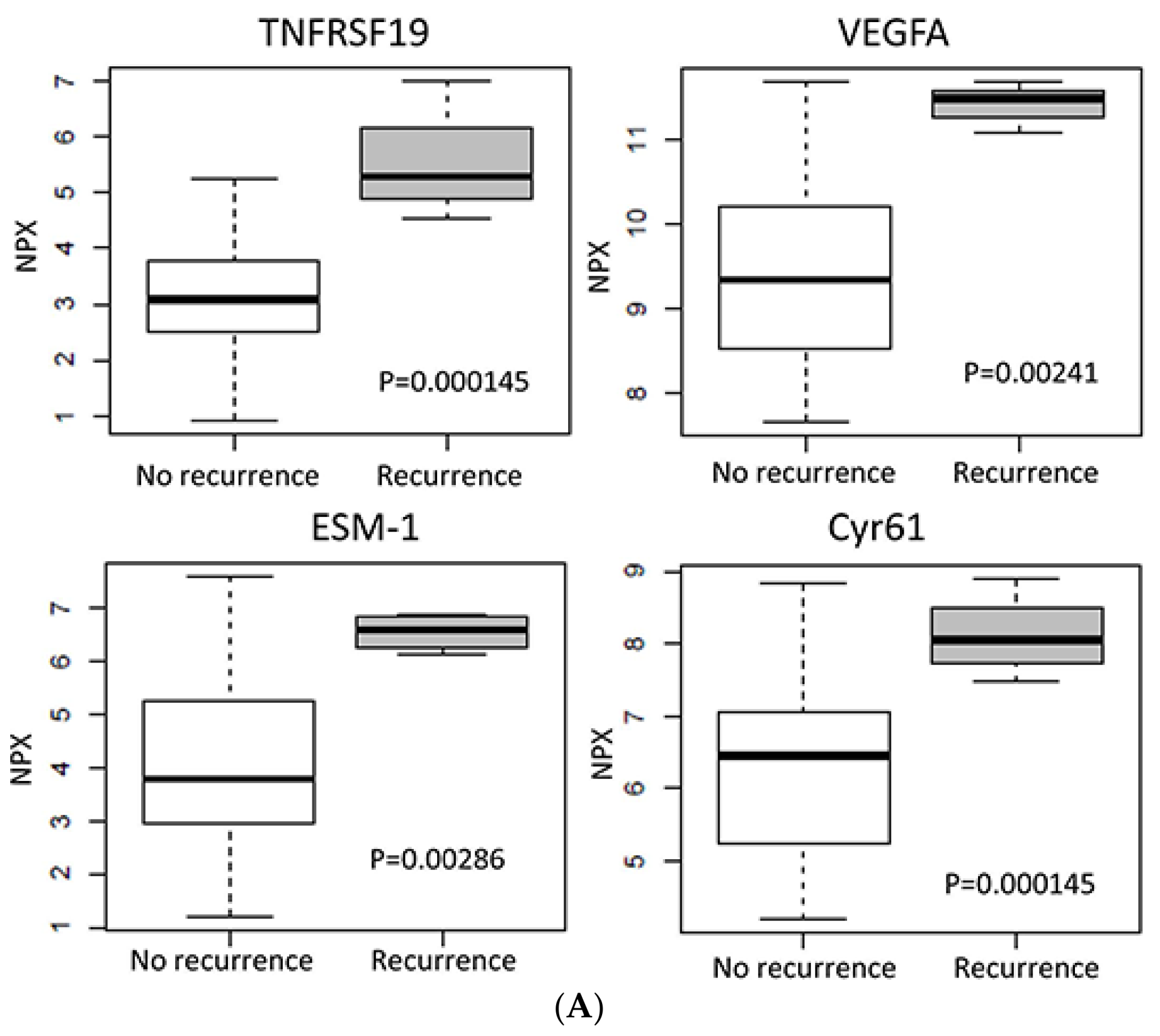
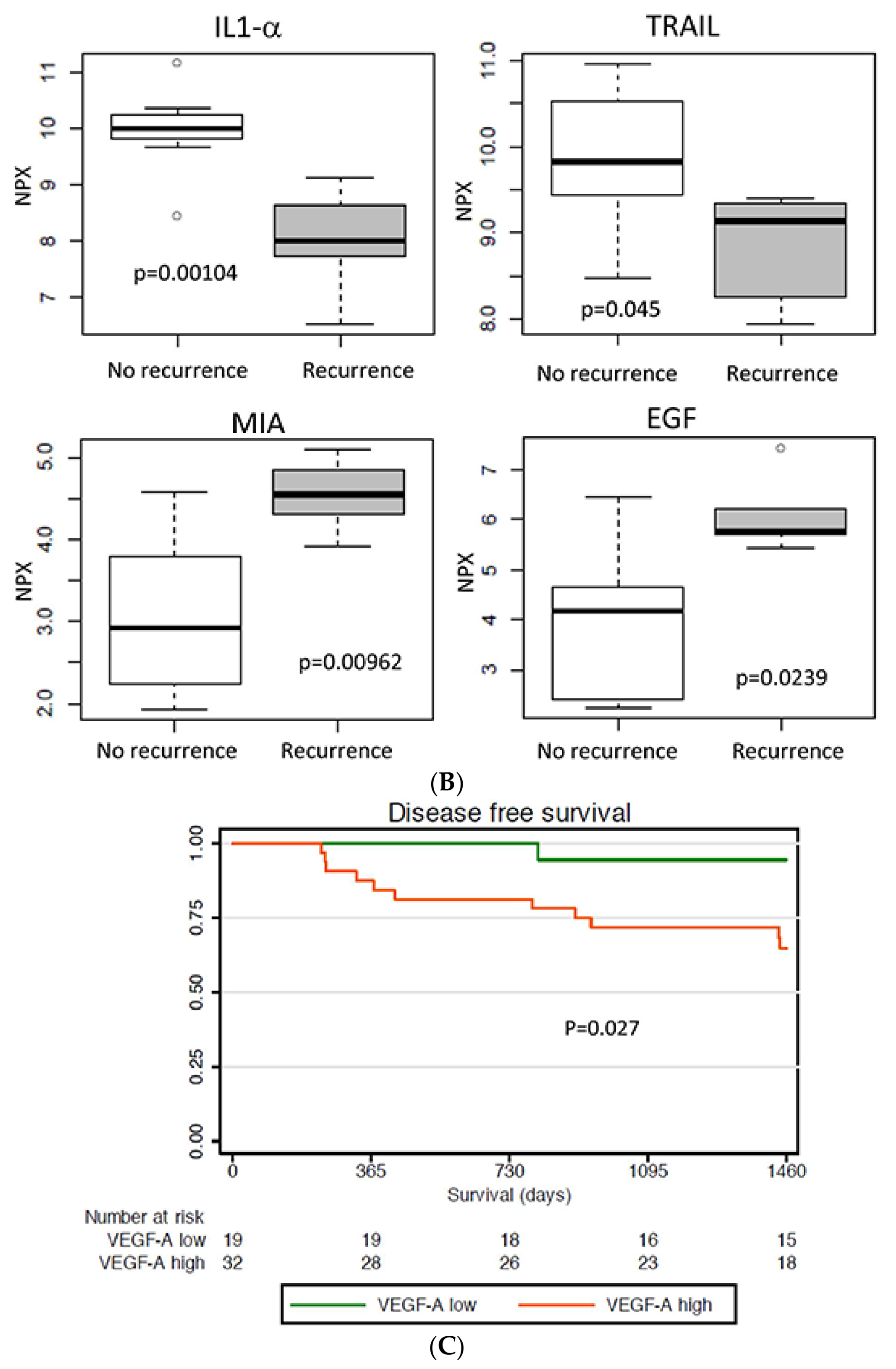
2.4. Protein Expression in Relation to Clinical Outcome
2.5. Proteins Related to High CD8A Expression
2.6. Protein Expression in Relation to Tumor T-Stage
2.7. Validation of VEGFA Expression in Relation to Prognosis
3. Discussion
4. Material and Methods
4.1. Patients and Tumor Biopsies
4.2. Sample Preparation
4.3. Analysis on Olink Panels
4.4. Evaluation of Data from Olink Panels
4.5. Heatmaps
4.6. Box Plots of Protein Expression
4.7. Immunohistochemistry for VEGFA
4.8. Evaluation of Survival
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Haeggblom, L.; Ramqvist, T.; Tommasino, M.; Dalianis, T.; Nasman, A. Time to change perspectives on hpv in oropharyngeal cancer. A systematic review of hpv prevalence per oropharyngeal sub-site the last 3 years. Papillomavirus Res. 2017, 4, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Marklund, L.; Näsman, A.; Ramqvist, T.; Dalianis, T.; Munck-Wikland, E.; Hammarstedt, L. Prevalence of human papillomavirus and survival in oropharyngeal cancer other than tonsil or base of tongue cancer. Cancer Med. 2012, 1, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Ramqvist, T.; Dalianis, T. Oropharyngeal cancer epidemic and human papillomavirus. Emerg. Infect. Dis. 2010, 16, 1671–1677. [Google Scholar] [CrossRef] [PubMed]
- Dahlstrand, H.; Dahlgren, L.; Lindquist, D.; Munck-Wikland, E.; Dalianis, T. Presence of human papillomavirus in tonsillar cancer is a favourable prognostic factor for clinical outcome. Anticancer Res. 2004, 24, 1829–1835. [Google Scholar] [PubMed]
- Lindquist, D.; Romanitan, M.; Hammarstedt, L.; Nasman, A.; Dahlstrand, H.; Lindholm, J.; Onelov, L.; Ramqvist, T.; Ye, W.; Munck-Wikland, E.; et al. Human papillomavirus is a favourable prognostic factor in tonsillar cancer and its oncogenic role is supported by the expression of e6 and e7. Mol. Oncol. 2007, 1, 350–355. [Google Scholar] [CrossRef] [PubMed]
- Gillison, M.L.; D’Souza, G.; Westra, W.; Sugar, E.; Xiao, W.; Begum, S.; Viscidi, R. Distinct risk factor profiles for human papillomavirus type 16-positive and human papillomavirus type 16-negative head and neck cancers. J. Natl. Cancer Inst. 2008, 100, 407–420. [Google Scholar] [CrossRef] [PubMed]
- Slebos, R.J.; Yi, Y.; Ely, K.; Carter, J.; Evjen, A.; Zhang, X.; Shyr, Y.; Murphy, B.M.; Cmelak, A.J.; Burkey, B.B.; et al. Gene expression differences associated with human papillomavirus status in head and neck squamous cell carcinoma. Clin. Cancer Res. 2006, 12, 701–709. [Google Scholar] [CrossRef] [PubMed]
- Pyeon, D.; Newton, M.A.; Lambert, P.F.; den Boon, J.A.; Sengupta, S.; Marsit, C.J.; Woodworth, C.D.; Connor, J.P.; Haugen, T.H.; Smith, E.M.; et al. Fundamental differences in cell cycle deregulation in human papillomavirus-positive and human papillomavirus-negative head/neck and cervical cancers. Cancer Res. 2007, 67, 4605–4619. [Google Scholar] [CrossRef] [PubMed]
- Dahlgren, L.; Mellin, H.; Wangsa, D.; Heselmeyer-Haddad, K.; Bjornestal, L.; Lindholm, J.; Munck-Wikland, E.; Auer, G.; Ried, T.; Dalianis, T. Comparative genomic hybridization analysis of tonsillar cancer reveals a different pattern of genomic imbalances in human papillomavirus-positive and -negative tumors. Int. J. Cancer 2003, 107, 244–249. [Google Scholar] [CrossRef] [PubMed]
- Lajer, C.B.; Garnaes, E.; Friis-Hansen, L.; Norrild, B.; Therkildsen, M.H.; Glud, M.; Rossing, M.; Lajer, H.; Svane, D.; Skotte, L.; et al. The role of mirnas in human papilloma virus (hpv)-associated cancers: Bridging between hpv-related head and neck cancer and cervical cancer. Br. J. Cancer 2012, 106, 1526–1534. [Google Scholar] [CrossRef] [PubMed]
- Smeets, S.J.; Hesselink, A.T.; Speel, E.J.; Haesevoets, A.; Snijders, P.J.; Pawlita, M.; Meijer, C.J.; Braakhuis, B.J.; Leemans, C.R.; Brakenhoff, R.H. A novel algorithm for reliable detection of human papillomavirus in paraffin embedded head and neck cancer specimen. Int. J. Cancer 2007, 121, 2465–2472. [Google Scholar] [CrossRef] [PubMed]
- Nasman, A.; Nordfors, C.; Grun, N.; Munck-Wikland, E.; Ramqvist, T.; Marklund, L.; Lindquist, D.; Dalianis, T. Absent/weak cd44 intensity and positive human papillomavirus (hpv) status in oropharyngeal squamous cell carcinoma indicates a very high survival. Cancer Med. 2013, 2, 507–518. [Google Scholar] [CrossRef] [PubMed]
- Nasman, A.; Bersani, C.; Lindquist, D.; Du, J.; Ramqvist, T.; Dalianis, T. Human papillomavirus and potentially relevant biomarkers in tonsillar and base of tongue squamous cell carcinoma. Anticancer Res. 2017, 37, 5319–5328. [Google Scholar] [PubMed]
- Badoual, C.; Hans, S.; Rodriguez, J.; Peyrard, S.; Klein, C.; Agueznay Nel, H.; Mosseri, V.; Laccourreye, O.; Bruneval, P.; Fridman, W.H.; et al. Prognostic value of tumor-infiltrating cd4+ t-cell subpopulations in head and neck cancers. Clin. Cancer Res. 2006, 12, 465–472. [Google Scholar] [CrossRef] [PubMed]
- Nasman, A.; Andersson, E.; Marklund, L.; Tertipis, N.; Hammarstedt-Nordenvall, L.; Attner, P.; Nyberg, T.; Masucci, G.V.; Munck-Wikland, E.; Ramqvist, T.; et al. Hla class i and ii expression in oropharyngeal squamous cell carcinoma in relation to tumor hpv status and clinical outcome. PLoS ONE 2013, 8, e77025. [Google Scholar] [CrossRef] [PubMed]
- Nasman, A.; Andersson, E.; Nordfors, C.; Grun, N.; Johansson, H.; Munck-Wikland, E.; Massucci, G.; Dalianis, T.; Ramqvist, T. Mhc class i expression in hpv positive and negative tonsillar squamous cell carcinoma in correlation to clinical outcome. Int. J. Cancer 2013, 132, 72–81. [Google Scholar] [CrossRef] [PubMed]
- Tertipis, N.; Haeggblom, L.; Grun, N.; Nordfors, C.; Nasman, A.; Dalianis, T.; Ramqvist, T. Reduced expression of the antigen processing machinery components tap2, lmp2, and lmp7 in tonsillar and base of tongue cancer and implications for clinical outcome. Transl. Oncol. 2015, 8, 10–17. [Google Scholar] [CrossRef] [PubMed]
- Tertipis, N.; Haeggblom, L.; Nordfors, C.; Grun, N.; Nasman, A.; Vlastos, A.; Dalianis, T.; Ramqvist, T. Correlation of lmp10 expression and clinical outcome in human papillomavirus (hpv) positive and hpv-negative tonsillar and base of tongue cancer. PLoS ONE 2014, 9, e95624. [Google Scholar] [CrossRef] [PubMed]
- Oguejiofor, K.; Galletta-Williams, H.; Dovedi, S.J.; Roberts, D.L.; Stern, P.L.; West, C.M. Distinct patterns of infiltrating cd8+ t cells in hpv+ and cd68 macrophages in hpv-oropharyngeal squamous cell carcinomas are associated with better clinical outcome but pd-l1 expression is not prognostic. Oncotarget 2017, 8, 14416–14427. [Google Scholar] [CrossRef] [PubMed]
- Assarsson, E.; Lundberg, M.; Holmquist, G.; Bjorkesten, J.; Thorsen, S.B.; Ekman, D.; Eriksson, A.; Rennel Dickens, E.; Ohlsson, S.; Edfeldt, G.; et al. Homogenous 96-plex pea immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 2014, 9, e95192. [Google Scholar] [CrossRef] [PubMed]
- Nasman, A.; Romanitan, M.; Nordfors, C.; Grun, N.; Johansson, H.; Hammarstedt, L.; Marklund, L.; Munck-Wikland, E.; Dalianis, T.; Ramqvist, T. Tumor infiltrating cd8+ and foxp3+ lymphocytes correlate to clinical outcome and human papillomavirus (hpv) status in tonsillar cancer. PLoS ONE 2012, 7, e38711. [Google Scholar] [CrossRef] [PubMed]
- Nordfors, C.; Grun, N.; Tertipis, N.; Ahrlund-Richter, A.; Haeggblom, L.; Sivars, L.; Du, J.; Nyberg, T.; Marklund, L.; Munck-Wikland, E.; et al. Cd8+ and cd4+ tumour infiltrating lymphocytes in relation to human papillomavirus status and clinical outcome in tonsillar and base of tongue squamous cell carcinoma. Eur. J. Cancer 2013, 49, 2522–2530. [Google Scholar] [CrossRef] [PubMed]
- Partlova, S.; Boucek, J.; Kloudova, K.; Lukesova, E.; Zabrodsky, M.; Grega, M.; Fucikova, J.; Truxova, I.; Tachezy, R.; Spisek, R.; et al. Distinct patterns of intratumoral immune cell infiltrates in patients with hpv-associated compared to non-virally induced head and neck squamous cell carcinoma. Oncoimmunology 2015, 4, e965570. [Google Scholar] [CrossRef] [PubMed]
- Messina, J.L.; Fenstermacher, D.A.; Eschrich, S.; Qu, X.; Berglund, A.E.; Lloyd, M.C.; Schell, M.J.; Sondak, V.K.; Weber, J.S.; Mule, J.J. 12-chemokine gene signature identifies lymph nod × 10−like structures in melanoma: Potential for patient selection for immunotherapy? Sci. Rep. 2012, 2, 765. [Google Scholar] [CrossRef] [PubMed]
- Sher, Y.P.; Chou, C.C.; Chou, R.H.; Wu, H.M.; Wayne Chang, W.S.; Chen, C.H.; Yang, P.C.; Wu, C.W.; Yu, C.L.; Peck, K. Human kallikrein 8 protease confers a favorable clinical outcome in non-small cell lung cancer by suppressing tumor cell invasiveness. Cancer Res. 2006, 66, 11763–11770. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Gilligan, B.M.; Yuan, J.; Li, T. Current status and perspectives in translational biomarker research for pd-1/pd-l1 immune checkpoint blockade therapy. J. Hematol. Oncol. 2016, 9, 47. [Google Scholar] [CrossRef] [PubMed]
- Honeychurch, J.; Cheadle, E.J.; Dovedi, S.J.; Illidge, T.M. Immuno-regulatory antibodies for the treatment of cancer. Expert Opin. Biol. Ther. 2015, 15, 787–801. [Google Scholar] [CrossRef] [PubMed]
- Hong, A.M.; Vilain, R.E.; Romanes, S.; Yang, J.; Smith, E.; Jones, D.; Scolyer, R.A.; Lee, C.S.; Zhang, M.; Rose, B. Pd-l1 expression in tonsillar cancer is associated with human papillomavirus positivity and improved survival: Implications for anti-pd1 clinical trials. Oncotarget 2016, 7, 77010–77020. [Google Scholar] [CrossRef] [PubMed]
- Felder, M.; Kapur, A.; Gonzalez-Bosquet, J.; Horibata, S.; Heintz, J.; Albrecht, R.; Fass, L.; Kaur, J.; Hu, K.; Shojaei, H.; et al. Muc16 (ca125): Tumor biomarker to cancer therapy, a work in progress. Mol. Cancer 2014, 13, 129. [Google Scholar] [CrossRef] [PubMed]
- Slebos, R.J.; Jehmlich, N.; Brown, B.; Yin, Z.; Chung, C.H.; Yarbrough, W.G.; Liebler, D.C. Proteomic analysis of oropharyngeal carcinomas reveals novel hpv-associated biological pathways. Int. J. Cancer 2013, 132, 568–579. [Google Scholar] [CrossRef] [PubMed]
- Sewell, A.; Brown, B.; Biktasova, A.; Mills, G.B.; Lu, Y.; Tyson, D.R.; Issaeva, N.; Yarbrough, W.G. Revers × 10−phase protein array profiling of oropharyngeal cancer and significance of pik3ca mutations in hpv-associated head and neck cancer. Clin. Cancer Res. 2014, 20, 2300–2311. [Google Scholar] [CrossRef] [PubMed]
- Pazina, T.; Shemesh, A.; Brusilovsky, M.; Porgador, A.; Campbell, K.S. Regulation of the functions of natural cytotoxicity receptors by interactions with diverse ligands and alterations in splice variant expression. Front. Immunol. 2017, 8, 369. [Google Scholar] [CrossRef] [PubMed]
- Freud, A.G.; Mundy-Bosse, B.L.; Yu, J.; Caligiuri, M.A. The broad spectrum of human natural killer cell diversity. Immunity 2017, 47, 820–833. [Google Scholar] [CrossRef] [PubMed]
- Mirghani, H.; Ugolin, N.; Ory, C.; Lefevre, M.; Baulande, S.; Hofman, P.; St Guily, J.L.; Chevillard, S.; Lacave, R. A predictive transcriptomic signature of oropharyngeal cancer according to hpv16 status exclusively. Oral Oncol. 2014, 50, 1025–1034. [Google Scholar] [CrossRef] [PubMed]
- Gaykalova, D.A.; Manola, J.B.; Ozawa, H.; Zizkova, V.; Morton, K.; Bishop, J.A.; Sharma, R.; Zhang, C.; Michailidi, C.; Considine, M.; et al. Nf-kappab and stat3 transcription factor signatures differentiate hpv-positive and hpv-negative head and neck squamous cell carcinoma. Int. J. Cancer 2015, 137, 1879–1889. [Google Scholar] [CrossRef] [PubMed]
- Keck, M.K.; Zuo, Z.; Khattri, A.; Stricker, T.P.; Brown, C.D.; Imanguli, M.; Rieke, D.; Endhardt, K.; Fang, P.; Bragelmann, J.; et al. Integrative analysis of head and neck cancer identifies two biologically distinct hpv and three non-hpv subtypes. Clin. Cancer Res. 2015, 21, 870–881. [Google Scholar] [CrossRef] [PubMed]
- Wilkie, M.D.; Emmett, M.S.; Santosh, S.; Lightbody, K.A.; Lane, S.; Goodyear, P.W.; Sheard, J.D.; Boyd, M.T.; Pritchard-Jones, R.O.; Jones, T.M. Relative expression of vascular endothelial growth factor isoforms in squamous cell carcinoma of the head and neck. Head Neck 2016, 38, 775–781. [Google Scholar] [CrossRef] [PubMed]
- Fei, J.; Hong, A.; Dobbins, T.A.; Jones, D.; Lee, C.S.; Loo, C.; Al-Ghamdi, M.; Harnett, G.B.; Clark, J.; O’Brien, C.J.; et al. Prognostic significance of vascular endothelial growth factor in squamous cell carcinomas of the tonsil in relation to human papillomavirus status and epidermal growth factor receptor. Ann. Surg. Oncol. 2009, 16, 2908–2917. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.P.; Chen, H.L. Increased vascular endothelial growth factor expression predicts a worse prognosis for laryngeal cancer patients: A meta-analysis. J. Laryngol. Otol. 2017, 131, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.F.; Yang, X.D.; Lu, M.X.; Sun, G.W.; Wang, Y.X.; Zhang, Y.K.; Pu, Y.M.; Tang, E.Y. Prognostic significance of vegf immunohistochemical expression in oral cancer: A meta-analysis of the literature. Tumour Biol. 2013, 34, 3165–3171. [Google Scholar] [CrossRef] [PubMed]
- Roudnicky, F.; Poyet, C.; Wild, P.; Krampitz, S.; Negrini, F.; Huggenberger, R.; Rogler, A.; Stohr, R.; Hartmann, A.; Provenzano, M.; et al. Endocan is upregulated on tumor vessels in invasive bladder cancer where it mediates vegf-a-induced angiogenesis. Cancer Res. 2013, 73, 1097–1106. [Google Scholar] [CrossRef] [PubMed]
- Sagara, A.; Igarashi, K.; Otsuka, M.; Kodama, A.; Yamashita, M.; Sugiura, R.; Karasawa, T.; Arakawa, K.; Narita, M.; Kuzumaki, N.; et al. Endocan as a prognostic biomarker of tripl × 10−negative breast cancer. Breast Cancer Res. Treat. 2017, 161, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Chen, C.; Wang, X.; Zhang, J.Y.; Ren, B.H.; Ma, D.W.; Xia, L.; Xu, X.Y.; Xu, L. Prognostic value of endocan expression in cancers: Evidence from meta-analysis. Oncol. Targets Ther. 2016, 9, 6297–6304. [Google Scholar] [CrossRef] [PubMed]
- Irshad, K.; Mohapatra, S.K.; Srivastava, C.; Garg, H.; Mishra, S.; Dikshit, B.; Sarkar, C.; Gupta, D.; Chandra, P.S.; Chattopadhyay, P.; et al. A combined gene signature of hypoxia and notch pathway in human glioblastoma and its prognostic relevance. PLoS ONE 2015, 10, e0118201. [Google Scholar] [CrossRef] [PubMed]
- Tudisco, L.; Orlandi, A.; Tarallo, V.; De Falco, S. Hypoxia activates placental growth factor expression in lymphatic endothelial cells. Oncotarget 2017, 8, 32873–32883. [Google Scholar] [CrossRef] [PubMed]
- van der Veeken, J.; Oliveira, S.; Schiffelers, R.M.; Storm, G.; van Bergen En Henegouwen, P.M.; Roovers, R.C. Crosstalk between epidermal growth factor receptor-and insulin-like growth factor-1 receptor signaling: Implications for cancer therapy. Curr. Cancer Drug Targets 2009, 9, 748–760. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yuan, J.L.; Zhang, Y.T.; Ma, J.J.; Xu, P.; Shi, C.H.; Zhang, W.; Li, Y.M.; Fu, Q.; Zhu, G.F.; et al. Inhibition of both egfr and igf1r sensitized prostate cancer cells to radiation by synergistic suppression of DNA homologous recombination repair. PLoS ONE 2013, 8, e68784. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, F.; Fujimaki, M.; Ohba, S.; Kojima, M.; Yokoyama, J.; Ikeda, K. Relationship between insulin-like growth factor-1 receptor and human papillomavirus in patients with oropharyngeal cancer. Head Neck 2015, 37, 977–981. [Google Scholar] [CrossRef] [PubMed]
- Bersani, C.; Mints, M.; Tertipis, N.; Haeggblom, L.; Sivars, L.; Ahrlund-Richter, A.; Vlastos, A.; Smedberg, C.; Grun, N.; Munck-Wikland, E.; et al. A model using concomitant markers for predicting outcome in human papillomavirus positive oropharyngeal cancer. Oral Oncol. 2017, 68, 53–59. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate -a practical and powerful approach to multiple testing. J. Roy. Stat. Soc. B Met. 1995, 57, 289–300. [Google Scholar]
| Patient and Tumor Characteristics | HPV+TSCC/BOTSCC (n = 42) | HPV-TSCC/BOTSCC (n = 17) | All TSCC/BOTSCC (n = 59) | Validation set (n = 49) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | n | % | |||
| Age | Mean (years) | 61.7 | 60.6 | 61.4 | 62.1 | |||||
| Median (years) | 61 | 64 | 61 | 61 | ||||||
| Range (years) | 46–84 | 32–83 | 32–84 | 42–84 | ||||||
| Diagnose | malignant neoplasm of the base of tongue (C01.9) | 9 | 21% | 9 | 53% | 18 | 31% | 13 | 22% | |
| malignant neoplasm of the tonsil (C09.0-9) | 33 | 79% | 8 | 47% | 41 | 69% | 36 | 61% | ||
| Sex | female | 8 | 19% | 3 | 18% | 11 | 19% | 13 | 22% | |
| male | 34 | 81% | 14 | 82% | 48 | 81% | 36 | 61% | ||
| Tumour differentiation | poorly | 21 | 50% | 9 | 53% | 30 | 51% | 29 | 49% | |
| moderatley | 17 | 40% | 8 | 47% | 25 | 42% | 18 | 31% | ||
| well | 2 | 5% | 0 | 0% | 2 | 3% | 1 | 2% | ||
| undefined | 2 | 5% | 0 | 0% | 2 | 3% | 1 | 2% | ||
| Tumour size | T1 | 6 | 14% | 3 | 18% | 9 | 15% | 8 | 14% | |
| T2 | 16 | 38% | 2 | 12% | 18 | 31% | 20 | 34% | ||
| T3 | 11 | 26% | 5 | 29% | 16 | 27% | 10 | 17% | ||
| T4 | 9 | 21% | 7 | 41% | 16 | 27% | 11 | 19% | ||
| Nodal disease | N0 | 5 | 12% | 6 | 35% | 11 | 19% | 5 | 8% | |
| N1 | 12 | 29% | 1 | 6% | 13 | 22% | 11 | 19% | ||
| N2a | 5 | 12% | 3 | 18% | 8 | 14% | 5 | 8% | ||
| N2b | 15 | 36% | 3 | 18% | 18 | 31% | 21 | 36% | ||
| N2c | 5 | 12% | 2 | 12% | 7 | 12% | 7 | 12% | ||
| N3 | 0 | 0% | 1 | 6% | 1 | 2% | 0 | 0% | ||
| NX | 0 | 0% | 1 | 6% | 1 | 2% | 0 | 0% | ||
| Distant metastasis | M0 | 42 | 100% | 16 | 94% | 58 | 98% | 48 | 81% | |
| M1 | 0 | 0% | 0 | 0% | 0 | 0% | 0 | 0% | ||
| MX | 0 | 0% | 1 | 6% | 1 | 2% | 1 | 2% | ||
| Tumour Stage | I | 0 | 0% | 3 | 18% | 3 | 5% | 0 | 0% | |
| II | 4 | 10% | 1 | 6% | 5 | 8% | 3 | 5% | ||
| III | 11 | 26% | 2 | 12% | 13 | 22% | 10 | 17% | ||
| IVa | 27 | 64% | 9 | 53% | 36 | 61% | 34 | 58% | ||
| IVb | 0 | 0% | 1 | 6% | 1 | 2% | 0 | 0% | ||
| IVc | 0 | 0% | 0 | 0% | 0 | 0% | 1 | 2% | ||
| Unknown | 0 | 0% | 1 | 6% | 1 | 2% | 0 | 0% | ||
| Treatment | Induction chemotherapy and radiation | conventional | 4 | 10% | 3 | 18% | 7 | 12% | 8 | 14% |
| accelerated | 11 | 26% | 9 | 53% | 20 | 34% | 14 | 24% | ||
| Radiation | conventional | 20 | 48% | 4 | 24% | 24 | 41% | 21 | 36% | |
| accelerated | 7 | 17% | 1 | 6% | 8 | 14% | 6 | 10% | ||
| Brachytherapy boost | Not administered | 34 | 81% | 12 | 71% | 46 | 78% | 38 | 64% | |
| Administered | 8 | 19% | 7 | 41% | 15 | 25% | 11 | 19% | ||
| Concomittant Cetuximab | Not administered | 35 | 83% | 13 | 76% | 48 | 81% | 42 | 71% | |
| Administered | 7 | 17% | 4 | 24% | 11 | 19% | 7 | 12% | ||
| Smoking | Never | 15 | 36% | 2 | 12% | 17 | 29% | 16 | 27% | |
| Former (>15 years ago) | 11 | 26% | 1 | 6% | 12 | 20% | 11 | 19% | ||
| Former (<15 years ago) | 7 | 17% | 3 | 18% | 10 | 17% | 11 | 19% | ||
| Current upon diagnosis | 9 | 21% | 11 | 65% | 20 | 34% | 11 | 19% | ||
| Protein | HPV Positive TSCC/BOTSCC vs. Normal | HPV Negative TSCC/BOTSCC vs. Normal | ||
|---|---|---|---|---|
| Ratio | p-Value | Ratio | p-Value | |
| CA9 | 55.72 | 1.10 × 10−21 | 50.56 | 1.80 × 10−8 |
| CXCL13 | 37.27 | 3.70 × 10−14 | 21.26 | 1.40 × 10−5 |
| CXCL10 | 36.25 | 1.40 × 10−19 | 15.89 | 1.50 × 10−6 |
| CCL4 | 35.75 | 9.30 × 10−24 | 20.25 | 1.80 × 10−9 |
| MMP-12 | 35.51 | 4.60 × 10−18 | 33.36 | 3.90 × 10−9 |
| CCL3 | 34.78 | 1.30 × 10−20 | 38.85 | 3.70 × 10−10 |
| CCL20 | 25.81 | 5.40 × 10−13 | 11.31 | 0.00011 |
| CXCL11 | 22.32 | 3.90 × 10−18 | 11.39 | 6.30 × 10−6 |
| IL-6 | 20.97 | 3.40 × 10−15 | 33.59 | 3.40 × 10−8 |
| CXCL9 | 19.43 | 2.50 × 10−18 | 9.78 | 7.70 × 10−6 |
| IL-8 | 19.16 | 5.90 × 10−15 | 24.42 | 1.20 × 10−7 |
| TNFRSF9 | 18.25 | 1.50 × 10−15 | 11 | 3.80 × 10−6 |
| WISP-1 | 16.34 | 2.30 × 10−23 | 13.64 | 2.10 × 10−9 |
| Protein | Ratio HPV-Positive/HPV-Negative (Linear) | p-Value | False Discovery Rate (FDR) |
|---|---|---|---|
| MUC-16 | 4.89 | 0.0016 | 0.062 |
| CXCL17 | 2.73 | 0.025 | 0.162 |
| CCL20 | 2.51 | 0.0046 | 0.101 |
| CD8A | 2.23 | 0.024 | 0.162 |
| FASL | 2.23 | 0.0065 | 0.101 |
| CCL4 | 2.20 | 0.00093 | 0.057 |
| CXCL10 | 2.19 | 0.037 | 0.186 |
| PD-L1 | 2.13 | 0.0024 | 0.062 |
| IL-12 | 2.10 | 0.025 | 0.162 |
| LY9 | 1.88 | 0.026 | 0.162 |
| CD27 | 1.83 | 0.025 | 0.162 |
| PDCD1 | 1.62 | 0.031 | 0.173 |
| KLRD1 | 1.61 | 0.018 | 0.148 |
| DKN1A | 1.53 | 0.031 | 0.173 |
| GZMH | 1.52 | 0.046 | 0.209 |
| NCR1 | 1.51 | 0.016 | 0.148 |
| IL12RB1 | 1.45 | 0.015 | 0.148 |
| CPE | 1.45 | 0.047 | 0.209 |
| IL-7 | 1.37 | 0.016 | 0.148 |
| VEGFC | 0.89 | 0.046 | 0.209 |
| ITGAV | 0.78 | 0.017 | 0.148 |
| HO-1 | 0.74 | 0.0096 | 0.120 |
| GZMB | 0.74 | 0.01 | 0.120 |
| FURIN | 0.73 | 0.035 | 0.186 |
| TXLNA | 0.72 | 0.027 | 0.162 |
| GPNMB | 0.70 | 0.0054 | 0.101 |
| TNFRSF12A | 0.66 | 0.046 | 0.209 |
| GAL-9 | 0.59 | 0.0011 | 0.057 |
| ADAMTS15 | 0.59 | 0.009 | 0.120 |
| HK8 | 0.45 | 0.037 | 0.186 |
| TFPI2 | 0.35 | 0.0061 | 0.101 |
| AR | 0.34 | 0.0024 | 0.062 |
| WIF-1 | 0.24 | 0.018 | 0.148 |
| HK14 | 0.16 | 0.00032 | 0.050 |
| Protein | Ratio * (Linear) | p-Value |
|---|---|---|
| DLL1 | 8.38 | 0.000422 |
| ESM-1 | 5.76 | 0.00286 |
| TNFRSF19 | 5.41 | 0.000145 |
| VEGFA | 4.01 | 0.00241 |
| CYR61 | 3.64 | 0.0044 |
| CCL7 | 3.23 | 0.0402 |
| PLGF | 2.89 | 0.0127 |
| MIC-A/B | 2.67 | 0.0452 |
| ANG-2 | 2.33 | 0.00218 |
| TNFRSF21 | 2.20 | 0.0411 |
| IGF1R | 1.78 | 0.00847 |
| CSF-1 | 1.74 | 0.0361 |
| GPNMB | 1.62 | 0.0347 |
| ICOSLG | 1.41 | 0.0441 |
| IFN-GAMMA-R1 | 1.40 | 0.037 |
| Protein | Ratio * (Linear) | p-Value |
|---|---|---|
| WFDC2 | 4.11 | 0.000339 |
| EGF | 4.03 | 0.0239 |
| MIA | 2.84 | 0.00962 |
| VEGFR-2 | 0.76 | 0.0275 |
| EPHA2 | 0.68 | 0.0167 |
| DKN1A | 0.58 | 0.0382 |
| ANG2 | 0.52 | 0.00641 |
| TNFRSF4 | 0.51 | 0.0449 |
| TRAIL | 0.50 | 0.045 |
| CXCL11 | 0.26 | 0.0462 |
| IL-1-ALPHA | 0.26 | 0.00104 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramqvist, T.; Näsman, A.; Franzén, B.; Bersani, C.; Alexeyenko, A.; Becker, S.; Haeggblom, L.; Kolev, A.; Dalianis, T.; Munck-Wikland, E. Protein Expression in Tonsillar and Base of Tongue Cancer and in Relation to Human Papillomavirus (HPV) and Clinical Outcome. Int. J. Mol. Sci. 2018, 19, 978. https://doi.org/10.3390/ijms19040978
Ramqvist T, Näsman A, Franzén B, Bersani C, Alexeyenko A, Becker S, Haeggblom L, Kolev A, Dalianis T, Munck-Wikland E. Protein Expression in Tonsillar and Base of Tongue Cancer and in Relation to Human Papillomavirus (HPV) and Clinical Outcome. International Journal of Molecular Sciences. 2018; 19(4):978. https://doi.org/10.3390/ijms19040978
Chicago/Turabian StyleRamqvist, Torbjörn, Anders Näsman, Bo Franzén, Cinzia Bersani, Andrey Alexeyenko, Susanne Becker, Linnea Haeggblom, Aeneas Kolev, Tina Dalianis, and Eva Munck-Wikland. 2018. "Protein Expression in Tonsillar and Base of Tongue Cancer and in Relation to Human Papillomavirus (HPV) and Clinical Outcome" International Journal of Molecular Sciences 19, no. 4: 978. https://doi.org/10.3390/ijms19040978





