A Neutrophil Proteomic Signature in Surgical Trauma Wounds
Abstract
1. Introduction
2. Results
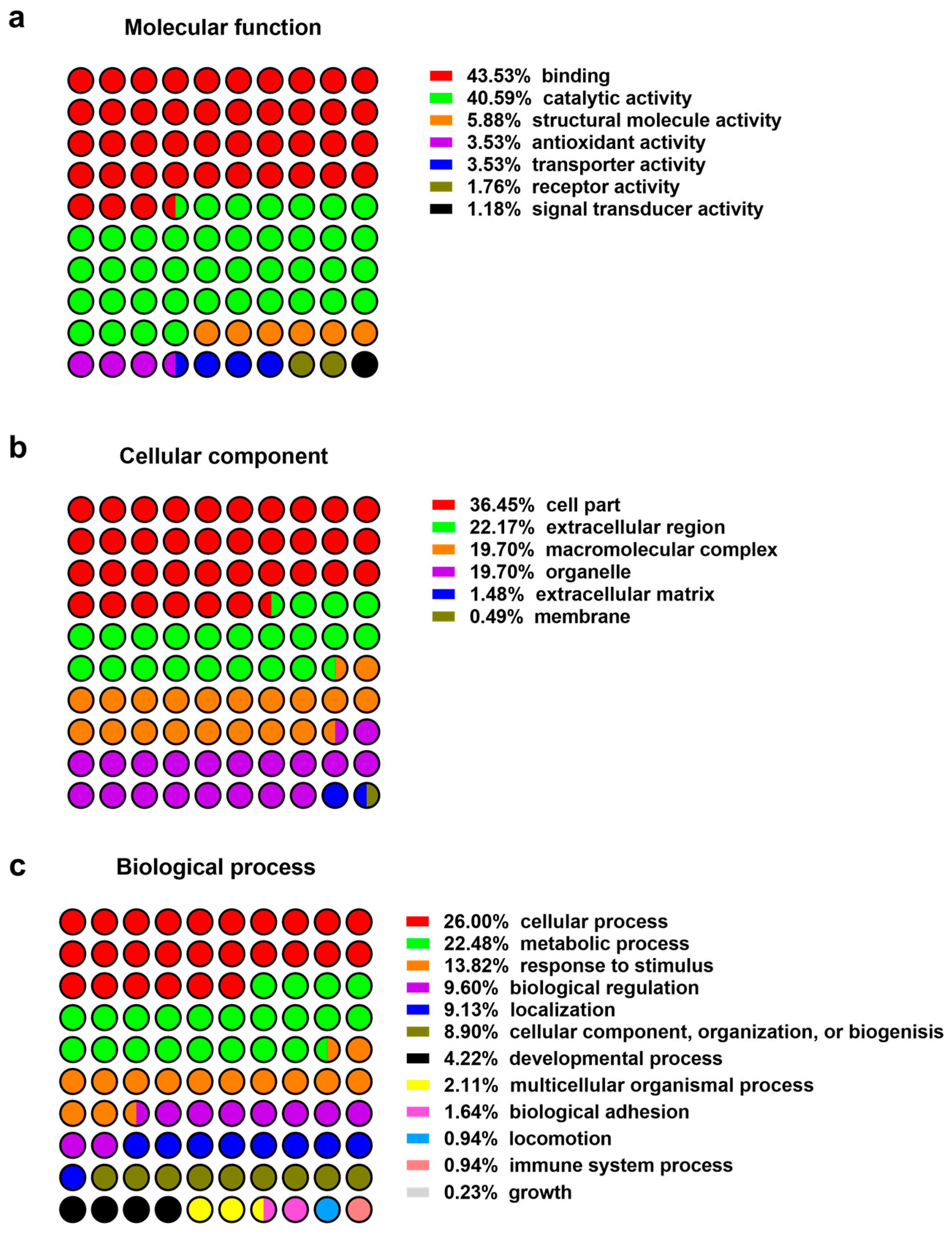
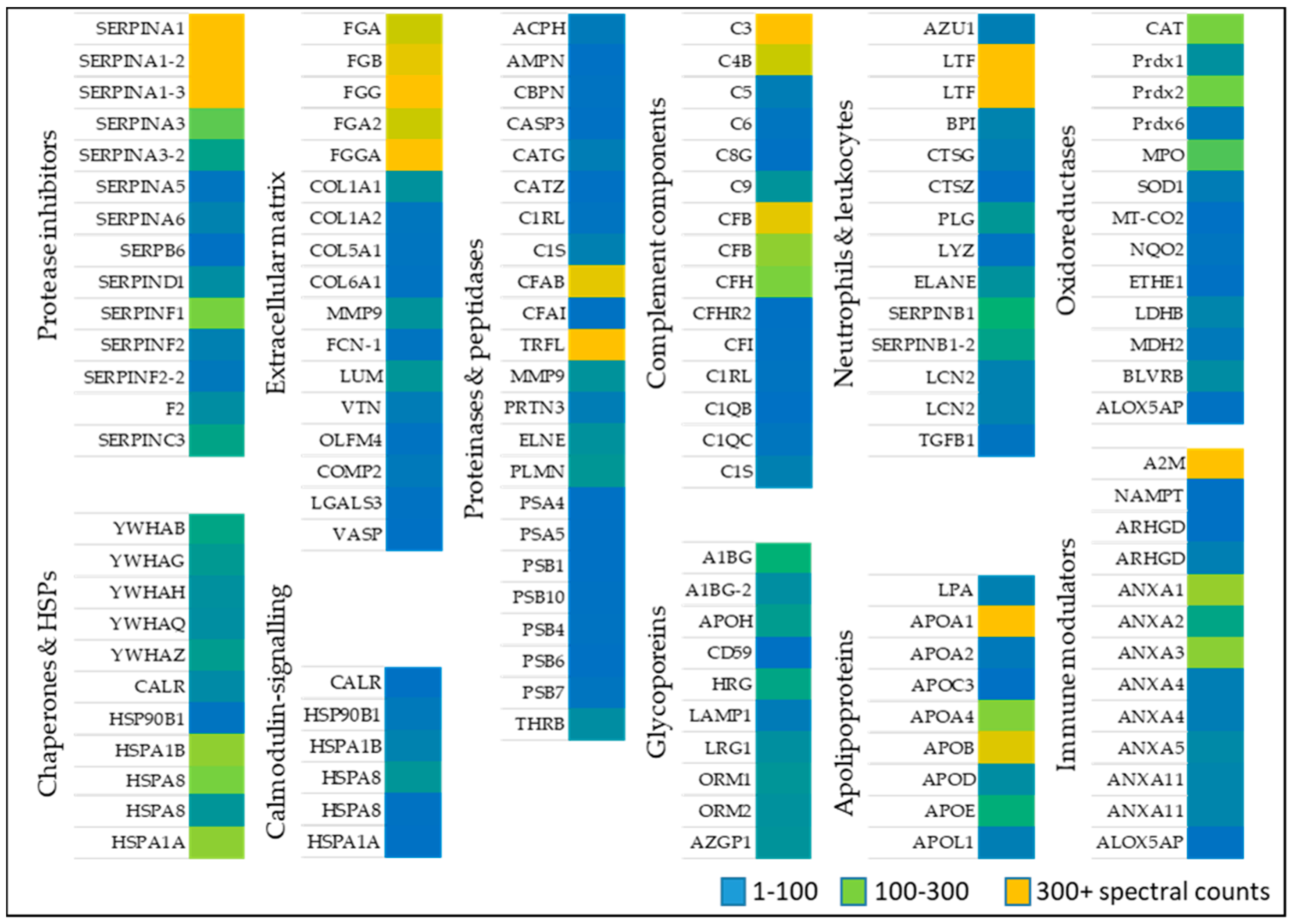
2.1. Proteomics of Wound Exudates
2.1.1. Oxidoreductases
2.1.2. Immune Modulators, Chaperones, and Heat Shock Proteins
2.1.3. Neutrophil and Leukocytes Associated Factors
2.1.4. Extracellular Matrix Proteins
2.1.5. Proteinases
2.1.6. Post-Translational Modifications
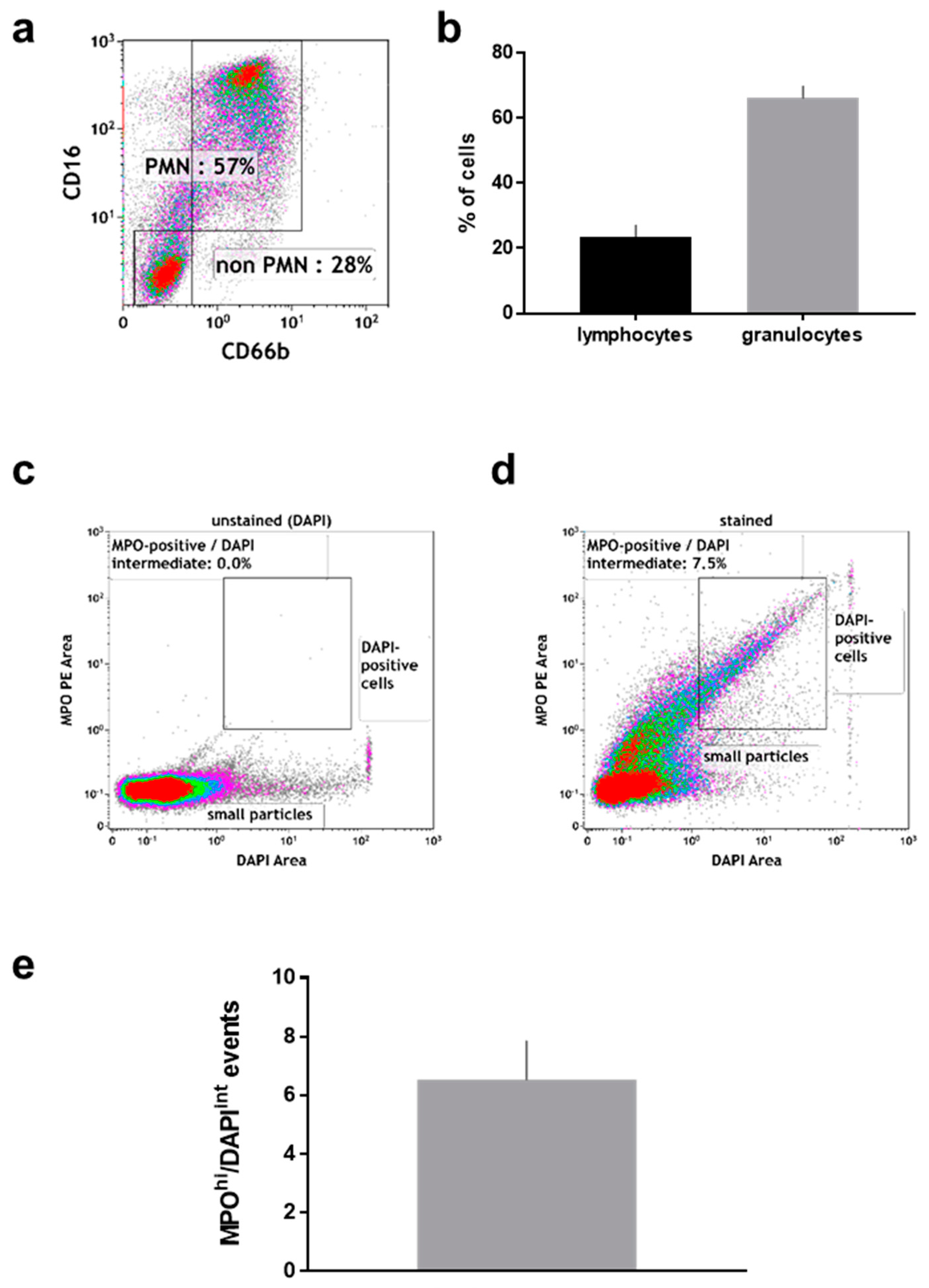
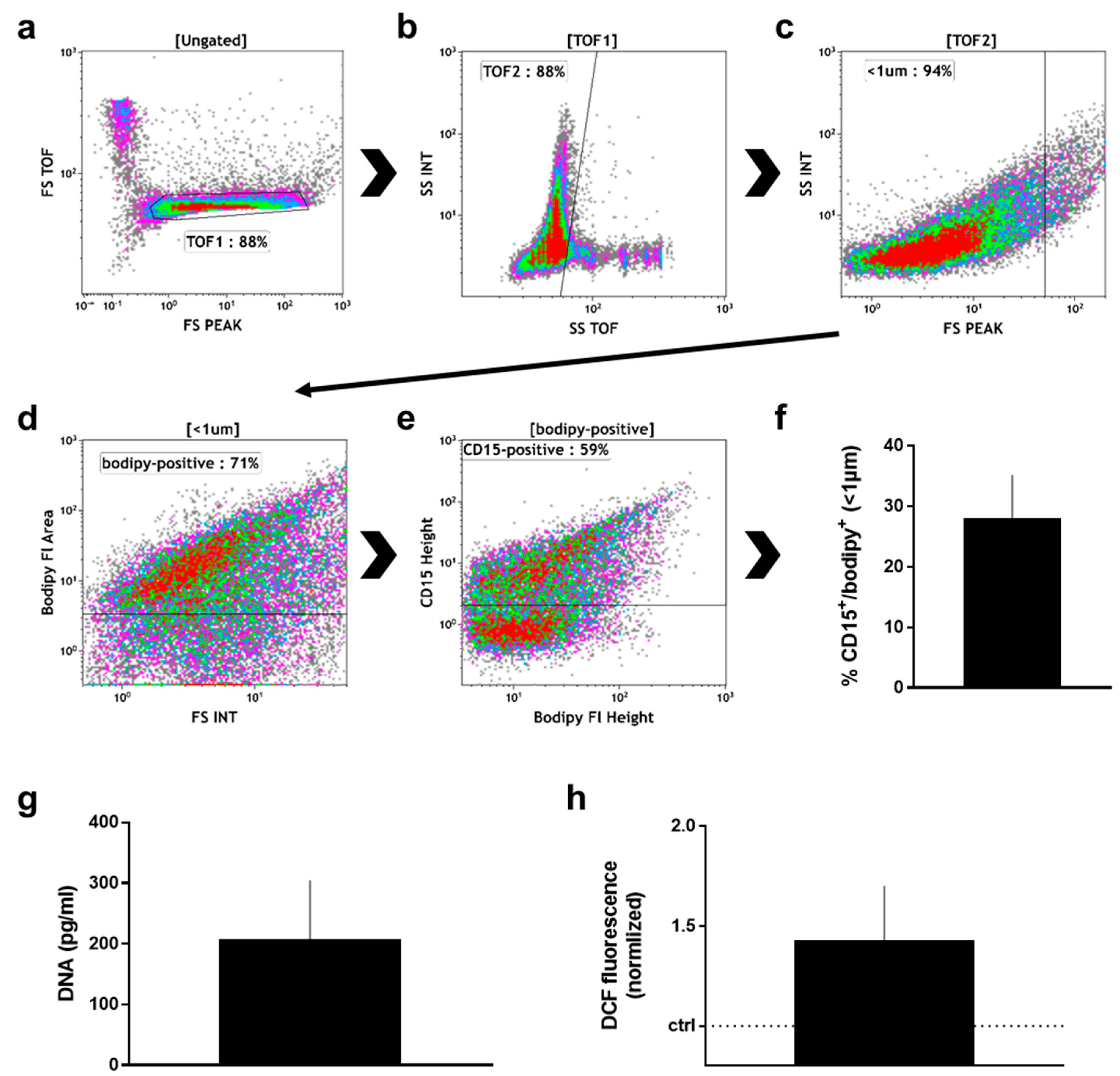
2.2. Manually Obtained Wound Material
3. Discussion
4. Materials and Methods
4.1. Wound Material
4.2. Protein Preparation from Material in Wound Sponge Dressings
4.3. Mass Spectrometry
4.4. Data Analysis
4.5. Analysis of Manually Collected Wound Material
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A
| Acronym | Protein Name | Acronym |
|---|---|---|
| P08519 | LPA | Apolipoprotein(a) |
| P02647 | APOA1 | Apolipoprotein A-I |
| P02652 | APOA2 | Apolipoprotein A-II |
| P02656 | APOC3 | Apolipoprotein C-III |
| P06727 | APOA4 | Apolipoprotein A-IV |
| P04114 | APOB | Apolipoprotein B-100 |
| P05090 | APOD | Apolipoprotein D |
| P02649 | APOE | Apolipoprotein E |
| O14791-3 | APOL1 | Isoform 3 of Apolipoprotein L1 |
| P04217 | A1BG | Alpha-1B-glycoprotein |
| P04217-2 | A1BG-2 | Isoform 2 of Alpha-1B-glycoprotein |
| P02749 | APOH | Beta-2-glycoprotein 1 |
| P13987 | CD59 | CD59 glycoprotein |
| P04196 | HRG | Histidine-rich glycoprotein |
| P11279 | LAMP1 | Lysosome-associated membrane glycoprotein |
| P02750 | LRG1 | Leucine-rich α-2-glycoprotein |
| P02763 | ORM1 | Alpha-1-acid glycoprotein |
| P02765 | ORM2 | Alpha-2-acid glycoprotein |
| P25311 | AZGP1 | Zinc-α-2-glycoprotein |
| P01009 | SERPINA1 | Alpha-1-antitrypsin |
| P01009-2 | SERPINA1-2 | Isoform 2 of α-1-antitrypsin |
| P01009-3 | SERPINA1-3 | Isoform 3 of α-1-antitrypsin |
| P01011 | SERPINA3 | Alpha-1-antichymotrypsin |
| P01011-2 | SERPINA3-2 | Isoform 2 of α-1-antichymotrypsin |
| P05154 | SERPINA5 | Plasma serine protease inhibitor |
| P08185 | SERPINA6 | Corticosteroid-binding globulin |
| P35237 | SERPB6 | Serpin B6 |
| P05546 | SERPIND1 | Heparin cofactor 2 |
| P08670 | SERPINF1 | Pigment epithelium-derived factor |
| P08697 | SERPINF2 | Alpha-2-antiplasmin |
| P08697-2 | SERPINF2-2 | Isoform 2 of α-2-antiplasmin |
| P00734 | F2 | Prothrombin |
| P01008 | SERPINC3 | Antithrombin-III |
| Acronym | Protein Name | Acronym |
|---|---|---|
| P01024 | C3 | Complement C3 |
| P0C0L5 | C4B | Complement C4-B |
| P01031 | C5 | Complement C5 |
| P13671 | C6 | Complement component C6 |
| P07360 | C8G | Complement component C8 γ chain |
| P02748 | C9 | Complement component C9 |
| P00751 | CFB | Complement factor B |
| P00751-2 | CFB | Isoform 2 of Complement factor B |
| P08603 | CFH | Complement factor H |
| P36980 | CFHR2 | Complement factor H-related protein 2 |
| P05156 | CFI | Complement factor I |
| Q9NZP8 | C1RL | Complement C1r subcomponent-like protein |
| P02746 | C1QB | Complement C1q subcomponent subunit B |
| P02747 | C1QC | Complement C1q subcomponent subunit C |
| P09871 | C1S | Complement C1s subcomponent |
| Protein ID | Acronym | Protein Name | PTMs a |
|---|---|---|---|
| P01834 | IGKC | Immunoglobulin kappa constant | Trioxidation [C87] |
| P15814 | IGLL1 | Immunoglobulin lambda-like polypeptide 1 | Trioxidation [C194]; Oxidation [M197] |
| Q04695 | KRT17 | Keratin, type I cytoskeletal 17 | Oxidation [M88] |
| P30043 | BLVRB | Flavin reductase (NADPH) | Oxidation [M87]; |
| P02549 | SPTA1 | Spectrin α chain, erythrocytic 1 | Oxidation [M647; M881] |
| P02549-2 | SPTA1 | Isoform 2 of Spectrin α chain, erythrocytic | Oxidation [M647; M881] |
| P11142 | HSPA8 | Heat shock cognate 71 kDa protein | Oxidation [M61] |
| P0DMV9 | HSPA1B | Heat shock 70 kDa protein 1B | Oxidation [M549] |
| P0DMV8-2 | HSPA1A | Isoform 2 of Heat shock 70 kDa protein 1A | Oxidation [M494] |
| Q14624 | ITIH4 | Inter-α-trypsin inhibitor heavy chain H4 | Oxidation [M491] |
| Q562R1 | ACTBL2 | Beta-actin-like protein 2 | Oxidation [M45; M48; M191]; |
| P26038 | MSN | Moesin | Oxidation [M433; M451] |
| P12429 | ANXA3 | Annexin A3 | Oxidation [M40] |
| P01871 | IGHM | Immunoglobulin heavy constant mu | Oxidation [M383]; |
| P01871-2 | IGHM | Isoform 2 of Immunoglobulin heavy constant mu | Oxidation [M383]; |
| P04004 | VTN | Vitronectin | Oxidation [M350]; |
| P35542 | SAA4 | Serum amyloid A-4 protein | Oxidation [M35] |
| P30101 | PDIA3 | Protein disulfide-isomerase A3 | Oxidation [M338] |
| P06727 | APOA4 | Apolipoprotein A-IV | Oxidation [M322] |
| P30740 | SERPINB1 | Leukocyte elastase inhibitor | Oxidation [M307] |
| P04114 | APOB | Apolipoprotein B-100 | Oxidation [M306; M3007; M3421] |
| P04264 | KRT1 | Keratin, type II cytoskeletal 1 | Oxidation [M296]; |
| P35237 | SERPINB6 | Serpin B6 | Oxidation [M291] |
| P40121 | CAPG | Macrophage-capping protein | Oxidation [M261] |
| P02671 | FGA | Fibrinogen α chain | Oxidation [M259] |
| P02671-2 | FGA | Isoform 2 of Fibrinogen α chain | Oxidation [M259] |
| P40121-2 | CAPG | Isoform 2 of Macrophage-capping protein | Oxidation [M246] |
| P61981 | YWHAG | 14-3-3 protein γ | Oxidation [M23; M27] |
| P02790 | HPX | Hemopexin | Oxidation [M229; M375; M409]; |
| P47756-2 | CAPZB | Isoform 2 of F-actin-capping protein subunit β | Oxidation [M220] |
| Q06830 | PRDX1 | Peroxiredoxin-1 | Oxidation [M21] |
| P27105 | STOM | Erythrocyte band 7 integral membrane protein | Oxidation [M207; M228] |
| O15145 | ARPC3 | Actin-related protein 2/3 complex subunit 3 | Oxidation [M19] |
| P01860 | IGHG3 | Immunoglobulin heavy constant γ 3 | Oxidation [M182; M288]; Trioxidation [C27; C297]; |
| P09525 | ANXA4 | Annexin A4 | Oxidation [M17] |
| P19823 | ITIH2 | Inter-α-trypsin inhibitor heavy chain H2 | Oxidation [M162] |
| P35527 | KRT9 | Keratin, type I cytoskeletal 9 | Oxidation [M157; M234; M245; M269] |
| P08670 | VIM | Vimentin | Oxidation [M154] |
| P13645 | KRT10 | Keratin, type I cytoskeletal 10 | Oxidation [M150] |
| P00915 | CA1 | Carbonic anhydrase 1 | Oxidation [M149; M242] |
| P01857 | IGHG1 | Immunoglobulin heavy constant γ 1 | Oxidation [M135]; Trioxidation [C27; C250]; Nitro [Y161] |
| P01861 | IGHG4 | Immunoglobulin heavy constant γ 4 | Oxidation [M132]; Trioxidation [C247]; |
| P01859 | IGHG2 | Immunoglobulin heavy constant γ 2 | Oxidation [M131; M237]; Trioxidation [C246]; |
| P02787 | TF | Serotransferrin | Oxidation [M128; M332; C374]; Trioxidation [C374] |
| P08779 | KRT16 | Keratin, type I cytoskeletal 16 | Oxidation [M121]; |
| P02533 | KRT14 | Keratin, type I cytoskeletal 14 | Oxidation [M119]; |
| P02647 | APOA1 | Apolipoprotein A-I | Oxidation [M110; M136; M172] |
| P37837 | TALDO1 | Transaldolase | Oxidation [M11] |
| P02679-2 | FGG | Isoform γ-A of Fibrinogen γ chain | Oxidation [M104; M290]; |
| P02679 | FGG | Fibrinogen γ chain | Oxidation [M104; M290]; |
| P02792 | FTL | Ferritin light chain | Oxidation [M101] |
| A0M8Q6 | IGLC7 | Immunoglobulin lambda constant 7 | Oxidation [C87]; Nitrosyl [C87]; Nitro [Y85] |
| A0A0C4DH72 | IGKV1 | Immunoglobulin kappa variable 1–6 | Oxidation [C22; M26] |
| B9A064 | IGLL5 | Immunoglobulin lambda-like polypeptide 5 | Oxidation [C195]; Trioxidation [C195] |
| P00450 | CP | Ceruloplasmin | Nitro [Y471]; Oxidation [M538] |
| P68871 | HBB | Hemoglobin subunit β | Nitro [Y36]; Oxidation [M56]; |
| P02042 | HBD | Hemoglobin subunit δ | Nitro [Y36] |
| P00738 | HP | Haptoglobin | Nitro [Y280]; Oxidation [M263; M300; M343] |
| P69905 | HBA1 | Hemoglobin subunit α | Nitro [Y25]; Oxidation [M33] |
| P00738-2 | HP | Isoform 2 of Haptoglobin | Nitro [Y221]; Oxidation [M204; M241; M284] |
| P01009 | SERPINA1 | Alpha-1-antitrypsin | Nitro [Y184]; Oxidation [M409] |
| P01009-3 | SERPINA1 | Isoform 3 of Alpha-1-antitrypsin | Nitro [Y184] |
| P01009-2 | SERPINA1 | Isoform 2 of Alpha-1-antitrypsin | Nitro [Y184] |
| P49913 | CAMP | Cathelicidin antimicrobial peptide | Nitro [Y56]; Oxidation [M109] |
| P02768 | ALB | Serum albumin | Oxidation [M111; C125; C289; M353; C500; C501; M572]; Trioxidation [C125; C192; C224; C289; C302; C303; C500; C501; C591]; Nitro [Y164; Y287]; Nitrosyl [C289] |
| P02768-3 | ALB | Isoform 3 of Serum albumin | Oxidation [M111; C125; C287; C288; M359]; Trioxidation [C125; C287; C288; C378] |
| P02768-2 | ALB | Isoform 2 of Serum albumin | Oxidation [C97; M161; C308; C309; M380]; Trioxidation [C97; C110; C111; C308; C309; C399]; Nitro [Y95]; Nitrosyl [C97] |
| P0C0L5 | C4B | Complement C4-B | Oxidation [M393] |
| P01023 | A2M | Alpha-2-macroglobulin | Oxidation [M713; M1378] |
| P01024 | C3 | Complement C3 | Oxidation [M164; M495; M581; M990; M1347; M1384] |
| P08603 | CFH | Complement factor H | Oxidation [M1174] |
| P02788 | LTF | Lactotransferrin | Oxidation [C28; M411]; Trioxidation [C28] |
| P02675 | FGB | Fibrinogen β chain | Oxidation [M254; M272] |
| P02788-2 | LTF | Isoform δLf of Lactotransferrin | Oxidation [M367] |
| P02763 | ORM1 | Alpha-1-acid glycoprotein 1 | Nitro [Y175] |
| P04406 | GAPDH | Glyceraldehyde-3-phosphate dehydrogenase | Oxidation [M328; M331] |
| P07738 | BPGM | Bisphosphoglycerate mutase | Oxidation [M103] |
| P13796 | LCP1 | Plastin-2 | Oxidation [M166] |
| P04406-2 | GAPDH | Isoform 2 of Glyceraldehyde-3-phosphate dehydrogenase | Oxidation [M286; M289] |
| P07738 | BPGM | Bisphosphoglycerate mutase | Oxidation [M103] |
| P13796 | LCP1 | Plastin-2 | Oxidation [M166] |
References
- Broughton, G., II; Janis, J.E.; Attinger, C.E. The basic science of wound healing. Plast. Reconstr. Surg. 2006, 117, 12S–34S. [Google Scholar] [CrossRef] [PubMed]
- Hunt, T.K.; Hopf, H.; Hussain, Z. Physiology of wound healing. Adv. Skin Wound Care 2000, 13, 6–11. [Google Scholar] [CrossRef]
- Gillitzer, R.; Goebeler, M. Chemokines in cutaneous wound healing. J. Leukoc. Biol. 2001, 69, 513–521. [Google Scholar] [PubMed]
- Behm, B.; Babilas, P.; Landthaler, M.; Schreml, S. Cytokines, chemokines and growth factors in wound healing. J. Eur. Acad. Dermatol. Venereol. 2012, 26, 812–820. [Google Scholar] [CrossRef] [PubMed]
- Okizaki, S.; Ito, Y.; Hosono, K.; Oba, K.; Ohkubo, H.; Amano, H.; Shichiri, M.; Majima, M. Suppressed recruitment of alternatively activated macrophages reduces TGF-β1 and impairs wound healing in streptozotocin-induced diabetic mice. Biomed. Pharmacother. 2015, 70, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Werner, S.; Krieg, T.; Smola, H. Keratinocyte-fibroblast interactions in wound healing. J. Investig. Dermatol. 2007, 127, 998–1008. [Google Scholar] [CrossRef] [PubMed]
- Heyer, K.; Herberger, K.; Protz, K.; Glaeske, G.; Augustin, M. Epidemiology of chronic wounds in Germany: Analysis of statutory health insurance data. Wound Repair Regen. 2016, 24, 434–442. [Google Scholar] [CrossRef] [PubMed]
- Spear, M. Acute or chronic? What’s the difference? Plast. Surg. Nurs. 2013, 33, 98–100. [Google Scholar] [CrossRef] [PubMed]
- Menke, N.B.; Ward, K.R.; Witten, T.M.; Bonchev, D.G.; Diegelmann, R.F. Impaired wound healing. Clin. Dermatol. 2007, 25, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Bowler, P.G. Wound pathophysiology, infection and therapeutic options. Ann. Med. 2002, 34, 419–427. [Google Scholar] [CrossRef] [PubMed]
- Eming, S.A.; Krieg, T.; Davidson, J.M. Inflammation in wound repair: Molecular and cellular mechanisms. J. Investig. Dermatol. 2007, 127, 514–525. [Google Scholar] [CrossRef] [PubMed]
- Falanga, V. Wound healing and its impairment in the diabetic foot. Lancet 2005, 366, 1736–1743. [Google Scholar] [CrossRef]
- Frank, D.N.; Wysocki, A.; Specht-Glick, D.D.; Rooney, A.; Feldman, R.A.; St Amand, A.L.; Pace, N.R.; Trent, J.D. Microbial diversity in chronic open wounds. Wound Repair Regen. 2009, 17, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Larson, B.J.; Longaker, M.T.; Lorenz, H.P. Scarless fetal wound healing: A basic science review. Plast. Reconstr. Surg. 2010, 126, 1172–1180. [Google Scholar] [CrossRef] [PubMed]
- Yager, D.R.; Kulina, R.A.; Gilman, L.A. Wound fluids: A window into the wound environment? Int. J. Lower Extrem. Wounds 2007, 6, 262–272. [Google Scholar] [CrossRef] [PubMed]
- Diegelmann, R.F. Excessive neutrophils characterize chronic pressure ulcers. Wound Repair Regen. 2003, 11, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Amulic, B.; Cazalet, C.; Hayes, G.L.; Metzler, K.D.; Zychlinsky, A. Neutrophil function: From mechanisms to disease. Annu. Rev. Immunol. 2012, 30, 459–489. [Google Scholar] [CrossRef] [PubMed]
- Korkmaz, B.; Moreau, T.; Gauthier, F. Neutrophil elastase, proteinase 3 and cathepsin G: Physicochemical properties, activity and physiopathological functions. Biochimie 2008, 90, 227–242. [Google Scholar] [CrossRef] [PubMed]
- Brinkmann, V.; Reichard, U.; Goosmann, C.; Fauler, B.; Uhlemann, Y.; Weiss, D.S.; Weinrauch, Y.; Zychlinsky, A. Neutrophil extracellular traps kill bacteria. Science 2004, 303, 1532–1535. [Google Scholar] [CrossRef] [PubMed]
- Kambas, K.; Chrysanthopoulou, A.; Vassilopoulos, D.; Apostolidou, E.; Skendros, P.; Girod, A.; Arelaki, S.; Froudarakis, M.; Nakopoulou, L.; Giatromanolaki, A. Tissue factor expression in neutrophil extracellular traps and neutrophil derived microparticles in antineutrophil cytoplasmic antibody associated vasculitis may promote thromboinflammation and the thrombophilic state associated with the disease. Ann. Rheum. Dis. 2014, 73, 1854–1863. [Google Scholar] [CrossRef] [PubMed]
- Dalli, J.; Montero-Melendez, T.; Norling, L.V.; Yin, X.; Hinds, C.; Haskard, D.; Mayr, M.; Perretti, M. Heterogeneity in neutrophil microparticles reveals distinct proteome and functional properties. Mol. Cell. Proteom. 2013, 12, 2205–2219. [Google Scholar] [CrossRef] [PubMed]
- El-Benna, J.; Dang, P.M.; Gougerot-Pocidalo, M.A. Priming of the neutrophil NADPH oxidase activation: Role of p47phox phosphorylation and NOX2 mobilization to the plasma membrane. Semin. Immunopathol. 2008, 30, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Winterbourn, C.C.; Kettle, A.J.; Hampton, M.B. Reactive oxygen species and neutrophil function. Annu. Rev. Biochem. 2016, 85, 765–792. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Khanna, S.; Nallu, K.; Hunt, T.K.; Sen, C.K. Dermal wound healing is subject to redox control. Mol. Ther. 2006, 13, 211–220. [Google Scholar] [CrossRef] [PubMed]
- Niethammer, P.; Grabher, C.; Look, A.T.; Mitchison, T.J. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 2009, 459, 996–999. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K. The general case for redox control of wound repair. Wound Repair Regen. 2003, 11, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Eming, S.A.; Koch, M.; Krieger, A.; Brachvogel, B.; Kreft, S.; Bruckner-Tuderman, L.; Krieg, T.; Shannon, J.D.; Fox, J.W. Differential proteomic analysis distinguishes tissue repair biomarker signatures in wound exudates obtained from normal healing and chronic wounds. J. Proteome Res. 2010, 9, 4758–4766. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Li, P.; Shi, Z.; Hou, T.; Chen, X.; Du, J. A prospective, randomized, controlled study of hyperbaric oxygen therapy: Effects on healing and oxidative stress of ulcer tissue in patients with a diabetic foot ulcer. Ostomy Wound Manag. 2013, 59, 18–24. [Google Scholar]
- Kwon, J.; Wang, A.; Burke, D.J.; Boudreau, H.E.; Lekstrom, K.J.; Korzeniowska, A.; Sugamata, R.; Kim, Y.S.; Yi, L.; Ersoy, I.; et al. Peroxiredoxin 6 (Prdx6) supports NADPH oxidase1 (Nox1)-based superoxide generation and cell migration. Free Radic. Biol. Med. 2016, 96, 99–115. [Google Scholar] [CrossRef] [PubMed]
- Soubhye, J.; Aldib, I.; Delporte, C.; Prévost, M.; Dufrasne, F.; Van Antwerpen, P. Myeloperoxidase as a target for the treatment of inflammatory syndromes: Mechanisms and structure activity relationships of inhibitors. Curr. Med. Chem. 2016, 23, 3975–4008. [Google Scholar] [CrossRef] [PubMed]
- Tomson, F.L.; Morgan, J.E.; Gu, G.; Barquera, B.; Vygodina, T.V.; Gennis, R.B. Substitutions for glutamate 101 in subunit II of cytochrome c oxidase from Rhodobacter sphaeroides result in blocking the proton-conducting K-channel. Biochemistry 2003, 42, 1711–1717. [Google Scholar] [CrossRef] [PubMed]
- Wilgus, T.A.; Bergdall, V.K.; Tober, K.L.; Hill, K.J.; Mitra, S.; Flavahan, N.A.; Oberyszyn, T.M. The impact of cyclooxygenase-2 mediated inflammation on scarless fetal wound healing. Am. J. Pathol. 2004, 165, 753–761. [Google Scholar] [CrossRef]
- Kato, G.J.; McGowan, V.; Machado, R.F.; Little, J.A.; Taylor, J.T.; Morris, C.R.; Nichols, J.S.; Wang, X.; Poljakovic, M.; Morris, S.M., Jr.; et al. Lactate dehydrogenase as a biomarker of hemolysis-associated nitric oxide resistance, priapism, leg ulceration, pulmonary hypertension, and death in patients with sickle cell disease. Blood 2006, 107, 2279–2285. [Google Scholar] [CrossRef] [PubMed]
- Bellaye, P.S.; Burgy, O.; Causse, S.; Garrido, C.; Bonniaud, P. Heat shock proteins in fibrosis and wound healing: Good or evil? Pharmacol. Ther. 2014, 143, 119–132. [Google Scholar] [CrossRef] [PubMed]
- Borth, W. Alpha 2-macroglobulin, a multifunctional binding protein with targeting characteristics. FASEB J. 1992, 6, 3345–3353. [Google Scholar] [CrossRef] [PubMed]
- Revollo, J.R.; Korner, A.; Mills, K.F.; Satoh, A.; Wang, T.; Garten, A.; Dasgupta, B.; Sasaki, Y.; Wolberger, C.; Townsend, R.R.; et al. Nampt/PBEF/Visfatin regulates insulin secretion in β cells as a systemic NAD biosynthetic enzyme. Cell Metab. 2007, 6, 363–375. [Google Scholar] [CrossRef] [PubMed]
- Ridley, A.J. Rho family proteins: Coordinating cell responses. Trends Cell Biol. 2001, 11, 471–477. [Google Scholar] [CrossRef]
- Watt, F.M.; Fujiwara, H. Cell-extracellular matrix interactions in normal and diseased skin. Cold Spring Harb. Perspect. Biol. 2011, 3, a005124. [Google Scholar] [CrossRef] [PubMed]
- Cheung, E.Y.; Weijers, E.M.; Tuk, B.; Scheffer, R.; Leebeek, F.W.; van Neck, J.W.; Koolwijk, P.; de Maat, M.P. Specific effects of fibrinogen and the γA and γ’-chain fibrinogen variants on angiogenesis and wound healing. Tissue Eng. Part A 2015, 21, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Klaas, M.; Kangur, T.; Viil, J.; Maemets-Allas, K.; Minajeva, A.; Vadi, K.; Antsov, M.; Lapidus, N.; Jarvekulg, M.; Jaks, V. The alterations in the extracellular matrix composition guide the repair of damaged liver tissue. Sci. Rep. 2016, 6, 27398. [Google Scholar] [CrossRef] [PubMed]
- Pozuelo Rubio, M.; Geraghty, K.M.; Wong, B.H.; Wood, N.T.; Campbell, D.G.; Morrice, N.; Mackintosh, C. 14-3-3-affinity purification of over 200 human phosphoproteins reveals new links to regulation of cellular metabolism, proliferation and trafficking. Biochem. J. 2004, 379, 395–408. [Google Scholar] [CrossRef] [PubMed]
- Meng, L.; Mohan, R.; Kwok, B.H.; Elofsson, M.; Sin, N.; Crews, C.M. Epoxomicin, a potent and selective proteasome inhibitor, exhibits in vivo antiinflammatory activity. Proc. Natl. Acad. Sci. USA 1999, 96, 10403–10408. [Google Scholar] [CrossRef] [PubMed]
- Luan, Y.; Xu, W. The structure and main functions of aminopeptidase N. Curr. Med. Chem. 2007, 14, 639–647. [Google Scholar] [CrossRef] [PubMed]
- Pulakazhi Venu, V.K.; Uboldi, P.; Dhyani, A.; Patrini, A.; Baetta, R.; Ferri, N.; Corsini, A.; Muro, A.F.; Catapano, A.L.; Norata, G.D. Fibronectin extra domain A stabilises atherosclerotic plaques in apolipoprotein E and in LDL-receptor-deficient mice. Thromb. Haemost. 2015, 114, 186–197. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, R.; Ichioka, S.; Sekiya, N.; Ohura, N.; Uchino, S.; Ojima, A.; Itoh, Y.; Ishihara, O.; Nakatsuka, T.; Ikebuchi, K. Elastic plasma protein film blended with platelet releasate accelerates healing of diabetic mouse skin wounds. Vox Sang. 2007, 93, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Herring, J.M.; McMichael, M.A.; Smith, S.A. Microparticles in health and disease. J. Vet. Intern. Med. 2013, 27, 1020–1033. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, C.Y.; Tsai, T.H.; Chak, K.F. The molecular basis of wound healing processes induced by lithospermi radix: A proteomics and biochemical analysis. Evid. Based Complement. Alternat. Med. 2012, 2012, 508972. [Google Scholar] [CrossRef] [PubMed]
- Kalkhof, S.; Forster, Y.; Schmidt, J.; Schulz, M.C.; Baumann, S.; Weissflog, A.; Gao, W.; Hempel, U.; Eckelt, U.; Rammelt, S.; et al. Proteomics and metabolomics for in situ monitoring of wound healing. Biomed. Res. Int. 2014, 2014, 934848. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, C.Y.; Hung, C.Y.; Tsai, T.H.; Chak, K.F. A Study of the Wound Healing Mechanism of a Traditional Chinese Medicine, Angelica sinensis, Using a Proteomic Approach. Evid. Based Complement. Alternat. Med. 2012, 2012, 467531. [Google Scholar] [CrossRef] [PubMed]
- Steinstrasser, L.; Jacobsen, F.; Hirsch, T.; Kesting, M.; Chojnacki, C.; Krisp, C.; Wolters, D. Immunodepletion of high-abundant proteins from acute and chronic wound fluids to elucidate low-abundant regulators in wound healing. BMC Res. Notes 2010, 3, 335. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K. Wound healing essentials: Let there be oxygen. Wound Repair Regen. 2009, 17, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Luo, B.; Wang, J.; Liu, Z.; Shen, Z.; Shi, R.; Liu, Y.Q.; Liu, Y.; Jiang, M.; Wu, Y.; Zhang, Z. Phagocyte respiratory burst activates macrophage erythropoietin signalling to promote acute inflammation resolution. Nat. Commun. 2016, 7, 12177. [Google Scholar] [CrossRef] [PubMed]
- Hüttemann, M.; Lee, I.; Grossman, L.I.; Doan, J.W.; Sanderson, T.H. Phosphorylation of mammalian cytochrome c and cytochrome c oxidase in the regulation of cell destiny: Respiration, apoptosis, and human disease. In Mitochondrial Oxidative Phosphorylation; Springer: Berlin, Germany, 2012; pp. 237–264. [Google Scholar]
- Winterbourn, C.C. Reconciling the chemistry and biology of reactive oxygen species. Nat. Chem. Biol. 2008, 4, 278–286. [Google Scholar] [CrossRef] [PubMed]
- Winterbourn, C.C.; Kettle, A.J. Reactions of superoxide with myeloperoxidase and its products. Jpn. J. Infect. Dis. 2004, 57, S31–S33. [Google Scholar] [PubMed]
- Metzler, K.D.; Fuchs, T.A.; Nauseef, W.M.; Reumaux, D.; Roesler, J.; Schulze, I.; Wahn, V.; Papayannopoulos, V.; Zychlinsky, A. Myeloperoxidase is required for neutrophil extracellular trap formation: Implications for innate immunity. Blood 2011, 117, 953–959. [Google Scholar] [CrossRef] [PubMed]
- Fuchs, T.A.; Abed, U.; Goosmann, C.; Hurwitz, R.; Schulze, I.; Wahn, V.; Weinrauch, Y.; Brinkmann, V.; Zychlinsky, A. Novel cell death program leads to neutrophil extracellular traps. J. Cell Biol. 2007, 176, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Bekeschus, S.; Winterbourn, C.C.; Kolata, J.; Masur, K.; Hasse, S.; Broker, B.M.; Parker, H.A. Neutrophil extracellular trap formation is elicited in response to cold physical plasma. J. Leukoc. Biol. 2016, 100, 791–799. [Google Scholar] [CrossRef] [PubMed]
- Wong, S.L.; Demers, M.; Martinod, K.; Gallant, M.; Wang, Y.; Goldfine, A.B.; Kahn, C.R.; Wagner, D.D. Diabetes primes neutrophils to undergo NETosis, which impairs wound healing. Nat. Med. 2015, 21, 815–819. [Google Scholar] [CrossRef] [PubMed]
- Meng, W.; Paunel-Gorgulu, A.; Flohe, S.; Witte, I.; Schadel-Hopfner, M.; Windolf, J.; Logters, T.T. Deoxyribonuclease is a potential counter regulator of aberrant neutrophil extracellular traps formation after major trauma. Mediat. Inflamm. 2012, 2012, 149560. [Google Scholar] [CrossRef] [PubMed]
- Akong-Moore, K.; Chow, O.A.; von Kockritz-Blickwede, M.; Nizet, V. Influences of chloride and hypochlorite on neutrophil extracellular trap formation. PLoS ONE 2012, 7, e42984. [Google Scholar] [CrossRef] [PubMed]
- Munafo, D.B.; Johnson, J.L.; Brzezinska, A.A.; Ellis, B.A.; Wood, M.R.; Catz, S.D. DNase I inhibits a late phase of reactive oxygen species production in neutrophils. J. Innate Immun. 2009, 1, 527–542. [Google Scholar] [CrossRef] [PubMed]
- Rada, B.; Jendrysik, M.A.; Pang, L.; Hayes, C.P.; Yoo, D.G.; Park, J.J.; Moskowitz, S.M.; Malech, H.L.; Leto, T.L. Pyocyanin-enhanced neutrophil extracellular trap formation requires the NADPH oxidase. PLoS ONE 2013, 8, e54205. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K.; Roy, S. Redox signals in wound healing. Biochim. Biophys. Acta 2008, 1780, 1348–1361. [Google Scholar] [CrossRef] [PubMed]
- Wood, Z.A.; Schroder, E.; Robin Harris, J.; Poole, L.B. Structure, mechanism and regulation of peroxiredoxins. Trends Biochem. Sci. 2003, 28, 32–40. [Google Scholar] [CrossRef]
- Serhan, C.N.; Chiang, N.; Dalli, J. The resolution code of acute inflammation: Novel pro-resolving lipid mediators in resolution. Semin. Immunol. 2015, 27, 200–215. [Google Scholar] [CrossRef] [PubMed]
- Gordts, S.C.; Muthuramu, I.; Amin, R.; Jacobs, F.; de Geest, B. The impact of lipoproteins on wound healing: Topical HDL therapy corrects delayed wound healing in apolipoprotein e deficient mice. Pharmaceuticals 2014, 7, 419–432. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Liu, L.; Song, Z.; Hu, Z.Y.; Liu, J.; Hou, D.R. Genetic variations of oxidative stress related genes ALOX5, ALOX5AP and MPO modulate ischemic stroke susceptibility through main effects and epistatic interactions in a Chinese population. Cell. Physiol. Biochem. 2017, 43, 1588–1602. [Google Scholar] [CrossRef] [PubMed]
- Barrientos, S.; Stojadinovic, O.; Golinko, M.S.; Brem, H.; Tomic-Canic, M. Growth factors and cytokines in wound healing. Wound Repair Regen. 2008, 16, 585–601. [Google Scholar] [CrossRef] [PubMed]
- Galkowska, H.; Wojewodzka, U.; Olszewski, W.L. Chemokines, cytokines, and growth factors in keratinocytes and dermal endothelial cells in the margin of chronic diabetic foot ulcers. Wound Repair Regen. 2006, 14, 558–565. [Google Scholar] [CrossRef] [PubMed]
- Barrientos, S.; Brem, H.; Stojadinovic, O.; Tomic-Canic, M. Clinical application of growth factors and cytokines in wound healing. Wound Repair Regen. 2014, 22, 569–578. [Google Scholar] [CrossRef] [PubMed]
- Bekeschus, S.; Schmidt, A.; Napp, M.; Kramer, A.; Kerner, W.; von Woedtke, T.; Wende, K.; Hasse, S.; Masur, K. Distinct cytokine and chemokine patterns in chronic diabetic ulcers and acute wounds. Exp. Dermatol. 2017, 26, 145–147. [Google Scholar] [CrossRef] [PubMed]
- Segal, A.W. How neutrophils kill microbes. Annu. Rev. Immunol. 2005, 23, 197–223. [Google Scholar] [CrossRef] [PubMed]
- Cole, A.M.; Shi, J.; Ceccarelli, A.; Kim, Y.H.; Park, A.; Ganz, T. Inhibition of neutrophil elastase prevents cathelicidin activation and impairs clearance of bacteria from wounds. Blood 2001, 97, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Odell, E.W.; Sarra, R.; Foxworthy, M.; Chapple, D.S.; Evans, R.W. Antibacterial activity of peptides homologous to a loop region in human lactoferrin. FEBS Lett. 1996, 382, 175–178. [Google Scholar] [CrossRef]
- Benarafa, C.; Priebe, G.P.; Remold-O’Donnell, E. The neutrophil serine protease inhibitor serpinb1 preserves lung defense functions in Pseudomonas aeruginosa infection. J. Exp. Med. 2007, 204, 1901–1909. [Google Scholar] [CrossRef] [PubMed]
- Sulniute, R.; Shen, Y.; Guo, Y.Z.; Fallah, M.; Ahlskog, N.; Ny, L.; Rakhimova, O.; Broden, J.; Boija, H.; Moghaddam, A.; et al. Plasminogen is a critical regulator of cutaneous wound healing. Thromb. Haemost. 2016, 115, 1001–1009. [Google Scholar] [CrossRef] [PubMed]
- Herren, T.; Burke, T.A.; Jardi, M.; Felez, J.; Plow, E.F. Regulation of plasminogen binding to neutrophils. Blood 2001, 97, 1070–1078. [Google Scholar] [CrossRef] [PubMed]
- Dentener, M.A.; Francot, G.J.M.; Smit, F.T.; Froon, A.H.M.; Pennings, H.J.; Wouters, E.F.M.; Buurman, W.A. Presence of bactericidal/permeability-increasing protein in disease: Detection by elisa. J. Infect. Dis. 1995, 171, 739–743. [Google Scholar] [CrossRef] [PubMed]
- Weiss, J.; Elsbach, P.; Olsson, I.; Odeberg, H. Purification and characterization of a potent bactericidal and membrane active protein from the granules of human polymorphonuclear leukocytes. J. Biol. Chem. 1978, 253, 2664–2672. [Google Scholar] [PubMed]
- Xue, M.; Jackson, C.J. Extracellular matrix reorganization during wound healing and its impact on abnormal scarring. Adv. Wound Care 2015, 4, 119–136. [Google Scholar] [CrossRef] [PubMed]
- De Boer, J.; Creasey, A.; Chang, A.; Abbink, J.; Roem, D.; Eerenberg, A.; Hack, C.; Taylor, F. Alpha-2-macroglobulin functions as an inhibitor of fibrinolytic, clotting, and neutrophilic proteinases in sepsis: Studies using a baboon model. Infect. Immun. 1993, 61, 5035–5043. [Google Scholar] [PubMed]
- Nguyen, T.N.; Uemura, A.; Shih, W.; Yamada, S. Zyxin-mediated actin assembly is required for efficient wound closure. J. Biol. Chem. 2010, 285, 35439–35445. [Google Scholar] [CrossRef] [PubMed]
- Ghiran, I.; Klickstein, L.B.; Nicholson-Weller, A. Calreticulin is at the surface of circulating neutrophils and uses CD59 as an adaptor molecule. J. Biol. Chem. 2003, 278, 21024–21031. [Google Scholar] [CrossRef] [PubMed]
- Lu, Q.; Xin, Y.; Ye, F.; Foulks, G.; Li, Q. 14-3-3sigma controls corneal epithelium homeostasis and wound healing. Investig. Ophthalmol. Vis. Sci. 2011, 52, 2389–2396. [Google Scholar] [CrossRef] [PubMed]
- Hansraj, N.Z.; Xiao, L.; Wu, J.; Chen, G.; Turner, D.J.; Wang, J.Y.; Rao, J.N. Posttranscriptional regulation of 14-3-3zeta by RNA-binding protein HuR modulating intestinal epithelial restitution after wounding. Physiol. Rep. 2016, 4, e12858. [Google Scholar] [CrossRef] [PubMed]
- Donato, R.; Cannon, B.R.; Sorci, G.; Riuzzi, F.; Hsu, K.; Weber, D.J.; Geczy, C.L. Functions of S100 proteins. Curr. Mol. Med. 2013, 13, 24–57. [Google Scholar] [CrossRef] [PubMed]
- Lerchenmueller, C.; Rengo, G.; Katus, H.A.; Koch, W.J.; Peppel, K.; Most, P. S100A1 deficiency impairs post-ischemic angiogenesis via compromised proangiogenic endothelial cell function and nitric oxide synthase regulation. Circulation 2012, 126. [Google Scholar] [CrossRef]
- Hsu, K.; Champaiboon, C.; Guenther, B.D.; Sorenson, B.S.; Khammanivong, A.; Ross, K.F.; Geczy, C.L.; Herzberg, M.C. Anti-infective protective properties of S100 calgranulins. Antiinflamm. Antiallergy Agents Med. Chem. 2009, 8, 290–305. [Google Scholar] [CrossRef] [PubMed]
- Du, M.; Wang, G.; Ismail, T.M.; Gross, S.; Fernig, D.G.; Barraclough, R.; Rudland, P.S. S100P dissociates myosin IIA filaments and focal adhesion sites to reduce cell adhesion and enhance cell migration. J. Biol. Chem. 2012, 287, 15330–15344. [Google Scholar] [CrossRef] [PubMed]
- Sakaguchi, M.; Sonegawa, H.; Murata, H.; Kitazoe, M.; Futami, J.; Kataoka, K.; Yamada, H.; Huh, N.H. S100A11, an dual mediator for growth regulation of human keratinocytes. Mol. Biol. Cell 2008, 19, 78–85. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, A.; von Woedtke, T.; Bekeschus, S. Periodic exposure of keratinocytes to cold physical plasma: an in vitro model for redox-related diseases of the skin. Oxid. Med. Cell. Longev. 2016, 2016, 9816072. [Google Scholar] [CrossRef] [PubMed]
- Van Genderen, H.O.; Kenis, H.; Hofstra, L.; Narula, J.; Reutelingsperger, C.P. Extracellular annexin A5: Functions of phosphatidylserine-binding and two-dimensional crystallization. Biochim. Biophys. Acta 2008, 1783, 953–963. [Google Scholar] [CrossRef] [PubMed]
- Gerke, V.; Creutz, C.E.; Moss, S.E. Annexins: Linking Ca2+ signalling to membrane dynamics. Nat. Rev. Mol. Cell Biol. 2005, 6, 449–461. [Google Scholar] [CrossRef] [PubMed]
- Gasser, O. Microparticles Released by Ectocytosis from Human Neutrophils: Characterisation, Properties and Functions. Ph.D. Thesis, University of Basel, Basel, Switzerland, 2004. [Google Scholar]
- Gasser, O.; Hess, C.; Miot, S.; Deon, C.; Sanchez, J.C.; Schifferli, J.A. Characterisation and properties of ectosomes released by human polymorphonuclear neutrophils. Exp. Cell Res. 2003, 285, 243–257. [Google Scholar] [CrossRef]
- Bekeschus, S.; Moritz, J.; Schmidt, A.; Wende, K. Redox regulation of leukocyte-derived microparticle release and protein content in response to cold physical plasma-derived oxidants. Clin. Plasma Med. 2017, 7–8, 24–35. [Google Scholar] [CrossRef]
- Trushin, S.A.; Pennington, K.N.; Algeciras-Schimnich, A.; Paya, C.V. Protein Kinase C and Calcineurin Synergize to Activate IκB Kinase and NF-κB in T Lymphocytes. J. Biol. Chem. 1999, 274, 22923–22931. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.K.; Starnes, T.W.; Deng, Q.; Huttenlocher, A. Lyn is a redox sensor that mediates leukocyte wound attraction in vivo. Nature 2011, 480, 109. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Santoriello, C.; Mione, M.; Hurlstone, A.; Martin, P. Live imaging of innate immune cell sensing of transformed cells in zebrafish larvae: Parallels between tumor initiation and wound inflammation. PLoS Biol. 2010, 8, e1000562. [Google Scholar] [CrossRef] [PubMed]
- Moreira, S.; Stramer, B.; Evans, I.; Wood, W.; Martin, P. Prioritization of competing damage and developmental signals by migrating macrophages in the Drosophila embryo. Curr. Biol. 2010, 20, 464–470. [Google Scholar] [CrossRef] [PubMed]
- Shigenaga, M.K.; Lee, H.H.; Blount, B.C.; Christen, S.; Shigeno, E.T.; Yip, H.; Ames, B.N. Inflammation and NOx-induced nitration: Assay for 3-nitrotyrosine by HPLC with electrochemical detection. Proc. Natl. Acad. Sci. USA 1997, 94, 3211–3216. [Google Scholar] [CrossRef] [PubMed]
- Birnboim, H.C.; Lemay, A.M.; Lam, D.K.Y.; Goldstein, R.; Webb, J.R. Cutting edge: MHC class II-restricted peptides containing the inflammation-associated marker 3-nitrotyrosine evade central tolerance and elicit a robust cell-mediated immune response. J. Immunol. 2014, 171, 528–532. [Google Scholar] [CrossRef]
- Nakazawa, H.; Fukuyama, N.; Takizawa, S.; Tsuji, C.; Yoshitake, M.; Ishida, H. Nitrotyrosine formation and its role in various pathological conditions. Free Radic. Res. 2000, 33, 771–784. [Google Scholar] [CrossRef] [PubMed]
- Hernansanz-Agustin, P.; Izquierdo-Alvarez, A.; Garcia-Ortiz, A.; Ibiza, S.; Serrador, J.M.; Martinez-Ruiz, A. Nitrosothiols in the immune system: Signaling and protection. Antioxid. Redox Signal. 2013, 18, 288–308. [Google Scholar] [CrossRef] [PubMed]
- Gostner, J.M.; Becker, K.; Fuchs, D.; Sucher, R. Redox regulation of the immune response. Redox Rep. 2013, 18, 88–94. [Google Scholar] [CrossRef] [PubMed]
- Krausz, A.; Friedman, A.J. Nitric oxide as a surgical adjuvant. Future Sci. 2015, 1. [Google Scholar] [CrossRef] [PubMed]
| Cohort Feature | Value |
|---|---|
| Number of Patients | 11 |
| Median patient age (years ± S.E.) | 54 ± 4 |
| Sex (m/f) | 9/2 |
| Wound type | 11 trauma wounds (1 thoracic, 3 upper extremity, 7 lower extremity) |
| Wound healing response | 11/11 |
| Median time to wound healing (days ± S.E.) | 21 ± 4 |
| Protein ID | Acronym | Protein Name | Protein Class |
|---|---|---|---|
| P04040 | CAT | Catalase | Peroxidase |
| Q06830 | Prdx1 | Peroxiredoxin 1 | Peroxidase |
| P32119 | Prdx2 | Peroxiredoxin 2 | Peroxidase |
| P30041 | Prdx6 | Peroxiredoxin 6 | Peroxidase |
| P05164-2 | MPO | Isoform H14 of Myeloperoxidase | Peroxidase |
| P00441 | SOD1 | Superoxide dismutase 1 | Oxidoreductase |
| P00403 | MT-CO2 | Cytochrome C oxidase 2 | Oxidoreductase |
| P16083 | NQO2 | Ribosyldihydronicotinamide dehydrogenase | Oxidoreductase |
| O95571 | ETHE1 | Persulfide dioxygenase | Oxidoreductase |
| P07195 | LDHB | l-lactate dehydrogenase β | Dehydrogenase |
| P40926 | MDH2 | Malate dehydrogenase 2 | Dehydrogenase |
| P30043 | BLVRB | Flavin reductase (NADPH) | Reductase |
| P20292 | ALOX5AP | Arachidonate 5-lipoxygenase activating protein | Transferase |
| Protein ID | Acronym | Protein Name |
|---|---|---|
| Calmodulin-signaling molecules | ||
| P26447 | S100A4 | Protein S100 A4 |
| P06703 | S100A6 | Protein S100 A6 |
| P05109 | S100A8 | Protein S100 A8 |
| P06702 | S100A9 | Protein S100 A9 |
| P31949 | S100A11 | Protein S100 A11 |
| P25815 | S100P | Protein S100 P |
| Chaperone and heat shock proteins | ||
| P31946 | YWHAB | 14-3-3 protein β/α |
| P61981 | YWHAG | 14-3-3 protein γ |
| P27348 | YWHAH | 14-3-3 protein τ |
| Q04917 | YWHAQ | 14-3-3 protein ε |
| P63104 | YWHAZ | 14-3-3 protein ζ |
| P27797 | CALR | calreticulin |
| P14625 | HSP90B1 | Endoplasmin |
| P0DMV9 | HSPA1B | Heat shock 70 kDa protein 1B |
| P11142 | HSPA8 | Heat shock cognate 71 kDa protein |
| P11142-2 | HSPA8 | Isoform 2 of Heat shock cognate 71 kDa protein |
| P0DMV8-2 | HSPA1A | Isoform 2 of Heat shock 70 kDa protein 1A |
| Immune modulators | ||
| P01023 | A2M | α2-macroglobulin |
| P43490 | NAMPT | Nicotinamide phosphoribosyltransferase |
| P52565 | ARHGD | Rho GDP-dissociation inhibitor 1 |
| P52566 | ARHGD | Rho GDP-dissociation inhibitor 2 |
| P04083 | ANXA1 | Annexin A1 |
| P07355 | ANXA2 | Annexin A2 |
| P12429 | ANXA3 | Annexin A3 |
| P09525 | ANXA4 | Annexin A4 |
| P09525-2 | ANXA4 | Annexin A4 Isoform 2 |
| P08758 | ANXA5 | Annexin A5 |
| P50995 | ANXA11 | Annexin A11 |
| P50995-2 | ANXA11 | Annexin A11 Isoform 2 |
| P20292 | ALOX5AP | Arachidonate 5-lipoxygenase activating protein |
| Protein ID | Acronym | Protein Name |
|---|---|---|
| P20160 | AZU1 | Azurocidin |
| P02788 | LTF | Lactotransferrin |
| P02788-2 | LTF | Isoform δLf of Lactotransferrin |
| P17213 | BPI | Bactericidal permeability-increasing protein |
| P08311 | CTSG | Cathepsin G |
| Q9UBR2 | CTSZ | Cathepsin Z |
| P00747 | PLG | Plasminogen |
| P61626 | LYZ | Lysozyme |
| P08246 | ELANE | Neutrophil elastase |
| P30740 | SERPINB1 | Leukocyte elastase inhibitor |
| P30740-2 | SERPINB1-2 | Leukocyte elastase inhibitor isoform 2 |
| P80188 | LCN2 | Neutrophil gelatinase-associated lipocalin |
| P80188-2 | LCN2 | Isoform 2 of Neutrophil gelatinase-associated lipocalin |
| Q15582 | TGFB1 | Transforming growth factor B1 |
| Protein ID | Acronym | Protein Name |
|---|---|---|
| P02671 | FGA | Fibrinogen α chain |
| P02675 | FGB | Fibrinogen β chain |
| P02679 | FGG | Fibrinogen γ chain |
| P02671-2 | FGA2 | Isoform 2 of Fibrinogen α chain |
| P02679-2 | FGGA | Isoform γ A of Fibrinogen γ chain |
| P02452 | COL1A1 | Collagen α-1(I) chain |
| P08123 | COL1A2 | Collagen α-2(I) chain |
| P20908 | COL5A1 | Collagen α-1(V) chain |
| P12109 | COL6A1 | Collagen α-1(VI) chain |
| O00602 | FCN-1 | Ficolin 1 |
| P51884 | LUM | Lumican |
| P04004 | VTN | Vitronectin |
| Q6UX06 | OLFM4 | Olfactomedin-4 |
| P49747-2 | COMP2 | Cartilage oligomeric matrix protein 2 |
| P17931 | LGALS3 | Galectin 3 |
| P50552 | VASP | Vasodilator-stimulated phosphoprotein |
| Protein ID | Acronym | Protein Name |
|---|---|---|
| P13798 | ACPH | Acylamino-acid-releasing enzyme |
| P15144 | AMPN | Aminopeptidase N |
| P15169 | CBPN | Carboxypeptidase N catalytic chain |
| P42574 | CASP3 | Caspase-3 |
| P08311 | CATG | Cathepsin G |
| Q9UBR2 | CATZ | Cathepsin Z |
| Q9NZP8 | C1RL | Complement C1r subcomponent-like protein |
| P09871 | C1S | Complement C1s subcomponent |
| P00751 | CFAB | Complement factor B |
| P05156 | CFAI | Complement factor I |
| P02788 | TRFL | Lactotransferrin |
| P14780 | MMP9 | Matrix metalloproteinase-9 |
| P24158 | PRTN3 | Myeloblastin |
| P08246 | ELNE | Neutrophil elastase |
| P00747 | PLMN | Plasminogen |
| P25789 | PSA4 | Proteasome subunit α type-4 |
| P28066 | PSA5 | Proteasome subunit α type-5 |
| P20618 | PSB1 | Proteasome subunit β type-1 |
| P40306 | PSB10 | Proteasome subunit β type-10 |
| P28070 | PSB4 | Proteasome subunit β type-4 |
| P28072 | PSB6 | Proteasome subunit β type-6 |
| Q99436 | PSB7 | Proteasome subunit β type-7 |
| P00734 | THRB | Prothrombin |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bekeschus, S.; Lackmann, J.-W.; Gümbel, D.; Napp, M.; Schmidt, A.; Wende, K. A Neutrophil Proteomic Signature in Surgical Trauma Wounds. Int. J. Mol. Sci. 2018, 19, 761. https://doi.org/10.3390/ijms19030761
Bekeschus S, Lackmann J-W, Gümbel D, Napp M, Schmidt A, Wende K. A Neutrophil Proteomic Signature in Surgical Trauma Wounds. International Journal of Molecular Sciences. 2018; 19(3):761. https://doi.org/10.3390/ijms19030761
Chicago/Turabian StyleBekeschus, Sander, Jan-Wilm Lackmann, Denis Gümbel, Matthias Napp, Anke Schmidt, and Kristian Wende. 2018. "A Neutrophil Proteomic Signature in Surgical Trauma Wounds" International Journal of Molecular Sciences 19, no. 3: 761. https://doi.org/10.3390/ijms19030761
APA StyleBekeschus, S., Lackmann, J.-W., Gümbel, D., Napp, M., Schmidt, A., & Wende, K. (2018). A Neutrophil Proteomic Signature in Surgical Trauma Wounds. International Journal of Molecular Sciences, 19(3), 761. https://doi.org/10.3390/ijms19030761







