Abstract
Graphene derivatives have immense potential in stem cell research. Here, we report a three-dimensional graphene/arginine-glycine-aspartic acid (RGD) peptide nanoisland composite effective in guiding the osteogenesis of human adipose-derived mesenchymal stem cells (ADSCs). Amine-modified silica nanoparticles (SiNPs) were uniformly coated onto an indium tin oxide electrode (ITO), followed by graphene oxide (GO) encapsulation and electrochemical deposition of gold nanoparticles. A RGD–MAP–C peptide, with a triple-branched repeating RGD sequence and a terminal cysteine, was self-assembled onto the gold nanoparticles, generating the final three-dimensional graphene–RGD peptide nanoisland composite. We generated substrates with various gold nanoparticle–RGD peptide cluster densities, and found that the platform with the maximal number of clusters was most suitable for ADSC adhesion and spreading. Remarkably, the same platform was also highly efficient at guiding ADSC osteogenesis compared with other substrates, based on gene expression (alkaline phosphatase (ALP), runt-related transcription factor 2), enzyme activity (ALP), and calcium deposition. ADSCs induced to differentiate into osteoblasts showed higher calcium accumulations after 14–21 days than when grown on typical GO-SiNP complexes, suggesting that the platform can accelerate ADSC osteoblastic differentiation. The results demonstrate that a three-dimensional graphene–RGD peptide nanoisland composite can efficiently derive osteoblasts from mesenchymal stem cells.
1. Introduction
Stem cells have emerged as a promising source in the field of regenerative medicine, because of their innate ability to produce a variety of desired cell types in the human body [1,2]. Among stem cells, mesenchymal stem cells (MSCs) have predominated in clinical trials and commercialization, since they are relatively easy to obtain from various sources (e.g., adipose tissue, bone marrow, dental tissue), free from ethical issues, and most importantly, have a low risk of teratoma formation after transplantation [3]. MSCs are also attractive because of their capability to generate various types of cells (e.g., osteoblasts, chondrocytes, adipocytes), all of which are fundamental for the reconstruction and regeneration of most body components [4,5,6]. However, to maximize regeneration efficiency, the characteristics of MSCs should be properly adjusted to the desired cell type through differentiation, because their multipotency may also result in the production of unwanted cell types [7,8,9].
The differentiation of stem cells is conventionally controlled by soluble cues; that is, the use of defined culture media containing multiple growth factors (e.g., bone morphogenetic proteins, insulin, peroxisome proliferator-activated receptor γ) and biomolecules (e.g., dexamethasone, ascorbic acid, glycerophosphate) [10,11,12,13,14]. However, recent findings suggest that biophysical cues, such as the physicochemical characteristics of the underlying substrate, to which the cells attach and actively interact with as they grow, can also play key roles in steering the transformation of MSCs into specific cell types [15,16,17,18]. Mechanical stimulation, a biophysical cue that can affect cell behavior, is generally initiated by cell–substrate interactions, which alter cytoskeletal dynamics and consequently affect gene expression in the cells [12,15,19,20,21,22,23,24,25]. To provide favorable biophysical stimulation to stem cells, nanostructures, micropatterns, and functional composite materials have been utilized to control stem cell behavior, particularly differentiation, with and without the use of soluble factors [26,27,28,29,30,31]. Among such materials, graphene and graphene oxide (GO) have recently been shown to effectively provide the necessary physical stimulation for stem cell differentiation [32,33,34,35,36]. The distinct physicochemical characteristics of graphene derivatives were found to successfully initiate several specific MSC differentiation routes, such as adipogenesis [37], osteogenesis [38,39], and neurogenesis [40]. The multifarious surface chemistry, controllable amphiphilicity, and unique honeycomb carbon structure were reported to affect several cellular aspects such as spreading, growth, and morphology, as well as the absorption kinetics of proteins and chemicals in the differentiation medium [32,33,41,42,43,44].
In addition to graphene derivatives, natural polymers such as extracellular matrix (ECM) proteins (e.g., fibronectin, laminin, and vitronectin) and peptides are also excellent candidates for controlling stem cell behavior [45,46]. Among the various available materials, peptides containing a repeating arginine–glycine–aspartic acid (RGD) sequence, which is often found in ECM proteins, have been widely applied not only to increase the uptake of materials and molecules of interest into specific types of cells, but also to enhance cell adhesion to artificial platforms such as synthetic polymers and metallic surfaces [47,48,49]. Interestingly, RGD-containing peptides immobilized onto substrates enhanced the adhesion strength of MSCs, ultimately resulting in increased osteogenic differentiation, as cell adhesion is a key factor guiding osteogenic lineage commitment [50,51].
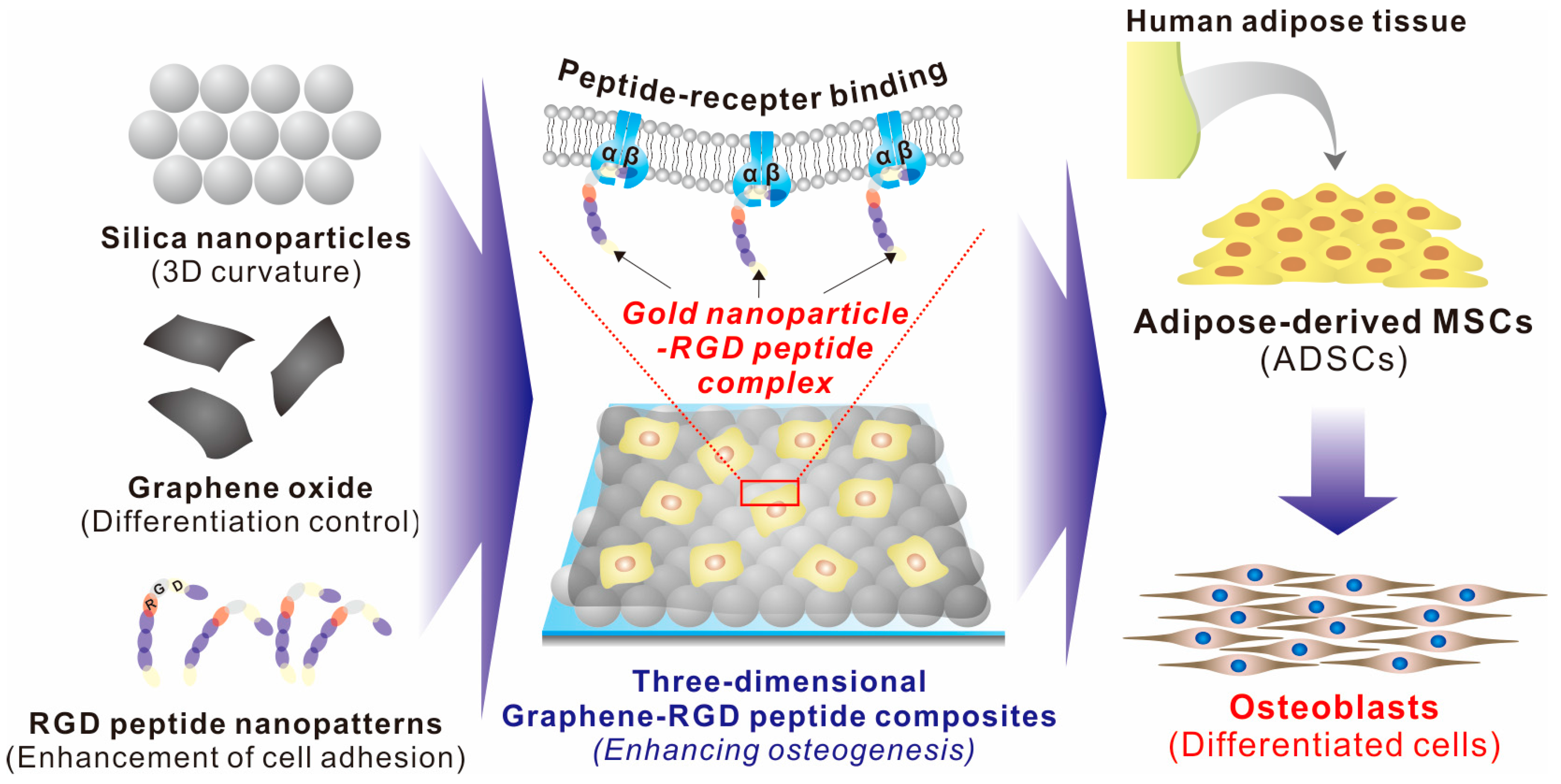
In this study, we report a three-dimensional graphene/RGD peptide nanoisland composite that is highly effective in controlling the osteogenic differentiation of human-adipose-derived mesenchymal stem cells (ADSCs; Figure 1). ADSC differentiation was controlled by four independent factors: (i) GOs that contribute to the absorption of several differentiation factors; (ii) RGD–MAP–C peptides that enhance cell adhesion; (iii) nanopatterns with randomly-distributed geometry and iv) height variations of underlying substrates that facilitate cell–GO and cell–peptide interactions (Figure 1). The physical and chemical properties of the three-dimensional nanocomposites were first characterized by X-ray photoelectron spectroscopy (XPS) and Raman spectroscopy. Thereafter, ADSCs were cultivated on GO surfaces with varying numbers of nanoparticle–RGD peptides and their adhesion, spreading, and proliferation were analyzed. Furthermore, the effects of nanoparticle–RGD peptides complexes on ADSC osteogenesis were characterized by assaying mRNA expression and calcification levels. Finally, to better understand the effects of nanoparticle–RGD peptides on the osteogenic differentiation of ADSCs, a four-week time course was performed in the presence of culture medium with and without differentiation factors.
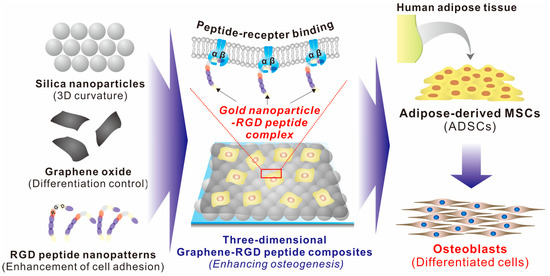
Figure 1.
Schematic diagram of three-dimensional graphene/arginine-glycine-aspartic acid (RGD) peptide composites for enhancement of adipose-derived mesenchymal stem cell (ADSC) osteogenesis.
2. Results
2.1. Fabrication of Silica-Nanoparticle–Graphene-Oxide Composites on an Indium Tin Oxide (ITO) Substrate
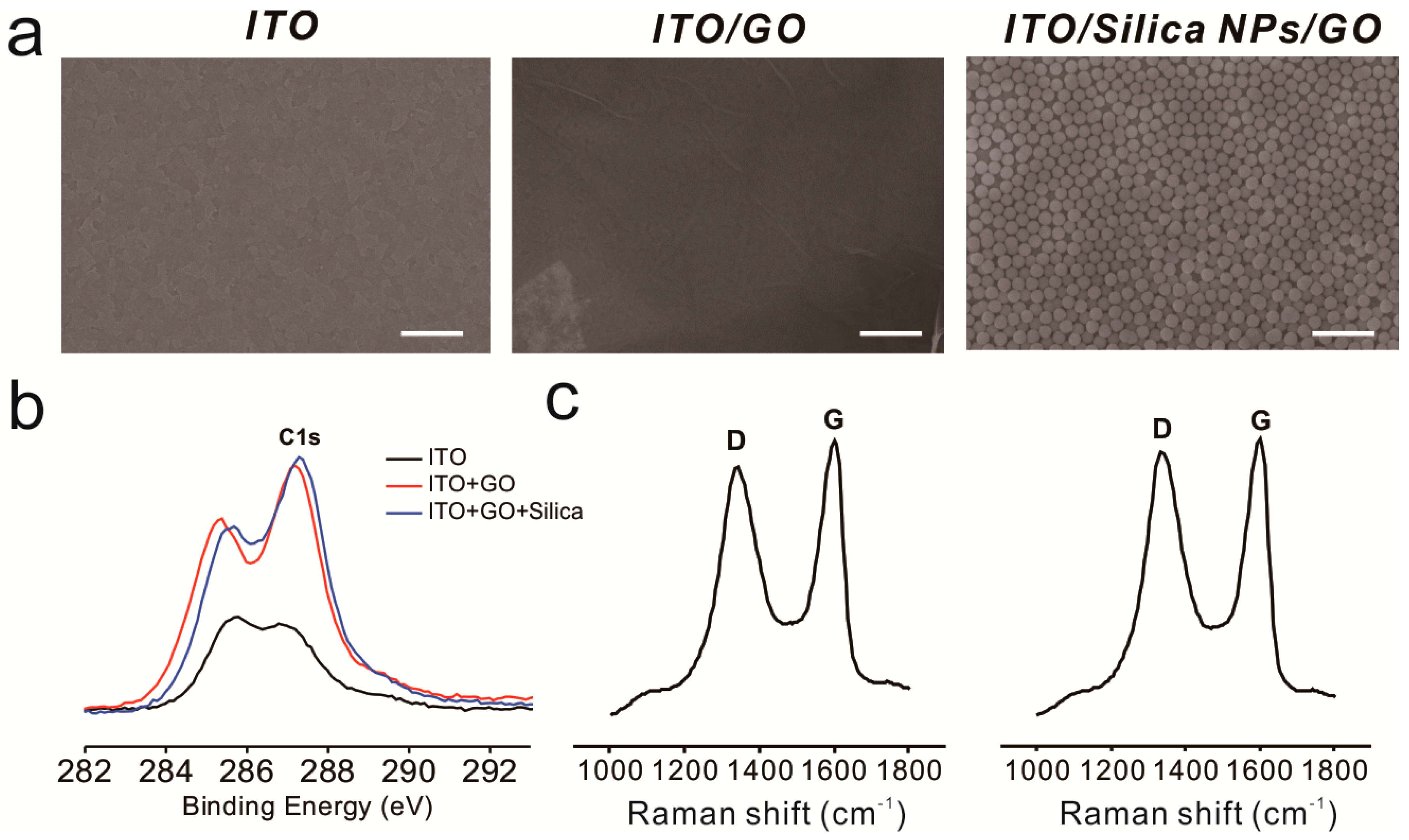
To enhance cell–substrate interactions, we hypothesized that the generation of RGD nanopattern ripples on the graphene surface would facilitate both αvβ3 ligand–RGD peptide and GO–cell membrane interactions, which ultimately affect stem cell behavior. There are several methods that could be utilized to fabricate three-dimensional environments on a GO surface. Photolithography is a representative tool suitable for the fabrication of various nano- and micro-structures with precisely controlled geometry [52,53]. However, this technique includes laborious and time-consuming steps such as photomask fabrication, photoresist coating, cleaning, and etching/metal deposition. Moreover, residual photoresist on the substrates could result in unexpected toxic effects on stem cells. Hence, silica nanoparticles (SiNPs), a material proven to be nontoxic and safe for use with most cell lines [54,55], were used as a coating material to generate height variations on the ITO substrate prior to GO modification. We then attached amine-functionalized SiNPs to the negatively charged GO via electrostatic interactions. As shown in Figure 2a, GO-coated SiNPs were uniformly coated onto the ITO substrate, and the ITO/GO substrate showed the typical wrinkle-like structure of graphene, different from the bare ITO substrate. In addition, XPS spectra revealed that the chemical structure of GO sheets on SiNPs were similar to those on the ITO substrate, showing C–C, C–O and C=O bonding, which are all typically found on the GO surface (Figure 2b and Figure S1). Raman spectroscopy was used to investigate GO oxidation. It has been reported that the G band represents the in-plane structure (sp2-bonded carbon) while the D band indicates structural defects, such as oxidized carbon. Based on this, the structural defects and carbon-to-oxygen ratio were analyzed. Figure 2c and Figure S2 show clear D (1350 cm−1) and G (1600 cm−1) peaks for ITO/SiNP/GOs, demonstrating the successful absorption of GO sheets onto amine-modified SiNP surfaces. The ID/IG values of ITO/GOs and ITO/SiNP/GOs were calculated to be 0.96 and 0.97, indicating that both GOs were highly oxidized, with C–C and C–O/C=O/O=C–O ratios of approximately 1:1. GO sheets embedded on the surface of SiNPs not only act as a conducting layer to enable electrochemical deposition of gold nanoparticles (GNPs) onto the SiNPs, but also helped attract gold chloride ions, resulting in the formation of the seed for GNP synthesis. Hence, it can be concluded that the fabricated GO-SiNP composite is a suitable platform for the generation of GNPs via electrochemical deposition, which could be further utilized to control stem cell behavior.
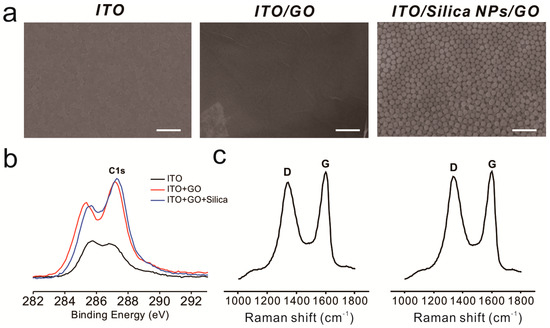
Figure 2.
Characterization of the basic substrates. (a) Scanning electron microscopy images of indium tin oxide (ITO) (left), ITO/graphene oxide (GO) (middle), and ITO/GO/silica nanoparticles (SiNPs) (right). Scale bars = 1 μm; (b) X-ray photoelectron spectroscopy (XPS) results for each substrate; (c) Raman spectroscopy data for ITO/GO (left) and ITO/GO/SiNPs (right).
2.2. Fabrication of a 3D Graphene–RGD Peptide Nanoisland Composite for Stem Cell Cultures
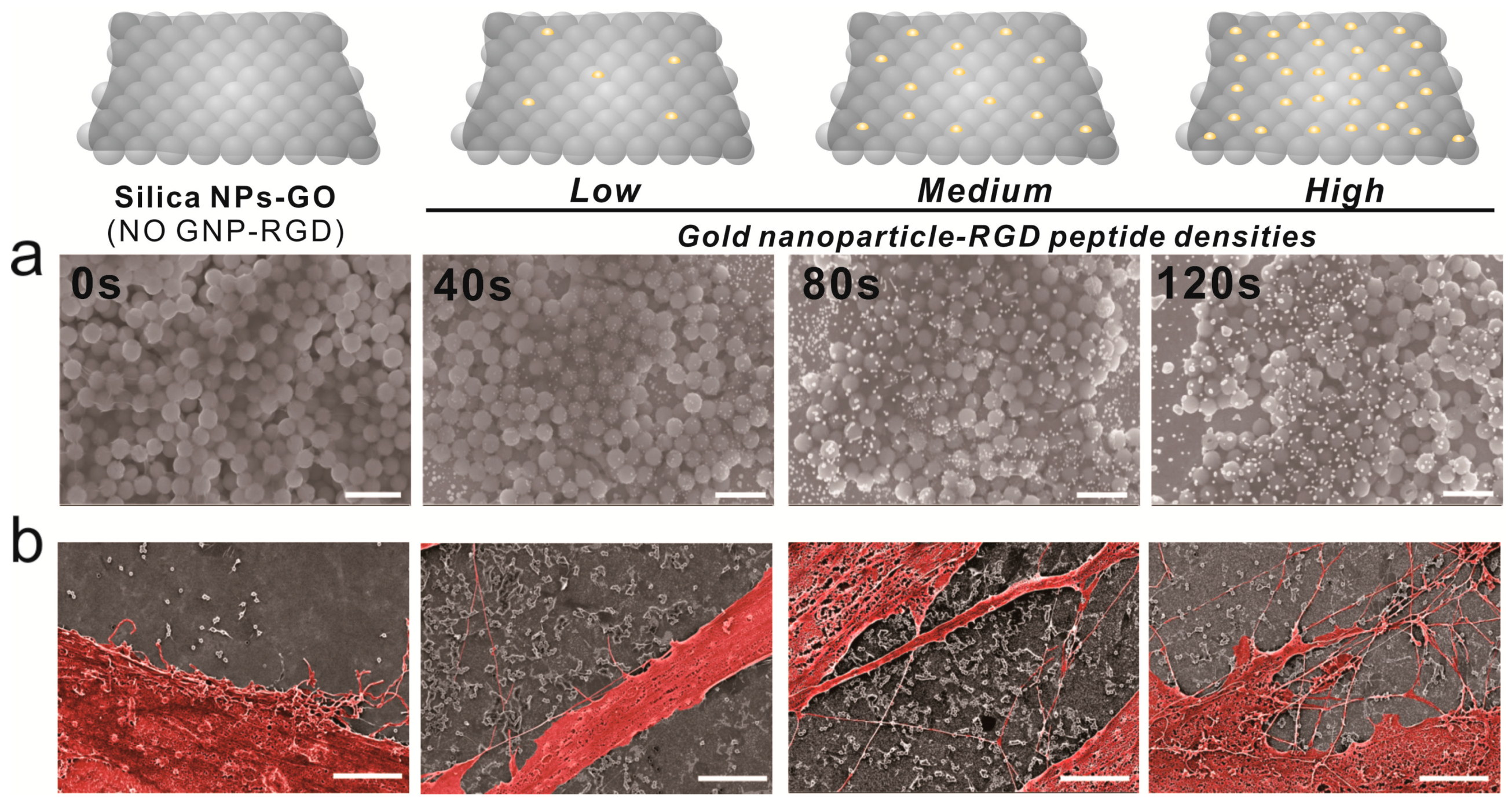
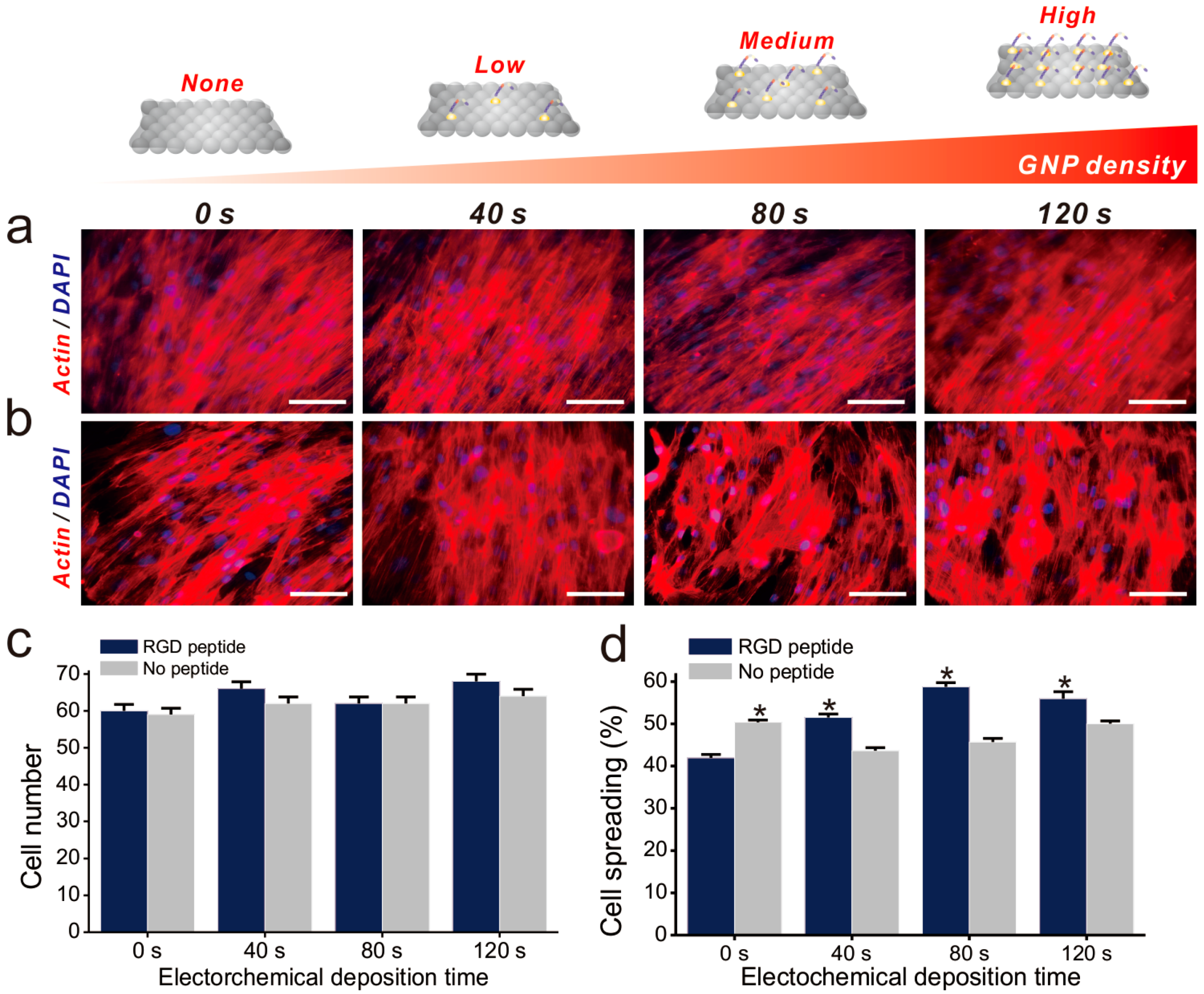
Cell adhesion is controlled by transmembrane glycoproteins, including integrins, selectins, and cadherins [56,57,58]. Integrins are core materials that actively regulate the anchoring of the cell membrane to the extracellular environment, and are hugely affected by the physicochemical properties of the underlying substrates [59,60]. A number of studies have reported that nanotopographic features influence many key cellular activities, including cell migration, proliferation, differentiation, and even apoptosis [61,62,63,64]. Specifically, since the size of the integrin subunits that are located in the outside of the cell membrane (α and β chains) is around 23nm and the final N-termini of each chain is around 5nm, the materials whose sizes are in the nanometer range could result in the changes of integrin-mediated cell anchoring [62,65]. Consequently, this causes cytoskeletal rearrangements of cells adhered to the nanomaterial surfaces, resulting in changes of cell signaling pathways, which ultimately affect cell growth, spreading and differentiation. In the case of MSCs, nanopattern geometry and arrangement are critical for both multipotency maintenance and differentiation. Dalby et al. reported that nanopatterns with random arrangements can effectively guide MSC osteogenesis, while the same nanopatterns with homogeneous arrangements maintain their multipotency in long-term cultures [65]. Based on this, we generated GNPs on GO-coated SiNPs via electrochemical deposition, as shown in Figure 3a. Due to the presence of the conducting GO layer, GNPs of 10–20 nm in diameter were successfully generated on the non-conducting SiNPs. Interestingly, the ID/IG values of Raman spectra were calculated to be 1.01, 1.04, 1.18, and 1.2 after 0, 40, 80, and 120 s of deposition, respectively, indicating that GO oxidation increased with increased electrochemical deposition (Figure S2). As the RGD-containing peptides (RGD–MAP–C) contain a terminal cysteine residue, they can be easily self-assembled on the surface of the GNPs. In addition, by varying the electrochemical deposition time, the size and density of GNPs could be varied, resulting in different RGD–MAP–C peptide arrangements, which would ultimately alter the cytoskeletal dynamics of stem cells. To investigate this, ADSCs were seeded onto each platform at different GNP densities, and therefore with different RGD–MAP–C peptide arrangements. As shown in Figure 3b, the GO-SiNP-GNP-RGD peptide composites with the highest GNP density (deposition time = 120 s) showed more active filopodia interaction with the underlying platform, indicating that this substrate might effectively enhance cell adhesion and migration, improving ADSC osteogenesis. To further study the effects of GNP-RGD peptide composites on ADSCs, the substrates were divided into two groups: (i) GO-SiNP-GNPs and (ii) GO-SiNP-GNP-RGD peptide composites. After two days of culture, the cytoskeletal structures and nuclei of ADSCs were visualized by phalloidin and Hoechst 33342 staining to calculate the rates of cell spreading and growth by fluorescence microscopy (Figure 4a,b and Figure S3). Cell growth was not affected by the presence of RGD–MAP–C peptides (Figure 4c), while cell spreading was highly enhanced on all the substrates possessing GNP-RGD peptide composites (Figure 4d). Cell spreading was 118, 128, and 110% higher on GNP-RGD composites compared to GNPs only, after 40, 80, and 120 s of deposition, respectively. Cell spreading seemed to increase with increased GNP density; however, owing to the limitations in the area in which cells could attach and grow, the enhancement of cell spreading was saturated with medium GNP density. Interestingly, cell spreading was not enhanced by RGD–MAP–C peptide modifications to the GO-SiNP platform, because of the absence of GNPs, which mediate RGD–MAP–C peptide immobilization via an Au–S bond. Accordingly, based on the cell adhesion, spreading, and proliferation results, we conclude that GNP-RGD peptide modifications to the GO-SiNP platform positively affect ADSC adhesion, and can thus be considered a candidate platform to control stem cell differentiation.
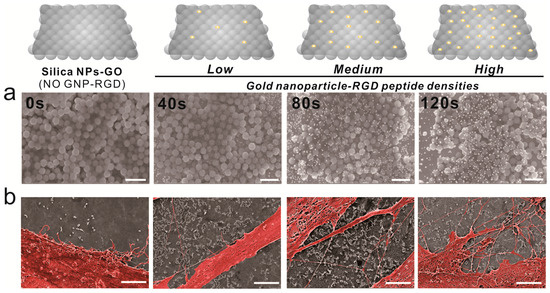
Figure 3.
Scanning electron microscopy (SEM) images of ADSCs on gold-deposited substrates. (a) From left to right, SEM images of gold deposition. Scale bars = 1 μm; (b) Pseudocolor SEM images of ADSCs showing cell spreading. Scale bars = 3 μm.
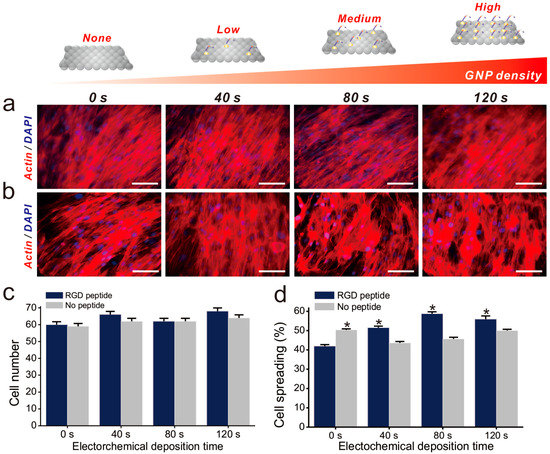
Figure 4.
Fluorescence images of all substrates Fluorescence images of each substrate with (a) and without (b) RGD peptide. Scale bars = 100 μm; (c) Number of cells on each substrate after 2 day of incubation; (d) Spreading of proliferated cells. * Student’s t-test, p < 0.05, n = 3.
2.3. Guiding ADSC Osteogenesis Using Graphene–RGD Peptide Nanoisland Composites
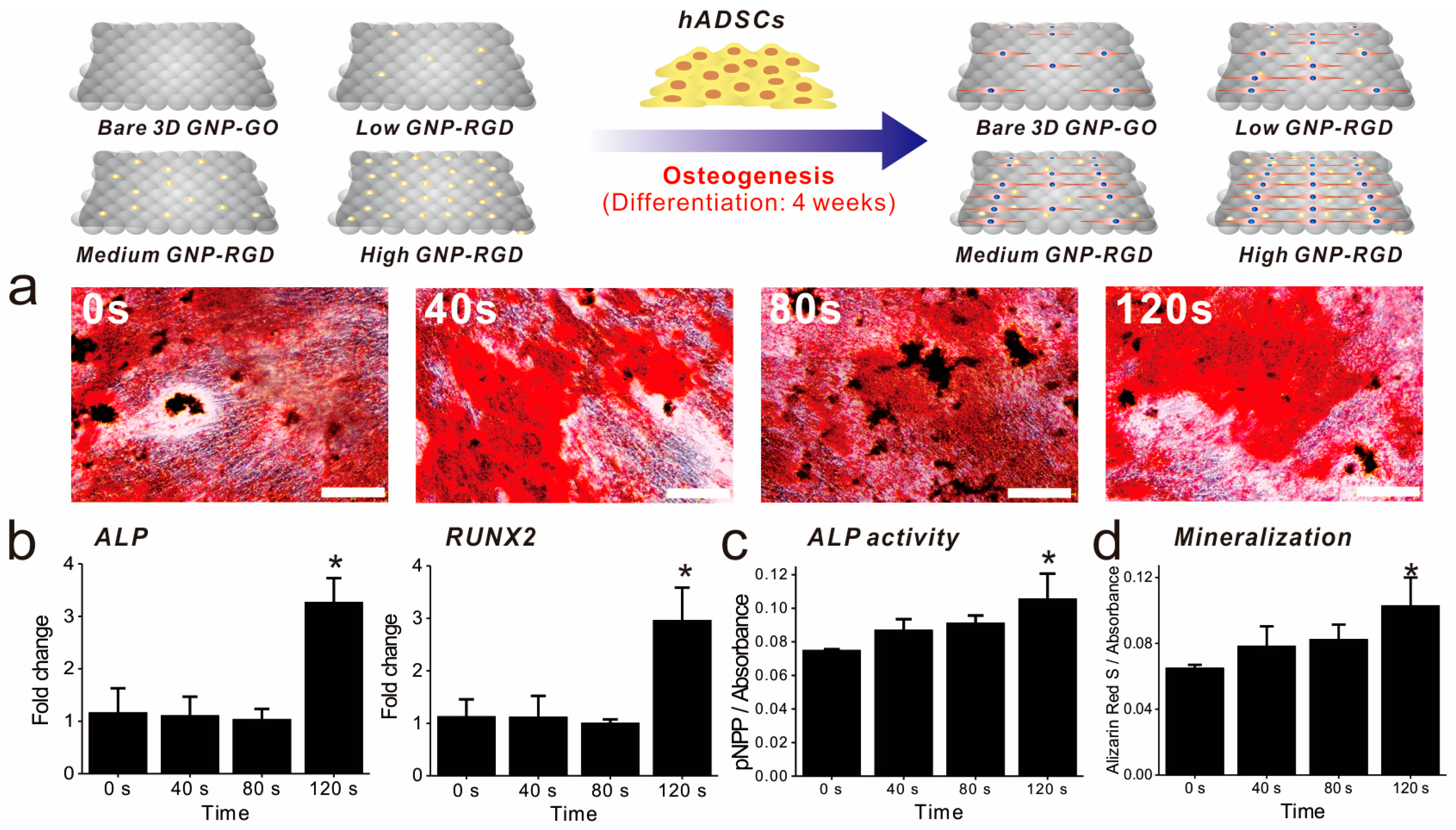
As cell adhesion and spreading were highly enhanced by GNP-RGD peptide modifications, we next investigated whether these differences in cell behavior affected ADSC osteogenesis. Differentiation was performed in medium containing well-known osteogenic differentiation factors (β-glycerophosphate, ascorbic acid, and dexamethasone; Figure 5). After four weeks of differentiation, ADSC osteogenesis levels were analyzed using several markers, including alkaline phosphatase (ALP) enzyme activity, ALP and runt-related transcription factor 2 (RUNX2) expression, and osteogenesis mineralization. ALP regulates the dephosphorylation of several biomolecules and is an indicator of pre-osteogenesis stem cells, while RUNX2 is critical for osteoblastic differentiation. Based on reverse transcription-quantitative polymerase chain reaction (RT-qPCR) results, remarkably, the expression of both genes was highly enhanced on GO-SiNPs with high levels of GNP-RGD peptides compared with bare 3D GO-SiNPs and the same substrate with low and medium GNP-RGD peptide densities (263% and 295% higher than low density of gold deposition for ALP and RUNX2, respectively (Figure 5b). Next, to confirm the superiority of 3D GO-RGD peptide nanoisland composites with high GNP density in ADSC osteogenic differentiation, the ALP enzyme activity and calcification levels were evaluated, using para-nitrophenylphosphate and Alizarin Red S (ARS) as colorimetric reagents, respectively. MSCs accumulation of calcium phosphate (hydroxyapatite mineral (Ca10(PO4)6)), an essential material for building bone structure, is an indicator of osteogenesis. As shown in Figure 5a, all ADSCs cultured in osteogenic medium successfully differentiated into cells displaying calcium deposits. Three-dimensional GO-RGD peptide nanoisland composites with high GNP density showed the best osteogenic differentiation efficiency based on ALP and ARS levels, which were 148% and 158% higher than with bare GO-SiNP platforms (Figure 5c,d). This is consistent with a previous study reporting that modifications to ECM-derived RGD-glycoproteins (e.g., fibronectin, vitronectin, and osteopontin) on cell culture substrates are critical for MSC osteogenic differentiation. Hence, it is highly likely that the increase in RGD–MAP–C peptide density in combination with the three-dimensional GO sheets on the SiNPs synergistically enhance ADSC osteogenesis via increased cell adhesion and absorption of differentiation factors. Based on these observations, we can logically conclude that the developed graphene–RGD peptide nanoislands are a promising platform to guide the differentiation of stem cells into specific lineages.
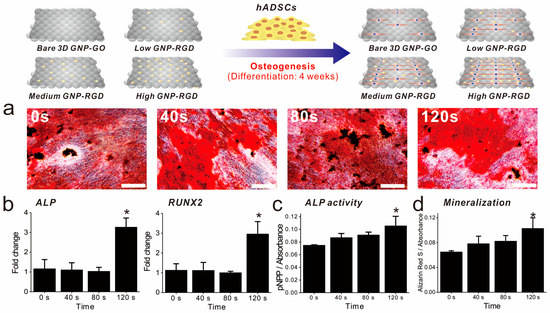
Figure 5.
Confirmation of ADSC osteogenic differentiation. (a) Alizarin Red S staining of all substrates. Scale bars = 200 μm; (b) RT-qPCR data for alkaline phosphatase (ALP), and RUNX2; (c) The ALP activity of each substrate; (d) Absorbance rates after Alizarin Red S (ARS) staining. * Student’s t-test, p < 0.05, n = 3.
2.4. Time Course of ADSC Osteogenic Differentiation on Graphene–RGD Peptide Nanoislands
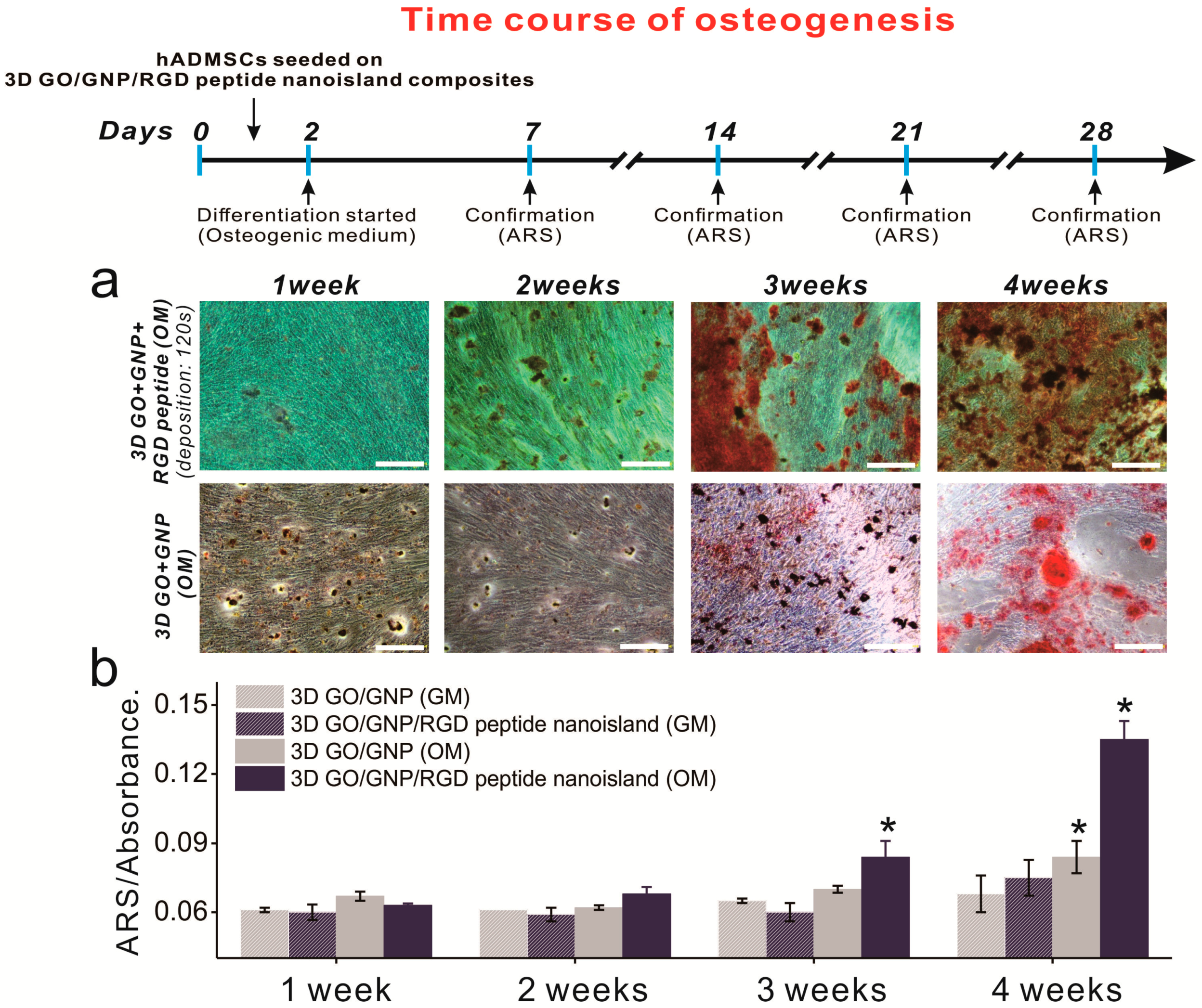
After confirming that the graphene–RGD peptide nanoislands with high GNP density are highly effective in guiding ADSC osteogenesis, we next investigated whether the platform could accelerate ADSC osteogenesis. This is important to study, because accelerated differentiation is needed to supply osteoblasts to the patients requiring urgent orthopedic surgery. In fact, it takes up to four weeks to generate bone cells from stem cells, and this is an obstacle in the clinical use of stem-cell-derived osteoblasts. Osteogenic ADSC differentiation was induced using typical osteogenic medium, and ARS staining was performed weekly to evaluate the osteoblastic differentiation of ADSCs grown on GO-SiNP/GNPs with and without RGD–MAP–C peptides. For the first 14 days, there was no discernable increase in osteoblast mineralization (Figure 6a). However, after two weeks of differentiation, the difference was apparent, especially between days 14 and 21 (Figure 6 and Figure S4). Specifically, the conversion of MSCs into osteoblasts on the GO-SiNP/GNP/RGD–MAP–C substrate was 120% and 160% higher than on the same substrate without GNP-RGD–MAP–C composites at days 21 and 28, respectively. Interestingly, the osteoblastic mineralization of ADSCs was highly enhanced at day 21 on the GO-SiNP/GNP/RGD–MAP–C substrate compared to on typical GO-SiNP substrates, indicating that the platform might be useful to more rapidly generate osteoblasts from stem cells, which could help meet timely clinical demands (Figure 6b). More studies regarding the gene expression levels of adhesion-related proteins (e.g., integrins, vinculin, focal adhesion kinase, paxillin, and talin) and osteogenic differentiation (e.g., osteocalcin, osteonectin, osteopontin) should be performed to validate the aforementioned results.
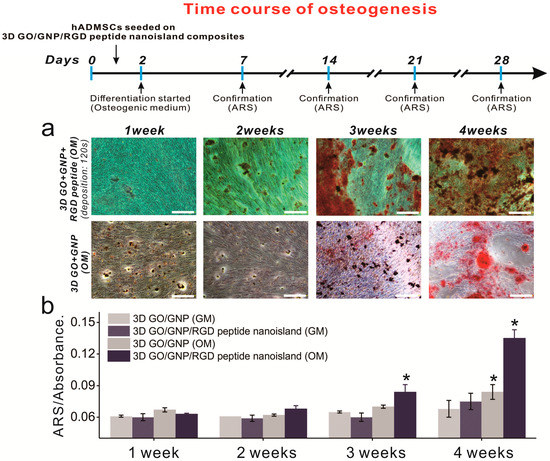
Figure 6.
Time course of osteogenesis with and without RGD–MAP–C peptides. (a) ADSC differentiation was monitored every 7 d. Scale bars = 200 μm; (b) ARS absorbance changes in the presence of gold and different media. * Student’s t-test, p < 0.05, n = 3.
3. Discussion
In this study, we report a new platform that enables the stable long-term culture of ADSCs and guides their osteogenic differentiation. Among the various substrates fabricated, the three-dimensional GO-SiNP platform with high GNP-RGD peptide density, which we have termed the graphene–RGD peptide nanoisland composite, was best at enhancing stem cell adhesion and spreading on the artificial surface. Furthermore, this substrate was superior to other platforms, including bare GO-SiNPs and GO-SiNPs with low and medium densities of GNP-RGD peptides, at guiding ADSC osteogenesis. Specifically, the RNA expression levels of ALP (a pre-osteogenic marker) and RUNX2 (an osteogenic marker) in ADSCs grown on graphene–RGD peptide nanoisland composite were 2.63- and 2.95-fold higher than on bare 3D GO-SiNPs after four weeks of differentiation. Other osteogenic differentiation indicators, such as ALP enzyme activity and calcium accumulation, were also highly enhanced, 1.48- and 1.58-fold higher than with bare 3D GO-SiNPs, proving the excellence of 3D graphene–RGD peptide nanoisland composite for steering ADSC osteogenesis. Moreover, ADSCs grown on the developed composites showed increased osteoblastic mineralization between days 14–21, 1.2-fold higher than on bare GO-SiNPs, exhibiting the potential of the developed platform to accelerate stem cell osteogenesis.
The 3D graphene–RGD peptide nanoisland composite shows great promise in stem cell research, and more advanced studies based on our findings will be required to improve the platform through the incorporation of different types of nanomaterials (e.g., polystyrene beads, glass beads) and peptides (cyclic RGD peptide, iRGD peptide). Other types of stem cells, such as pluripotent stem cells and neural stem cells, could also be applied to fully investigate the potential of 3D graphene–peptide nanoislands as a stable platform for long-term stem cell culture [66,67]. The desired cell types achieved from various stem cell lines using 3D graphene–peptide nanoisland composites as the culture platforms would then be usefully applied for clinical study, especially for the regeneration of damaged parts of human organs/tissues. Taken together, we can logically conclude that the developed platform, graphene–peptide complexes, could make remarkable contributions in the field of stem-cell-based regenerative medicine.
4. Materials and Methods
4.1. Materials
SiNPs were purchased from NanoComposix (San Diego, CA, USA). GO was obtained from Graphene Supermarket (Calverton, NY, USA). ADSCs and cell engineering for origin (CEFO) ADSC growth media supplemented with 10% fetal bovine serum and 1% penicillin/streptomycin were obtained from CEFO Bio (Seoul, Korea). Dexamethasone, L-ascorbic acid, β-glycerophosphate, and gold chloride trihydrate were purchased from Sigma Aldrich (St. Louis, MO, USA). RGD peptide was purchased from Peptron (Daejeon, Korea). ITO was purchased from UID (Sejong, Korea).
4.2. Fabrication of Three-Dimensional Hybrid Structures
First, ITO was prepared by sonication in 1% Triton X-100 in distilled water (DW) and pure ethanol for 20 min each. The ITO was spin-coated at 2000 rpm for 20 sec using 300 nm silica nanoparticles in 70% ethanol. After coating the substrates, 250 μg/mL GO solution was dropped onto the ITO and baked in a 70 °C oven for 1 h, after which the substrates were washed with DW. A chamber of 0.8 cm diameter was placed on the ITO substrates, using Polydimethylsilozane (PDMS) for cold deposition. A 0.5 mM gold solution was electrochemically treated at an initial voltage of −1.3 at 0, 40, 80, and 120 s. After electrical deposition, the chamber was washed with PBS, and coated with 0.05 mg/mL RGD peptide overnight at 4 °C.
4.3. Substrate Characterization
The silica-coated substrates were characterized by a variety of analytical techniques, including scanning electron microscopy (SEM, SIGMA, Carl Zeiss, Oberkochen, Germany), XPS (K-alpha+, Waltham, MA, USA; Thermo Fisher Scientific, Waltham, MA, USA.), and Raman spectroscopy.
4.4. Cell Proliferation and Spreading Analysis
ADSCs (6000 cells/cm2) were cultured on the 3D hybrid substrates in growth medium for 2 day. Before staining, cells were fixed in neutral buffered formalin (Sigma Aldrich) and permeabilized with 0.1% Triton X-100 for 5 min. After staining with Alexa Fluor 568 phalloidin (Life Technology, Carlsbad, CA, USA) and Hoechst 33342 (Thermo Scientific, Waltham, MA, U.S.A), cells were imaged by fluorescence microscopy. Cell proliferation and spreading were analyzed using ImageJ software.
4.5. Osteogenesis of ADSCs
The osteogenic differentiation medium contained 50 μM l-ascorbic acid, 10 mM β-glycerophosphate, and 100 nM dexamethasone. ADSCs were seeded in chambers, and the medium was changed every 2 days for 4 weeks. After the induction of osteogenesis, differentiation was confirmed by assaying calcium deposition with ARS. Cells were stained with ARS for 20 min, washed with DW, and imaged on an optical microscope. Subsequently, 10% cetylpyridinium chloride in 10 mM sodium phosphate was added, and the solution was incubated at 25 °C for 15 min. ARS extraction was measured at 562 nm on a microplate reader. In the case of alkaline phosphatase (ALP) analysis, ALP activity was measured based on the conversion of p-nitrophenyl phosphate (pNPP) into p-nitrophenol. First, the cell lysates achieved and were added in to 96 well plate with reaction reagent (pNPP). The plates were then incubated at 25 °C for 15 min while protecting the plates from light. Finally, the reaction was terminated with the addition of 3 M NaOH, followed by reading the absorbance at 405 nm using the plate reader.
4.6. Osteogenic Marker Expression Analysis
Total RNA was extracted from osteogenic cells using an RNeasy Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions, and cDNA was synthesized from 1 μg of total RNA with SuperScript II Reverse Transcriptase (Invitrogen, Carlsbad, CA, U.S.A). RT-qPCR was performed with SYBR Premix Ex Taq (Takara, Shiga, Japan) in a StepOnePlus RT-PCR system.
Supplementary Materials
Supplementary materials can be found at http://www.mdpi.com/1422-0067/19/3/669/s1.
Acknowledgments
This work was supported by the Nano Material Technology Development Program through the NRF of Korea funded by the Korea Government (MSIP) (NRF-2014M3A7B4051907) and by the Chung-Ang University Research Scholarship Grants in 2016.
Author Contributions
Tae-Hyung Kim and Junhong Min conceived and designed the experiments; Ee-Seul Kang, Da-Seul Kim, and Yoojoong Han performed the experiments; Tae-Hyung Kim, Ee-Seul Kang and Hyungbin Son analyzed the data; Tae-Hyung Kim, Hyungbin Son and Junhong Min contributed reagents/materials/analysis tools; Ee-Seul Kang, Tae-Hyung Kim, Yoojoong Han, Yong-Ho Chung, and Junhong Min wrote the paper.
Conflicts of Interest
The authors declare no conflict of interest. The founding sponsors had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, and in the decision to publish the results.
Abbreviations
| ADSC | Adipose-derived mesenchymal stem cells |
| ALP | Alkaline phosphatase |
| ARS | Alizarin red S |
| DW | Distilled water |
| ECM | Extracellular matrix |
| ITO | Indium tin oxide |
| GNP | Gold nanoparticle |
| GO | Graphene oxide |
| PDMS | Polydimethylsilozane |
| RGD | Arginine-glycine-aspartic acid |
| RT-qPCR | Reverse transcription quantitative polymerase chain reaction |
| RUNX2 | Runt-related transcription factor 2 |
| SEM | Scanning electron microscopy |
| SiNPs | Silica nanoparticles |
| XPS | X-ray photoelectron spectroscopy |
References
- Segers, V.F.; Lee, R.T. Stem-cell therapy for cardiac disease. Nature 2008, 451, 937–942. [Google Scholar] [CrossRef] [PubMed]
- Strauer, B.E.; Kornowski, R. Stem cell therapy in perspective. Circulation 2003, 107, 929–934. [Google Scholar] [CrossRef] [PubMed]
- Ankrum, J.; Karp, J.M. Mesenchymal stem cell therapy: Two steps forward, one step back. Trends Mol. Med. 2010, 16, 203–209. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.B.; Hui, J.H.; Song, I.C.; Ardany, L.; Lee, E.H. Injectable mesenchymal stem cell therapy for large cartilage defects—A porcine model. Stem Cells 2007, 25, 2964–2971. [Google Scholar] [CrossRef] [PubMed]
- Crisan, M.; Yap, S.; Casteilla, L.; Chen, C.-W.; Corselli, M.; Park, T.S.; Andriolo, G.; Sun, B.; Zheng, B.; Zhang, L.; et al. A perivascular origin for mesenchymal stem cells in multiple human organs. Cell Stem Cell 2008, 3, 301–313. [Google Scholar] [CrossRef] [PubMed]
- Campagnoli, C.; Roberts, I.A.; Kumar, S.; Bennett, P.R.; Bellantuono, I.; Fisk, N.M. Identification of mesenchymal stem/progenitor cells in human first-trimester fetal blood, liver, and bone marrow. Blood 2001, 98, 2396–2402. [Google Scholar] [CrossRef] [PubMed]
- Karnoub, A.E.; Dash, A.B.; Vo, A.P.; Sullivan, A.; Brooks, M.W.; Bell, G.W.; Richardson, A.L.; Polyak, K.; Tubo, R.; et al. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature 2007, 449, 557–563. [Google Scholar] [CrossRef] [PubMed]
- Curran, J.M.; Chen, R.; Hunt, J.A. The guidance of human mesenchymal stem cell differentiation in vitro by controlled modifications to the cell substrate. Biomaterials 2006, 27, 4783–4793. [Google Scholar] [CrossRef] [PubMed]
- Suhito, I.R.; Han, Y.; Min, J.; Son, H.; Kim, T.-H. In situ label-free monitoring of human adipose-derived mesenchymal stem cell differentiation into multiple lineages. Biomaterials 2018, 154, 223–233. [Google Scholar] [CrossRef] [PubMed]
- Majumdar, M.K.; Keane-Moore, M.; Buyaner, D.; Hardy, W.B.; Moorman, M.A.; McIntosh, K.R.; Mosca, J.D. Characterization and functionality of cell surface molecules on human mesenchymal stem cells. J. Biomed. Sci. 2003, 10, 228–241. [Google Scholar] [CrossRef] [PubMed]
- Caplan, A.I.; Bruder, S.P. Mesenchymal stem cells: Building blocks for molecular medicine in the 21st century. Trends Mol. Med. 2001, 7, 259–264. [Google Scholar] [CrossRef]
- Engler, A.J.; Sen, S.; Sweeney, H.L.; Discher, D.E. Matrix elasticity directs stem cell lineage specification. Cell 2006, 126, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Mackay, A.M.; Beck, S.C.; Murphy, J.M.; Barry, F.P.; Chichester, C.O.; Pittenger, M.F. Chondrogenic differentiation of cultured human mesenchymal stem cells from marrow. Tissue Eng. 1998, 4, 415–428. [Google Scholar] [CrossRef] [PubMed]
- Oh, M.; Kim, Y.J.; Son, Y.J.; Yoo, H.S.; Park, J.H. Promotive effects of human induced pluripotent stem cell-conditioned medium on the proliferation and migration of dermal fibroblasts. Biotechnol. Bioproc. Eng. 2017, 22, 561–568. [Google Scholar] [CrossRef]
- Shih, Y.R.V.; Tseng, K.F.; Lai, H.Y.; Lin, C.H.; Lee, O.K. Matrix stiffness regulation of integrin-mediated mechanotransduction during osteogenic differentiation of human mesenchymal stem cells. J. Bone Miner Res. 2011, 26, 730–738. [Google Scholar] [CrossRef] [PubMed]
- Kilian, K.A.; Bugarija, B.; Lahn, B.T.; Mrksich, M. Geometric cues for directing the differentiation of mesenchymal stem cells. Proc. Natl. Acad. Sci. USA 2010, 107, 4872–4877. [Google Scholar] [CrossRef] [PubMed]
- Knippenberg, M.; Helder, M.N.; Zandieh, D.B.; Semeins, C.M.; Wuisman, P.I.; Klein-Nulend, J. Adipose tissue-derived mesenchymal stem cells acquire bone cell-like responsiveness to fluid shear stress on osteogenic stimulation. Tissue Eng. 2005, 11, 1780–1788. [Google Scholar] [CrossRef] [PubMed]
- Shin, Y.M.; Park, J.-S.; Jeong, S.I.; An, S.-J.; Gwon, H.-J.; Lim, Y.-M.; Nho, Y.-C.; Kim, C.-Y. Promotion of human mesenchymal stem cell differentiation on bioresorbable polycaprolactone/biphasic calcium phosphate composite scaffolds for bone tissue engineering. Biotechnol. Bioproc. Eng. 2014, 19, 341–349. [Google Scholar] [CrossRef]
- Nava, M.M.; Raimondi, M.T.; Pietrabissa, R. Controlling self-renewal and differentiation of stem cells via mechanical cues. Biom. Res. Int. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Connelly, J.T.; Gautrot, J.E.; Trappmann, B.; Tan, D.W.-M.; Donati, G.; Huck, W.T.; Watt, F.M. Actin and serum response factor transduce physical cues from the microenvironment to regulate epidermal stem cell fate decisions. Nat. Cell Biol. 2010, 12, 711–718. [Google Scholar] [CrossRef] [PubMed]
- Kuo, S.-W.; Lin, H.-I.; Ho, J.H.-C.; Shih, Y.-R.V.; Chen, H.-F.; Yen, T.-J.; Lee, O.K. Regulation of the fate of human mesenchymal stem cells by mechanical and stereo-topographical cues provided by silicon nanowires. Biomaterials 2012, 33, 5013–5022. [Google Scholar] [CrossRef] [PubMed]
- Lutolf, M.P.; Gilbert, P.M.; Blau, H.M. Designing materials to direct stem-cell fate. Nature 2009, 462, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Chen, C.S.; Fu, J. Forcing stem cells to behave: A biophysical perspective of the cellular microenvironment. Ann. Rev. Biophys. 2012, 41, 519–542. [Google Scholar] [CrossRef] [PubMed]
- Xue, R.; Li, J.Y.S.; Yeh, Y.; Yang, L.; Chien, S. Effects of matrix elasticity and cell density on human mesenchymal stem cells differentiation. J. Orthop. Res. 2013, 31, 1360–1365. [Google Scholar] [CrossRef] [PubMed]
- Suhito, I.R.; Han, Y.; Kim, D.-S.; Son, H.; Kim, T.-H. Effects of two-dimensional materials on human mesenchymal stem cell behaviors. Biochem. Biophys. Res. Commun. 2017, 493, 578–584. [Google Scholar] [CrossRef] [PubMed]
- Ragetly, G.R.; Griffon, D.J.; Lee, H.-B.; Fredericks, L.P.; Gordon-Evans, W.; Chung, Y.S. Effect of chitosan scaffold microstructure on mesenchymal stem cell chondrogenesis. Acta Biomater. 2010, 6, 1430–1436. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Kim, P.; Helen, W.; Engler, A.J.; Levchenko, A.; Kim, D.-H. Control of stem cell fate and function by engineering physical microenvironments. Integr. Biol. 2012, 4, 1008–1018. [Google Scholar] [CrossRef]
- Huri, P.Y.; Cook, C.; Hutton, D.; Goh, B.; Gimble, J.; DiGirolamo, D.; Grayson, W.L. Biophysical cues enhance myogenesis of human adipose derived stem/stromal cells. Biochem. Biophys. Res. Commun. 2013, 438, 180–185. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Dai, J.; Zhang, X.A. Environmental physical cues determine the lineage specification of mesenchymal stem cells. Biochim. Biophys. Acta Gen. Subj. 2015, 1850, 1261–1266. [Google Scholar] [CrossRef] [PubMed]
- Keung, A.J.; Healy, K.E.; Kumar, S.; Schaffer, D.V. Biophysics and dynamics of natural and engineered stem cell microenvironments. Wiley Interdiscip. Rev. Syst. Biol. Med. 2010, 2, 49–64. [Google Scholar] [CrossRef] [PubMed]
- Chueng, S.-T.D.; Yang, L.; Zhang, Y.; Lee, K.-B. Multidimensional nanomaterials for the control of stem cell fate. Nano. Converg. 2016, 3, 23. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.C.; Lim, C.H.Y.; Shi, H.; Tang, L.A.; Wang, Y.; Lim, C.T.; Loh, K.P. Origin of enhanced stem cell growth and differentiation on graphene and graphene oxide. ACS Nano 2011, 5, 7334–7341. [Google Scholar] [CrossRef] [PubMed]
- Nayak, T.R.; Andersen, H.; Makam, V.S.; Khaw, C.; Bae, S.; Xu, X.; Ee, P.-L.R.; Ahn, J.-H.; Hong, B.H.; Pastorin, G.; et al. Graphene for controlled and accelerated osteogenic differentiation of human mesenchymal stem cells. ACS Nano 2011, 5, 4670–4678. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Park, J.; Sim, S.H.; Sung, M.G.; Kim, K.S.; Hong, B.H.; Hong, S. Enhanced differentiation of human neural stem cells into neurons on graphene. Adv. Mater. 2011, 23. [Google Scholar] [CrossRef] [PubMed]
- Kang, E.-S.; Kim, D.-S.; Suhito, I.R.; Choo, S.-S.; Kim, S.-J.; Song, I.; Kim, T.H. Guiding osteogenesis of mesenchymal stem cells using carbon-based nanomaterials. Nano Converg. 2017, 4, 2. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.-H.; Lee, T.; El-Said, W.A.; Choi, J.-W. Graphene-based materials for stem cell applications. Materials 2015, 8, 8674–8690. [Google Scholar] [CrossRef] [PubMed]
- Janderová, L.; McNeil, M.; Murrell, A.N.; Mynatt, R.L.; Smith, S.R. Human mesenchymal stem cells as an in vitro model for human adipogenesis. Obesity 2003, 11, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Luu, Y.K.; Capilla, E.; Rosen, C.J.; Gilsanz, V.; Pessin, J.E.; Judex, S.; Rubin, C.T. Mechanical stimulation of mesenchymal stem cell proliferation and differentiation promotes osteogenesis while preventing dietary-induced obesity. J. Bone. Miner Res. 2009, 24, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Bruder, S.P.; Fink, D.J.; Caplan, A.I. Mesenchymal stem cells in bone development, bone repair, and skeletal regenaration therapy. J. Cell Biochem. 1994, 56, 283–294. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Fraser, J.L.; Lu, Z.-Y.; Hu, X.; Yu, S.P. Transplantation of hypoxia preconditioned bone marrow mesenchymal stem cells enhances angiogenesis and neurogenesis after cerebral ischemia in rats. Neurobiol. Dis. 2012, 46, 635–645. [Google Scholar] [CrossRef] [PubMed]
- Mooney, E.; Dockery, P.; Greiser, U.; Murphy, M.; Barron, V. Carbon nanotubes and mesenchymal stem cells: Biocompatibility, proliferation and differentiation. Nano Lett. 2008, 8, 2137–2143. [Google Scholar] [CrossRef] [PubMed]
- Park, S.Y.; Namgung, S.; Kim, B.; Im, J.; Kim, J.; Sun, K.; Lee, K.B.; Nam, J.M.; Park, Y.; Hong, S. Carbon nanotube monolayer patterns for directed growth of mesenchymal stem cells. Adv. Mater. 2007, 19, 2530–2534. [Google Scholar] [CrossRef]
- Baik, K.Y.; Park, S.Y.; Heo, K.; Lee, K.B.; Hong, S. Carbon nanotube monolayer cues for osteogenesis of mesenchymal stem cells. Small 2011, 7, 741–745. [Google Scholar] [CrossRef] [PubMed]
- Akhavan, O.; Ghaderi, E.; Emamy, H.; Akhavan, F. Genotoxicity of graphene nanoribbons in human mesenchymal stem cells. Carbon 2013, 54, 419–431. [Google Scholar] [CrossRef]
- Trappmann, B.; Gautrot, J.E.; Connelly, J.T.; Strange, D.G.; Li, Y.; Oyen, M.L.; Cohen Stuart, M.A.; Boehm, H.; Li, B.; Vogel, V.; et al. Extracellular-matrix tethering regulates stem-cell fate. Nat. Mater. 2012, 11, 642–649. [Google Scholar] [CrossRef] [PubMed]
- Watt, F.M.; Huck, W.T. Role of the extracellular matrix in regulating stem cell fate. Nat. Rev. Mol. Cell Biol. 2013, 14, 467–473. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, A.; Hennessy, K.; Bellis, S. Regulation of mesenchymal stem cell attachment and spreading on hydroxyapatite by RGD peptides and adsorbed serum proteins. Biomaterials 2005, 26, 1467–1475. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.G.; Park, T.G. Biomimicking extracellular matrix: Cell adhesive RGD peptide modified electrospun poly (d, l-lactic-co-glycolic acid) nanofiber mesh. Tissue Eng. 2006, 12, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.-H.; Lee, D.; Choi, J.-W. Live cell biosensing platforms using graphene-based hybrid nanomaterials. Biosens. Bioelectron. 2017, 94, 485–499. [Google Scholar] [CrossRef] [PubMed]
- You, M.; Peng, G.; Li, J.; Ma, P.; Wang, Z.; Shu, W.; Peng, S.; Chen, G.-Q. Chondrogenic differentiation of human bone marrow mesenchymal stem cells on polyhydroxyalkanoate (PHA) scaffolds coated with PHA granule binding protein PhaP fused with RGD peptide. Biomaterials 2011, 32, 2305–2313. [Google Scholar] [CrossRef] [PubMed]
- Sawyer, A.; Weeks, D.; Kelpke, S.; McCracken, M.S.; Bellis, S. The effect of the addition of a polyglutamate motif to RGD on peptide tethering to hydroxyapatite and the promotion of mesenchymal stem cell adhesion. Biomaterials 2005, 26, 7046–7056. [Google Scholar] [CrossRef] [PubMed]
- Cao, H.; Tegenfeldt, J.O.; Austin, R.H.; Chou, S.Y. Gradient nanostructures for interfacing microfluidics and nanofluidics. Appl. Phys. Lett. 2002, 81, 3058–3060. [Google Scholar] [CrossRef]
- Lowe, C.R. Nanobiotechnology: The fabrication and applications of chemical and biological nanostructures. Curr. Opin. Struct. Biol. 2000, 10, 428–434. [Google Scholar] [CrossRef]
- Lu, C.-W.; Hung, Y.; Hsiao, J.-K.; Yao, M.; Chung, T.-H.; Lin, Y.-S.; Wu, S.H.; Hsu, S.C.; Liu, H.M.; Mou, C.Y.; et al. Bifunctional magnetic silica nanoparticles for highly efficient human stem cell labeling. Nano Lett. 2007, 7, 149–154. [Google Scholar] [CrossRef] [PubMed]
- Bharali, D.J.; Klejbor, I.; Stachowiak, E.K.; Dutta, P.; Roy, I.; Kaur, N.; Bergey, E.J.; Prasad, P.N.; Stachowiak, M.K. Organically modified silica nanoparticles: A nonviral vector for in vivo gene delivery and expression in the brain. Proc. Natl. Acad. Sci. USA 2005, 102, 11539–11544. [Google Scholar] [CrossRef] [PubMed]
- Hynes, R.O. Integrins: Versatility, modulation, and signaling in cell adhesion. Cell 1992, 69, 11–25. [Google Scholar] [CrossRef]
- Grinnell, B.W.; Hermann, R.B.; Yan, S.B. Human protein C inhibits selectin-mediated cell adhesion: Role of unique fucosylated oligosaccharide. Glycobiology 1994, 4, 221–225. [Google Scholar] [CrossRef] [PubMed]
- Takeichi, M. Cadherins: A molecular family important in selective cell-cell adhesion. Annu. Rev. Biochem. 1990, 59, 237–252. [Google Scholar] [CrossRef] [PubMed]
- Albelda, S. Role of integrins and other cell adhesion molecules in tumor progression and metastasis. Lab. Investig. 1993, 68, 4–17. [Google Scholar] [PubMed]
- Chapman, H.A. Plasminogen activators, integrins, and the coordinated regulation of cell adhesion and migration. Curr. Opin. Cell Biol. 1997, 9, 714–724. [Google Scholar] [CrossRef]
- Kim, D.-H.; Provenzano, P.P.; Smith, C.L.; Levchenko, A. Matrix nanotopography as a regulator of cell function. J. Cell Biol. 2012, 197, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Yim, E.K.; Darling, E.M.; Kulangara, K.; Guilak, F.; Leong, K.W. Nanotopography-induced changes in focal adhesions, cytoskeletal organization, and mechanical properties of human mesenchymal stem cells. Biomaterials 2010, 31, 1299–1306. [Google Scholar] [CrossRef] [PubMed]
- McNamara, L.E.; McMurray, R.J.; Biggs, M.J.; Kantawong, F.; Oreffo, R.O.; Dalby, M.J. Nanotopographical control of stem cell differentiation. J. Tissue Eng. Regen. Med. 2010, 1, 120623. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.N.; Jiao, A.; Hwang, N.S.; Kim, M.S.; Kim, D.-H.; Suh, K.-Y. Nanotopography-guided tissue engineering and regenerative medicine. Adv. Drug Deliv. Rev. 2013, 65, 536–558. [Google Scholar] [CrossRef] [PubMed]
- Dalby, M.J.; Gadegaard, N.; Tare, R.; Andar, A.; Riehle, M.O.; Herzyk, P.; Wilkinson, C.D.W.; Oreffo, R.O.C. The control of human mesenchymal cell differentiation using nanoscale symmetry and disorder. Nat. Mater. 2007, 6, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Shah, S. The nanomaterial tookit for neuroengineering. Nano Converg. 2016, 3, 25. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.-C.; Choo, S.-S.; Kim, K.-T.; Hong, K.-S.; Moon, S.-H.; Cha, H.-J.; Kim, T.-H. Conductive hybrid matrigel layer to enhance electrochemical signals of human embryonic stem cells. Sens. Actuator B Chem. 2017, 424, 224–230. [Google Scholar] [CrossRef]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).