Analysis of DNA Methylation Status in Bodily Fluids for Early Detection of Cancer
Abstract
:1. Introduction
2. Methods Used in Methylation Analyses
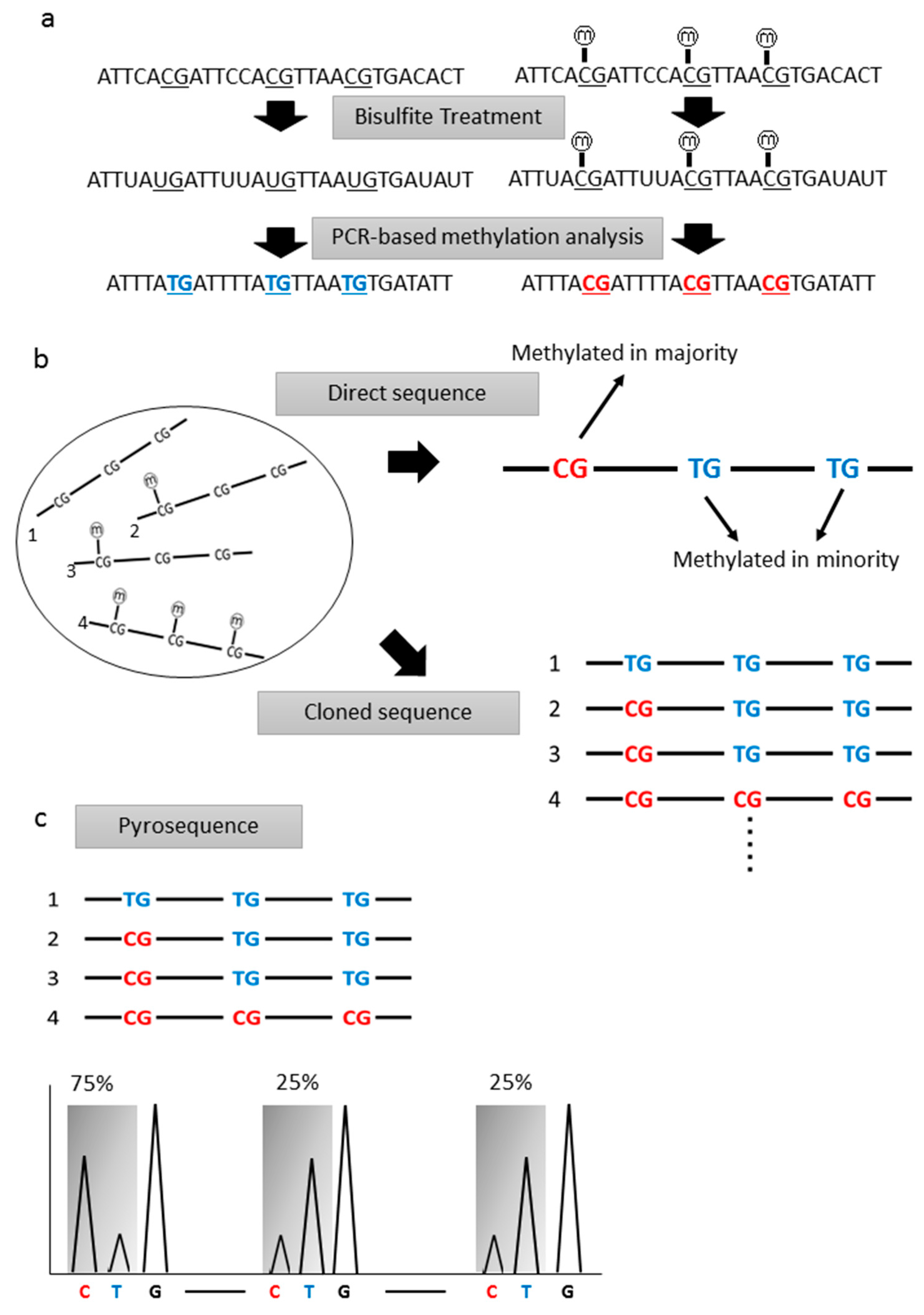
2.1. Sodium Bisulfite Treatment
2.2. Bisulfite Sequence Analyses
2.3. Pyrosequencing
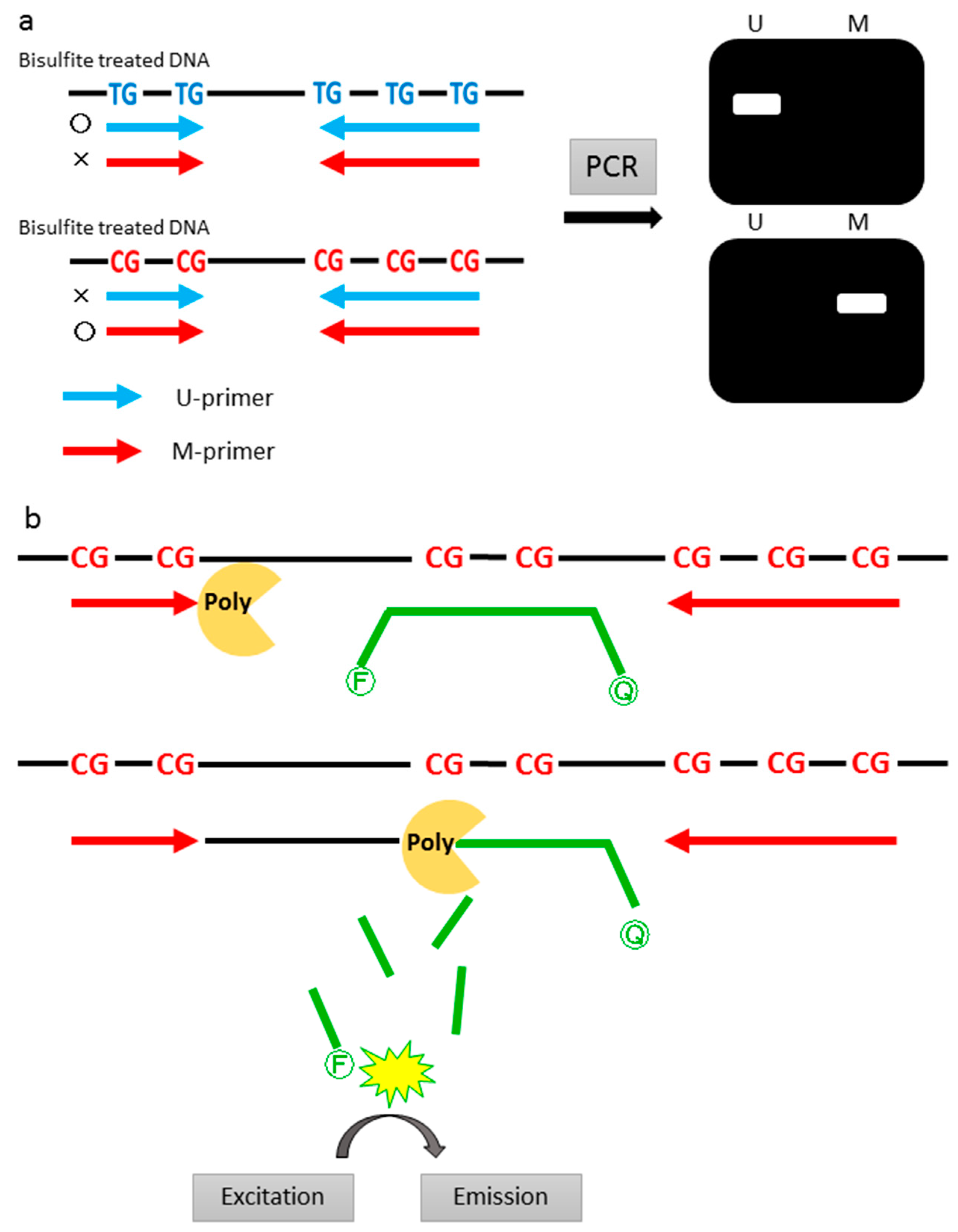
2.4. Methylation-Specific Polymerase Chain Reaction (PCR)
2.5. Quantitative Methylation Specific PCR (qMSP)
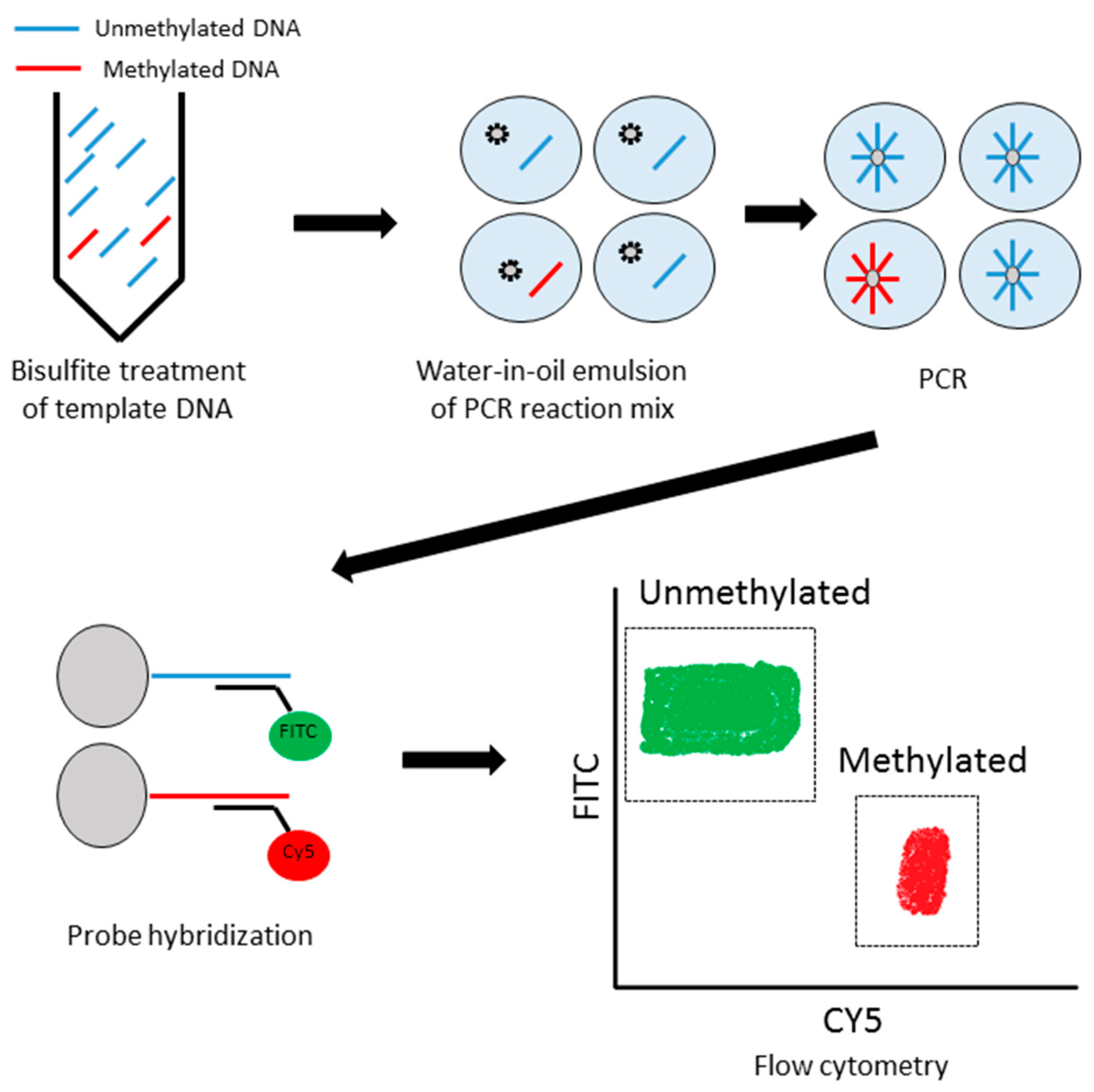
2.6. Methyl BEAMing
3. Use of Bodily Fluids for Cancer Detection
3.1. Saliva
3.2. Sputum
3.3. Stool
3.4. Urine (For the Detection of Bladder Cancer)
3.5. Urine (For the Detection of Prostate Cancer)
4. Conclusions
Supplementary Materials
Conflicts of Interest
References
- Hegi, M.E.; Diserens, A.C.; Gorlia, T.; Hamou, M.F.; de Tribolet, N.; Weller, M.; Kros, J.M.; Hainfellner, J.A.; Mason, W.; Mariani, L.; et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 2005, 352, 997–1003. [Google Scholar] [PubMed]
- Katoh, H.; Yamashita, K.; Waraya, M.; Margalit, O.; Ooki, A.; Tamaki, H.; Sakagami, H.; Kokubo, K.; Sidransky, D.; Watanabe, M. Epigenetic silencing of HOPX promotes cancer progression in colorectal cancer. Neoplasia 2012, 14, 559–571. [Google Scholar] [CrossRef]
- Yamashita, K.; Waraya, M.; Kim, M.S.; Sidransky, D.; Katada, N.; Sato, T.; Nakamura, T.; Watanabe, M. Detection of methylated CDO1 in plasma of colorectal cancer; a PCR study. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Minatani, N.; Waraya, M.; Yamashita, K.; Kikuchi, M.; Ushiku, H.; Kojo, K.; Ema, A.; Nishimiya, H.; Kosaka, Y.; Katoh, H.; et al. Prognostic significance of promoter DNA hypermethylation of cysteine dioxygenase 1 (CDO1) gene in primary breast cancer. PLoS ONE. 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Ushiku, H.; Yamashita, K.; Kawamata, H.; Waraya, M.; Katoh, H.; Yokoi, K.; Tanaka, T.; Ishii, S.; Nishizawa, N.; Kikuchi, M.; et al. Homeobox-only protein expression is a critical prognostic indicator of pancreatic neuroendocrine tumor and is regulated by promoter DNA hypermethylation. Pancreas 2016, 45, 1255–1262. [Google Scholar] [CrossRef] [PubMed]
- Wong, I.H.; Lo, Y.D.; Zhang, J.; Liew, C.T.; Ng, M.H.; Wong, N.; Lai, P.B.; Lau, W.Y.; Hjelm, N.M.; Johnson, P.J. Advances in brief detection of aberrant p16 methylation in the plasma and serum of liver cancer patients. Cancer Res. 1999, 59, 71–73. [Google Scholar] [PubMed]
- Lo, Y.D.; Chan, L.Y.; Lo, K.W.; Leung, S.F.; Zhang, J.; Chan, A.T.; Lee, J.C.; Hjelm, N.M.; Johnson, P.J.; Huang, D.P. Quantitative analysis of cell-free Epstein-Barr virus DNA in plasma of patients with nasopharyngeal carcinoma. Cancer Res. 1999, 59, 1188–1191. [Google Scholar] [PubMed]
- Diehl, F.; Li, M.; Dressman, D.; He, Y.; Shen, D.; Szabo, S.; Diaz, L.A.; Goodman, S.N.; David, K.A.; Juhl, H.; et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc. Natl. Acad. Sci. USA 2005, 102, 16368–16373. [Google Scholar] [CrossRef] [PubMed]
- Stroun, M.; Lyautey, J.; Lederrey, C.; Olson-Sand, A.; Anker, P. About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin. Chim. Acta 2001, 313, 139–142. [Google Scholar] [CrossRef]
- Bettegowda, C.; Sausen, M.; Leary, R.J.; Kinde, I.; Wang, Y.; Agrawal, N.; Bartlett, B.R.; Wang, H.; Luber, B.; Alani, R.M.; et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl. Med. 2014, 6. [Google Scholar] [CrossRef] [PubMed]
- Cheng, F.; Su, L.; Qian, C. Circulating tumor DNA: A promising biomarker in the liquid biopsy of cancer. Oncotarget 2016, 7, 48832–48841. [Google Scholar] [CrossRef] [PubMed]
- Anker, P.; Mulcahy, H.; Chen, X.Q.; Stroun, M. Detection of circulating tumour DNA in the blood (plasma/serum) of cancer patients. Cancer Metastasis Rev. 1999, 18, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Jahr, S.; Hentze, H.; Englisch, S.; Hardt, D.; Fackelmayer, F.O.; Hesch, R.D.; Knippers, R. DNA fragments in the blood plasma of cancer patients: Quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001, 61, 1659–1665. [Google Scholar] [PubMed]
- Dawson, S.J.; Tsui, D.W.; Murtaza, M.; Biggs, H.; Rueda, O.M.; Chin, S.F.; Dunning, M.J.; Gale, D.; Forshew, T.; Mahler-Araujo, B.; et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N. Engl. J. Med. 2013, 368, 1199–1209. [Google Scholar] [CrossRef] [PubMed]
- Siravegna, G.; Mussolin, B.; Buscarino, M.; Corti, G.; Cassingena, A.; Crisafulli, G.; Ponzetti, A.; Cremolini, C.; Amatu, A.; Lauricella, C.; et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat. Med. 2015, 21, 795–801. [Google Scholar] [CrossRef] [PubMed]
- Leary, R.J.; Kinde, I.; Diehl, F.; Schmidt, K.; Clouser, C.; Duncan, C.; Antipova, A.; Lee, C.; McKernan, K.; Francisco, M.; et al. Development of personalized tumor biomarkers using massively parallel sequencing. Sci. Transl. Med. 2010, 2. [Google Scholar] [CrossRef] [PubMed]
- Leary, R.J.; Sausen, M.; Kinde, I.; Papadopoulos, N.; Carpten, J.D.; Craig, D.; O’Shaughnessy, J.; Kinzler, K.W.; Parmigiani, G.; Vogelstein, B.; et al. Detection of chromosomal alterations in the circulation of cancer patients with whole-genome sequencing. Sci. Transl. Med. 2012, 4. [Google Scholar] [CrossRef] [PubMed]
- Dressman, D.; Yan, H.; Traverso, G.; Kinzler, K.W.; Vogelstein, B. Transforming single DNA molecules into fluorescent magnetic particles for detection and enumeration of genetic variations. Proc. Natl. Acad. Sci. USA 2003, 100, 8817–8822. [Google Scholar] [CrossRef] [PubMed]
- Esposito, A.; Criscitiello, C.; Locatelli, M.; Milano, M.; Curigliano, G. Liquid biopsies for solid tumors: Understanding tumor heterogeneity and real time monitoring of early resistance to targeted therapies. Pharmacol. Ther. 2016, 157, 120–124. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.; Ljubimov, V.A.; Zhou, C.; Tong, Y.; Liang, J. Cell-free circulating tumor DNA in cancer. Chin. J. Cancer 2016, 35. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, L.S.; Hansen, L.L. PCR-based methods for detecting single-locus DNA methylation biomarkers in cancer diagnostics, prognostics, and response to treatment. Clin. Chem. 2009, 55, 1471–1483. [Google Scholar] [CrossRef] [PubMed]
- Taylor, K.H.; Kramer, R.S.; Davis, J.W.; et al. Ultradeep bisulfite sequencing analysis of DNA methylation patterns in multiple gene promoters by 454 sequencing. Cancer Res. 2007, 67, 8511–8518. [Google Scholar] [CrossRef] [PubMed]
- Frommer, M.; McDonald, L.E.; Millar, D.S.; Collis, C.M.; Watt, F.; Grigg, G.W.; Molloy, P.L.; Paul, C.L. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl. Acad. Sci. USA 1992, 89, 1827–1831. [Google Scholar] [CrossRef] [PubMed]
- Herman, J.G.; Graff, J.R.; Myöhänen, S.; Nelkin, B.D.; Baylin, S.B. Methylation-specific PCR: A novel PCR assay for methylation status of CpG islands. Proc. Natl. Acad. Sci. USA 1996, 93, 9821–9826. [Google Scholar] [CrossRef] [PubMed]
- Eads, C.A.; Danenberg, K.D.; Kawakami, K.; Saltz, L.B.; Blake, C.; Shibata, D.; Danenberg, P.V.; Laird, P.W. MethyLight: A high-throughput assay to measure DNA methylation. Nucleic Acids Res. 2000, 28, E32. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Chen, W.D.; Papadopoulos, N.; Goodman, S.N.; Bjerregaard, N.C.; Laurberg, S.; Levin, B.; Juhl, H.; Arber, N.; Moinova, H.; et al. Sensitive digital quantification of DNA methylation in clinical samples. Nat. Biotechnol. 2009, 27, 858–863. [Google Scholar] [CrossRef] [PubMed]
- Rosas, S.L.; Koch, W.; da Costa Carvalho, M.D.; Wu, L.; Califano, J.; Westra, W.; Jen, J.; Sidransky, D. Promoter hypermethylation patterns of p16, O6-methylguanine-DNA-methyltransferase, and death-associated protein kinase in tumors and saliva of head and neck cancer patients. Cancer Res. 2001, 61, 939–942. [Google Scholar] [PubMed]
- Righini, C.A.; de Fraipont, F.; Timsit, J.F.; Faure, C.; Brambilla, E.; Reyt, E.; Favrot, M.C. Tumor-specific methylation in saliva: A promising biomarker for early detection of head and neck cancer recurrence. Clin. Cancer Res. 2007, 13, 1179–1185. [Google Scholar] [CrossRef] [PubMed]
- Demokan, S.; Chang, X.; Chuang, A.; Mydlarz, W.K.; Kaur, J.; Huang, P.; Khan, Z.; Khan, T.; Ostrow, K.L.; Brait, M.; et al. KIF1A and EDNRB are differentially methylated in primary HNSCC and salivary rinses. Int. J. Cancer 2010, 127, 2351–2359. [Google Scholar] [CrossRef] [PubMed]
- Pattani, K.M.; Zhang, Z.; Demokan, S.; Glazer, C.; Loyo, M.; Goodman, S.; Sidransky, D.; Bermudez, F.; Jean-Charles, G.; McCaffrey, T.; et al. Endothelin receptor type B gene promoter hypermethylation in salivary rinses is independently associated with risk of oral cavity cancer and premalignancy. Cancer Prev. Res. 2010, 3, 1093–1103. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.L.; Henrique, R.; Jeronimo, C.; Nayak, C.S.; Reddy, A.N.; Hoque, M.O.; Chang, S.; Brait, M.; Jiang, W.W.; Kim, M.M.; et al. Detection of promoter hypermethylation in salivary rinses as a biomarker for head and neck squamous cell carcinoma surveillance. Clin. Cancer Res. 2011, 17, 4782–4789. [Google Scholar] [CrossRef] [PubMed]
- Rettori, M.M.; de Carvalho, AC.; Longo, A.L.; de Oliveira, C.Z.; Kowalski, L.P.; Carvalho, A.L.; Vettore, A.L. Prognostic significance of TIMP3 hypermethylation in post-treatment salivary rinse from head and neck squamous cell carcinoma patients. Carcinogenesis 2013, 34, 20–27. [Google Scholar] [CrossRef] [PubMed]
- Schussel, J.; Zhou, X.C.; Zhang, Z.; Pattani, K.; Bermudez, F.; Jean-Charles, G.; McCaffrey, T.; Padhya, T.; Phelan, J.; Spivakovsky, S.; et al. EDNRB and DCC salivary rinse hypermethylation has a similar performance as expert clinical examination in discrimination of oral cancer/dysplasia versus benign lesions. Clin. Cancer Res. 2013, 19, 3268–3275. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, A.L.; Jeronimo, C.; Kim, M.M.; Henrique, R.; Zhang, Z.; Hoque, M.O.; Chang, S.; Brait, M.; Nayak, C.S.; Jiang, W.W.; et al. Evaluation of promoter hypermethylation detection in body fluids as a screening/diagnosis tool for head and neck squamous cell carcinoma. Clin. Cancer Res. 2008, 14, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Van Klaveren, R.J. Lung cancer screening. Eur. J. Cancer 2011, 47, S147–S155. [Google Scholar] [CrossRef]
- Rivera, M.P.; Mehta, A.C.; Wahidi, M.M. Establishing the diagnosis of lung cancer: Diagnosis and management of lung cancer, 3rd ed: American college of chest physicians evidence-based clinical practice guidelines. CHEST J. 2013, 143, e142S–e165S. [Google Scholar] [CrossRef] [PubMed]
- Belinsky, S.A.; Nikula, K.J.; Palmisano, W.A.; Michels, R.; Saccomanno, G.; Gabrielson, E.; Baylin, S.B.; Herman, J.G. Aberrant methylation of p16INK4a is an early event in lung cancer and a potential biomarker for early diagnosis. Proc. Natl. Acad. Sci. USA 1998, 95, 11891–11896. [Google Scholar] [CrossRef] [PubMed]
- Honorio, S.; Agathanggelou, A.; Schuermann, M.; Pankow, W.; Viacava, P.; Maher, E.R.; Latif, F. Detection of RASSF1A aberrant promoter hypermethylation in sputum from chronic smokers and ductal carcinoma in situ from breast cancer patients. Oncogene 2003, 22, 147–150. [Google Scholar] [CrossRef] [PubMed]
- Konno, S.; Morishita, Y.; Fukasawa, M.; Shu, Y.; Wang, D.; Tanaka, R.; Minami, Y.; Iijima, T.; Noguchi, M. Anthracotic index and DNA methylation status of sputum contents can be used for identifying the population at risk of lung carcinoma. Cancer 2004, 102, 348–354. [Google Scholar] [CrossRef] [PubMed]
- Cirincione, R.; Lintas, C.; Conte, D.; Mariani, L.; Roz, L.; Vignola, A.M.; Pastorino, U.; Sozzi, G. Methylation profile in tumor and sputum samples of lung cancer patients detected by spiral computed tomography: A nested case-control study. Int. J. Cancer 2006, 118, 1248–1253. [Google Scholar] [CrossRef] [PubMed]
- Belinsky, S.A.; Liechty, K.C.; Gentry, F.D.; Wolf, H.J.; Rogers, J.; Vu, K.; Haney, J.; Kennedy, T.C.; Hirsch, F.R.; Miller, Y.; et al. Promoter hypermethylation of multiple genes in sputum precedes lung cancer incidence in a high-risk cohort. Cancer Res. 2006, 66, 3338–3344. [Google Scholar] [CrossRef] [PubMed]
- Shivapurkar, N.; Stastny, V.; Suzuki, M.; Wistuba, I.I.; Li, L.; Zheng, Y.; Feng, Z.; Hol, B.; Prinsen, C.; Thunnissen, F.B.; et al. Application of a methylation gene panel by quantitative PCR for lung cancers. Cancer Lett. 2007, 247, 56–71. [Google Scholar] [CrossRef] [PubMed]
- Belinsky, S.A.; Grimes, M.J.; Casas, E.; Stidley, C.A.; Franklin, W.A.; Bocklage, T.J.; Johnson, D.H.; Schiller, J.H. Predicting gene promoter methylation in non-small-cell lung cancer by evaluating sputum and serum. Br. J. Cancer 2007, 96, 1278–1283. [Google Scholar] [CrossRef] [PubMed]
- Shivapurkar, N.; Stastny, V.; Okumura, N.; Girard, L.; Xie, Y.; Prinsen, C.; Thunnissen, F.B.; Wistuba, I.I.; Czerniak, B.; Frenkel, E.; et al. Cytoglobin, the newest member of the globin family, functions as a tumor suppressor gene. Cancer Res. 2008, 68, 7448–7456. [Google Scholar] [CrossRef] [PubMed]
- Guzmán, L.; Depix, M.S.; Salinas, A.M.; Roldán, R.; Aguayo, F.; Silva, A.; Vinet, R. Analysis of aberrant methylation on promoter sequences of tumor suppressor genes and total DNA in sputum samples: A promising tool for early detection of COPD and lung cancer in smokers. Diagn. Pathol. 2012, 7, 87. [Google Scholar] [CrossRef] [PubMed]
- Leng, S.; Do, K.; Yingling, C.M.; Picchi, M.A.; Wolf, H.J.; Kennedy, T.C.; Feser, W.J.; Baron, A.E.; Franklin, W.A.; Brock, M.V.; et al. Defining a gene promoter methylation signature in sputum for lung cancer risk assessment. Clin. Cancer Res. 2012, 18, 3387–3395. [Google Scholar] [CrossRef] [PubMed]
- Hubers, A.J.; van der Drift, M.A.; Prinsen, C.F.; Witte, B.I.; Wang, Y.; Shivapurkar, N.; Stastny, V.; Bolijn, A.S.; Hol, B.E.; Feng, Z.; et al. Methylation analysis in spontaneous sputum for lung cancer diagnosis. Lung Cancer 2014, 84, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Hubers, A.J.; Brinkman, P.; Boksem, R.J.; Rhodius, R.J.; Witte, B.I.; Zwinderman, A.H.; Heideman, D.A.; Duin, S.; Koning, R.; Steenbergen, R.D.; et al. Combined sputum hypermethylation and eNose analysis for lung cancer diagnosis. J. Clin. Pathol. 2014, 67, 707–711. [Google Scholar] [CrossRef] [PubMed]
- Hubers, A.J.; Heideman, D.A.; Burgers, S.A.; Herder, G.J.; Sterk, P.J.; Rhodius, R.J.; Smit, H.J.; Krouwels, F.; Welling, A.; Witte, B.I.; et al. DNA hypermethylation analysis in sputum for the diagnosis of lung cancer: Training validation set approach. Br. J. Cancer 2015, 112, 1105–1113. [Google Scholar] [CrossRef] [PubMed]
- Hulbert, A.; Torres, I.J.; Stark, A.; Chen, C.; Rodgers, K.; Lee, B.; Griffin, C.; Yang, A.; Huang, P.; Wrangle, J.; et al. Early detection of lung cancer using DNA promoter hypermethylation in plasma and sputum. Clin. Cancer Res. 2016. [Google Scholar] [CrossRef] [PubMed]
- Mandel, J.S.; Bond, J.H.; Church, T.R.; Snover, D.C.; Bradley, G.M.; Schuman, L.M.; Ederer, F. Reducing mortality from colorectal cancer by screening for fecal occult blood. N. Engl. J. Med. 1993, 328, 1365–1371. [Google Scholar] [CrossRef] [PubMed]
- Hardcastle, J.D.; Chamberlain, J.O.; Robinson, M.H.; Moss, S.M.; Amar, S.S.; Balfour, T.W.; James, P.D.; Mangham, C.M. Randomised controlled trial of faecal-occult-blood screening for colorectal cancer. Lancet 1996, 348, 1472–1477. [Google Scholar] [CrossRef]
- Kronborg, O.; Fenger, C.; Olsen, J.; Jørgensen, O.D.; Søndergaard, O. Randomised study of screening for colorectal cancer with faecal-occult-blood test. Lancet 1996, 348, 1467–1471. [Google Scholar] [CrossRef]
- Ahlquist, D.A.; Sargent, D.J.; Loprinzi, C.L.; Levin, T.R.; Rex, D.K.; Ahnen, D.J.; Knigge, K.; Lance, M.P.; Burgart, L.J.; Hamilton, S.R.; et al. Stool DNA and occult blood testing for screen detection of colorectal neoplasia. Ann. Intern. Med. 2008, 149, 441–450. [Google Scholar] [CrossRef] [PubMed]
- Park, D.I.; Ryu, S.; Kim, Y.H.; Lee, S.H.; Lee, C.K.; Eun, C.S.; Han, D.S.; et al. Comparison of guaiac-based and quantitative immunochemical fecal occult blood testing in a population at average risk undergoing colorectal cancer screening. Am. J. Gastroenterol. 2010, 105, 2017–2025. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.K.; Liles, E.G.; Bent, S.; Levin, T.R.; Corley, D.A. Accuracy of fecal immunochemical tests for colorectal cancer: Systematic review and meta-analysis. Ann. Intern. Med. 2014, 160, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Van Rossum, L.G.; van Rijn, A.F.; Laheij, R.J.; van Oijen, M.G.; Fockens, P.; van Krieken, H.H.; Verbeek, A.L.; Jansen, J.B.; Dekker, E. Random comparison of guaiac and immunochemical fecal occult blood tests for colorectal cancer in a screening population. Gastroenterology 2008, 135, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Parra-Blanco, A.; Gimeno-García, A.Z.; Quintero, E.; Nicolás, D.; Moreno, S.G.; Jiménez, A.; Hernández-Guerra, M.; Carrillo-Palau, M.; Eishi, Y.; López-Bastida, J. Diagnostic accuracy of immunochemical versus guaiac faecal occult blood tests for colorectal cancer screening. J. Gastroenterol. 2010, 45, 703–712. [Google Scholar] [CrossRef] [PubMed]
- Müller, H.M.; Oberwalder, M.; Fiegl, H.; Morandell, M.; Goebel, G.; Zitt, M.; Mühlthaler, M.; Öfner, D.; Margreiter, R.; Widschwendter, M. Methylation changes in faecal DNA: A marker for colorectal cancer screening? Lancet 2004, 363, 1283–1285. [Google Scholar] [CrossRef]
- Song, J.; Park, K.U.; Park, H.D.; Yoon, Y.; Kim, J.Q. High-throughput liquid chromatography-tandem mass spectrometry assay for plasma theophylline and its metabolites. Clin. Chem. 2004, 50, 2176–2179. [Google Scholar] [CrossRef] [PubMed]
- Lenhard, K.; Bommer, G.T.; Asutay, S.; Schauer, R.; Brabletz, T.; Göke, B.; Lamerz, R.; Kolligs, F.T. Analysis of promoter methylation in stool: A novel method for the detection of colorectal cancer. Clin. Gastroenterol. Hepatol. 2005, 3, 142–149. [Google Scholar] [CrossRef]
- Chen, W.D.; Han, Z.J.; Skoletsky, J.; Olson, J.; Sah, J.; Myeroff, L.; Platzer, P.; Lu, S.; Dawson, D.; Willis, J.; et al. Detection in fecal DNA of colon cancer-specific methylation of the nonexpressed vimentin gene. J. Natl. Cancer Inst. 2005, 97, 1124–1132. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.H.; Li, L.H.; Yang, F.; Wang, J.F. Detection of aberrant methylation in fecal DNA as a molecular screening tool for colorectal cancer and precancerous lesions. World J. Gastroenterol. 2007, 13, 950–954. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Bauer, M.; Croner, R.S.; Pelz, J.O.; Lodygin, D.; Hermeking, H.; Stürzl, M.; Hohenberger, W.; Matzel, K.E. DNA stool test for colorectal cancer: Hypermethylation of the secreted frizzled-related protein-1 gene. Dis. Colon Rectum. 2007, 50, 1618–1627. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.R.; Tang, D. Hypermethylated SFRP2 gene in fecal DNA is a high potential biomarker for colorectal cancer noninvasive screening. World J. Gastroenterol. 2008, 14, 524–531. [Google Scholar] [CrossRef] [PubMed]
- Baek, Y.H.; Chang, E.; Kim, Y.J.; Kim, B.K.; Sohn, J.H.; Park, D.I. Stool methylation-specific polymerase chain reaction assay for the detection of colorectal neoplasia in Korean patients. Dis. Colon Rectum. 2009, 52, 1452–1463. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Louwagie, J.; Carvalho, B.; sive Droste, J.S.; Park, H.L.; Chae, Y.K.; Yamashita, K.; Liu, J.; Ostrow, K.L.; Ling, S.; et al. Promoter DNA methylation of oncostatin M receptor-β as a novel diagnostic and therapeutic marker in colon cancer. PLoS ONE 2009, 4, e6555. [Google Scholar] [CrossRef] [PubMed]
- Glöckner, S.C.; Dhir, M.; Yi, J.M.; McGarvey, K.E.; van Neste, L.; Louwagie, J.; Chan, T.A.; Kleeberger, W.; de Bruïne, A.P.; Smits, K.M.; et al. Methylation of TFPI2 in stool DNA: A potential novel biomarker for the detection of colorectal cancer. Cancer Res. 2009, 69, 4691–4699. [Google Scholar] [CrossRef] [PubMed]
- Nagasaka, T.; Tanaka, N.; Cullings, H.M.; Sun, D.S.; Sasamoto, H.; Uchida, T.; Koi, M.; Nishida, N.; Naomoto, Y.; Boland, C.R.; et al. Analysis of fecal DNA methylation to detect gastrointestinal neoplasia. J. Natl. Cancer Inst. 2009, 101, 1244–1258. [Google Scholar] [CrossRef] [PubMed]
- Hellebrekers, D.M.; Lentjes, M.H.; van den Bosch, S.M.; Melotte, V.; Wouters, K.A.; Daenen, K.L.; Smits, K.M.; Akiyama, Y.; Yuasa, Y.; Sanduleanu, S.; et al. GATA4 and GATA5 are potential tumor suppressors and biomarkers in colorectal cancer. Clin. Cancer Res. 2009, 15, 3990–3997. [Google Scholar] [CrossRef] [PubMed]
- Melotte, V.; Lentjes, M.H.; van den Bosch, S.M.; Hellebrekers, D.M.; de Hoon, J.P.; Wouters, K.A.; Daenen, K.L.; Partouns-Hendriks, I.E.; Stessels, F.; Louwagie, J.; et al. N-Myc Downstream-Regulated Gene 4 (NDRG4): A candidate tumor suppressor gene and potential biomarker for colorectal cancer. J. Natl. Cancer Inst. 2009, 101, 916–927. [Google Scholar] [CrossRef] [PubMed]
- Ahlquist, D.A.; Zou, H.; Domanico, M.; Mahoney, D.W.; Yab, T.C.; Taylor, W.R.; Butz, M.L.; Thibodeau, S.N.; Rabeneck, L.; Paszat, L.F.; et al. Next-generation stool DNA test accurately detects colorectal cancer and large adenomas. Gastroenterology 2012, 142, 248–256. [Google Scholar] [CrossRef] [PubMed]
- Imperiale, T.F.; Ransohoff, D.F.; Itzkowitz, S.H.; Levin, T.R.; Lavin, P.; Lidgard, G.P.; Ahlquist, D.A.; Berger, B.M. Multitarget stool DNA testing for colorectal-cancer screening. N. Engl. J. Med. 2014, 370, 1287–1297. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhu, Y.Q.; Wu, Y.Q.; Zhang, P.; Qi, J. Detection of promoter hypermethylation of Wnt antagonist genes in fecal samples for diagnosis of early colorectal cancer. World J. Gastroenterol. 2014, 20, 6329–6335. [Google Scholar] [CrossRef] [PubMed]
- Têtu, B. Diagnosis of urothelial carcinoma from urine. Mod. Pathol. 2009, 2, S53–S59. [Google Scholar] [CrossRef] [PubMed]
- Chan, M.W.; Chan, L.W.; Tang, N.L.; Tong, J.H.; Lo, K.W.; Lee, T.L.; Cheung, H.Y.; Wong, W.S.; Chan, P.S.; Lai, F.M.; et al. Hypermethylation of multiple genes in tumor tissues and voided urine in urinary bladder cancer patients. Clin. Cancer Res. 2002, 8, 464–470. [Google Scholar] [PubMed]
- Chan, M.W.; Chan, L.W.; Tang, N.L.; Lo, K.W.; Tong, J.H.; Chan, A.W.; Cheung, H.Y.; Wong, W.S.; Chan, P.S.; Lai, F.M.; et al. Frequent hypermethylation of promoter region of RASSF1A in tumor tissues and voided urine of urinary bladder cancer patients. Int. J. Cancer 2003, 104, 611–616. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, M.G.; Weisenberger, D.J.; Cheng, J.C.; Chandrasoma, S.; Siegmund, K.D.; Gonzalgo, M.L.; Toma, M.I.; Huland, H.; Yoo, C.; Tsai, Y.C.; et al. Detection of methylated apoptosis-associated genes in urine sediments of bladder cancer patients. Clin. Cancer Res. 2004, 10, 7457–7465. [Google Scholar] [CrossRef] [PubMed]
- Sathyanarayana, U.G.; Maruyama, R.; Padar, A.; Suzuki, M.; Bondaruk, J.; Sagalowsky, A.; Minna, J.D.; Frenkel, E.P.; Grossman, H.B.; Czerniak, B.; et al. Molecular detection of noninvasive and invasive bladder tumor tissues and exfoliated cells by aberrant promoter methylation of laminin-5 encoding genes. Cancer Res. 2004, 64, 1425–1430. [Google Scholar] [CrossRef] [PubMed]
- Dulaimi, E.; Uzzo, R.G.; Greenberg, R.E.; Al-Saleem, T.; Cairns, P. Detection of bladder cancer in urine by a tumor suppressor gene hypermethylation panel. Clin. Cancer Res. 2004, 10, 1887–1893. [Google Scholar] [CrossRef] [PubMed]
- Yates, D.R.; Rehman, I.; Meuth, M.; Cross, S.S.; Hamdy, F.C.; Catto, J.W. Methylational urinalysis: A prospective study of bladder cancer patients and age stratified benign controls. Oncogene 2006, 25, 1984–1988. [Google Scholar] [CrossRef] [PubMed]
- Hoque, M.O.; Begum, S.; Topaloglu, O.; Chatterjee, A.; Rosenbaum, E.; van Criekinge, W.; Westra, W.H.; Schoenberg, M.; Zahurak, M.; Goodman, S.N.; et al. Quantitation of promoter methylation of multiple genes in urine DNA and bladder cancer detection. J. Natl. Cancer Inst. 2006, 98, 996–1004. [Google Scholar] [CrossRef] [PubMed]
- Urakami, S.; Shiina, H.; Enokida, H.; Kawakami, T.; Kawamoto, K.; Hirata, H.; Tanaka, Y.; Kikuno, N.; Nakagawa, M.; Igawa, M.; et al. Combination analysis of hypermethylated Wnt-antagonist family genes as a novel epigenetic biomarker panel for bladder cancer detection. Clin. Cancer Res. 2006, 12, 2109–2116. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Zhu, T.; Wang, Z.; Zhang, H.; Qian, Z.; Xu, H.; Gao, B.; Wang, W.; Gu, L.; Meng, J.; et al. A novel set of DNA methylation markers in urine sediments for sensitive/specific detection of bladder cancer. Clin. Cancer Res. 2007, 13, 7296–7304. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Chen, Z.; Zhu, T.; Yu, J.; Ma, K.; Zhang, H.; He, Y.; Luo, X.; Zhu, J. Hypermethylated SFRP1, but none of other nine genes “informative” for western countries, is valuable for bladder cancer detection in Mainland China. J. Cancer Res. Clin. Oncol. 2009, 135, 1717–1727. [Google Scholar] [CrossRef] [PubMed]
- Renard, I.; Joniau, S.; van Cleynenbreugel, B.; Collette, C.; Naômé, C.; Vlassenbroeck, I.; Nicolas, H.; de Leval, J.; Straub, J.; van Criekinge, W.; et al. Identification and validation of the methylated TWIST1 and NID2 genes through real-time methylation-specific polymerase chain reaction assays for the noninvasive detection of primary bladder cancer in urine samples. Eur. Urol. 2010, 58, 96–104. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.H.; Ke, H.L.; Huang, S.P.; Wu, W.J.; Chen, Y.K.; Chang, L.L. Increase sensitivity in detecting superficial, low grade bladder cancer by combination analysis of hypermethylation of E-cadherin, p16, p14, RASSF1A genes in urine. Urol. Oncol. Semin. Orig. Investig. 2010, 28, 597–602. [Google Scholar] [CrossRef] [PubMed]
- Eissa, S.; Swellam, M.; El-Khouly, I.M.; Kassim, S.K.; Shehata, H.; Mansour, A.; Esmat, M.; Nossier, A.I.; Hamdy, M.A.; Awad, N.M.; et al. Aberrant methylation of RARβ2 and APC genes in voided urine as molecular markers for early detection of bilharzial and nonbilharzial bladder cancer. Cancer Epidemiol. Biomark. Prev. 2011, 20, 1657–1664. [Google Scholar] [CrossRef] [PubMed]
- Reinert, T.; Modin, C.; Castano, F.M.; Lamy, P.; Wojdacz, T.K.; Hansen, L.L.; Wiuf, C.; Borre, M.; Dyrskjøt, L.; Ørntoft, T.F. Comprehensive genome methylation analysis in bladder cancer: Identification and validation of novel methylated genes and application of these as urinary tumor markers. Clin. Cancer Res. 2011, 17, 5582–5592. [Google Scholar] [CrossRef] [PubMed]
- Vinci, S.; Giannarini, G.; Selli, C.; Kuncova, J.; Villari, D.; Valent, F.; Orlando, C. Quantitative methylation analysis of BCL2, hTERT, and DAPK promoters in urine sediment for the detection of non-muscle-invasive urothelial carcinoma of the bladder: A prospective, two-center validation study. Urol. Oncol. Semin. Orig. Investig. 2011, 29, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.C.; Tsai, M.H.; Yip, S.K.; Jou, Y.C.; Ng, C.F.; Chen, Y.; Wang, X.; Huang, W.; Tung, C.L.; Chen, G.C.; et al. Distinct DNA methylation epigenotypes in bladder cancer from different Chinese sub-populations and its implication in cancer detection using voided urine. BMC Med. Genom. 2011, 4. [Google Scholar] [CrossRef] [PubMed]
- Serizawa, R.R.; Ralfkiær, U.; Steven, K.; Lam, G.W.; Schmiedel, S.; Schüz, J.; Hansen, A.B.; Horn, T.; Guldberg, P. Integrated genetic and epigenetic analysis of bladder cancer reveals an additive diagnostic value of FGFR3 mutations and hypermethylation events. Int. J. Cancer 2011, 129, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Costa, V.L.; Henrique, R.; Danielsen, S.A.; Eknaes, M.; Patrício, P.; Morais, A.; Oliveira, J.; Lothe, R.A.; Teixeira, M.R.; Lind, G.E.; et al. TCF21 and PCDH17 methylation: An innovative panel of biomarkers for a simultaneous detection of urological cancers. Epigenetics 2011, 6, 1120–1130. [Google Scholar] [CrossRef] [PubMed]
- Chung, W.; Bondaruk, J.; Jelinek, J.; Lotan, Y.; Liang, S.; Czerniak, B.; Issa, J.P. Detection of bladder cancer using novel DNA methylation biomarkers in urine sediments. Cancer Epidemiol. Biomark. Prev. 2011, 20, 1483–1491. [Google Scholar] [CrossRef] [PubMed]
- Reinert, T.; Borre, M.; Christiansen, A.; Hermann, G.G.; Ørntoft, T.F.; Dyrskjøt, L. Diagnosis of bladder cancer recurrence based on urinary levels of EOMES, HOXA9, POU4F2, TWIST1, VIM, and ZNF154 hypermethylation. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Chihara, Y.; Kanai, Y.; Fujimoto, H.; Sugano, K.; Kawashima, K.; Liang, G.; Jones, P.A.; Fujimoto, K.; Kuniyasu, H.; Hirao, Y. Diagnostic markers of urothelial cancer based on DNA methylation analysis. BMC Cancer 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, M.; Bernert, H.; Kagohara, L.T.; Maldonado, L.; Brait, M.; Schoenberg, M.; Bivalacqua, T.; Netto, G.J.; Koch, W.; Sidransky, D.; et al. Epigenetic inactivation of VGF associated with Urothelial Cell Carcinoma and its potential as a non-invasive biomarker using urine. Oncotarget 2014, 5, 3350–3361. [Google Scholar] [CrossRef] [PubMed]
- Abern, M.R.; Owusu, R.; Inman, B.A. Clinical performance and utility of a DNA methylation urine test for bladder cancer. Urol. Oncol. 2014, 32, 51.e21–51.e26. [Google Scholar] [CrossRef] [PubMed]
- Yeh, C.M.; Chen, P.C.; Hsieh, H.Y.; Jou, Y.C.; Lin, C.T.; Tsai, M.H.; Huang, W.Y.; Wang, Y.T.; Lin, R.I.; Chen, S.S.; et al. Methylomics analysis identifies ZNF671 as an epigenetically repressed novel tumor suppressor and a potential non-invasive biomarker for the detection of urothelial carcinoma. Oncotarget 2015, 6, 29555–29572. [Google Scholar] [PubMed]
- Fantony, J.J.; Abern, M.R.; Gopalakrishna, A.; Owusu, R.; Tay, K.J.; Lance, R.S.; Inman, B.A. Multi-institutional external validation of urinary TWIST1 and NID2 methylation as a diagnostic test for bladder cancer. Urol. Oncol. 2015, 33, 387.e1–387.e6. [Google Scholar] [CrossRef] [PubMed]
- Roperch, J.P.; Grandchamp, B.; Desgrandchamps, F.; Mongiat-Artus, P.; Ravery, V.; Ouzaid, I.; Roupret, M.; Phe, V.; Ciofu, C.; Tubach, F.; et al. Promoter hypermethylation of HS3ST2, SEPTIN9 and SLIT2 combined with FGFR3 mutations as a sensitive/specific urinary assay for diagnosis and surveillance in patients with low or high-risk non-muscle-invasive bladder cancer. BMC Cancer 2016, 16. [Google Scholar] [CrossRef] [PubMed]
- Dahmcke, C.M.; Steven, K.E.; Larsen, L.K.; Poulsen, A.L.; Abdul-Al, A.; Dahl, C.; Guldberg, P. A prospective blinded evaluation of urine-DNA testing for detection of urothelial bladder carcinoma in patients with gross hematuria. Eur. Urol. 2016, 70, 916–919. [Google Scholar] [CrossRef] [PubMed]
- Catalona, W.J.; Partin, A.W.; Slawin, K.M.; Brawer, M.K.; Flanigan, R.C.; Patel, A.; Richie, J.P.; Walsh, P.C.; Scardino, P.T.; Lange, P.H.; et al. Use of the percentage of free prostate-specific antigen to enhance differentiation of prostate cancer from benign prostatic disease: A prospective multicenter clinical trial. JAMA 1998, 279, 1542–1547. [Google Scholar] [CrossRef] [PubMed]
- Thompson, I.M.; Ankerst, D.P.; Chi, C.; Lucia, M.S.; Goodman, P.J.; Crowley, J.J.; Parnes, H.L.; Coltman, C.A. Operating characteristics of prostate-specific antigen in men with an initial PSA level of 3.0 ng/mL or lower. JAMA 2005, 294, 66–70. [Google Scholar] [CrossRef] [PubMed]
- Goessl, C.; Krause, H.; Müller, M.; Heicappell, R.; Schrader, M.; Sachsinger, J.; Miller, K. Fluorescent methylation-specific polymerase chain reaction for DNA-based detection of prostate cancer in bodily fluids. Cancer Res. 2000, 60, 5941–5945. [Google Scholar] [PubMed]
- Cairns, P.; Esteller, M.; Herman, J.G.; Schoenberg, M.; Jeronimo, C.; Sanchez-Cespedes, M.; Chow, N.H.; Grasso, M.; Wu, L.; Westra, W.B.; et al. Molecular detection of prostate cancer in urine by GSTP1 hypermethylation. Clin. Cancer Res. 2001, 7, 2727–2730. [Google Scholar] [PubMed]
- Goessl, C.; Müller, M.; Heicappell, R.; Krause, H.; Straub, B.; Schrader, M.; Miller, K. DNA-based detection of prostate cancer in urine after prostatic massage. Urology 2001, 58, 335–338. [Google Scholar] [CrossRef]
- Goessl, C.; Muller, M.; Heicappell, R.; Krause, H.; Miller, K. DNA-based detection of prostate cancer in blood, urine, and ejaculates. Ann. N. Y. Acad. Sci. 2001, 945, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Jernimo, C.; Usadel, H.; Henrique, R.; Silva, C.; Oliveira, J.; Lopes, C.; Sidransky, D. Quantitative GSTP1 hypermethylation in bodily fluids of patients with prostate cancer. Urology 2002, 60, 1131–1135. [Google Scholar] [CrossRef]
- Hoque, M.O.; Topaloglu, O.; Begum, S.; Henrique, R.; Rosenbaum, E.; van Criekinge, W.; Westra, W.H.; Sidransky, D. Quantitative methylation-specific polymerase chain reaction gene patterns in urine sediment distinguish prostate cancer patients from control subjects. J. Clin. Oncol. 2005, 23, 6569–6575. [Google Scholar] [CrossRef] [PubMed]
- Rouprêt, M.; Hupertan, V.; Yates, D.R.; Catto, J.W.; Rehman, I.; Meuth, M.; Ricci, S.; Lacave, R.; Cancel-Tassin, G.; de la Taille, A.; et al. Molecular detection of localized prostate cancer using quantitative methylation-specific PCR on urinary cells obtained following prostate massage. Clin. Cancer Res. 2007, 13, 1720–1725. [Google Scholar] [CrossRef] [PubMed]
- Baden, J.; Green, G.; Painter, J.; Curtin, K.; Markiewicz, J.; Jones, J.; Astacio, T.; Canning, S.; Quijano, J.; Guinto, W.; et al. Multicenter evaluation of an investigational prostate cancer methylation assay. J. Urol. 2009, 182, 1186–1193. [Google Scholar] [CrossRef] [PubMed]
- Daniunaite, K.; Jarmalaite, S.; Kalinauskaite, N.; Petroska, D.; Laurinavicius, A.; Lazutka, J.R.; Jankevicius, F. Prognostic value of RASSF1 promoter methylation in prostate cancer. J. Urol. 2014, 192, 1849–1855. [Google Scholar] [CrossRef] [PubMed]
- Vener, T.; Derecho, C.; Baden, J.; Wang, H.; Rajpurohit, Y.; Skelton, J.; Mehrotra, J.; Varde, S.; Chowdary, D.; Stallings, W.; et al. Development of a multiplexed urine assay for prostate cancer diagnosis. Clin. Chem. 2008, 54, 874–882. [Google Scholar] [CrossRef] [PubMed]
| Author | Year | Method | Prospective Study | Sample Size (Number of Patients) | Gene | Sensitivity (%) | Specificity (%) |
|---|---|---|---|---|---|---|---|
| Rosas et al. [27] | 2001 | MSP | No | HNSCC (30) Healthy control (30) | DAPK, MGMT, p16 | At least 1 gene (37%) | At least 1 gene (97%) |
| Righini et al. [28] | 2007 | MSP | No | HNSCC (60) Non Malignant (30) | CDH1, DAPK, MGMT, p16, RASSF1, TIMP3 | At least 1 gene (79%) | At least 1 gene (100%) |
| Carvalho et al. [34] | 2008 | qMSP | No | HNSCC (211) Control (527) | AIM1, CCNA1, CCND2, CDH1, DAPK, DCC, ESR1, MGMT, MINT1, MINT31, PGP9.5, p16, TIMP3 | At least 1 of 4 gene (31%) | At least 1 of 4 gene (90%) |
| Demokan et al. [29] | 2010 | qMSP | No | HNSCC (71) Healthy Control (61) | EDNRB, KIF1A | EDRNB + KIF1A (77%) | EDRNB + KIF1A (93%) |
| Pattani et al. [30] | 2010 | qMSP | Yes | Clinically high risk patients Total (191) Malignant (35) Premalignant (43) Benign (113) | EDNRB | EDNRB (65%) | EDNRB (51%) |
| Carvalho et al. [31] | 2011 | qMSP | No | HNSCC (61) | CCNA1, DAPK, DCC, MGMT, MINT31, p16, TIMP3 | At least 1 gene (54%) | |
| Schussel et al. [33] | 2013 | qMSP | Yes | Clinically high risk patients HNSCC or dysplasia (48) Benign (113) | DCC, EDNRB | EDRNB + DCC + risk classification (75%) | EDRNB + DCC + risk classification (48%) |
| Rettori et al. [32] | 2013 | qMSP | No | HNSCC (146) Healthy control (60) | CCNA1, DAPK, DCC, MGMT, TIMP3, and other 19 genes | At least 1 gene (55%) | At least 1 gene (76%) |
| Author | Year | Method | Prospective Study | Sample Size (Number of Patients) | Gene | Sensitivity (%) | Specificity (%) |
|---|---|---|---|---|---|---|---|
| Belinsky et al. [37] | 1998 | MSP | No | LC (7) Smokers (26) | p16 | p16 (43%) | p16 (81%) |
| Honorio et al. [38] | 2003 | MSP | No | SCLC (8) NSCLC (24) Chronic Smokers (13) | RASSF1A | SCLC; RASSF1A (50%) NSCLC; RASSF1A (21%) Chronic Smokers; RASSF1A (31%) | |
| Konno et al. [39] | 2004 | MSP | No | LC (78) None LC (52) | APC, p16, RARβ | APC (28%), p16 (22%), RARβ (27%) | APC (96%), p16 (100%), RARβ (93%) |
| Belinsky et al. [41] | 2006 | Nested MSP | No | LC (98) Healthy Controls (92) | BETA3, DAPK, GATA4, GATA5, HCAD, HLHP, IGFBP3, LAMC2, MGMT, PAX5α, PAX5β, p16, RASSF1A, SFRP1 | GATA5 (74%), LAMC2 (72%), SFRP1 (68%) | GATA5 (74%), LAMC2 (30%), SFRP1 (29%) |
| Cirincione et al. [40] | 2006 | MSP | No | LC (18) Healthy Controls (smoker) (112) | p16, RARβ2, RASSF1A | At least 1 gene (50%) | At least 1 gene (38%) |
| Belinsky et al. [43] | 2007 | MSP | No | LC (Stage III) (72) | DAPK, GATA5, HCAD, MGMT, PAX5α, PAX5β, p16, RASSF1A | GATA5 (43%), MGMT (32%), p16 (40%) | |
| Shivapurkar et al. [42] | 2007 | qMSP | No | NSCLC (13) Controls without LC (25) | APC, p16, RASSF1A, HS3ST2 | At least 1 gene (62%) | At least 1 gene (100%) |
| Shivapurkar et al. [44] | 2008 | qMSP | No | LC (13) Non Cancer (25) | CYGB | CYGB (30%) | CYGB (100%) |
| Guzmán et al. [45] | 2012 | MSP | No | LC (26) COPD (23) Healthy Controls (33) | CDH1, MGMT, p16 | LC CDH1 (35%), MGMT (65%), p16 (73%) COPD CDH1 (45%), MGMT (65%), p16 (70%) Healthy controls CDH1 (32%), MGMT (6%), p16 (9%) | |
| Leng et al. [46] | 2012 | Nested MSP (cohort1) qMSP (cohort2) | No | Cohort 1 LC (64) Non Cancer (64) Cohort 2 LC (40) Non Cancer (90) | GATA5, PAX5α, SULF2 | Cohort 1 GATA5 (33%), PAX5α (25%), SULF2 (34%) Cohort 2 GATA5 (78%), PAX5α (63%), SULF2 (78%) | Cohort 1 GATA5 (74%), PAX5α (80%), SULF2 (75%) Cohort 2 GATA5 (53%), PAX5α (67%), SULF2 (45%) |
| Hubers et al. [48] | 2014 | qMSP | No | LC (20) COPD (31) | APC, CYGB, FAM19A4, HS3ST2, PHACTR3, PRDM14, RASSF1A | RASSF1A + 3OST2 (85%) | RASSF1A + 3OST2 (74%) |
| Hubers et al. [47] | 2014 | qMSP | No | Set1 LC (98) None LC (90) Set2 LC (60) none LC (445) | APC, CYGB, RASSF1A | Set1; At least 1 gene (63%) Set2; At least 1 gene (90%) | Set1; At least 1 gene (78%) Set2; At least 1 gene (47%) |
| Hubers et al. [49] | 2015 | qMSP | No | Learning set LC (73) none LC (86) Validation set LC (159) none LC (154) | APC, CYGB, FA19A4, HS3ST2, PHACTR3, PRDM14, RASSF1A | Learning Set HS3ST2 + PHACTR3 + RASSF1A (82%) Validation Set HS3ST2 + PHACTR3 + RASSF1A (79%) | Learning Set HS3ST2 + PHACTR3 + RASSF1A (66%) Validation Set HS3ST2 + PHACTR3 + RASSF1A (64%) |
| Hulbert et al. [50] | 2016 | qMSP | No | LC (90) none LC (24) | CDO1, HOXA7, HOXA9, SOX17, TAC1, ZFP42 | HOXA7 + SOX17 + TAC1 (98%) | HOXA7 + SOX17 + TAC1 (71%) |
| Author | Year | Method | Prospective Study | Sample Size (Number of Patients) | Gene | Sensitivity (%) | Specificity (%) |
|---|---|---|---|---|---|---|---|
| Song et al. [60] | 2004 | MSP | No | CRC (20) Normal CF (20) | APC, ATM, HLTF, MGMT, hMLH-1 | At least 1 gene (70%) | |
| Müller et al. [59] | 2004 | qMSP | No | Training Set CRC (10) Healthy Control (13) Validation Set CRC (13) Healthy Control (13) | SFRP2 | Training Set; SFRP2 (90%) Validation Set; SFRP2 (77%) | Training Set; SFRP2 (77%) Validation Set; SFRP2 (77%) |
| Lenhard et al. [61] | 2005 | MSP | No | CRC (26) Adenoma (13) Hyperplastic Polyp (9) CIBD (9) Normal control (32) | HIC1 | CRC; HIC1 (42%) Adenoma; HIC1 (31%) | CRC + Adenoma; HIC1 (98%) |
| Chen et al. [62] | 2005 | MSP | No | CRC (94) Normal Control (198) | VIM | All Stages; VIM (46%) Stage I and II; VIM (43%) | VIM (90%) |
| Huang et al. [63] | 2007 | MSP | No | CRC (52) Adenoma (21) Hyperplastic Polyp (8) Ulcerative Colitis (6) Healthy Control (24) | HPP1, MGMT,SFRP2 | CRC; At least 1 gene (96%) Adenoma; At least 1 gene (71%) | CRC + Adenoma; At least 1 gene (96%) |
| Zhang et al. [64] | 2007 | MSP | No | CRC (29) Adenoma (7) Healthy Control (17) | SFRP1 | CRC + Adenoma; SFRP1 (89%) | CRC + Adenoma; SFRP1 (86%) |
| Wang et al. [65] | 2008 | qMSP | No | CRC (69) Adenoma (34) Hyperplastic Polyp (26) Healthy Control (30) | SFRP2 | CRC; SFRP2 (87%) Adenoma; SFRP2 (62%) Hyperplastic Polyp; SFRP2 (42%) | CRC + Adenoma; SFRP2 (93%) |
| Ahlquist et al. [54] | 2008 | OBT SDT | Yes | Total (3764) CRC (39) Adenoma (251) | SDT-1 SDT-2 | CRC + Adenoma Hemoccult (11%), HemoccultSensa (21%), SDT-1 (20%), SDT-2 (40%) | CRC + Adenoma; Hemoccult (98%), HemoccultSensa (97%), SDT-1 (96%) |
| Nagasaka et al. [69] | 2009 | Hi-SA | No | CRC (84) Adenoma (56) Hyperplastic Polyp (12) Without Neoplasms (113) Other Disease (31) | RASSF2, SFRP2 | CRC RASSF2 (27%), SFRP2 (31%) Adenoma RASSF2 (11%), SFRP2 (25%) | CRC + Adenoma; RASSF2 (95%), SFRP2 (92%) |
| Glöckner et al. [68] | 2009 | MSP | No | CRC (84) Adenoma (26) CF negative control (87) | TFPI2 | Training Set CRC; TFPI2 (89%) Validation Set CRC; TFPI2 (76%) Adenoma; TFPI2 (21%) | Training Set CRC; TFPI2 (79%) Validation Set CRC; TFPI2 (93%) Adenoma; TFPI2 (93%) |
| Hellebrekers et al. [70] | 2009 | qMSP | No | Set1 CRC (28) Healthy Control (45) Set2 CRC (47) Healthy Controls (30) | GATA4 GATA5 | Set1; GATA4 (71%) Set2; GATA4 (51%) | Set1; GATA4 (84%) Set2; GATA4 (93%) |
| Melotte et al. [71] | 2009 | qMSP | No | Training Set CRC (28) CF Negative Control (45) Validation Set CRC (47) CF Negative Control (30) | NDRG4 | Training Set; NDRG4 (61%) Validation Set; NDRG4 (53%) | Training Set; NDRG4 (93%) Validation Set; NDRG4 (100%) |
| Baek et al. [66] | 2009 | MSP | No | CRC (60) Adenoma (52) CF Negative (37) | MGMT, hMLH1, VIM | CRC; At least 1 gene (75%) Adenoma; At least 1 gene (60%) | CRC + Adenoma; MGMT (86%), hMLH1 (100%), VIM (100%) |
| Kim et al. [67] | 2009 | qMSP | No | CRC (20) Adenoma (17) CF Normal (15) | B4GALT, OSMR, SFRP1 | CRC; OSMR + SFRP (60%) Adenoma; OSMR + SFRP1 (35%) | CRC + Adenoma; OSMR + SFRP1 (100%) |
| Ahlquist et al. [72] | 2012 | SDT | No | CRC (252) Adenoma (133) CF Negative Control (293) | BMP3, NDRG4, TFPI2, VIM, kras (mutation) | CRC; SDT (85%) Adenoma (>1 cm); SDT (63%) | CRC; SDT (89%) Adenoma (>1 cm); SDT (89%) |
| Imperiale et al. [73] | 2014 | SDT | Yes | Total (9989) CRC (65) | BMP3, NDRG4, kras (mutation) | SDT (92%), FIT (74%) | SDT (90%), FIT (96%) |
| Zhang et al. [74] | 2014 | MSP | No | CRC (48) Adenoma (35) Hyperplastic Polyp (32) Healthy Control (30) | SFRP2, WIF-1 | CRC; SFRP2 + WIF-1 (81%) Adenoma; SFRP2 + WIF-1 (81%) | CRC + Adenoma; SFRP2 + WIF-1 (97%) |
| Author | Year | Method | Prospective Study | Sample Size (Number of Patients) | Gene | Sensitivity (%) | Specificity (%) |
|---|---|---|---|---|---|---|---|
| Chan et al. [76] | 2002 | MSP | No | BC (22) Normal Control (17) | DAPK, E-cadherin, p16, RARβ | At least 1 gene (91%) | At least 1 gene (77%) |
| Chan et al. [77] | 2003 | MSP | No | BC (14) Normal control (10) | RASSF1A | RASSF1A (50%) | RASSF1A (100%) |
| Sathyanarayana et al. [79] | 2004 | MSP | No | BC (71) Bladder Wash (28) Voided Urine (43) None Malignant (6) | LAMA3, LAMB3, LAMC2 | At least 1 gene (49%) | At least 1 gene (100%) |
| Friedrich et al. [78] | 2004 | qMSP | No | BC (37) Normal Control (20) | BCL2, DAPK, TERT | BCL2 (65%), DAPK (22%), TERT (51%) | At least 1 gene (100%) |
| Dulaimi et al. [80] | 2004 | MSP | No | BC (45) normal (12) Inflammatory Urinary Disease (9) | APC, p14, RASSF1A | At least 1 gene (87%) | At least 1 gene (100%) |
| Hoque et al. [82] | 2006 | qMSP | No | BC (175) Normal Control (94) | ARF, GSTP1, MGMT, p16 | At least 1 gene (69%) | At least 1 gene (100%) |
| Urakami S et al. [83] | 2006 | MSP | No | BC (24) Normal Control (20) | DKK3, SFRP1, SFRP2, SFRP4, SFRP5, WIF1 | At least 1 gene (61%) | At least 1 gene (93%) |
| Yates et al. [81] | 2006 | qMSP | Yes | BC (35) Benign Control (35) Healthy Volunteer (34) | APC, DAPK, E-cadherin, GSTP1, p14, p16, RARB, RASSF1A | APC + E-cadherin + RASSF1A (69%) | APC + E-cadherin + RASSF1A (60%) |
| Yu et al. [84] | 2007 | MSP | No | BC (132) Normal Control (7) Noncancerous Urinary Lesion (23) Other disease(6) | ABCC6, ALX4, BCL2, BMP3, BRCA1, CCNA1, CDH13, CFTR, DRM, HPR1, ITGA4, MINT1, MTA1, MYOD1, RASSF1A, RPRM, RUNX3, SALL3 | Combination of 11 genes (92%) | Combination of 11 genes (87%) |
| Sun et al. [85] | 2009 | MSP | No | BC (82) Noncancerous Urinary Lesion (15) Normal Control (5) | CDH1, FANCF, LOXL1, LOXL4, p16, SFRP1, SOX9, TIG1, TIMP3, XAF1 | LOXL1 (40%), SFRP1 (37%), XAF1 (71%) | LOXL1 (73%), SFRP1 (93%), XAF1 (33%) |
| Lin et al. [87] | 2010 | MSP | No | BC (57) Normal Control (20) | E-cadherin, p14, p16, RASSF1A | At least 1 gene (83%) | |
| Renard et al. [86] | 2010 | qMSP | Yes | Symptomatic patients Training Set BC (48) Normal Control (121) Validation Set BC (35) Normal Control (57) | NID2, TWIST1 | Training Set; NID2 and TWIST1 (88%) Validation Set; NID2 and TWIST1 (94%) | Training Set; NID2 and TWIST1 (94%) Validation Set; NID2 and TWIST1 (91%) |
| Reinert et al. [89] | 2011 | MS-HRM | No | BC (115) BPH or Bladder Stone (59) | EOMES, HOXA9, POU4F2, ZNF154 | At least 3 genes (84%) | At least 3 genes (96%) |
| Eissa et al. [88] | 2011 | MSP | No | BC (210) Benign Urological Disease (61) Normal Control (49) | APC, RARβ2 | APC (60%), RARβ2 (63%) | APC (84%), RARβ2 (95%) |
| Chen et al. [91] | 2011 | qMSP | No | BC (30) None Cancer Control (19) | DAPK, IRF8, p14, RASSF1A, SFRP1 | IRF8 (57%), p14 (28%), SFRP1 (41%) | IRF8 (95%), p14 (100%), SFRP1 (100%) |
| Vinci et al. [90] | 2011 | qMSP | Yes | Bladder cancer (108) Control (105) BPH (29) Urinary tract infection (17) Bladder Stone (16) Normal Volunteer(43) | BCL2, DAPK, hTERT | At least 1 gene (79%) | At least 1 gene (90%) |
| Serizawa et al. [92] | 2011 | qMSP | No | BC (113) Normal Control (33) | APC, DBC1, RARB, RASSF1A, SFRP1, SFRP2, SFRP4, SFRP5 FGFR (mutation), PIK3CA (mutation), RAS (mutation), TP53 (mutation) | Total (70%) | Total (94%) |
| Chung et al. [94] | 2011 | qMSP | No | BC (128) None Cancer Control (110) | A2BP1, CA10, DBC1, MYO3A, NKX6-2, NPTX2, PENK, SOX11 | CA10 + MYO3A + NKX6-2 + SOX11 (81%) | CA10 + MYO3A + NKX6-2 + SOX11 (97%) |
| Costa et al. [93] | 2011 | qMSP | No | BC (50) RCC (50) PC (50) Healthy Control (48) | PCDH17, TCF21 | BC; PCDH17 + TCF21 (60%) RCC; PCDH17 + TCF21 (32%) PC; PCDH17 + TCF21 (26%) | BC; PCDH17 + TCF21 (100%) RCC; PCDH17 + TCF21 (100%) PC; PCDH17 + TCF21 (100%) |
| Reinert et al. [95] | 2012 | qMSP | No | BC (184) BPH or Bladder Stone (35) | EOMES, HOXA9, POU4F2, TWIST1, VIM, ZNF154 | EOMES (88%), TWIST1 (88%), VIM (89%) | EOMES (97%), TWIST1 (100%), VIM (100%) |
| Chihara et al. [96] | 2013 | Pyrosequencing | No | BC (73) Healthy Volunteer (18) | Hypermethylation; HOXA9_1, HOXA9_2, MYOD, SOX1, TJP2 Hypomethylation; CAPG, CASP8, HLADPA1, IFNG, RIPK3, SPP1, VAMP8 | HOXA9_1 (86%), HOXA9_2 (86%), MYOD (87%), TJP2 (93%) | HOXA9_1 (89%), HOXA9_2 (62%), MYOD (88%), TJP2 (56%) |
| Abern et al. [98] | 2014 | qMSP | Yes | Hematuria or on surveillance for prior NMIBC Total (111) BC (24) None Cancer Control (87) | NID2, TWIST1 | NID2, TWIST1 (75%) | NID2, TWIST1 (71%) |
| Hayashi et al. [97] | 2014 | qMSP | No | BC (20) Normal Control (20) | VGF | VGF (40%) | VGF (95%) |
| Fantony et al. [100] | 2015 | qMSP | Yes | Hematuria or on surveillance for prior NMIBC, or NMIBC treated with BCG Total (209) BC (52) Suspicious of BC (12) Negative for Cystoscopy (145) | NID2, TWIST1 | Believe the positive (67%) Believe the negative (37%) | Believe the positive (61%) Believe the negative (86%) |
| Yeh et al. [99] | 2015 | qMSP | No | Training set BC (69) None Cancer Control (28) Test set BC (33) None Cancer Control (28) | ZNF671 | Training Set; ZNF671 (42%) Test Set; ZNF671 (48%) | Training set; ZNF671 (93%) Test set; ZNF671 (89%) |
| Dahmcke et al. [102] | 2016 | qMSP | Yes | Hematuria Total (475) BC (99) None Cancer Control (376) | BCL2, CCNA1, EOMES, ONECUT2, SALL3, VIM, FGFR (mutation), TERT (mutation) | Total (97%) | Total (77%) |
| Roperch et al. [101] | 2016 | qMSP | No | BC (167) None Cancer Control (105) | HS3ST2, SEPT9, SLIT2, FGFR3 (mutation) | Total (Methylation + Mutation) (98%) | Total (Methylation + Mutation) (85%) |
| Author | Year | Method | Prospective Study | Sample Size (Number of Patients) | Gene | Sensitivity (%) | Specificity (%) |
|---|---|---|---|---|---|---|---|
| Goessl et al. [105] | 2000 | MSP | No | PC (33) BPH (26) | GSTP1 | GSTP1 (36%) | GSTP1 (100%) |
| Goessl et al. [108] | 2001 | MSP | No | PC (29) BPH (40) | GSTP1 | GSTP1 (77%) | GSTP1 (97%) |
| Goessl et al. [107] | 2001 | MSP | No | PC (40) PIN (7) BPH (45) | GSTP1 | PC; GSTP1 (73%) PIN; GSTP1 (29%) | PC and PIN; GSTP1 (98%) |
| Cairns et al. [106] | 2001 | MSP | No | PC (22) | GSTP1 | GSTP1 (27%) | |
| Jeronimo et al. [109] | 2002 | MSP qMSP | No | PC (69) BPH (31) | GSTP1 | qMSP; GSTP1 (19%) MSP; GSTP1 (30%) | qMSP and MSP; GSTP1 (67%) |
| Hoque et al. [110] | 2005 | qMSP | No | PC (52) None Cancer Control (91) | APC, ARF, E-cadherin, GSTP1, MGMT, p16, RARβ2, RASSF1A, TIMP3 | At least 1 of 4 genes (87%) | At least 1 of 4 genes (100%) |
| Roupret et al. [111] | 2007 | qMSP | No | PC (95) None Cancer Control (38) | APC, CDH1, DAPK, GSTP1, MGMT, p14, p16, RARβ2, RASSF1A, TIMP3 | At least 1 of 4 genes (89%) | At least 1 of 4 genes (89%) |
| Venar et al. [114] | 2008 | qMSP | Yes | PSA > 2.5 ng/mL Biopsy Positive (111) Biopsy Negative (123) | APC, GSTP1, RARβ2 | Cohort1; At least 1 gene (55%) Cohort2; At least 1 gene (53%) | Cohort1; At least 1 gene (80%) Cohort2; At least 1 gene (76%) |
| Baden et al. [112] | 2009 | qMSP | Yes | PSA 2–10 ng/mL PC(178) None Cancer Control (159) | APC, GSTP1, RARβ2 | At least 1 Gene (60%) | At least 1 Gene (81%) |
| Daniunaite et al. [113] | 2014 | qMSP | No | PC (253) BPH (32) | GSTP1, RARB, RASSF1 | GSTP1 (11%), RARB (29%), RASSF1 (45%) | GSTP1 (97%), RARB (81%), RASSF1 (84%) |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yokoi, K.; Yamashita, K.; Watanabe, M. Analysis of DNA Methylation Status in Bodily Fluids for Early Detection of Cancer. Int. J. Mol. Sci. 2017, 18, 735. https://doi.org/10.3390/ijms18040735
Yokoi K, Yamashita K, Watanabe M. Analysis of DNA Methylation Status in Bodily Fluids for Early Detection of Cancer. International Journal of Molecular Sciences. 2017; 18(4):735. https://doi.org/10.3390/ijms18040735
Chicago/Turabian StyleYokoi, Keigo, Keishi Yamashita, and Masahiko Watanabe. 2017. "Analysis of DNA Methylation Status in Bodily Fluids for Early Detection of Cancer" International Journal of Molecular Sciences 18, no. 4: 735. https://doi.org/10.3390/ijms18040735
APA StyleYokoi, K., Yamashita, K., & Watanabe, M. (2017). Analysis of DNA Methylation Status in Bodily Fluids for Early Detection of Cancer. International Journal of Molecular Sciences, 18(4), 735. https://doi.org/10.3390/ijms18040735






