Elucidating the Diversity of Aquatic Microdochium and Trichoderma Species and Their Activity against the Fish Pathogen Saprolegnia diclina
Abstract
:1. Introduction
2. Results and Discussion
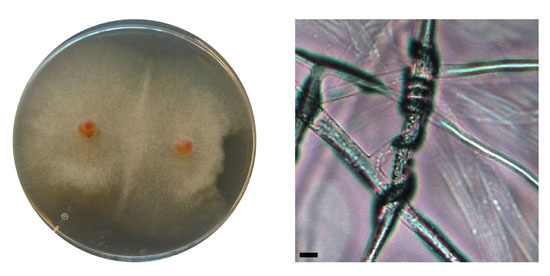
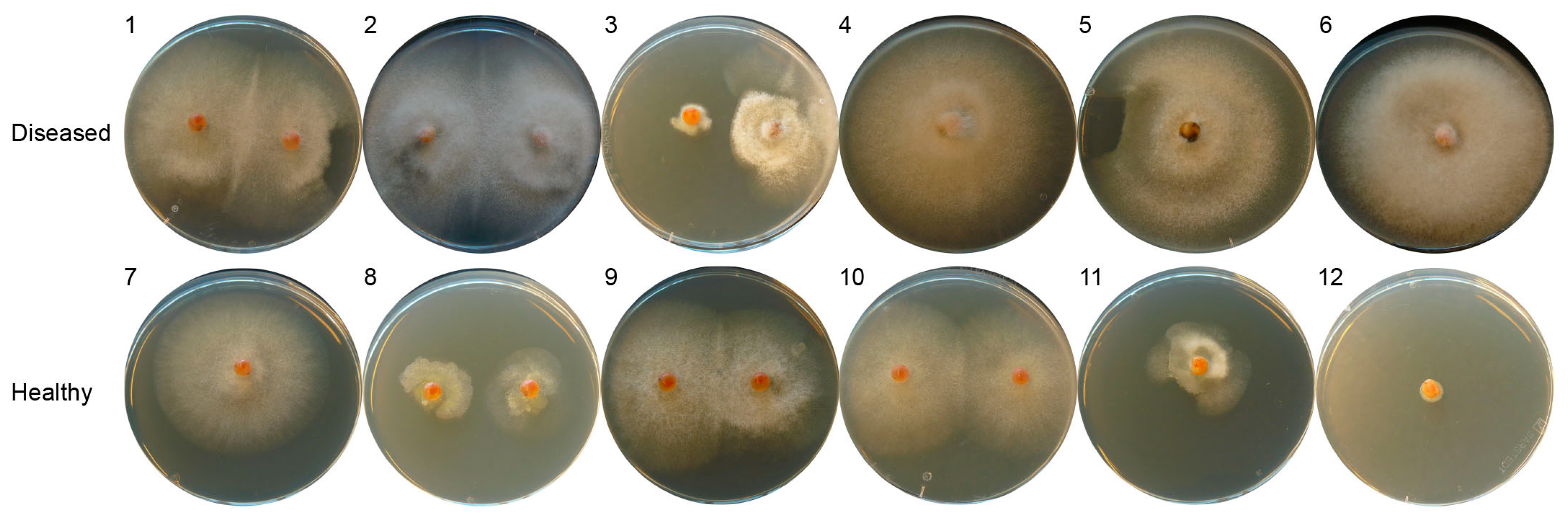
2.1. Isolation of Fungi from Diseased and Healthy Salmon Eggs
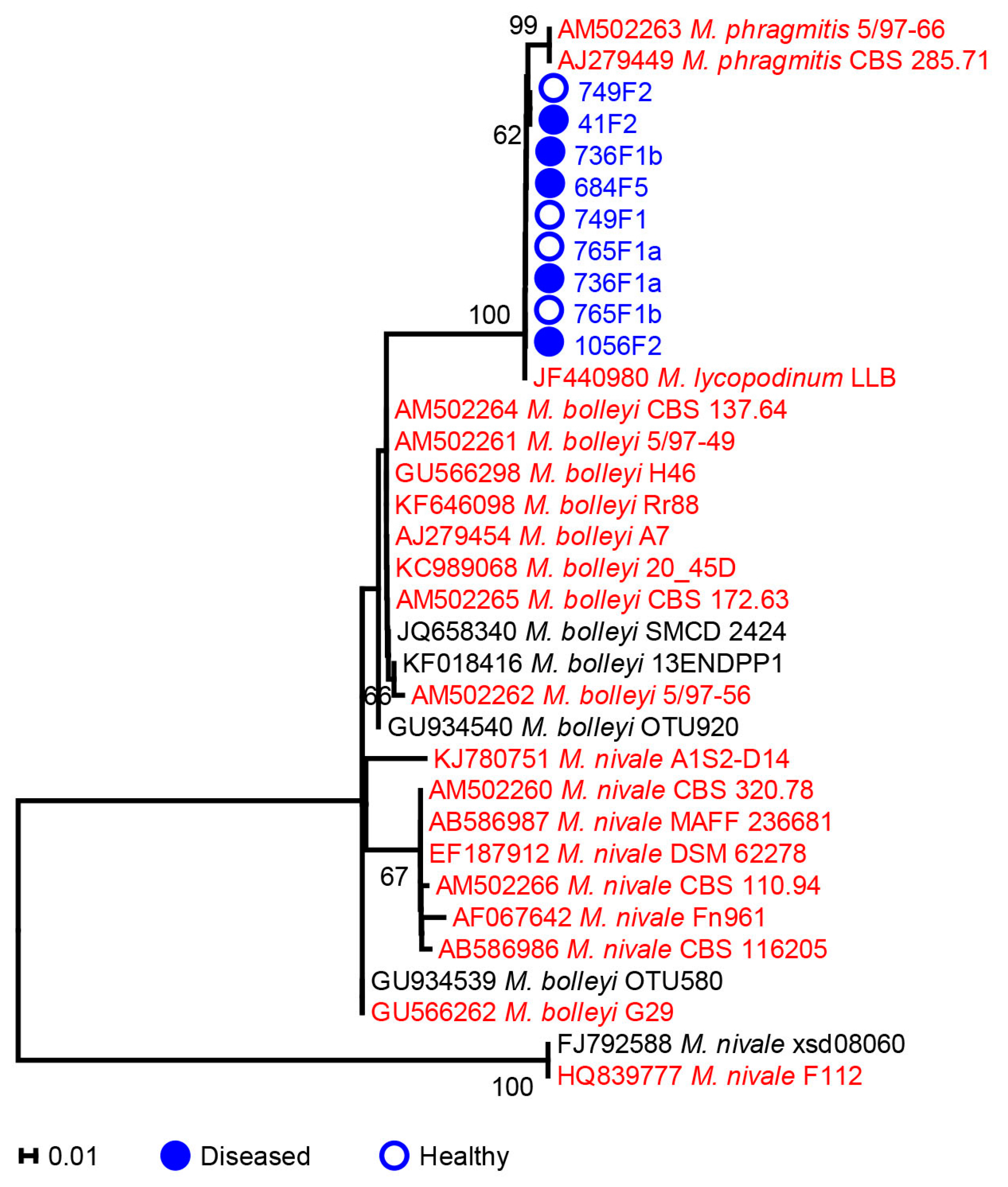
2.2. Microdochium
| Genus | Strain No. | Salmon Egg Sample | Activity of Culture Filtrate | Dual Culture Assay on 1/5PDA | Hyphal Interaction with S. diclina Microscopically |
|---|---|---|---|---|---|
| Microdochium | 41F2 | Diseased | Not inhibitory | Not inhibitory | Not observed |
| 684F5 | Diseased | Not inhibitory | Not inhibitory | Not observed | |
| 736F1a | Diseased | Not inhibitory | Not inhibitory | Not observed | |
| 736F1b | Diseased | Not inhibitory | Not inhibitory | Not observed | |
| 1056F2 | Diseased | Not inhibitory | Not inhibitory | Not observed | |
| 765F1a | Healthy | Not inhibitory | Not inhibitory | Not observed | |
| 765F1b | Healthy | Not inhibitory | Not inhibitory | Not observed | |
| 749F1 | Healthy | Not inhibitory | Inhibitory | Not observed | |
| 749F2 | Healthy | Not inhibitory | Not inhibitory | Not observed | |
| Trichoderma | 684F1 | Diseased | Not inhibitory | Not inhibitory | Coiling, papilla-like structure |
| 1056F1 | Diseased | Not inhibitory | Not inhibitory | Papilla-like structure | |
| 1152F1 | Diseased | Not inhibitory | Not inhibitory | Inconclusive | |
| 762F1a | Healthy | Not inhibitory | Not inhibitory | Coiling | |
| 762F1b | Healthy | Not inhibitory | Not inhibitory | Coiling | |
| 762F2 | Healthy | Not inhibitory | Not inhibitory | Papilla-like structure | |
| 764F1 | Healthy | Inhibitory | Not inhibitory | Papilla-like structure | |
| 764F2 | Healthy | Not inhibitory | Not inhibitory | Coiling, papilla-like structure | |
| 764F3 | Healthy | Not inhibitory | Not inhibitory | Coiling, papilla-like structure |
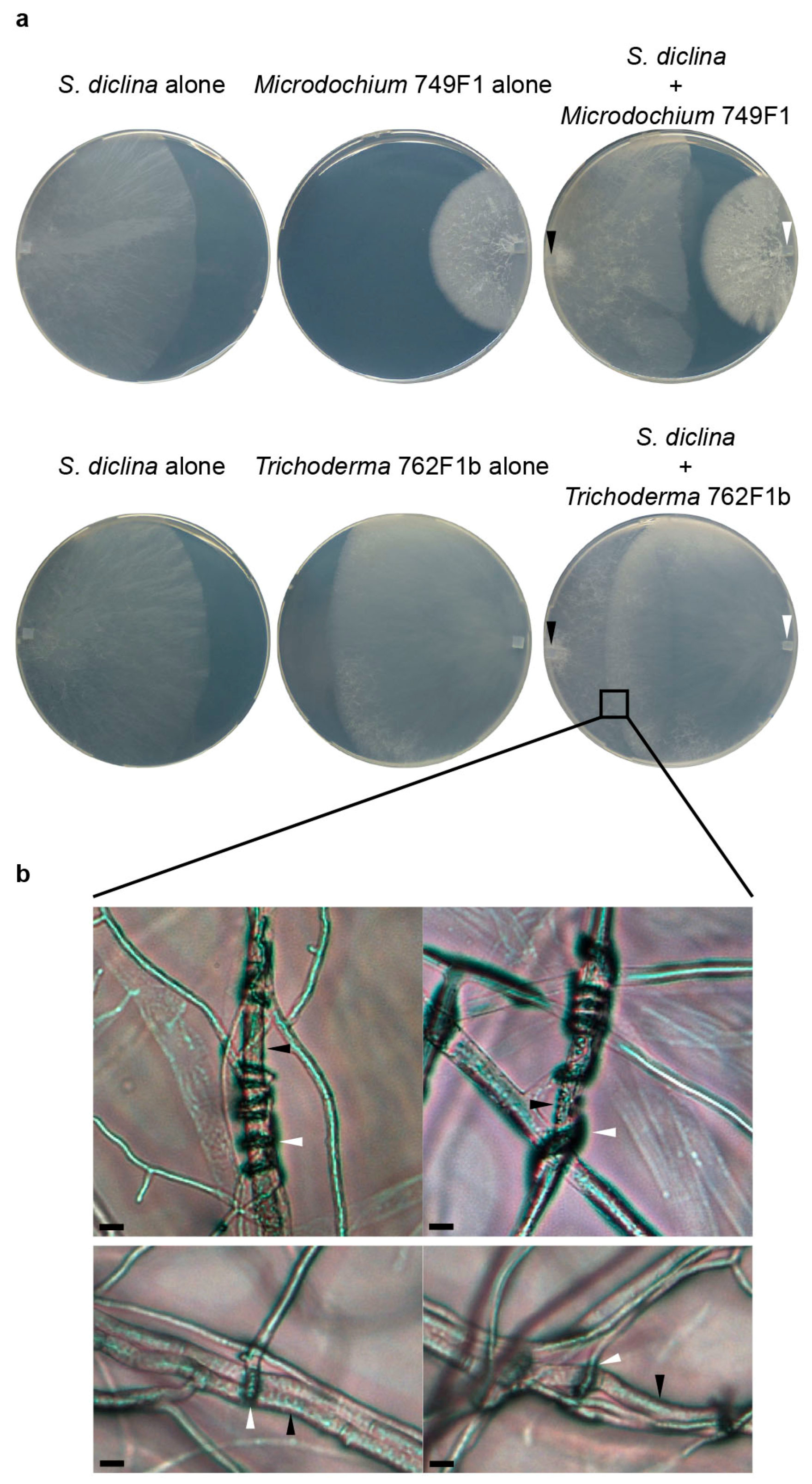
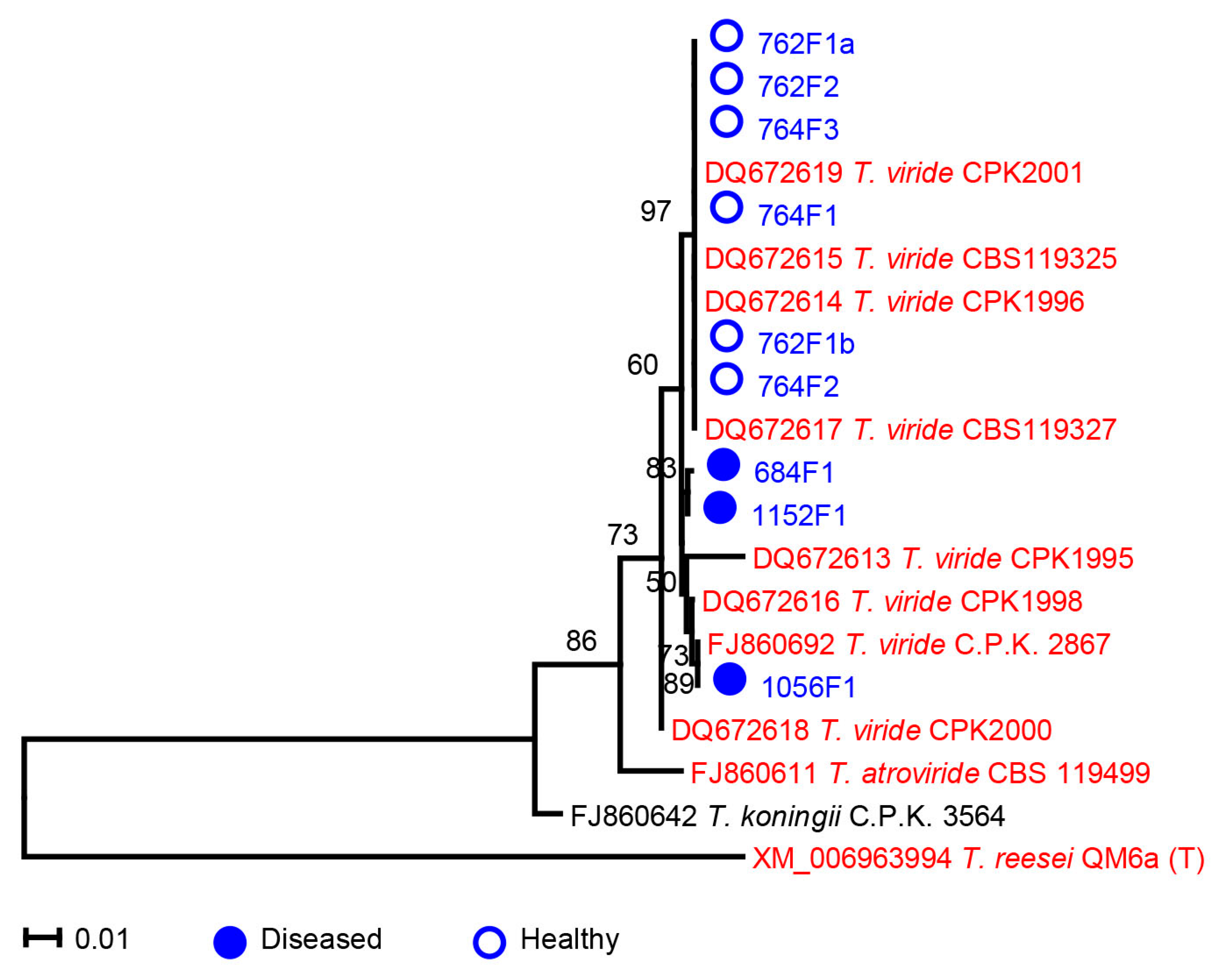
2.3. Trichoderma
3. Experimental Section
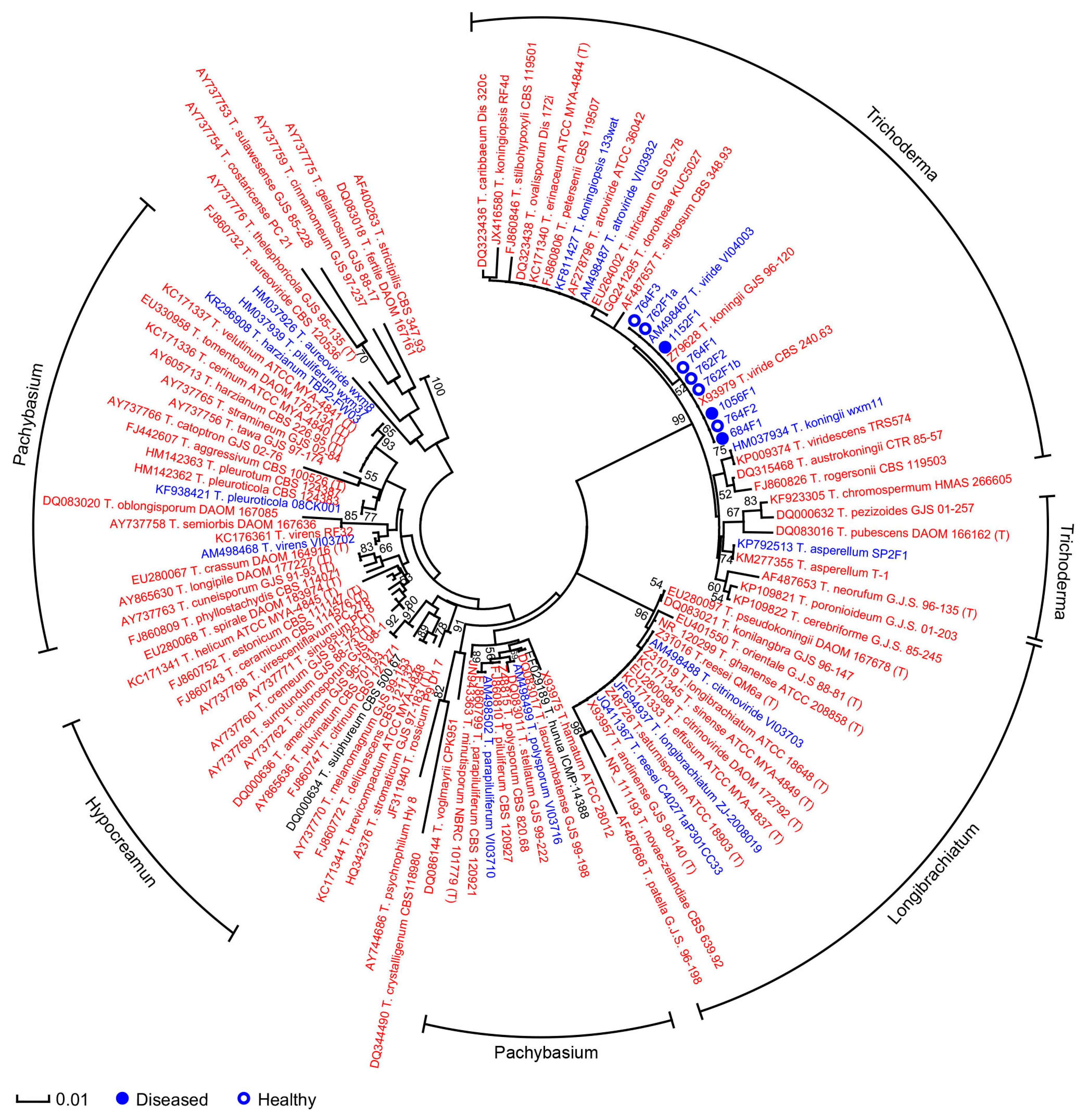
3.1. Phylogenetic Analysis of Microdochium and Trichoderma Isolates
3.2. Culture Filtrate Activity of Microdochium and Trichoderma Isolates
3.3. Dual Culture Assay
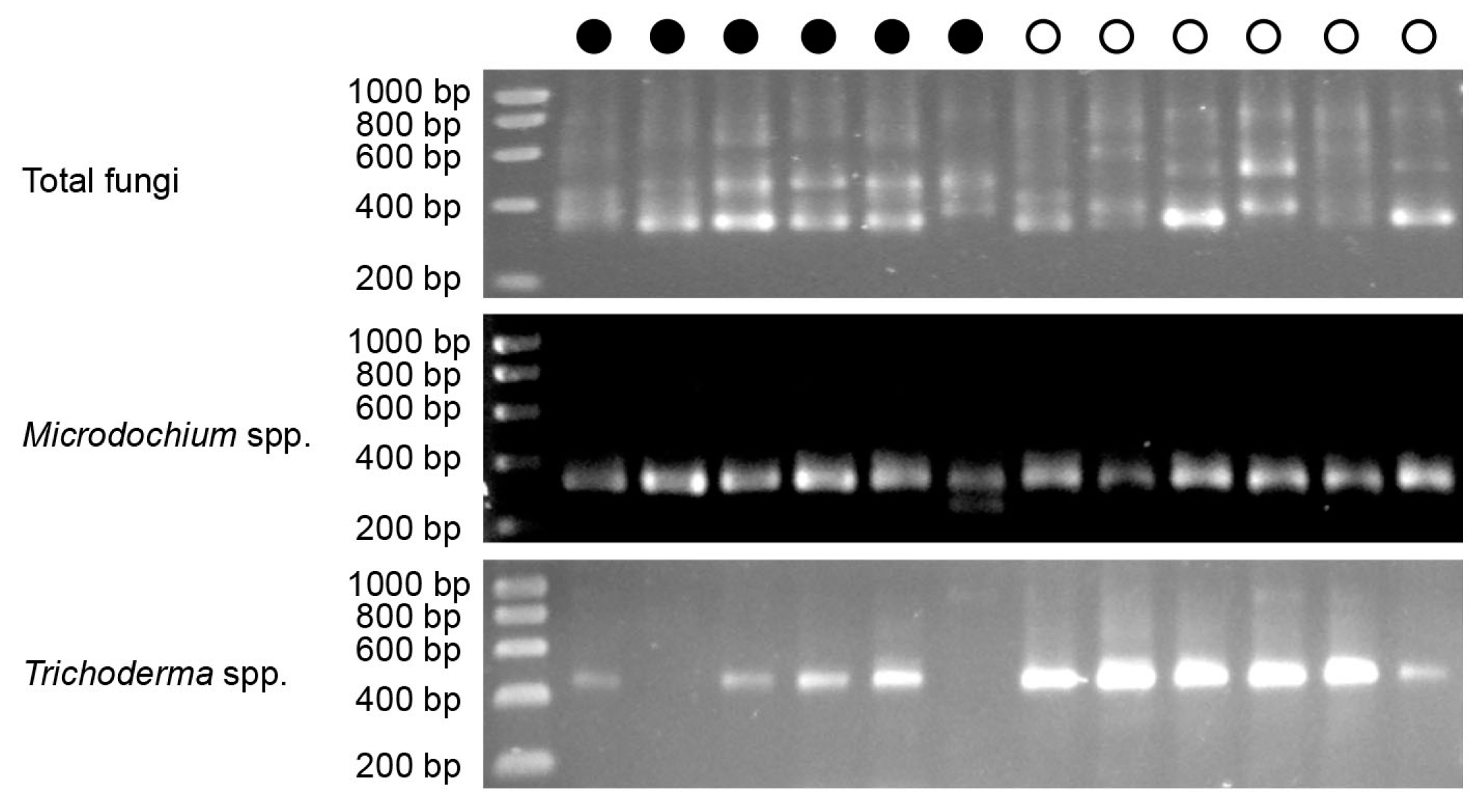
3.4. Quantification of Total Fungi, Microdochium and Trichoderma in Salmon Egg Incubation Water
3.5. Nucleotide Sequence Accession Numbers
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bruno, D.; van West, P.; Beakes, G. Saprolegnia and other oomycetes. In Fish Diseases and Disorders, Viral, Bacterial and Fungal Infections; Woo, P., Bruno, D., Eds.; CABI: Wallingford, UK, 2011; Volume 3, pp. 669–720. [Google Scholar]
- Van West, P. Saprolegnia parasitica, an oomycete pathogen with a fishy appetite: New challenges for an old problem. Mycologist 2006, 20, 99–104. [Google Scholar] [CrossRef]
- Hatai, K.; Hoshiai, G. Mass mortality in cultured coho salmon (Oncorhynchus kisutch) due to Saprolegnia parasitica coker. J. Wildl. Dis. 1992, 28, 532–536. [Google Scholar] [CrossRef] [PubMed]
- Hatai, K.; Hoshiai, G.-I. Characteristics of two Saprolegnia species isolated from coho salmon with Saprolegniosis. J. Aquat. Anim. Health 1993, 5, 115–118. [Google Scholar] [CrossRef]
- Van den Berg, A.H.; McLaggan, D.; Diéguez-Uribeondo, J.; van West, P. The impact of the water moulds Saprolegnia diclina and Saprolegnia parasitica on natural ecosystems and the aquaculture industry. Fungal Biol. Rev. 2013, 27, 33–42. [Google Scholar] [CrossRef]
- Irianto, A.; Austin, B. Probiotics in aquaculture. J. Fish Dis. 2002, 25, 633–642. [Google Scholar] [CrossRef]
- Chang, C.I.; Liu, W.Y. An evaluation of two probiotic bacterial strains, Enterococcus faecium SF68 and Bacillus toyoi, for reducing edwardsiellosis in cultured European eel, Anguilla anguilla L. J. Fish Dis. 2002, 25, 311–315. [Google Scholar] [CrossRef]
- Taoka, Y.; Maeda, H.; Jo, J.-Y.; Jeon, M.-J.; Bai, S.C.; Lee, W.-J.; Yuge, K.; Koshio, S. Growth, stress tolerance and non-specific immune response of Japanese flounder Paralichthys olivaceus to probiotics in a closed recirculating system. Fish. Sci. 2006, 72, 310–321. [Google Scholar] [CrossRef]
- Gatesoupe, F.-J. Updating the importance of lactic acid bacteria in fish farming: Natural occurrence and probiotic treatments. J. Mol. Microbiol. Biotechnol. 2008, 14, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Lara-Flores, M.; Olvera-Novoa, M.A.; Guzmán-Méndez, B.E.; López-Madrid, W. Use of the bacteria Streptococcus faecium and Lactobacillus acidophilus, and the yeast Saccharomyces cerevisiae as growth promoters in Nile tilapia (Oreochromis niloticus). Aquaculture 2003, 216, 193–201. [Google Scholar] [CrossRef]
- Hatai, K.; Willoughby, L.G. Saprolegnia parasitica from rainbow trout inhibited by the bacterium Pseudomonas fluorescens. Bull. Eur. Ass. Fish Pathol. 1988, 8, 27–29. [Google Scholar]
- Bly, J.E.; Quiniou, S.M.A.; Lawson, L.A.; Clem, L.W. Inhibition of Saprolegnia pathogenic for fish by Pseudomonas fluorescens. J. Fish Dis. 1997, 20, 35–40. [Google Scholar] [CrossRef]
- Hussein, M.M.A.; Hatai, K. In vitro inhibition of Saprolegnia by bacteria isolated from lesions of salmonids with saprolegniasis. Fish Pathol. 2001, 36, 73–78. [Google Scholar] [CrossRef]
- Lategan, M.J.; Gibson, L.F. Antagonistic activity of Aeromonas media strain A199 against Saprolegnia sp., an opportunistic pathogen of the eel, Anguilla australis Richardson. J. Fish Dis. 2003, 26, 147–153. [Google Scholar] [CrossRef] [PubMed]
- Lategan, M.J.; Torpy, F.R.; Gibson, L.F. Biocontrol of saprolegniosis in silver perch Bidyanus bidyanus (Mitchell) by Aeromonas media strain A199. Aquaculture 2004, 235, 77–88. [Google Scholar] [CrossRef]
- Lategan, M.J.; Torpy, F.R.; Gibson, L.F. Control of saprolegniosis in the eel Anguilla australis Richardson, by Aeromonas media strain A199. Aquaculture 2004, 240, 19–27. [Google Scholar] [CrossRef]
- Carbajal-González, M.T.; Fregeneda-Grandes, J.M.; Suárez-Ramos, S.; Rodríguez-Cadenas, F.; Aller-Gancedo, J.M. Bacterial skin flora variation and in vitro inhibitory activity against Saprolegnia parasitica in brown and rainbow trout. Dis. Aquat. Org. 2011, 96, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; de Bruijn, I.; Jack, A.L.H.; Drynan, K.; van den Berg, A.H.; Thoen, E.; Sandoval-Sierra, V.; Skaar, I.; van West, P.; Dieguez-Uribeondo, J.; et al. Deciphering microbial landscapes of fish eggs to mitigate emerging diseases. ISME J. 2014, 8, 2002–2014. [Google Scholar] [CrossRef] [PubMed]
- Campbell, W.A. A new species of Coniothyrium parasitic on sclerotia. Mycologia 1947, 39, 190–195. [Google Scholar] [CrossRef]
- Sandys-Winsch, C.; Whipps, J.M.; Gerlagh, M.; Kruse, M. World distribution of the sclerotial mycoparasite Coniothyrium minitans. Mycol. Res. 1993, 97, 1175–1178. [Google Scholar] [CrossRef]
- Vargas Gil, S.; Pastor, S.; March, G.J. Quantitative isolation of biocontrol agents Trichoderma spp., Gliocladium spp. and actinomycetes from soil with culture media. Microbiol. Res. 2009, 164, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Kasa, P.; Modugapalem, H.; Battini, K. Isolation, screening, and molecular characterization of plant growth promoting rhizobacteria isolates of Azotobacter and Trichoderma and their beneficial activities. J. Nat. Sci. Biol. Med. 2015, 6, 360–363. [Google Scholar] [PubMed]
- Lopez, L.L.M.A.; Aganon, C.P.; Juico, P.P. Isolation of Trichoderma species from carabao manure and evaluation of its beneficial uses. Int. J. Sci. Technol. Res. 2014, 3, 190–199. [Google Scholar]
- Mulaw, T.B.; Druzhinina, I.S.; Kubicek, C.P.; Atanasova, L. Novel endophytic Trichoderma spp. isolated from healthy Coffea arabica roots are capable of controlling coffee Tracheomycosis. Diversity 2013, 5, 750–766. [Google Scholar] [CrossRef]
- Sallenave-Namont, C.; Pouchus, Y.F.; Robiou du Pont, T.; Lassus, P.; Verbist, J.F. Toxigenic saprophytic fungi in marine shellfish farming areas. Mycopathologia 2000, 149, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Druzhinina, I.S.; Seidl-Seiboth, V.; Herrera-Estrella, A.; Horwitz, B.A.; Kenerley, C.M.; Monte, E.; Mukherjee, P.K.; Zeilinger, S.; Grigoriev, I.V.; Kubicek, C.P. Trichoderma: The genomics of opportunistic success. Nat. Rev. Microbiol. 2011, 9, 749–759. [Google Scholar] [CrossRef] [PubMed]
- Wolken, W.A.M.; Tramper, J.; van der Werf, M.J. What can spores do for us? Trends Biotechnol. 2003, 21, 338–345. [Google Scholar] [CrossRef]
- De Vrije, T.; Antoine, N.; Buitelaar, R.M.; Bruckner, S.; Dissevelt, M.; Durand, A.; Gerlagh, M.; Jones, E.E.; Luth, P.; Oostra, J.; et al. The fungal biocontrol agent Coniothyrium minitans: Production by solid-state fermentation, application and marketing. Appl. Microbiol. Biotechnol. 2001, 56, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Kmet, V.; Flint, H.J.; Wallace, R.J. Probiotics and manipulation of rumen development and function. Arch. Anim. Nutri. 1993, 44, 1–10. [Google Scholar] [CrossRef]
- BioZyme, Inc. Available online: http://vitaferm.com/products/amaferm-digest-more/ (accessed on 4 December 2015).
- Abdelhamid, A.M.; Shabana, Y.M.; Gomaa, S.S.A. Aquatic Fungi and Fish Production in Egypt, II—In Vivo Studies. Available online: http://en.engormix.com/MA-aquaculture/articles/aquatic-fungi-fish-production-t460/MYC-p0.htm (accessed on 4 December 2015).
- Neospark. Available online: http://www.neospark.com/hetronex-abtp.html (accessed on 4 December 2015).
- Passarini, M.R.Z.; Santos, C.; Lima, N.; Berlinck, R.G.S.; Sette, L.D. Filamentous fungi from the Atlantic marine sponge Dragmacidon reticulatum. Arch. Microbiol. 2013, 195, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Dörr, A.J.M.; Elia, A.C.; Rodolfi, M.; Garzoli, L.; Picco, A.M.; D’Amen, M.; Scalici, M. A model of co-occurrence: Segregation and aggregation patterns in the mycoflora of the crayfish Procambarus clarkii in Lake Trasimeno (central Italy). J. Limnol. 2012, 71, 135–143. [Google Scholar] [CrossRef] [Green Version]
- Alford, D.V. Pest and Disease Management Handbook; WILEY: Hoboken, NJ, USA, 2008. [Google Scholar]
- Horst, R.K. Westcott’s Plant Disease Handbook; Springer: Berlin, Germany; Heidelberg, Germany, 2008. [Google Scholar]
- Cook, R.J. Fusarium diseases of wheat and other small grains in North America. In Fusarium Diseases, Biology, and Taxonomy; Nelson, P.E., Toussoun, T.A., Cook, R.J., Eds.; Pennsylvania State University Press: University Park, PA, USA, 1981; pp. 39–52. [Google Scholar]
- Simpson, D.R.; Rezanoor, H.N.; Parry, D.W.; Nicholson, P. Evidence for differential host preference in Microdochium nivale var. majus and Microdochium nivale var. nivale. Plant Pathol. 2000, 49, 261–268. [Google Scholar] [CrossRef]
- Hong, S.K.; Kim, W.G.; Choi, H.W.; Lee, S.Y. Identification of Microdochium bolleyi associated with basal rot of creeping bent grass in Korea. Mycobiology 2008, 36, 77–80. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, N. Snow molds: A group of fungi that prevail under snow. Microbes Environ. 2009, 24, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Rapacz, M.; Ergon, A.; Hoglind, M.; Jorgensen, M.; Jurczyk, B.; Ostrem, L.; Rognli, O.A.; Tronsmo, A.M. Overwintering of herbaceous plants in a changing climate. Still more questions than answers. Plant Sci. 2014, 225, 34–44. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Nicholson, P. Community ecology of fungal pathogens causing wheat head blight. Annu. Rev. Phytopathol. 2009, 47, 83–103. [Google Scholar] [CrossRef] [PubMed]
- Daamen, R.A.; Langerak, C.J.; Stol, W. Surveys of cereal diseases and pests in the netherlands. 3. Monographella nivalis and Fusarium spp. in winter-wheat fields and seed lots. Neth. J. Plant Pathol. 1991, 97, 105–114. [Google Scholar] [CrossRef]
- Al-Hashimi, M.H.; Perry, D.A. Fungal antagonists of Gerlachia nivalis. J. Phytopathol. 1986, 116, 106–112. [Google Scholar] [CrossRef]
- Ernst, M.; Neubert, K.; Mendgen, K.W.; Wirsel, S.G.R. Niche differentiation of two sympatric species of Microdochium colonizing the roots of common reed. BMC Microbiol. 2011, 11. [Google Scholar] [CrossRef] [PubMed]
- Jaklitsch, W.; Voglmayr, H. Phylogenetic relationships of five genera of Xylariales and Rosasphaeria gen. nov. (Hypocreales). Fungal Divers. 2012, 52, 75–98. [Google Scholar] [CrossRef]
- Berg, G.; Zachow, C.; Lottmann, J.; Gotz, M.; Costa, R.; Smalla, K. Impact of plant species and site on rhizosphere-associated fungi antagonistic to Verticillium dahliae kleb. Appl. Environ. Microbiol. 2005, 71, 4203–4213. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Bhosale, S.H.; Patil, K.B.; Parameswaran, P.S.; Naik, C.G.; Jagtap, T.G. Active pharmaceutical ingredient (api) from an estuarine fungus, Microdochium nivale (Fr.). J. Environ. Biol. 2011, 32, 653–658. [Google Scholar] [PubMed]
- Santiago, I.F.; Alves, T.M.A.; Rabello, A.; Sales Junior, P.A.; Romanha, A.J.; Zani, C.L.; Rosa, C.A.; Rosa, L.H. Leishmanicidal and antitumoral activities of endophytic fungi associated with the Antarctic angiosperms Deschampsia antarctica Desv. and Colobanthus quitensis (Kunth) Bartl. Extremophiles 2012, 16, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Schuster, A.; Schmoll, M. Biology and biotechnology of Trichoderma. Appl. Microbiol. Biotechnol. 2010, 87, 787–799. [Google Scholar] [CrossRef] [PubMed]
- Verma, M.; Brar, S.K.; Tyagi, R.D.; Surampalli, R.Y.; Valero, J.R. Antagonistic fungi, Trichoderma spp.: Panoply of biological control. Biochem. Eng. J. 2007, 37, 1–20. [Google Scholar] [CrossRef]
- Schirmbӧck, M.; Lorito, M.; Wang, Y.L.; Hayes, C.K.; Arisanatac, I.; Scala, F.; Harman, G.E.; Kubicek, C.P. Parallel formation and synergism of hydrolytic enzymes and peptaibol antibiotics, molecular mechanisms involved in the antagonistic action of Trichoderma harzianum against phytopathogenic fungi. Appl. Environ. Microbiol. 1994, 60, 4364–4370. [Google Scholar]
- Harman, G.E. Multifunctional fungal plant symbionts: New tools to enhance plant growth and productivity. New Phytol. 2011, 189, 647–649. [Google Scholar] [CrossRef] [PubMed]
- Harman, G.E.; Howell, C.R.; Viterbo, A.; Chet, I.; Lorito, M. Trichoderma species—Opportunistic, avirulent plant symbionts. Nat. Rev. Microbiol. 2004, 2, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Hageskal, G.; Vrålstad, T.; Knutsen, A.K.; Skaar, I.D.A. Exploring the species diversity of Trichoderma in Norwegian drinking water systems by DNA barcoding. Mol. Ecol. Resour. 2008, 8, 1178–1188. [Google Scholar] [CrossRef] [PubMed]
- Gal-Hemed, I.; Atanasova, L.; Komon-Zelazowska, M.; Druzhinina, I.S.; Viterbo, A.; Yarden, O. Marine isolates of Trichoderma spp. As potential halotolerant agents of biological control for arid-zone agriculture. Appl. Environ. Microbiol. 2011, 77, 5100–5109. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.; Oesker, V.; Wiese, J.; Schmaljohann, R.; Imhoff, J.F. Two new antibiotic pyridones produced by a marine fungus, Trichoderma sp. strain MF106. Mar. Drugs 2014, 12, 1208–1219. [Google Scholar] [CrossRef] [PubMed]
- Zhou, K.; Zhang, X.; Zhang, F.; Li, Z. Phylogenetically diverse cultivable fungal community and polyketide synthase (PKS), non-ribosomal peptide synthase (NRPS) genes associated with the South China Sea sponges. Microb. Ecol. 2011, 62, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Miao, L.; Qian, P.Y. Antagonistic antimicrobial activity of marine fungi and bacteria isolated from marine biofilm and seawaters of Hong Kong. Aquat. Microb. Ecol. 2005, 38, 231–238. [Google Scholar] [CrossRef]
- Yamazaki, H.; Saito, R.; Takahashi, O.; Kirikoshi, R.; Toraiwa, K.; Iwasaki, K.; Izumikawa, Y.; Nakayama, W.; Namikoshi, M. Trichoketides A and B, two new protein tyrosine phosphatase 1B inhibitors from the marine-derived fungus Trichoderma sp. J. Antibiot. 2015, 68, 628–632. [Google Scholar] [CrossRef] [PubMed]
- Yamada, T.; Umebayashi, Y.; Kawashima, M.; Sugiura, Y.; Kikuchi, T.; Tanaka, R. Determination of the chemical structures of Tandyukisins B–D, isolated from a marine sponge-derived fungus. Mar. Drugs 2015, 13, 3231–3240. [Google Scholar] [CrossRef] [PubMed]
- Citarasu, T. Natural antimicrobial compounds for use in aquaculture. In Infectious Disease in Aquaculture; Austin, B., Ed.; Woodhead Publishing: Cambridge, UK, 2012; pp. 419–456. [Google Scholar]
- Druzhinina, I.; Kopchinskiy, A. International Subcommission on Trichoderma and Hypocrea Taxonomy. Available online: http://isth.info/ (accessed on 4 December 2015).
- Jaklitsch, W.M.; Samuels, G.J.; Dodd, S.L.; Lu, B.S.; Druzhinina, I.S. Hypocrea rufa/Trichoderma viride: A reassessment, and description of five closely related species with and without warted conidia. Stud. Mycol. 2006, 56, 135–177. [Google Scholar] [CrossRef] [PubMed]
- Jaklitsch, W.M. European species of Hypocrea part II: Species with hyaline ascospores. Fungal Divers. 2011, 48, 1–250. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.X.; Tombolini, R.; Woo, S.; Zeilinger, S.; Lorito, M.; Jansson, J.K. In vivo study of Trichoderma-pathogen-plant interactions, using constitutive and inducible green fluorescent protein reporter systems. Appl. Environ. Microbiol. 2004, 70, 3073–3081. [Google Scholar] [CrossRef] [PubMed]
- Chacón, M.R.; Rodríguez-Galán, O.; Benítez, T.; Sousa, S.; Rey, M.; Llobell, A.; Delgado-Jarana, J. Microscopic and transcriptome analyses of early colonization of tomato roots by Trichoderma harzianum. Int. Microbiol. 2007, 10, 19–27. [Google Scholar] [PubMed]
- Rocha-Ramírez, V.; Omero, C.; Chet, I.; Horwitz, B.A.; Herrera-Estrella, A. Trichoderma atroviride G-protein alpha-subunit gene tga1 is involved in mycoparasitic coiling and conidiation. Eukaryot. Cell 2002, 1, 594–605. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K. Estimation of the number of nucleotide substitutions when there are strong transition-transversion and G + C-content biases. Mol. Biol. Evol. 1992, 9, 678–687. [Google Scholar] [PubMed]
- Tamura, K.; Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [PubMed]
- De Miguel, D.; Gomez, P.; Gonzalez, R.; Garcia-Suarez, J.; Cuadros, J.A.; Banas, M.H.; Romanyk, J.; Burgaleta, C. Nonfatal pulmonary Trichoderma viride infection in an adult patient with acute myeloid leukemia: Report of one case and review of the literature. Diagn. Microbiol. Infect. Dis. 2005, 53, 33–37. [Google Scholar] [CrossRef] [PubMed]
- Loeppky, C.B.; Sprouse, R.F.; Carlson, J.V.; Everett, E.D. Trichoderma viride peritonitis. South. Med. J. 1983, 76, 798–799. [Google Scholar] [CrossRef] [PubMed]
- Escudero Gil, M.R.; Pino, C.E.; Munoz, R. Pulmonary mycoma cause by Trichoderma viride. Actas Dermo-Sifiliogr. 1976, 67, 673–680. [Google Scholar]
- Jacobs, F.; Byl, B.; Bourgeois, N.; Coremans-Pelseneer, J.; Florquin, S.; Depre, G.; van de Stadt, J.; Adler, M.; Gelin, M.; Thys, J.P. Trichoderma viride infection in a liver transplant recipient. Mycoses 1992, 35, 301–303. [Google Scholar] [CrossRef] [PubMed]
- Rota, S.; Marchesi, D.; Farina, C.; de Bievre, C. Trichoderma pseudokoningii peritonitis in automated peritoneal dialysis patient successfully treated by early catheter removal. Perit. Dial. Int. 2000, 20, 91–93. [Google Scholar] [PubMed]
- Guiserix, J.; Ramdane, M.; Finielz, P.; Michault, A.; Rajaonarivelo, P. Trichoderma harzianum peritonitis in peritoneal dialysis. Nephron 1996, 74, 473–474. [Google Scholar] [CrossRef] [PubMed]
- Campos-Herrero, M.; Bordes, A.; Perera, A.; Ruiz, M.; Fernández, A. Trichoderma koningii peritonitis in a patient undergoing peritoneal dialysis. Clin. Microbiol. Newsl. 1996, 18, 150–152. [Google Scholar] [CrossRef]
- Tanis, B.C.; van der Pijl, H.; van Ogtrop, M.L.; Kibbelaar, R.E.; Chang, P.C. Fatal fungal peritonitis by Trichoderma longibrachiatum complicating peritoneal dialysis. Nephrol. Dial. Transplant. 1995, 10, 114–116. [Google Scholar] [PubMed]
- Richter, S.; Cormican, M.G.; Pfaller, M.A.; Lee, C.K.; Gingrich, R.; Rinaldi, M.G.; Sutton, D.A. Fatal disseminated Trichoderma longibrachiatum infection in an adult bone marrow transplant patient: Species identification and review of the literature. J. Clin. Microbiol. 1999, 37, 1154–1160. [Google Scholar] [PubMed]
- Gautheret, A.; Dromer, F.; Bourhis, J.H.; Andremont, A. Trichoderma pseudokoningii as a cause of fatal infection in a bone marrow transplant recipient. Clin. Infect. Dis. 1995, 20, 1063–1064. [Google Scholar] [CrossRef] [PubMed]
- Chouaki, T.; Lavarde, V.; Lachaud, L.; Raccurt, C.P.; Hennequin, C. Invasive infections due to Trichoderma species: Report of 2 cases, findings of in vitro susceptibility testing, and review of the literature. Clin. Infect. Dis. 2002, 35, 1360–1367. [Google Scholar] [CrossRef] [PubMed]
- Guarro, J.; Antolín-Ayala, M.I.; Gené, J.; Gutiérrez-Calzada, J.; Nieves-Díez, C.; Ortoneda, M. Fatal case of Trichoderma harzianum infection in a renal transplant recipient. J. Clin. Microbiol. 1999, 37, 3751–3755. [Google Scholar] [PubMed]
- Seguin, P.; Degeilh, B.; Grulois, I.; Gacouin, A.; Maugendre, S.; Dufour, T.; Dupont, B.; Camus, C. Successful treatment of a brain abscess due to Trichoderma longibrachiatum after surgical resection. Eur. J. Clin. Microbiol. Infect. Dis. 1995, 14, 445–448. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, H.; Kusne, S.; Sutton, D.A.; Manez, R.; Carrau, R.; Nichols, L.; Abu-Elmagd, K.; Skedros, D.; Todo, S.; Rinaldi, M.G. Acute invasive sinusitis due to Trichoderma longibrachiatum in a liver and small bowel transplant recipient. Clin. Infect. Dis. 1998, 26, 487–489. [Google Scholar] [CrossRef] [PubMed]
- Munoz, F.M.; Demmler, G.J.; Travis, W.R.; Ogden, A.K.; Rossmann, S.N.; Rinaldi, M.G. Trichoderma longibrachiatum infection in a pediatric patient with aplastic anemia. J. Clin. Microbiol. 1997, 35, 499–503. [Google Scholar] [PubMed]
- Myoken, Y.; Sugata, T.; Fujita, Y.; Asaoku, H.; Fujihara, M.; Mikami, Y. Fatal necrotizing stomatitis due to Trichoderma longibrachiatum in a neutropenic patient with malignant lymphoma: A case report. Int. J. Oral Maxillofac. Surg. 2002, 31, 688–691. [Google Scholar] [CrossRef] [PubMed]
- Marrouchi, R.; Benoit, E.; le Caer, J.-P.; Belayouni, N.; Belghith, H.; Molgó, J.; Kharrat, R. Toxic C17-sphinganine analogue mycotoxin, contaminating Tunisian mussels, causes flaccid paralysis in rodents. Mar. Drugs 2013, 11, 4724. [Google Scholar] [CrossRef] [PubMed]
- Cortinas, M.-N.; Crous, P.W.; Wingfield, B.D.; Wingfield, M.J. Multi-gene phylogenies and phenotypic characters distinguish two species within the Colletogloeopsis zuluensis complex associated with Eucalyptus stem cankers. Stud. Mycol. 2006, 55, 133–146. [Google Scholar] [CrossRef] [PubMed]
- Nagy, V.; Seidl, V.; Szakacs, G.; Komofl-Zelazowska, M.; Kubicek, C.P.; Druzhinina, I.S. Application of DNA bar codes for screening of industrially important fungi: The haplotype of Trichoderma harzianum sensu stricto indicates superior chitinase formation. Appl. Environ. Microbiol. 2007, 73, 7048–7058. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., White, T.J., Eds.; Academic Press: New York, NY, USA, 1990; pp. 315–322. [Google Scholar]
- Ihrmark, K.; Bödeker, I.T.M.; Cruz-Martinez, K.; Friberg, H.; Kubartova, A.; Schenck, J.; Strid, Y.; Stenlid, J.; Brandström-Durling, M.; Clemmenssen, K.E.; et al. New primers to amplify the fungal ITS2 region—Evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol. Ecol. 2012, 82, 666–677. [Google Scholar] [CrossRef] [PubMed]
- Meincke, R.; Weinert, N.; Radl, V.; Schloter, M.; Smalla, K.; Berg, G. Development of a molecular approach to describe the composition of Trichoderma communities. J. Microbiol. Methods 2010, 80, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Food and Agriculture Organization. The State of World Fisheries and Aquaculture 2014; Food and Agriculture Organization of the United Nations: Rome, Italy, 2014; p. 223. [Google Scholar]
- Woodhams, D.C.; Brandt, H.; Baumgartner, S.; Kielgast, J.; Küpfer, E.; Tobler, U.; Davis, L.R.; Schmidt, B.R.; Bel, C.; Hodel, S.; et al. Interacting symbionts and immunity in the amphibian skin mucosome predict disease risk and probiotic effectiveness. PLoS ONE 2014. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, Y.; Zachow, C.; Raaijmakers, J.M.; De Bruijn, I. Elucidating the Diversity of Aquatic Microdochium and Trichoderma Species and Their Activity against the Fish Pathogen Saprolegnia diclina. Int. J. Mol. Sci. 2016, 17, 140. https://doi.org/10.3390/ijms17010140
Liu Y, Zachow C, Raaijmakers JM, De Bruijn I. Elucidating the Diversity of Aquatic Microdochium and Trichoderma Species and Their Activity against the Fish Pathogen Saprolegnia diclina. International Journal of Molecular Sciences. 2016; 17(1):140. https://doi.org/10.3390/ijms17010140
Chicago/Turabian StyleLiu, Yiying, Christin Zachow, Jos M. Raaijmakers, and Irene De Bruijn. 2016. "Elucidating the Diversity of Aquatic Microdochium and Trichoderma Species and Their Activity against the Fish Pathogen Saprolegnia diclina" International Journal of Molecular Sciences 17, no. 1: 140. https://doi.org/10.3390/ijms17010140
APA StyleLiu, Y., Zachow, C., Raaijmakers, J. M., & De Bruijn, I. (2016). Elucidating the Diversity of Aquatic Microdochium and Trichoderma Species and Their Activity against the Fish Pathogen Saprolegnia diclina. International Journal of Molecular Sciences, 17(1), 140. https://doi.org/10.3390/ijms17010140