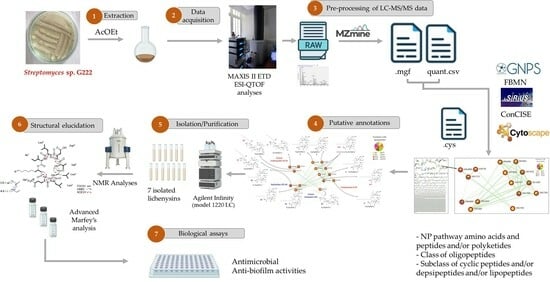
Computational Methods Reveal a Series of Cyclic and Linear Lichenysins and Surfactins from the Vietnamese Marine Sediment-Derived Streptomyces Strain G222
Abstract
1. Introduction
2. Results
2.1. Phylogenetic Affiliation of the G222 Bacteria Based on 16S rRNA
2.2. LC-ESI-MS/MS Analysis of the Streptomyces G222 Derived Marine Sponge Crude Extract and Detection of Lichenysin and Surfactin Analogues
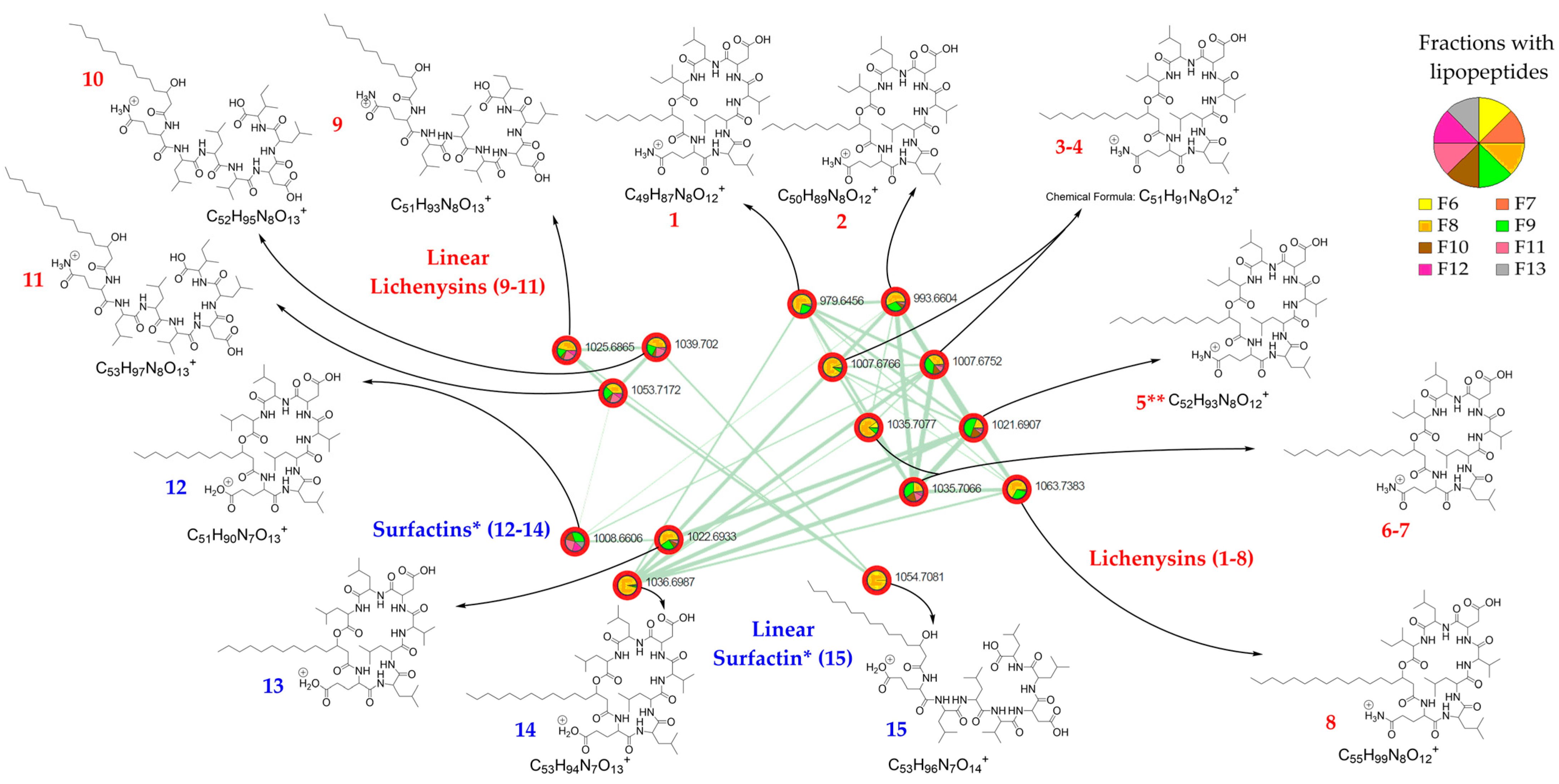
2.3. Characterization of Lichenysin Linear and Cyclic Lipopeptides: 1D and 2D NMR Spectroscopy

2.4. Absolute Configuration Using Modified Marfey’s Reagent of the Lichenysin Lipopeptides
2.5. Antibiofilm Activities of the Lichenysin Lipopeptides
2.6. Antimicrobial Activity
3. Discussion
4. Materials and Methods
4.1. General Experimental Procedures
4.2. Sampling and Strain Isolation
4.3. Isolation of Strain G222
4.4. Strain Identification Using 16S rRNA Gene Sequencing
4.5. Fermentation of the Streptomyces sp G222
4.6. Preparation of the Crude Extract
4.7. LC-MS2 Analyzes of the AcOEt Extract
4.8. Mass Spectrometry: LC-MS/MS Data Processing
4.9. Mass Spectrometry: Molecular Networking
4.10. ConCISE (Consensus Classifications of In Silico Elucidations)
4.11. Isolation and Purification of the Lichenysin Lipopetides
4.12. Advanced Marfey’s Analysis
4.13. Biofilm Formation Activity
4.14. Antimicrobial Assays
4.15. Real-Time Growth Curves
4.16. ATP Efflux Measurement
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhang, S.; Chen, Y.; Zhu, J.; Lu, Q.; Cryle, M.J.; Zhang, Y.; Yan, F. Structural Diversity, Biosynthesis, and Biological Functions of Lipopeptides from Streptomyces. Nat. Prod. Rep. 2023, 40, 557–594. [Google Scholar] [CrossRef]
- Jakubiec-Krzesniak, K.; Rajnisz-Mateusiak, A.; Guspiel, A.; Ziemska, J.; Solecka, J. Secondary Metabolites of Actinomycetes and Their Antibacterial, Antifungal and Antiviral Properties. Pol. J. Microbiol. 2018, 67, 259–272. [Google Scholar] [CrossRef]
- Schneemann, I.; Kajahn, I.; Ohlendorf, B.; Zinecker, H.; Erhard, A.; Nagel, K.; Wiese, J.; Imhoff, J.F. Mayamycin, a Cytotoxic Polyketide from a Streptomyces Strain Isolated from the Marine Sponge Halichondria panicea. J. Nat. Prod. 2010, 73, 1309–1312. [Google Scholar] [CrossRef]
- Cao, D.D.; Trinh, T.T.V.; Mai, H.D.T.; Vu, V.N.; Le, H.M.; Thi, Q.V.; Nguyen, M.A.; Duong, T.T.; Tran, D.T.; Chau, V.M.; et al. Antimicrobial Lavandulylated Flavonoids from a Sponge-Derived Streptomyces sp. G248 in East Vietnam Sea. Mar. Drugs 2019, 17, 529. [Google Scholar] [CrossRef] [PubMed]
- Reimer, A.; Blohm, A.; Quack, T.; Grevelding, C.G.; Kozjak-Pavlovic, V.; Rudel, T.; Hentschel, U.; Abdelmohsen, U.R. Inhibitory Activities of the Marine Streptomyces-Derived Compound SF2446A2 against Chlamydia trachomatis and Schistosoma mansoni. J. Antibiot. 2015, 68, 674–679. [Google Scholar] [CrossRef]
- Tenebro, C.P.; Trono, D.J.V.L.; Vicera, C.V.B.; Sabido, E.M.; Ysulat, J.A.; Macaspac, A.J.M.; Tampus, K.A.; Fabrigar, T.A.P.; Saludes, J.P.; Dalisay, D.S. Multiple Strain Analysis of Streptomyces Species from Philippine Marine Sediments Reveals Intraspecies Heterogeneity in Antibiotic Activities. Sci. Rep. 2021, 11, 17544. [Google Scholar] [CrossRef]
- Wolfender, J.-L.; Litaudon, M.; Touboul, D.; Queiroz, E.F. Innovative Omics-Based Approaches for Prioritisation and Targeted Isolation of Natural Products–New Strategies for Drug Discovery. Nat. Prod. Rep. 2019, 36, 855–868. [Google Scholar] [CrossRef]
- Olivon, F.; Allard, P.-M.; Koval, A.; Righi, D.; Genta-Jouve, G.; Neyts, J.; Apel, C.; Pannecouque, C.; Nothias, L.-F.; Cachet, X.; et al. Bioactive Natural Products Prioritization Using Massive Multi-Informational Molecular Networks. ACS Chem. Biol. 2017, 12, 2644–2651. [Google Scholar] [CrossRef]
- Quinlan, Z.A.; Koester, I.; Aron, A.T.; Petras, D.; Aluwihare, L.I.; Dorrestein, P.C.; Nelson, C.E.; Wegley Kelly, L. ConCISE: Consensus Annotation Propagation of Ion Features in Untargeted Tandem Mass Spectrometry Combining Molecular Networking and In Silico Metabolite Structure Prediction. Metabolites 2022, 12, 1275. [Google Scholar] [CrossRef] [PubMed]
- Lilge, L.; Ersig, N.; Hubel, P.; Aschern, M.; Pillai, E.; Klausmann, P.; Pfannstiel, J.; Henkel, M.; Morabbi Heravi, K.; Hausmann, R. Surfactin Shows Relatively Low Antimicrobial Activity against Bacillus subtilis and Other Bacterial Model Organisms in the Absence of Synergistic Metabolites. Microorganisms 2022, 10, 779. [Google Scholar] [CrossRef] [PubMed]
- De Oliveira, A.; Cataneli Pereira, V.; Pinheiro, L.; Moraes Riboli, D.; Benini Martins, K.; Ribeiro De Souza Da Cunha, M. Antimicrobial Resistance Profile of Planktonic and Biofilm Cells of Staphylococcus aureus and Coagulase-Negative Staphylococci. Int. J. Mol. Sci. 2016, 17, 1423. [Google Scholar] [CrossRef]
- Nothias, L.-F.; Petras, D.; Schmid, R.; Dührkop, K.; Rainer, J.; Sarvepalli, A.; Protsyuk, I.; Ernst, M.; Tsugawa, H.; Fleischauer, M.; et al. Feature-Based Molecular Networking in the GNPS Analysis Environment. Nat. Methods 2020, 17, 905–908. [Google Scholar] [CrossRef]
- Dührkop, K.; Fleischauer, M.; Ludwig, M.; Aksenov, A.A.; Melnik, A.V.; Meusel, M.; Dorrestein, P.C.; Rousu, J.; Böcker, S. SIRIUS 4: A Rapid Tool for Turning Tandem Mass Spectra into Metabolite Structure Information. Nat. Methods 2019, 16, 299–302. [Google Scholar] [CrossRef]
- Dührkop, K.; Nothias, L.-F.; Fleischauer, M.; Reher, R.; Ludwig, M.; Hoffmann, M.A.; Petras, D.; Gerwick, W.H.; Rousu, J.; Dorrestein, P.C.; et al. Systematic Classification of Unknown Metabolites Using High-Resolution Fragmentation Mass Spectra. Nat. Biotechnol. 2021, 39, 462–471. [Google Scholar] [CrossRef]
- Djoumbou Feunang, Y.; Eisner, R.; Knox, C.; Chepelev, L.; Hastings, J.; Owen, G.; Fahy, E.; Steinbeck, C.; Subramanian, S.; Bolton, E.; et al. ClassyFire: Automated Chemical Classification with a Comprehensive, Computable Taxonomy. J. Cheminformatics 2016, 8, 61. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.W.; Wang, M.; Leber, C.A.; Nothias, L.-F.; Reher, R.; Kang, K.B.; Van Der Hooft, J.J.J.; Dorrestein, P.C.; Gerwick, W.H.; Cottrell, G.W. NPClassifier: A Deep Neural Network-Based Structural Classification Tool for Natural Products. J. Nat. Prod. 2021, 84, 2795–2807. [Google Scholar] [CrossRef] [PubMed]
- Ono, K.; Demchak, B.; Ideker, T. Cytoscape Tools for the Web Age: D3.Js and Cytoscape.Js Exporters. F1000Research 2014, 3, 143. [Google Scholar] [CrossRef] [PubMed]
- Trischman, J.A.; Jensen, P.R.; Fenical, W. Halobacillin: A Cytotoxic Cyclic Acylpeptide of the Iturin Class Produced by a Marine Bacillus. Tetrahedron Lett. 1994, 35, 5571–5574. [Google Scholar] [CrossRef]
- Ma, Y.; Kong, Q.; Qin, C.; Chen, Y.; Chen, Y.; Lv, R.; Zhou, G. Identification of Lipopeptides in Bacillus megaterium by Two-Step Ultrafiltration and LC–ESI–MS/MS. AMB Express 2016, 6, 79. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.C.; Minton, M.A.; Sharma, M.M.; Georgiou, G. Structural and Immunological Characterization of a Biosurfactant Produced by Bacillus licheniformis JF-2. Appl. Environ. Microbiol. 1994, 60, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Grangemard, I.; Bonmatin, J.-M.; Bernillon, J.; Das, B.C.; Peypoux, F. Lichenysins G, a Novel Family of Lipopeptide Biosurfactants from Bacillus licheniformis IM 1307: Production, Isolation and Structural Evaluation by NMR and Mass Spectrometry. J. Antibiot. 1999, 52, 363–373. [Google Scholar] [CrossRef] [PubMed]
- Kowall, M.; Vater, J.; Kluge, B.; Stein, T.; Franke, P.; Ziessow, D. Separation and Characterization of Surfactin Isoforms Produced Bacillus subtilis. J. Colloid Interface Sci. 1998, 204, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Boukerb, A.M.; Simon, M.; Pernet, E.; Jouault, A.; Portier, E.; Persyn, E.; Bouffartigues, E.; Bazire, A.; Chevalier, S.; Feuilloley, M.G.J.; et al. Draft Genome Sequences of Four Pseudomonas aeruginosa Clinical Strains with Various Biofilm Phenotypes. Microbiol. Resour. Announc. 2020, 9, e01286-19. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Halary, S.; Duval, C.; Bernard, C.; Troussellier, M.; Beniddir, M.A.; Brunel, J.-M.; Castaldi, A.; Caudal, F.; Golléty, C.; et al. Diversity, Metabolome Profiling and Bioactivities of Benthic Filamentous Cyanobacteria Isolated from Coastal Mangroves of Mayotte. Front. Mar. Sci. 2023, 10, 1201594. [Google Scholar] [CrossRef]
- Tang, J.; Gao, H.; Hong, K.; Yu, Y.; Jiang, M.; Lin, H.; Ye, W.; Yao, X. Complete Assignments of 1H and 13C NMR Spectral Data of Nine Surfactin Isomers. Magn. Reson. Chem. 2007, 45, 792–796. [Google Scholar] [CrossRef] [PubMed]
- Pang, X.; Zhao, J.; Fang, X.; Liu, H.; Zhang, Y.; Cen, S.; Yu, L. Surfactin Derivatives from Micromonospora sp. CPCC 202787 and Their Anti-HIV Activities. J. Antibiot. 2017, 70, 105–108. [Google Scholar] [CrossRef]
- Madslien, E.H.; Rønning, H.T.; Lindbäck, T.; Hassel, B.; Andersson, M.A.; Granum, P.E. Lichenysin Is Produced by Most Bacillus licheniformis Strains. J. Appl. Microbiol. 2013, 115, 1068–1080. [Google Scholar] [CrossRef]
- Yeak, K.Y.C.; Perko, M.; Staring, G.; Fernandez-Ciruelos, B.M.; Wells, J.M.; Abee, T.; Wells-Bennik, M.H.J. Lichenysin Production by Bacillus licheniformis Food Isolates and Toxicity to Human Cells. Front. Microbiol. 2022, 13, 831033. [Google Scholar] [CrossRef]
- Yakimov, M.M.; Timmis, K.N.; Wray, V.; Fredrickson, H.L. Characterization of a New Lipopeptide Surfactant Produced by Thermotolerant and Halotolerant Subsurface Bacillus licheniformis BAS50. Appl. Environ. Microbiol. 1995, 61, 1706–1713. [Google Scholar] [CrossRef]
- Park, T.; Jeon, M.-K.; Yoon, S.; Lee, K.S.; Kwon, T.-H. Modification of Interfacial Tension and Wettability in Oil–Brine–Quartz System by in Situ Bacterial Biosurfactant Production at Reservoir Conditions: Implications for Microbial Enhanced Oil Recovery. Energy Fuels 2019, 33, 4909–4920. [Google Scholar] [CrossRef]
- Zammuto, V.; Rizzo, M.G.; De Pasquale, C.; Ferlazzo, G.; Caccamo, M.T.; Magazù, S.; Guglielmino, S.P.P.; Gugliandolo, C. Lichenysin-like Polypeptide Production by Bacillus licheniformis B3-15 and Its Antiadhesive and Antibiofilm Properties. Microorganisms 2023, 11, 1842. [Google Scholar] [CrossRef] [PubMed]
- Zhen, C.; Ge, X.-F.; Lu, Y.-T.; Liu, W.-Z. Chemical Structure, Properties and Potential Applications of Surfactin, as Well as Advanced Strategies for Improving Its Microbial Production. AIMS Microbiol. 2023, 9, 195–217. [Google Scholar] [CrossRef] [PubMed]
- Schmid, R.; Heuckeroth, S.; Korf, A.; Smirnov, A.; Myers, O.; Dyrlund, T.S.; Bushuiev, R.; Murray, K.J.; Hoffmann, N.; Lu, M.; et al. Integrative Analysis of Multimodal Mass Spectrometry Data in MZmine 3. Nat. Biotechnol. 2023, 41, 447–449. [Google Scholar] [CrossRef] [PubMed]
- Myers, O.D.; Sumner, S.J.; Li, S.; Barnes, S.; Du, X. One Step Forward for Reducing False Positive and False Negative Compound Identifications from Mass Spectrometry Metabolomics Data: New Algorithms for Constructing Extracted Ion Chromatograms and Detecting Chromatographic Peaks. Anal. Chem. 2017, 89, 8696–8703. [Google Scholar] [CrossRef]
- Biniarz, P.; Łukaszewicz, M. Direct Quantification of Lipopeptide Biosurfactants in Biological Samples via HPLC and UPLC-MS Requires Sample Modification with an Organic Solvent. Appl. Microbiol. Biotechnol. 2017, 101, 4747–4759. [Google Scholar] [CrossRef]
- Vijayasarathy, S.; Prasad, P.; Fremlin, L.J.; Ratnayake, R.; Salim, A.A.; Khalil, Z.; Capon, R.J. C3 and 2D C3 Marfey’s Methods for Amino Acid Analysis in Natural Products. J. Nat. Prod. 2016, 79, 421–427. [Google Scholar] [CrossRef]
- Coffey, B.M.; Anderson, G.G. Biofilm Formation in the 96-Well Microtiter Plate. In Pseudomonas Methods and Protocols; Filloux, A., Ramos, J.-L., Eds.; Methods in Molecular Biology; Springer: New York, NY, USA, 2014; Volume 1149, pp. 631–641. ISBN 978-1-4939-0472-3. [Google Scholar]
- Chen, D.; Cadelis, M.M.; Rouvier, F.; Troia, T.; Edmeades, L.R.; Fraser, K.; Gill, E.S.; Bourguet-Kondracki, M.-L.; Brunel, J.M.; Copp, B.R. α,ω-Diacyl-Substituted Analogues of Natural and Unnatural Polyamines: Identification of Potent Bactericides That Selectively Target Bacterial Membranes. Int. J. Mol. Sci. 2023, 24, 5882. [Google Scholar] [CrossRef]

| Cpd | Exact Mass | Chemical Formula | Lichenysin Precursor Ion [M + H]+ at m/z | Retention Time in Min | Cyclic Lichenysin Containing a Cx β-Hydroxy Fatty Acid Chain |
|---|---|---|---|---|---|
| 1 | 979.64380 | C49H87N8O12+ | 979.6434 | 17.6 | C11 |
| 2 | 993.65945 | C50H89N8O12+ | 993.6581 | 18.4 | C12 |
| 3 | 1007.67510 | C51H91N8O12+ | 1007.6734 | 17.9 | C13 |
| 4 | 1007.67510 | C51H91N8O12+ | 1007.6733 | 18.5 | C13 |
| 5 | 1021.69075 | C52H93N8O12+ | 1021.6896 | 19.4 | C14 |
| 6 | 1035.70640 | C53H95N8O12+ | 1035.7063 | 20.0 | C15 |
| 7 | 1035.70640 | C53H95N8O12+ | 1035.7063 | 20.2 | C15 |
| 8 | 1063.73770 | C55H99N8O12+ | 1063.7369 | 21.2 | C17 |
| Cpd | Exact Mass | Chemical Formula | Lichenysin Precursor Ion [M + H]+ at m/z | Retention Time in Min | Cyclic Surfactin Containing a Cx β-Hydroxy Fatty Acid Chain |
|---|---|---|---|---|---|
| 12 | 1008.65911 | C51H90N7O13+ | 1008.6618 | 19.3 | C13 |
| 13 | 1022.67476 | C52H92N7O13+ | 1022.6767 | 19.9 | C14 |
| 14 | 1036.69041 | C53H94N7O13+ | 1036.6906 | 20.5 | C15 |
| Cpd | Exact Mass | Chemical Formula | Lichenysin Precursor Ion [M + H]+ at m/z | Retention Time in Min | Linear Lichenysin Containing a Cx β-Hydroxy Fatty Acid Chain |
|---|---|---|---|---|---|
| 9 | 1025.68566 | C51H93N8O13+ | 1025.6832 | 17.4 | C13 |
| 10 | 1039.70131 | C52H95N8O13+ | 1039.6990 | 18.1 | C14 |
| 11 | 1053.71696 | C53H97N8O13+ | 1053.7147 | 18.8 | C15 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Castaldi, A.; Truong, B.N.; Vu, Q.T.; Le, T.H.M.; Marie, A.; Le Pennec, G.; Rouvier, F.; Brunel, J.-M.; Longeon, A.; Pham, V.C.; et al. Computational Methods Reveal a Series of Cyclic and Linear Lichenysins and Surfactins from the Vietnamese Marine Sediment-Derived Streptomyces Strain G222. Molecules 2024, 29, 1458. https://doi.org/10.3390/molecules29071458
Castaldi A, Truong BN, Vu QT, Le THM, Marie A, Le Pennec G, Rouvier F, Brunel J-M, Longeon A, Pham VC, et al. Computational Methods Reveal a Series of Cyclic and Linear Lichenysins and Surfactins from the Vietnamese Marine Sediment-Derived Streptomyces Strain G222. Molecules. 2024; 29(7):1458. https://doi.org/10.3390/molecules29071458
Chicago/Turabian StyleCastaldi, Andrea, Bich Ngan Truong, Quyen Thi Vu, Thi Hong Minh Le, Arul Marie, Gaël Le Pennec, Florent Rouvier, Jean-Michel Brunel, Arlette Longeon, Van Cuong Pham, and et al. 2024. "Computational Methods Reveal a Series of Cyclic and Linear Lichenysins and Surfactins from the Vietnamese Marine Sediment-Derived Streptomyces Strain G222" Molecules 29, no. 7: 1458. https://doi.org/10.3390/molecules29071458
APA StyleCastaldi, A., Truong, B. N., Vu, Q. T., Le, T. H. M., Marie, A., Le Pennec, G., Rouvier, F., Brunel, J.-M., Longeon, A., Pham, V. C., Doan, T. M. H., & Bourguet-Kondracki, M.-L. (2024). Computational Methods Reveal a Series of Cyclic and Linear Lichenysins and Surfactins from the Vietnamese Marine Sediment-Derived Streptomyces Strain G222. Molecules, 29(7), 1458. https://doi.org/10.3390/molecules29071458