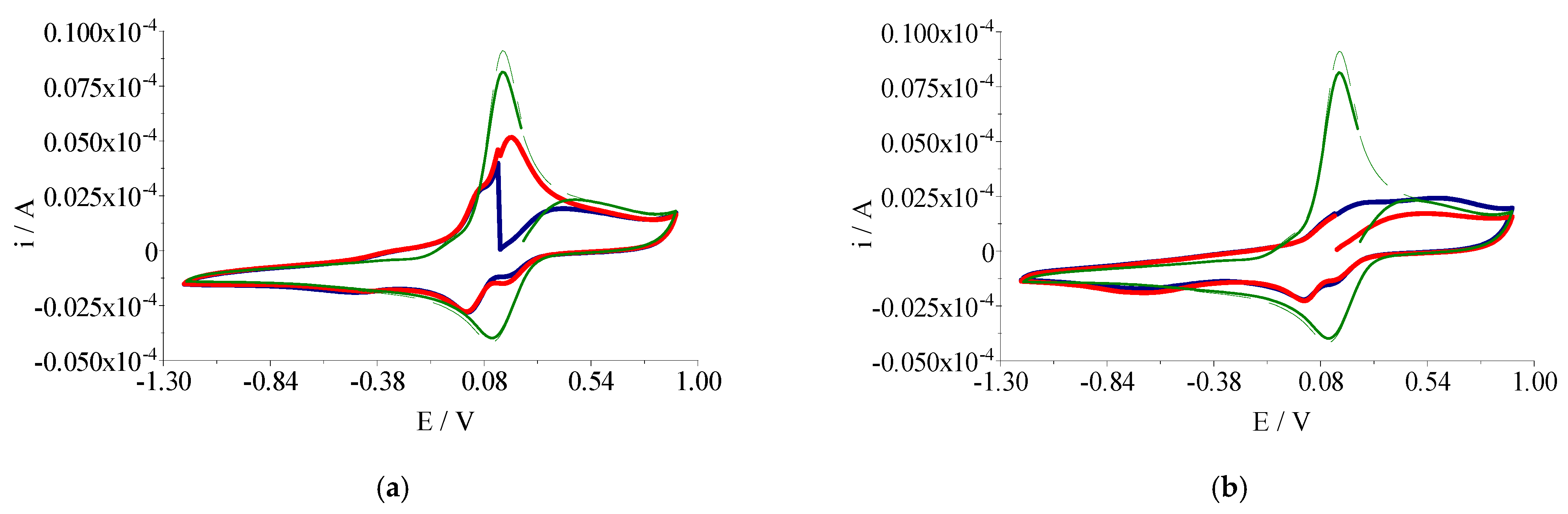1. Introduction
The triazolopyrimidine derivatives are valuable pharmacophores based on their resemblance to purine bases. As result, several compounds bearing diverse substituents were developed as species with relevant biological activity [
1]. The most recent findings in the field describe triazolopyrimidines as agents with potent antiproliferative activity by inducing G0/G1 phase arrest [
2] or cellular apoptosis [
3], as species with anti-HIV activity [
4], as SARS-CoV-2 main protease inhibitors [
5], and as antimicrobials [
6,
7]. Additionally, it is believed that triazolopyrimidines are promising candidates for treating Alzheimer’s disease and related neurodegenerative tauopathies [
8].
A promising activity, comparable with that of standard anticancer drugs, against a panel of cancer cell lines was reported for [
1,
2,
4]triazolo[1,5-
a]pyrimidine derivatives (tpds). Among these, some pyrimidine ring-disubstituted species exhibit a good activity against colon (HCT116) [
9], human promyelocytic leukemia (HL60) [
10], gastric (MGC-803), esophageal (EC 109), lung (A549 and PC-9) [
11], liver (HepG2) [
12] and breast (MCF-7) [
12,
13] cancer cell lines. It is worth mentioning that most of these lines are not sensitive to inorganic drugs such as cisplatin and related derivatives.
A significant antibacterial activity, comparable with that of standard antimicrobials against
Staphylococcus aureus,
Escherichia coli [
14,
15],
Bacillus subtilis [
14] and
Pseudomonas aeruginosa [
15] strains was also reported for some substituted tpds.
Considering its biological potency, the tp scaffold is often used to develop biologically active complexes, with a net preference for Cu(II) against other metal ions. Copper(II) exhibits particular traits, such as borderline acid characteristics or stereochemical and oxidation state versatility [
16]. Moreover, most of the Cu(II) complexes with tps were reported as having antiparasitic [
17], antitumor, or antimicrobial activity [
18].
Among the antiparasitic compounds, [Cu
2(μ-7atp)
4Cl
2]Cl
2·4H
2O, [Cu
2(μ-7atp)
4(H
2O)
2](NO
3)·4H
2O (7atp: 7-amino-1,2,4-triazolo[1,5-
a]pyrimidine) [
19], [Cu
2(dmtp)
4Cl
4]2H
2O [
20], [Cu(dmtp)
4(H
2O)
2](ClO
4)
2·2H
2O [
21], and [Cu(dmtp)
2(NO
3)
2(H
2O)] (dmtp: 5,7-dimethyl-1,2,4-triazolo[1,5-
a]pyrimidine) [
22] showed a substantial in vitro activity against
Leishmania spp. and
Trypanosoma cruzi. Additionally, low toxicity against macrophage host cells was observed for all complexes.
A good antiproliferative activity against
L. infantum and
L. braziliensis was also observed for [Cu(HmtpO)
2(H
2O)
3](ClO
4)
2H
2O and {[Cu(HmtpO)
2(H
2O)
2](ClO
4)
22HmtpO}
n (HmtpO: 5-methyl-1,2,4-triazolo[1,5-
a]pyrimidin-7(4H)-one)—complexes that affect the metabolism of the parasites at the level of both NAD+/NADH balance and organelle membranes, causing the cell death [
23].
Moreover, complexes with mixed ligands such as [Cu(H
2O)(phen)(tp)
2](ClO
4)
2·H
2O and [Cu(NO
3)(H
2O)(phen)(tp)](NO
3) (phen: 1,10-phenanthroline) [
24] inhibit in vitro epimastigote forms of
T. cruzi and promastigotes of
L. peruviana, while [Cu(dmtp)
2(bpym)(H
2O)
2](ClO
4)
22H
2O and [Cu
2(tp)
2(bpym)
2(μ-bpym)(ClO
4)
2](ClO
4)
2 (bpym: 2,2′-bipyrimidine) exhibit activity against the same parasites, which cause leishmaniasis and Chagas disease, respectively [
25].
A broad spectrum of antimicrobial activity was demonstrated by [Cu
2(pmtp)
2Cl
4(OH
2)
2] [
26], [Cu(pmtp)(CH
3COO)
2]0.5H
2O [
27] and [Cu(pmtp)(OH
2)
3](ClO
4)3H
2O (pmtp: 5-phenyl-7-methyl-1,2,4-triazolo[1,5-
a]pyrimidine) [
28] against both planktonic and biofilm-embedded strains. It is worth mentioning the activity of all complexes against methicillin-resistant
S. aureus (MRSA). Moreover, complexes with mixed ligands [Cu(N-N)
2(pmtp)](ClO
4)
2 (N-N: 2,2′-bipyridine (bpy) and phen) showed antibacterial potential against several bacterial strains, including MRSA, the extended-spectrum beta-lactamase (ESBL)-producing
E. coli 5, and the multi-drug-resistant
P. aeruginosa 9027, both in planktonic and biofilm growth state [
18].
Among these species, [Cu(pmtp)(CH
3COO)
2]0.5H
2O and [Cu(pmtp)(OH
2)
3](ClO
4)3H
2O also induce a decrease in the DNA content of cells in the G0/G1 phase in a human colon adenocarcinoma cell line (HT 29) [
27], while [Cu(N-N)
2(pmtp)](ClO
4)
2 exhibited excellent activity against B16 murine melanoma cells, that in addition, is accompanied by a lack of cytotoxicity against healthy BJ cells [
18].
Concerning species with N-N derivatives, we have recently shown that {[Cu(bpy)
2(μ
2OClO
3)]ClO
4}
n exhibits a selective cytotoxicity against B16 cells while [Cu(phen)
2(OH
2)](ClO
4)
2 proved to be a very potent antibacterial agent against both susceptible and resistant Gram-positive and Gram-negative strains, in planktonic or biofilm growth states [
29]. By combining these N-N heterocycles with a disubstituted tp derivative we succeded in improving the antimicrobial activity of Cu(II) complexes [
18].
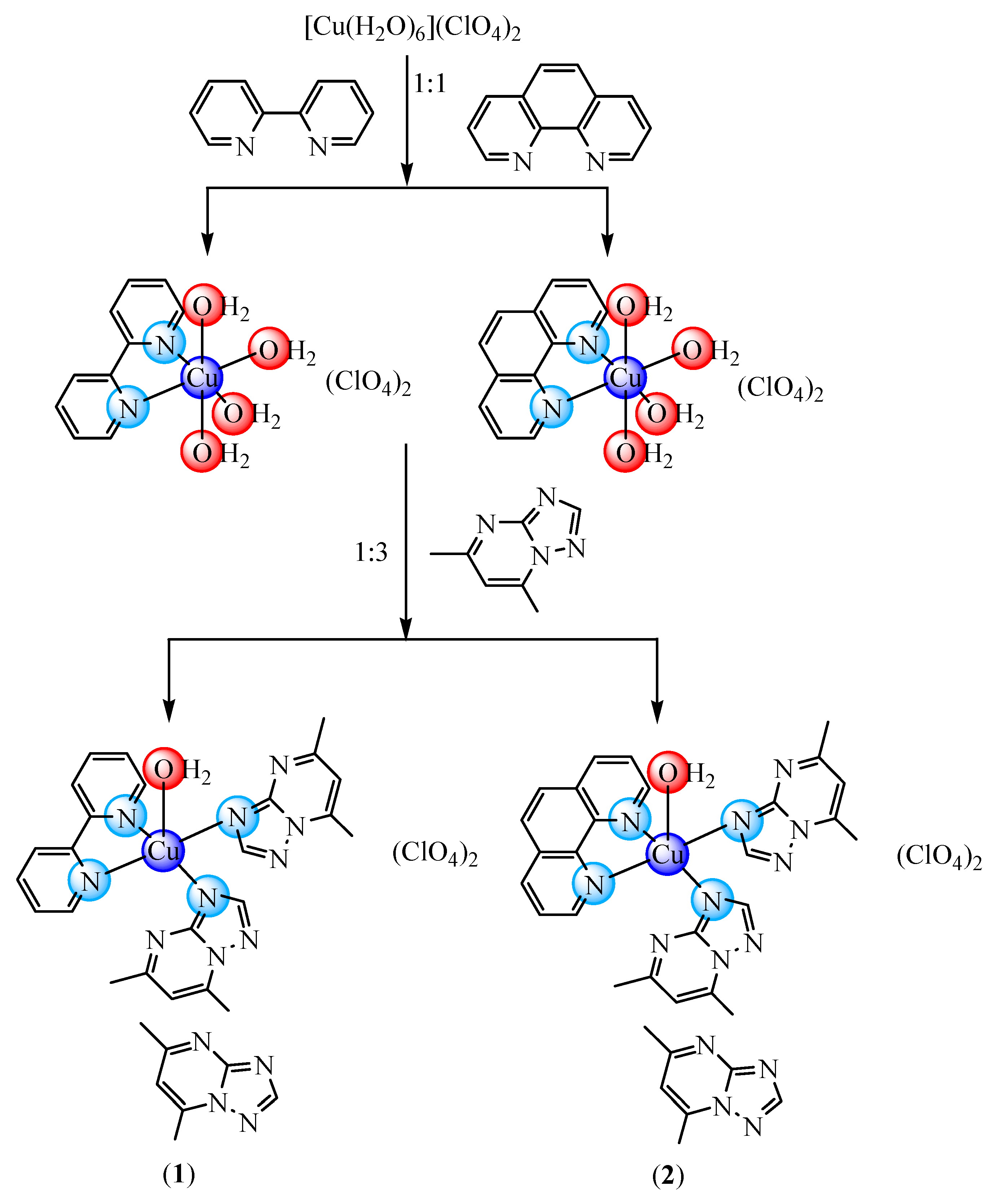
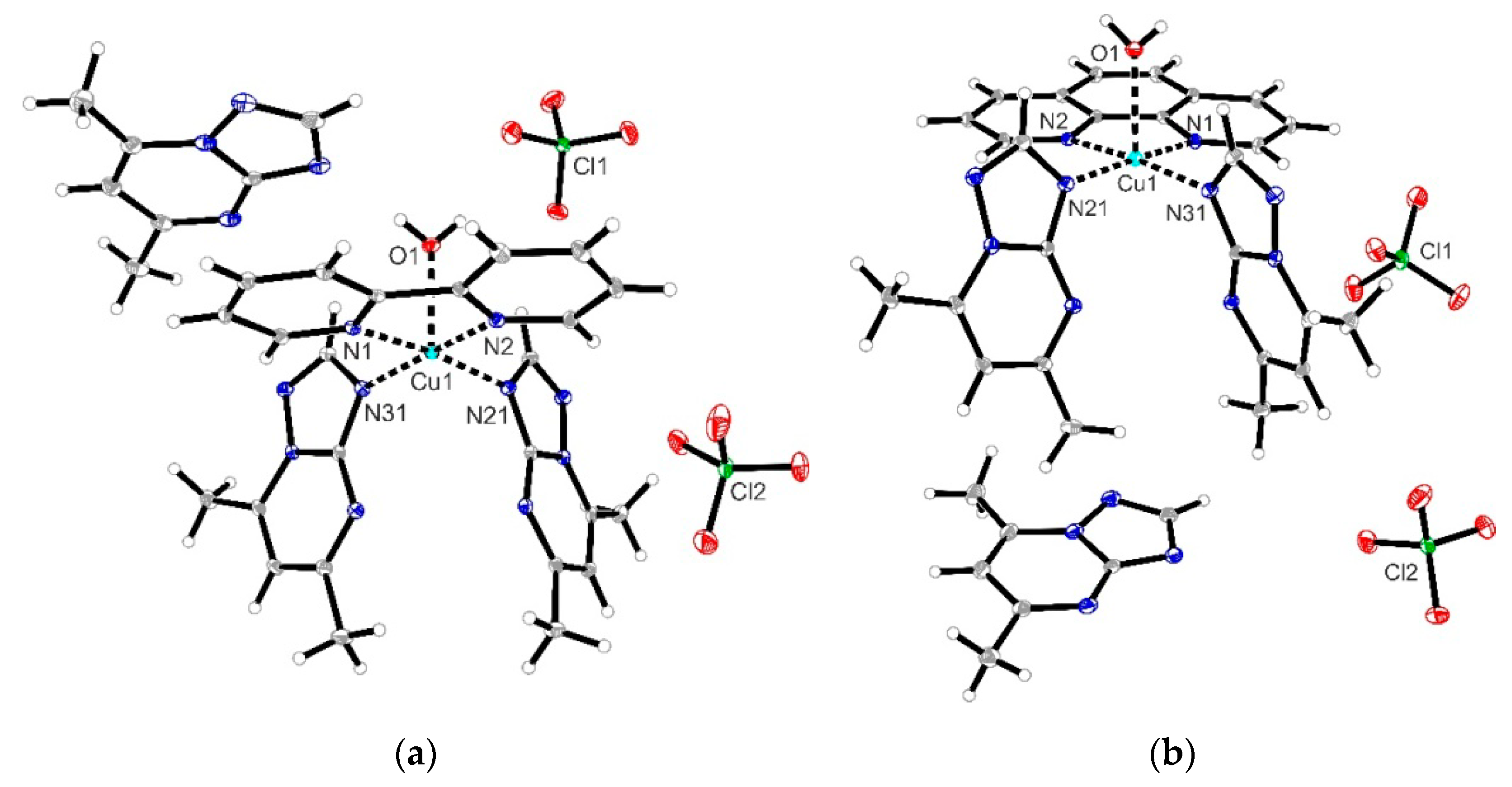
In an attempt to modulate the biological activity of dmtp Cu(II) complexes through such N-N derivatives, we present here new complexes with mixed ligands of [Cu(N-N)(dmtp)2(OH2)](ClO4)2dmtp (N-N: bpy (1) and phen (2)) type that were fully characterized by single-crystal X-ray analysis and spectroscopic methods. In addition, data regarding the antitumor and antimicrobial activity of these complexes are also included, together with that concerning their potential mechanisms of action.
3. Materials and Methods
3.1. Reagents
The chemicals for the synthesis of the complexes were purchased from Sigma-Aldrich (Darmstadt, Germany) (copper(II) perchlorate hexahydrate (≥99.99% trace metals basis), 2,2′-biyridine (bpy, 99%), 1,10-phenantroline (phen, 99%), 2,3-pentanedione (97%) and 3-amino-4H-1,2,4-triazole (96%)) and Merck (Darmstadt, Germany; dibenzo-18-crown-6-ether, potassium superoxide) at reagent grade and were used as received, without further purification. The 5,7-dimethyl-1,2,4-triazolo[1,5-a]pyrimidine (dmtp) was synthesized by [1+1} condensation of 3-amino-4H-1,2,4-triazole and 2,3-pentanedione.
3.2. Physical Measurements
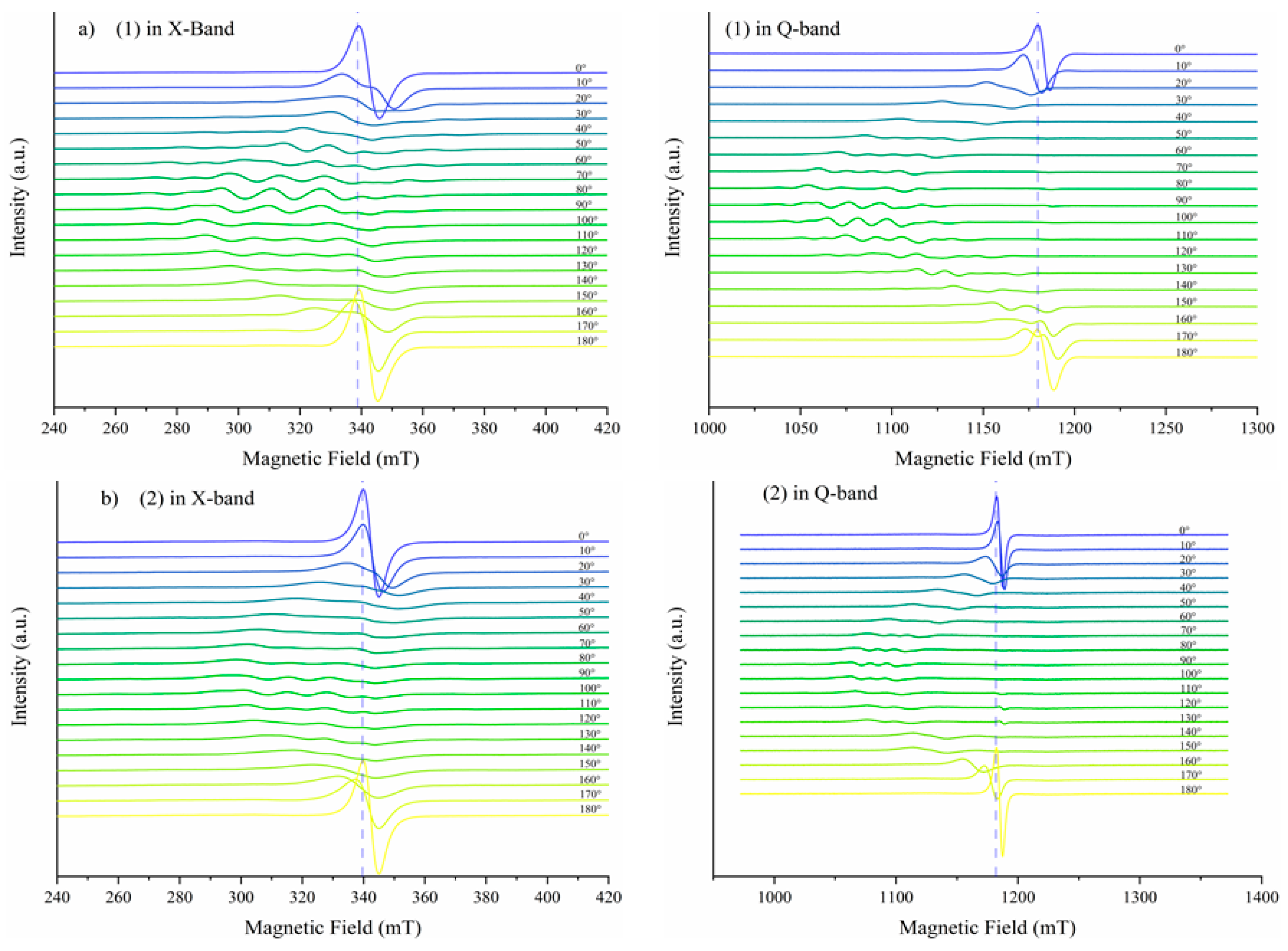
A EuroEA elemental analyzer (Perkin Elmer, Waltham, MA, USA) was used for chemical analyses (C, N, and H). Fourier Transform Infrared spectroscopy (FTIR) spectra were recorded in KBr pellets with a Tensor 37 spectrometer (Bruker, Billerica, MA, USA) in the 400–4000 cm−1 range. UV-Vis spectroscopy was performed in the solid state on a V 670 spectrophotometer (Jasco, Easton, MD, USA) with Spectralon as standard in the 200–1500 nm range. The X-band Electron Paramagnetic Resonance (EPR) spectroscopy measurements were carried out with a continuous wave X-Band EMX plus EPR spectrometer (Bruker AXS GmbH, Karlsruhe, Germany) equipped with a Bruker X-SHQ 4119HS-W1 X-Band resonator. The measurement parameters for the X-Band measurements, if not otherwise mentioned, were set as follows: microwave frequency 9.879 GHz, microwave power 2 mW, modulation amplitude 0.1 mT, conversion time 26.67 ms, time constant 10.24 ms at one scan. A continuous-wave Q-band Bruker ELEXSYS E500Q EPR spectrometer and an ER 5106 QT-W resonator were used for the Q-Band measurements. The measurement parameters for the Q-Band measurements, if not otherwise mentioned, were set as follows: microwave frequency 34.1507 GHz, microwave power 0.59 mW, modulation amplitude 0.1 mT, conversion time 40 ms, time constant 20.48 ms at one scan. All measurements were carried out at room temperature. Cyclic voltammograms were recorded at room temperature with an electrochemical system (potentiostat/galvanostat) Autolab PGSTAT 12 using the GPES (General Purpose Electrochemical System) software. These studies were carried out in inert atmosphere (Ar 99.9999%) in DMSO containing tetrabutylammonium perchlorate (Bu4NClO4) 0.1 M as supporting electrolyte. The reference electrode was Ag/AgCl separated from the solution by a bridge filled with a 0.1 M Bu4NClO4 solution in DMSO. The counter electrode was the platinum wire. A platinum disk electrode with 0.3 mm diameter was used as working electrode. The complexes concentration was 0.3 mM in all measurements.
X-ray single crystal diffraction data sets for (
1) were collected with a Bruker APEX II CCD diffractometer (Bruker AXS, Karlsruhe, Germany). Programs used: data collection:
APEX3 V2016.1-0 [
54]; cell refinement:
SAINT V8.37A [
54]; data reduction:
SAINT V8.37A [
54]; absorption correction,
SADABS V2014/7 [
54]; structure solution
SHELXT-2015 [
55]; structure refinement
SHELXL-2015 [
56]. For compound (
2), data sets were collected with a Nonius Kappa CCD diffractometer (Bruker Nonius B.V., Delft, Holland). Programs used: data collection,
COLLECT [
57]; data reduction Denzo-SMN [
58]; absorption correction
Denzo [
59]; structure solution
SHELXT-2015 [
55]; structure refinement
SHELXL-2015 [
56] and graphics,
XP [
60].
R-values are given for observed reflections, and
wR
2 values are given for all reflections.
Exceptions and special features: For compound (
2), one perchlorate anion was found to be disordered over two positions in the asymmetric unit. Several restraints (SADI, SAME, ISOR, and SIMU) were used to improve refinement stability. Additionally, for compounds (
1) and (
2), two badly disordered water molecules were found in the asymmetrical unit and could not be satisfactorily refined. The
SQUEEZE program [
61] was therefore used to remove the effects of the solvent mathematically. The quoted formula and derived parameters do not include the squeezed solvent molecules. The crystallographic data for complexes (
1) and (
2) were deposited at the Cambridge Crystallographic Data Centre as a supplementary publication with CCDC numbers 2067846 and 2067847, respectively. These data can be obtained free of charge via
http://www.ccdc.cam.ac.uk/data_request/cif (accessed on 29 August 2021), by e-mailing
data_request@ccdc.cam.ac.uk, or by contacting the Cambridge Crystallographic Data Centre, 12 Union Road, Cambridge CB2 1EZ, UK; Fax: +44-1223-336033.
3.3. Synthesis and Characterization of the Complexes
[Cu(dpy)(dmtp)2(OH2)](ClO4)2·dmtp (1): To a solution, containing copper(II) perchlorate hexahydrate (0.186 g, 0.5 mmol) in 50 mL ethanol, a solution containing dpy (0.078 g, 0.5 mmol) in 25 mL ethanol was added. This mixture was magnetically stirred at 50 °C for 2 h, until the color turned blue, and then a solution containing dmtp (0.222 g, 1.5 mmol) in 50 mL ethanol was added. The stirring of the reaction mixture continued for 4 h, while the blue color intensified. Crystals suitable for X-ray analysis were obtained after three weeks by slow evaporation of this solution. Yield: 78% (0.34 g). Analysis, found: C, 42.37; H, 3.75; N, 22.31%; calculated forCuC31H34N14O9Cl2 (Mw: 881.15 g mol−1): C, 42.26; H, 3.89; N, 22.25%, IR (KBr pellet, cm−1): υ(H2O), 3511 w; υ(CH), 3136 w, 3112 w, 3083 w; υ(C=N)trp, 1632 m; υ(C=N)pym, 1556 m; υ(C=N)+υ(C=C), 1602m, 1496m; υ3(ClO4), 1095 vs; υ4(ClO4), 623 m; υ(Cu-N), 417 w, UV-Vis (solid, nm): π → π*, 290, 345; dxz, dyz → dz2, 590; dxy → dz2, 675.
[Cu(phen)(dmtp)2(OH2)](ClO4)2·dmtp (2): To a solution, containing copper(II) perchlorate hexahydrate (0.186 g, 0.5 mmol) in 50 mL water, a solution containing phen (0.090 g, 1 mmol) in 25 mL ethanol was added. This mixture was magnetically stirred at 50 °C for 2 h, until the color turned green, and then a solution containing dmtp (0.222 g, 1.5 mmol) in 25 mL ethanol was added. The stirring of the reaction mixture continued for 4 h, until the blue color was formed. Crystals suitable for X-ray analysis were obtained after two weeks by slow evaporation of this solution. Yield: 75% (0.34 g). Analysis, found: C, 43.83; H, 3.75; N, 21.68%; calculated forCuC33H34N14O9Cl2 (Mw: 905.17 g mol−1): C, 43.79; H, 3.79; N, 21.66%, IR (KBr pellet, cm−1): υ(H2O), 3517 w; υ(CH), 3137 w, 3110 w, 3078 w; υ(C=N)trp, 1630 m; υ(C=N)pym, 1553 m; υ(C=N)+υ(C=C), 1474 m, 1443 m; υ3(ClO4), 1099 vs; υ4(ClO4), 624 m; υ(Cu-N), 432 w, UV-Vis (solid, nm): π → π*, 265, 360; dxz, dyz →dz2, 610; dxy → dz2, 680.
3.4. Biological Characterization of Compounds
Screening of the Antibacterial Properties
The antibacterial assays were carried out using two Gram-negative (Escherichia coli ATCC 25922, Pseudomonas aeruginosa ATCC 27853) and two Gram-positive (Staphylococcus aureus ATCC 25923, MRSA (meticillin-Resistant S. aureus) 388 (ATCC, American Type Culture Collection) bacteria.
The antimicrobial activity of the two complexes (
1) and (
2) was assessed using their minimum inhibitory concentration (MIC) by the broth microdilution method, as previously described by the Clinical and Laboratory Standard Institute (CLSI; M07-A9 document) [
62]. A broth microdilution assay was performed in sterile 96-well plates using Muller-Hinton broth (Scharlau; MHB). The compounds solubilized in DSO (1,000 μg/mL) were serially diluted in 90 μL MHB, in concentrations ranging between 500 and 0.97 μg/mL. Overnight cultures were used to prepare suspension in phosphate-buffered saline (pH 7.0) to match a 0.5 McFarland density. The bacterial suspensions were further diluted to 1:100 by adding 0.1 mL of bacterial suspension to 9.9 mL of MHB. A volume of 10 μL of this dilution was added to each well of columns 1–10, containing different concentrations of the tested compounds. This resulted in the final desired inoculum of 105 cfu/mL. Additionally, 10 μL of the diluted bacterial suspension was added in column 11 (the growth control). A volume of 100 μL MHB was added to the microtiter plates’ sterility control wells (column 12). After inoculation, the plates were incubated at 35 ± 2 ºC for 18 h. Growth was measured as light absorbance (620 nm) in comparison to an uninoculated well (negative control). It was detected using a microtiter plate reader (Apollo LB 911ELISA (Berthold Technologies GmbH & Co. KG, Waltham, MA, USA)). The MIC of each compound was defined as the lowest concentration required to inhibit bacterial growth.
The method for assessment of biofilm formation on polystyrene microtiter plates was based on previously published protocols [
18,
63,
64]. The absorbance at 490 nm was measured with an Apollo LB 911ELISA (Berthold Technologies GmbH & Co. KG, Waltham, MA, USA) reader. The minimal biofilm-eradication concentration (MBEC) was defined as the lowest concentration of antibiotics required to eradicate the biofilm. All experiments were performed in triplicate.
3.5. In Vitro Cytotoxicity Assay
3.5.1. Cell Culture Conditions
Human fibroblast cells (BJ—ATCC CRL-2522, USA) were grown in minimal essential medium (MEM) supplemented with 2 mM L-Glutamine, 10% fetal calf serum (FCS), 100 units/mL of penicillin, and 100 µg/mL of streptomycin at 37 °C in a humidified incubator under an atmosphere containing 5% CO2. Mouse melanoma cells (B16—ATCC CRL—6475, USA) were grown in DMEM (Dulbecco’s Modified Eagle Medium) supplemented with similar reagents as MEM. All cell cultivation media and reagents were purchased from Biochrom AG (Berlin, Germany) and Sigma-Aldrich (Darmstadt, Germany).
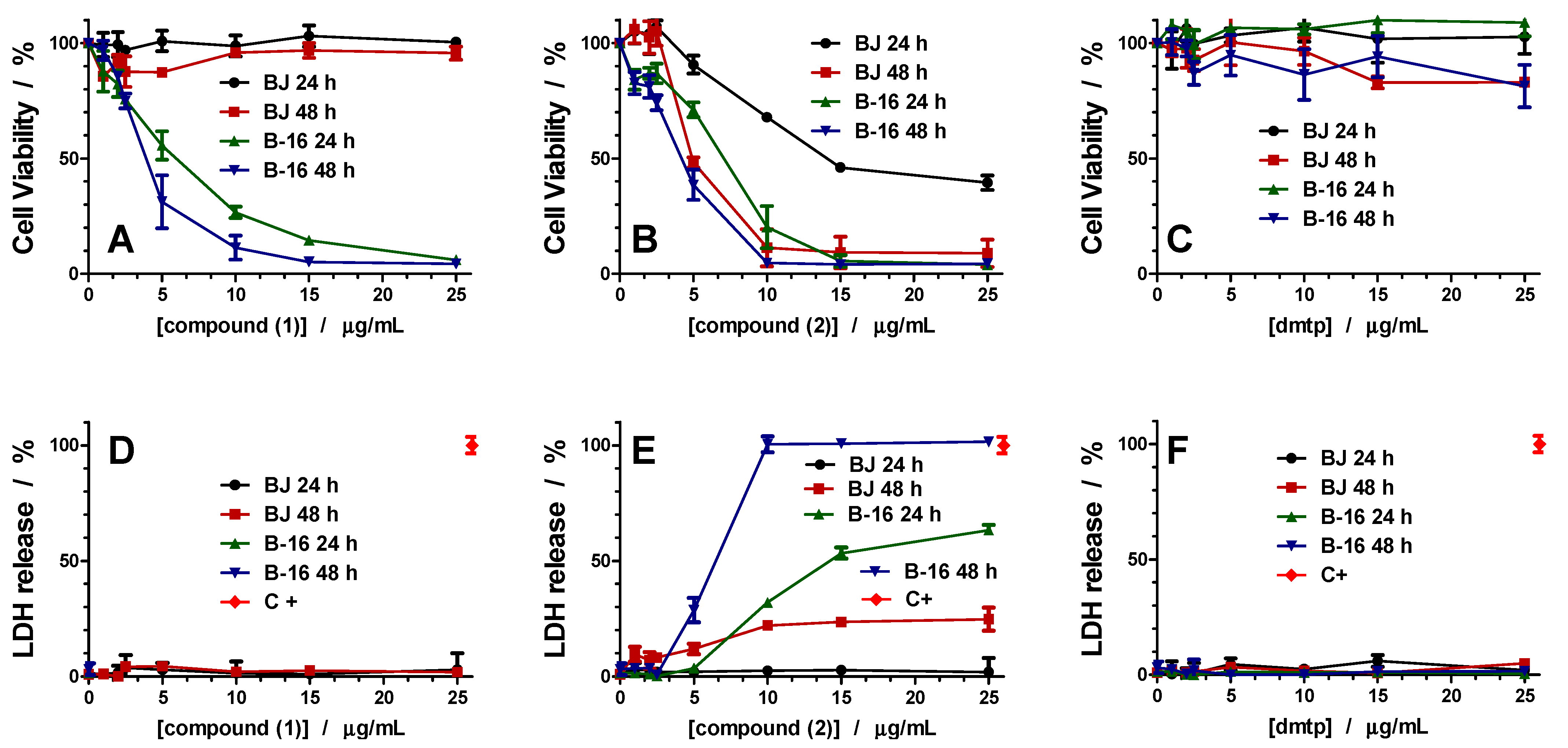
3.5.2. Cell Viability Assay
Cell viability was evaluated using 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay as described previously [
18]. First, the cells were seeded in 96-well plates (20000 cells/well) and cultured for 24 h in medium. After overnight incubation, the medium was changed, and the investigated sample at a concentration varying from 0.03 to 0.25 mg/mL was added for 24 h. The negative control was represented by cells cultivated in a medium without the investigated compounds. Following incubation, the medium was changed, and the MTT solution was added to each well to a final concentration of 1 mg/mL and incubated for an additional 4 h, at 37 °C. Finally, the medium was collected, and DMSO was used to dissolve the insoluble formazan product. The absorbance of the samples was recorded at 570 nm using a plate reader Mithras 940 (Berthold). The data were corrected for the background, and the percentage of viable cells was obtained using the equation:
The half-maximal inhibitory concentration (IC50) was determined by fitting the data with a sigmoidal logistical equation using the software Origin 8.1 (Microcal Inc., Los Angeles, CA, USA).
3.5.3. Lactate Dehydrogenase (LDH) Release Assay
Membrane integrity after peptide treatment was assessed based on LDH release using a CytoTox 96 Non-Radioactive Cytotoxicity Assay (Promega), as previously described [
65]. Cells were treated with different concentrations for 24 h, and the medium was used for the LDH assay. Maximal LDH release was obtained by complete cell lysis induced by using 1% of Triton X-100. The absorption resulting from LDH activity was measured in a microplate reader Mithras 940 (Berthold) at 490 nm, and the % LDH was calculated as: [(corrected absorbance of the LDH released in treated cells)/(corrected absorbance of the total LDH released)] * 100%.
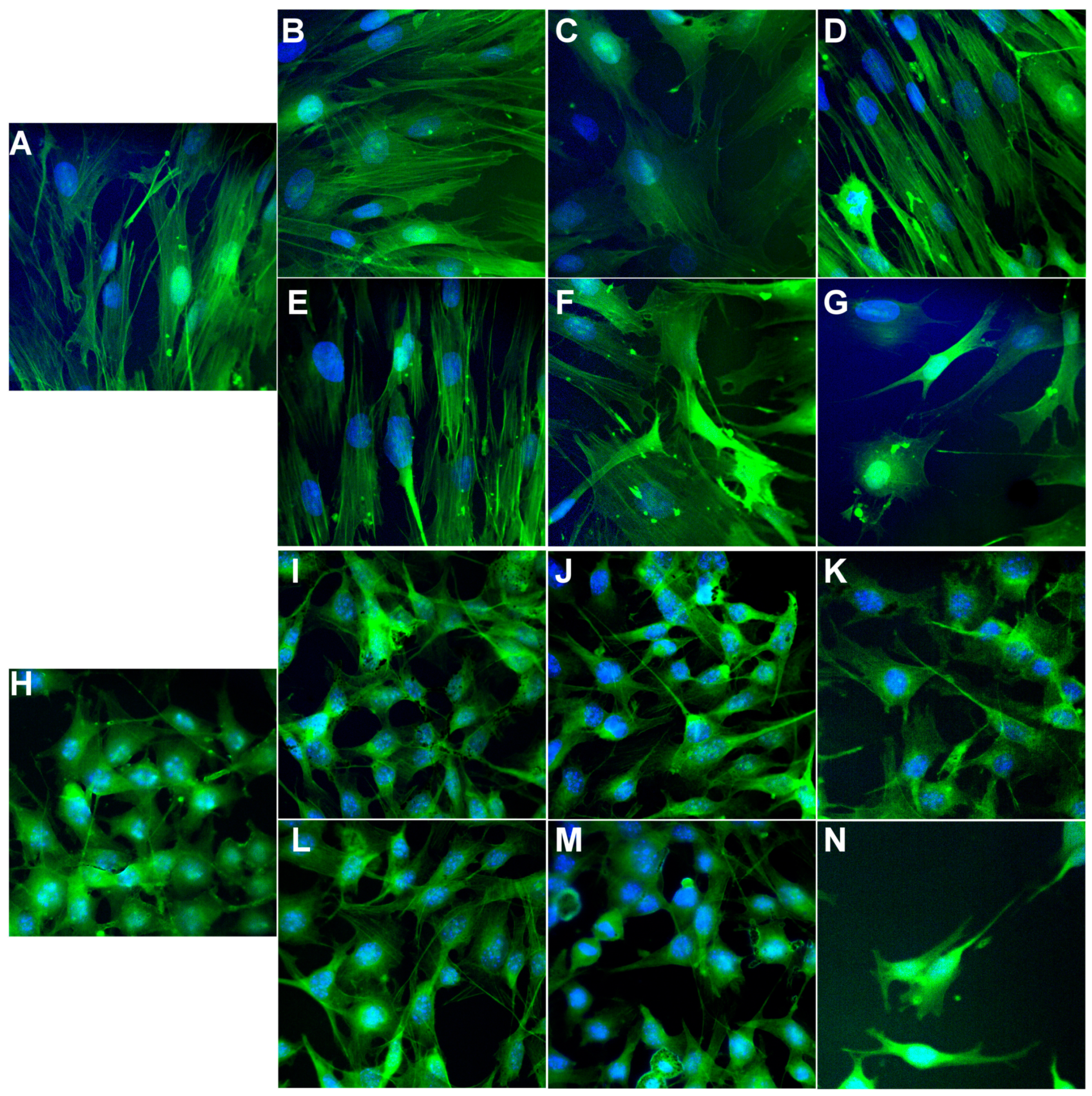
3.5.4. Phalloidin Staining and Cell Imaging
According to the manufacturer protocol, the cytoskeleton actin filaments of BJ and B16 cells were stained with phalloidin-FITC (Sigma-Aldrich, St. Louis, MO, USA). Briefly, cells were washed with PBS (5 min, 3 times), fixed for 5 min with 3% paraformaldehyde, washed three times with PBS, permeabilized with 0.1% Triton X-100 in PBS for 15 min, washed three times with PBS, stained with 20 μg/mL phalloidin-FITC at room temperature for 1 h and rewashed three times with PBS. The cell nucleus was stained with 8 μM of Hoechst 33342 solution for 10 min, washed three times with PBS, and finally, mounted and sealed on glass slides with FluorSave™ Reagent (Merck Millipore, Germany). The fluorescence images were acquired using an Andor DSD2 Confocal Unit (Andor, Ireland), mounted on an epifluorescence microscope, Olympus BX-51 (Olympus, Germany), equipped with a 40x and 100x objective and an appropriate DAPI/Hoechst filter cube (excitation filter 390/40 m, dichroic mirror 405 nm and emission filter 452/45 nm) and GFP/FITC filter cube (excitation filter 466/40 nm, dichroic mirror 488 nm and emission filter 525/54 nm).
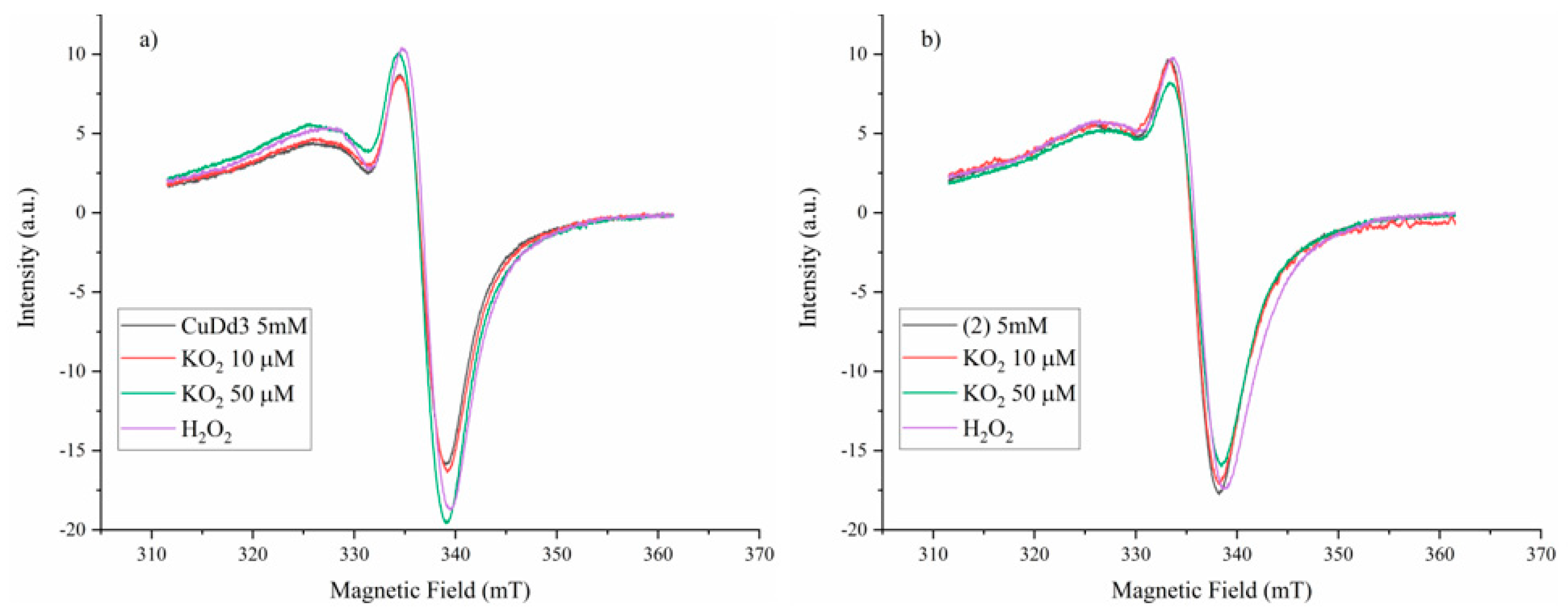
3.5.5. Superoxide Scavenging Ability
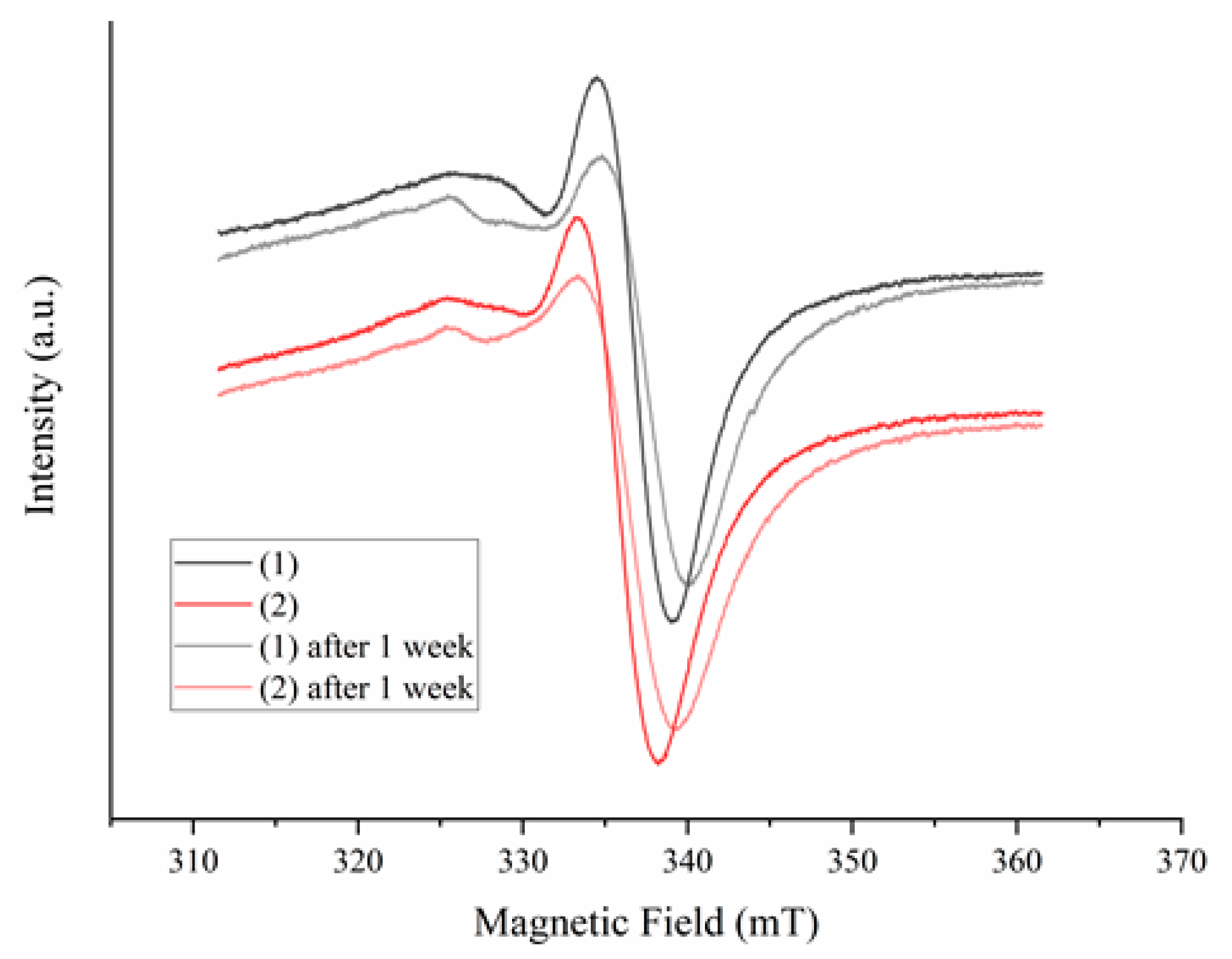
The superoxide scavenging ability of the complexes was tested using the KO2 compound as a superoxide source combined with EPR spectroscopy. To carry out the experiments, a 10 µM in DMSO solution of the complexes was mixed with different concentrations of KO2 in DMSO solution, and the EPR signal intensity changes were monitored. The KO2 was dissolved by complexation with dibenzo-18-crown-6-ether.
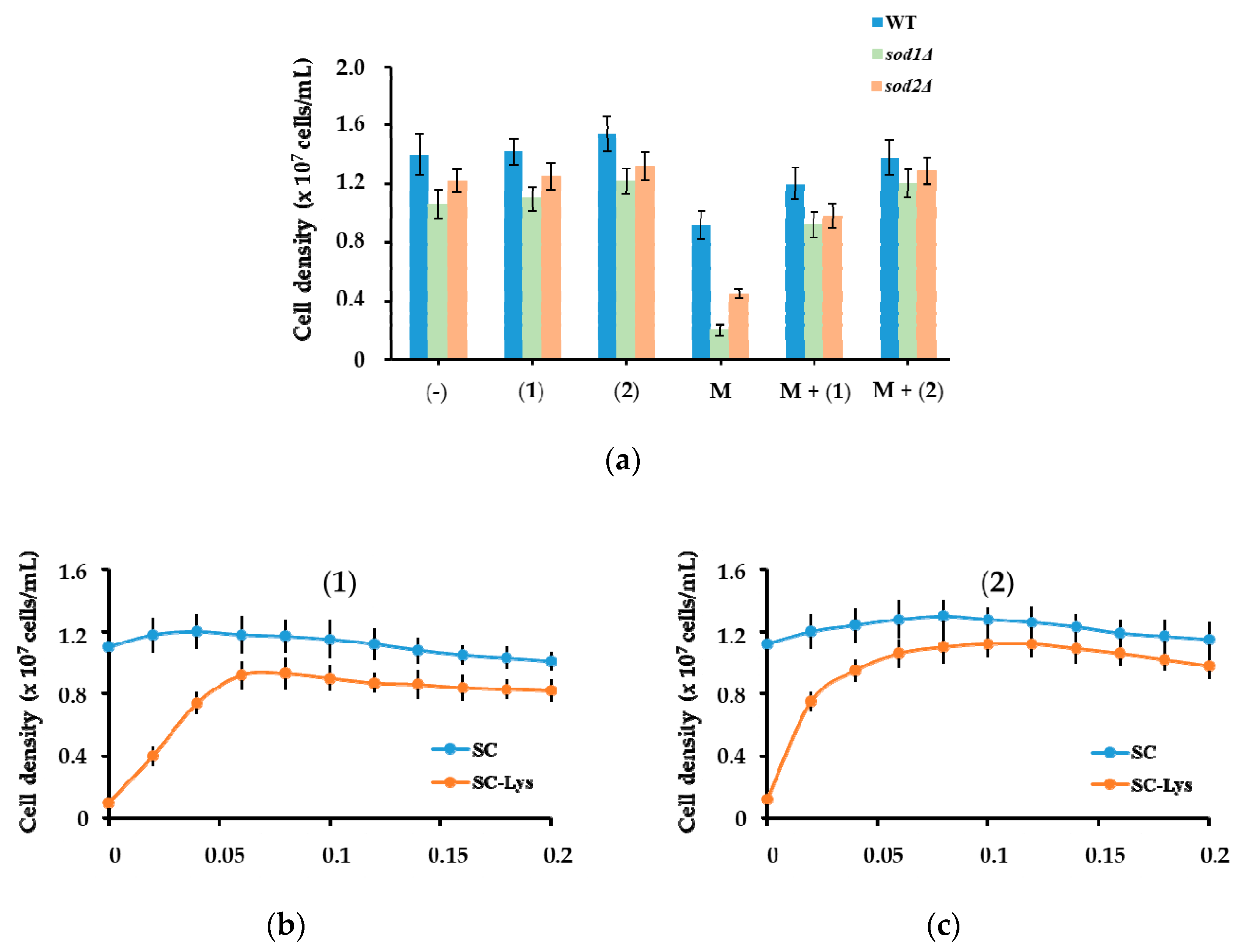
3.5.6. Yeast Cells Experiments
The
Saccharomyces cerevisiae laboratory strain BY4741 (
MATa;
his3Δ1;
leu2Δ0;
met15Δ0;
ura3Δ0) considered wild type (WT), and the isogenic strains
sod1Δ and
sod2Δ lacking the genes
SOD1 (encoding Cu/Zn-superoxide dismutase) and
SOD2 (encoding Mn-superoxide dismutase) were obtained from EUROSCARF (Frankfurt, Germany). Cells were maintained, manipulated and grown in a rich medium (YPD, 1%
w/
v yeast extract, 2%
w/
v peptone, 2%
w/
v glucose; 2%
w/
v agar was added for solid media) or on synthetic complete medium (SC, 0.67%
w/
v yeast nitrogen base without amino acids, 2%
w/
v glucose) supplemented with the necessary amino acids [
66]. Lysine drop-out medium (SC-Lys) was prepared similarly to SC, omitting lysine from the recipe. We added the complexes to autoclaved media from DMSO sterile stocks (10 mM). An overnight culture was 10
3-fold diluted in fresh medium and incubated (200 rpm, 30 °C) until the culture reached a density of 5 × 10
5 cells. The compounds were added to the desired concentrations, and culture growth was monitored at 660 nm [
67] in a 96-well multi-scanner plate reader equipped with a thermostat and shaker (Varioskan, Thermo Fischer Scientific, Vantaa, Finland). Three biological replicates were set for all determinations, and data were presented as average ± standard error (SE). The
Saccharomyces cerevisiae laboratory strain BY4741 (
MATa;
his3Δ1;
leu2Δ0;
met15Δ0;
ura3Δ0) considered wild type (WT), and the isogenic strains
sod1Δ and
sod2Δ lacking the genes
SOD1 (encoding Cu/Zn-superoxide dismutase) and
SOD2 (encoding Mn-superoxide dismutase) were purchased from EUROSCARF (Frankfurt, Germany). Cells were maintained and manipulated and grown in YPD medium (1%
w/
v yeast extract, 2%
w/
v peptone, 2%
w/
v glucose; 2%
w/
v agar was added for solid media) [
66]. Complexes were added to autoclaved media from DMSO sterile stocks (10 mM). To determine the effect of compounds on yeast growth, a fresh overnight pre-culture was 10
3-fold diluted in YPD and incubated (200 rpm, 30 °C) until the culture reached a density of 5 × 10
5 cells/mL. The compounds were added at the desired concentrations, and cell proliferation was monitored at 660 nm [
60] in a 96-well multi-scanner auto reader plate reader equipped with a thermostat and shaker (Varioskan Thermo Fischer Scientific, Vantaa, Finland). All determinations were done on three biological replicates.
3.5.7. Spectroscopic Determination of the Compounds’ Interactions with DNA
To determine if the compounds can interact with DNA, the 48,500 base-pairs genomic DNA from bacteriophage lambda (λ-DNA, Promega, MA, USA) was used. A stock solution of 10 µM λ-DNA (base-pair concentration) was prepared in 10 mMTris/HCl buffer containing 1 mM NaCl, pH 8.
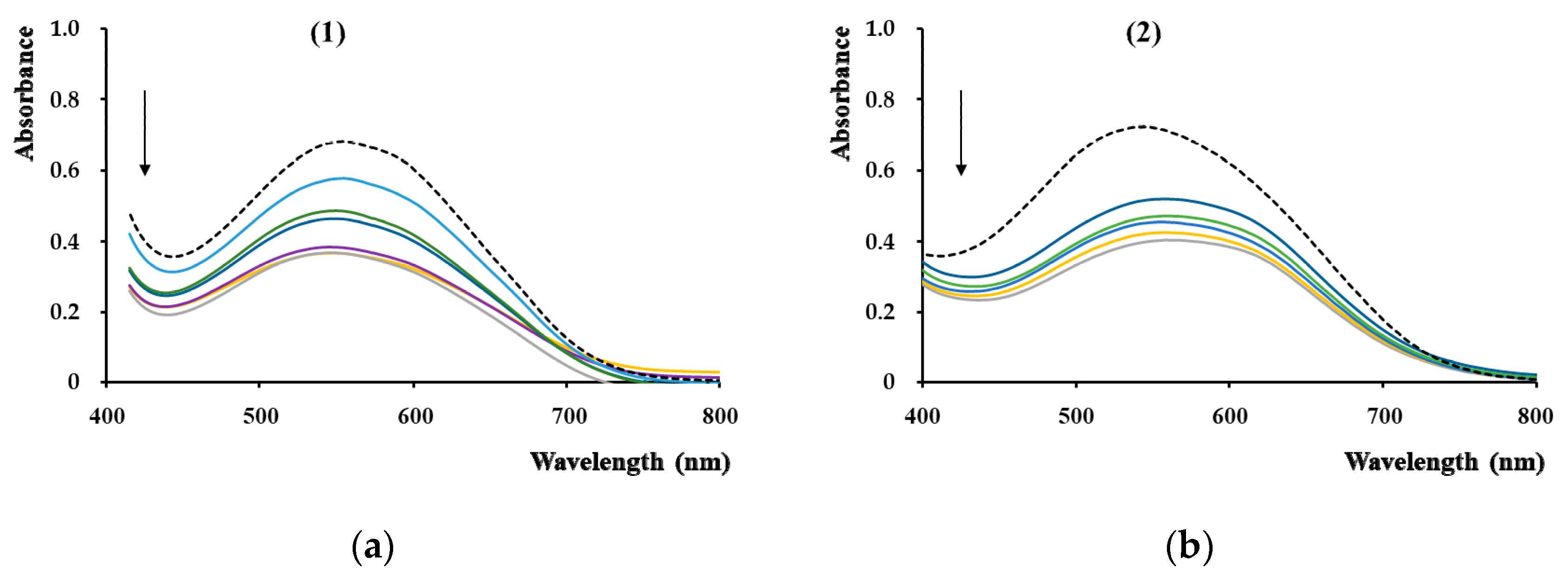
3.5.8. Effect of DNA on Spectral Characteristics of Compounds
To determine the DNA intercalating ability of complexes (
1) and (
2), we used the 48,500 base-pairs genomic DNA from the bacteriophage lambda (λ-DNA, Promega, Madison, USA). A 10 µM λ-DNA (base-pair concentration) stock solution was prepared in 10 mM Tris/HCl buffer, pH 8, containing 1 mM NaCl. The UV-visible absorption titration experiments were done using a constant complex concentration (5 µM). The concentrations of the λ-DNA solutions were gradually increased until a saturation state was achieved [
68,
69]. After DNA addition, the solutions were left to equilibrate for 5 min at room temperature before recording the spectra with a Jasco V-630 spectrophotometer. Stock solutions of compounds (
1) and (
2) were prepared in DMSO (1 mM) and diluted to 5 µM in 10 mM Tris/HCl containing 1 mM NaCl, pH 8.
The UV-Vis absorption titration experiments were done, keeping the complex concentration constant (3 µM) and increasing the concentration of λ-DNA in Tris-HCl/NaCl buffer solution at room temperature until a saturation state was achieved [
63,
64]. After DNA addition, the solutions were allowed to equilibrate for 5 min at room temperature. The UV-Vis absorption spectra were recorded with a Jasco V-630 spectrophotometer, using a 10 mm quartz cell. Stock solutions of compounds were prepared in DMSO (1 mM) and diluted to 3 µM in 10 mMTris/HCl containing 1 mM NaCl, pH 8.
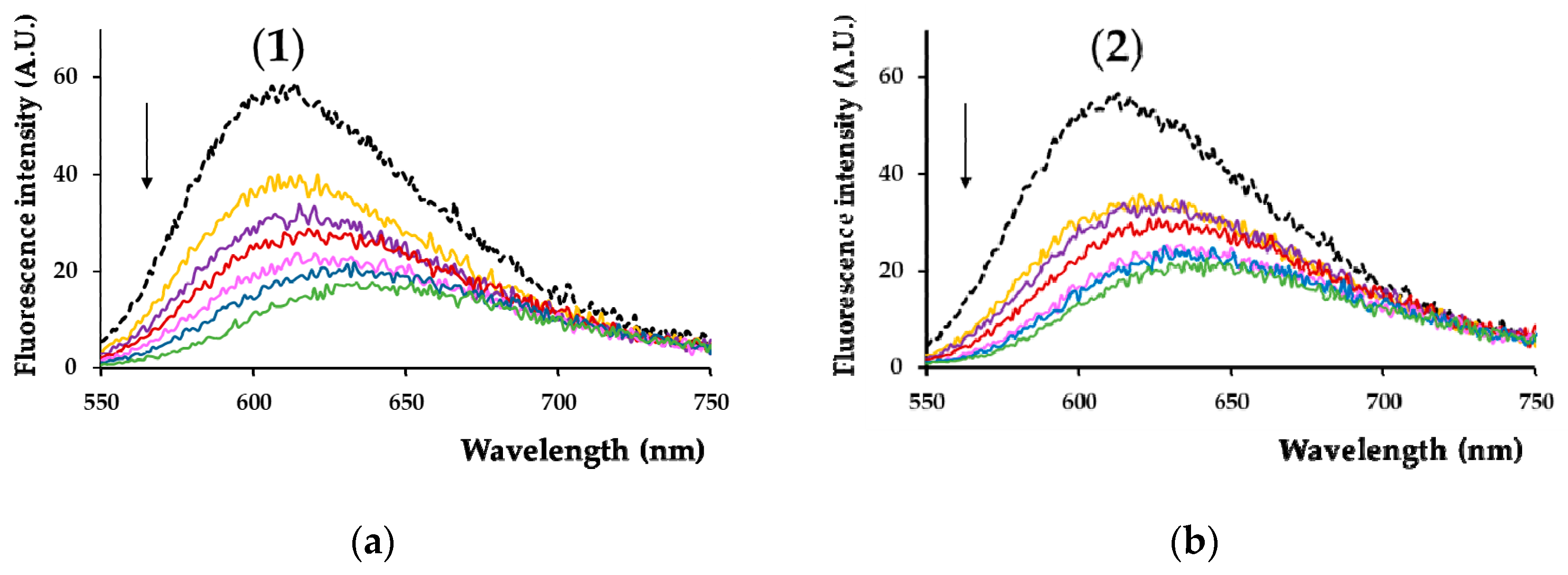
3.5.9. Effect of Compounds on the Fluorescence of λ-DNA/Ethidium Bromide Adduct
The binding of complexes (
1) and (
2) to DNA was assayed by monitoring the quenching of fluorescence emitted by λ-DNA/ethidium bromide (EB) adduct (λ
excit = 510 nm, maximum emission at λ
em = 601 nm) in the presence of different concentrations of the tested compounds [
68,
69]. The fluorescence spectra were recorded in the range of 550–750 nm using a Thermo Fischer Scientific Varioskan Flash spectral scanning multimode reader (Vantaa, Finland). The spectra were recorded in suitable plates using 5 nm excitation and emission slits for all measurements.
The binding of compounds to λ-DNA was assayed by monitoring the quenching of fluorescence emitted by λ-DNA/ethidium bromide (EB) adduct (λ
excit = 510, maximum emission at λ
em = 601). The λ-DNA solution was added to the EB solution, prepared in the same buffer, then increasing concentrations of compounds were added to EB-DNA [
52,
70]. The fluorescence spectra were recorded in the range of 550–750 nm using a Thermo Fischer Scientific Varioskan Flash spectral scanning multimode reader (Vantaa, Finland). The spectra were recorded in suitable plates using 5 nm excitation and emission slits for all measurements.
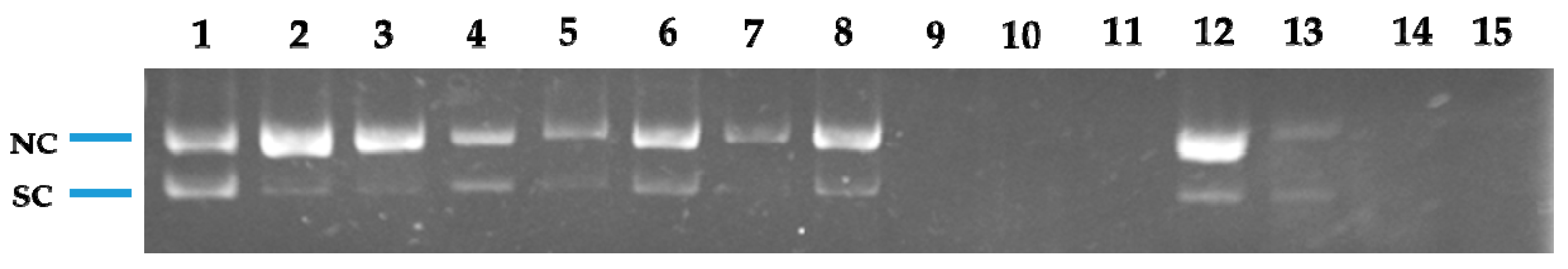
3.5.10. Nuclease Activity Assay
The nuclease activity of the compounds was determined using plasmid pRS325II DNA, which was a gift from Steven Haase (Addgene plasmid #35467, 6835 base pairs) [
71]. The plasmid was amplified in
Escherichia coli by transforming One Shot
® TOP10 chemically competent
E. coli (Invitrogen, Thermo Fisher Scientific). The plasmid was isolated from positive colonies using a PureLink™ HiPure Plasmid Miniprep Kit (Invitrogen, Thermo Fisher Scientific). Plasmid (200 ng/sample) was exposed to complexes (
1) and (
2) in the presence of 1 mM hydrogen peroxide (to exploit the Cu(II/III) redox couple) or 1 mM ascorbic acid (to exploit the Cu(I/II) redox couple) [
46]. The DNA cleavage experiments were done in a 9/1 (
v/
v) ratio of 50 mM Tris-HCl, pH 8, and DMSO. The samples were incubated for 1 h at 37 °C before a bromophenol blue/xylene cyanol-based loading dye (Roth, Germany) was added. The samples were loaded onto a 1% (
w/
v) agarose gel containing 1 μg/mL EtBr in 1 × TBE (Tris–boric acid–EDTA) buffer. Electrophoresis was performed at 50 V for 60 min in 1 × TBE buffer. The images of the fluorescent ethidium bromide-stained gels were captured using a gel documentation system (Doc-Print II, VilberLourmat, France). The cleavage experiments were done three times, with similar results. One representative gel is shown.
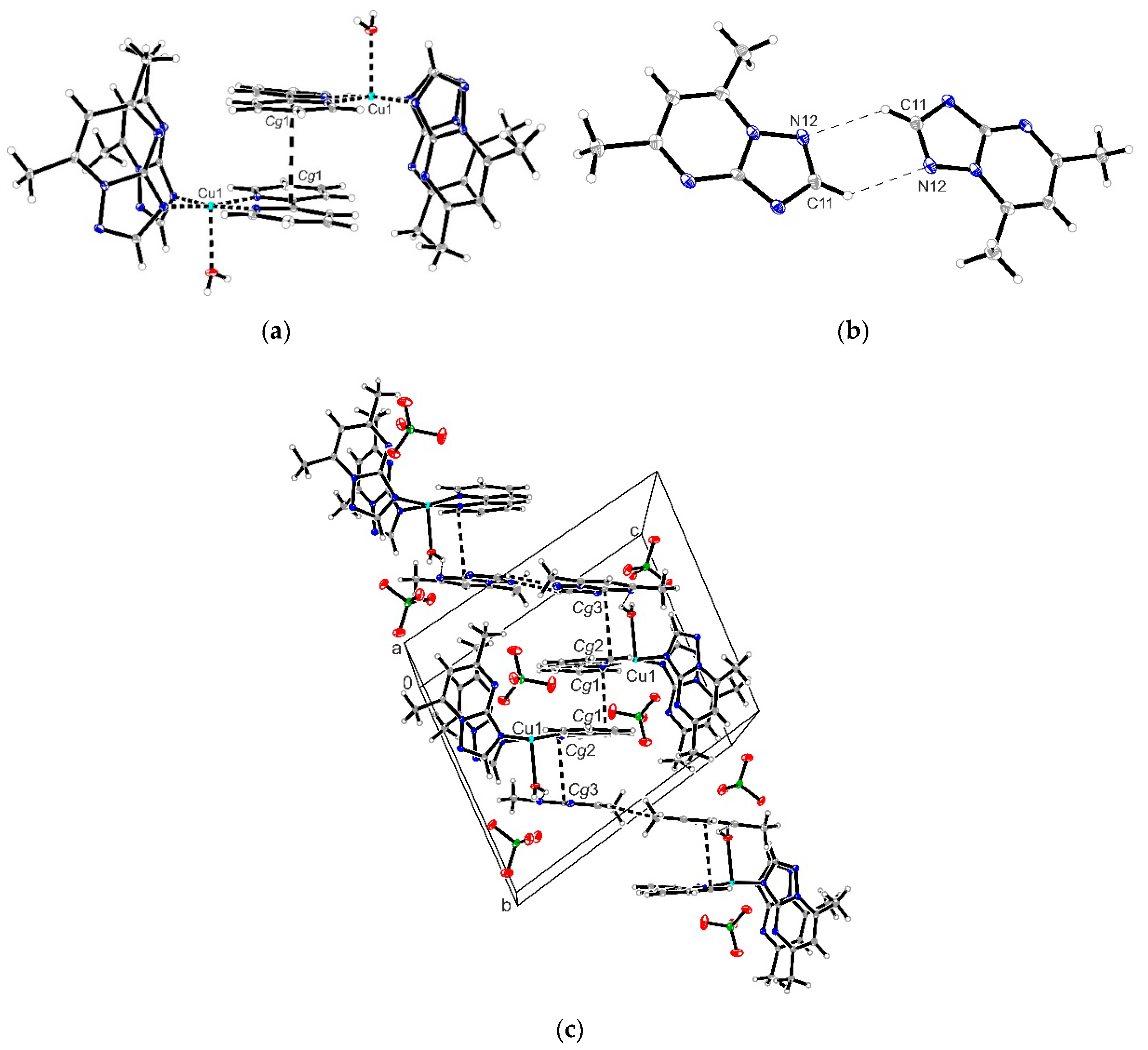
4. Conclusions
A novel series of copper(II) complexes with mixed ligands (5,7-dimethyl-1,2,4-triazolo[1,5-a]pyrimidine and 2,2′-bipyridine/1,10-phenanthroline) were stepwise synthesized, and structurally characterized. A complex supramolecular network is generated through π-π stacking interactions realized between both 2,2′-bipyridine/1,10-phenanthroline belonging to different cationic units as well as between these and free triazolopyrimidine moieties. Complex (1) exhibits the ability to reduce the viability of B16 cells in a dose-dependent manner and furthermore does not show any significant toxicity against BJ and Saccharomyces cerevisiae cells in the concentration ranges proven for the manifestation of biological activity. The value of the therapeutic index value and morphological modifications indicate that complex (1) was more susceptible than (2). Both complexes exhibited improved antimicrobial activity, as compared to the triazolopyrimidine ligand. Complex (2) was by far more active than (1) against the Gram-positive strains, susceptible or resistant, growing in planktonic or in biofilm states. Both MIC and MBEC values decreased in the case of complex (2) by 4 to 17 times compared to (1). The results suggest that both the antitumor and antimicrobial activity of these complexes involves intercalation into DNA strands as well as its damage through metallonuclease activity. Nuclease-like activity is enhanced in the presence of hydrogen peroxide, the complexes’ interactions with this ROS also being proved by EPR and cyclic voltammetry data. Overall studies indicate the antitumor potential of complex (1) and the antibacterial potential of complex (2), especially against the Gram positive strains. As a result, these species will be further studied in order to improve their solubility, and consequently their activity, by their inglobation in organic matrix.