1. Introduction
When newly translated secretory proteins are inserted into the ER, quality control must occur to ensure that misfolded proteins do not accumulate and disturb ER function. In the ER, protein folding is continuously monitored by molecular chaperones. When a protein fails to acquire its correct conformation, it is transferred to the ER-associated protein degradation (ERAD) pathway. Substrates of ERAD are transported over the ER membrane towards the cytosol, where they are ubiquitinated and degraded by the proteasome.
With over 70 diseases associated with ERAD [
1,
2], including cystic fibrosis and Parkinson’s disease, a better understanding of this protein degradation pathway is required. Because viruses depend on and manipulate their host cells, they provide useful models to study a wide range of cellular processes. Indeed, many viruses exploit ERAD to facilitate virus replication [
3] or to evade immune recognition [
4,
5]. These manipulation strategies can be exploited to study protein degradation.
Human cytomegalovirus (HCMV) is a herpesvirus that causes severe congenital defects when it infects pregnant women [
6]. The virus can successfully evade the immune system, allowing it to persist in the body lifelong. HCMV induces accelerated ERAD of HLA class I molecules to prevent recognition of virus-infected cells by CD8
+ T lymphocytes. The viral proteins responsible for this, US2 and US11, serve as important models to study the degradation of ER-resident proteins. These HCMV proteins have allowed the identification of many key mammalian ERAD factors, including Derlins, VIMP [
7,
8], and the ubiquitin E3 ligases TRC8 [
9] and TMEM129 [
10,
11].
Despite the identification of a number of factors involved in ERAD, many questions remain to be answered. It is thought that the protein complexes required for ERAD are (partially) specific to the substrate that is degraded, in combination with some general ERAD players such as p97/VCP and the proteasome. While the ubiquitination machinery for US2-mediated degradation of HLA class I has been identified [
9,
10,
11,
12,
13], knowledge about other ERAD factors is lacking.
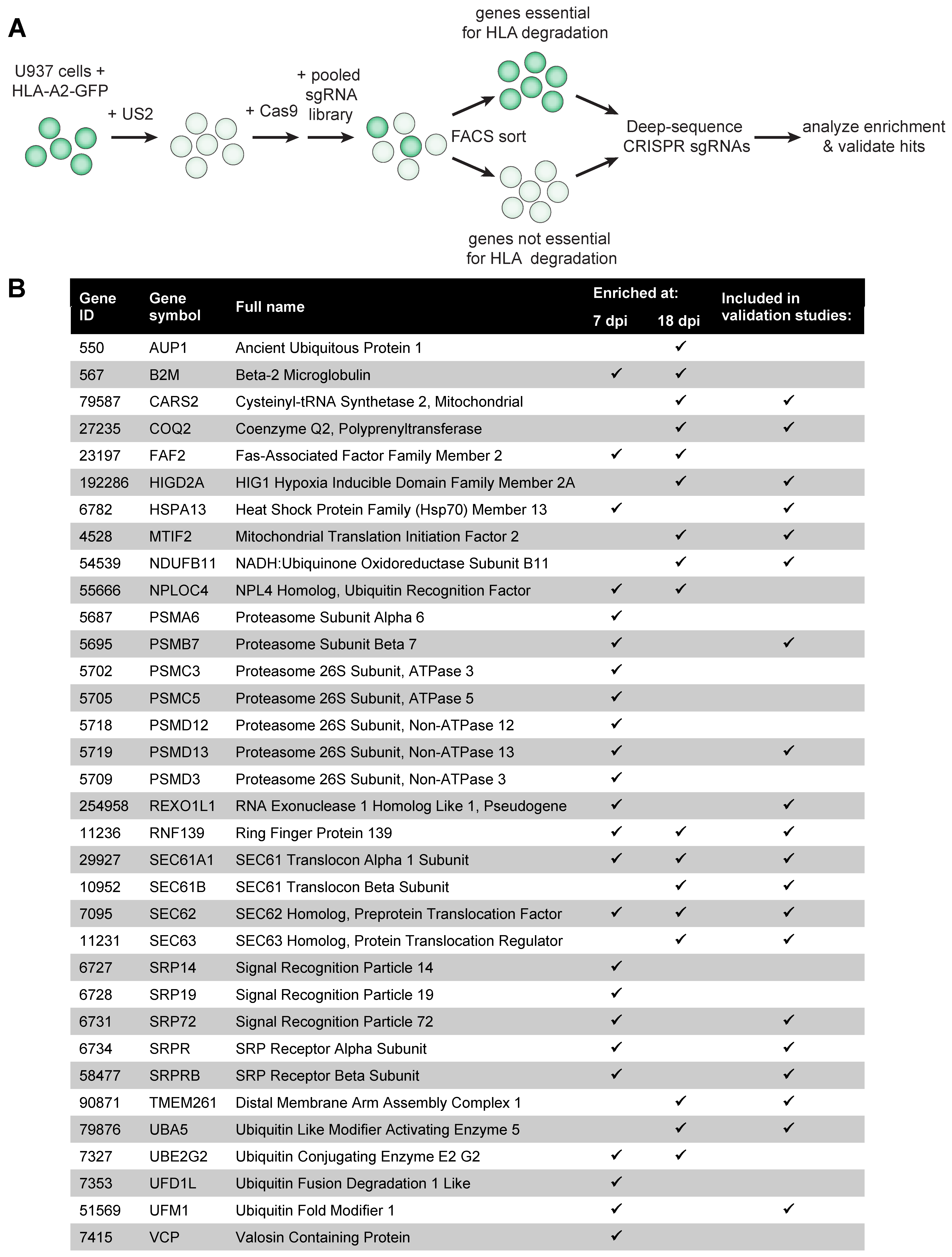
Here, we performed a genome-wide CRISPR/Cas9 library screen to identify novel host genes involved in US2-mediated HLA class I degradation. Notably, besides known ERAD-related factors, we identified all known factors of the UFMylation pathway to affect HLA-I degradation via US2. UFMylation is a ubiquitin-like post-translational modification that involves covalent attachment of ubiquitin-fold modifier 1 (UFM1), a ~9.1 kDA protein, to target proteins. Under the experimental conditions used, however, we did not detect direct UFMylation of HLA-I or other proteins involved in ERAD. A mass spectrometry analysis showed that UFMylation predominantly occurs on ribosomal proteins in US2-expressing cells. We therefore speculate that UFM1 may affect US2-mediated protein degradation indirectly, via a mechanism potentially involving the ribosome.
3. Discussion
Here, we describe a genome-wide CRISPR/Cas9 screen to identify cellular factors involved in HCMV US2-mediated ERAD of HLA-I. We identified multiple genes that were previously linked to US2-mediated HLA-I downregulation, including the ubiquitin ligase TRC8, the E2 enzyme UBE2G2, and p97, the ATPase facilitating dislocation of ERAD substrates. Some genes, such as UBXD8 (FAF2) and the p97 co-factors Npl4 and Ufd1, have previously been described in ERAD, but did not affect US2 function in those studies [
14,
18]. Although we have not validated these hits in detail, targeting them with CRISPR sgRNAs results in abrogation of US2-mediated HLA class I downregulation [
16] (unpublished). The discrepancy with previous studies may arise from the different techniques used: in contrast to our knockout approach, Npl4 and Ufd1 were previously knocked down using siRNAs [
14], which does not result in full target knockdown, while UBXD8 was studied in a pulse-chase approach to assess HLA class I dislocation [
18].
We show that knockout of genes involved in the UFM1 pathway moderately, yet consistently, hamper US2-mediated HLA-I degradation. A link between UFM1 and the ER has previously been described [
27], yet a role in protein degradation has been reported only recently [
25,
28]. UFM1 is a post-translational modifier structurally related to ubiquitin [
29]. Similar to ubiquitin, UFM1 is conjugated to its substrates via an iso-peptide bond between the C-terminal glycine of UFM1 and a lysine residue of the substrate [
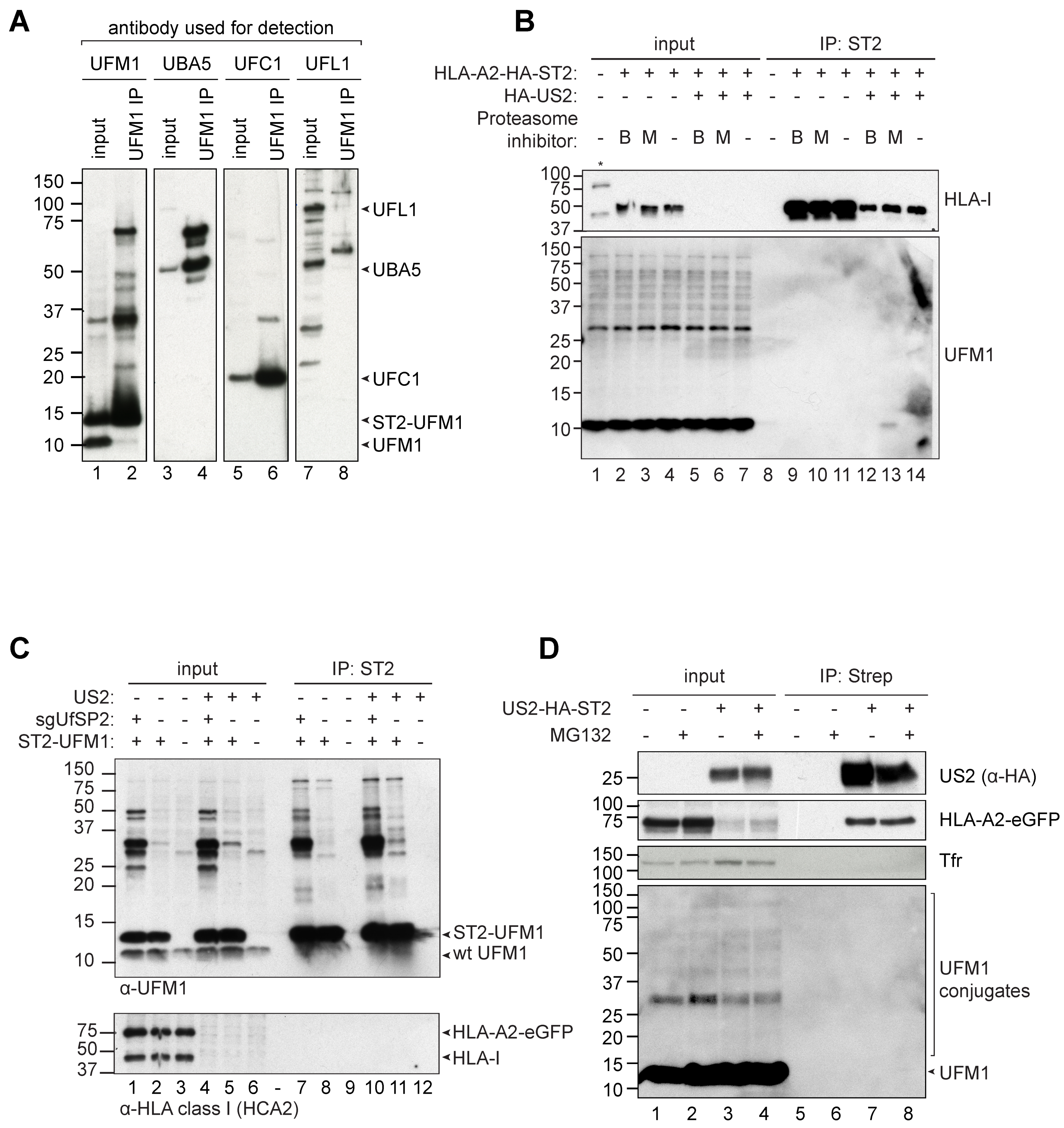
30]. For both modifiers, the conjugation to substrates is facilitated by E1, E2 and E3 enzymes. However, UFM1 does not seem to function in ERAD the same way as ubiquitin: while the degradation substrate HLA-I becomes ubiquitinated, UFMylation of HLA-I could not be detected. Similarly, UFMylation did not take place on US2 or proteins directly related to US2-mediated ERAD. Therefore, the mechanism behind UFM1′s impact on protein degradation remains to be clarified. This, in combination with the subtle HLA-I rescue phenotypes observed upon knocking out players of the UFMylation pathway, suggests that UFM1 may play an indirect role in protein degradation, potentially by affecting the protein sorting mechanisms of the cell.
The results from our genome-wide CRISPR/Cas9 library screen are in agreement with screens that have been performed using different model systems. One of these recently described genome-wide CRISPR/Cas9 library screens has identified the UFM1 pathway to regulate SQSTM1 expression in an ER-stress-dependent manner [
31]. Another screen, aimed to identify genes involved in ERAD, identified a similarly modest role for UFM1 [
25]. Several additional studies have attempted to identify targets for UFMylation, mostly by mass spectrometry-based approaches. Despite these efforts, only few UFM1 substrates have been identified to date. The first-identified UFM1 target [
21], UFBP1, was later suggested to play a role in the UFMylation pathway itself [
20,
27,
32]. Other targets include LZAP, a binding partner of UFL1, the ribosome [
24,
25], and, interestingly, multiple chaperones of the Hsp40 and -70 families, such as DNAJC1, HSPA8, and BiP [
20,
32]. While our genome-wide screen also identifies a Hsp70 (HSPA13) and Hsp40 chaperone (SEC63), they did not associate with UFM1 in our mass spectrometry analysis. Although UFM1 is ubiquitously expressed in many tissues [
21], its target proteins may differ between cell types and the experimental context used. In B cells in particular, UFMylation of UFBP1 has been shown to act upon different branches of the unfolded protein response (UPR) to promote the differentiation and function of plasma cells [
33].
We identified many ribosome subunits as potential targets for UFMylation in US2-expressing cells. UFMylation of RPS3, RPS20 and RPL10 [
24], and of RPL26 [
25,
28] has been described previously, suggesting that these ribosomal proteins are genuine targets of UFMylation. Because RPS3, RPS20 and RPL10 are located near the mRNA entry channel in the large ribosomal subunit, it has been suggested that UFMylation may affect mRNA entry into the ribosome [
24]. RPL26 on the other hand is located closely to the polypeptide exit tunnel. This, in combination with the finding that UFMylated RPL26 interacts with the SEC61 complex, suggests that UFMylation of the ribosome may regulate translation and translocation efficiency, although no direct effect on protein translation could be observed [
25,
28]. The changes US2 promotes in the ER upon directing HLA class I for degradation might consequently affect protein sorting indirectly, in a UFM1-dependent manner. As UFMylation of the ribosome may affect translation efficiency [
24,
25,
28], and US2 may be particularly sensitive to changes in translational efficiency as a consequence of its inefficient signal peptide [
16] (preprint), the rescue phenotype could alternatively arise from altered US2 expression. Due to the low expression levels of US2 and the lack of a suitable detection antibody, we were unable to assess this option. Finally we cannot exclude that changes in the UFMylation pathway may alter the translation efficiency of HLA-I itself, and of ERAD-related proteins.
In the same screen that identified RPS3, RPS20 and RPL10, UFMylation of eIF6 was observed [
24]; eIF6 is a translation initiation factor that prevents association between the 40S and 60S ribosomal subunits. We identified eIF1AX, another translation initiation factor associated with the 40S ribosome. None of the proteins found to interact with UFM1 in our mass spectrometry studies are functionally related to ER-associated degradation, and it was recently suggested that the identification of UFM1 in the context of ERAD may be an indirect consequence of UFMylation being related to maintaining ER homeostasis [
25], which may be perturbed when inducing protein degradation. Upon chemically-induced ER stress, UFM1 is upregulated via the transcription factor Xbp1s [
34], and UFMylation allows cells to survive ER stress by suppressing apoptosis [
20,
27,
32,
35]. More specifically, UFBP1 (also known as DDRGK1 or C20orf116) is an ER membrane protein that binds the ER stress protein IRE1α in an UFM1-dependent manner [
27]. IRE1α in turn cleaves Xbp1, that functions as a transcription factor to activate ER chaperones as well as UFM1 [
34]. A positive feedback loop may arise during ER stress, as the elevated expression of UFM1 potentially stabilizes additional IRE1α molecules.
The UFMylation pathway could be related to ER stress via UFBP1. In the absence of UFM1, IRE1α is a substrate for SEL1/HRD1-mediated ERAD. IRE1α is not UFMylated itself, but it is rescued from degradation by binding to UFMylated UFBP1. Hence, depending on the UFMylation status of UFBP1, the protein can rescue ERAD substrates from degradation. Also the ER chaperone BiP has been shown to be an indirect target of UFMylation by interacting with UFBP1 [
32]. UFBP1 may thus act as a regulator of protein stability, depending on its UFMylation status. By switching the UFMylation status of UFBP1, a far larger number of targets may be regulated via protein-protein interactions without the need for direct UFMylation of these target proteins themselves. In fact, it has been recently shown that UFBP1/DDRGK1 brings the E3 ligase UFL1 to the ER surface, where RPN1 and RPL26 are UFMylated. This UFMylation at the ER seems to facilitate ER-phagy in conditions of starvation and to repress the unfolded protein response [
36].
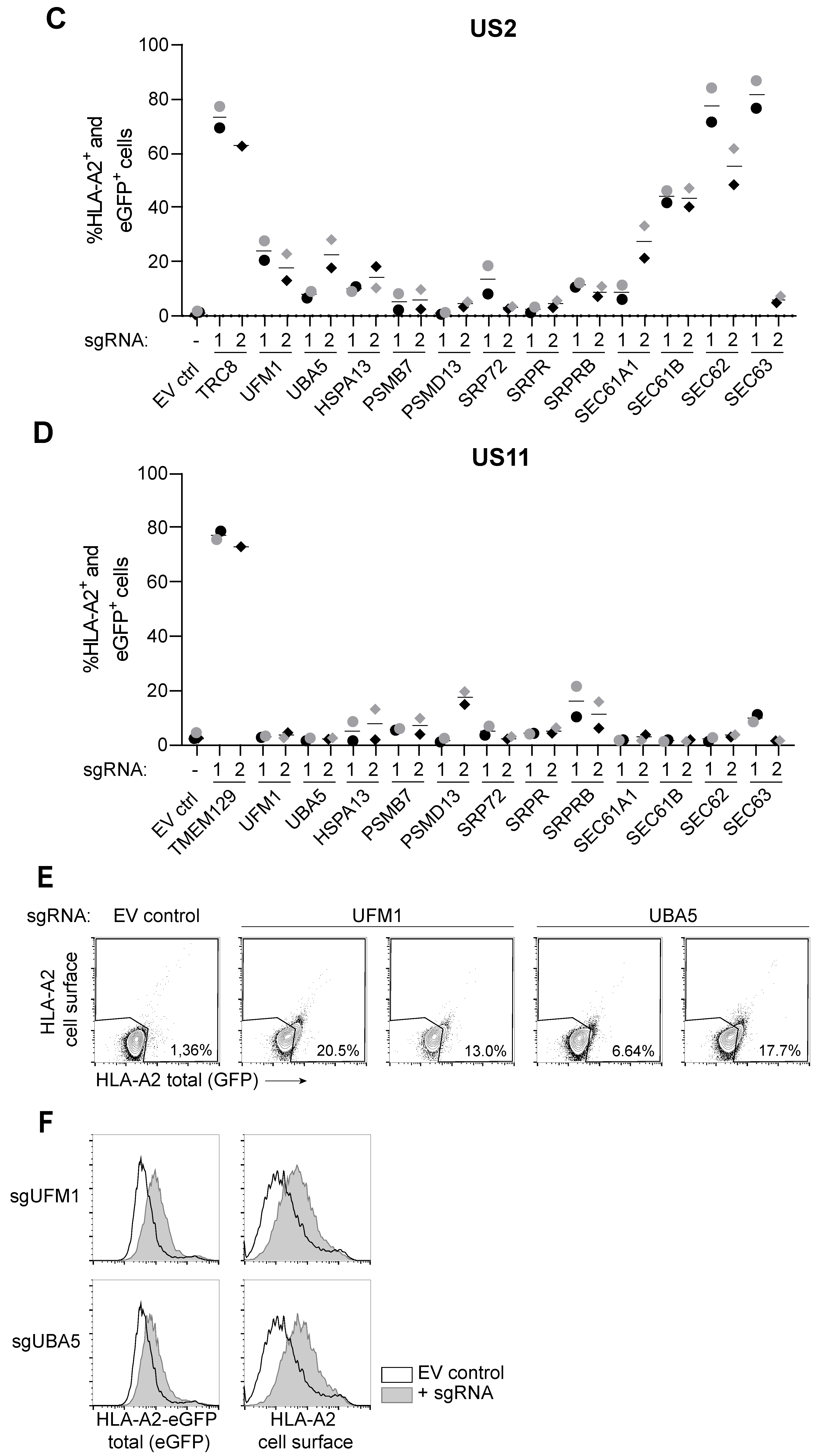
In addition to the underlying mechanism, the substrate specificity of the UFM1 pathway remains to be established. In our unbiased CRISPR/Cas9 screen, all known factors of the UFMylation pathway were found to affect HLA-I degradation via US2, but not US11, suggesting substrate specificity. The contribution of the UFM1 pathway to degradation of other cellular proteins remains to be established.
US2-mediated HLA-I downregulation at the cell surface is partly restored upon knockout of genes involved in the UFMylation pathway. Also total levels of HLA-I increase within the cell upon inhibition of the UFM1 pathway. In the absence of UFMylation, US2-dependent ER-to-cytosol dislocation appears to be compromised. However, nor HLA class I, nor US2, nor proteins previously shown to be involved in US2-mediated ERAD, were found to be UFMylated. The impact of the UFM1 pathway on HLA-I degradation may, therefore, be indirect: UFMylation is known to play an important role in maintaining ER homeostasis, and US2-mediated protein degradation likely acts as a disturbing factor. The ins-and-outs of this dysregulation require further investigation.
4. Materials and Methods
4.1. Cell Culture and Lentiviral Transduction
U937 cells (ATCC) were cultured in RPMI 1640 culture medium (Gibco) supplemented with penicillin/streptomycin (Gibco), Ultraglutamine-1 (Gibco) and 10% fetal calf serum (BioWest). Wildtype U937 cells were lentivirally transduced with HLA-A2-eGFP (kindly provided by Dr. Louise Boyle, University of Cambridge UK) expressed from a lentiviral pSicoR vector containing a hygromycin B resistance gene. Successfully transduced cells were selected with Hygromycin B at 3 days post-infection, and subcloned. The HLA-A2-eGFP cells were subsequently transduced with US2 under control of an EF1a promoter (pSicoR-EF1A-US2 (RP-549)) and a clonal cell line was established by fluorescence-activated cell sorting (FACS) of the HLA-A2-eGFPlow cells. Knock-out (KO) of genes involved in the UFMylation pathway was achieved by CRISPR/Cas9 guided genome editing (see below for details). Rescue of protein expression in KO cells was achieved by expression of sgRNA-resistant cDNAs from the BIC-PGK-Zeo vector.
U373 cells (ATCC) were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Gibco) supplemented with penicillin/streptomycin (Gibco), Ultraglutamine-1 (Gibco) and 10% fetal calf serum (BioWest). U373 cells were transduced with the abovementioned HLA-A2-eGFP vector and selected in the same manner (Hygromycin B at 3 days post-infection). A polyclonal cell line was established after fluorescence-activated cell sorting (FACS) of the HLA-A2-eGFP+ cells. Cells were subsequently transduced with an HA-tagged US2 and selected with Blasticidin at day 3 post-infection. CRISPR/Cas9 genome editing was performed as described below.
4.2. Plasmids
HLA-A2-eGFP (kindly provided by Dr. Louise Boyle, University of Cambridge UK) was subcloned in a bidirectional lentiviral expression vector derived from no.2025.pCCLsin.PPT.pA.CTE.4 × -scrT.eGFP.mCMV.hPGK.NG-FR.pre (kindly provided by L. Naldini, San Raffaele Scientific Institute, Milan, Italy) as described elsewhere [
10]. This lentiviral vector (BIC HLA-A2-EGFP_HygB-T2A-mCD4 (RP-31)) contains a human EF1A promoter to facilitate potent expression of HLA-A2-eGFP and expresses an Hygromycin B resistance gene fused with a T2A sequence to a tail-less mouse CD4 under control of a different promoter (PGK).
US2 was expressed from several backbone vectors. Untagged US2 was expressed in U937 cells from the pSicoR-EF1A-US2 vector (RP-549)). This vector was constructed from the pSicoR vector [
37], from which the U6 promoter was removed and the CMV promoter was replaced by an EF1A promotor. US2leader-HA-US2 was expressed in U373 cells from BIC-PGK-Blast (RP138), derived from no.2025.pCCLsin.PPT.pA.CTE.4 × -scrT.eGFP.mCMV.hPGK.NG-FR.pre. This lentiviral vector contains a human EF1A promoter to facilitate potent expression of the downstream cloned gene and expresses the BlastR selection marker from a different promoter (PGK).
For CRISPR/Cas9 genome engineering, single guide RNAs (sgRNAs) were introduced in the lentiviral pSicoR-CRISPR-PuroR vector (RP-557) [
38] as described previously. To allow for rescue of protein expression in KO cells, we cloned cDNAs, UFM1 and UFC1 inactive mutant controls in dual promoter lentiviral vectors (BIC-PGK-Zeo (RP137), which is the same vector as BIC-PGK-Blast (RP138), yet with ZeoR selection instead of BlastR. We introduced silent mutations in the sgRNA target sites to prevent CRISPR/Cas9 mediated editing of lentiviral cDNA sequences in CRISPR/Cas9-harbouring KO cells.
4.3. Genome-Wide CRISPR/Cas9 Library Screen
150 million U937 HLA-A2-eGFP cells co-expressing US2 and SpCas9 were transduced at an MOI of 2 in duplicate with the human GeCKOv2 CRISPR knockout pooled library (obtained from Feng Zhang (Addgene, Watertown, NY, USA). The library targets 19,050 genes with 6 sgRNAs/gene. Transduced cells were selected by puromycin treatment (2 μg/mL) at 2 days post-infection (d.p.i.) and maintained at high complexity for the duration of the screen. At 7 and 18 d.p.i., 1 billion cells were harvested and subjected to cell sorting via a two-step sort-protocol using a Becton Dickinson Influx cell sorter. First, PE
+ cells were sorted using an ‘enrichment-protocol’, which allowed for high-speed cell sorting of the entire population of cells in a short timeframe. Next, we sorted the top ±1% of eGFP
+/PE
+ cells to purity selecting for cells that display enhanced levels of eGFP and HLA-A2 surface staining (stained with BB7.2-PE; BD Biosciences, #558570). As control, the eGFPlow/HLA-A2low were sorted. We next isolated genomic DNA from all sorted cell populations by standard phenol/chloroform extraction protocols using the Phase Lock gel heavy tubes (Quantabio; 10847-802) according to manufacturer’s instructions. Next, the lentiviral sgRNA inserts were PCR-amplified for 27 cycles using Fw primer 5′-
AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCTNNNNNNcttgtggaaaggacgaaacacc-3′ and Rev primer 5′-
CAAGCAGAAGACGGCATACGAGATgactcggtgccactttttcaag-3′ and Phusion polymerase (NEB) in the presence of buffer GC supplemented with DMSO. Both primers contain an Illumina adapter sequence (displayed in bold) that allows for direct loading on an Illumina NextSeq500 sequencer and a lentiviral-specific primer binding site (lowercase letter) to facilitate amplification of the integrated lentiviral sgRNA sequence. The Fw primer also contains a unique 6nt barcode sequence (NNNNNN) allowing for multiplexed sequencing, and a primer binding site for the Illumina sequencing primer (underlined). The PCR products were purified/concentrated using a PCR purification kit (Qiagen, Hilden, DE), and subsequently loaded on a 20% polyacrylamide gel in 0.5 × TBE. Bands of the correct size were excised, electro-eluted, purified by phenol-chloroform extraction and subsequently quantified using a Nanodrop quantification device (Nanodrop, Rockland, DE, USA) and an Agilent bioanalyzer (Agilent Technologies, Palo Alto, CA, USA). Deep sequencing was carried out as single 75bp run on a Illumina NextSeq500 machine (performed by the Utrecht Sequencing facility USEQ) using the sequencing primer 5′-ACACTCTTTCCCTACACGACGCTCTTCCGATCT-5′, in which the Index and sgRNA sequence was sequenced simultaneously. Due to the low complexity at the start of the sequence, a Phix library was mixed with the libraries to 20% of total reads. Sequences were aligned to the sgRNA library by using Bowtie2 [
39] and the counts per sgRNA were calculated. We used the MaGeCk package [
40] (available from
https://sourceforge.net/projects/mageck/) as a computational tool to identify genes that were significantly enriched in the screens by comparing sgRNA read counts of control sorted cells to cells displaying enhanced HLA-A2-eGFP levels. The overlap between the top 300 of the two duplicates was compared and used to select genes for further validation. The hits in this list were ranked, based on the number of sgRNAs that was enriched >5-fold, and the number of sgRNAs that showed >20-fold enrichment. As every gene was targeted with 6 sgRNAs in duplicate, the genes that showed >5-fold enrichment in at least 6 out of the total 12 sgRNAs (from both duplicates) were selected for further validation. In total, this list contained 46 genes. For initial validation studies, two sgRNAs/gene from the GeCKOv2human library that yielded the highest enrichment were selected and cloned into a pSicoR lentiviral vector with an EFS-PuroR-T2A-Cas9 cassette. sgRNA sequences are listed in
Supplementary information 1. sgRNAs were transduced in target cells, and transduced cells were selected for by puromycin selection (2 μg/mL). HLA-A2-eGFP rescue of these hits was validated based on eGFP intensity and an HLA-A2-specific antibody staining on the cell surface, using a flow cytometric readout (BD FACS Canto II). When setting up the validation, some gene-knockouts resulted in strong autofluorescence, indicating that these were false-positive hits. We therefore included the irrelevant PE-Texas Red channel to omit PE-Texas Red
+ cells prior to assessing HLA-A2-eGFP rescue. HLA-A2-eGFP expression was measured using the FITC channel for eGFP and the PE channel for HLA-A2 cell surface expression (using a PE-conjugated HLA-A2-specific antibody, BB7.2, see
Section 4.5). Genes that showed only autofluorescent signal were omitted from further analysis and are not shown in the list from
Figure 1B. Validation was performed on day 7, 11, 14, 18 and 28 post infection. Target site sequences of these additional sgRNAs are also listed in
Supplementary information 1.
4.4. Clonal Knockout Cell Lines
sgRNAs targeting UFM1-, UBA5-, and UFC1 were introduced in target cell lines by lentiviral transduction. At 3 days post-infection, transduced cells were selected for by Puromycin treatment (2 μg/mL). At 10 d.p.i., the sgUFM1 and sgUBA5 cells were stained for HLA-A2 cell surface expression and PE+/GFP+ cells were single-cell sorted by fluorescence-activated cell sorting (FACS) on a FACSAria III. Cells were allowed to recover for ~8 weeks and analyzed by flow cytometry to select cells that displayed enhanced HLA-A2-eGFP and endogenous HLA-A3 expression. The knock-out status was confirmed by Western blot and the genomic target sites of both alleles were sequenced by Sanger sequencing.
4.5. Antibodies
Antibodies used in this study were: rabbit anti-UFM1 [EPR4264(2)] (Abcam, ab109305), rabbit anti-UBA5 (Abcam, ab177507), rabbit anti-UFC1 [EPR15014-102] (Abcam, Cambridge, UK, AB189252), rabbit anti-UFL1 (Atlas Antibodies, Bromma, Sweden, HPA030559), mouse anti-human transferrin receptor (H68.4) (Invitrogen, Carlsbad, CA, USA, 13-6800), mouse anti-p97 (VCP) (BD Transduction Laboratories, San Jose, CA, USA, 612183), rat anti-HA (3F10) (Roche, Woerden, NL, 11867423001), mouse anti-HLA class I HCA2, mouse anti-HLA class I HC10, mouse anti-ubiquitin (P4D1) (Santa Cruz, Dallas, TX, USA, sc-8017), goat anti-rabbit-HRP (light chain-specific) (Jackson Immunoresearch, Ely, UK, 211-032-171), goat anti-rat-HRP (light chain-specific) (Jackson Immunoresearch, Ely, UK, 112-035-175), goat anti-mouse-HRP (light chain-specific (Jackson Immunoresearch, Ely, UK, 115-035-174), mouse anti-HLA-A2-PE (BB7.2) (BD Biosciences, San Jose, CA, USA, 558570), human anti-HLA-A3 (OK2F3), goat anti-human-PE (Jackson Immunoresearch, Ely, UK, 109-116-127).
4.6. Immunoblotting
When indicated, cells were incubated overnight with 500 nM MG132 (Sigma-Aldrich, Zwijndrecht, NL, C2211-5MG), 5 nM Bortezomib (New England Biolabs, Ipswich, MA, USA, 2204S) or DMSO control, prior to preparing cell lysates. To make lysates, cells were counted using a Casy cell counter and an equal number of live cells was subjected to two washes in PBS containing 20 mM N-ethylmaleimide (Sigma-Aldrich, Zwijndrecht, The Netherlands, E3876-5G) to block de-ubiquitinating and de-UFMylating activity. Subsequently, cells were lysed on ice in Triton X-100 lysis buffer (1% Triton X-100, Applichem Panreac, Darmstadt, DE, A13880500) 100 mM NaCl, 50 mM Tris, pH 7.5) supplemented with 20 mM N-ethylmaleimide. Samples were spun down at 12,000× g for 20 min at 4 °C to pellet cell debris and nuclei. Supernatant was transferred to a clean tube and mixed with Laemmli sample buffer containing DTT. Lysates were stored at −80 °C until further use. For Western blot analysis, proteins were separated using SDS-PAGE (Thermo Bolt 4-12% or self-made gels) and subsequently transferred to PVDF membranes (Merck Millipore, Darmstadt DE, IPVH00010). Membranes were blocked using 5% milk and incubated with the respective antibodies for specific protein detection. Protein bands were visualized using ECL (Thermo Scientific Pierce, Landsmeer, The Netherlands) on Amersham Hyperfilm ECL films (GE Healthcare, Buckinghamshire, UK). Samples subjected to either EndoH or PNGaseF treatment were incubated at 37oC for 1 h with EndoH (New England Biolabs, Ipswich, MA, USA, P0702L) or GlycosidaseF (Roche, Woerden, The Netherlands), respectively, in the appropriate glycobuffer. Quantification of total and deglycosylated HLA-I was performed using ImageJ software (National Institutes of Health, Bethesda, MD, USA).
4.7. Co-immunoprecipitation
Cells were lysed in 1% Digitonin (Calbiochem, Darmstadst, DE) lysis buffer (pH 7.5) containing 50 mM Tris-HCl, 5 mM MgCl2 and 150 mM NaCl, supplemented with 1 mM Pefabloc SC (Roche), 10 μM Leupeptin (Roche) and 20 mM N-ethylmaleimide (Sigma-Aldrich, Zwijndrecht, NL). Lysates were incubated on ice for 60 min and subsequently centrifuged at 12,000× g for 20 min at 4 °C to remove nuclei and cell debris. Post-nuclear lysates were incubated overnight with StrepTactin beads (GE Healthcare, Buckinghamshire, UK,). Beads were washed 4 times with 0.1% Digitonin lysis buffer, after which they were eluted for 45 min on ice. Elution buffer contained 2.5 mM d-Desthiobiotin, 150 mM NaCl, 100 mM Tris-HCl and 1 mM EDTA, at a pH of 8.0. The eluate was collected from the beads using SpinX columns (Corning Costar, Amsterdam, The Netherlands) and was denatured in Laemmli sample buffer containing DTT. Immunoblotting was performed as described before.
4.8. Mass Spectrometry
U937 cells containing HLA-A2-eGFP and US2 were transduced with either StrepII-tagged UFM1 wildtype cDNA or one of two control constructs: StrepII-tagged UFM1 ΔVGSC (inactive mutant lacking the 4 C-terminal residues) or StrepII-tagged mCherry. Per cell line, 50 million cells were pelleted in quadruplicate. Cell pellets were stored at −80 °C until lysis and immunoprecipitation. For affinity enrichment, pellets were lysed on ice using TAP lysis buffer (50 mM Tris pH 7.5, 100 mM NaCl, 5% (
v/
v) glycerol, 0.2 % (
v/
v) Nonidet-P40, 1.5 mM MgCl2, 1µg/mL Avidin (2-0204-015; IBA) and protease inhibitor cocktail (EDTA-free, cOmplete; Roche, Woerden, NL) for 15 min followed by a 5 min sonication step at 4 °C. StrepTactin agarose was added to the clarified cell lysates and incubated for 3 h at 4 °C on a rotary wheel. Beads were washed four times in TAP lysis buffer to reduce the concentration of unspecific proteins and to separate specific binders from background ones. Samples were washed five additional times with TAP washing buffer (50 mM Tris pH 7.5, 100 mM NaCl, 5% (
v/
v) glycerol, 1.5 mM MgCl2), to remove remaining detergents. Beads were re-suspended in 20 µl guanidinium chloride buffer (6 M GdmCl, 10 mM TCEP, 40 mM CAA, 100 mM Tris/HCl pH 8), boiled at 95 °C for 5 min and digested by adding 20 µl LysC-Protease-Mix (100 mM Tris/HCl pH 8 and 0.5 µg LysC) for 3 h at 30 °C. Samples were diluted 1:5 with Trypsin-Protease-Mix (100 mM Tris/HCl pH 8 and 1 µg Trypsin) (Promega, Madison, WI, USA) and incubated for 12h at room temperature. TFA and acetonitrile was added to a final concentration of 0.6% and 2%, respectively. Peptides were desalted and concentrated using C18 Empore filter discs (3M, Eagan, MN, USA). After elution, peptides were analysed employing an EASY-nanoLC1200 system (Thermo Fisher Scientific, Leiden, NL), which was directly coupled to a Q-Exactive plus HF mass spectrometer (Thermo Fisher Scientific, Leiden, The Netherlands). Peptides were loaded on an analytical 50 cm C18 column (50 cm, 75 µm column diameter; ReproSil-Pur C18-AQ 1.9 µM resin; Dr. Maisch, Ammerbuch-Entringen, DE) and eluted using an 120 min acetonitrile gradient starting with 5% to 30% (95 min), 30% to 60% (5 min), 60% to 95% (5 min), a wash out period of 5 min at 95%, 95% to 5% (5 min) and a re-adjustment phase at 5% of organic acetonitrile buffer (80% acetonitrile, 0.1% Formic acid) (5 min) at a constant flow of 300 nL/min. The mass spectrometer was used in a data-dependent acquisition mode with one full MS scan, followed by 15 MS/MS scans. Raw files were processed with MaxQuant version 1.5.3.34 (Max-Planck Institut, Martinsried, DE) using label-free quantification (LFQ) and match between run options and searched against forward and reverse sequences of the human proteome (UniprotKB, release 03.2016, The UniProt Consortium,
https://www.uniprot.org) by the built-in Andromeda search engine. Carbamidometylation was set as fixed, methionine oxidation and
N-acetylation as variable modification. Peptide and protein identification were controlled by a False Discovery Rate (FDR) of 0.01. Perseus version 1.5.3.0 (Max-Planck Institut, Martinsried, DE) was used to analyze the output of MaxQuant. Protein groups identified as known contaminants or reverse sequence matches were excluded from the analysis. Only proteins with a minimum of 2 LFQ quantifications in at least one group of replicate experiments (N = 4) for a specific condition were considered for the analysis. Missing values were imputed using normal distribution, whose standard deviation was defined as 30% and the mean was offset by −1.8 standard deviations of the data distribution of the real intensities observed in the corresponding MS run, respectively. The significance of the protein enrichment in the pulldowns of a bait
versus the other condition was determined by Welch’s t-test (two-sided, S0 = 1) and corrected for multiple hypothesis testing using permutation-based false discovery rate statistics (FDR = 0.01, 250 permutations).