Antiviral Activity of Metabolites from Peruvian Plants against SARS-CoV-2: An In Silico Approach
Abstract
1. Introduction
2. Results
2.1. Literature Search
2.2. Virtual Screening
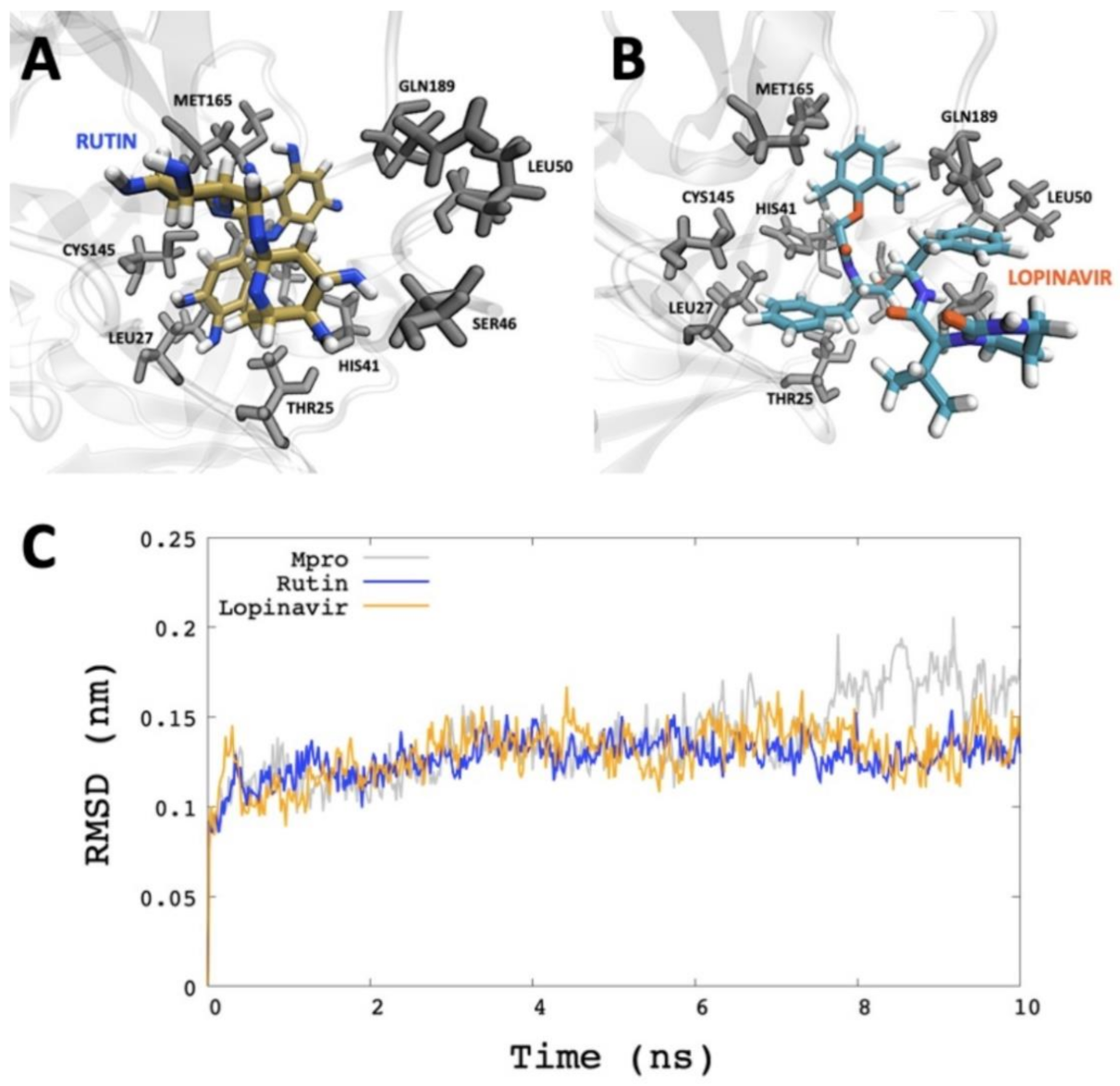
2.3. Molecular Dynamics Simulation Analysis
2.4. MM/GBSA Estimation
3. Discussion
4. Materials and Methods
4.1. Literature Search Strategy and Data Collection
4.2. Virtual Screening
4.3. Molecular Dynamics Simulations and Molecular Mechanics Generalized Born Surface Area Calculations
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Reperant, L.A.; Osterhaus, A.D.M.E. AIDS Avian flu, SARS, MERS, Ebola, Zika…What next? Vaccine 2017, 35, 4470–4474. [Google Scholar] [CrossRef]
- Piret, J.; Boivin, G. Pandemics Throughout History. Front. Microbiol. 2021, 11, 3594. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. WHO Coronavirus Disease (COVID-19) Dashboard. 2021. Available online: https://apps.who.int/iris/bitstream/handle/10665/336034/nCoV-weekly-sitrep11Oct20-eng.pdf (accessed on 30 March 2021).
- Pan, L.; Mu, M.; Yang, P.; Sun, Y.; Wang, R.; Yan, J.; Li, P.; Hu, B.; Wang, J.; Hu, C.; et al. Clinical, Characteristics of, COVID-19 Patients with Digestive Symptoms in Hubei China: A Descriptive Cross-Sectional Multicenter Study. Am. J. Gastroenterol. 2020, 115, 766–773. [Google Scholar] [CrossRef]
- Tang, Y.-W.; Schmitz, J.E.; Persing, D.H.; Stratton, C.W. Laboratory Diagnosis of COVID-19: Current Issues and Challenges. J. Clin. Microbiol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Feng, W.; Newbigging, A.M.; Le, C.; Pang, B.; Peng, H.; Cao, Y.; Wu, J.; Abbas, G.; Song, J.; Wang, D.-B.; et al. Molecular Diagnosis of COVID-19: Challenges and Research Needs. Anal. Chem. 2020, 92, 10196–10209. [Google Scholar] [CrossRef] [PubMed]
- Food and Drug Administration. COVID-19 Vaccines. 2020. Available online: https://www.fda.gov/emergency-preparedness-and-response/coronavirus-disease-2019-covid-19/covid-19-vaccines (accessed on 29 March 2021).
- Gobal Change Data. Coronavirus (COVID-19) Vaccinations. 2021. Available online: https://ourworldindata.org/covid-vaccinations?country=OWID_WRL (accessed on 5 March 2021).
- Llover, M.N.; Jiménez, M.C. Estado actual de los tratamientos para la COVID-19. FMC Form Medica Contin Aten Primaria 2021, 28, 40–56. [Google Scholar] [CrossRef]
- Food and Drug Administration. Coronavirus (COVID-19) Update: FDA Authorizes, Monoclonal, Antibody for Treatment of COVID-19. 2021. Available online: https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-authorizes-monoclonal-antibody-treatment-covid-19 (accessed on 29 March 2021).
- Teralı, K.; Baddal, B.; Gülcan, H.O. Prioritizing potential ACE2 inhibitors in the COVID-19 pandemic: Insights from a molecular mechanics-assisted structure-based virtual screening experiment. J. Mol. Graph. Model. 2020, 100, 107697. [Google Scholar] [CrossRef]
- Brown, A.S.; Patel, C.J. A standard database for drug repositioning. Sci. Data 2017, 4, 170029. [Google Scholar] [CrossRef]
- Tachoua, W.; Kabrine, M.; Mushtaq, M.; Ul-Haq, Z. An in-silico evaluation of COVID-19 main protease with clinically approved drugs. J. Mol. Graph. Model. 2020, 101, 107758. [Google Scholar] [CrossRef]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A Trial of Lopinavir–Ritonavir in Adults Hospitalized with Severe Covid-19. N. Engl. J. Med. 2020, 382, 1787–1799. [Google Scholar] [CrossRef]
- Kearney, J. Chloroquine as a Potential Treatment and Prevention Measure for the 2019 Novel Coronavirus: A Review. Preprints 2020. [Google Scholar] [CrossRef]
- National Institutes of Health. COVID-19 Clinical Trials. 2021. Available online: https://clinicaltrials.gov/ct2/results?cond=Covid19&recrs=d&age_v=&gndr=&type=&rslt=&phase=2&phase=3&Search=Apply (accessed on 29 March 2021).
- Muhammed, Y. Molecular targets for COVID-19 drug development: Enlightening Nigerians about the pandemic and future treatment. Biosaf. Health 2020, 2, 210–216. [Google Scholar] [CrossRef]
- Wang, H.; Xue, S.; Yang, H.; Chen, C. Recent progress in the discovery of inhibitors targeting coronavirus proteases. Virol. Sin. 2016, 31, 24–30. [Google Scholar] [CrossRef] [PubMed]
- Niemeyer, D.; Mösbauer, K.; Klein, E.M.; Sieberg, A.; Mettelman, R.C.; Mielech, A.M.; Dijkman, R.; Baker, S.C.; Drosten, C.; Müller, M.A. The papain-like protease determines a virulence trait that varies among members of the SARS-coronavirus species. PLoS Pathog. 2018, 14, e1007296. [Google Scholar] [CrossRef] [PubMed]
- McBride, R.; Van Zyl, M.; Fielding, B.C. The coronavirus nucleocapsid is a multifunctional protein. Viruses 2014, 6, 2991–3018. [Google Scholar] [CrossRef]
- Da Silva, A.; Wiedemann, L.S.M.; Veiga-Junior, V.F. Natural “products” role against COVID-19. RSC Adv. 2020, 10, 23379–23393. [Google Scholar]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J. Nat. Prod. 2020, 83, 770–803. [Google Scholar] [CrossRef]
- Grifoni, A.; Sidney, J.; Zhang, Y.; Scheuermann, R.H.; Peters, B.; Sette, A. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host. Microbe 2020, 27, 671–680. [Google Scholar] [CrossRef]
- Ministerio de Educación. Guía de Aplicación de Arquitectura Bioclimática en Locales Educativos. 2008. Available online: https://www2.congreso.gob.pe/sicr/cendocbib/con4_uibd.nsf/9A45F1BED1AB7C6705257CCA00550ABD/$FILE/GuiaBioclimática2008.pdf (accessed on 5 March 2021).
- Ruiz, H. Florae Peruvianae et Chilensis: Sive Descriptiones et Icones Plantarum Peruvianarum et Chilensium Secundum Systema Linnaeanum digestae cum Characteribus Plurium Generum Evulgatorum Reformatis; Sancha: Madrid, Spain, 1798. [Google Scholar]
- De Salud, O.A.; Unanue, C.H. Plantas, Medicinales de la, Subregión. In Andina. Plantas Medicinales de la Subregión Andina; Organismo Andino De Salud Convenio: Lima, Peru, 2014; pp. 1–200. [Google Scholar]
- Bussmann, R.W.; Sharon, D. Plantas medicinales de los Andes y la Amazonía-La flora mágica y medicinal del Norte del Perú. Ethnobot. Res. Appl. 2016, 15, 1–293. [Google Scholar] [CrossRef]
- Navarro-Hoyos, M.; Lebrón-Aguilar, R.; Quintanilla-López, J.E.; Cueva, C.; Hevia, D.; Quesada, S.; Azofeifa, G.; Moreno-Arribas, M.V.; Monagas, M.; Bartolomé, B. Proanthocyanidin Characterization and Bioactivity of Extracts from Different Parts of Uncaria tomentosa L. (Cat’s Claw). Antioxidants 2017, 6, 12. [Google Scholar] [CrossRef]
- Cai, Y.; Evans, F.J.; Roberts, M.F.; Phillipson, J.D.; Zenk, M.H.; Gleba, Y.Y. Polyphenolic compounds from Croton lechleri. Phytochemistry 1991, 30, 2033–2040. [Google Scholar] [CrossRef]
- Perdue, G.P.; Blomster, R.N.; Blake, D.A.; Farnsworth, N.R. South american plants II: Taspine isolation and anti-inflammatory activity. J. Pharm. Sci. 1979, 68, 124–126. [Google Scholar] [CrossRef]
- Sistema Integrado de Información de Comercio Exterior—SIICEX. Informes de las Exportaciones Peruanas por Sectores. 2020. Available online: https://www.siicex.gob.pe/siicex/portal5ES.asp?_page_=217.00000&_portletid_=sFichaProductoRegistro&scriptdo=sc_fp_productosector (accessed on 5 March 2021).
- Fanali, C.; Dugo, L.; Cacciola, F.; Beccaria, M.; Grasso, S.; Dachà, M.; Dugo, P.; Mondello, L. Chemical characterization of Sacha inchi (Plukenetia volubilis L.) oil. J. Agric Food Chem. 2011, 59, 13043–13049. [Google Scholar] [CrossRef] [PubMed]
- Bussmann, R.; Malca, G.; Glenn, A.; Sharon, D.; Nilsen, B.; Parris, B.; Dubose, D.; Ruiz, D.; Saleda, J.; Martinez, M.; et al. Toxicity of medicinal plants used in traditional medicine in Northern Peru. J. Ethnopharmacol. 2011, 137, 121–140. [Google Scholar] [CrossRef] [PubMed]
- Gonzales, G.F. Medicinal Plants from Peru: A Review of Plants as Potential Agents Against Cancer. Anti Cancer Agents Med. Chem. 2006, 6, 429–444. [Google Scholar] [CrossRef]
- Mercorelli, B.; Palù, G.; Loregian, A. Drug repurposing for viral infectious diseases: How far are we? Trends Microbiol. 2018, 26, 865–876. [Google Scholar] [CrossRef] [PubMed]
- Caetano, B.F.R.; de Moura, N.A.; Almeida, A.P.S.; Dias, M.C.; Sivieri, K.; Barbisan, L.F. Yacon (Smallanthus sonchifolius) as a, Food Supplement: Health-Promoting Benefits of Fructooligosaccharides. Nutrients 2016, 8, 436. [Google Scholar] [CrossRef]
- Peres, N.; Da, S.L.; Bortoluzzi, L.C.P.; Marques, L.L.M.; Formigoni, M.; Fuchs, R.H.B.; Droval, A.A.; Cardoso, F.A.R. Medicinal effects of Peruvian maca (Lepidium meyenii): A review. Food Funct. 2020, 11, 83–92. [Google Scholar] [CrossRef]
- Pedreschi, R.; Cisneros-Zevallos, L. Phenolic profiles of Andean purple corn (Zea mays L.). Food Chem. 2007, 100, 956–963. [Google Scholar] [CrossRef]
- Park, J.-S.; Yang, J.-S.; Hwang, B.-Y.; Yoo, B.-K.; Han, K. Hypoglycemic effect of yacon tuber extract and its constituent chlorogenic acid in streptozotocin-induced diabetic rats. Biomol. Ther. 2009, 17, 256–262. [Google Scholar] [CrossRef][Green Version]
- Ramos-Escudero, F.; Muñoz, A.M.; Alvarado-Ortíz, C.; Alvarado, A.; Yánez, J.A. Purple corn (Zea mays L.) phenolic compounds profile and its assessment as an agent against oxidative stress in isolated mouse organs. J. Med. Food 2012, 15, 206–215. [Google Scholar] [CrossRef]
- Lin, F.; Hasegawa, M.; Kodama, O. Purification and identification of antimicrobial sesquiterpene lactones from yacon (Smallanthus sonchifolius) leaves. Biosci. Biotechnol. Biochem. 2003, 67, 2154–2159. [Google Scholar] [CrossRef]
- Dou, D.-Q.; Tian, F.; Qiu, Y.-K.; Xiang, Z.; Xu, B.X.; Kang, T.G.; Dong, F. Studies on chemical constituents of the leaves of Smallantus sonchifolius (yacon): Structures of two new diterpenes. Nat. Prod. Res. 2010, 24, 40–47. [Google Scholar] [CrossRef]
- Inoue, A.; Tamogami, S.; Kato, H.; Nakazato, Y.; Akiyama, M.; Kodama, O.; Akatsuka, T.; Hashidoko, Y. Antifungal melampolides from leaf extracts of Smallanthus sonchifolius. Phytochemistry 1995, 39, 845–848. [Google Scholar] [CrossRef]
- Padla, E.P.; Solis, L.T.; Ragasa, C.Y. Antibacterial and antifungal properties of ent-kaurenoic acid from Smallanthus sonchifolius. Chin. J. Nat. Med. 2012, 10, 408–414. [Google Scholar] [CrossRef]
- De Andrade, E.F.; de Souza, R.L.; Ellendersen, L.N.; Masson, M.L. Phenolic profile and antioxidant activity of extracts of leaves and flowers of yacon (Smallanthus sonchifolius). Ind. Crops. Prod. 2014, 62, 499–506. [Google Scholar] [CrossRef]
- Alonso-Castro, A.J.; Ortiz-Sánchez, E.; Dominguez, F.; López-Toledo, G.; Chávez, M.; Ortiz-Tello, A.D.J.; García-Carrancá, A. Antitumor effect of Croton lechleri mull. arg.(euphorbiaceae). J. Ethnopharmacol. 2012, 140, 438–442. [Google Scholar] [CrossRef]
- Mayorga, H.; Duque, C.; Knapp, H.; Winterhalter, P. Hydroxyester disaccharides from fruits of cape gooseberry (Physalis peruviana). Phytochemistry 2002, 59, 439–445. [Google Scholar] [CrossRef]
- Lock, O.; Perez, E.; Villar, M.; Flores, D.; Rojas, R. Bioactive compounds from plants used in Peruvian traditional medicine. Nat. Prod. Commun. 2016, 11, 315–337. [Google Scholar]
- Latza, S.; Ganßer, D.; Berger, R.G. Carbohydrate esters of cinnamic acid from fruits of Physalis peruviana Psidium guajava and Vaccinium vitis-idaea. Phytochemistry 1996, 43, 481–485. [Google Scholar] [CrossRef]
- Al-Olayan, E.M.; El-Khadragy, M.F.; Aref, A.M.; Othman, M.S.; Kassab, R.B.; Moneim, A.E. The potential protective effect of Physalis peruviana L. against carbon tetrachloride-induced hepatotoxicity in rats is mediated by suppression of oxidative stress and downregulation of M.MP-9 expression. Oxid. Med. Cell. Longev. 2014, 2014, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.; Suzuki, M.; Ohnishi-Kameyama, M.; Sada, Y.; Nakanishi, T.; Nagata, T. Extraction and Identification of Antioxidants in the Roots of Yacon (Smallanthus s onchifolius). J. Agric. Food Chem. 1999, 47, 4711–4713. [Google Scholar] [CrossRef]
- Meissner, H.O.; Mscisz, A.; Mrozikiewicz, M.; Baraniak, M.; Mielcarek, S.; Kedzia, B.; Piatkowska, E.; Jolkowska, J.; Pisulewski, P. Peruvian, Maca (Lepidium peruvianum): (I) Phytochemical and genetic differences in three Maca phenotypes. Int. J. Biomed. Sci. 2015, 11, 131. [Google Scholar]
- Yábar, E.; Pedreschi, R.; Chirinos, R.; Campos, D. Glucosinolate content and myrosinase activity evolution in three maca (Lepidium meyenii Walp.) ecotypes during preharvest harvest and postharvest drying. Food Chem. 2011, 127, 1576–1583. [Google Scholar] [CrossRef]
- Li, G.; Ammermann, U.; Quirós, C.F. Glucosinolate contents in maca (Lepidium peruvianum Chacón) seeds, sprouts, mature plants and several derived commercial products. Econ. Bot. 2001, 55, 255–262. [Google Scholar] [CrossRef]
- Piacente, S.; Carbone, V.; Plaza, A.; Zampelli, A.; Pizza, C. Investigation of the tuber constituents of maca (Lepidium meyenii, Walp.). J. Agric. Food Chem. 2002, 50, 5621–5625. [Google Scholar] [CrossRef] [PubMed]
- Lim, T.K. Lepidium Meyenii. Edible Medicinal and Non Medicinal Plants; Springer: Berlin/Heidelberg, Germany, 2015; pp. 801–828. [Google Scholar]
- Zheng, B.L.; He, K.; Kim, C.H.; Rogers, L.; Shao, Y.; Huang, Z.Y.; Lu, Y.; Yan, S.J.; Qien, L.C.; Zheng, Q.Y. Effect of a lipidic extract from Lepidium meyenii on sexual behavior in mice and rats. Urology 2000, 55, 598–602. [Google Scholar] [CrossRef]
- Almukadi, H.; Wu, H.; Böhlke, M.; Kelley, C.J.; Maher, T.J.; Pino-Figueroa, A. The macamide N-3-methoxybenzyl-linoleamide is a time-dependent fatty acid amide hydrolase (FAAH) inhibitor. Mol. Neurobiol. 2013, 48, 333–339. [Google Scholar] [CrossRef]
- McCollom, M.M.; Villinski, J.R.; McPhail, K.L.; Craker, L.E.; Gafner, S. Analysis of macamides in samples of Maca (Lepidium meyenii) by HPLC-UV-MS/MS. Phytochem. Anal. 2005, 16, 463–469. [Google Scholar] [CrossRef] [PubMed]
- Ramadan, M.F.; Mörsel, J.-T. Oil goldenberry (Physalis peruviana, L.). J. Agric. Food Chem. 2003, 51, 969–974. [Google Scholar] [CrossRef]
- Dini, A.; Migliuolo, G.; Rastrelli, L.; Saturnino, P.; Schettino, O. Chemical composition of Lepidium meyenii. Food Chem. 1994, 49, 347–349. [Google Scholar] [CrossRef]
- Marín, F.R.; Frutos, M.J.; Pérez-Alvarez, J.A.; Martinez-Sánchez, F.; Del Río, J.A. Flavonoids as nutraceuticals: Structural related antioxidant properties and their role on ascorbic acid preservation. Stud. Nat. Prod. Chem. 2002, 26, 741–778. [Google Scholar] [CrossRef]
- Negahdari, R.; Bohlouli, S.; Sharifi, S.; Dizaj, S.M.; Saadat, Y.R.; Khezri, K.; Jafari, S.; Ahmadian, E.; Jahandizi, N.G.; Raeesi, S. Therapeutic benefits of rutin and its nanoformulations. Phyther. Res. 2021, 35, 1719–1738. [Google Scholar] [CrossRef] [PubMed]
- Wen, C.-C.; Kuo, Y.-H.; Jan, J.-T.; Liang, P.-H.; Wang, S.-Y.; Liu, H.-G.; Lee, C.-K.; Chang, S.-T.; Kuo, C.-J.; Lee, S.-S.; et al. Specific plant terpenoids and lignoids possess potent antiviral activities against severe acute respiratory syndrome coronavirus. J. Med. Chem. 2007, 50, 4087–4095. [Google Scholar] [CrossRef]
- Jo, S.; Kim, S.; Shin, D.H.; Kim, M.-S. Inhibition of SARS-CoV 3CL protease by flavonoids. J. Enzyme Inhib. Med. Chem. 2020, 35, 145–151. [Google Scholar] [CrossRef]
- Verma, S.; Twilley, D.; Esmear, T.; Oosthuizen, C.B.; Reid, A.-M.; Nel, M. Anti-SARS-CoV natural products with the potential to inhibit SARS-CoV-2 (COVID-19). Front. Pharmacol. 2020, 11, 1514. [Google Scholar] [CrossRef]
- Akinboro, A.; Bakare, A.A. Cytotoxic and genotoxic effects of aqueous extracts of five medicinal plants on Allium cepa Linn. J. Ethnopharmacol. 2007, 112, 470–475. [Google Scholar] [CrossRef]
- Padilla-González, G.F.; Amrehn, E.; Frey, M.; Gómez-Zeledón, J.; Kaa, A.; Da Costa, F.B.; Spring, O. Metabolomic and gene expression studies reveal the diversity distribution and spatial regulation of the specialized metabolism of yacon (Smallanthus sonchifolius Asteraceae). Int. J. Mol. Sci. 2020, 21, 4555. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.G.; Kang, O.H.; Lee, Y.S.; Oh, Y.C.; Chae, H.S.; Obiang-Obounou, B.; Park, S.C.; Shin, D.W.; Hwang, B.Y.; Kwon, D.Y. Antimicrobial activity of the constituents of Smallanthus sonchifolius leaves against methicillin-resistant Staphylococcus aureus. Eur. Rev. Med. Pharmacol. Sci. 2010, 14, 1005–1009. [Google Scholar]
- Ullrich, S.; Nitsche, C. The SARS-CoV-2 main protease as drug target. Bioorg. Med. Chem. Lett. 2020, 30, 127377. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Du, X.; Xu, Y.; Deng, Y.; Liu, M.; Zhao, Y.; Zhang, B.; Li, X.; Zhang, L.; Peng, C.; et al. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020, 582, 289–293. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, M.A.; Abdelrahman, A.H.; Hussien, T.A.; Badr, E.A.; Mohamed, T.A.; El-Seedi, H.R.; Pare, P.W.; Efferth, T.; Hegazy, M.-E.F. In silico drug discovery of major metabolites from spices as SARS-CoV-2 main protease inhibitors. Comput. Biol. Med. 2020, 126, 104046. [Google Scholar] [CrossRef]
- Manandhar, A.; Blass, B.E.; Colussi, D.J.; Almi, I.; Abou-Gharbia, M.; Klein, M.L.; Elokely, K.M. Targeting, SARS-CoV-2 M3CLpro by HCV NS3/4a Inhibitors: In Silico Modeling and In Vitro Screening. J. Chem. Inf. Model. 2021, 61, 1020–1032. [Google Scholar] [CrossRef]
- Kitamura, N.; Sacco, M.D.; Ma, C.; Hu, Y.; Townsend, J.A.; Meng, X.; Zhang, F.; Zhang, X.; Ba, M.; Szeto, T.; et al. Expedited Approach toward the Rational Design of Noncovalent SARS-CoV-2 Main Protease Inhibitors. J. Med. Chem. 2021. [Google Scholar] [CrossRef] [PubMed]
- Chenot, C.; Robiette, R.; Collin, S. First evidence of the cysteine and glutathione conjugates of 3-sulfanylpentan-1-ol in hop (Humulus lupulus L.). J. Agric. Food Chem. 2019, 67, 4002–4010. [Google Scholar] [CrossRef]
- Qiao, J.; Li, Y.-S.; Zeng, R.; Liu, F.-L.; Luo, R.-H.; Huang, C.; Wang, Y.-F.; Zhang, J.; Quan, B.; Shen, C.; et al. SARS-CoV-2 Mpro inhibitors with antiviral activity in a transgenic mouse model. Science 2021, 371, 1374–1378. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Qin, B.; Chen, P.; Zhu, K.; Hou, P.; Wojdyla, J.A.; Wang, M.; Cui, S. Crystal structure of SARS-CoV-2 papain-like protease. Acta Pharm. Sin. B 2021, 11, 237–245. [Google Scholar] [CrossRef]
- Rogers, F.B. Medical subject headings. Bull. Med. Libr. Assoc. 1963, 51, 114–116. [Google Scholar]
- Van Eck, N.J.; Waltman, L. Software survey: VOSviewer a computer program for bibliometric mapping. Scientometrics 2010, 84, 523–538. [Google Scholar] [CrossRef]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A.; et al. PubChem Substance and Compound databases. Nucleic Acid Res. 2016, 44, D1202–D1213. [Google Scholar] [CrossRef]
- Wishart, D.S.; Feunang, Y.D.; Guo, A.C.; Lo, E.J.; Marcu, A.; Grant, J.R.; Sajed, T.; Johnson, D.; Li, C.; Sayeeda, Z.; et al. DrugBank 5.0: A Major Update to the DrugBank Database for 2018. Nucleic Acids Res. 2018, 46, D1074–D1082. [Google Scholar] [CrossRef] [PubMed]
- Gaulton, A.; Hersey, A.; Nowotka, M.; Bento, A.P.; Chambers, J.; Mendez, D.; Mutowo, P.; Atkinson, F.; Bellis, L.J.; Cibrián-Uhalte, E.; et al. The ChEMBL database in 2017. Nucleic Acids Res. 2017, 45, D945–D954. [Google Scholar] [CrossRef] [PubMed]
- Sander, T.; Freyss, J.; von Korff, M.; Rufener, C. DataWarrior: An open-source program for chemistry aware data visualization and analysis. J. Chem. Inf. Model. 2015, 55, 460–473. [Google Scholar] [CrossRef] [PubMed]
- Biasini, M.; Bienert, S.; Waterhouse, A.; Arnold, K.; Studer, G.; Schmidt, T.; Kiefer, F.; Cassarino, T.G.; Bertoni, M.; Bordoli, L.; et al. SWISS-MODEL: Modelling protein tertiary and quaternary structure using evolutionary information. Nucleic Acids Res. 2014, 42, W252–W258. [Google Scholar] [CrossRef] [PubMed]
- Dallakyan, S.; Olson, A.J. Small-molecule library screening by docking with PyRx. Methods Mol. Biol. 2015, 1263, 243–250. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock, Vina: Improving the speed and accuracy of docking with a new scoring function efficient optimization and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, A.W.S.; Vranken, W.F. ACPYPE—AnteChamber PYthon Parser interfacE. BMC Res. Notes 2012, 5, 1–8. [Google Scholar] [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, HJC. GROMACS: Fast flexible and free. J. Comput. Chem. 2005, 26, 1701–1718. [Google Scholar] [CrossRef]
- Racine, J. gnuplot 4.0: A portable interactive plotting utility. J. Appl. Econ. 2006, 21, 133–141. [Google Scholar] [CrossRef]
- Tresanco, M.S.V.; Valdes-Tresanco, M.E.; Valiente, P.A.; Frías, E.M. gmx\_MMPBSA. Zenodo 2021. [Google Scholar] [CrossRef]
- Miller, I.B.R.; McGee, J.T.D.; Swails, J.M.; Homeyer, N.; Gohlke, H.; Roitberg, A.E. MMPBSA. py: An Efficient Program for End-State Free Energy Calculations. J. Chem. Theory Comput. 2012, 8, 3314–3321. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. Sartorius products. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Stierand, K.; Maaß, P.C.; Rarey, M. Molecular complexes at a glance: Automated generation of two-dimensional complex diagrams. Bioinformatics 2006, 22, 1710–1716. [Google Scholar] [CrossRef] [PubMed]
| PubChem ID | Structure | Name | Part of the Plant/Extract | TMW | cLogP | HBA | HBB | TPSA | References |
|---|---|---|---|---|---|---|---|---|---|
| 689043 | 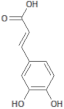 | Caffeic acid | Roots/Methanolic extract | 180.16 | 0.78 | 4 | 3 | 136.56 | [38] |
| 1794427 |  | Chlorogenic acid | Roots/Methanolic extract | 354.31 | −0.768 | 9 | 6 | 245.29 | [39] |
| 445858 | 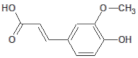 | Ferulic acid | Roots | 194.19 | 1.06 | 4 | 2 | 152.47 | [40] |
| 131753040 |  | Sonchifolin | Leaves/Methanolic extract | 374.43 | 4.06 | 6 | 0 | 294.45 | [41] |
| 101324862 |  | Polymatin B | Leaves/NA | 432.47 | 3.52 | 8 | 0 | 333.2 | [42] |
| 92043370 |  | Uvedalin | Leaves/dichloromethane extract | 448.47 | 2.09 | 9 | 0 | 335.55 | [41] |
| 101250074 |  | Fluctuanin | Leaves/dichloromethane extract | 448.47 | 2.09 | 9 | 0 | 335.55 | [43] |
| 73062 | 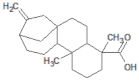 | Kaurenoic acid | Leaves/NA | 302.46 | 4.12 | 2 | 1 | 223.44 | [44] |
| 370 |  | Gallic acid | Leaves/NA | 170.12 | 0.11 | 5 | 4 | 116.41 | [45] |
| 5280805 |  | Rutin | Leaves/NA | 610.52 | -1.26 | 16 | 10 | 397.71 | [45,46,47,48] |
| 5281672 | 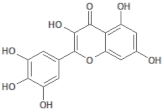 | Myricetin | Leaves/NA | 318.24 | 1.14 | 8 | 6 | 208.29 | [49] |
| 5280863 | 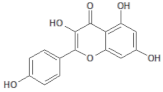 | Kaempferol | Leaves/NA | 286.24 | 1.84 | 6 | 4 | 195.59 | [50] |
| 5280343 | 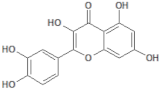 | Quercetin | Leaves/NA | 302.24 | 1.49 | 7 | 5 | 201.94 | [51] |
| PubChemID | Structure | Name | Part of the Plant/Extract | TMW | cLogP | HBA | HBB | TPSA | References |
|---|---|---|---|---|---|---|---|---|---|
| 656498 |  | Glucotropaeolin | Hypocotyl, roots/ethanol extract | 409.44 | −0.75 | 10 | 5 | 270.57 | [52,53] |
| 6602400 |  | Glucosinalbin | Hypocotyl/methanol extract | 425.43 | −1.09 | 11 | 6 | 276.92 | [53,54] |
| 5485207 | 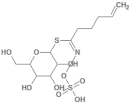 | Glucobrassicanapin | Hypocotyl/methanol extract | 387.43 | −0.559 | 10 | 5 | 262.87 | [53,55] |
| 5317667 | 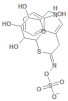 | Glucobrassicin | Hypocotyl/methanol extract | 447.46 | −1.957 | 11 | 5 | 295.92 | [53,55] |
| 11198769 | 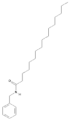 | N-Benzylpalmitamide | Hypocotyl/hexane-ethanol extract | 345.57 | 7.43 | 2 | 1 | 321.13 | [56] |
| 220495 | 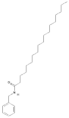 | N-Benzyloctadecanamide | Hypocotyl/hexane-ethanol extract | 373.62 | 8.34 | 2 | 1 | 348.65 | [57] |
| 68742556 | 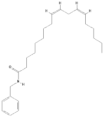 | N-Benzyl-linoleamide | Hypocotyl/hexane-ethanol extract | 369.59 | 7.84 | 2 | 1 | 346.61 | [58] |
| 68741582 |  | N-Benzyl-(9z,12z,15z)-octadecatrienamide | Hypocotyl/hexane-ethanol extract | 367.58 | 7.59 | 2 | 1 | 345.59 | [59] |
| 5280450 | 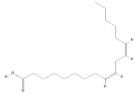 | Linoleic acid | Hypocotyl/ether extracts | 280.45 | 6.47 | 2 | 1 | 267.98 | [48,60] |
| 445639 | 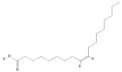 | Oleic acid | Hypocotyl/ether extracts | 282.47 | 6.72 | 2 | 1 | 269 | [48,61] |
| 71386083 |  | 7-Tridecanoic acid | Hypocotyl/hexane extracts | 212.33 | 4.45 | 2 | 1 | 200.2 | [61] |
| Mpro | Mpro * | Plpro | PLpro * | N | N * | |
|---|---|---|---|---|---|---|
| RMSD (nm) | 0.18 ± 0.04 | 0.14 ± 0.01 | 0.52 ± 0.12 | 0.58 ± 0.14 | 0.17 ± 0.04 | 0.13 ± 0.02 |
| RG (nm) | 2.55 ± 0.01 | 2.55 ± 0.01 | 3.28 ± 0.05 | 3.24 ± 0.05 | 1.52 ± 0.01 | 1.53 ± 0.01 |
| Energy Component | Mpro-Rutin | Plpro-Rutin | N-Rutin | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Average | Std Dev | Std Err of Mean | Average | Std Dev | Std Err of Mean | Average | Std Dev | Std Err of Mean | |
| VDWAALS | −53.481 | 4.310 | 0.193 | −21.713 | 6.213 | 0.278 | −34.342 | 3.510 | 0.157 |
| EEL | −33.983 | 10.924 | 0.488 | −47.024 | 23.706 | 1.059 | −19.027 | 7.432 | 0.332 |
| EGB | 53.900 | 7.243 | 0.324 | 60.491 | 18.393 | 0.822 | 31.047 | 5.453 | 0.244 |
| ESURF | −6.730 | 0.402 | 0.018 | −3.451 | 0.816 | 0.036 | −4.684 | 0.371 | 0.017 |
| Δ Ggas | −87.463 | 11.289 | 0.504 | −68.737 | 22.061 | 0.986 | −53.370 | 8.381 | 0.374 |
| Δ Gsolv | 47.170 | 7.075 | 0.316 | 57.041 | 18.012 | 0.805 | 26.363 | 5.246 | 0.234 |
| Δ TOTAL | −40.293 | 5.740 | 0.256 | −11.697 | 6.628 | 0.296 | −27.007 | 4.115 | 0.184 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goyzueta-Mamani, L.D.; Barazorda-Ccahuana, H.L.; Mena-Ulecia, K.; Chávez-Fumagalli, M.A. Antiviral Activity of Metabolites from Peruvian Plants against SARS-CoV-2: An In Silico Approach. Molecules 2021, 26, 3882. https://doi.org/10.3390/molecules26133882
Goyzueta-Mamani LD, Barazorda-Ccahuana HL, Mena-Ulecia K, Chávez-Fumagalli MA. Antiviral Activity of Metabolites from Peruvian Plants against SARS-CoV-2: An In Silico Approach. Molecules. 2021; 26(13):3882. https://doi.org/10.3390/molecules26133882
Chicago/Turabian StyleGoyzueta-Mamani, Luis Daniel, Haruna Luz Barazorda-Ccahuana, Karel Mena-Ulecia, and Miguel Angel Chávez-Fumagalli. 2021. "Antiviral Activity of Metabolites from Peruvian Plants against SARS-CoV-2: An In Silico Approach" Molecules 26, no. 13: 3882. https://doi.org/10.3390/molecules26133882
APA StyleGoyzueta-Mamani, L. D., Barazorda-Ccahuana, H. L., Mena-Ulecia, K., & Chávez-Fumagalli, M. A. (2021). Antiviral Activity of Metabolites from Peruvian Plants against SARS-CoV-2: An In Silico Approach. Molecules, 26(13), 3882. https://doi.org/10.3390/molecules26133882