Targeting Melanoma-Initiating Cells by Caffeine: In Silico and In Vitro Approaches
Abstract
1. Introduction
2. Results
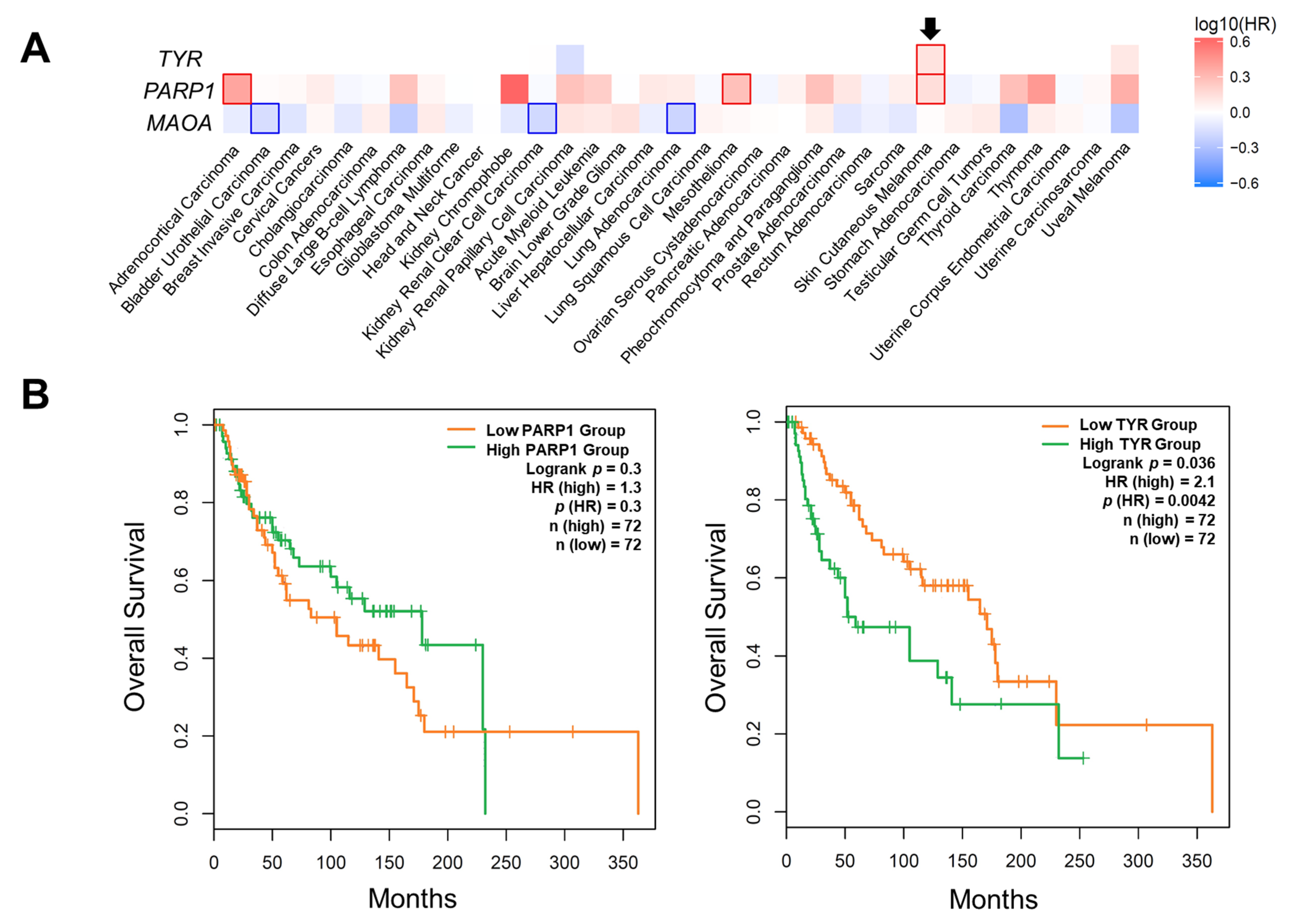
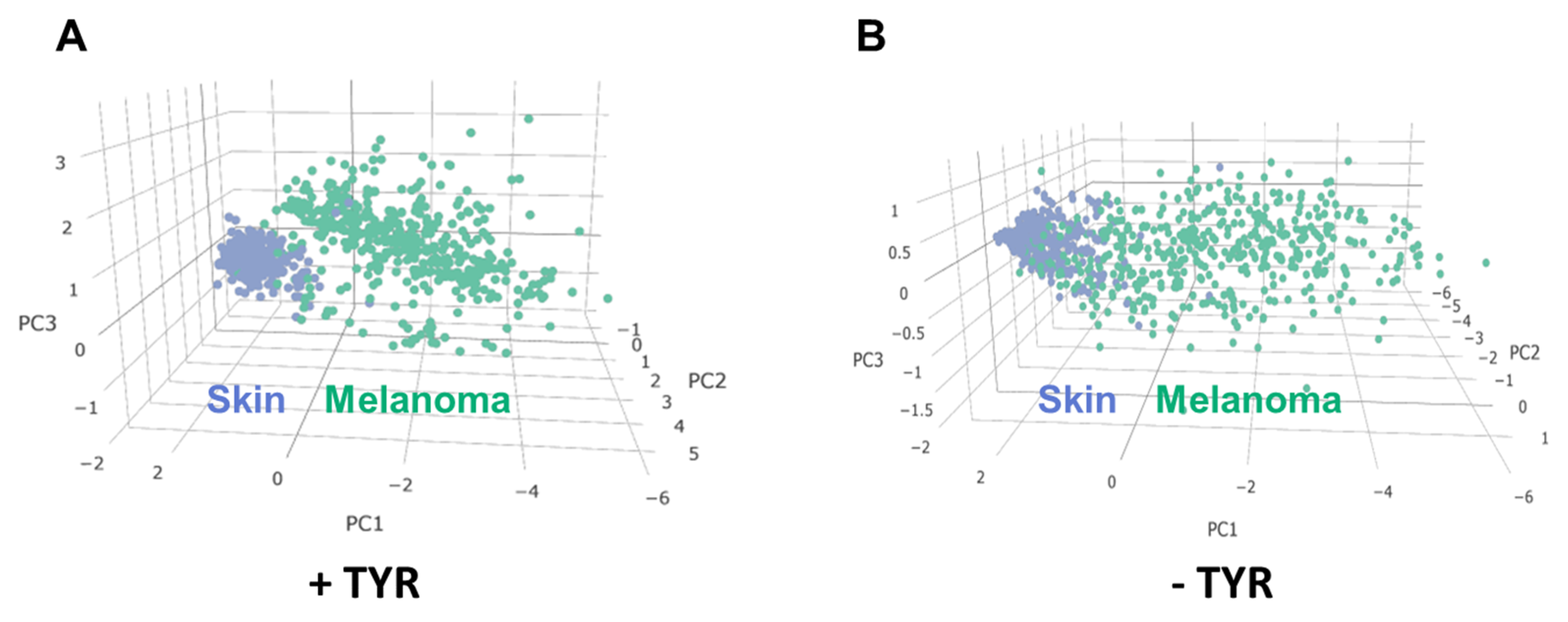
2.1. In Silico Analysis of Caffeine Molecular Targets (CMT) in Melanoma
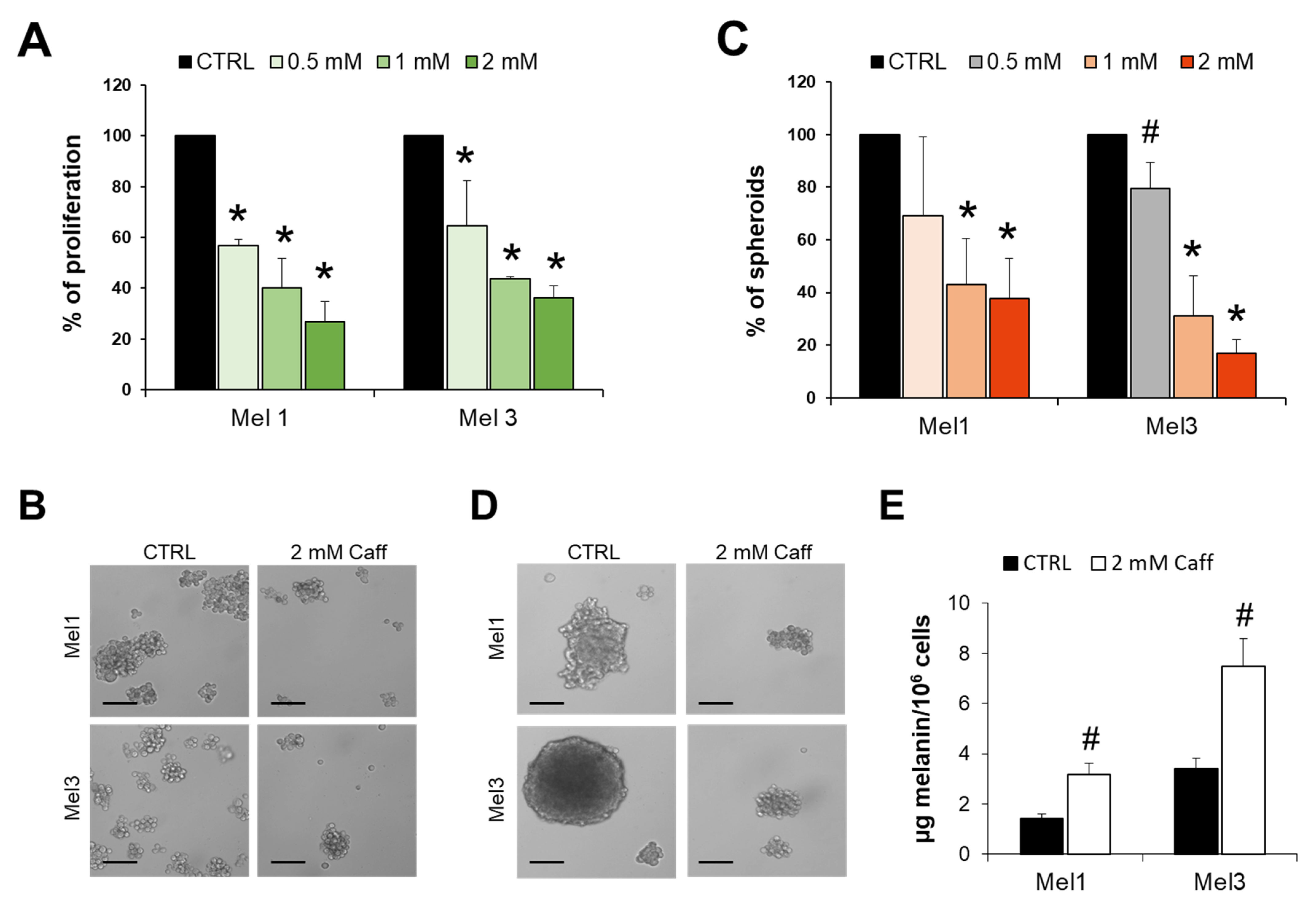
2.2. Effects of Caffeine on MICs Proliferation, Melanospheres Formation and Melanin Production
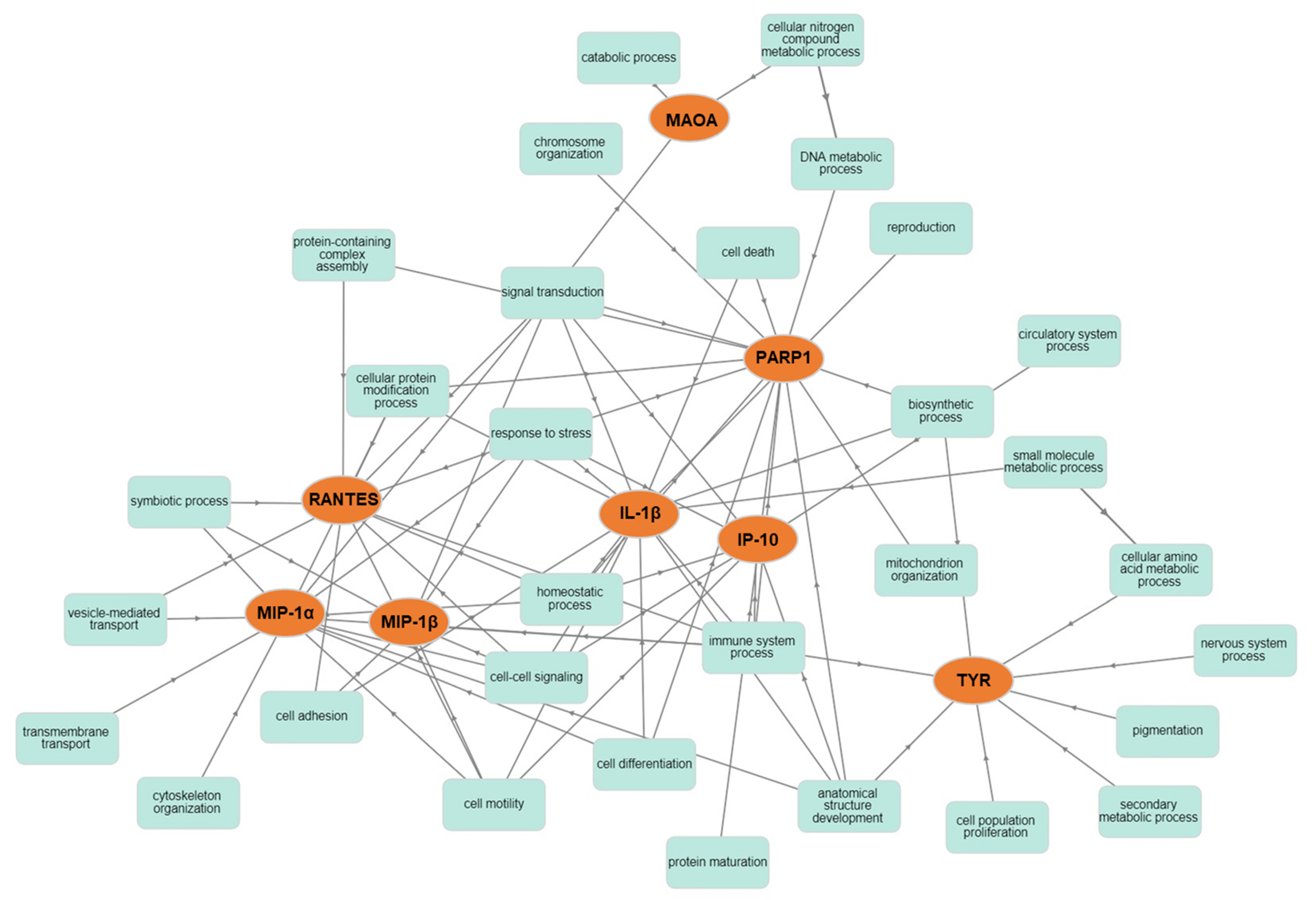
2.3. Network Analysis
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Identification of Caffeine Molecular Targets (CMT) Through Bioinformatic Analyses
4.3. Cell Culture and Proliferation Studies
4.4. Evaluation of Melanin Content
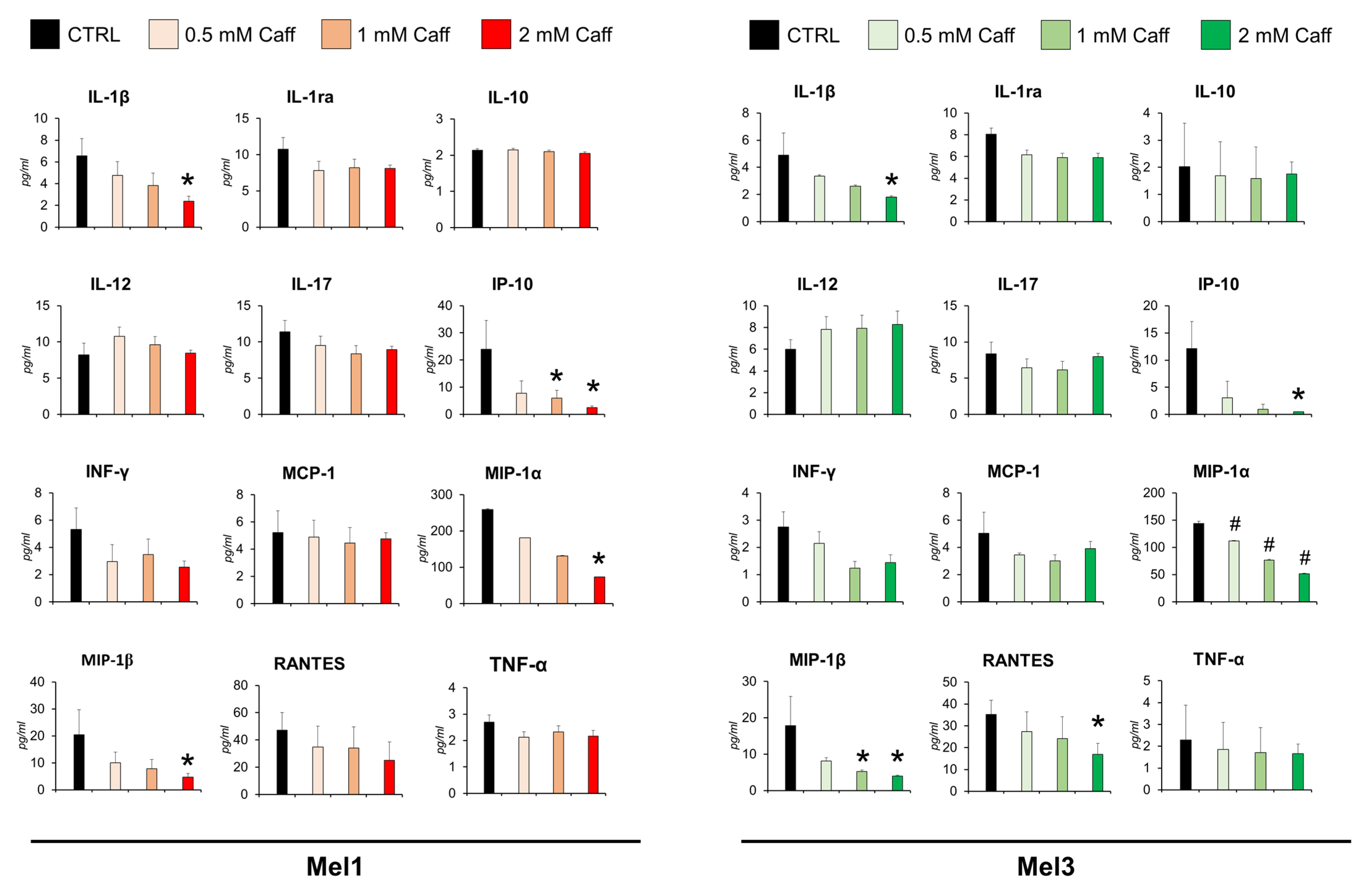
4.5. Cytokines/Chemokines Profile in Melanospheres Supernatants
4.6. Network Analysis
4.7. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Grosso, G.; Godos, J.; Galvano, F.; Giovannucci, E.L. Coffee, Caffeine, and Health Outcomes: An Umbrella Review. Annu. Rev. Nutr. 2017, 37, 131–156. [Google Scholar] [CrossRef]
- Poole, R.; Kennedy, O.J.; Roderick, P.; Fallowfield, J.A.; Hayes, P.C.; Parkes, J. Coffee consumption and health: Umbrella re-view of meta-analyses of multiple health outcomes. BMJ 2017, 359, 5024. [Google Scholar] [CrossRef]
- Socała, K.; Szopa, A.; Serefko, A.; Poleszak, E.; Wlaź, P. Neuroprotective Effects of Coffee Bioactive Compounds: A Review. Int. J. Mol. Sci. 2020, 22, 107. [Google Scholar] [CrossRef]
- Caini, S.; Masala, G.; Saieva, C.; Kvaskoff, M.; Savoye, I.; Sacerdote, C.; Hemmingsson, O.; Hammer Bech, B.; Overvad, K.; Tjonneland, A.; et al. Coffee, tea and melanoma risk: Findings from the European Prospective Investigation into Cancer and Nutrition. Int. J. Cancer 2017, 140, 2246–2255. [Google Scholar] [CrossRef]
- Mackintosh, C.; Yuan, C.; Ou, F.-S.; Zhang, S.; Niedzwiecki, D.; Chang, I.-W.; O’Neil, B.H.; Mullen, B.C.; Lenz, H.-J.; Blanke, C.D.; et al. Association of Coffee Intake With Survival in Patients With Advanced or Metastatic Colorectal Cancer. JAMA Oncol. 2020, 6, 1713. [Google Scholar] [CrossRef]
- Pounis, G.; Tabolacci, C.; Costanzo, S.; Cordella, M.; Bonaccio, M.; Rago, L.; D’Arcangelo, D.; Di Castelnuovo, A.F.; de Gaetano, G.; Donati, M.B.; et al. Reduction by coffee consumption of prostate cancer risk: Evidence from the Moli-sani cohort and cellular models. Int. J. Cancer 2017, 141, 72–82. [Google Scholar] [CrossRef] [PubMed]
- Kudwongsa, W.; Promthet, S.; Suwanrungruang, K.; Phunmanee, A.; Vatanasapt, P. Coffee Consumption and Lung Cancer Risk: A Prospective Cohort Study in Khon Kaen Thailand. Asian Pac. J. Cancer Prev. 2020, 21, 2367–2371. [Google Scholar] [CrossRef] [PubMed]
- Bhurwal, A.; Ratta, P.; Yoshitake, S.; Pioppo, L.; Reja, D.; Dellatore, P.; Rustgi, V. Inverse Association of Coffee with Liver Cancer Development: An Updated Systematic Review and Meta-analysis. J. Gastrointest. Liver Dis. 2020, 29, 421–428. [Google Scholar] [CrossRef]
- Bhoo-Pathy, N.; Peeters, P.H.; Uiterwaal, C.S.; Bueno-De-Mesquita, H.B.; Bulgiba, A.M.; Bech, B.H.; Overvad, K.; Tjønneland, A.; Olsen, A.; Clavel-Chapelon, F.; et al. Coffee and tea consumption and risk of pre- and postmenopausal breast cancer in the European Prospective Investigation into Cancer and Nutrition (EPIC) cohort study. Breast Cancer Res. 2015, 17, 15. [Google Scholar] [CrossRef] [PubMed]
- de Melo Pereira, G.V.; de Carvalho Neto, D.P.; Magalhães Júnior, A.I.; do Prado, F.G.; Pagnoncelli, M.G.B.; Karp, S.G.; Soccol, C.R. Chemical composition and health properties of coffee and coffee by-products. Adv. Food Nutr. Res. 2020, 91, 65–96. [Google Scholar] [CrossRef]
- Pohanka, M. The perspective of caffeine and caffeine derived compounds in therapy. Bratisl. Lek. Listy 2015, 116, 520–530. [Google Scholar] [CrossRef]
- Farias-Pereira, R.; Park, C.-S.; Park, Y. Mechanisms of action of coffee bioactive components on lipid metabolism. Food Sci. Biotechnol. 2019, 28, 1287–1296. [Google Scholar] [CrossRef]
- Tangella, L.P.; Clark, M.E.; Gray, E.S. Resistance mechanisms to targeted therapy in BRAF-mutant melanoma—A mini re-view. Biochim. Biophys. Acta Gen. Subj. 2020, 1865, 129736. [Google Scholar] [CrossRef]
- Damsky, W.E.; Rosenbaum, L.E.; Bosenberg, M. Decoding Melanoma Metastasis. Cancers 2010, 3, 126–163. [Google Scholar] [CrossRef] [PubMed]
- Patel, M.; Eckburg, A.; Gantiwala, S.; Hart, Z.; Dein, J.; Lam, K.; Puri, N. Resistance to Molecularly Targeted Therapies in Melanoma. Cancers 2021, 13, 1115. [Google Scholar] [CrossRef] [PubMed]
- Steininger, J.; Gellrich, F.F.; Schulz, A.; Westphal, D.; Beissert, S.; Meier, F. Systemic Therapy of Metastatic Melanoma: On the Road to Cure. Cancers 2021, 13, 1430. [Google Scholar] [CrossRef] [PubMed]
- Mazurkiewicz, J.; Simiczyjew, A.; Dratkiewicz, E.; Ziętek, M.; Matkowski, R.; Nowak, D. Stromal Cells Present in the Mela-noma Niche Affect Tumor Invasiveness and Its Resistance to Therapy. Int. J. Mol. Sci. 2021, 22, 529. [Google Scholar] [CrossRef]
- Tucci, M.; Passarelli, A.; Mannavola, F.; Felici, C.; Stucci, L.S.; Cives, M.; Silvestris, F. Immune System Evasion as Hallmark of Melanoma Progression: The Role of Dendritic Cells. Front. Oncol. 2019, 9, 1148. [Google Scholar] [CrossRef] [PubMed]
- Jamal, S.M.; Alamodi, A.; Wahl, R.U.; Grada, Z.; Shareef, M.A.; Hassan, S.-Y.; Murad, F.E.; Hassan, S.-L.; Santourlidis, S.; Gomez, C.R.; et al. Melanoma stem cell maintenance and chemo-resistance are mediated by CD133 signal to PI3K-dependent pathways. Oncogene 2020, 39, 5468–5478. [Google Scholar] [CrossRef]
- Zhang, S.; Yang, Z.; Qi, F. Aldehyde dehydrogenase-positive melanoma stem cells in tumorigenesis, drug resistance and anti-neoplastic immunotherapy. Mol. Biol. Rep. 2020, 47, 1435–1443. [Google Scholar] [CrossRef]
- Ravindran, S.; Rasool, S.; Maccalli, C. The Cross Talk between Cancer Stem Cells/Cancer Initiating Cells and Tumor Micro-environment: The Missing Piece of the Puzzle for the Efficient Targeting of these Cells with Immunotherapy. Cancer Microenviron. 2019, 12, 133–148. [Google Scholar] [CrossRef]
- Cordella, M.; Tabolacci, C.; Senatore, C.; Rossi, S.; Mueller, S.; Lintas, C.; Eramo, A.; D’Arcangelo, D.; Valitutti, S.; Facchiano, A.; et al. Theophylline induces differentiation and modulates cytoskeleton dynamics and cytokines secretion in hu-man melanoma-initiating cells. Life Sci. 2019, 230, 121–131. [Google Scholar] [CrossRef]
- Hida, T.; Kamiya, T.; Kawakami, A.; Ogino, J.; Sohma, H.; Uhara, H.; Jimbow, K. Elucidation of Melanogenesis Cascade for Identifying Pathophysiology and Therapeutic Approach of Pigmentary Disorders and Melanoma. Int. J. Mol. Sci. 2020, 21, 6129. [Google Scholar] [CrossRef] [PubMed]
- Horrigan, L.A.; Kelly, J.P.; Connor, T.J. Immunomodulatory effects of caffeine: Friend or foe? Pharmacol. Ther. 2006, 111, 877–892. [Google Scholar] [CrossRef]
- Loftfield, E.; Shiels, M.S.; Graubard, B.I.; Katki, H.A.; Chaturvedi, A.K.; Trabert, B.; Pinto, L.A.; Kemp, T.J.; Shebl, F.M.; Mayne, S.T.; et al. Associations of Coffee Drinking with Systemic Immune and Inflammatory Markers. Cancer Epidemiol. Biomark. Prev. 2015, 24, 1052–1060. [Google Scholar] [CrossRef]
- Pauwels, E.K.J.; Volterrani, D. Coffee Consumption and Cancer Risk; An Assessment of the Health Implications Based on Recent Knowledge. Med. Princ. Pract. 2021. [Google Scholar] [CrossRef]
- Vignoli, J.A.; Viegas, M.C.; Bassoli, D.G.; Benassi, M.T. Roasting process affects differently the bioactive compounds and the antioxidant activity of arabica and robusta coffees. Food Res. Int. 2014, 61, 279–285. [Google Scholar] [CrossRef]
- Bułdak, R.J.; Hejmo, T.; Osowski, M.; Bułdak, Ł.; Kukla, M.; Polaniak, R.; Birkner, E. The Impact of Coffee and Its Selected Bioactive Compounds on the Development and Progression of Colorectal Cancer In Vivo and In Vitro. Molecules 2018, 23, 3309. [Google Scholar] [CrossRef] [PubMed]
- Ross, G.W.; Petrovitch, H. Current Evidence for Neuroprotective Effects of Nicotine and Caffeine Against Parkinson’s Disease. Drugs Aging 2001, 18, 797–806. [Google Scholar] [CrossRef] [PubMed]
- Van Schaik, L.; Kettle, C.; Green, R.; Irving, H.R.; Rathner, J.A. Effects of Caffeine on Brown Adipose Tissue Thermogenesis and Metabolic Homeostasis: A Review. Front. Neurosci. 2021, 15, 621356. [Google Scholar] [CrossRef]
- Ohishi, T.; Fukutomi, R.; Shoji, Y.; Goto, S.; Isemura, M. The Beneficial Effects of Principal Polyphenols from Green Tea, Coffee, Wine, and Curry on Obesity. Molecules 2021, 26, 453. [Google Scholar] [CrossRef]
- Davies, D.B.; Veselkov, D.A.; Djimant, L.N.; Veselkov, A.N. Hetero-association of caffeine and aromatic drugs and their competitive binding with a DNA oligomer. Eur. Biophys. J. 2001, 30, 354–366. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, N.; Ujihara, T.; Kohata, K. Binding energy of tea catechin/caffeine complexes in water evaluated by titration ex-periments with 1H-NMR. Biosci. Biotechnol. Biochem. 2004, 68, 2512–2518. [Google Scholar] [CrossRef] [PubMed]
- Syed, T.A.; Gaikar, V.G.; Mukherjee, S. Stability of co-crystals of caffeine with gallic acid in presence of coformers. J. Food Process Eng. 2019, 42, e13066. [Google Scholar] [CrossRef]
- Król, K.; Gantner, M.; Tatarak, A.; Hallmann, E. The content of polyphenols in coffee beans as roasting, origin and storage effect. Eur. Food Res. Technol. 2020, 246, 33–39. [Google Scholar] [CrossRef]
- Cui, W.-Q.; Wang, S.-T.; Pan, D.; Chang, B.; Sang, L.-X. Caffeine and its main targets of colorectal cancer. World J. Gastrointest. Oncol. 2020, 12, 149–172. [Google Scholar] [CrossRef] [PubMed]
- Lentini, A.; Kleinman, H.K.; Mattioli, P.; Autuori-Pezzoli, V.; Nicolini, L.; Pietrini, A.; Abbruzzese, A.; Cardinali, M.; Beninati, S. Inhibition of melanoma pulmonary metastasis by methylxanthines due to decreased invasion and proliferation. Melanoma Res. 1998, 8, 131–138. [Google Scholar] [CrossRef]
- Gude, R.P.; Menon, L.G.; Rao, S.G. Effect of Caffeine, a xanthine derivative, in the inhibition of experimental lung metastasis induced by B16F10 melanoma cells. J. Exp. Clin. Cancer Res. 2001, 20, 287–292. [Google Scholar]
- Carballo-Carbajal, I.; Laguna, A.; Romero-Giménez, J.; Cuadros, T.; Bové, J.; Martinez-Vicente, M.; Parent, A.; Gonzalez-Sepulveda, M.; Peñuelas, N.; Torra, A.; et al. Brain tyrosinase overexpression implicates age-dependent neuromelanin production in Parkinson’s disease pathogenesis. Nat. Commun. 2019, 10, 1–19. [Google Scholar] [CrossRef]
- Sette, G.; Fecchi, K.; Salvati, V.; Lotti, F.; Pilozzi, E.; Duranti, E.; Biffoni, M.; Pagliuca, A.; Martinetti, D.; Memeo, L.; et al. Mek inhibition results in marked antitumor activity against metastatic melanoma patient-derived melanospheres and in melanosphere-generated xenografts. J. Exp. Clin. Cancer Res. 2013, 32, 91. [Google Scholar] [CrossRef]
- Turdo, A.; Veschi, V.; Gaggianesi, M.; Chinnici, A.; Bianca, P.; Todaro, M.; Stassi, G. Meeting the Challenge of Targeting Cancer Stem Cells. Front. Cell Dev. Biol. 2019, 7, 16. [Google Scholar] [CrossRef]
- Ailles, L.E.; Weissman, I.L. Cancer stem cells in solid tumors. Curr. Opin. Biotechnol. 2007, 18, 460–466. [Google Scholar] [CrossRef] [PubMed]
- Arima, Y.; Nobusue, H.; Saya, H. Targeting of cancer stem cells by differentiation therapy. Cancer Sci. 2020, 111, 2689–2695. [Google Scholar] [CrossRef] [PubMed]
- Eun Lee, K.; Bharadwaj, S.; Yadava, U.; Gu Kang, S. Evaluation of caffeine as inhibitor against collagenase, elastase and ty-rosinase using in silico and in vitro approach. J. Enzyme Inhib. Med. Chem. 2019, 34, 927–936. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Sun, X.; Ni, H.; Du, X.; Chen, F.; Jiang, Z.; Li, Q.; Yuanfan, Y. Identification and Characterization of the Tyrosinase Inhibitory Activity of Caffeine from Camellia Pollen. J. Agric. Food Chem. 2019, 67, 12741–12751. [Google Scholar] [CrossRef]
- Al-Maaieh, A.; Flanagan, D.R. Salt Effects On Caffeine Solubility, Distribution, And Self-Association. J. Pharm. Sci. 2002, 91, 1000–1008. [Google Scholar] [CrossRef]
- Park, H.Y.; Wu, C.; Yonemoto, L.; Murphy-Smith, M.; Wu, H.; Stachur, C.M.; Gilchrest, B.A. MITF mediates cAMP-induced protein kinase C-beta expression in human melanocytes. Biochem. J. 2006, 395, 571–578. [Google Scholar] [CrossRef]
- Vargas, A.J.; Sittadjody, S.; Thangasamy, T.; Mendoza, E.E.; Limesand, K.H.; Burd, R. Exploiting tyrosinase expression and activity in melanocytic tumors: Quercetin and the central role of p53. Integr. Cancer Ther. 2011, 10, 328–340. [Google Scholar] [CrossRef]
- Vereecken, P.; Cornelis, F.; Van Baren, N.; Vandersleyen, V.; Baurain, J.F. A synopsis of serum biomarkers in cutaneous mel-anoma patients. Dermatol. Res. Pract. 2012, 2012, 260643. [Google Scholar] [CrossRef] [PubMed]
- Paolino, G.; Bearzi, P.; Pampena, R.; Longo, C.; Frascione, P.; Rizzo, N.; Raucci, M.; Carbone, A.; Cantisani, C.; Ricci, F.; et al. Clinicopathological and dermoscopic features of amelanotic and hypomelanotic melanoma: A retrospective multicentric study. Int. J. Dermatol. 2020, 59, 1371–1380. [Google Scholar] [CrossRef]
- Rohatgi, A.; Kirkwood, J.M. Beyond PD-1: The Next Frontier for Immunotherapy in Melanoma. Front. Oncol. 2021, 11, 640314. [Google Scholar] [CrossRef]
- Gonçalves, R.C.; Kitagawa, R.R.; Raddi, M.S.; Carlos, I.Z.; Pombeiro-Sponchiado, S.R. Inhibition of Nitric Oxide and Tumour Necrosis Factor-α Production in Peritoneal Macrophages by Aspergillus nidulans Melanin. Biol. Pharm. Bull. 2013, 36, 1915–1920. [Google Scholar] [CrossRef] [PubMed]
- Li, L.; Shi, F.; Li, J.; Huang, Q.; Xu, C.; Yang, L.; Yang, Q.; Shaikh, F.; Ye, M. Immunoregulatory effect assessment of a novel melanin and its carboxymethyl derivative. Bioorg. Med. Chem. Lett. 2017, 27, 1831–1834. [Google Scholar] [CrossRef]
- Burkahart, C.G.; Burkhart, C.N. The mole theory: Primary function of melanocytes and melanin may be antimicrobial defense and immunomodulation (not solar protection). Int. J. Dermatol. 2005, 44, 340–342. [Google Scholar] [CrossRef] [PubMed]
- Fu, C.; Chen, J.; Lu, J.; Yi, L.; Tong, X.; Kang, L.; Pei, S.; Ouyang, Y.; Jiang, L.; Ding, Y.; et al. Roles of inflammation factors in melanogenesis (Review). Mol. Med. Rep. 2020, 21, 1421–1430. [Google Scholar] [CrossRef]
- Baruch, E.N.; Youngster, I.; Ben-Betzalel, G.; Ortenberg, R.; Lahat, A.; Katz, L.; Adler, K.; Dick-Necula, D.; Raskin, S.; Bloch, N.; et al. Fecal microbiota transplant promotes response in immunotherapy-refractory melanoma patients. Science 2021, 371, 602–609. [Google Scholar] [CrossRef]
- Davar, D.; Dzutsev, A.K.; McCulloch, J.A.; Rodrigues, R.R.; Chauvin, J.-M.; Morrison, R.M.; Deblasio, R.N.; Menna, C.; Ding, Q.; Pagliano, O.; et al. Fecal microbiota transplant overcomes resistance to anti–PD-1 therapy in melanoma patients. Science 2021, 371, 595–602. [Google Scholar] [CrossRef]
- Kleber Silveira, A.; Moresco, K.S.; Mautone Gomes, H.; da Silva Morrone, M.; Kich Grun, L.; Pens Gelain, D.; de Mattos Pereira, L.; Giongo, A.; Rodrigues De Oliveira, R.; Fonseca Moreira, J.C. Guarana (Paullinia cupanaMart.) alters gut microbiota and modulates redox status, partially via caffeine in Wistar rats. Phytother. Res. 2018, 32, 2466–2474. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.; Chen, J.-F. Caffeine and Parkinson’s Disease: Multiple Benefits and Emerging Mechanisms. Front. Neurosci. 2020, 14, 602697. [Google Scholar] [CrossRef]
- Gaascht, F.; Dicato, M.; Diederich, M. Coffee provides a natural multitarget pharmacopeia against the hallmarks of cancer. Genes Nutr. 2015, 10, 1–17. [Google Scholar] [CrossRef]
- Zulli, A.; Smith, R.M.; Kubatka, P.; Novak, J.; Uehara, Y.; Loftus, H.; Qaradakhi, T.; Pohanka, M.; Kobyliak, N.; Zagatina, A.; et al. Caffeine and cardiovascular diseases: Critical review of current research. Eur. J. Nutr. 2016, 55, 1331–1343. [Google Scholar] [CrossRef]
- Pérez-Pérez, D.; Reyes-Vidal, I.; Chávez-Cortez, E.G.; Sotelo, J.; Magaña-Maldonado, R. Methylxanthines: Potential Thera-peutic Agents for Glioblastoma. Pharmaceuticals 2019, 12, 130. [Google Scholar] [CrossRef]
- D’Arcangelo, D.; Giampietri, C.; Muscio, M.; Scatozza, F.; Facchiano, F.; Facchiano, A. WIPI1, BAG1, and PEX3 Autophagy-Related Genes Are Relevant Melanoma Markers. Oxidative Med. Cell. Longev. 2018, 2018, 1–12. [Google Scholar] [CrossRef]
- D’Arcangelo, D.; Facchiano, F.; Nassa, G.; Stancato, A.; Antonini, A.; Rossi, S.; Senatore, C.; Cordella, M.; Tabolacci, C.; Salvati, A.; et al. PDGFR-alpha inhibits melanoma growth via CXCL10/IP-10: A multi-omics approach. Oncotarget 2016, 7, 77257–77275. [Google Scholar] [CrossRef]
- Tang, Z.; Kang, B.; Li, C.; Chen, T.; Zhang, Z. GEPIA2: An enhanced web server for large-scale expression profiling and in-teractive analysis. Nucleic Acids Res. 2019, 47, W556–W560. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, G.; Venugopal, A.; Ramamoorthy, P.; Standing, D.; Subramaniam, D.; Umar, S.; Jensen, R.A.; Anant, S.; Mammen, J.M. Honokiol inhibits melanoma stem cells by targeting notch signaling. Mol. Carcinog. 2015, 54, 1710–1721. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Pomaznoy, M.; Ha, B.; Peters, B. GOnet: A tool for interactive Gene Ontology analysis. BMC Bioinform. 2018, 19, 470. [Google Scholar] [CrossRef] [PubMed]
- Cesati, M.; Scatozza, F.; D’Arcangelo, D.; Antonini-Cappellini, G.C.; Rossi, S.; Tabolacci, C.; Nudo, M.; Palese, E.; Lembo, L.; Di Lella, G.; et al. Investigating Serum and Tissue Expression Identified a Cytokine/Chemokine Signature as a Highly Effective Melanoma Marker. Cancers 2020, 12, 3680. [Google Scholar] [CrossRef] [PubMed]
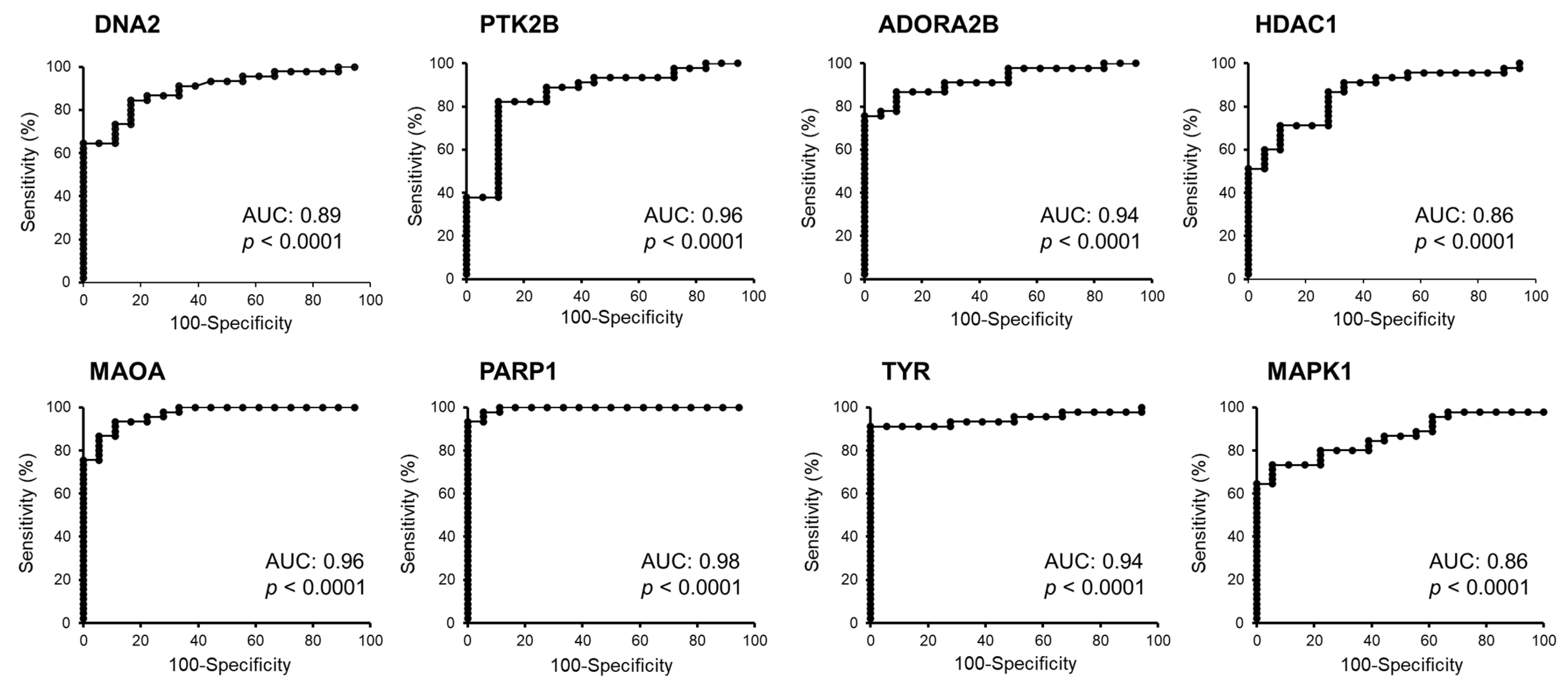
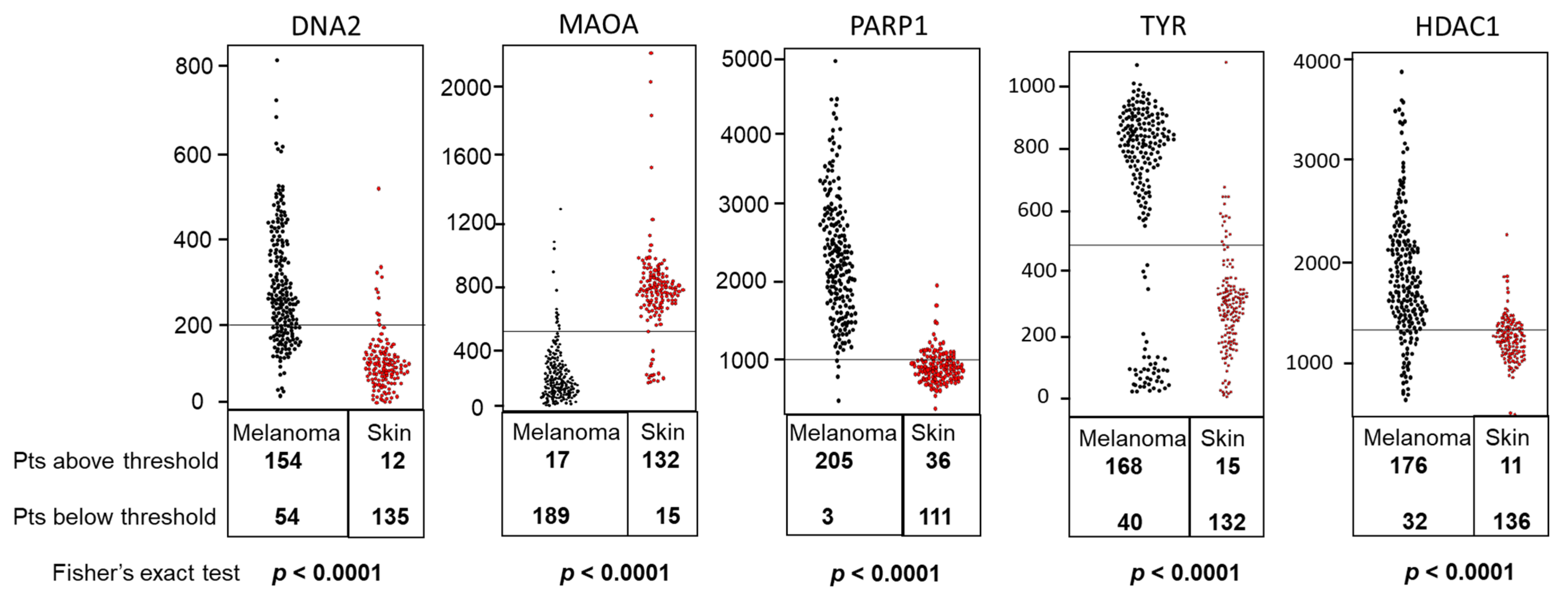
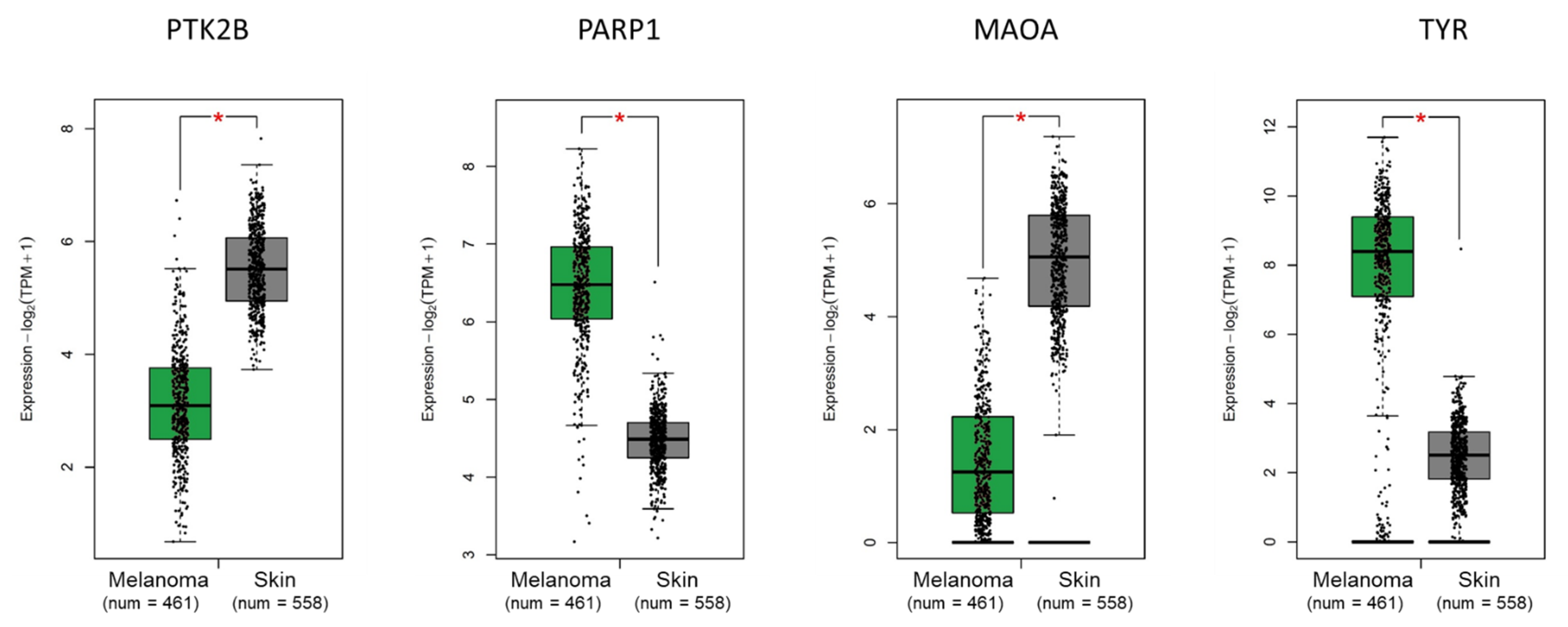
| Symbol | Mean Expression in Melanoma | Mean Expression in Nevi | t-Test (p Value) | Fold Change (Melanoma/Nevi) | AUC (ROC Value) | p Value of AUC |
|---|---|---|---|---|---|---|
| DNA2 | 149.7 | 75.49 | 2.22 × 10−6 | 1.98 | 0.89 | <0.0001 |
| IL1RN (IL-1ra) | 440.1 | 827.11 | 9.00 × 10−5 | 0.53 | 0.80 | 0.0001 |
| SCL2A1 (GLUT-1) | 1213.99 | 1985.62 | 1.00 × 10−4 | 0.61 | 0.80 | 0.0001 |
| MAPK1 | 733.11 | 1361.07 | 2.30 × 10−6 | 0.53 | 0.87 | 0.0001 |
| PTK2B | 1411.28 | 784.15 | 4.40 × 10−13 | 1.8 | 0.96 | <0.0001 |
| ADORA1 | 299.01 | 201.01 | 3.70 × 10−3 | 1.5 | 0.75 | 0.001 |
| ADORA2B | 217.84 | 564.00 | 2.06 × 10−11 | 0.39 | 0.94 | 0.0001 |
| MAOA | 46.83 | 291.86 | 2.48 × 10−11 | 0.16 | 0.95 | 0.0001 |
| PDE2A | 212.43 | 427.64 | 2.10 × 10−4 | 0.49 | 0.78 | 0.0004 |
| PARP1 | 2182.16 | 978.73 | 1.20 × 10−11 | 2.23 | 0.98 | 0.0001 |
| TYR | 22881.3 | 7144.98 | 1.25 × 10−10 | 3.2 | 0.94 | 0.0001 |
| HDAC1 | 1614.36 | 1146.88 | 2.19 × 10−5 | 1.41 | 0.86 | <0.0001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tabolacci, C.; Cordella, M.; Rossi, S.; Bonaccio, M.; Eramo, A.; Mischiati, C.; Beninati, S.; Iacoviello, L.; Facchiano, A.; Facchiano, F. Targeting Melanoma-Initiating Cells by Caffeine: In Silico and In Vitro Approaches. Molecules 2021, 26, 3619. https://doi.org/10.3390/molecules26123619
Tabolacci C, Cordella M, Rossi S, Bonaccio M, Eramo A, Mischiati C, Beninati S, Iacoviello L, Facchiano A, Facchiano F. Targeting Melanoma-Initiating Cells by Caffeine: In Silico and In Vitro Approaches. Molecules. 2021; 26(12):3619. https://doi.org/10.3390/molecules26123619
Chicago/Turabian StyleTabolacci, Claudio, Martina Cordella, Stefania Rossi, Marialaura Bonaccio, Adriana Eramo, Carlo Mischiati, Simone Beninati, Licia Iacoviello, Antonio Facchiano, and Francesco Facchiano. 2021. "Targeting Melanoma-Initiating Cells by Caffeine: In Silico and In Vitro Approaches" Molecules 26, no. 12: 3619. https://doi.org/10.3390/molecules26123619
APA StyleTabolacci, C., Cordella, M., Rossi, S., Bonaccio, M., Eramo, A., Mischiati, C., Beninati, S., Iacoviello, L., Facchiano, A., & Facchiano, F. (2021). Targeting Melanoma-Initiating Cells by Caffeine: In Silico and In Vitro Approaches. Molecules, 26(12), 3619. https://doi.org/10.3390/molecules26123619