Nonhost Disease Resistance in Pea: Chitosan’s Suggested Role in DNA Minor Groove Actions Relative to Phytoalexin-Eliciting Anti-Cancer Compounds
Abstract
:1. Introduction
1.1. AT-Rich DNA Sequences in Pre-Promoter Region of Pea HMG A Gene
- TCAATTAAAA AATCAATTTA TTTATTTCAT TTCATAAATA TATTCATAA
- AATTAAATAC AATGAGTAGA ATTTCAAAC TCTCAATAAATTTTAGT
1.2. Sequences Used in Crystal Modeling of Drug/DNA Minor Groove Complexes
- Netropsin: d(CGCAATTCGCG)
- Distamycin: d(CGCAAATTTGCG)
- Hoechst 33258: d(CGCGAATTCGCG)
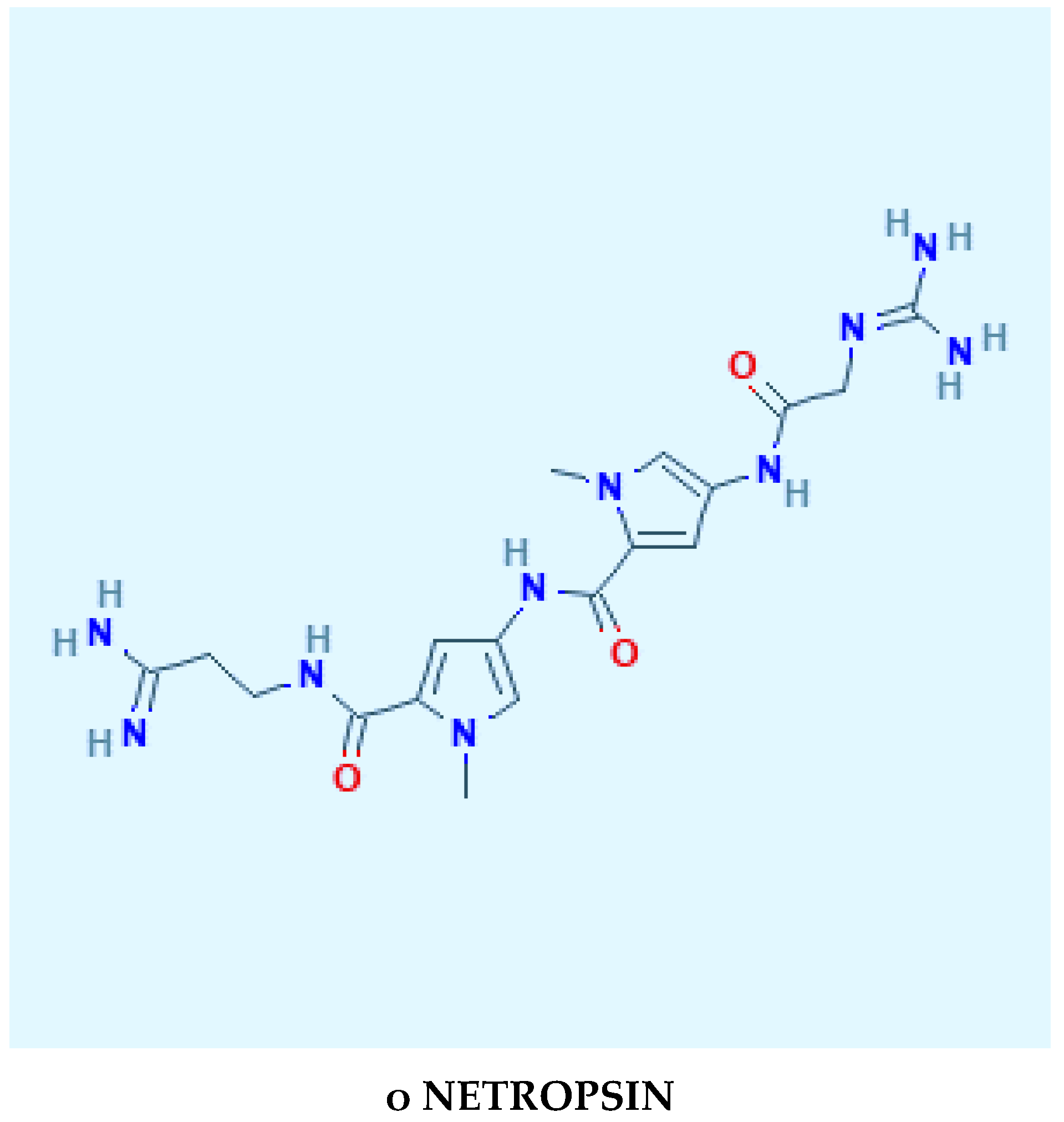
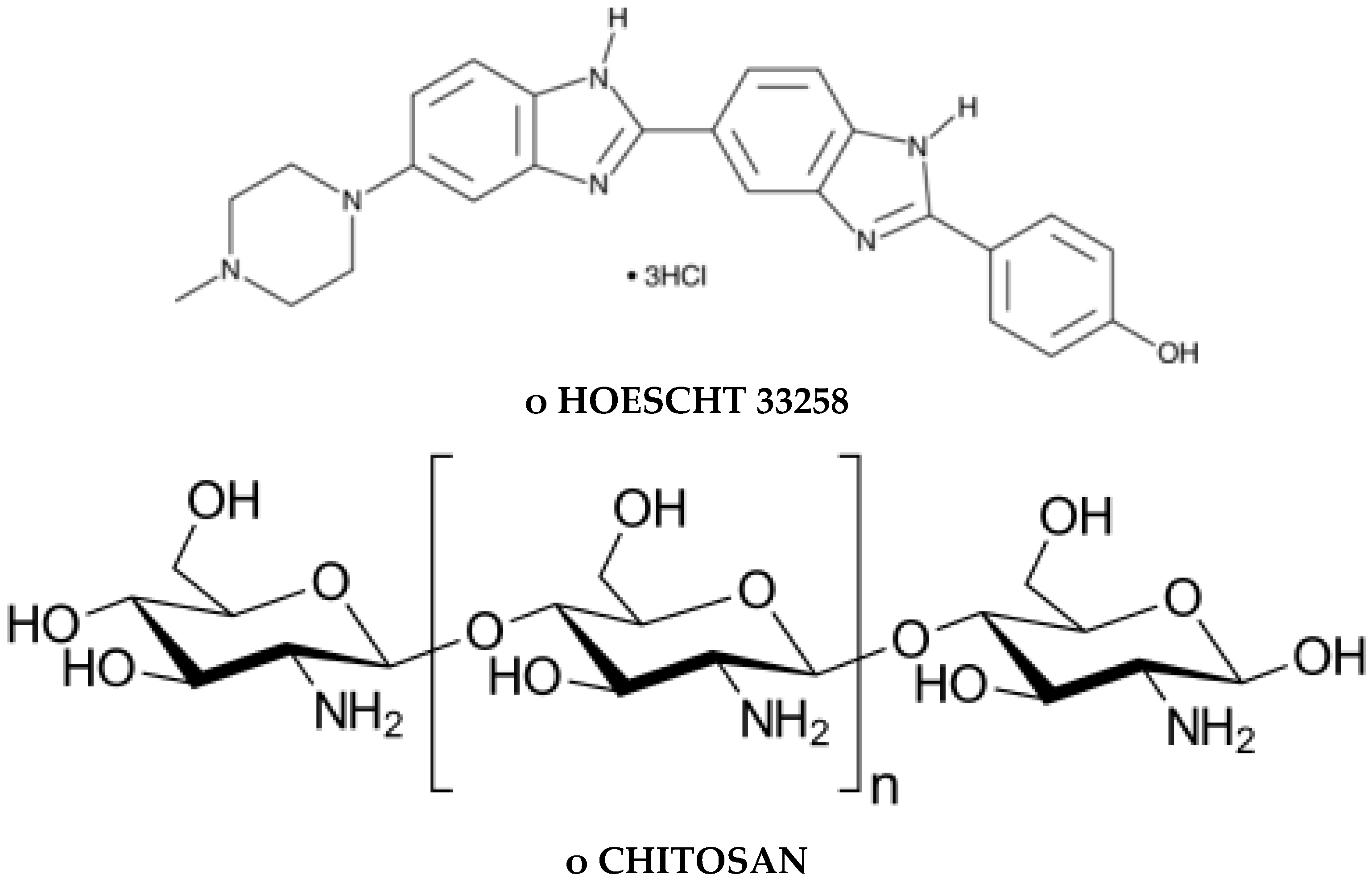
1.3. DNA Minor Groove Compounds Evaluated as Pisatin Elicitors Include Chitosan, Chromomycin A3, Distamycin, Hoecsht 33258, Netropsin, and Methyl Blue
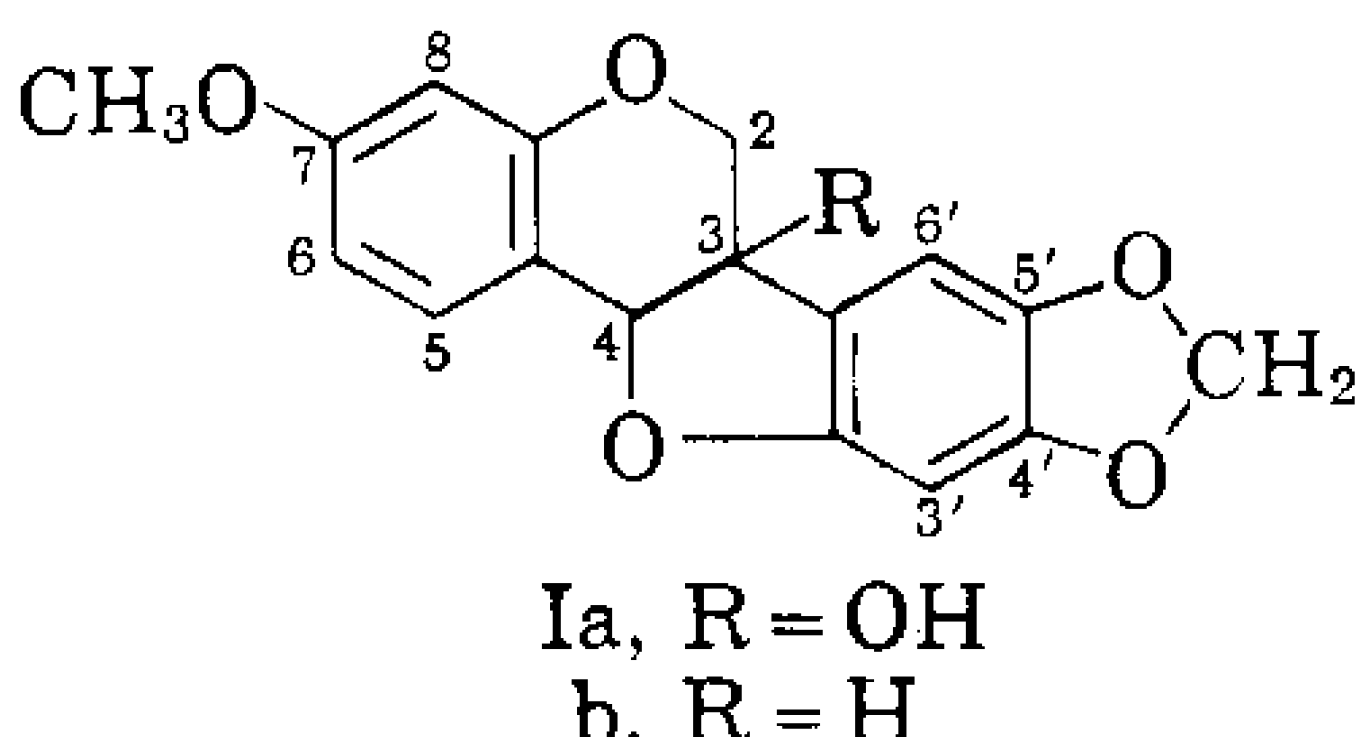
1.4. Production of PR Gene Proteins and Pisatin
1.5. Natural Components
1.6. Effects of Chitosan-Oligomer and Pathogen Challenge on Animal Cells
1.7. Chitosan Signaling in Pea Tissue
2. Results and Discussion
2.1. Chromomycin
2.2. Other DNA Minor Groove Localizing Compounds
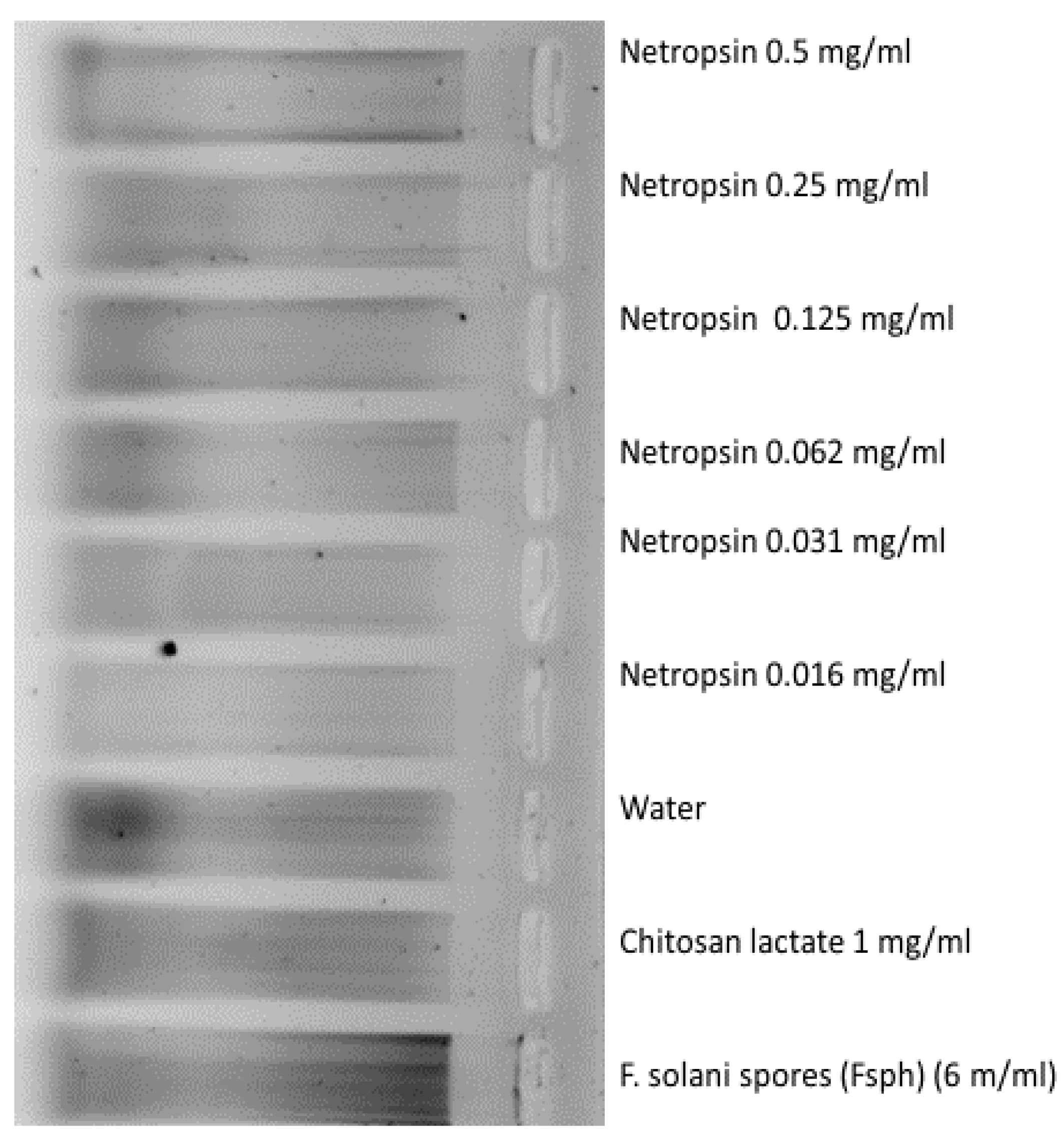
2.3. DNA-Fragmentation Assciated with Netropsin Treatment
2.4. The Specifics of Netropsin/DNA Dodecamer Attachment
2.5. Signal Reception in the Hypothesized Role of Host Chromatin
3. Experimental Section
3.1. Elicitor Compound Sources
3.2. Plant and Pathogen
3.3. Elicitor Treatments and Pisatin Quantization
3.4. DNA Damage Assessment
4. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Hadwiger, L.A.; Beckman, J.M.; Adams, M.J. Localization of fungal components in the Pea-Fusarium interaction detected immunochemically with anti-chitosan and anti-fungal cell wall antisera. Plant Physiol. 1981, 57, 170–175. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hadwiger, L.A.; Chiang, C.; Victory, S.; Horovitz, D. The molecular biology of chitosan in plant/pathogen interaction and its application in agriculture. In Chitin and Chitosan, Sources, Chemistry, Biochemistry, Physical Properties and Applications; Skjat-Braek, G., Sandford, P.A., Eds.; Elsevier Applied Science: London, UK, 1989; pp. 119–137. [Google Scholar]
- Isaac, J.; Hartney, S.L.; Druffel, K.; Hadwiger, L.A. The non-host disease resistance in peas; alterations in phosphorylation and ubiquitination of HMGA and histone H2A/H2B. Plant Sci. 2009, 177, 439–449. [Google Scholar] [CrossRef]
- Hartney, S.; Carson, J.; Hadwiger, L.A. The use of chemical genomics to detect functional systems affecting the non-host disease resistance of pea to Fusarium solani f. sp. phaseoli. Plant Sci. 2007, 172, 45–56. [Google Scholar] [CrossRef]
- Kendra, D.F.; Christian, D.A.; Hadwiger, L.A. Chitosan oligomers from Fusarium solani/pea interactions, chitinase/β-glucanase digestion of sporelings and from fungal wall chitin actively inhibit fungal growth and enhance disease resistance. Physiol. Mol. Plant Path. 1989, 35, 215–230. [Google Scholar] [CrossRef]
- Hadwiger, L.A.; Tanaka, T. DNA damage and chromatin conformation changes confer nonhost resistance: A hypothesis based on effects of anti-cancer agents on plant defense responses. Front. Plant Sci. 2018, 9, 1056–1072. [Google Scholar] [CrossRef]
- Hadwiger, L.A. Anatomy of a nonhost disease resistance response of pea to Fusarium solani: PR gene elicitation via DNase, chitosan and chromatin alterations. Front. Plant Sci. 2015, 6, 373–400. [Google Scholar] [CrossRef]
- Hadwiger, L.A.; Tanaka, K. Nonhost resistance: DNA damage is associated with SA signaling for induction of PR genes and contributes to the growth suppression of a pea pathogen on pea endocarp tissue. Front. Plant Sci. 2017, 8, 446–458. [Google Scholar] [CrossRef] [Green Version]
- Hadwiger, L.A.; Polashock, J. Fungal mitochondrial DNases. Effectors with the potential to activate plant defenses in nonhost resistance. Phytopathology 2013, 1031, 81–90. [Google Scholar] [CrossRef] [Green Version]
- Tanaka, K.; Hadwiger, L.A. Nonhost resistance: Reactive oxygen species (ROS) signal causes DNA damage and the induction of PR genes and disease resistance in pea tissue. Physiol. Mol. Plant Pathol. 2017, 98, 18–24. [Google Scholar] [CrossRef]
- Almeida, M.S.; Cabral, K.M.S.; Zingali, R.B.; Kurtenbach, E. Characterization of two novel defense peptides from pea (Pisum sativum) seeds. Arch. Biochem. Biophys. 2000, 378, 278–286. [Google Scholar] [CrossRef]
- Chiang, C.C.; Hadwiger, L.A. The Fusarium solani-induced expression of a pea gene family encoding cysteine content proteins. Mol. Plant Microbe Interact. 1991, 4, 324–331. [Google Scholar] [CrossRef] [PubMed]
- Teasdale, J.; Daniels, D.; Davis, W.C.; Eddy, R.J.; Hadwiger, L.A. Physiological and cytological similarities between disease resistance and cellular incompatibility responses. Plant Physiol. 1974, 54, 690–695. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lui, D.; Jiao, S.; Cheng, G.; Li, X.; Pei, Z.; Pei, Y.; Yin, H.; Du, Y. Identification of chitosan oligosaccharides binding proteins from the plasma membrane of wheat leaf cell. Int. J. Biol. Macromol. 2018, 111, 1083–1090. [Google Scholar]
- Mauch, F.; Hadwiger, L.A.; Boller, T. Ethylene: Symptom not signal for induction of chitinase and β-1, 3-glucanase in pea pods by pathogens and elicitors. Plant Physiol. 1984, 76, 607–616. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kopka, M.L.; Yoon, D.; Goodsell, D.; Pjura, P.; Dickerson, R.E. The molecular origin of DNA-drug specificity in netropsin and distamycin. Proc. Natl. Acad. Sci. USA 1985, 82, 1376–1386. [Google Scholar] [CrossRef] [Green Version]
- Nunn, M.N.; Garman, E.; Neidle, S. Crystal structure of the DNA decamer d(CGCAATTGCG) complexed with the minor groove binding drug netropsin. Biochemistry 1997, 36, 4792–4799. [Google Scholar] [CrossRef]
- Bhaduri, S.; Ranjan, N.; Arya, D.P. An overview of recent advances in duplex DNA recognition by small molecules. Beilstein J. Org. Chem. 2018, 14, 1051–1086. [Google Scholar] [CrossRef]
- Lewis, E.A.; Munde, M.; Wang, S.; Rettig, M.; Le, V.; Machha, V.; Wilson, W.D. Complexity in the binding of minor groove agents: Netropsin has two thermodynamically different DNA binding modes at a single site. Nucleic Acids Res. 2011, 39, 9649–9658. [Google Scholar] [CrossRef] [Green Version]
- Rohs, R.; West, S.M.; Sosinsky, A.; Lui, P.; Mann, R.S.; Honig, B. The Role of DNA shape in protein-DNA recognition. Nature 2009, 461, 1248–1253. [Google Scholar] [CrossRef] [Green Version]
- Neidle, S. DNA minor-groove recognition by small molecules. Nat. Prod. Rep. 2001, 18, 291–309. [Google Scholar] [CrossRef]
- Druffel, K.; Carson, J.A.; Hartney, S.; Hadwiger, L.A. Inverse PCR to identify DNA sequence upstream of the pea HMY I/Y open reading frame. Pisum Genet. 2005, 37, 15–18. [Google Scholar]
- Hadwiger, L.A.; Webster, D.M. Phytoalexin production in five cultivars of peas differentially resistance to three races of Pseudomonas syringe pv. pisi. Phytopathology 1984, 74, 1312–1314. [Google Scholar] [CrossRef]
- Somssich, I.E.; Hahlbrock, K. Pathogen defence in plants—A paradigm of biological complexity. Trends Plant Sci. 1998, 3, S1360–S1385. [Google Scholar] [CrossRef]
- Fristenski, B.; Riggleman, R.D.; Wagoner, W.; Hadwiger, L.A. Gene expression in susceptible and disease resistant interactions of Peas induced with Fusarium solani pathogens and chitosan. Physiol. Plant Pathol. 1985, 27, 15–28. [Google Scholar] [CrossRef]
- Daniels, D.H.; Fristensky, B.; Wagoner, W.; Hadwiger, L.A. Pea genes associated with non-host disease to Fusarium are also active in race-specific disease resistance to Pseudomononas. Plant Mol. Biol. 1987, 8, 309–316. [Google Scholar] [CrossRef] [PubMed]
- Pilet-Nayel, M.L.; Muehlbauer, F.J.; McGee, R.J.; Kraft, J.M.; Baranger, A.; Coyne, C.J. Quantitive trait loci for partial resistance to Aphanomyces root rot in pea. Theor. Appl. Genet. 2002, 106, 28–39. [Google Scholar] [CrossRef] [PubMed]
- Hadwiger, L.A. How Plants Resist Disease-A focus on “nonhost resistance” of pea to a bean pathogen. Openaccess ebooks 2018, 2, 4. [Google Scholar]
- Hadwiger, L.A. Nonhost resistance: Self-inflicted DNA damage by fungal DNase accumulation is a major factor in terminating fungal growth in the pea-Fusarium solani f. sp. phaseoli interaction. Physiol. Mol. Plant Pathol. 2015, 92, 79–87. [Google Scholar] [CrossRef]
- Gerhold, D.I.; Pettinger, A.J.; Hadwiger, L.A. Characterization of a plant stimulated nuclease from Fusarium solani. Physiol. Mol. Plant Pathol. 1993, 43, 33–46. [Google Scholar] [CrossRef]
- Williamson-Benavides, B.A.; Sharpe, R.M.; Nelson, G.; Bodah, E.T.; Porter, L.D.; Dhingra, A. Identification of Fusarium solani f. sp. pisi (Fsp) responsive genes in Pisum sativum. Front. Genet. 2020, 11, 950–970. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Van Strien, E.A. The families of pathogenesis-related proteins, their activities and comparative analysis of PR-1 type proteins. Physiol. Mol. Plant Pathol. 1999, 55, 85–97. [Google Scholar] [CrossRef]
- Perrin, D.R.; Bottomley, W. Studies on phytoalexins. V. The structure of pisatin from Pisum sativum L. Aust. J. Biol. Sci. 1962, 14, 336–340. [Google Scholar]
- Parsons, M.A.; Hadwiger, L.A. Photoactivated psoralens elicit defense genes and phytoalexins production in the pea plant. Photochem. Photobiol. 1998, 67, 438–445. [Google Scholar] [CrossRef]
- Weake, V.M.; Workman, J.L. Histone ubiquitination triggering gene activity. Mol. Cell 2008, 29, 653–663. [Google Scholar] [CrossRef]
- Kendra, D.F.; Hadwiger, L.A. Characterization of the smallest chitosan oligomer that is maximally antifungal to Fusarium solani and elicits pisatin formation in Pisum sativum. Exp. Mycol. 1984, 8, 276–281. [Google Scholar] [CrossRef]
- Paul, A.; Guo, P.; Boykin, D.W.; Wilson, W.D. A new generation of minor-groove-binding heterocyclic diamidines that recognize G-C base pairs in an AT sequence context. Molecules 2019, 24, 946. [Google Scholar] [CrossRef] [Green Version]
- Nanjunda, R.; Wilson, W.D. Binding to the DNA minor groove by heterocyclic dications: From AT-specific monomers to GC recognition with dimers. Corr. Protoc. Nucleic Acid Chem. 2012, 8, 1–24. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Ronge, X.; Lui, S.; Qin, Y.; Li, K.; Yu, H.; Li, P. Immunostimulatory effects of chitooligosaccharides on RAW 264.7 mouse macrophages via regulation of the MAPK and PI3K/Akt signaling pathways. Mar. Drugs 2019, 17, 36. [Google Scholar] [CrossRef] [Green Version]
- Hadwiger, L.A. Changes in plant metabolism associated with phytoalexin production. Neth. J. Plant Path. 1968, 74, 163–169. [Google Scholar] [CrossRef]
- Allan, C.R.; Hadwiger, L.A. The fungicidal effect of chitosan on fungi of varying cell wall composition. Exp. Mycol. 1979, 3, 285–290. [Google Scholar] [CrossRef]
- Matica, A.M.; Aachmann, F.L.; Tondervik, A.; Sletta, H.; Ostafe, V. Chitosan as a wound dressing starting material: Antimicrobial properties and mode of action. Int. J. Mol. Sci. 2019, 20, 5889. [Google Scholar] [CrossRef] [Green Version]
- Yin, H.; Du, Y.; Dong, Z. Chitin oligosaccharide and chitosan oligosaccharide: Two similar but different plant elicitors. Front. Plant Sci. 2016, 7, 522–526. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miya, A.; Albert, P.; Shinya, T.; Desaki, Y.; Ichimura, K.; Shirasu, K. CERK1, a LysM receptor kinase, is essential for chitin elicitor signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 2007, 104, 19613–19618. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reeves, R.; Beckerbauer, L. HMG A proteins: Flexible regulators of transcription and chromatin structure. Biochem. Biophys. Acta 2001, 1519, 13–19. [Google Scholar] [PubMed]
- Klosterman, S.J.; Choi, J.J.; Hadwiger, L.A. Analysis of pea HMG I/Y suggests a role in defense gene regulation. Mol. Plant Pathol. 2003, 4, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Bewley, C.A.; Gronenbern, A.M.; Clore, M.G. Minor groove-binding architectural proteins: Structure, function and RNA recognition. Ann. Rev. Biophys. Biomed. Struct. 1998, 27, 105–131. [Google Scholar] [CrossRef]
- Tavis, D.S.; Kumar, A.; Stephens, D.E.; Boykin, D.W.; Wilson, W.D. Large, sequence-dependent effects on DNA conformation by minor groove binding compounds. Nucleic Acids Res. 2009, 37, 5550–5558. [Google Scholar] [CrossRef] [Green Version]
- Ma, J.; Bai, L.; Wang, M.D. The supercoiling of DNA is a removable barrier to the RNA polymerase complex (RNAP) transcription of genes. Science 2013, 340, 1580. [Google Scholar] [CrossRef] [Green Version]
| Pre-Treatment | Secondary Trmt in 30 min | Pisatin µg/g fr. wt. 22 h | Hypersensitive Response at 24 h |
|---|---|---|---|
| Netropsin 0.5 mg/mL | Fsph spores | 40.8 ± 5.0 | +++ |
| Netropsin 0.5 mg/mL | water | 26.3 ± 1.6 | +++ |
| Netropsin 0.25 mg/mL | Fsph spores | 22.6 ± 2.5 | +++ |
| Netropsin 0.25 mg/mL | water | 2.1 ± 0.1 | +++ |
| Netropsin 0.125 mg/mL | Fsph spores | 36.1 ± 24.0 | ++ |
| Netropsin 0.125 mg/mL | water | 2.7 ± 1.6 | ++ |
| Water | Fsph spores | 34.7± 8.2 | ++ |
| Pre-Treatment 20 µL/Pod Half | Secondary Trmt in 30 min 20 µL/Pod Half | Pisatin µg/g fr. wt. 24 h | Hyper. Response 24 h |
|---|---|---|---|
| Netropsin 0.5 mg/mL | Fsph spores (3 × 106/mL) | 56.7 ± 3.8 | ++ |
| Water | Fsph spores (3 × 106/mL) | 101.3 ± 22.7 | ++ |
| Netropsin 0.5 mg/mL | Fsph spores (7.5 × 105/mL) | 39.0 ± 3.9 | +++ |
| Water | Fsph spores (7.5 × 105/mL) | 57.0 ± 15.8 | +++ |
| Netropsin 0.5 mg/mL | Fsph spores (3.7 × 105/mL) | 43.8 ± 2.6 | + |
| Water | Fsph spores (3.7 × 105/mL) | 35.3 ± 6.1 | ++ |
| Netropsin 0.5 mg/mL | Fsph spores (1.8 × 105/mL) | 24.2 ± 5.0 | ++ |
| Water | Fsph spores (1.8 × 105/mL) | 52.6 ± 10.1 | ++ |
| Netropsin 0.5 mg/mL | |||
| Fsph spores 3 × 106/mL) | -------------------------------- | 48.9 ± 7.0 | +++ |
| Water | -------------------------------- | --------------- | + |
| Netropsin 0.5 mg/mL | |||
| Chitosan 0.5 mg/mL | ------------------------------- | 10.7 ± 4.9 | ++ |
| Secondary Trmt in 30 min | Pisatin µg/g fr. wt. 24 h | Hypersensitivity Response 24 h | |
|---|---|---|---|
| Chitosan 0.2 mg/mL | Netropsin 1 mg/mL | 50.2 ± 3.2 | +++ |
| Water | Netropsin 1 mg/mL | 22.5 ± 0.9 | +++ |
| Chitosan 0.2 mg/mL | Netropsin 0.5 mg/mL | 38.7 ± 0.9 | +++ |
| Water | Netropsin 0.5 mg/mL | 13.7 ± 0.1 | ++ |
| Chitosan 0.2 mg/mL | Netropsin 0.25 mg/mL | 27.2 ± 3.4 | ++ |
| Water | Netropsin 0.25 mg/mL | 14.2 ± 5.2 | +++ |
| Chitosan 0.2 mg/mL | Netropsin 0.12 mg/mL | 9.8 ± 1.3 | ++ |
| Water | Netropsin 0.12 mg/mL | ---------- | + |
| Fsph spores (2.4 × 106) | ---------------------------- | 244.4 ± 27.4 | +++ |
| Water | ---------------------------- | ---------------- | Light |
| Chitosan 0.2 mg/mL | ---------------------------- | 0.9 ± 9.27 | ++ |
| Treatment (20 µL Volume) | Pisatin µg/g fr. wt. | Hypersensitivity |
|---|---|---|
| Chromomycin 2.0 mg/mL | 69.9 ± 1.0 | +++ |
| Chromomycin 1.0 mg/mL | 133.5 ± 11.2 | ++ |
| Chromomycin 0.5 mg/mL | 188.3 ± 3.5 | + |
| Chromomycin 0.25 mg/mL | 70.9 ± 33.5 | +++ |
| Chromomycin 0.125 mg/mL | 106.1 ± 16.1 | ++ |
| Chromomycin 0.06 mg/mL | 77.3 ± 7.9 | +++ |
| Chromomycin 0.03 mg/mL | 10.9 ± 10.9 | + |
| Chromomycin 0.015 mg/mL | 8.5 ± 7.1 | + |
| Water | ---------------- | + |
| Fsph spores (4.5 × 106) | 184.7 ± 42.7 | ++ |
| Chitosan 1 mg/mL | 8.6 ± 4.6 | +++ |
| Netropsin 0.5 mg/mL | 17.8 ± 1.9 | +++ |
| Pre-Treatment 20 µL/Pod Half | Secondary Trmt in 30 min 20 µL/Pod Half | Pisatin µg/g fr. wt. 24 h |
|---|---|---|
| Chromomycin 0.5 mg/mL | Fsph spores (6.4 × 106/mL) | 271.1 ± 23.3 |
| Chromomycin 0.5 mg/mL | Water | 111.2 ± 9.2 |
| Chromomycin 0.25 mg/mL | Fsph spores (6.4 × 106/mL) | 248.8 ± 10.8 |
| Chromomycin 0.25 mg/mL | Water | 109.8 ± 3.8 |
| Chromomycin 0.12 mg/mL | Fsph spores (6.4 × 106/mL) | 291.0 ± 9.5 |
| Chromomycin 0.12 mg/mL | Water | 58.9 ± 9.9 |
| Chromomycin 0.06 mg/mL | Fsph spores (6.4 × 106/mL) | 270.9 ± 17.5 |
| Chromomycin 0.06 mg/mL | Water | 53.7 ± 6.3 |
| Chromomycin 0.03 mg/mL | Fsph spores (6.4 × 106/mL) | 259.7 ± 28.4 |
| Chromomycin 0.03 mg/mL | Water | 78.3 ± 43.2 |
| Water | Fsph spores (6.4 × 106/mL) | 210.6 ± 34.9 |
| Water | Water | ---------------- |
| Pre-Treatment 20 µL/Pod Half | Secondary Trmt in 30 min 20 µL/Pod Half | Pisatin µg/g fr. wt. |
|---|---|---|
| Chromomycin 0.12 mg/mL | Chitosan 1 mg/mL | 153.0 ± 87.1 |
| Chromomycin 0.12 mg/mL | Water | 113.2 ± 22.1 |
| Chromomycin 0.06 mg/mL | Chitosan 1 mg/mL | 81.9 ± 1.2 |
| Chromomycin 0.06 mg/mL | Water | 29.9 ± 8.9 |
| Chromomycin 0.03 mg/mL | Chitosan 1 mg/mL | 33.4 ± 3.5 |
| Chromomycin 0.03 mg/mL | Water | 29.7 ± 20.6 |
| Chromomycin 0.015 mg/mL | Chitosan 1 mg/mL | 20.6 ± 3.0 |
| Chromomycin 0.015 mg/mL | Water | 2.1 ± 0.0 |
| Chromomycin 0.007 mg/mL | Chitosan 1 mg/mL | 43.2 ± 22.9 |
| Chromomycin 0.007 mg/mL | Water | 2.4 ± 0.5 |
| Water | Chitosan 1 mg/mL | 9.4 ± 4.5 |
| Water | Water | 0.0 ± 0.0 |
| Pre-Treatment | Secondary Treatment 30 min | Pisatin µg/g fr. wt. 24 h |
|---|---|---|
| Hoechst 33258 1 mg/mL | Fsph spores | 102 ± 7 |
| Hoechst 33258 1 mg/mL | Water | 9 ± 1 |
| Hoechst 33258 0.5 mg/mL | Fsph spores | 91 ± 7 |
| Hoechst 33258 0.5 mg/mL | Water | 3 ± 2 |
| Hoechst 33258 0.25 mg/mL | Fsph spores | 116 ± 26 |
| Hoechst 33258 0.25 mg/mL | Water | 1 ± 0.6 |
| Hoechst 33258 0.125 mg/mL | Fsph spores | 90 ± 5 |
| Hoechst 33258 0.125 mg/mL | Water | 2 ± 0.9 |
| Water | Fsph spores | 84 ± 21 |
| Water | Water | --------- |
| Chitosan lactate 1 mg/mL | Fsph spores | 125 ± 62 |
| Chitosan lactate | Water | 9 ± 2 |
Sample Availability: Samples of the compounds are not available from the authors. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hadwiger, L.A. Nonhost Disease Resistance in Pea: Chitosan’s Suggested Role in DNA Minor Groove Actions Relative to Phytoalexin-Eliciting Anti-Cancer Compounds. Molecules 2020, 25, 5913. https://doi.org/10.3390/molecules25245913
Hadwiger LA. Nonhost Disease Resistance in Pea: Chitosan’s Suggested Role in DNA Minor Groove Actions Relative to Phytoalexin-Eliciting Anti-Cancer Compounds. Molecules. 2020; 25(24):5913. https://doi.org/10.3390/molecules25245913
Chicago/Turabian StyleHadwiger, Lee A. 2020. "Nonhost Disease Resistance in Pea: Chitosan’s Suggested Role in DNA Minor Groove Actions Relative to Phytoalexin-Eliciting Anti-Cancer Compounds" Molecules 25, no. 24: 5913. https://doi.org/10.3390/molecules25245913
APA StyleHadwiger, L. A. (2020). Nonhost Disease Resistance in Pea: Chitosan’s Suggested Role in DNA Minor Groove Actions Relative to Phytoalexin-Eliciting Anti-Cancer Compounds. Molecules, 25(24), 5913. https://doi.org/10.3390/molecules25245913





