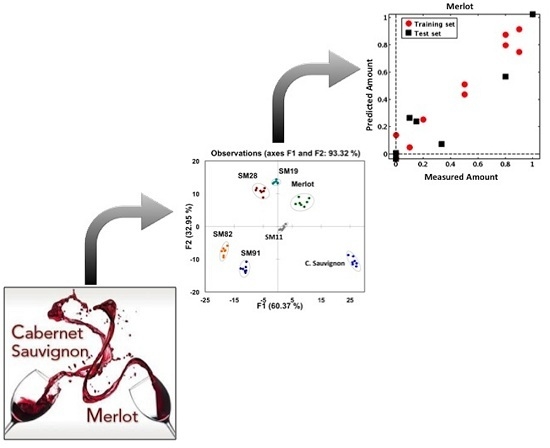
Predicting the Composition of Red Wine Blends Using an Array of Multicomponent Peptide-Based Sensors
Abstract
:1. Introduction
2. Results and Discussion
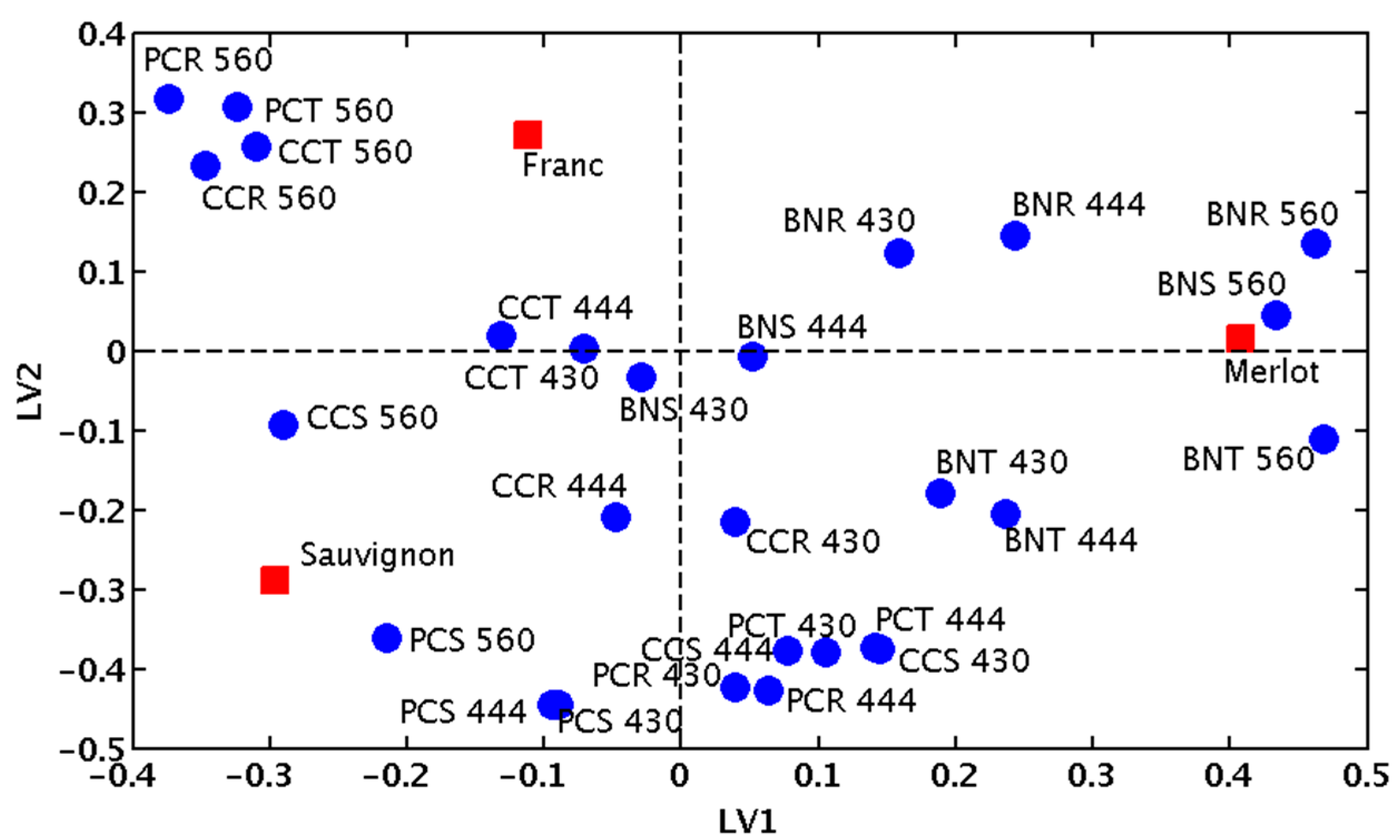
2.1. Fingerprinting Red Wine Blends
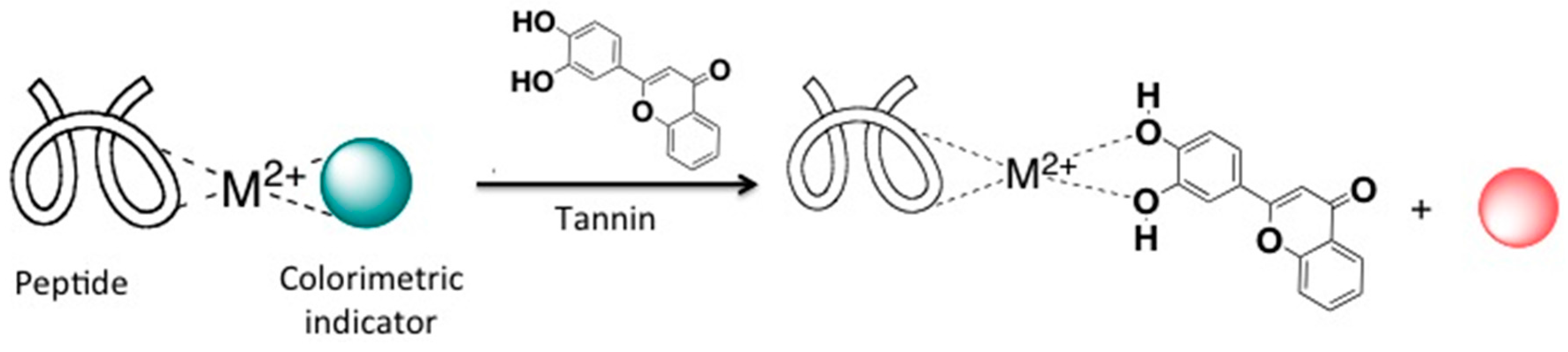
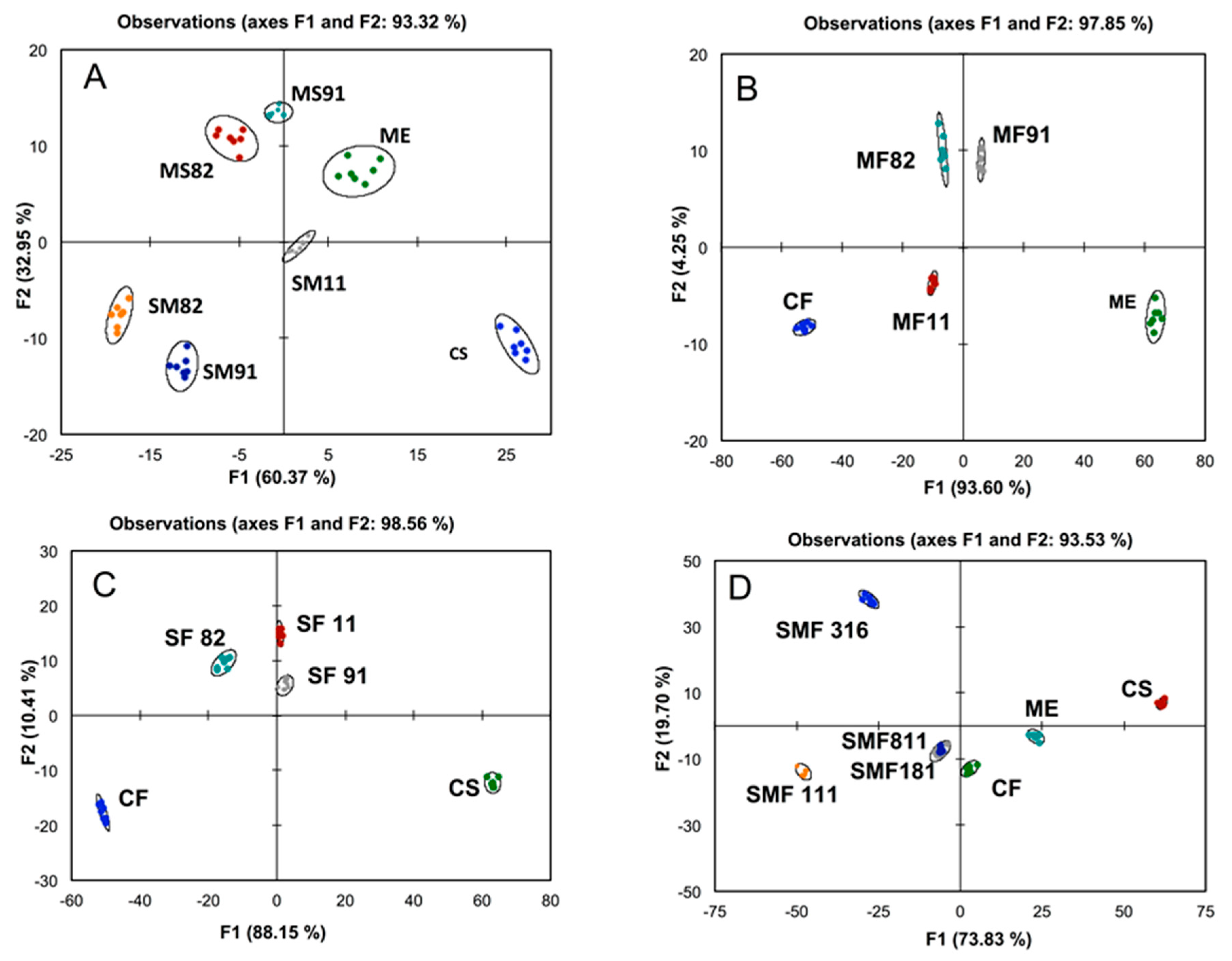
2.2. Predicting the Composition of Tri-Varietal Red Wine Blends
| C. Sauvignon | Merlot | C. Franc | ||||
|---|---|---|---|---|---|---|
| Calibration | CV 2 | Calibration | CV 2 | Calibration | CV 2 | |
| RMSE 1 | 0.10 | 0.16 | 0.065 | 0.14 | 0.067 | 0.15 |
| Bias | 0.000 | −0.007 | 0.000 | 0.003 | 0.000 | −0.004 |
| C. Sauvignon | Merlot | C. Franc | |
|---|---|---|---|
| RMSEP 1 | 0.18 | 0.15 | 0.16 |
| Bias | −0.02 | −0.01 | 0.03 |
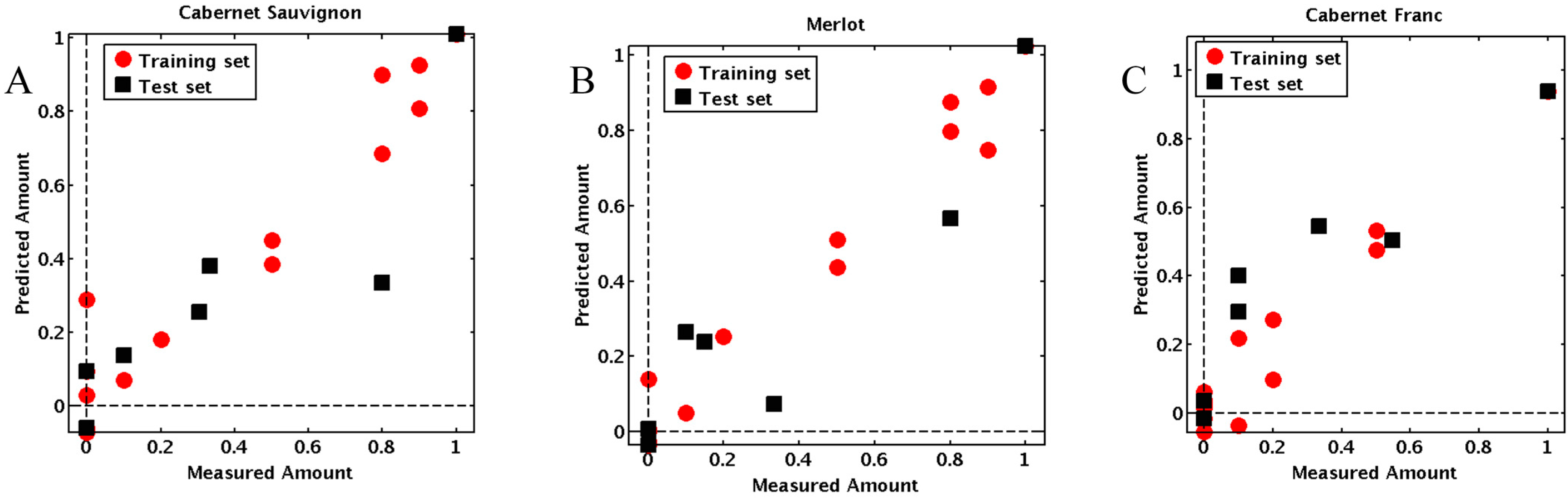
2.3. Correlation of the Peptide Receptors and Sensory Attributes of Red Wine
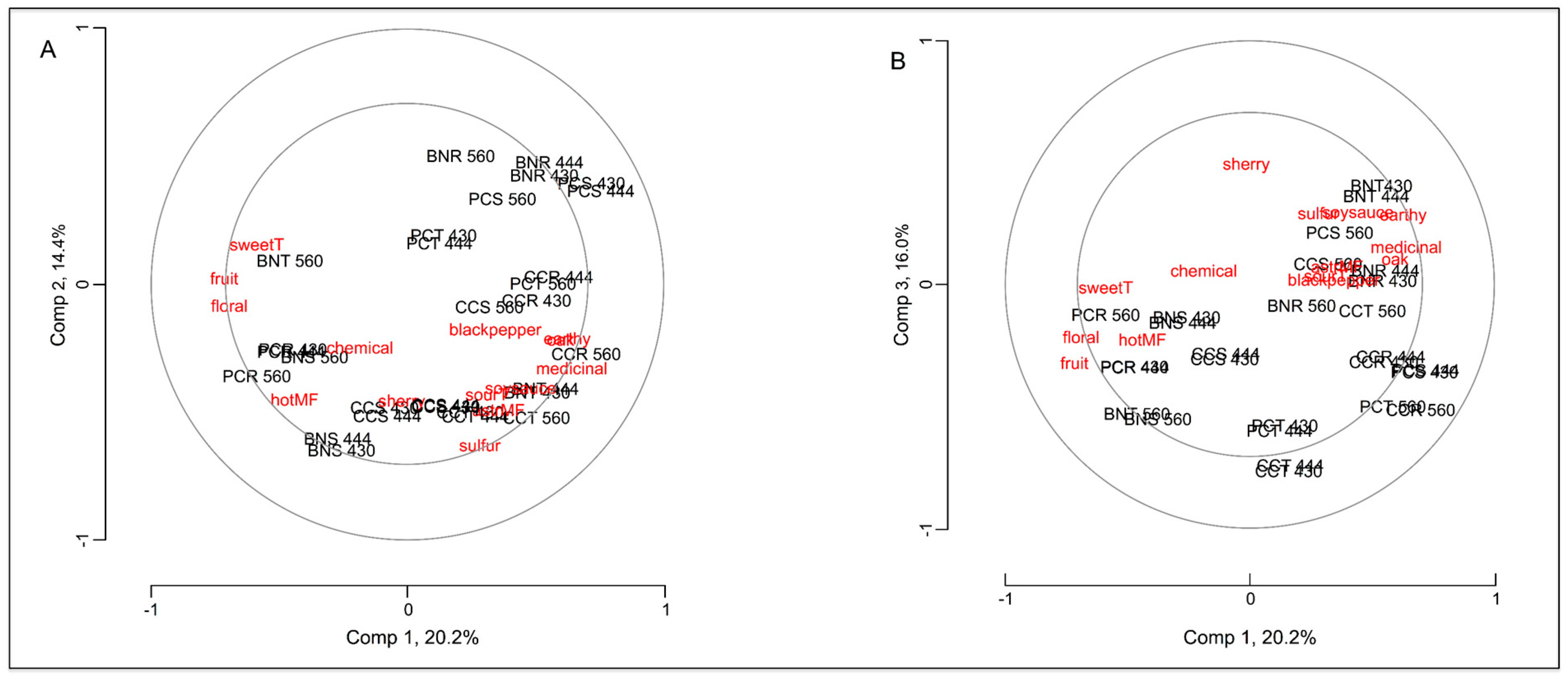
3. Experimental Section
3.1. General
| Number | Label | S | M | F |
|---|---|---|---|---|
| 1 | S | 100% | ||
| 2 | M | 100% | ||
| 3 | F | 100% | ||
| 4 | SM11 | 49.6% | 50.4% | |
| 5 | SM91 | 90.1% | 9.9% | |
| 6 | SM82 | 80.2% | 19.8% | |
| 7 | SF11 | 49.4% | 50.6% | |
| 8 | SF91 | 90.1% | 9.9% | |
| 9 | SF82 | 80.3% | 19.7% | |
| 10 | MS91 | 9.9% | 90.1% | |
| 11 | MS82 | 19.7% | 80.3 | |
| 12 | MF11 | 50.7% | 49.3% | |
| 13 | MF91 | 90.1% | 9.9% | |
| 14 | MF82 | 80.0% | 20.0% | |
| 15 | SMF111 | 33.4% | 33.2% | 33.4% |
| 16 | SMF811 | 80.0% | 10.0% | 10.0% |
| 17 | MSF811 | 10.1% | 79.9% | 10.1% |
| 18 | Blend | 30.4% | 14.9% | 54.8% |
3.2. Indicator Displacement Assay of Wine Blends
| Assembly | Code | Binding Ratio (Indicator:M2+:Peptide) | Final Indicator Concentration (mM) |
|---|---|---|---|
| PCV-Cu2+-SEL1 | PCS | 1:1:1 | 0.075 |
| PCV-Cu2+-TT2 | PCT | 1:1:0.5 | 0.075 |
| PCV-Cu2+-RN8 | PCR | 1:1:0.5 | 0.075 |
| CAS-Cu2+-SEL1 | CCS | 1:1:0.5 | 0.06 |
| CAS-Cu2+-TT2 | CCT | 1:1:0.4 | 0.06 |
| CAS-Cu2+-RN8 | CCR | 1:1:0.4 | 0.06 |
| PBR-Ni2+-SEL1 | PNS | 1:1:0.75 | 0.018 |
| PBR-Ni2+-TT2 | PNT | 1:1:0.4 | 0.018 |
| PBR-Ni2+-RN8 | PNR | 1:1:1 | 0.018 |
3.3. Statistical Data Analysis
3.4. Prediction of the Tri-Blends Composition
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hopfer, H.; Ebeler, S.E.; Heymann, H. How Blending Affects the Sensory and Chemical Properties of Red Wine. Am. J. Enol. Vitic. 2012, 63, 313–324. [Google Scholar] [CrossRef]
- Hjelmeland, A.K.; King, E.S.; Ebeler, S.E.; Heymann, H. Characterizing the Chemical and Sensory Profiles of United States Cabernet Sauvignon Wines and Blends. Am. J. Enol. Vitic. 2013, 64, 169–179. [Google Scholar] [CrossRef]
- Dooley, L.M.; Threlfall, R.T.; Jean-Francois, M.; Howard, L.R. Compositional and Sensory Impacts from Blending Red Wine Varietals. Am. J. Enol. Vitic. 2012, 63, 241–250. [Google Scholar] [CrossRef]
- Frank, Mitch. Georges Duboeuf’s Company Convicted of Fraud. Wine Spectator, 14 February 2015. Available online: http://www.winespectator.com/webfeature/show/id/Georges-Duboeufs-Company-Convicted-of-Fraud_3133 (accessed on 14 February 2015).
- Agence France-Presse. Italian police foil counterfeit Tuscan red wine scam in biggest food fraud. The Guardian. Available online: http://www.theguardian.com/world/2014/sep/11/italian-police-foil-brunello-di-montalcino-wine-scam (accessed on 14 February 2015).
- Codinachs, L.M.; Kloock, J.P.; Schoning, M.J.; Baldi, A.; Ipatov, A.; Bratova, A.; Jimenez-Jorquera, C. Electronic integrated multisensor tongue applied to grape juice and wine analysis. Analyst 2008, 133, 1440–1448. [Google Scholar] [CrossRef] [PubMed]
- Ciosek, P.; Wróblewski, W. Potentiometric electronic tongues for foodstuff and biosample recognition—An overview. Sensors 2011, 11, 4688–4701. [Google Scholar] [CrossRef] [PubMed]
- Tahara, Y.; Toko, K. Electronic tongues—A review. IEEE Sens. J. 2013, 13, 3001–3011. [Google Scholar] [CrossRef]
- Aleixandre, M.; Lozano, J.; Gutiérrez, J.; Sayago, I.; Fernandez, M.J.; Horrillo, M.C. Portable e-nose to classify different kinds of wine. Sens. Actuators B Chem. 2008, 131, 71–76. [Google Scholar] [CrossRef]
- López de Lerma, M.N.; Bellincontro, A.; García-Martínez, T.; Mencarelli, F.; Moreno, J.J. Feasibility of an electronic nose to differentiate commercial Spanish wines elaborated from the same grape variety. Food Res. Int. 2013, 51, 790–796. [Google Scholar] [CrossRef]
- Garcıa, M.; Aleixandre, M.; Guti´errez, J.; Horrillo, M.C. Electronic nose for wine discrimination. Sens. Actuators B Chem. 2006, 113, 911–916. [Google Scholar] [CrossRef]
- Garcia, M.; Fernandez, M.J.; Fontecha, J.L.; Lozano, J.; Santos, J.P.; Aleixandre, M.; Sayago, I.; Gutierrez, J.; Horrillo, M.C. Differentiation of red wines using an electronic nose based on surface acoustic wave devices. Talanta 2006, 68, 1162–1165. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, J.; Lei, Y.; Zhou, Q.; Sun, S.; Noda, I. Discrimination of different red wine by Fourier-transform infrared and two-dimensional infrared correlation spectroscopy. J. Mol. Struct. 2010, 974, 144–150. [Google Scholar] [CrossRef]
- Fernandez, K.; Agosin, E. Quantitative analysis of red wine tannins using fourier-transform mid-infrared spectrometry. J. Agric. Food Chem. 2007, 55, 7294–7300. [Google Scholar] [CrossRef] [PubMed]
- Weekley, A.J.; Bruins, P.; Sisto, M.; Augustine, M.P. Using NMR to study full intact wine bottles. J. Magn. Reson. 2003, 161, 91–98. [Google Scholar]
- Pereira, G.E.; Gaudillere, J.P.; Leeuwen, C.V.; Hilbert, G.; Lavialle, O.; Maucourt, M.; Deborde, C.; Moing, A.; Rolin, D. 1H-NMR and Chemometrics to Characterize Mature Grape Berries in Four Wine-Growing Areas in Bordeaux, France. J. Agric. Food Chem. 2005, 53, 6382–6389. [Google Scholar] [CrossRef] [PubMed]
- Lopez, R.; Aznar, M.; Cacho, J.; Ferreira, V. Determination of minor and trace volatile compounds in wine by solid-phase extraction and gas chromatography with mass spectrometric detection. J. Chromatogr. A 2002, 966, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Umali, A.P.; Anslyn, E.V. A General Approach to Differential Sensing Using Synthetic Molecular Receptors. Curr. Opin. Chem. Biol. 2010, 14, 685–692. [Google Scholar] [CrossRef] [PubMed]
- Umali, A.P.; LeBoeuf, S.E.; Newberry, R.W.; Kim, S.; Tran, L.; Rome, W.A.; Tian, T.; Taing, D.; Hong, J.; Kwan, M.; et al. Discrimination of flavonoids and red wine varietals by arrays of differential peptidic sensors. Chem. Sci. 2011, 2, 439–445. [Google Scholar] [CrossRef]
- Gallagher, L.T.; Heo, J.H.; Lopez, M.A.; Ray, B.M.; Xiao, J.; Umali, A.P.; Zhang, A.; Dharmarajan, S.; Heymann, H.; Anslyn, E.V. Pattern-based discrimination of organic acids and red wine varietals by arrays of synthetic receptors. Supramol. Chem. 2012, 24, 143–148. [Google Scholar] [CrossRef]
- Nguyen, B.T.; Anslyn, E.V.; Indicator Displacement Assays. Coord. Chem. Rev. 2006, 250, 3118–3127. [CrossRef]
- Umali, A.P.; Ghanem, E.; Hopfer, H.; Hussain, A.; Kao, Y.; Zabanal, L.G.; Wilkins, B.J.; Hobza, C.; Quach, D.K.; Fredell, M.; et al. Grape and wine sensory attributes correlate with pattern-based discrimination of Cabernet Sauvignon wines by a peptidic sensor array. Tetrahedron 2014, 1–5. [Google Scholar]
- Cala, O.; Pinaud, N.; Simon, C.; Fouquet, E.; Laguerre, M.; Dufourc, E.J.; Pianet, I. NMR and Molecular Modeling of Wine Tannins Binding to Saliva Proteins: Revisiting Astringency from Molecular and Colloidal Prospects. FASEB J. 2010, 24, 4281–4290. [Google Scholar] [CrossRef] [PubMed]
- Abdi, H. Partial Least Squares (PLS) Regression. In Encyclopedia of Social Sciences Research Methods; Lewis-Beck, M.S., Bryman, A., Liao, T.F., Eds.; Sage Publications: Thousand Oaks, CA, USA, 2003; Volumes 1–3. [Google Scholar]
- Mattivi, F.; Vrhovesk, U.; Masuero, D.; Trainotti, D. Differences in the Amount and Structure of Extractable Skin and Seed Tannins amongst Red Grape Varieties. Aust. J. Grape Wine Res. 2009, 15, 27–35. [Google Scholar] [CrossRef]
- Vidal1, S.; Francis, L.; Guyot, S.; Marnet, N.; Kwiatkowski, M.; Gawel, R.; Cheynier, V.; Waters, E.J. The Mouth-feel Properties of Grape and Apple Proanthocyanidins in a Wine-like Medium. J. Sci. Food Agric. 2003, 83, 564–573. [Google Scholar] [CrossRef]
- Yamane, T.; Jeong, S.T.; Goto-Yamamoto, N.; Koshita, Y.; Kobayashi, S. Effects of Temperature on Anthocyanin Biosynthesis in Grape Berry Skins. Am. J. Enol. Vitic. 2006, 57, 54–59. [Google Scholar]
- R: A Language and Environment for Statistical Computing; The R Foundation for Statistical Computing. R Development Core Team: Vienna, Austria, 2011. Available online: http://www.R-project.org/ (accessed on 18 May 2015).
- MATLAB 8.0 and Statistics Toolbox 8.1; The MathWorks, Inc.: Natick, MA, USA, 2012.
- Sample Availability: Samples of the three peptides used in this investigation, WAHEDEFF (TT2), WEEHEE (RN8), and FHFPHHF (SEL1) are available from the authors.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ghanem, E.; Hopfer, H.; Navarro, A.; Ritzer, M.S.; Mahmood, L.; Fredell, M.; Cubley, A.; Bolen, J.; Fattah, R.; Teasdale, K.; et al. Predicting the Composition of Red Wine Blends Using an Array of Multicomponent Peptide-Based Sensors. Molecules 2015, 20, 9170-9182. https://doi.org/10.3390/molecules20059170
Ghanem E, Hopfer H, Navarro A, Ritzer MS, Mahmood L, Fredell M, Cubley A, Bolen J, Fattah R, Teasdale K, et al. Predicting the Composition of Red Wine Blends Using an Array of Multicomponent Peptide-Based Sensors. Molecules. 2015; 20(5):9170-9182. https://doi.org/10.3390/molecules20059170
Chicago/Turabian StyleGhanem, Eman, Helene Hopfer, Andrea Navarro, Maxwell S. Ritzer, Lina Mahmood, Morgan Fredell, Ashley Cubley, Jessica Bolen, Rabia Fattah, Katherine Teasdale, and et al. 2015. "Predicting the Composition of Red Wine Blends Using an Array of Multicomponent Peptide-Based Sensors" Molecules 20, no. 5: 9170-9182. https://doi.org/10.3390/molecules20059170
APA StyleGhanem, E., Hopfer, H., Navarro, A., Ritzer, M. S., Mahmood, L., Fredell, M., Cubley, A., Bolen, J., Fattah, R., Teasdale, K., Lieu, L., Chua, T., Marini, F., Heymann, H., & Anslyn, E. V. (2015). Predicting the Composition of Red Wine Blends Using an Array of Multicomponent Peptide-Based Sensors. Molecules, 20(5), 9170-9182. https://doi.org/10.3390/molecules20059170