Proteomic Analysis of Differentially Expressed Plasma Exosome Proteins in Heat-Stressed Holstein cows
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Plasma Exosome Isolation
2.3. Identification of Plasma Exosomes
2.3.1. Transmission Electron Microscopy
2.3.2. Nanoparticle Tracking Analysis (NTA)
2.3.3. Western Blotting (WB)
2.3.4. Sample Processing
2.3.5. Liquid Chromatography–Mass Spectrometry (LC–MS)
2.3.6. Label-Free Analysis of MaxQuant
2.4. Bioinformatic Analysis
2.4.1. Cluster Analysis of Phosphorylated Peptides
2.4.2. Subcellular Localization
2.4.3. Annotation and Enrichment Analysis
2.4.4. PPI Analysis
2.5. Data Analysis
3. Results
3.1. Identification of Pasma Exosomes
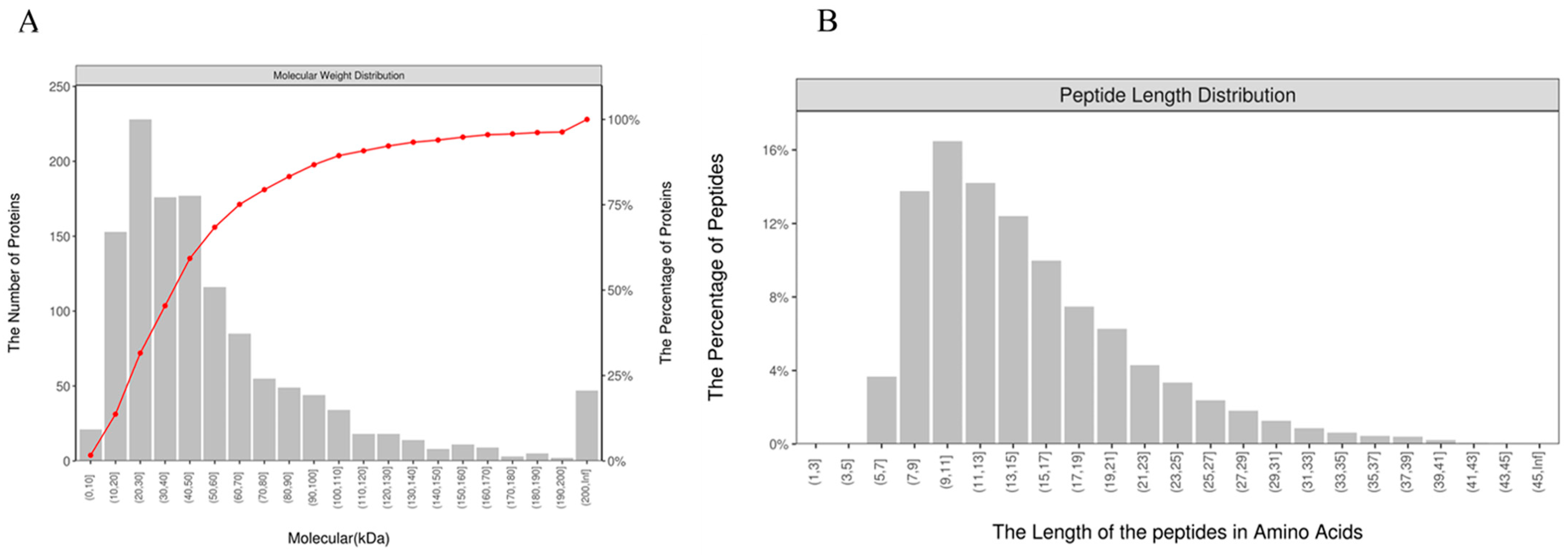
3.2. Identification of Plasma Exosome Proteins and Peptides
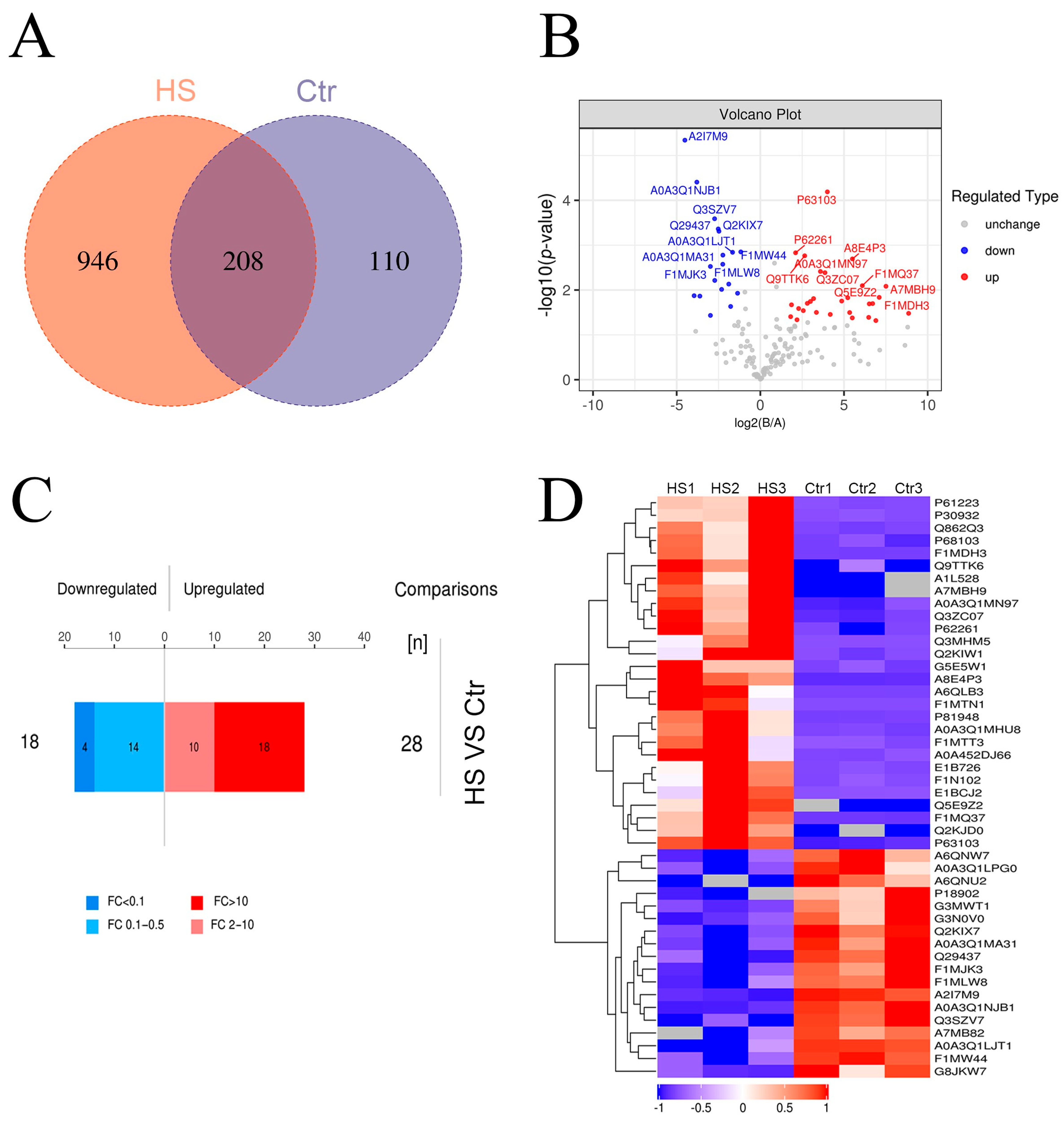
3.3. Analysis of Differential Proteins in Plasma Exosomes
3.4. Subcellular Localization Analysis
3.5. GO and KEGG Enrichment Analyses for Differentially Expressed Proteins
3.6. PPI Network of Differentially Expressed Proteins
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jordan, E.R. Effects of Heat Stress on Reproduction. J. Dairy Sci. 2003, 86, E104–E114. [Google Scholar] [CrossRef]
- West, J.W.; Mullinix, B.G.; Bernard, J.K. Effects of Hot, Humid Weather on Milk Temperature, Dry Matter Intake, and Milk Yield of Lactating Dairy Cows. J. Dairy Sci. 2003, 86, 232–242. [Google Scholar] [CrossRef]
- Tao, S.; Orellana Rivas, R.M.; Marins, T.N.; Chen, Y.C.; Gao, J.; Bernard, J.K. Impact of Heat Stress on Lactational Performance of Dairy Cows. Theriogenology 2020, 150, 437–444. [Google Scholar] [CrossRef]
- St-Pierre, N.R.; Cobanov, B.; Schnitkey, G. Economic Losses from Heat Stress by US Livestock Industries1. J. Dairy Sci. 2003, 86, E52–E77. [Google Scholar] [CrossRef]
- Baumgard, L.H.; Rhoads, R.P., Jr. Effects of Heat Stress on Postabsorptive Metabolism and Energetics. Annu. Rev. Anim. Biosci. 2013, 1, 311–337. [Google Scholar] [CrossRef]
- Li, G.; Yu, X.; Portela Fontoura, A.B.; Javaid, A.; de la Maza-Escolà, V.S.; Salandy, N.S.; Fubini, S.L.; Grilli, E.; McFadden, J.W.; Duan, J.E. Transcriptomic Regulations of Heat Stress Response in the Liver of Lactating Dairy Cows. BMC Genom. 2023, 24, 410. [Google Scholar] [CrossRef] [PubMed]
- Becker, C.A.; Collier, R.J.; Stone, A.E. Invited Review: Physiological and Behavioral Effects of Heat Stress in Dairy Cows. J. Dairy Sci. 2020, 103, 6751–6770. [Google Scholar] [CrossRef]
- Gu, Z.T.; Wang, H.; Li, L.; Liu, Y.S.; Deng, X.B.; Huo, S.F.; Yuan, F.F.; Liu, Z.F.; Tong, H.S.; Su, L. Heat Stress Induces Apoptosis through Transcription-Independent P53-Mediated Mitochondrial Pathways in Human Umbilical Vein Endothelial Cell. Sci. Rep. 2014, 4, 4469. [Google Scholar] [CrossRef]
- Sun, H.; Xia, T.; Ma, S.; Lv, T.; Li, Y. Intercellular Communication Is Crucial in the Regulation of Healthy Aging via Exosomes. Pharmacol. Res. 2025, 212, 107591. [Google Scholar] [CrossRef]
- Ciardiello, C.; Cavallini, L.; Spinelli, C.; Yang, J.; Reis-Sobreiro, M.; De Candia, P.; Minciacchi, V.R.; Di Vizio, D. Focus on Extracellular Vesicles: New Frontiers of Cell-to-Cell Communication in Cancer. Int. J. Mol. Sci. 2016, 17, 175. [Google Scholar] [CrossRef]
- da Silveira, J.C.; Veeramachaneni, D.N.R.; Winger, Q.A.; Carnevale, E.M.; Bouma, G.J. Cell-Secreted Vesicles in Equine Ovarian Follicular Fluid Contain Mirnas and Proteins: A Possible New Form of Cell Communication within the Ovarian Follicle. Biol. Reprod. 2012, 86, 71–72. [Google Scholar] [CrossRef]
- Tkach, M.; Théry, C. Communication by Extracellular Vesicles: Where We Are and Where We Need to Go. Cell 2016, 164, 1226–1232. [Google Scholar] [CrossRef]
- Chen, H.; Han, Z.; Ma, Y.; Meng, Q. Advances in Macrophage-Derived Exosomes as Immunomodulators in Disease Progression and Therapy. Int. Immunopharmacol. 2024, 142, 113248. [Google Scholar] [CrossRef]
- Saeed-Zidane, M.; Linden, L.; Salilew-Wondim, D.; Held, E.; Neuhoff, C.; Tholen, E.; Hoelker, M.; Schellander, K.; Tesfaye, D. Cellular and Exosome Mediated Molecular Defense Mechanism in Bovine Granulosa Cells Exposed to Oxidative Stress. PLoS ONE 2017, 12, e0187569. [Google Scholar] [CrossRef] [PubMed]
- Xiong, M.; Chen, Z.; Tian, J.; Peng, Y.; Song, D.; Zhang, L.; Jin, Y. Exosomes Derived from Programmed Cell Death: Mechanism and Biological Significance. Cell Commun. Signal. 2024, 22, 156. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Guo, J.; Han, C.; Yang, M.; Cao, X. Heat Shock Protein 70, Released from Heat-Stressed Tumor Cells, Initiates Antitumor Immunity by Inducing Tumor Cell Chemokine Production and Activating Dendritic Cells via TLR4 Pathway. J. Immunol. 2009, 182, 1449–1459. [Google Scholar] [CrossRef]
- Chen, T.; Guo, J.; Yang, M.; Zhu, X.; Cao, X. Chemokine-Containing Exosomes Are Released from Heat-Stressed Tumor Cells via Lipid Raft-Dependent Pathway and Act as Efficient Tumor Vaccine. J. Immunol. 2011, 186, 2219–2228. [Google Scholar] [CrossRef]
- Clayton, A.; Turkes, A.; Navabi, H.; Mason, M.D.; Tabi, Z. Induction of Heat Shock Proteins in B-Cell Exosomes. J. Cell Sci. 2005, 118, 3631–3638. [Google Scholar] [CrossRef]
- Chen, H.S.; Tong, H.S.; Zhao, Y.; Hong, C.Y.; Bin, J.P.; Su, L. Differential Expression Pattern of Exosome Long Non-Coding RNAs (LncRNAs) and MicroRNAs (MiRNAs) in Vascular Endothelial Cells Under Heat Stroke. Med. Sci. Monit. 2018, 24, 7965–7974. [Google Scholar] [CrossRef]
- Wang, Y.; Fang, J.; Zeng, H.F.; Zhong, J.F.; Li, H.X.; Chen, K.L. Identification and Bioinformatics Analysis of Differentially Expressed Milk Exosomal MicroRNAs in Milk Exosomes of Heat-Stressed Holstein Cows. Funct. Integr. Genom. 2022, 22, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Li, S.; Du, C.; Zhang, Y.; Li, Y.; Chu, L.; Han, X.; Galons, H.; Zhang, Y.; Sun, H.; et al. Exosomes from Different Cells: Characteristics, Modifications, and Therapeutic Applications. Eur. J. Med. Chem. 2020, 207, 112784. [Google Scholar] [CrossRef] [PubMed]
- Vlassov, A.V.; Magdaleno, S.; Setterquist, R.; Conrad, R. Exosomes: Current Knowledge of Their Composition, Biological Functions, and Diagnostic and Therapeutic Potentials. Biochim. Biophys. Acta-Gen. Subj. 2012, 1820, 940–948. [Google Scholar] [CrossRef]
- Yu, J.; Yin, P.; Liu, F.; Cheng, G.; Guo, K.; Lu, A.; Zhu, X.; Luan, W.; Xu, J. Effect of Heat Stress on the Porcine Small Intestine: A Morphological and Gene Expression Study. Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2010, 156, 119–128. [Google Scholar] [CrossRef]
- Pearce, S.C.; Sanz-Fernandez, M.V.; Hollis, J.H.; Baumgard, L.H.; Gabler, N.K. Short-Term Exposure to Heat Stress Attenuates Appetite and Intestinal Integrity in Growing Pigs1. J. Anim. Sci. 2014, 92, 5444–5454. [Google Scholar] [CrossRef]
- Polsky, L.; von Keyserlingk, M.A.G. Invited Review: Effects of Heat Stress on Dairy Cattle Welfare. J. Dairy Sci. 2017, 100, 8645–8657. [Google Scholar] [CrossRef]
- Liu, J.; Li, L.; Chen, X.; Lu, Y.; Wang, D. Effects of Heat Stress on Body Temperature, Milk Production, and Reproduction in Dairy Cows: A Novel Idea for Monitoring and Evaluation of Heat Stress—A Review. Asian-Australas. J. Anim. Sci. 2019, 32, 1332–1339. [Google Scholar] [CrossRef] [PubMed]
- Luigi, P.; Claudia, F.; Mauro, C.; Giampiero, C. Heat Stress Affects the Cytoskeleton and the Delivery of Sucrose Synthase in Tobacco Pollen Tubes. Planta Int. J. Plant Biol. 2016, 243, 43–63. [Google Scholar]
- Weis, F.; Moullintraffort, L.; Heichette, C.; Chrétien, D.; Garnier, C. The 90-KDa Heat Shock Protein Hsp90 Protects Tubulin against Thermal Denaturation. J. Biol. Chem. 2010, 285, 9525–9534. [Google Scholar] [CrossRef] [PubMed]
- Aitken, A.; Collinge, D.B.; van Heusden, B.P.H.; Isobe, T.; Roseboom, P.H.; Rosenfeld, G.; Soll, J. 14-3-3 Proteins: A Highly Conserved, Widespread Family of Eukaryotic Proteins. Trends Biochem. Sci. 1992, 17, 498–501. [Google Scholar] [CrossRef]
- Yano, M.; Nakamuta, S.; Wu, X.; Okumura, Y.; Kido, H. A Novel Function of 14-3-3 Protein: 14-3-3ζ Is a Heat-Shock-Related Molecular Chaperone That Dissolves Thermal-Aggregated Proteins. Mol. Biol. Cell 2006, 17, 4769–4779. [Google Scholar] [CrossRef]
- Kumsta, C.; Ching, T.T.; Nishimura, M.; Davis, A.E.; Gelino, S.; Catan, H.H.; Yu, X.; Chu, C.C.; Ong, B.; Panowski, S.H.; et al. Integrin-Linked Kinase Modulates Longevity and Thermotolerance in C. Elegans through Neuronal Control of HSF-1. Aging Cell 2014, 13, 419–430. [Google Scholar] [CrossRef] [PubMed]
- Bos, M.A.; van ‘t Veer, C.; Reitsma, P.H. Molecular biology and biochemistry of the coagulation factors and pathways of hemostasis. In Williams Hematology, 9th ed.; Kaushansky, K., Lichtman, M.A., Prchal, J.T., Levi, M.M., Press, O.W., Burns, L.J., Caligiuri, M., Eds.; McGraw-Hill Education: New York, NY, USA, 2015; pp. 614–616. [Google Scholar]
- Palta, S.; Saroa, R.; Palta, A. Overview of the Coagulation System. Indian J. Anaesth. 2014, 58, 515. [Google Scholar] [CrossRef]
- Iba, T.; Connors, J.M.; Levi, M.; Levy, J.H. Heatstroke-Induced Coagulopathy: Biomarkers, Mechanistic Insights, and Patient Management. eClinicalMedicine 2022, 44, 101276. [Google Scholar] [CrossRef]
- Negrutskii, B.S.; Shalak, V.F.; Novosylna, O.V.; Porubleva, L.V.; Lozhko, D.M.; El’skaya, A.V. The EEF1 Family of Mammalian Translation Elongation Factors. BBA Adv. 2023, 3, 100067. [Google Scholar] [CrossRef]
- Vera, M.; Pani, B.; Griffiths, L.A.; Muchardt, C.; Abbott, C.M.; Singer, R.H.; Nudler, E. The Translation Elongation Factor EEF1A1 Couples Transcription to Translation during Heat Shock Response. eLife 2014, 3, e03164. [Google Scholar] [CrossRef]
- Yun, C.W.; Kim, H.J.; Lim, J.H.; Lee, S.H. Heat Shock Proteins: Agents of Cancer Development and Therapeutic Targets in Anti-Cancer Therapy. Cells 2019, 9, 60. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.; Yang, J.; Qi, Z.; Wu, H.; Wang, B.; Zou, F.; Mei, H.; Liu, J.; Wang, W.; Liu, Q. Heat Shock Proteins: Biological Functions, Pathological Roles, and Therapeutic Opportunities. MedComm 2022, 3, e161. [Google Scholar] [CrossRef]
- Mazurakova, A.; Solarova, Z.; Koklesova, L.; Caprnda, M.; Prosecky, R.; Khakymov, A.; Baranenko, D.; Kubatka, P.; Mirossay, L.; Kruzliak, P.; et al. Heat Shock Proteins in Cancer—Known but Always Being Rediscovered: Their Perspectives in Cancer Immunotherapy. Adv. Med. Sci. 2023, 68, 464–473. [Google Scholar] [CrossRef]
- Min, L.; Zheng, N.; Zhao, S.; Cheng, J.; Yang, Y.; Zhang, Y.; Yang, H.; Wang, J. Long-Term Heat Stress Induces the Inflammatory Response in Dairy Cows Revealed by Plasma Proteome Analysis. Biochem. Biophys. Res. Commun. 2016, 471, 296–302. [Google Scholar] [CrossRef]
- Choi, Y.; Park, H.; Kim, J.; Lee, H.; Kim, M. Heat Stress Induces Alterations in Gene Expression of Actin Cytoskeleton and Filament of Cellular Components Causing Gut Disruption in Growing–Finishing Pigs. Animals 2024, 14, 2476. [Google Scholar] [CrossRef] [PubMed]
- Doshi, B.M.; Hightower, L.E.; Lee, J. The Role of Hsp27 and Actin in the Regulation of Movement in Human Cancer Cells Responding to Heat Shock. Cell Stress Chaperones 2009, 14, 445–457. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, Z.; Huang, S.; Wang, X.; He, S.; Liu, L.; Hu, Y.; Chen, L.; Chen, P.; Liu, S.; et al. Adipocyte IRE1α Promotes PGC1α MRNA Decay and Restrains Adaptive Thermogenesis. Nat. Metab. 2022, 4, 1166–1184. [Google Scholar] [CrossRef]
- Malini, M.K.; Lekshmy, V.S.; Pal, M.; Chinnusamy, V.; Kumar, M.N. Unfolded Protein Response (UPR) Mediated under Heat Stress in Plants. Plant Physiol. Rep. 2020, 25, 569–582. [Google Scholar] [CrossRef]
- Singh, M.B.; Lohani, N.; Bhalla, P.L. The Role of Endoplasmic Reticulum Stress Response in Pollen Development and Heat Stress Tolerance. Front. Plant Sci. 2021, 12, 661062. [Google Scholar] [CrossRef]
- Lai, E.; Teodoro, T.; Volchuk, A. Endoplasmic Reticulum Stress: Signaling the Unfolded Protein Response. Physiology 2007, 22, 193–201. [Google Scholar] [CrossRef]
- Cao, S.S.; Kaufman, R.J. Unfolded Protein Response. Curr. Biol. 2012, 22, R622–R626. [Google Scholar] [CrossRef]
- Wang, S.; Deng, L.; Zhao, J.; Zhu, X.; Shu, G.; Wang, L.; Gao, P.; Jiang, Q. Effects of Heat Stress on Antioxidant Capacity and Gastrocnemius Musclefiber Types of Broilers. J. South China Agric. Univ. 2015, 36, 23–28. [Google Scholar]
- Liu, Y.; Cai, H.; Guo, X.; Aierken, A.; Hua, J.; Ma, B.; Peng, S. Melatonin Alleviates Heat Stress-Induced Testicular Damage in Dairy Goats by Inhibiting the PI3K/AKT Signaling Pathway. Stress Biol. 2022, 2, 47. [Google Scholar] [CrossRef]
- Thompson, S.M.; Callstrom, M.R.; Jondal, D.E.; Butters, K.A.; Knudsen, B.E.; Anderson, J.L.; Lien, K.R.; Sutor, S.L.; Lee, J.S.; Thorgeirsson, S.S.; et al. Heat Stress-Induced PI3K/MTORC2-Dependent AKT Signaling Is a Central Mediator of Hepatocellular Carcinoma Survival to Thermal Ablation Induced Heat Stress. PLoS ONE 2016, 11, e0162634. [Google Scholar] [CrossRef]




| Group | Parity | Lactation Days (d) | Average Rectal Temperature/°C | Temperature and Humidity Index (THI) | Average Breathing Rate (Times/Min) |
|---|---|---|---|---|---|
| Ctr1 | 2 | 126 | 38.40 ± 0.22 | 55.41 ± 1.49 | 47 ± 2.03 |
| Ctr2 | 2 | 128 | 38.43 ± 0.09 | 55.41 ± 1.49 | 52 ± 1.78 |
| Ctr3 | 2 | 132 | 38.60 ± 0.08 | 57.87 ± 3.33 | 53 ± 2.10 |
| HS1 | 2 | 125 | 39.40 ± 0.36 | 79.57 ± 0.89 | 106 ± 2.65 |
| HS2 | 2 | 130 | 39.50 ± 0.24 | 77.44 ± 3.43 | 96 ± 2.00 |
| HS3 | 2 | 131 | 39.83 ± 0.34 | 79.95 ± 3.32 | 108 ± 2.65 |
| Item | Value |
|---|---|
| Enzyme | Trypsin |
| Max missed cleavages | 2 |
| Main search | 6 ppm |
| First search | 20 ppm |
| MS/MS tolerance | 20 ppm |
| Fixed modifications | Carbamidomethyl (C) |
| Variable modifications | Oxidation (M) |
| Database | uniport_Bovine_46480_20200217.fasta |
| Database pattern | Reverse |
| Include contaminants | True |
| Protein FDR | ≤0.01 |
| Peptide FDR | ≤0.01 |
| Peptides used for protein quantification | Used razor and unique peptides |
| Time window (match between runs) | 2 min |
| Protein quantification | LFQ |
| Min. ratio count | 1 |
| Upregulated | Downregulated | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Protein Symbol | Protein Name | Gene Name | FC * | p-Value | Protein Symbol | Protein Name | Gene Name | FC * | p-Value |
| F1MTN1 | Integrin beta | ITGB3 | 465.04 | 0.0331196 | A2I7M9 | Serpin A3-2 | SERPINA3-2 | 0.04 | 4.56 × 10−6 |
| A7MBH9 | G protein subunit alpha i2 | GNAI2 | 181.86 | 0.0082595 | G8JKW7 | Serpin A3-7 | SERPINA3-7 | 0.06 | 0.013396 |
| F1MDH3 | Talin 1 | TLN1 | 137.56 | 0.0145823 | A0A3Q1NJB1 | Ceruloplasmin | CP | 0.07 | 3.93 × 10−5 |
| E1BCJ2 | Complement factor H related 5 | CFHR5 | 119.67 | 0.047979 | G3MWT1 | Ig-like domain-containing protein | IGLDCPs | 0.08 | 0.013672 |
| A1L528 | small monomeric GTPase | RAB1A | 104.68 | 0.0200544 | A6QNW7 | CD5 molecule-like | CD5L | 0.1 | 0.006112 |
| A6QLB3 | Integrin subunit alpha 2b | ITGA2B | 91.14 | 0.020363 | F1MJK3 | Uncharacterized protein | LOC506828 | 0.12 | 0.002985 |
| Q3MHM5 | Tubulin beta-4B chain | TUBB4B | 88.96 | 0.0407212 | P18902 | Retinol-binding protein 4 | RBP4 | 0.13 | 0.03687 |
| F1MQ37 | Myosin-9 | MYH9 | 68.31 | 0.0079969 | Q3SZV7 | Hemopexin | HPX | 0.15 | 0.000258 |
| A8E4P3 | Stomatin | STOM | 45.32 | 0.0020229 | Q29437 | Primary amine oxidase, liver isozyme | LOC100138645 | 0.17 | 0.000435 |
| P30932 | CD9 antigen | CD9 | 45.12 | 0.0418379 | Q2KIX7 | Protein HP-25 homolog 1 | LOC511240 | 0.18 | 0.000489 |
| A0A452DJ66 | Tubulin alpha chain | TUBA1D | 40.44 | 0.0317772 | G3N0V0 | Ig-like domain-containing protein | IGLDCPs | 0.2 | 0.009688 |
| Q5E9Z2 | Hyaluronan-binding protein 2 | HABP2 | 37.37 | 0.0148506 | A0A3Q1MA31 | Inter-alpha-trypsin inhibitor heavy chain H4 | ITIH4 | 0.21 | 0.001666 |
| P81948 | Tubulin alpha-4A chain | TUBA4A | 29.09 | 0.017609 | F1MLW8 | Uncharacterized protein | - | 0.21 | 0.002664 |
| P61223 | Ras-related protein Rap-1b | RAP1B | 18.11 | 0.0350055 | A6QNU2 | IVL protein | IVL | 0.27 | 0.007375 |
| P63103 | 14-3-3 protein zeta/delta | YWHAZ | 16.06 | 6.474 × 10−5 | A7MB82 | Complement C1q tumor necrosis factor-related protein 3 | C1QTNF3 | 0.29 | 0.023276 |
| Q3ZC07 | Actin, alpha cardiac muscle 1 | ACTC1 | 14.54 | 0.0041034 | A0A3Q1LJT1 | Ig-like domain-containing protein | IGLDCPs | 0.32 | 0.001441 |
| A0A3Q1MN97 | Vinculin | VCL | 12.05 | 0.0038576 | A0A3Q1LPG0 | Ig-like domain-containing protein | - | 0.39 | 0.0118 |
| E1B726 | Plasminogen | PLG | 10.19 | 0.0315644 | F1MW44 | Coagulation factor XIII A chain | F13A1 | 0.44 | 0.001412 |
| P68103 | Elongation factor 1-alpha 1 | EEF1A1 | 9.07 | 0.015542 | |||||
| Q2KJD0 | Tubulin beta-5 chain | TUBB5 | 7.96 | 0.0180104 | |||||
| Q862Q3 | Beta-2-microglobulin | B2M | 6.99 | 0.0198202 | |||||
| Q9TTK6 | Membrane primary amine oxidase | AOC3 | 6.28 | 0.0017246 | |||||
| G5E5W1 | Coagulation factor VIII | F8 | 5.93 | 0.0288047 | |||||
| Q2KIW1 | Paraoxonase 1 | PON1 | 4.88 | 0.0258445 | |||||
| F1MTT3 | Coagulation factor XII | F12 | 4.57 | 0.0460769 | |||||
| P62261 | 14-3-3 protein epsilon | YWHAE | 4.30 | 0.0014777 | |||||
| A0A3Q1MHU8 | Complement factor properdin | CFP | 3.63 | 0.0213611 | |||||
| F1N102 | Complement C8 beta chain | C8B | 3.511 | 0.0392651 | |||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xia, S.; Jiang, Y.; Li, W.; An, Z.; Shen, Y.; Ding, Q.; Chen, K. Proteomic Analysis of Differentially Expressed Plasma Exosome Proteins in Heat-Stressed Holstein cows. Animals 2025, 15, 2286. https://doi.org/10.3390/ani15152286
Xia S, Jiang Y, Li W, An Z, Shen Y, Ding Q, Chen K. Proteomic Analysis of Differentially Expressed Plasma Exosome Proteins in Heat-Stressed Holstein cows. Animals. 2025; 15(15):2286. https://doi.org/10.3390/ani15152286
Chicago/Turabian StyleXia, Shuwen, Yingying Jiang, Wenjie Li, Zhenjiang An, Yangyang Shen, Qiang Ding, and Kunlin Chen. 2025. "Proteomic Analysis of Differentially Expressed Plasma Exosome Proteins in Heat-Stressed Holstein cows" Animals 15, no. 15: 2286. https://doi.org/10.3390/ani15152286
APA StyleXia, S., Jiang, Y., Li, W., An, Z., Shen, Y., Ding, Q., & Chen, K. (2025). Proteomic Analysis of Differentially Expressed Plasma Exosome Proteins in Heat-Stressed Holstein cows. Animals, 15(15), 2286. https://doi.org/10.3390/ani15152286






