A New Methodology to Comprehend the Effect of El Niño and La Niña Oscillation in Early Warning of Anthrax Epidemic Among Livestock
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.1.1. Disease Incidence and ENSO Events Classification
2.1.2. Livestock Data
2.1.3. Meteorological Data
2.1.4. Remote Sensing Data
2.1.5. Soil Profile
2.2. Spatial Endemicity
2.3. Getis-Ord Gi* Spatial Statistics to Identify Hotspots (Spatial Autocorrelation)
2.4. Space-Time Cluster Analysis
2.5. Identifying Risk Factors by Linear Discriminant Analysis
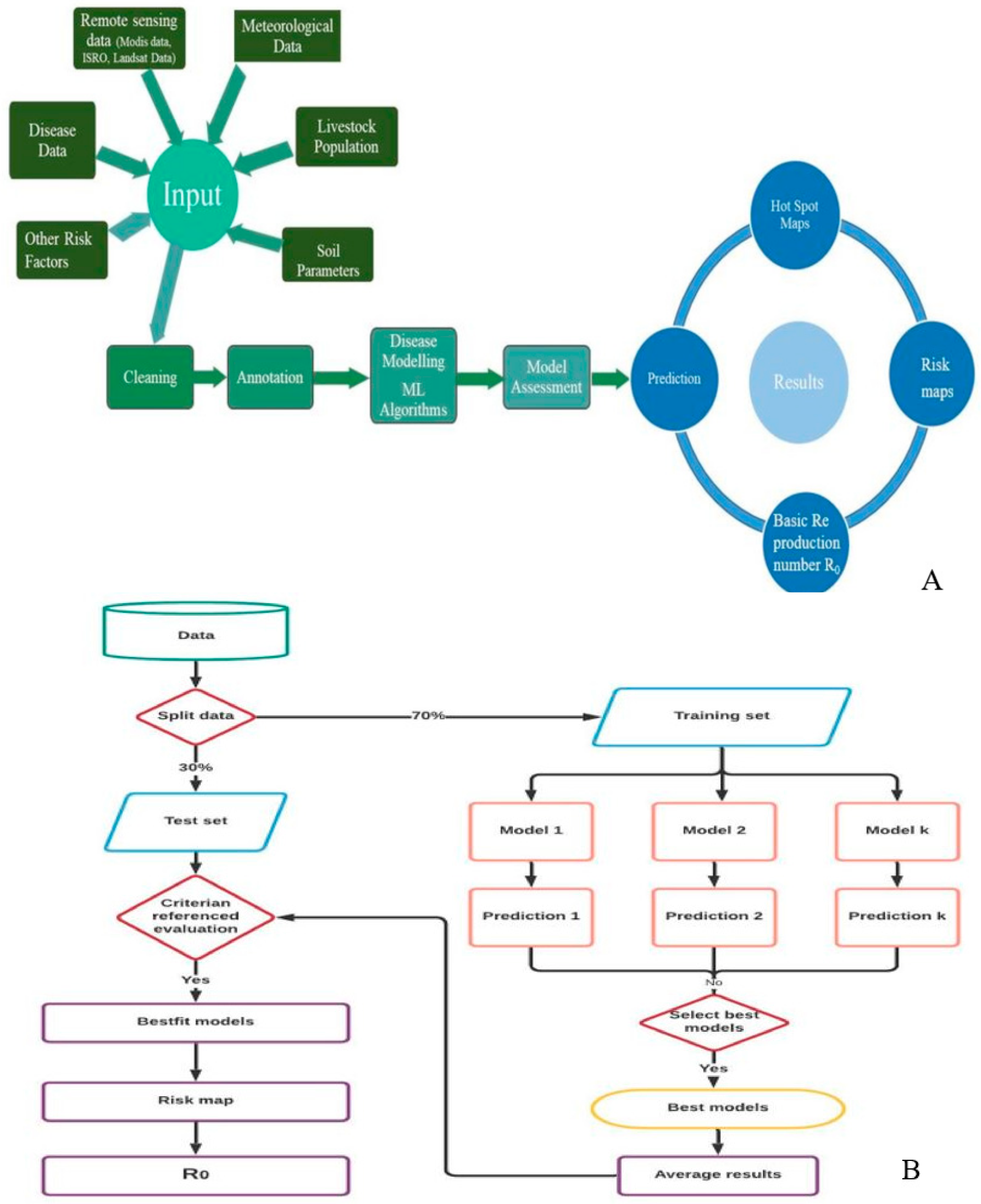
2.6. Risk Modelling and Mapping
2.7. Basic Reproduction Number (R0)
2.7.1. Exponential Growth Rate (EG)
2.7.2. Maximum Likelihood Estimate (ML)
2.7.3. Attack Rate Estimate (AR)
2.8. Statistical Software
3. Results
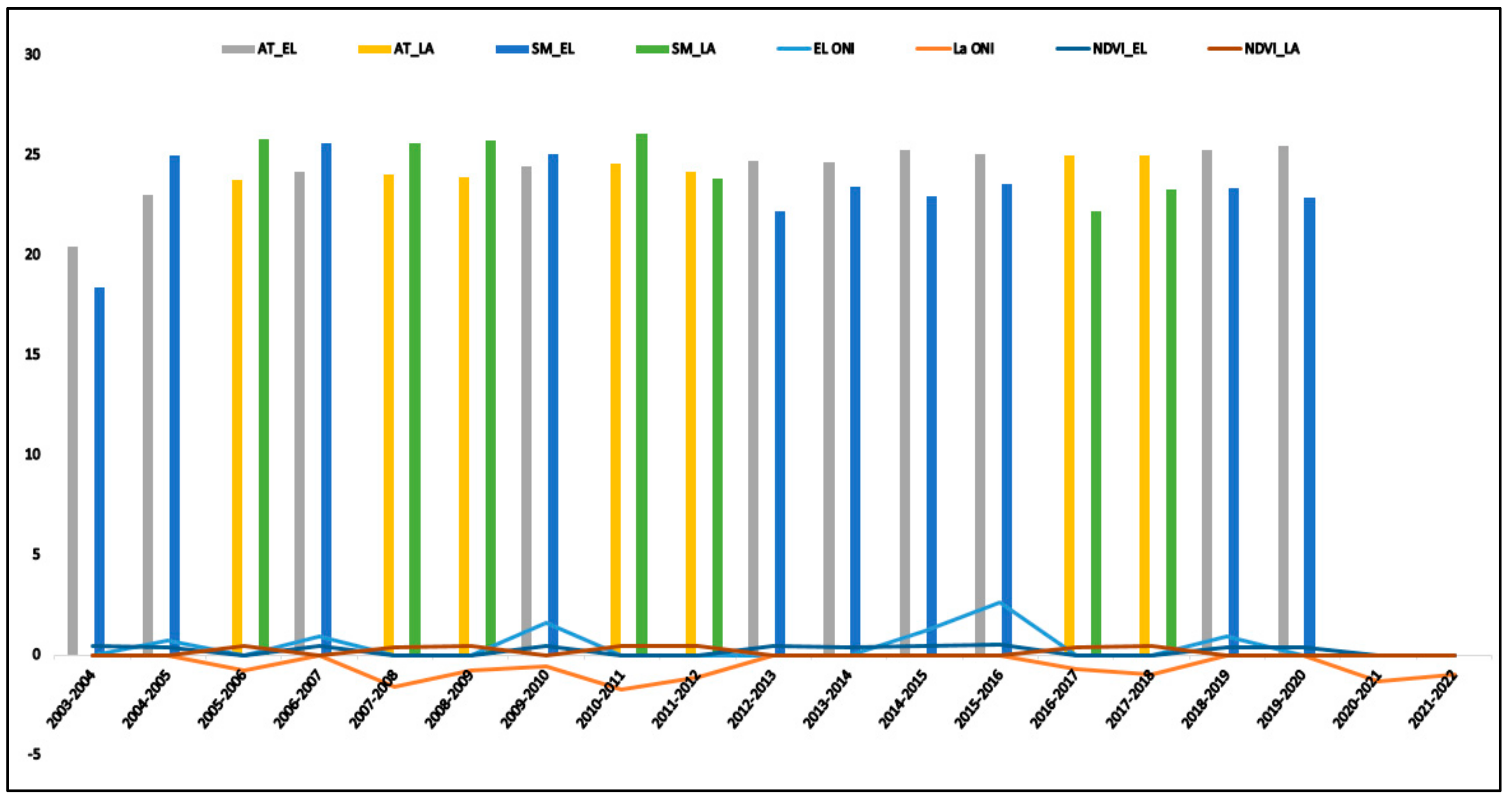
3.1. Temporal Distribution of Weather Parameters
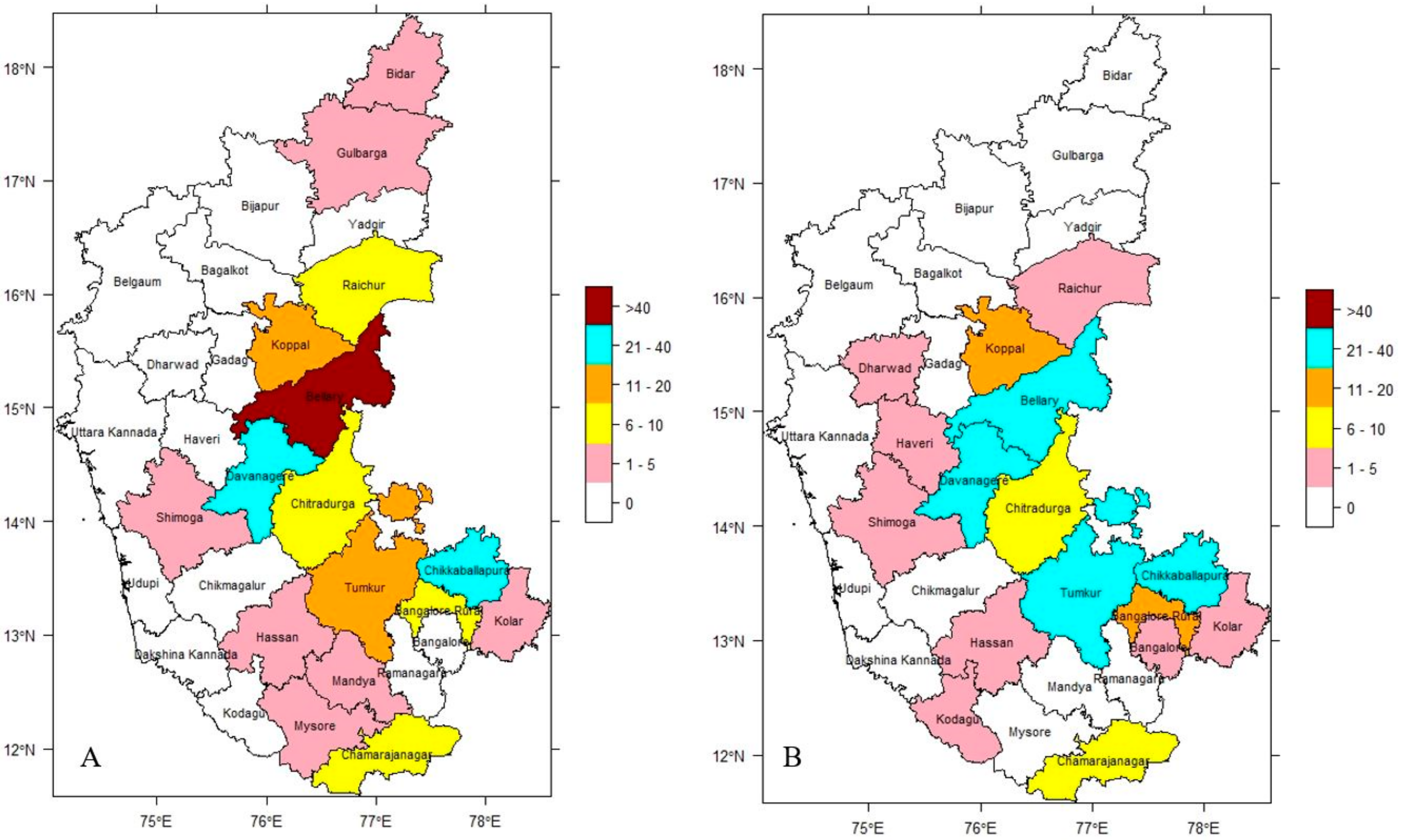
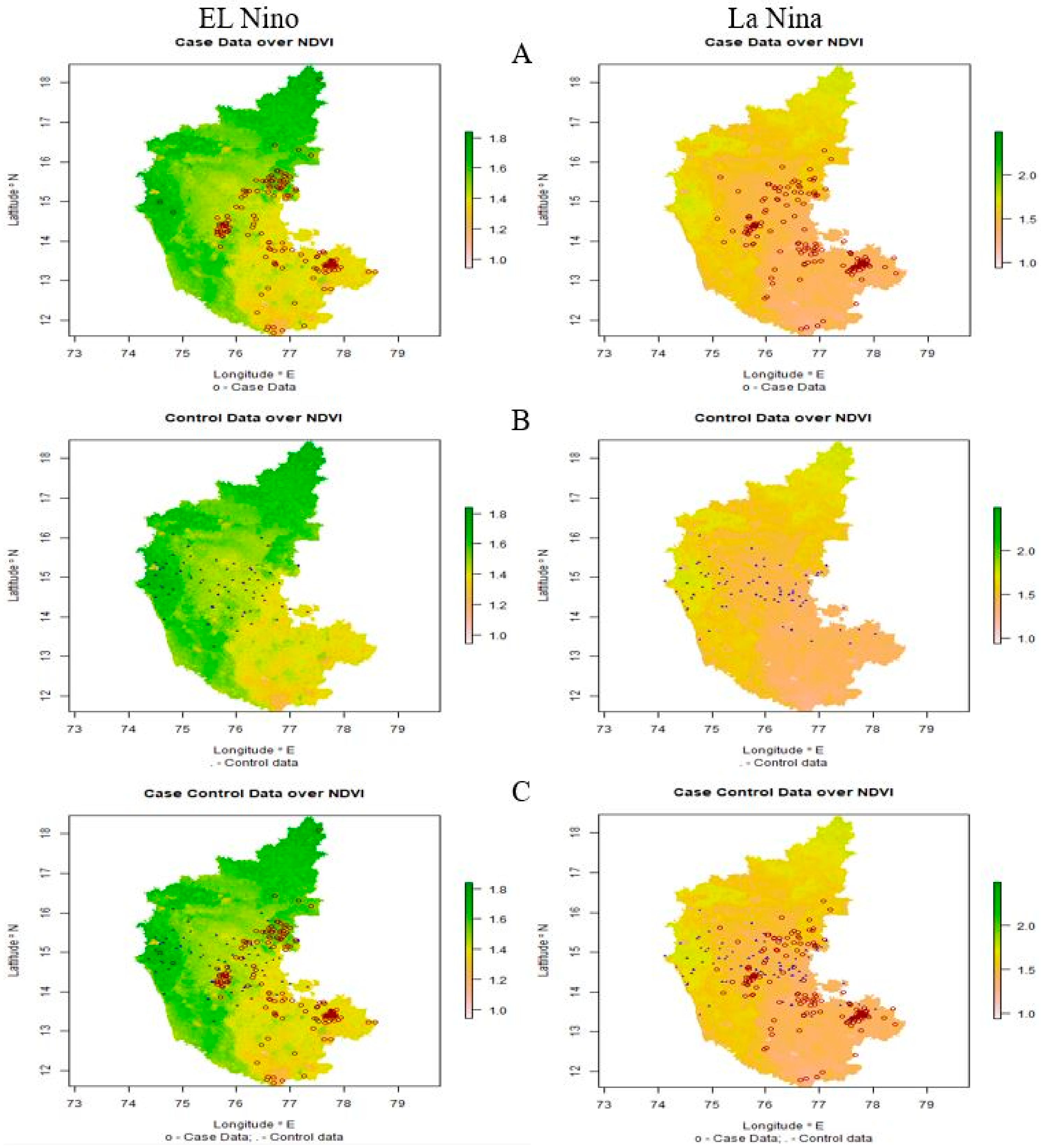
3.2. Spatial Endemicity of Anthrax
3.3. Spatial Autocorrelation of Anthrax
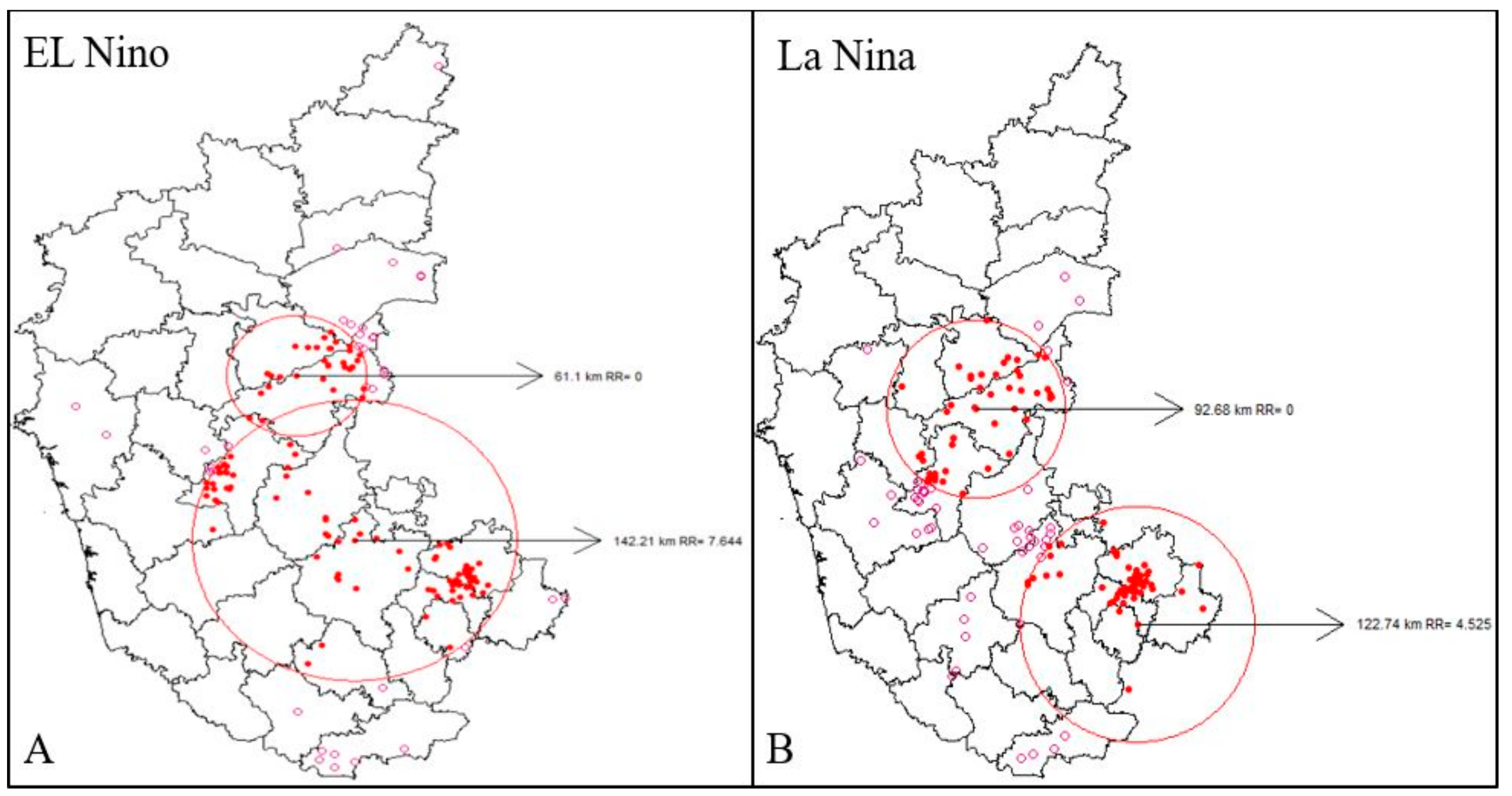
3.4. Space-Time Cluster Analysis of Anthrax
3.5. Linear Discriminant Analysis of Anthrax
3.6. Anthrax Risk Assessment and Estimation
3.7. Anthrax Risk Prediction and Mapping
3.8. Estimation of Basic Reproduction Number (R0) of Anthrax
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Serkan, O.; Selcen, O.; Serhan, S. Anthrax—An overview. Med. Sci. Monit. 2003, 9, 276–283. [Google Scholar]
- Jayachandran, R. Anthrax: Biology of Bacillus anthracis. Curr. Sci. 2002, 82, 1220–1226. [Google Scholar]
- Jocelyn, M.; Larissa, L.; Alim, A.; Yerlan, P.; Mathew, V.E.; Jason, K.B. Ecological Niche Modelling of the Bacillus anthracis A1.a sub-lineage in Kazakhstan. BMC Ecol. 2011, 11, 32. [Google Scholar]
- Turnbull, P.C.B. Guidelines for the Surveillance and Control of Anthrax in Humans and Animals; WHO/EMC/ZDI/98.6, Wiltshire SP4 0JG; World Health Organization: Geneva, Switzerland, 1998. [Google Scholar]
- Gombe, N.T.; Nkomo, B.M.M.; Chadambuka, A.; Shambira, G.; Tshimanga, M. Risk factors for contracting anthrax in Kuwirirana ward, Gokwe North, Zimbabwe. Afr. Health Sci. 2010, 10, 159–164. [Google Scholar]
- Madeleine, C.T.; Angel, G.M.; Remi, C.; Joy, S.G. Climate drivers of vector-borne diseases in Africa and their relevance to control programmes. Infect. Dis. Poverty 2018, 7, 81. [Google Scholar]
- Myers, M.F.; Rogers, D.J.; Cox, J.; Flahault, A.; Hay, S.I. Forecasting Disease Risk for Increased Epidemic Preparedness in Public. Health Adv. Parasitol. 2000, 47, 309–330. [Google Scholar]
- Michael, G.W.; de Smalen, A.W.; Siobhan, M.M. Climatic influence on anthrax suitability in warming northern latitudes. Sci. Rep. 2018, 8, 9269. [Google Scholar]
- Troccoli, A. Review Seasonal climate forecasting. Metrol. Appl. 2010, 17, 251–268. [Google Scholar] [CrossRef]
- Kriss, A.B.; Paul, P.A.; Madden, L.V. Variability in Fusarium Head Blight Epidemics in Relation to Global Climate Fluctuations as Represented by the El Niño-Southern Oscillation and Other Atmospheric Patterns. Ecol. Epidemiol. 2012, 55, 64. [Google Scholar] [CrossRef]
- Malay, P.; Poonam, S.; Kumar, G.; Ojha, V.P.; Dhiman, R.C. El Niño Southern Oscillation as an early warning tool for dengue outbreak in India. BMC Public Health 2020, 20, 1498. [Google Scholar]
- Cane, M.A. The evolution of El Niño, past and future. Earth Planet. Sci. Lett. 2005, 230, 227–240. [Google Scholar] [CrossRef]
- Saini, S.; Ashok, G. El Niño and Indian Droughts: A Scoping Exercise; No. 276, Working Paper; Indian Council for Research on International Economic Relations (ICRIER): New Delhi, Indian, 2014. [Google Scholar]
- Cherian, S.; Sridhara, S.; Manoj, K.N.; Gopakkali, P.; Ramesh, N.; Alrajhi, A.A.; Dewidar, A.Z.; Mattar, M.A. Impact of El Niño Southern Oscillation on Rainfall and Rice Production: A Micro-Level Analysis. Agronomy 2021, 11, 1021. [Google Scholar] [CrossRef]
- Anna, M.S.; Rachel, L. Climate and Non-Climate Drivers of Dengue Epidemics in Southern Coastal Ecuador. Am. J. Trop. Med. Hyg. 2013, 88, 971–981. [Google Scholar]
- Pinto, J.; Bonacic, C.; Hamilton-West, C.; Romero, J.; Lubroth, J. Climate change and animal diseases in South America. Rev. Sci. Tech. 2008, 27, 599–613. [Google Scholar] [CrossRef] [PubMed]
- Debashish, D. Machine learning algorithms for disease prediction: A methodological Review in Biomedical. In Proceedings of the 3rd Global Conference on Computing and Media Technology, Kuala Lumpur, Malaysia, 17–18 July 2019. [Google Scholar]
- Carella, E.; Orusa, T.; Viani, A.; Meloni, D.; Borgogno-Mondino, E.; Orusa, R. An Integrated, Tentative Remote-Sensing Approach Based on NDVI Entropy to Model Canine Distemper Virus in Wildlife and to Prompt Science-Based Management Policies. Animals 2022, 12, 1049. [Google Scholar] [CrossRef]
- Orusa, T.; Mondino, E.B. Exploring Short-term climate change effects on rangelands and broad-leaved forests by free satellite data in Aosta Valley (Northwest Italy). Climate 2021, 9, 47. [Google Scholar] [CrossRef]
- Orusa, T.; Orusa, R.; Viani, A.; Carella, E.; Mondino, E.B. Geomatics and EO data to support wildlife diseases assessment at landscape level: A pilot experience to map infectious keratoconjunctivitis in chamois and phenological trends in Aosta Valley (NW Italy). Remote Sens. 2020, 12, 3542. [Google Scholar] [CrossRef]
- Ward, M.M. Estimating disease prevalence and incidence using administrative data: Some assembly required. J. Rheumatol. 2013, 40, 1241–1243. [Google Scholar] [CrossRef]
- Ranganath, R.N.; Jayaraman, V.; Roy, P.S. Remote sensing applications: An overview. Curr. Sci. 2007, 93, 1747–1766. [Google Scholar]
- Suma, A.P.; Suresh, K.P.; Gajendragad, M.R.; Kavya, B.A. Forecasting Anthrax in Livestock in Karnataka State using Remote Sensing and Climatic Variables. Int. J. Sci. Res. 2015, 6, 1891–1897. [Google Scholar]
- Justice, C.O.; Townshend, J.R.G.; Vermote, E.F.; Masuoka, E.; Wolf, R.E.; Saleous, N.; Roy, D.P.; Morisette, J.T. An overview of MODIS Land data processing and product status. Remote Sens. Environ. 2002, 83, 3–15. [Google Scholar] [CrossRef]
- Hay, S.I.; Tatem, A.J.; Graham, A.J.; Goetz, S.J.; Rogers, D.J. Global Environmental Data for Mapping Infectious Disease Distribution. Adv Parasitol. 2006, 62, 37–77. [Google Scholar] [PubMed]
- Margaret, D.; Rwego, I.B.; Ndimuligo, S.A.; Travis, D.A.; Mwakapeje, E.R.; Craft, M.; Asiimwe, B.; Alvarez, J.; Ayebare, S.; Pelican, K. Environmental determinants influencing anthrax distribution in Queen Elizabeth Protected Area, Western Uganda. PLoS ONE 2020, 15, e0237223. [Google Scholar]
- Horvath, D.J.; Reid, R.L. Indirect effects of soil and water on Animal health. Sci. Total Environ. 1984, 34, 143–156. [Google Scholar] [CrossRef]
- Lawrence, N.K.; Adamson, S.M.; Christopher, C.A.; Immo, K. Modelling the effect of malaria endemicity on spatial variations in childhood fever, diarrhoea and pneumonia in Malawi. Int. J. Health Geogr. 2007, 6, 33. [Google Scholar]
- Songchitruksa, P.; Zeng, X. Getis–Ord Spatial Statistics to Identify Hot Spots by Using Incident Management Data. Transp. Res. Rec. 2010, 2165, 42–51. [Google Scholar] [CrossRef]
- Tobler, W.R. A Computer movie simulating urban growth in the detroit region. Econ. Geogr. 1970, 46, 234–240. [Google Scholar] [CrossRef]
- Michael, J.; Sara, G.; Caitlin, K. Spatial Modeling in Environmental and Public Health Research. Int. J. Environ. Res. Public Health. 2010, 7, 1302–1329. [Google Scholar]
- Muluken, A.; Abera, K.; Alemayehu, W.; Bagtzoglou, A.C. Childhood Diarrhea Exhibits Spatiotemporal Variation in Northwest Ethiopia: A SaTScan Spatial Statistical Analysis. PLoS ONE 2015, 10, e0144690. [Google Scholar]
- Marlize, C.; Michael, C.; Aaron, M.M.; Gerdalize, K.; Maureen, C.; David, N.D. Using the SaTScan method to detect local malaria clusters for guiding malaria control programmes. Malar J. 2009, 8, 68. [Google Scholar]
- Shahera, B.; Wenbiao, H.; Cameron, H.; Yuming, G.; Mohammad, Z.I.; Shilu, T. Space-time clusters of dengue fever in Bangladesh. Trop. Med. Int. Health 2012, 17, 1086–1091. [Google Scholar]
- Izenman, A.J. Linear Discriminant Analysis. In Springer Texts in Statistics; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Pankaj, M.; Ching-Hao, W.; Alexandre Day, G.R.; Clint, R.; Marin, B.; Charles, K.F.; David, J.S. A high-bias, low-variance introduction to Machine Learning for physicists. Phys. Rep. 2019, 810, 1–124. [Google Scholar]
- Omri, A.; Asaf, T.; Ronen, K. Assessing the accuracy of species distribution models: Prevalence, kappa and the true skill statistic (TSS). J. Appl. 2006, 43, 1223–1232. [Google Scholar]
- Andrius, V.; Emma, G.; Ellen, P.; Alexander, J.C. Machine learning algorithm validation with a limited sample size. PLoS ONE 2019, 14, e0224365. [Google Scholar]
- Johnson, K.; Vinay, K.K.; Phridviraj, M.S.B.; Shaik, R. Reducing Overfitting Problem in Machine Learning Using Novel L1/4 Regularization Method. In Proceedings of the Fourth International Conference on Trends in Electronics and Informatics, Tirunelveli, India, 16–18 April 2020. [Google Scholar]
- Jake, L.; Martin, K.; Naomi, A. Model selection and overfitting. Nat. Methods 2016, 13, 703–704. [Google Scholar]
- Farzin, S.; Lalit, K.; Mohsen, A. Assessing Accuracy Methods of Species Distribution Models: AUC, Specificity, Sensitivity and the True Skill Statistic. Glob. J. 2018, 18, 1–13. [Google Scholar]
- Ferri, C.; Hernández-Orallo, J.; Modroiu, R. An experimental comparison of performance measures for classification. Pattern Recognit. Lett. 2009, 30, 27–38. [Google Scholar] [CrossRef]
- Li, Y.; Wang, L.-W.; Peng, Z.-H.; Shen, H.-B. Basic reproduction number and predicted trends of coronavirus disease 2019 epidemic in the mainland of China. Infect. Dis. Poverty. 2020, 9, 1–13. [Google Scholar] [CrossRef]
- Mahmud, R.; Patwari, H.A.F. Estimation of the Basic Reproduction Number of SARS-CoV-2 in Bangladesh Using Exponential Growth Method. Infect Dis. Poverty 2020, 9, 94. [Google Scholar]
- Froda, S.; Leduc, H. Estimating the basic reproduction number from surveillance data on past epidemics. Math. Biosci. 2014, 256, 89–101. [Google Scholar] [CrossRef]
- Obadia, T.; Haneef, R.; Boëlle, P.-Y. The R0 package: A toolbox to estimate reproduction numbers for epidemic outbreaks. BMC Med. Inform. Decis. Mak. 2012, 12, 147. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Bingham, D.; Braun, W.J.; Hu, X.J. Moran’s I statistic-based nonparametric test with spatio-temporal observations. J. Nonparametric Stat. 2019, 31, 244–267. [Google Scholar] [CrossRef]
- Hamid, R.K.; Seyed, H.H.; Mehrdad, F.; Mohammad, A.H.; Nasir, A. Early warning system models and components in emergency and disaster: A systematic literature review protocol. Syst. Rev. 2019, 8, 315. [Google Scholar]
- Fekri, D.; Kamran, A.; Claudia, B.; Claire, J.S.; Ali, A.; Albrecht, J. Assessment of electronic disease early warning system for improved disease surveillance and outbreak response in Yemen. BMC Public Health 2020, 20, 1422. [Google Scholar]
- Ian, T.K.; Lile, M.; Nikoloz, T.; Julietta, M.; Lela, B.; Paata, I.; Shota, T.; Jason, K.B. Evidence of Local Persistence of Human Anthrax in the Country of Georgia Associated with Environmental and Anthropogenic Factors. PLoS Negl. Trop. Dis. 2013, 7, 9. [Google Scholar]
- Sean, B.; Thuillier, G.; Barlier, F. The DTM-2000 empirical thermosphere model with new data assimilation and constraints at lower boundary: Accuracy and properties. J. Atmos. Sol.-Terr. Phys. 2003, 65, 1053–1070. [Google Scholar]
- Dragon, D.C.; Rennie, R.P. The ecology of anthrax spores: Tough but not invincible. Can. Vet. J. 1995, 36, 295–301. [Google Scholar]
- Gunaseelan, L.; Rishikesavan, R.; Adarsh, T.; Baskar, R.; Hamilton, E.; Kaneene, J.B. Temporal and geographical distribution of animal anthrax in Tamil Nadu state, India. Tamilnadu J. Vet. Anim. Sci. 2011, 7, 277–284. [Google Scholar]
- Turner, W.C.; Imologhome, P.; Havarua, Z.; Kaaya, G.P.; Mfune, J.K.E.; Mpofu, I.D.T.; Getz, W.M. Soil ingestion, nutrition and the seasonality of anthrax in herbivores of Etosha National Park. Ecosphere 2013, 4, 1–19. [Google Scholar] [CrossRef]
- Bylaiah, S.; Shedole, S.; Suresh, K.P.; Gowda, L.; Shivananda, B.; Shivamallu, C.; Patil, S.S. Disease Prediction Model to Assess the Impact of Changes in Precipitation Level on the Risk of Anthrax Infectiousness among the Livestock Hosts in Karnataka, India. Int. J. Spec. Educ. 2022, 37, 711–727. [Google Scholar]
| Parameters | El Niño | La Niña | ||||||
|---|---|---|---|---|---|---|---|---|
| Mean | SD | Max. | Min. | Mean | SD | Max. | Min. | |
| Air temperature (k) | 23.91 | 1.69 | 27.43 | 13.21 | 23.16 | 1.69 | 27.13 | 13.21 |
| Soil moisture (kg/m2) | 23.37 | 2.31 | 28.87 | 13.21 | 23.94 | 2.42 | 30.15 | 13.21 |
| Rainfall (kg/m2/s) | 1.07 | 0.96 | 14.74 | 0.89 | 1.06 | 0.96 | 14.74 | 0.89 |
| Vegetative index (−1 to +1) | 0.41 | 0.09 | 0.71 | -0.03 | 0.43 | 0.09 | 0.73 | −0.09 |
| Parameters | Mean (Presence) | SD | F Value | p Value | 95% CI |
|---|---|---|---|---|---|
| Air_Temperature | 23.91 | 1.69 | 37.83 | <0.05 * | 23.13 to 24.68 |
| EVI | 1.32 | 0.11 | 0.54 | 0.466 | 1.31 to 1.326 |
| LAI | 2.01 | 2.44 | 0.78 | 0.379 | 1.84 to 2.17 |
| LST | 27.9 | 3.26 | 1.04 | 0.309 | 27.65 to 28.14 |
| NDVI | 1.44 | 0.12 | 0.00 | 0.947 | 1.43 to 1.441 |
| PET | 943.96 | 259.63 | 2.49 | 0.116 | 913.31 to 974.60 |
| Potential_evaporation_rate | 245.87 | 33.59 | 4.66 | 0.032 * | 240.45 to 251.28 |
| Rain_precipitation_rate | 1.07 | 0.96 | 0.65 | 0.423 | 1.01 to 1.12 |
| Soil_moisture | 23.37 | 2.31 | 0.58 | 0.449 | 23.23 to 23.50 |
| Specific_humidity | 1.08 | 0.96 | 2.35 | 0.128 | 0.97 to 1.19 |
| Surface_Pressure | 85,809.19 | 6584.11 | 1.83 | 0.178 | 85,143.10 to 86,475.27 |
| Wind_speed | 4.19 | 0.99 | 22.32 | <0.05 * | 3.84 to 4.54 |
| Parameter | Mean (Presence) | SD | F Value | p Value | 95% CI |
|---|---|---|---|---|---|
| Air_Temperature | 23.16 | 1.69 | 22.535 | <0.05 * | 22.59 to 23.73 |
| EVI | 1.31 | 0.1 | 3.922 | 0.049 | 1.30 to 1.32 |
| LAI | 2.03 | 2.44 | 2.576 | 0.111 | 1.75 to 2.31 |
| LST | 26.96 | 3.32 | 0.622 | 0.431 | 26.77 to 27.15 |
| NDVI | 1.45 | 0.12 | 7.959 | 0.005 * | 1.43 to 1.47 |
| PET | 1015.63 | 249.43 | 0.383 | 0.537 | 1004.69 to 1026.57 |
| Potential_evaporation_rate | 218.82 | 32.09 | 0.770 | 0.382 | 216.82 to 220.82 |
| Rain_precipitation_rate | 1.06 | 0.96 | 2.732 | 0.100 | 0.95 to 1.17 |
| Soil_moisture | 23.94 | 2.42 | 3.087 | 0.081 ** | 23.64 to 24.24 |
| Specific_humidity | 1.07 | 0.96 | 9.857 | 0.002 * | 0.86 to 1.28 |
| Surface_Pressure | 85,602.37 | 6581.58 | 3.074 | 0.082 | 84,784.33 to 86,420.41 |
| Wind_speed | 3.72 | 0.95 | 22.909 | <0.05 * | 3.40 to 4.04 |
| Sl.No | Model | Model Specification | KAPPA | ROC | TSS | AUC | ACCURACY | ERROR RATE | F1 SCORE | LOGLOSS |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | GLM | Y—Expected Value, X—Conditional, —Linear Predicator, g—Link Function | 0.698 | 0.943 | 0.769 | 0.9426 | 0.875 | 0.125 | 0.885 | 0.295 |
| 2 | GAM | Y—Response Variable, g—Link Function, fi—Specified Parametric Form, xi—Predicator Variable | 0.698 | 0.943 | 0.769 | 0.9426 | 0.875 | 0.125 | 0.885 | 0.295 |
| 3 | RF | Y f(tn) Y—Average of aggregated predictions of the multiple decision trees, tn—multiple decision trees trained on different subset of the same training data | 0.849 | 0.999 | 0.972 | 0.9987 | 0.982 | 0.018 | 0.980 | 0.107 |
| 4 | GBM | m—Iteration, —Learning Rate, —Step length | 0.634 | 0.966 | 0.863 | 0.9660 | 0.932 | 0.068 | 0.936 | 0.254 |
| 5 | NNET | Y—Output, —Inputs, —Weights, —Bias | 0.004 | 0.527 | 0 | 0.5269 | 0.629 | 0.371 | 0.500 | NA |
| 6 | MARS | —Constant Coefficient, —Basis Function | 0.595 | 0.942 | 0.773 | 0.9416 | 0.885 | 0.115 | 0.902 | 0.301 |
| 7 | FDA | 0.625 | 0.805 | 0.609 | 0.8047 | 0.832 | 0.168 | 0.873 | 5.818 | |
| 8 | CT | 0.764 | 0.95 | 0.787 | 0.9504 | 0.889 | 0.111 | 0.921 | 0.255 | |
| 9 | SVM | 0.645 | 0.931 | 0.759 | 0.9307 | 0.871 | 0.129 | 0.878 | 0.439 | |
| 10 | NB | —Posterior Probability, —Likelihood, —Class Prior Probability, —Predictor Prior Probability | −0.46 | 0.819 | −0.309 | 0.8189 | 0.319 | 0.681 | 0.143 | 9.099 |
| 11 | ADA | —Weak Learner, —Input,th Positive or Negative Classifier | 0.838 | 0.924 | 0.847 | 0.9237 | 0.925 | 0.075 | 0.941 | 2.600 |
| Sl.No | Model | Model Specification | KAPPA | ROC | TSS | AUC | ACCURACY | ERROR RATE | F1SCORE | LOGLOSS |
| 1 | GLM | Y—Expected Value, X—Conditional, —Linear Predicator, g—Link Function | 0.472 | 0.867 | 0.624 | 0.8670 | 0.803 | 0.197 | 0.886 | 0.419 |
| 2 | GAM | Y—Response Variable, g—Link Function, fi—Specified Parametric Form, xi—Predicator Variable | 0.472 | 0.867 | 0.624 | 0.8670 | 0.803 | 0.197 | 0.886 | 0.419 |
| 3 | RF | Y f(tn) Y—Average of aggregated predictions of the multiple decision trees, tn—multiple decision trees trained on different subset of the same training data | 0.765 | 1 | 0.99 | 0.9998 | 0.997 | 0.003 | 0.995 | 0.093 |
| 4 | GBM | m—Iteration, —Learning Rate, —Step length | 0.629 | 0.973 | 0.85 | 0.9725 | 0.926 | 0.074 | 0.939 | 0.238 |
| 5 | NNET | Y—Output, —Inputs, —Weights, —Bias | 0.018 | 0.512 | 0.025 | 0.5125 | 0.671 | 0.329 | 0.500 | NA |
| 6 | MARS | —Constant Coefficient, —Basis Function | 0.595 | 0.952 | 0.779 | 0.9524 | 0.893 | 0.107 | 0.912 | 0.274 |
| 7 | FDA | 0.543 | 0.75 | 0.5 | 0.7498 | 0.813 | 0.187 | 0.870 | 6.469 | |
| 8 | CT | 0.69 | 0.937 | 0.784 | 0.9367 | 0.893 | 0.107 | 0.918 | 0.270 | |
| 9 | SVM | 0.689 | 0.938 | 0.749 | 0.9379 | 0.876 | 0.124 | 0.914 | 0.388 | |
| 10 | NB | —Posterior Probability, —Likelihood, —Class Prior Probability, —Predictor Prior Probability | −0.302 | 0.714 | −0.199 | 0.7136 | 0.301 | 0.699 | 0.076 | 9.829 |
| 11 | ADA | Weak Learner, —Input,th Positive or Negative Classifier | 0.886 | 0.94 | 0.88 | 0.9399 | 0.950 | 0.050 | 0.963 | 1.733 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Suresh, K.P.; Bylaiah, S.; Patil, S.; Kumar, M.; Indrabalan, U.B.; Panduranga, B.A.; Srinivas, P.T.; Shivamallu, C.; Kollur, S.P.; Cull, C.A.; et al. A New Methodology to Comprehend the Effect of El Niño and La Niña Oscillation in Early Warning of Anthrax Epidemic Among Livestock. Zoonotic Dis. 2022, 2, 267-290. https://doi.org/10.3390/zoonoticdis2040022
Suresh KP, Bylaiah S, Patil S, Kumar M, Indrabalan UB, Panduranga BA, Srinivas PT, Shivamallu C, Kollur SP, Cull CA, et al. A New Methodology to Comprehend the Effect of El Niño and La Niña Oscillation in Early Warning of Anthrax Epidemic Among Livestock. Zoonotic Diseases. 2022; 2(4):267-290. https://doi.org/10.3390/zoonoticdis2040022
Chicago/Turabian StyleSuresh, Kuralayanapalya Puttahonnappa, Sushma Bylaiah, Sharanagouda Patil, Mohan Kumar, Uma Bharathi Indrabalan, Bhavya Anenahalli Panduranga, Palya Thimmaiah Srinivas, Chandan Shivamallu, Shiva Prasad Kollur, Charley A. Cull, and et al. 2022. "A New Methodology to Comprehend the Effect of El Niño and La Niña Oscillation in Early Warning of Anthrax Epidemic Among Livestock" Zoonotic Diseases 2, no. 4: 267-290. https://doi.org/10.3390/zoonoticdis2040022
APA StyleSuresh, K. P., Bylaiah, S., Patil, S., Kumar, M., Indrabalan, U. B., Panduranga, B. A., Srinivas, P. T., Shivamallu, C., Kollur, S. P., Cull, C. A., & Amachawadi, R. G. (2022). A New Methodology to Comprehend the Effect of El Niño and La Niña Oscillation in Early Warning of Anthrax Epidemic Among Livestock. Zoonotic Diseases, 2(4), 267-290. https://doi.org/10.3390/zoonoticdis2040022









