Heart Rate Variability by Dynamical Patterns in Windows of Holter Electrocardiograms: A Method to Discern Left Ventricular Hypertrophy in Heart Transplant Patients Shortly after the Transplant
Abstract
1. Introduction
2. Materialsand Methods
2.1. The Dynamical Landscape Method of ECG Processing
- 1.
- Extraction of R-event time moments from QRS complexes in the acquired ECG recording and their annotation as normal or abnormal;
- 2.
- Preprocessing annotated series of R-event time moments to a signal with normal-to-normal RR intervals, i.e., to a signal with lengths of time intervals between two consecutive heart contractions annotated as normal; in case the resulting RR interval was to short ( ms) or too long ( ms), the RR interval was replaced by the median of the surrounding 7 RR intervals, i.e., 3 preceding, 3 following, and by itself;
- 3.
- Estimates of HRV on RR intervals by the chosen HRV measures.
2.2. Complexity Measures Used for HRV Assessment
- —
- probability of inflection points: ;
- —
- probability of alternation segments: ;
- —
- probability of short segments: .
2.3. Visualization of Complexity of RR Increments
- –
- probability matrix with elementsThus .
- –
- transition matrix T where an element is the probability that increment occurs given increment happened:Thus , and
- –
- entropic matrices:matrix of Shannon entropy with elements:matrix of entropy of transition rates with elements:tensor of self-transfer entropy with elements
2.4. Study Population
- –
- LVM, calculated according to the linear `cube’ method formula of Devereux and Reichek;
- –
- LVMI (LVM index): the ratio of LVM with respect to the body surface area (BSA) to normalize heart mass measurement in subjects with different body sizes;
- –
- RWT (relative wall thickness): to report the relationship between the wall thickness and ventricle size.
- NG:
- when RWT < 0.42 and LVMI <115 g/m in the case of a man and LVMI < 95 g/m in the case of a woman;
- CR:
- when RWT ≥ 0.42 and LVMI <115 g/m in the case of a man and LVMI < 95 g/m in the case of a woman;
- H:
- when LVMI ≥115 g/m in the case of a man and LVMI ≥ 95 g/m in the case of a woman, independently of RWT value.
2.5. ECG Signals Processing
2.6. HRV Measures Estimates
2.7. HRV Analysis of Segmented Signals
- (I)
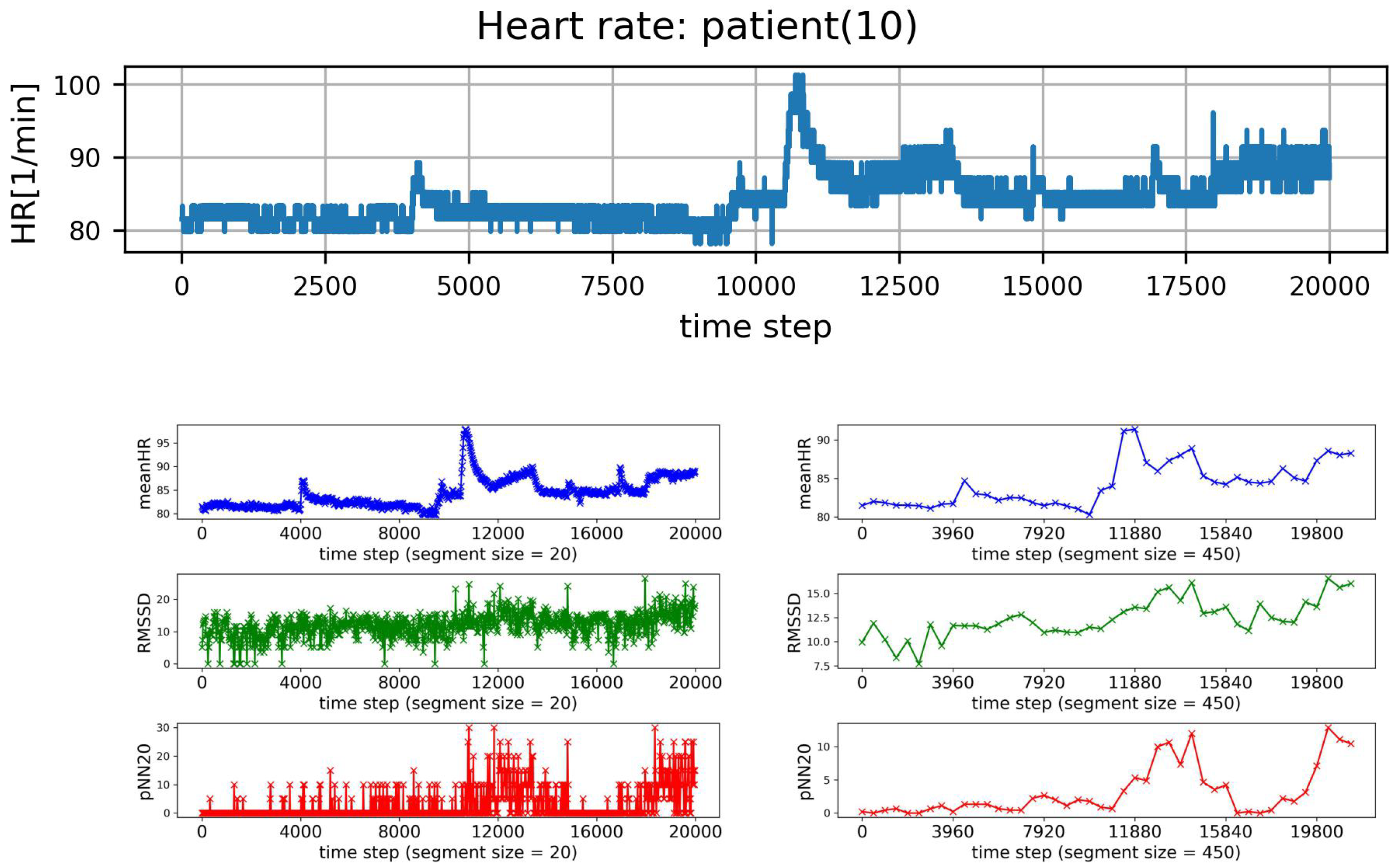
- HRV of an individual segment: segments corresponding to the lowest and the greatest value of HR were extracted, and then HRV analysis was performed for each of these special segments only;
- (II)
- complexity in HRV measure values for a given window size: the variability among the HRV series was investigated by the standard deviation of HRV series and by SampEn to assess whether two similar consecutive L points from a series remain similar if we add the next th point to each subsequence. The estimates of SampEn were performed assuming and
2.8. Statistical Analysis of Data
3. Results
3.1. HRV of Whole Signals
- (1)
- Probability of the no-change event p(zero) is the greatest for the CR group.
- (2)
- Patterns consisting of two–three elements of alternating a and d, i.e., the probabilities: p(ad), p(da), p(ada), p(dad), and corresponding entropies: e(ad), e(da), e(ada), e(dad), are less prevalent in the series of the CR group than in the other groups.
- (3)
- Above observations are in agreement with the lowest values of the probability of points-of-inflection PIP = p(ad) + p(da). The entropic measures: ShE_L, L=1,2,3, S_T and sTE attain the lowest values for signals from the CR group.
- (4)
- The short-term variability measures: pNN20, pNN50, RMSSD display group properties similar to p(da).
- (5)
- The highest values of the studied pattern measures were attained for signals of the NG group. The frequency domain measures, the long-range variability measure, such as SDNN, and the short-range time-domain measures pNN20, and pNN50 took the highest values for signals of H group.
- (6)
- In all groups, the medians of S_T were lower than ShE_1 and sTE were significantly greater than 0, which means that the dynamics of changes in RR increments is richer than in a simple Markov chain. The greatest memory effects were revealed in the signals from the NG group, the smallest in the signals from the CR group.
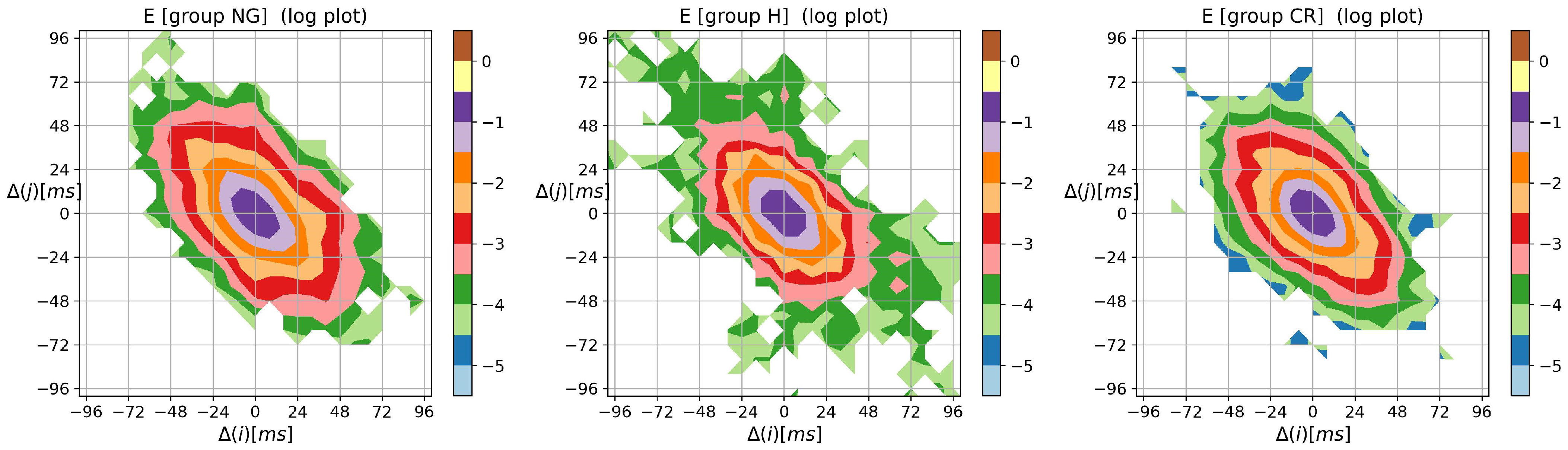
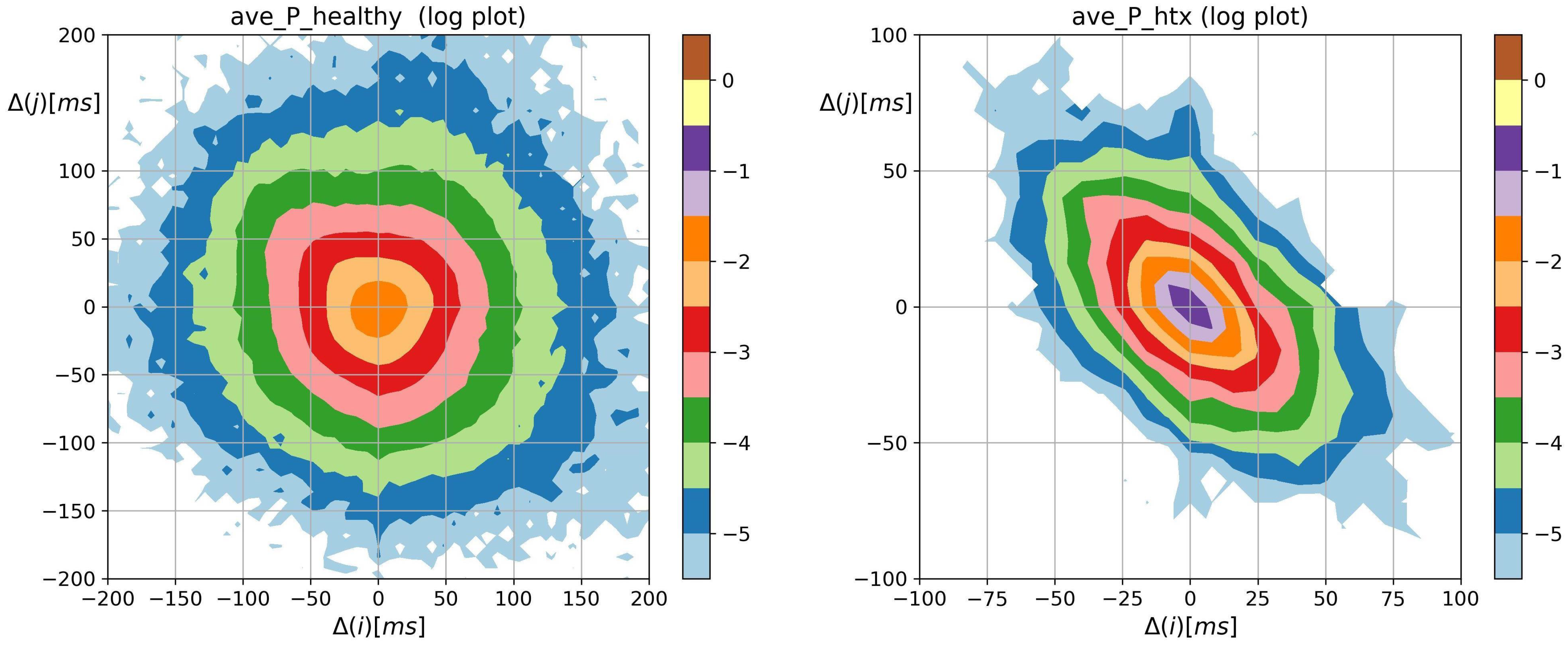
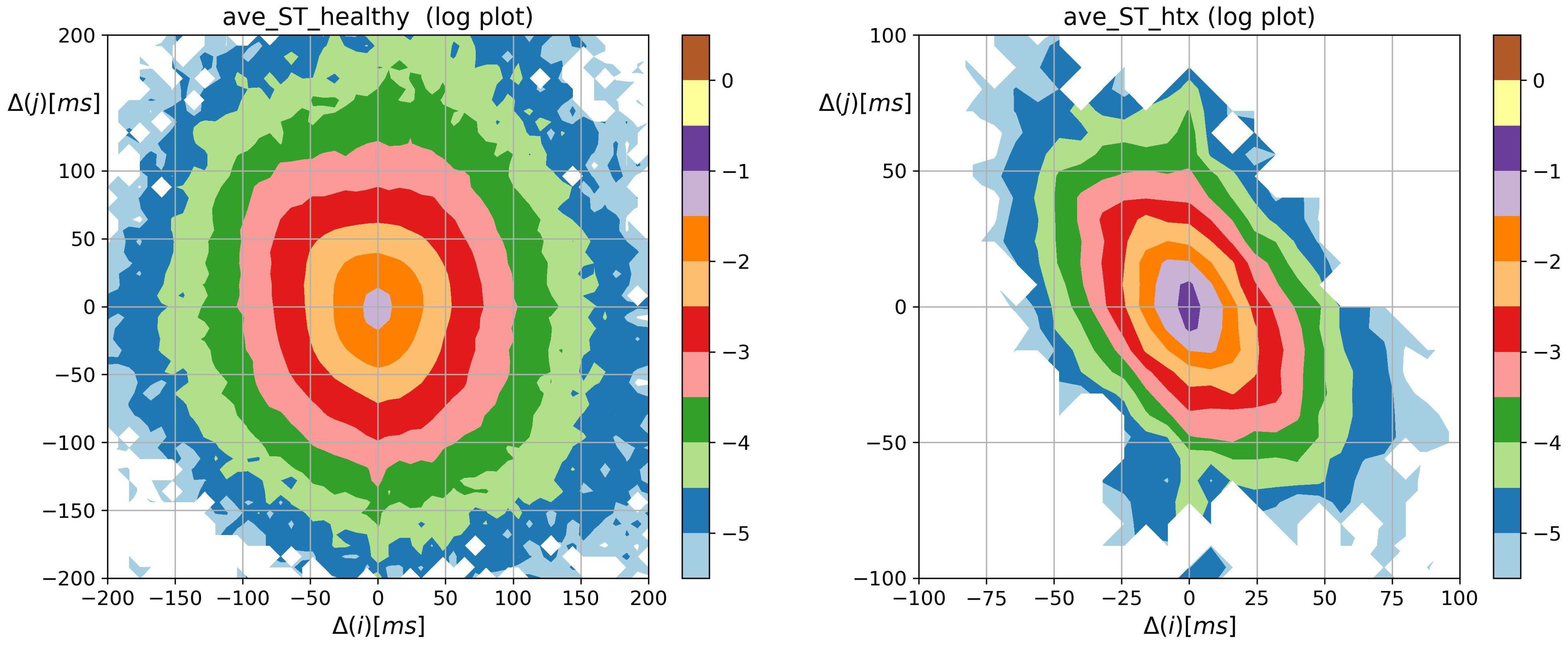
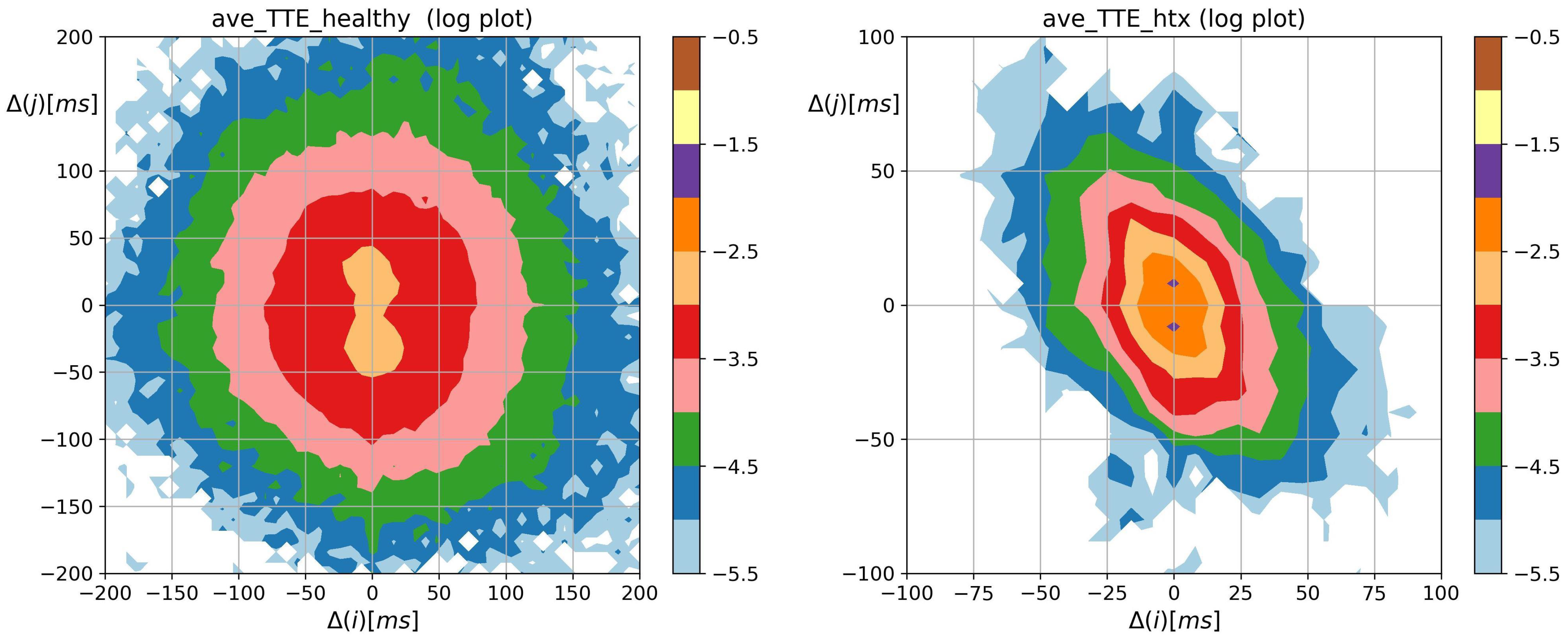
3.2. Visualization of HRV by Matrices of Dynamical Dependence
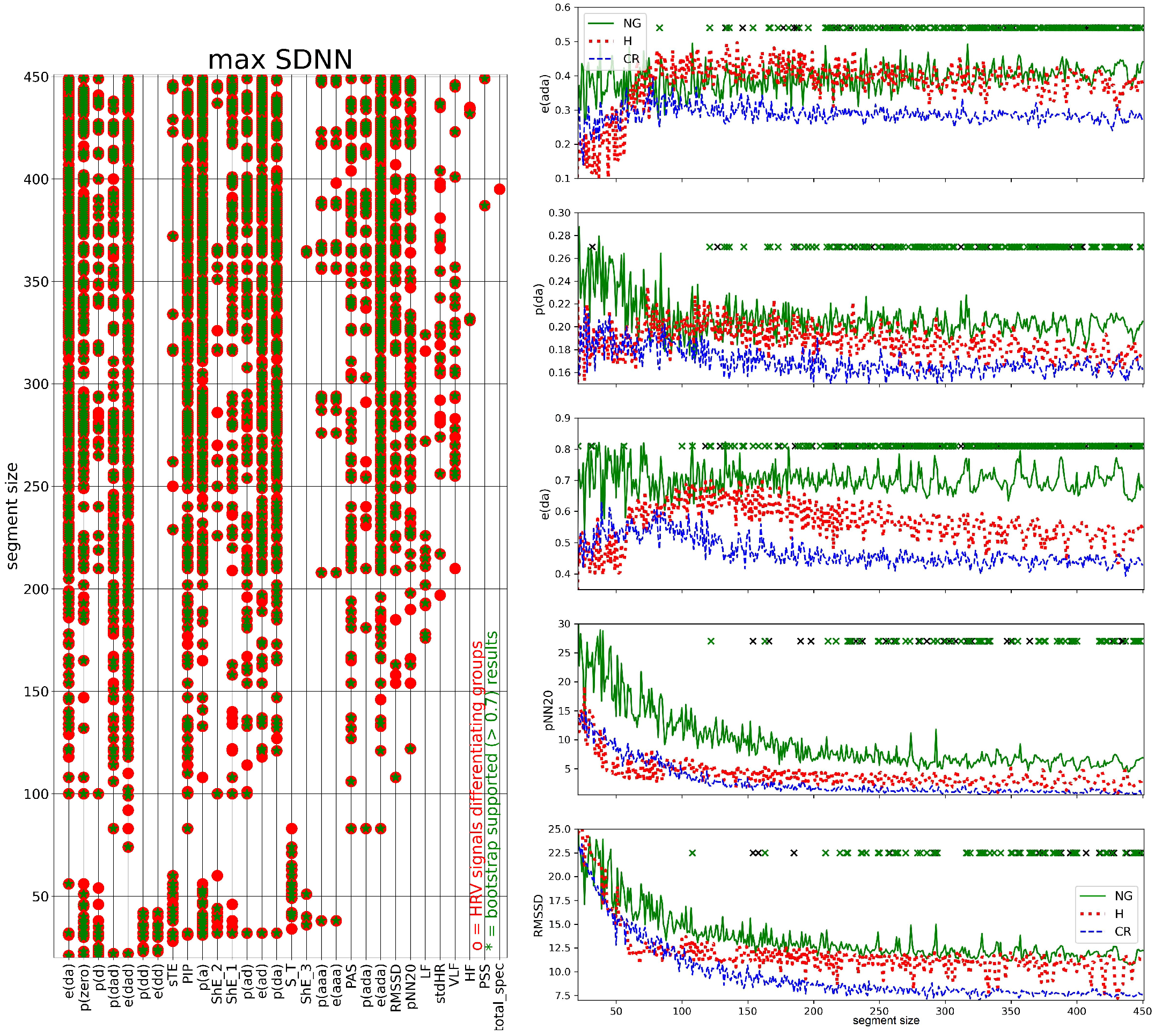
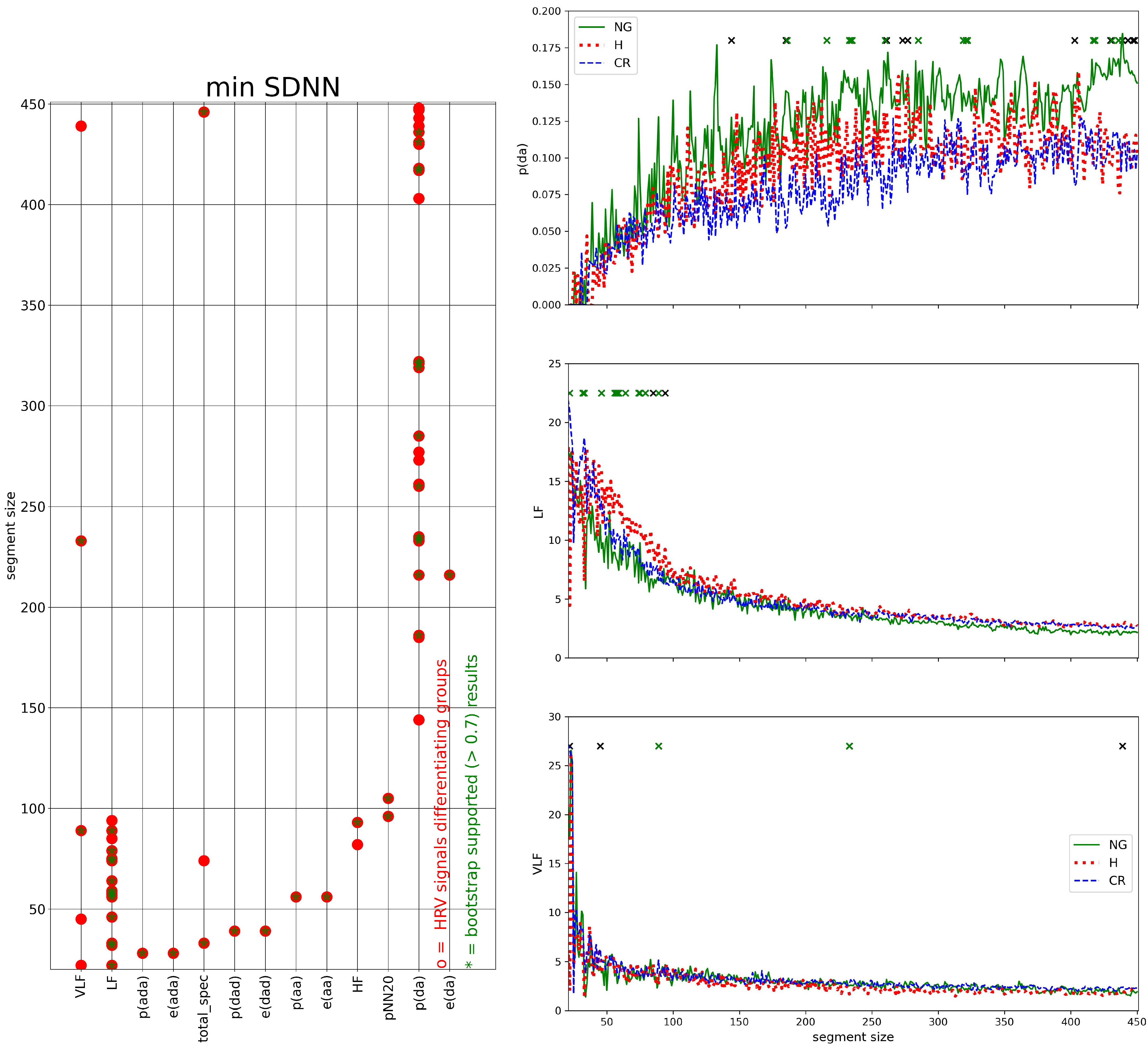
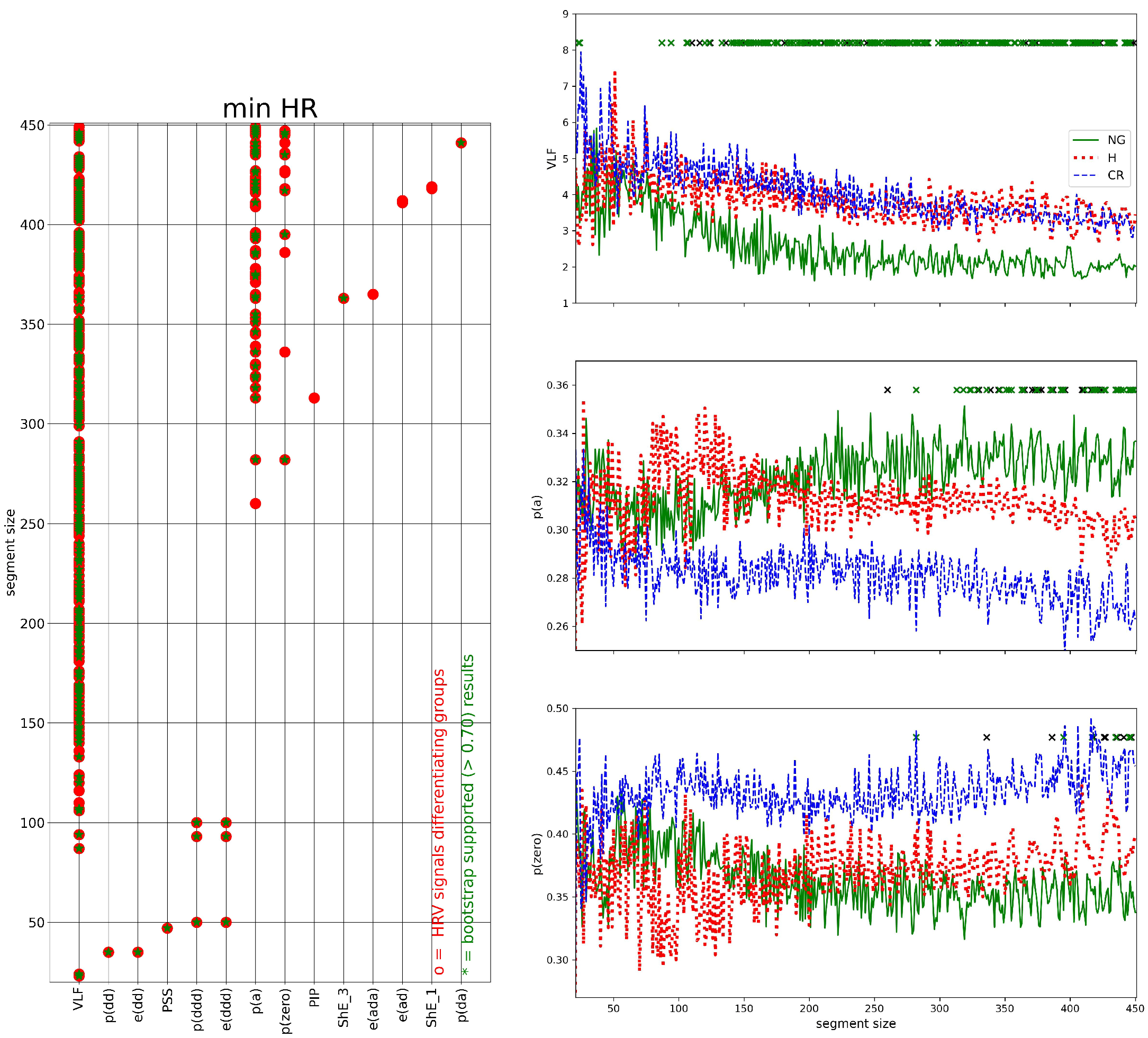
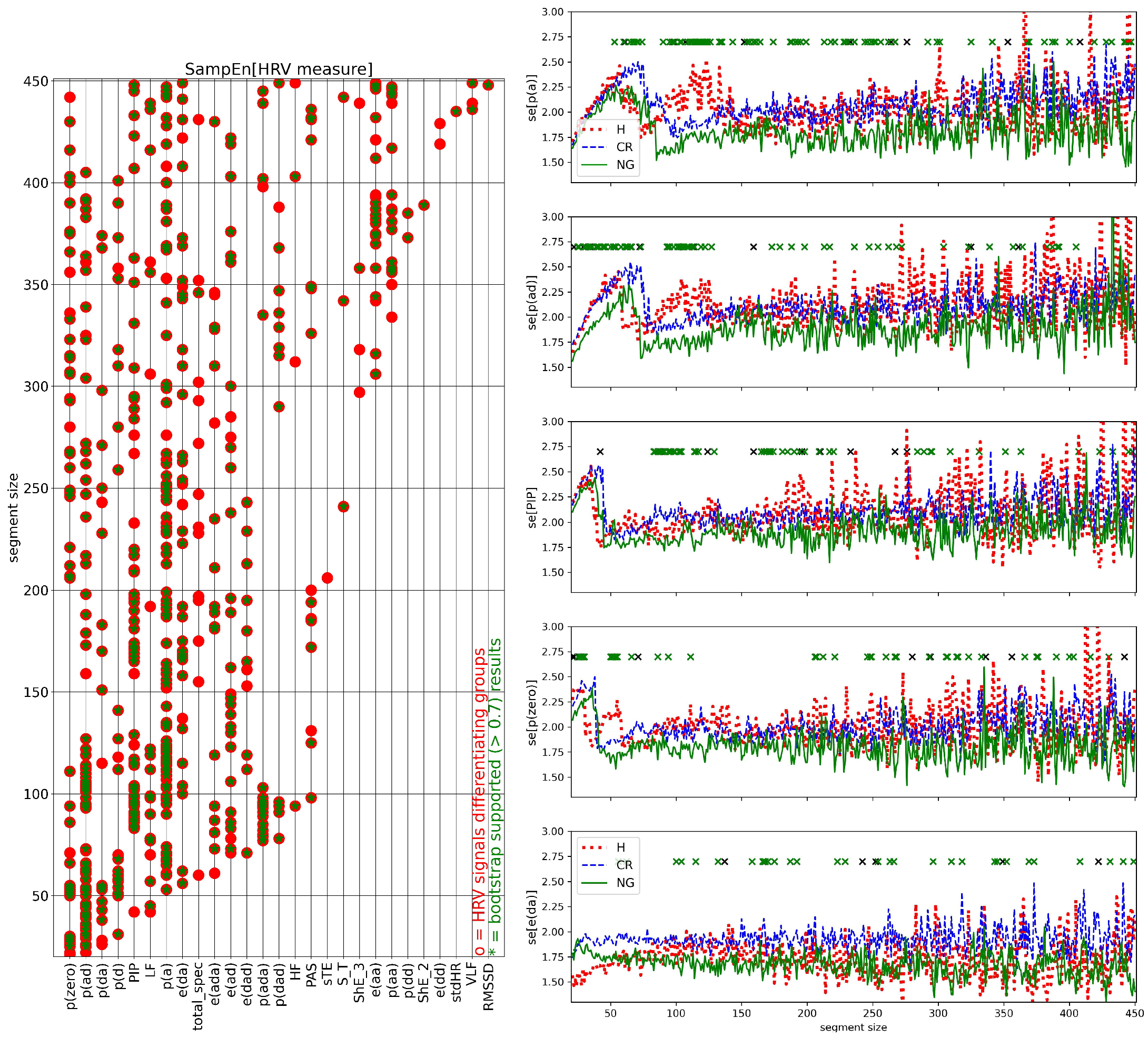
3.3. Segmented Signal HRV Analysis
4. Discussion and Summary
- The probability distribution of HTX patients was strongly steep, and sharply peaked at pattern m where the basic transitions involving accelerations and/or decelerations of magnitude covered more than 90%.
- Accelerations and decelerations were likely to occur alternately, which affected RR intervals (i.e., to change the mean value in a pendulum-type motion rather than as a stochastic walk). Alternating patterns were observed 100 times more frequently than monotonic patterns. This pendulum-type motion was damped in HTX patients.
- Similar to the healthy coevals, the strongest memory effects in patients after HTX were associated with transitions opposite to damped alternating dynamics.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A




References
- Malik, M.; Bigger, J.T.; Camm, A.J.; Kleiger, R.E.; Malliani, A.; Moss, A.J.; Schwartz, P.J. Heart rate variability: Standards of measurement, physiological interpretation, and clinical use. Eur. Heart J. 1996, 17, 354–381. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Stein, P.K. Evaluation of Heart Rate Variability. 2020. Available online: http://www.uptodate.com/contents/evaluation-of-heart-rate-variability (accessed on 29 December 2022).
- Shaffer, F.; Ginberg, J. An Overview of Heart Rate Variability Metrics and Norms. Front. Public Health 2017, 5, 258. [Google Scholar] [CrossRef] [PubMed]
- Saul, J.P.; Valenza, G. Heart rate variability and the dawn of complex physiological signal analysis: Methodological and clinical perspectives. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2021, 379, 20200255. [Google Scholar] [CrossRef] [PubMed]
- Pikkujämsä, S.M.; Mäkikallio, T.H.; Sourander, L.B.; Räihä, I.J.; Puukka, P.; Skyttä, J.; Peng, C.K.; Goldberger, A.L.; Huikuri, H.V. Cardiac interbeat interval dynamics from childhood to senescence: Comparison of conventional and new measures based on fractals and chaos theory. Circulation 1999, 100, 393–399. [Google Scholar] [CrossRef]
- Meersman, R.E.D.; Stein, P.K. Vagal modulation and aging. Biol. Psychol. 2007, 74, 165–173. [Google Scholar] [CrossRef]
- Tsuji, H.; Larson, M.G.; Venditti, F.J.; Manders, E.S.; Evans, J.C.; Feldman, C.L.; Levy, D. Impact of Reduced Heart Rate Variability on Risk for Cardiac Events. Circulation 1996, 94, 2850–2855. [Google Scholar] [CrossRef]
- Nolan, J.; Batin, P.D.; Andrews, R.; Lindsay, S.J.; Brooksby, P.; Mullen, M.; Baig, W.; Flapan, A.D.; Cowley, A.; Prescott, R.J.; et al. Prospective Study of Heart Rate Variability and Mortality in Chronic Heart Failure. Circulation 1998, 98, 1510–1516. [Google Scholar] [CrossRef]
- Huikuri, H.V.; Stein, P.K. Heart Rate Variability in Risk Stratification of Cardiac Patients. Prog. Cardiovasc. Dis. 2013, 56, 153–159. [Google Scholar] [CrossRef]
- Alter, P.; Grimm, W.; Vollrath, A.; Czerny, F.; Maisch, B. Heart rate variability in patients with cardiac hypertrophy—Relation to left ventricular mass and etiology. Am. Heart J. 2006, 151, 829–836. [Google Scholar] [CrossRef]
- Vuoti, A.O.; Tulppo, M.P.; Ukkola, O.H.; Junttila, M.J.; Huikuri, H.V.; Kiviniemi, A.M.; Perkiömäki, J.S. Prognostic value of heart rate variability in patients with coronary artery disease in the current treatment era. PLoS ONE 2021, 16, e0254107. [Google Scholar] [CrossRef]
- Thayer, J.F.; Yamamoto, S.S.; Brosschot, J.F. The relationship of autonomic imbalance, heart rate variability and cardiovascular disease risk factors. Int. J. Cardiol. 2010, 141, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Tracy, E.; Rowe, G.; LeBlanc, A.J. Cardiac tissue remodeling in healthy aging: The road to pathology. Am. J.-Physiol.-Cell Physiol. 2020, 319, C166–C182. [Google Scholar] [CrossRef] [PubMed]
- Frantz, S.; Hundertmark, M.J.; Schulz-Menger, J.; Bengel, F.M.; Bauersachs, J. Left ventricular remodelling post-myocardial infarction: Pathophysiology, imaging, and novel therapies. Eur. Heart J. 2022, 43, 2549–2561. [Google Scholar] [CrossRef] [PubMed]
- Stein, P.K.; Le, Q.; Domitrovich, P.P. Development of more erratic heart rate patterns is associated with mortality post-myocardial infarction. J. Electrocardiol. 2008, 41, 110–115. [Google Scholar] [CrossRef][Green Version]
- Lang, R.M.; Badano, L.P.; Mor-Avi, V.; Afilalo, J.; Armstrong, A.; Ernande, L.; Flachskampf, F.A.; Foster, E.; Goldstein, S.A.; Kuznetsova, T.; et al. Recommendations for Cardiac Chamber Quantification by Echocardiography in Adults: An Update from the American Society of Echocardiography and the European Association of Cardiovascular Imaging. Eur. Heart J.-Cardiovasc. Imaging 2015, 16, 233–271. [Google Scholar] [CrossRef] [PubMed]
- Coelho-Filho, O.R.; Shah, R.; Lavagnoli, C.F.R.; Barros, J.C.; Neilan, T.G.; Murthy, V.L.; de Oliveira, P.P.M.; Souza, J.R.M.; de Oliveira Severino, E.S.B.; de Souza Vilarinho, K.A.; et al. Myocardial tissue remodeling after orthotopic heart transplantation: A pilot cardiac magnetic resonance study. Int. J. Cardiovasc. Imaging 2018, 34, 15–24. [Google Scholar] [CrossRef] [PubMed]
- Liebo, M.; Newman, J.; Joshi, A.; Lowes, B.; Peled-Potashnik, Y.; Basha, H.; Zolty, R.; Um, J.; McGee, E.J.; Heroux, A.; et al. Elevated Heart Rate Following Heart Transplantation Is Associated With Increased Graft Vasculopathy and Mortality. J. Card. Fail. 2019, 25, 249–256. [Google Scholar] [CrossRef]
- Masarone, D.; Kittleson, M.; Gravino, R.; Valente, F.; Petraio, A.; Pacileo, G. The Role of Echocardiography in the Management of Heart Transplant Recipients. Diagnostics 2021, 11, 2338. [Google Scholar] [CrossRef]
- Olymbios, M.; Kwiecinski, J.; Berman, D.S.; Kobashigawa, J.A. Imaging in Heart Transplant Patients. JACC Cardiovasc. Imaging 2018, 11, 1514–1530. [Google Scholar] [CrossRef]
- Billman, G.E.; Huikuri, H.V.; Sacha, J.; Trimmel, K. An introduction to heart rate variability: Methodological considerations and clinical applications. Front. Physiol. 2015, 6, 55. [Google Scholar] [CrossRef]
- Wdowczyk, J.; Makowiec, D.; Gruchała, M.; Wejer, D.; Struzik, Z.R. Dynamical Landscape of Heart Rhythm in Long-Term Heart Transplant Recipients: A Way to Discern Erratic Rhythms. Front. Physiol. 2018, 9, 274. [Google Scholar] [CrossRef] [PubMed]
- Kleiger, R.E.; Stein, P.K.; Bigger, J.T. Heart Rate Variability: Measurement and Clinical Utility. Ann. Noninvasive Electrocardiol. 2005, 10, 88–101. [Google Scholar] [CrossRef] [PubMed]
- Soares-Miranda, L.; Sattelmair, J.; Chaves, P.; Duncan, G.E.; Siscovick, D.S.; Stein, P.K.; Mozaffarian, D. Physical activity and heart rate variability in older adults: The cardiovascular health study. Circulation 2014, 129, 2100–2110. [Google Scholar] [CrossRef]
- Stein, P.K.; Yanez, D.; Domitrovich, P.P.; Gottdiener, J.; Chaves, P.; Kronmal, R.; Rautaharju, P. Heart rate variability is confounded by the presence of erratic sinus rhythm. In Proceedings of the Computers in Cardiology 2002, Memphis, TN, USA, 22–25 September 2002; pp. 669–672. [Google Scholar] [CrossRef]
- Costa, M.; Goldberger, A.L.; Peng, C.K. Multiscale entropy analysis of biological signals. Phys. Rev. E 2005, 71, 021906. [Google Scholar] [CrossRef] [PubMed]
- Nicolini, P.; Ciula, M.M.; de Asmundus, C.; Magrini, F.; Brugada, P. The prognostic value of heart rate variability in the elderly, changing the perspective: From sympathovagal balance to chaos theory. PACE 2012, 35, 622–638. [Google Scholar] [CrossRef]
- Lerma, C.; Krogh-Madsen, T.; Guevara, M.; Glass, L. Stochastic Aspects of Cardiac Arrhythmias. J. Stat. Phys. 2007, 128, 347–374. [Google Scholar] [CrossRef]
- Glass, L. Dynamical disease: Challenges for nonlinear dynamics and medicine. Chaos 2015, 25, 097603. [Google Scholar] [CrossRef]
- Sassi, R.; Cerutti, S.; Lombardi, F.; Malik, M.; Huikuri, H.V.; Peng, C.K.; Schmidt, G.; Yamamoto, Y.; Gorenek, B.; Lip, G.H.; et al. Advances in heart rate variability signal analysis: Joint position statement by the e-Cardiology ESC Working Group and the European Heart Rhythm Association co-endorsed by the Asia Pacific Heart Rhythm Society. Europace 2015, 17, 1341–1353. [Google Scholar] [CrossRef]
- Spellenberg, C.; Heusser, P.; Büssing, A.; Savelsbergh, A.; Cysarz, D. Binary symbolic dynamics analysis to detect stress-associated changes of nonstationary heart rate variability. Sci. Rep. 2020, 10, 15440. [Google Scholar] [CrossRef] [PubMed]
- Cysarz, D.; Bettermann, H.; van Leeuwen, P. Entropies of short binary sequences in heart period dynamics. Am. J.-Physiol.-Heart Circ. Physiol. 2000, 278, H2163–H2172. [Google Scholar] [CrossRef]
- Boccaletti, S.; Latora, V.; Moreno, Y.; Chavez, M.; Hwang, D.U. Complex networks: Structure and dynamics. Phys. Rep. 2006, 424, 175–308. [Google Scholar] [CrossRef]
- Porta, A.; Faes, L.; Masé, M.; D’Addio, G.; Pinna, G.D.; Maestri, R.; Montano, N.; Furlan, R.; Guzzetti, S.; Nollo, G.; et al. An integrated approach based on uniform quantization for the evaluation of complexity of short-term heart period variability: Application to 24h Holter recordings in healthy and heart failure humans. Chaos Interdiscip. J. Nonlinear Sci. 2007, 17, 015117. [Google Scholar] [CrossRef] [PubMed]
- Alcaraz, R. Symbolic Entropy Analysis and Its Applications. Entropy 2018, 20, 568. [Google Scholar] [CrossRef] [PubMed]
- Cysarz, D.; Van Leeuwen, P.; Edelhäuser, F.; Montano, N.; Porta, A. Binary symbolic dynamics classifies heart rate variability patterns linked to autonomic modulations. Comput. Biol. Med. 2012, 42, 313–318. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.; Davis, R.; Goldberger, A. Heart Rate Fragmentation: A New Approach to the Analysis of Cardiac Interbeat Interval Dynamics. Front. Physiol. 2017, 8, 255. [Google Scholar] [CrossRef]
- Makowiec, D.; Kaczkowska, A.; Wejer, D.; Żarczyńska-Buchowiecka, M.; Struzik, Z.R. Entropic measures of complexity of short-term dynamics of nocturnal heartbeats in an aging population. Entropy 2015, 17, 1253–1272. [Google Scholar] [CrossRef]
- Makowiec, D.; Graff, B.; Kaczkowska, A.; Graff, G.; Wejer, D.; Wdowczyk, J.; Zarczynska-Buchowiecka, M.; Gruchala, M.; Struzik, Z.R. Chapter 6—Visualization of short-term heart period variability with network tools as a method for quantifying autonomic drive. In ECG Time Series Variability Analysis. Engineering and Medicine, 1st ed.; Jedlinek, H.F., Cornforth, D.J., Khandoker, A.H., Eds.; CRC Press, Taylor & Francis Group: Boca Raton, FL, USA, 2018; pp. 141–157. [Google Scholar]
- Makowiec, D.; Wdowczyk, J.; Gruchała, M.; Struzik, Z.R. Network tools for tracing the dynamics of heart rate after cardiac transplantation. Chaos Solitons Fractals 2016, 90, 101–110. [Google Scholar] [CrossRef]
- Zhang, H.; Zhu, M.; Zheng, Y.; Li, G. Toward Capturing Momentary Changes of Heart Rate Variability by a Dynamic Analysis Method. PLoS ONE 2015, 10, e133148. [Google Scholar] [CrossRef]
- Bonnet, M.; Arand, D. Heart rate variability: Sleep stage, time of night, and arousal influences. Electroencephalogr. Clin. Neurophysiol. 1997, 102, 390–396. [Google Scholar] [CrossRef]
- Bartsch, R.P.; Kantelhardt, J.W.; Penzel, T.; Havlin, S. Experimental evidence for phase synchronization transitions in the human cardiorespiratory system. Phys. Rev. Lett. 2007, 98, 054102. [Google Scholar] [CrossRef]
- Viola, A.U.; Tobaldini, E.; Chellappa, S.L.; Casali, K.R.; Porta, A.; Montano, N. Short-term complexity of cardiac autonomic control during sleep: REM as a potential risk factor for cardiovascular system in aging. PLoS ONE 2011, 6, e19002. [Google Scholar] [CrossRef] [PubMed]
- Reinertsen, E.; Nemati, S.; Vest, A.N.; Vaccarino, V.; Lampert, R.; Shah, A.J.; Clifford, G.D. Heart rate-based window segmentation improves accuracy of classifying posttraumatic stress disorder using heart rate variability measures. Physiol. Meas. 2017, 38, 1061–1076. [Google Scholar] [CrossRef] [PubMed]
- Makowiec, D.; Kaczkowska, A.; Wejer, D.; Zarczynska-Buchowiecka, M.; Struzik, Z.R. Chronographic imprint of age-induced alterations in heart rate dynamical organization. Front. Physiol. 2015, 6, 201. [Google Scholar] [CrossRef] [PubMed]
- Porta, A.; Guzzetti, S.; Montano, N.; Furlan, R.; Pagani, M.; Malliani, A.; Cerutti, S. Entropy, entropy rate, and pattern classification as tools to typify complexity in short heart period variability series. IEEE Trans. Biomed. Eng. 2001, 48, 1282–1291. [Google Scholar] [CrossRef]
- Cysarz, D.; Porta, A.; Montano, N.; Leeuwen, P.; Kurths, J.; Wessel, N. Quantifying heart rate dynamics using different approaches of symbolic dynamics. Eur. Phys. J. Spec. Top. 2013, 222, 487–500. [Google Scholar] [CrossRef]
- Pincus, S.M. Approximate entropy as a measure of system complexity. Proc. Natl. Acad. Sci. USA 1991, 88, 2297–2301. [Google Scholar] [CrossRef]
- Hlavackova-Schindler, K.; Palus, M.; Vejmelka, M.; Bhattacharya, J. Causality detection based on information-theoretic approaches in time series analysis. Phys. Rep. 2007, 441, 1–46. [Google Scholar] [CrossRef]
- Shannon, C.E. A Mathematical Theory of Communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Kaiser, A.; Schreiber, T. Information transfer in continuous processes. Phys. D Nonlinear Phenom. 2002, 166, 43–62. [Google Scholar] [CrossRef]
- Delgado-Bonal, A.; Marshak, A. Approximate Entropy and Sample Entropy: A Comprehensive Tutorial. Entropy 2019, 21, 541. [Google Scholar] [CrossRef]
- Richman, J.S.; Moorman, J.R. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol.-Heart Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [CrossRef] [PubMed]
- Ciuperca, G.; Girardin, V. On the estimation of the entropy rate of finite Markov chains. In Proceedings of the Applied Stochastic Modes and Data Analysis (ASMDA2005), Brest, France, 17–20 May 2005; pp. 1109–1117. [Google Scholar]
- Schreiber, T. Measuring Information Transfer. Phys. Rev. Lett. 2000, 85, 461–464. [Google Scholar] [CrossRef] [PubMed]
- Makowiec, D.; Wdowczyk, J. Patterns of Heart Rate Dynamics in Healthy Aging Population: Insights from Machine Learning Methods. Entropy 2019, 21, 1206. [Google Scholar] [CrossRef]
- Donner, R.V.; Zou, Y.; Donges, J.F.; Marwan, N.; Kurths, J. Recurrence networks—A novel paradigm for nonlinear time series analysis. New J. Phys. 2010, 12, 033025. [Google Scholar] [CrossRef]
- Piskorski, J.; Guzik, P. Geometry of Poincare plot of RR intervals and its asymmetry in healthy adults. Physiol. Meas. 2007, 28, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Henriques, T.S.; Mariani, S.; Burykin, A.; Rodrigues, F.; Silva, T.F.; Goldberger, A.L. Multiscale Poincaré plots for visualizing the structure of heartbeat time series. BMC Med. Inform. Decis. Mak. 2016, 16, 17. [Google Scholar] [CrossRef]
- Costanzo, M.R.; Costanzo, M.R.; Dipchand, A.; Starling, R.; Anderson, A.; Chan, M.; Desai, S.; Fedson, S.; Fisher, P.; Gonzales-Stawinski, G.; et al. The International Society of Heart and Lung Transplantation Guidelines for the care of heart transplant recipients. J. Heart Lung Transplant. 2010, 29, 914–956. [Google Scholar] [CrossRef]
- Badano, L.P.; Miglioranza, M.H.; Edvardsen, T.; Colafranceschi, A.S.; Muraru, D.; Bacal, F.; Nieman, K.; Zoppellaro, G.; Braga, F.G.M.; Binder, T.; et al. European Association of Cardiovascular Imaging/Cardiovascular Imaging Department of the Brazilian Society of Cardiology recommendations for the use of cardiac imaging to assess and follow patients after heart transplantation. Eur. Heart J. Cardiovasc. Imaging 2015, 16, 919–948. [Google Scholar] [CrossRef]
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; Haberland, M.; Reddy, T.; Cournapeau, D.; Burovski, E.; Peterson, P.; Weckesser, W.; Bright, J.; et al. SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python. Nat. Methods 2020, 17, 261–272. [Google Scholar] [CrossRef]
- Wes McKinney. Data Structures for Statistical Computing in Python. In Proceedings of the 9th Python in Science Conference, Austin, TX, USA, 28 June–3 July 2010; van der Walt, S., Millman, J., Eds.; pp. 56–61. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Stein, P.K.; Pu, Y. Heart rate variability, sleep and sleep disorders. Sleep Med. Rev. 2012, 16, 47–66. [Google Scholar] [CrossRef] [PubMed]
- Burchfield, J.S.; Xie, M.; Hill, J.A. Pathological Ventricular Remodeling. Circulation 2013, 128, 388–400. [Google Scholar] [CrossRef] [PubMed]
- Chouchou, F.; Desseilles, M. Heart rate variability: A tool to explore the sleeping brain? Front. Neurosci. 2014, 8, 402. [Google Scholar] [CrossRef] [PubMed]
- Guyton, A.C.; Hall, J.E. Textbook of Medical Physiology; Elsevier Saunders Company: Philadelphia, PA, USA, 2006. [Google Scholar]
- Monti, A.; Medigue, C.; Nedelcoux, H.; Escourrou, P. Autonomic control of the cardiovascular system during sleep in normal subjects. Eur. J. Appl. Physiol. 2002, 87, 174–181. [Google Scholar]
- Tobaldini, E.; Nobili, L.; Strada, S.; Casali, K.R.; Braghiroli, A.; Montano, N. Heart rate variability in normal and pathological sleep. Front. Physiol. 2013, 4, 294. [Google Scholar] [CrossRef]
- de Boer, R.; Karemaker, J.; Strackee, J. Hemodynamic fluctuations and baroreflex sensitivity in humans: A beat-to-beat model. Am. J. Physiol. 1987, 253, H680–H689. [Google Scholar]
- Dudkowska, A.; Makowiec, D. Seidel-Herzel model of human baroreflex in cardiorespiratory system with stochastic delays. J. Math. Biol. 2008, 57, 111–137. [Google Scholar] [CrossRef]
- Karemaker, J.M. How the vagus nerve produces beat-to-beat heart rate variability; experiments in rabbits to mimic in vivo vagal patterns. J. Clin. Transl. Res. 2015, 1, 190–204. [Google Scholar] [CrossRef]




| Characteristic | NG | CR | H | Difference between Groups |
|---|---|---|---|---|
| number of patients | 12 (29%) | 22 (52%) | 8 (19%) | |
| age at transplantation, | 43 ± 14 | 49 ± 11 | 48 ± 11 | NS |
| male gender, n | 10 (83%) | 20 (91%) | 8 (88%) | |
| BMI | 26± 4 | 26 ± 6 | 27 ± 5 | NS |
| BSA | 1.9 ± 0.1 | 2.0 ± 0.2 | 2.0 ± 0.2 | NS |
| LVMI (g/m) | 82 ± 10 | 92 ± 14 | 122 ± 9 | (NG,H) (CR,H) |
| LVM (g) | 156 ± 25 | 184 ± 28 | 245 ± 33 | all pairs are different |
| RWT | 0.39 ± 0.03 | 0.53 ± 0.07 | 0.50 ± 0.07 | (NG,H) (NG,CR) |
| EF % | 64 ± 3 | 64 ± 4 | 67 ± 10 | NS |
| LV SV (mL) | 66 ± 7 | 60 ± 11 | 76 ± 16 | NS |
| standard measures: | long–term time domain | meanRR, meanHR, |
| based on a series of | SDNN, stdHR | |
| {} | ||
| frequency | PS, VLF, LF, HF | |
| short-term time domain | RMSSD, pNN50, pNN20 | |
| increment pattern measures: | probability of patterns | p(zero), p(a), p(d), |
| based on series of increments | p(aa), p(ad), p(da), p(dd), | |
| between consecutive RR intervals: | p(aaa), p(ada), p(dad), p(ddd) | |
| { } | ||
| a is an acceleration if ms | fragmentation measures | PIP = p(ad)+p(da) |
| d is a deceleration if ms. | PAS = p(ada) + p(dad) | |
| Otherwise an increment is zero | PSS = 1 – [p(aaa) +p(ddd)] | |
| entropic measures: | ||
| –Shannon entropy of L-length patterns | ShE_L for L=1,2,3 | |
| –dynamics by entropy patterns, | S_T = ShE_1 - ShE_2, | |
| sTE = ShE_2 - ShE_3 – S_T | ||
| –partial entropy of patterns | e(aa), e(ad), e(da), e(dd), | |
| e(aaa), e(ada), e(dad), e(ddd) |
| HRV | Healthy | HTX Groups | Kruskal–Wallis | ||
|---|---|---|---|---|---|
| Index | Coveals | NG | CR | H | Test p for HTX |
| meanRR | 958 [870, 1067] | 697 [679, 819] | 688 [635, 791] | 714 [683, 757] | p = 0.587 |
| meanHR | 62.85 [56.95, 69.80] | 86.23 [73.35, 88.39] | 87.62 [75.88, 94.72] | 84.16 [79.36, 88.04] | p = 0.542 |
| SDNN | 87.10 [72.18, 102.30] | 22.17 [20.10, 31.66] | 28.44 [20.77, 35.89] | 31.15 [25.65, 35.17] | p = 0.415 |
| stdHR | 6.577 [5.849, 7.253] | 2.853 [1.953, 3.330] | 3.807 [2.322, 4.162] | 3.266 [3.193, 4.185] | p = 0.128 |
| PS | 3.384 [3.033, 3.755] | 2.432 [2.114, 2.916] | 2.558 [2.373, 2.671] | 2.569 [2.358, 2.750] | p = 0.938 |
| VLF | 1.392 [1.308, 1.507] | 1.112 [0.929, 1.167] | 1.180 [1.054, 1.323] | 1.110 [1.066, 1.200] | p = 0.338 |
| LF | 1.000 [0.862, 1.281] | 0.265 [0.224, 0.366] | 0.297 [0.244, 0.369] | 0.302 [0.257, 0.399] | p = 0.603 |
| HF | 0.868 [0.668, 1.051] | 1.005 [0.708, 1.235] | 0.958 [0.786, 1.192] | 1.035 [0.855, 1.326] | p = 0.859 |
| RMSSD | 32.12 [22.85, 46.87] | 9.879 [7.308, 11.29] | 7.189 [6.593, 9.669] | 9.575 [7.012, 12.24] | p = 0.328 |
| pNN50 | 6.215 [2.435, 18.76] | 0.025 [0.004, 0.059] | 0.015 [0.005, 0.025] | 0.040 [0.020, 0.185] | p = 0.099 (0.05) |
| pNN20 | 39.68 [29.89, 55.69] | 3.815 [2.001, 5.566] | 0.583 [0.184, 3.109] | 3.908 [0.320, 4.885] | p = 0.244 |
| p(zero) | 0.145 [0.101, 0.181] | 0.358 [0.317, 0.456] | 0.448 [0.405, 0.469] | 0.430 [0.352, 0.489] | p = 0.289 |
| p(a) | 0.441 [0.415, 0.465] | 0.325 [0.274, 0.353] | 0.273 [0.265, 0.296] | 0.294 [0.257, 0.318] | p = 0.174 |
| p(d) | 0.423 [0.398, 0.446] | 0.307 [0.272, 0.340] | 0.276 [0.264, 0.301] | 0.276 [0.257, 0.323] | p = 0.415 |
| PSS | 0.883 [0.853, 0.919] | 1.000 [0.996, 1.000] | 1.000 [0.998, 1.000] | 0.999 [0.997, 1.000] | p = 0.862 |
| p(aa) | 0.183 [0.165, 0.207] | 0.028 [0.007, 0.049] | 0.017 [0.009, 0.029] | 0.021 [0.009, 0.038] | p = 0.915 |
| e(aa) | 1.019 [0.859, 1.175] | 0.137 [0.038, 0.232] | 0.077 [0.045, 0.118] | 0.097 [0.043, 0.169] | p = 0.857 |
| p(aaa) | 0.058 [0.039, 0.078] | 0.000 [0.000, 0.003] | 0.000 [0.000, 0.001] | 0.001 [0.000, 0.001] | p = 0.625 |
| e(aaa) | 0.461 [0.312, 0.621] | 0.003 [0.001, 0.020] | 0.002 [0.000, 0.008] | 0.005 [0.000, 0.008] | p = 0.672 |
| p(dd) | 0.176 [0.152, 0.195] | 0.021 [0.005, 0.038] | 0.016 [0.009, 0.028] | 0.012 [0.008, 0.035] | p = 0.992 |
| e(dd) | 1.006 [0.871, 1.127] | 0.109 [0.027, 0.187] | 0.069 [0.046, 0.118] | 0.064 [0.039, 0.157] | p = 0.999 |
| p(ddd) | 0.058 [0.039, 0.069] | 0.000 [0.000, 0.001] | 0.000 [0.000, 0.001] | 0.000 [0.000, 0.002] | p = 0.996 |
| e(ddd) | 0.477 [0.302, 0.574] | 0.002 [0.000, 0.007] | 0.001 [0.000, 0.007] | 0.001 [0.000, 0.013] | p = 0.993 |
| PAS | 0.125 [0.082, 0.189] | 0.165 [0.132, 0.207] | 0.156 [0.136, 0.165] | 0.161 [0.146, 0.170] | p = 0.704 |
| PIP | 0.373 [0.321, 0.448] | 0.341 [0.328, 0.385] | 0.325 [0.313, 0.333] | 0.330 [0.314, 0.369] | p = 0.084 (0.07) |
| p(ad) | 0.188 [0.170, 0.229] | 0.168 [0.154, 0.198] | 0.159 [0.152, 0.169] | 0.165 [0.160, 0.173] | p = 0.418 |
| e(ad) | 1.025 [0.848, 1.309] | 0.557 [0.448, 0.709] | 0.427 [0.387, 0.493] | 0.516 [0.392, 0.611] | p = 0.209 |
| p(da) | 0.185 [0.154, 0.212] | 0.175 [0.171, 0.203] | 0.162 [0.158, 0.175] | 0.164 [0.157, 0.185] | p = 0.045 (0.72) |
| e(da) | 0.938 [0.834, 1.281] | 0.594 [0.462, 0.706] | 0.433 [0.399, 0.593] | 0.528 [0.391, 0.621] | p = 0.225 |
| p(ada) | 0.055 [0.040, 0.092] | 0.086 [0.073, 0.105] | 0.080 [0.068, 0.083] | 0.079 [0.074, 0.087] | p = 0.618 |
| e(ada) | 0.441 [0.338, 0.682] | 0.355 [0.308, 0.549] | 0.279 [0.252, 0.323] | 0.293 [0.257, 0.409] | p = 0.131 (0.04) |
| p(dad) | 0.070 [0.035, 0.100] | 0.077 [0.066, 0.104] | 0.077 [0.069, 0.082] | 0.080 [0.072, 0.086] | p = 0.755 |
| e(dad) | 0.534 [0.282, 0.785] | 0.325 [0.280, 0.513] | 0.273 [0.249, 0.326] | 0.307 [0.267, 0.393] | p = 0.229 |
| ShE_3 | 7.321 [6.915, 8.250] | 4.049 [3.137, 4.546] | 3.388 [3.079, 4.036] | 3.546 [3.016, 4.361] | p = 0.52 |
| ShE_2 | 5.146 [4.792, 5.849] | 2.945 [2.267, 3.123] | 2.374 [2.169, 2.869] | 2.542 [2.147, 3.073] | p = 0.437 |
| ShE_1 | 2.663 [2.421, 3.019] | 1.613 [1.280, 1.637] | 1.279 [1.190, 1.525] | 1.404 [1.179, 1.629] | p = 0.27 |
| S_T | 2.507 [2.352, 2.844] | 1.339 [0.970, 1.511] | 1.099 [0.988, 1.307] | 1.138 [0.970, 1.438] | p = 0.716 |
| sTE | 0.302 [0.204, 0.487] | 0.138 [0.093, 0.205] | 0.097 [0.082, 0.120] | 0.131 [0.108, 0.160] | p = 0.106 (0.02) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Makowiec, D.; Wdowczyk, J.; Gruchała, M. Heart Rate Variability by Dynamical Patterns in Windows of Holter Electrocardiograms: A Method to Discern Left Ventricular Hypertrophy in Heart Transplant Patients Shortly after the Transplant. BioMedInformatics 2023, 3, 220-251. https://doi.org/10.3390/biomedinformatics3010015
Makowiec D, Wdowczyk J, Gruchała M. Heart Rate Variability by Dynamical Patterns in Windows of Holter Electrocardiograms: A Method to Discern Left Ventricular Hypertrophy in Heart Transplant Patients Shortly after the Transplant. BioMedInformatics. 2023; 3(1):220-251. https://doi.org/10.3390/biomedinformatics3010015
Chicago/Turabian StyleMakowiec, Danuta, Joanna Wdowczyk, and Marcin Gruchała. 2023. "Heart Rate Variability by Dynamical Patterns in Windows of Holter Electrocardiograms: A Method to Discern Left Ventricular Hypertrophy in Heart Transplant Patients Shortly after the Transplant" BioMedInformatics 3, no. 1: 220-251. https://doi.org/10.3390/biomedinformatics3010015
APA StyleMakowiec, D., Wdowczyk, J., & Gruchała, M. (2023). Heart Rate Variability by Dynamical Patterns in Windows of Holter Electrocardiograms: A Method to Discern Left Ventricular Hypertrophy in Heart Transplant Patients Shortly after the Transplant. BioMedInformatics, 3(1), 220-251. https://doi.org/10.3390/biomedinformatics3010015






