Interplay Between Membrane Adhesion and Distribution of Lipid Rafts
Abstract
1. Introduction
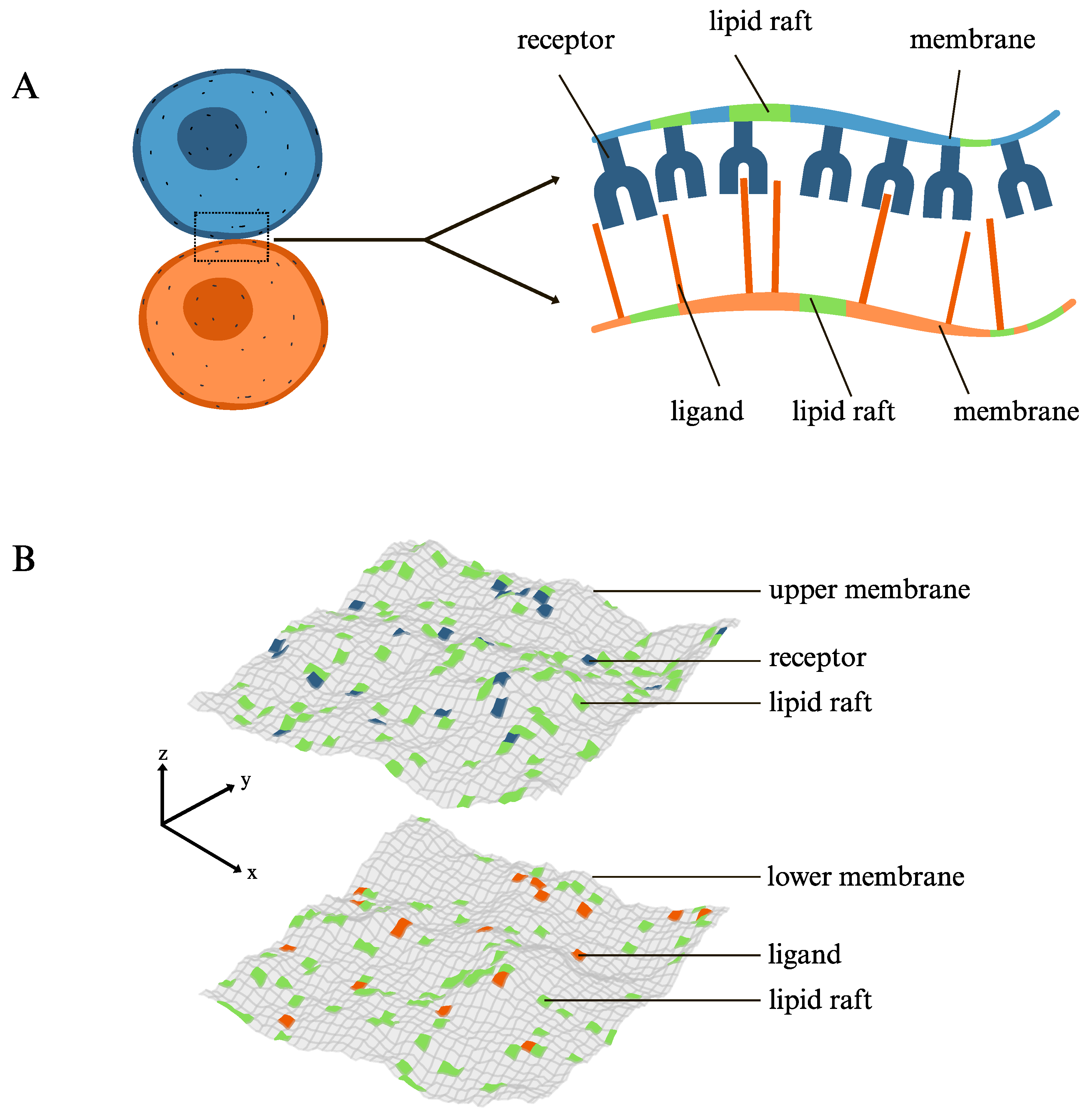
2. Theoretical Model and Simulation Methods
3. Results and Discussion
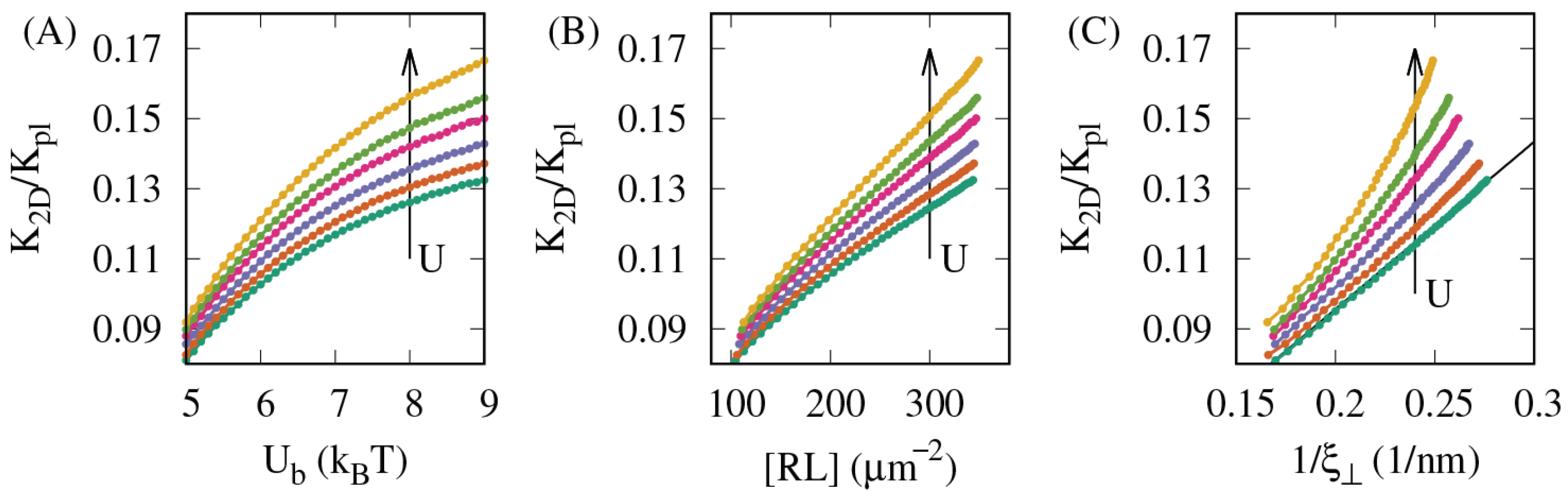
3.1. The Lateral Attraction Between Lipid Rafts Enhances Both the Affinity and the Membrane-Mediated Cooperativity of the Receptor–Ligand Binding
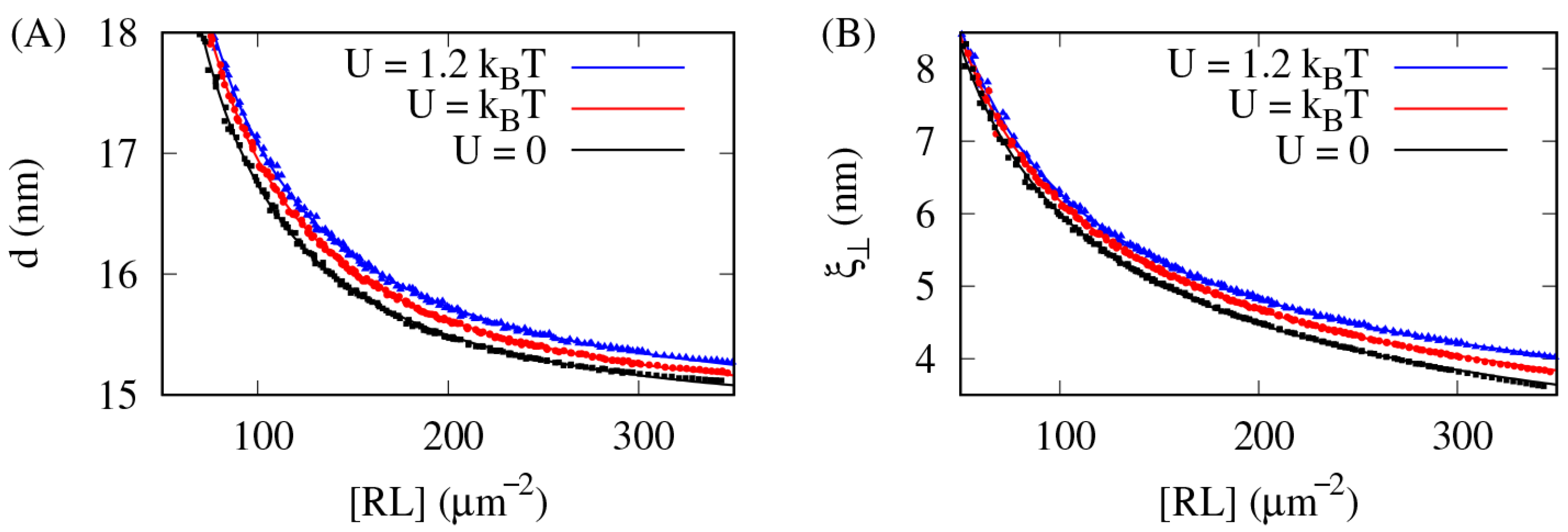
3.2. The Lateral Attraction Between Lipid Rafts Broadens the Inter-Membrane Distance Distribution
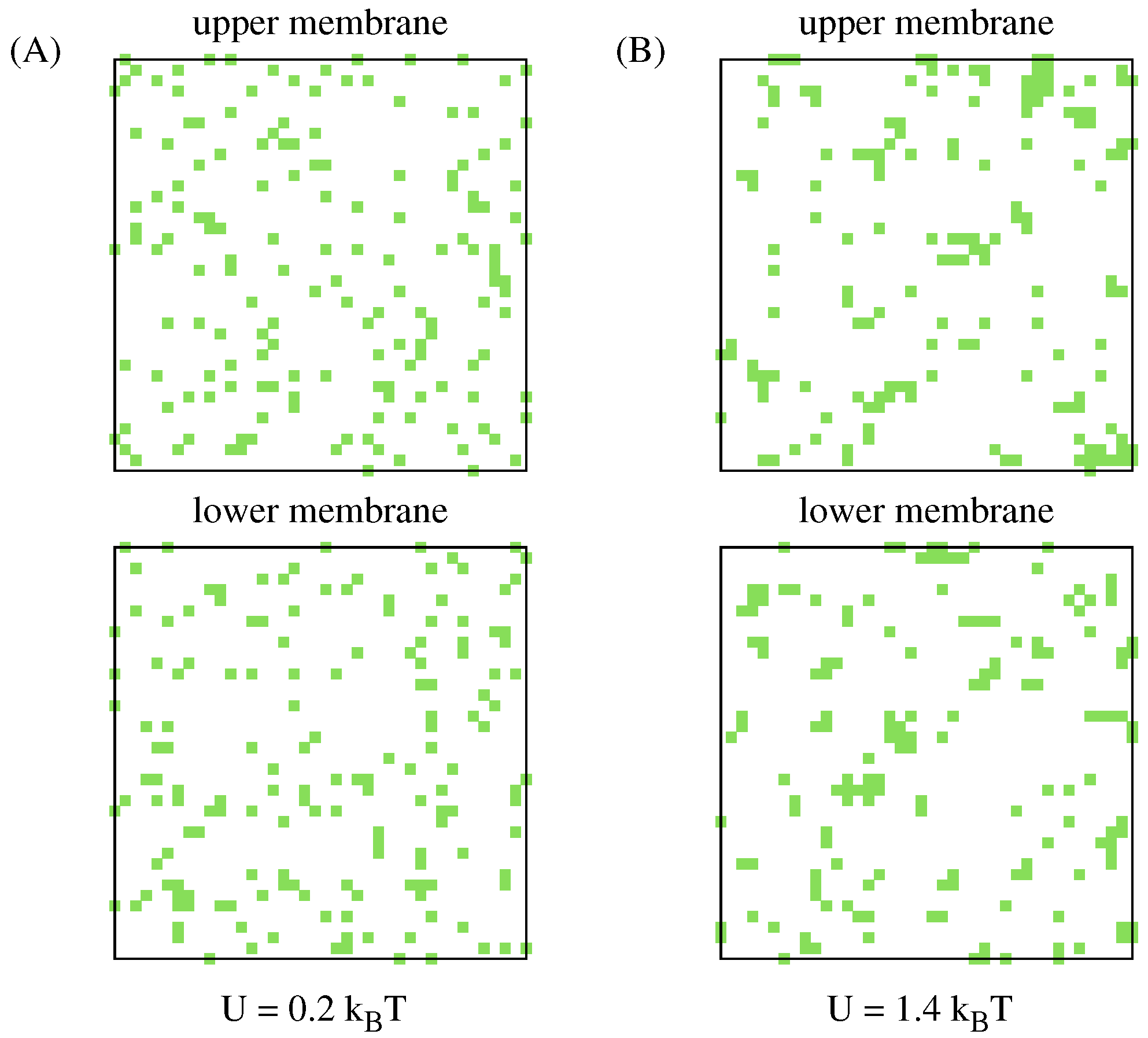
3.3. Formation of the Receptor–Ligand Complexes Causes Lipid Rafts to Be Distributed Less Uniformly Within the Adhered Membranes
3.4. Formation of Receptor–Ligand Complexes Enhances Co-Localization of Adhesion Proteins with Lipid Rafts
4. Summary and Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Correction Statement
References
- Alberts, B.; Bray, D.; Hopkin, K.; Johnson, A.D.; Lewis, J.; Raff, M.; Roberts, K.; Walter, P. Essential Cell Biology; Garland Science: New York, NY, USA, 2015. [Google Scholar]
- Dustin, M.L. The immunological synapse. Cancer Immunol. Res. 2014, 2, 1023–1033. [Google Scholar] [CrossRef]
- Bassereau, P.; Jin, R.; Baumgart, T.; Deserno, M.; Dimova, R.; Frolov, V.A.; Bashkirov, P.V.; Grubmüller, H.; Jahn, R.; Risselada, H.J.; et al. The 2018 biomembrane curvature and remodeling roadmap. J. Phys. D Appl. Phys. 2018, 51, 343001. [Google Scholar] [CrossRef]
- Weikl, T.R. Membrane-mediated cooperativity of proteins. Annu. Rev. Phys. Chem. 2018, 69, 521–539. [Google Scholar] [CrossRef]
- Hurley, J.H.; Boura, E.; Carlson, L.A.; Różycki, B. Membrane budding. Cell 2010, 143, 875–887. [Google Scholar] [CrossRef]
- Lingwood, D.; Simons, K. Lipid rafts as a membrane-organizing principle. Science 2010, 327, 46–50. [Google Scholar] [CrossRef]
- Simons, K.; Sampaio, J.L. Membrane organization and lipid rafts. Cold Spring Harb. Perspect. Biol. 2011, 3, a004697. [Google Scholar] [CrossRef]
- Levental, I.; Veatch, S.L. The continuing mystery of lipid rafts. J. Mol. Biol. 2016, 428, 4749–4764. [Google Scholar] [CrossRef]
- Li, L.; Hu, J.; Różycki, B.; Song, F. Intercellular receptor–ligand binding and thermal fluctuations facilitate receptor aggregation in adhering membranes. Nano Lett. 2020, 20, 722–728. [Google Scholar] [CrossRef]
- Li, L.; Hu, J.; Shi, X.; Różycki, B.; Song, F. Interplay between cooperativity of intercellular receptor–ligand binding and coalescence of nanoscale lipid clusters in adhering membranes. Soft Matter 2021, 17, 1912–1920. [Google Scholar] [CrossRef]
- Hu, J.; Lipowsky, R.; Weikl, T.R. Binding constants of membrane-anchored receptors and ligands depend strongly on the nanoscale roughness of membranes. Proc. Natl. Acad. Sci. USA 2013, 110, 15283–15288. [Google Scholar] [CrossRef]
- Hu, J.; Xu, G.K.; Lipowsky, R.; Weikl, T.R. Binding kinetics of membrane-anchored receptors and ligands: Molecular dynamics simulations and theory. J. Chem. Phys. 2015, 143, 243137. [Google Scholar] [CrossRef]
- Xu, G.K.; Hu, J.; Lipowsky, R.; Weikl, T.R. Binding constants of membrane-anchored receptors and ligands: A general theory corroborated by Monte Carlo simulations. J. Chem. Phys. 2015, 143, 243136. [Google Scholar] [CrossRef]
- Dustin, M.L.; Bromley, S.K.; Davis, M.M.; Zhu, C. Identification of self through two-dimensional chemistry and synapses. Annu. Rev. Cell Dev. Biol. 2001, 17, 133–157. [Google Scholar] [CrossRef]
- Zhu, D.M.; Dustin, M.L.; Cairo, C.W.; Golan, D.E. Analysis of two-dimensional dissociation constant of laterally mobile cell adhesion molecules. Biophys. J. 2007, 92, 1022–1034. [Google Scholar] [CrossRef] [PubMed]
- Krobath, H.; Różycki, B.; Lipowsky, R.; Weikl, T.R. Binding cooperativity of membrane adhesion receptors. Soft Matter 2009, 5, 3354–3361. [Google Scholar] [CrossRef]
- Li, L.; Gui, C.; Hu, J.; Różycki, B. Membrane-mediated cooperative interactions of CD47 and SIRPα. Membranes 2023, 13, 871. [Google Scholar] [CrossRef]
- Steinkühler, J.; Różycki, B.; Alvey, C.; Lipowsky, R.; Weikl, T.R.; Dimova, R.; Discher, D.E. Membrane fluctuations and acidosis regulate cooperative binding of ‘marker of self’protein CD47 with the macrophage checkpoint receptor SIRPα. J. Cell Sci. 2019, 132, jcs216770. [Google Scholar] [CrossRef]
- Weikl, T.R.; Asfaw, M.; Krobath, H.; Różycki, B.; Lipowsky, R. Adhesion of membranes via receptor–ligand complexes: Domain formation, binding cooperativity, and active processes. Soft Matter 2009, 5, 3213–3224. [Google Scholar] [CrossRef]
- Carbone, C.B.; Kern, N.; Fernandes, R.A.; Hui, E.; Su, X.; Garcia, K.C.; Vale, R.D. In vitro reconstitution of T cell receptor-mediated segregation of the CD45 phosphatase. Proc. Natl. Acad. Sci. USA 2017, 114, E9338–E9345. [Google Scholar] [CrossRef]
- Schmid, E.M.; Bakalar, M.H.; Choudhuri, K.; Weichsel, J.; Ann, H.S.; Geissler, P.L.; Dustin, M.L.; Fletcher, D.A. Size-dependent protein segregation at membrane interfaces. Nat. Phys. 2016, 12, 704–711. [Google Scholar] [CrossRef]
- Rouhiparkouhi, T.; Weikl, T.R.; Discher, D.E.; Lipowsky, R. Adhesion-induced phase behavior of two-component membranes and vesicles. Int. J. Mol. Sci. 2013, 14, 2203–2229. [Google Scholar] [CrossRef]
- Weikl, T.R.; Andelman, D.; Komura, S.; Lipowsky, R. Adhesion of membranes with competing specific and generic interactions. Eur. Phys. J. E 2002, 8, 59–66. [Google Scholar] [CrossRef]
- Asfaw, M.; Różycki, B.; Lipowsky, R.; Weikl, T.R. Membrane adhesion via competing receptor/ligand bonds. Europhys. Lett. 2006, 76, 703. [Google Scholar] [CrossRef]
- Różycki, B.; Lipowsky, R.; Weikl, T.R. Segregation of receptor–ligand complexes in cell adhesion zones: Phase diagrams and the role of thermal membrane roughness. New J. Phys. 2010, 12, 095003. [Google Scholar] [CrossRef]
- Weikl, T.R.; Lipowsky, R. Adhesion-induced phase behavior of multicomponent membranes. Phys. Rev. E 2001, 64, 011903. [Google Scholar] [CrossRef]
- Li, L.; Hu, J.; Shi, X.; Shao, Y.; Song, F. Lipid rafts enhance the binding constant of membrane-anchored receptors and ligands. Soft Matter 2017, 13, 4294–4304. [Google Scholar] [CrossRef]
- Li, L.; Hu, J.; Xu, G.; Song, F. Binding constant of cell adhesion receptors and substrate-immobilized ligands depends on the distribution of ligands. Phys. Rev. E 2018, 97, 012405. [Google Scholar] [CrossRef]
- Huang, J.; Zarnitsyna, V.I.; Liu, B.; Edwards, L.J.; Jiang, N.; Evavold, B.D.; Zhu, C. The kinetics of two-dimensional TCR and pMHC interactions determine T-cell responsiveness. Nature 2010, 464, 932–936. [Google Scholar] [CrossRef]
- Murai, T.; Sato, C.; Sato, M.; Nishiyama, H.; Suga, M.; Mio, K.; Kawashima, H. Membrane cholesterol modulates the hyaluronan-binding ability of CD44 in T lymphocytes and controls rolling under shear flow. J. Cell Sci. 2013, 126, 3284–3294. [Google Scholar] [CrossRef]
- Anderson, H.A.; Roche, P.A. MHC class II association with lipid rafts on the antigen presenting cell surface. Biochim. Biophys. Acta (BBA)-Mol. Cell Res. 2015, 1853, 775–780. [Google Scholar] [CrossRef]
- Pandey, P.R.; Różycki, B.; Lipowsky, R.; Weikl, T.R. Structural variability and concerted motions of the T cell receptor–CD3 complex. Elife 2021, 10, e67195. [Google Scholar] [CrossRef]
- Pandey, P.R.; Różycki, B.; Weikl, T.R. Molecular Dynamics Simulations of Immune Receptors and Ligands. In The Immune Synapse: Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2023; pp. 51–59. [Google Scholar]
- Hou, R.; Ren, S.; Wang, R.; Różycki, B.; Hu, J. Multiscale Simulations of Membrane Adhesion Mediated by CD47-SIRPα Complexes. J. Chem. Theory Comput. 2025, 21, 2030–2042. [Google Scholar] [CrossRef]
- Hou, R.; Gao, J.; Chen, J.; Wang, R.; Różycki, B.; Hu, J. Binding of Soluble Ligands to Membrane Receptors: A Molecular Dynamics Simulation Study. J. Phys. Chem. B 2025, 129, 7475–7482. [Google Scholar] [CrossRef]
- Duncan, A.L.; Pezeshkian, W. Mesoscale simulations: An indispensable approach to understand biomembranes. Biophys. J. 2023, 122, 1883–1889. [Google Scholar] [CrossRef]
- Bahrami, A.H.; Weikl, T.R. Curvature-mediated assembly of janus nanoparticles on membrane vesicles. Nano Lett. 2018, 18, 1259–1263. [Google Scholar] [CrossRef]
- Helfrich, W. Elastic properties of lipid bilayers: Theory and possible experiments. Z. Naturforschung C 1973, 28, 693–703. [Google Scholar] [CrossRef]
- Li, L.; Gao, J.; Milewski, Ł.; Hu, J.; Różycki, B. Lattice-based mesoscale simulations and mean-field theory of cell membrane adhesion. In Methods in Enzymology; Elsevier: Amsterdam, The Netherlands, 2024; Volume 701, pp. 425–455. [Google Scholar]
- Williams, T.; Kelley, C. Gnuplot 5.4: An Interactive Plotting Program. 2020. Available online: http://gnuplot.sourceforge.net/ (accessed on 28 June 2025).
- Speck, T.; Reister, E.; Seifert, U. Specific adhesion of membranes: Mapping to an effective bond lattice gas. Phys. Rev. E 2010, 82, 021923. [Google Scholar] [CrossRef]
- Fenz, S.F.; Bihr, T.; Schmidt, D.; Merkel, R.; Seifert, U.; Sengupta, K.; Smith, A.S. Membrane fluctuations mediate lateral interaction between cadherin bonds. Nat. Phys. 2017, 13, 906–913. [Google Scholar] [CrossRef]
- Gao, J.; Shen, Y.; Komura, S.; Hu, W.; Shen, L.; Hu, J. Polymer sailing on rafts within lipid membranes. Proc. Natl. Acad. Sci. USA 2025, 122, e2503203122. [Google Scholar] [CrossRef]
- Pettmann, J.; Awada, L.; Różycki, B.; Huhn, A.; Faour, S.; Kutuzov, M.; Limozin, L.; Weikl, T.R.; van der Merwe, P.A.; Robert, P.; et al. Mechanical forces impair antigen discrimination by reducing differences in T-cell receptor/peptide–MHC off-rates. EMBO J. 2023, 42, e111841. [Google Scholar] [CrossRef]
- Różycki, B.; Lipowsky, R. Spontaneous curvature of bilayer membranes from molecular simulations: Asymmetric lipid densities and asymmetric adsorption. J. Chem. Phys. 2015, 142, 054101. [Google Scholar] [CrossRef]
- Stachowiak, J.C.; Schmid, E.M.; Ryan, C.J.; Ann, H.S.; Sasaki, D.Y.; Sherman, M.B.; Geissler, P.L.; Fletcher, D.A.; Hayden, C.C. Membrane bending by protein–protein crowding. Nat. Cell Biol. 2012, 14, 944–949. [Google Scholar] [CrossRef]
- Stone, M.B.; Shelby, S.A.; Veatch, S.L. Super-resolution microscopy: Shedding light on the cellular plasma membrane. Chem. Rev. 2017, 117, 7457–7477. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Thalakodan, I.B.H.; Różycki, B. Interplay Between Membrane Adhesion and Distribution of Lipid Rafts. Surfaces 2025, 8, 62. https://doi.org/10.3390/surfaces8030062
Thalakodan IBH, Różycki B. Interplay Between Membrane Adhesion and Distribution of Lipid Rafts. Surfaces. 2025; 8(3):62. https://doi.org/10.3390/surfaces8030062
Chicago/Turabian StyleThalakodan, Iyad Bin Hussain, and Bartosz Różycki. 2025. "Interplay Between Membrane Adhesion and Distribution of Lipid Rafts" Surfaces 8, no. 3: 62. https://doi.org/10.3390/surfaces8030062
APA StyleThalakodan, I. B. H., & Różycki, B. (2025). Interplay Between Membrane Adhesion and Distribution of Lipid Rafts. Surfaces, 8(3), 62. https://doi.org/10.3390/surfaces8030062





