1. Introduction
Diabetes is a chronic condition that requires careful management to maintain blood glucose levels (BGLs) within a safe range [
1]. Several factors, including diet, physical activity, insulin administration, stress, and illness, influence glucose fluctuations, making regulation challenging [
2]. Therefore, self-care, adherence to lifestyle recommendations, and timely blood glucose monitoring play a crucial role in effective diabetes management [
3,
4]. To facilitate glucose monitoring and minimize the risk of both short- and long-term complications, continuous glucose monitoring (CGM) systems have become increasingly prevalent [
5]. These systems measure glucose levels in the interstitial fluid beneath the skin, estimating plasma glucose concentrations with a high sampling rate. The vast amount of data generated by CGM can be leveraged in both physiological and data-driven models for blood glucose prediction, each offering distinct advantages that aid early intervention and complication prevention [
6]. Accurate and timely predictions enable proactive decision-making to mitigate the risks of hyperglycemia and hypoglycemia while optimizing dietary choices, exercise routines, and treatment plans [
7,
8].
Physiological models mathematically describe glucose metabolism and kinetics. However, they require detailed knowledge of an individual’s physiological processes and prior configuration of numerous parameters [
9,
10]. Estimating and fine-tuning these parameters is often complex, error-prone, and time-consuming due to limited observed data. In contrast, data-driven models rely solely on self-monitored historical data, requiring minimal knowledge of glucose metabolism [
10,
11,
12]. These models, often referred to as black-box approaches, have demonstrated superior performance over physiological models. However, their lack of physiological grounding results in less generalizable predictions, making interpretation difficult. Another challenge with these models is their dependence on large amounts of labeled data, which is often scarce in clinical settings [
9].
A variety of machine learning (ML) architectures have been explored for blood glucose prediction, ranging from traditional regression-based algorithms such as autoregressive moving average (ARMA), autoregressive integrated moving average (ARIMA), random forest (RF), extreme gradient boosting (XGBoost), and support vector regression (SVR) to more advanced models [
9,
10,
11,
12,
13]. These range from basic feedforward neural networks (FNNs) to deep learning architectures such as recurrent neural networks (RNNs), convolutional neural networks (CNNs), temporal CNNs (TCNs), and attention-based networks. Due to their high predictive accuracy, deep learning works have rapidly emerged as a powerful tool for blood glucose prediction. Among these, long short-term memory (LSTM) networks are widely used in different time series applications [
14,
15,
16,
17] due to their architecture, which includes memory cells and input, forget, and output gates. These components dynamically regulate information flow, preserving critical patterns and insights across extended sequences, making LSTMs particularly effective for time series predictions like BGLs [
12,
13].
Attention mechanisms, originally designed for natural language processing and computer vision tasks [
18], have also been applied to blood glucose prediction [
19]. Transformers, in particular, have demonstrated significant success in handling sequential data. In the context of BGL prediction, studies [
20,
21,
22] have utilized attention-based recurrent networks to enhance predictive performance. Additionally, other studies [
7,
23] have implemented and evaluated the effectiveness of self-attention networks in predicting BGLs.
Even though CNNs have not been extensively used for time series forecasting or BGL prediction, there are instances where they have been used, either independently in some studies [
7,
24] or for feature extraction in conjunction with other models [
2,
14]. These applications have demonstrated CNNs to be suitable for BGL prediction or any other time series forecasting. Given their ability to address long-term dependencies within time series data, TCNs are also particularly useful for predicting BGL. BGLs are responsive to factors that manifest hours or even days prior, necessitating models capable of learning such extended dependencies, a capability TCNs possess [
23]. Few studies have considered TCN for BGL prediction [
11,
25]; nevertheless, these networks have been successfully applied for other time series forecasting tasks.
A potential limitation in existing data-driven BGL predictive models is their ability to accurately represent insulin and meal dynamics. When bolus insulin is administered, its effect on BGLs is not immediate; rather, it follows a time-dependent pattern, entering the bloodstream, peaking at a certain point, and gradually declining [
26]. A simplistic model that relies solely on insulin dose and administration time might not fully capture this gradual influence on BGLs. Instead, a more detailed approach that considers fluctuations in insulin concentrations over time could offer a clearer representation of glucose responses. Likewise, meal-related glucose appearance rates could play a more significant role than just the total meal amount and timing. Incorporating insulin kinetics, meal-related glucose appearance rates, and other physiological parameters could enable predictive models to better reflect real-world glucose dynamics.
Several studies have developed models that integrate physiological models of blood glucose dynamics with data-driven predictive models [
1,
27,
28,
29,
30,
31,
32]. These studies have demonstrated performance improvements through the integration of various physiological knowledge. For example, Ref. [
1] enhanced the performance of the SVR model by incorporating features such as the rate of glucose appearance from meal intake, plasma insulin levels, and cumulative glucose appearance over a specific period. These features, obtained from meal and insulin models of blood glucose dynamics, serve as inputs, providing a straightforward way to incorporate domain knowledge into learning models. Similarly, Ref. [
31] included plasma insulin and glucose appearance rate from meals as input features through insulin kinetics and meal models. Additional features such as insulin on board, carbohydrates on board, glucose appearance rate from carbs, rate of glucose appearance in the blood from the gut, and activity on board were employed in [
27,
28,
29,
32] to refine machine learning models. Ref. [
30] proposed a hybrid method that sequentially combines predictions from both mathematical and machine learning models, where the machine learning model predicts residuals based on phenotypic features, which are then subtracted from the predictions made by the mathematical model. The results demonstrated that personalized physiological models consistently outperformed data-driven and hybrid model approaches. Here, personalized physiological models may inherently capture individualized critical processes and features that data-driven models struggle to individualize or interpret effectively. While these studies have demonstrated performance enhancements through the integration of various physiological insights specifically as static input features, they have yet to explore integration at a personalized level and lack interpretability in this context, a limitation that this study aims to address. Furthermore, they have not systematically analyzed the potential benefits of such integration, representing an additional gap that this study seeks to fill.
Hybrid modeling approaches combining physiological ordinary differential equation (ODE) knowledge with neural networks have emerged as a promising direction for glucose–insulin dynamics modeling. Ref. [
33] proposed systems biology-informed neural networks (SBINNs), a general methodological framework for inferring hidden dynamics and unknown ODE parameters from sparse noisy observations by embedding ODE residuals into the neural network loss. Applied to three benchmark biological systems including an ultradian glucose–insulin model, the study demonstrates that unobserved states and unknown parameters can be recovered from minimal observations and that the model can additionally infer hidden inputs such as unknown meal content. However, the approach is validated exclusively on synthetic data generated from known models and does not explicitly address patient-specific personalization, making it primarily a methodological contribution rather than a clinically validated glucose management tool. The authors of [
34] introduce a biology-informed recurrent neural network (BI-RNN) that employs a gated recurrent unit (GRU) trained with a three-component loss function enforcing data fidelity, ODE-based state consistency, and auxiliary constraints such as non-negativity. While the model predicts glucose dynamics, its primary goal is system identification within a model predictive control framework, enabling the reconstruction of unmeasured physiological states such as insulin-on-board and rate of glucose appearance. Physiological parameters are pre-identified and kept fixed, contributing to the loss formulation as constraints rather than being learned by the network. The authors of [
35] apply physics-informed neural networks to estimate physiological parameters of the Bergman minimal model, recovering hidden insulin dynamics from glucose-only intravenous glucose tolerance test (IVGTT) data via ODE residual minimization and biologically informed parameter bounds. Most parameters are estimated with reasonable accuracy, though some exhibit notable bias. The approach is validated on simulations from a single subject with multiple noise realizations and is not evaluated on real-world or free-living data.
Ref. [
36] proposes hybrid graph sparsification (HGS), a method for automatically pruning redundant latent states and interactions from hybrid neural ODEs that combine mechanistic physiological models with neural networks, particularly useful in data-scarce healthcare settings. Blood glucose forecasting in Type 1 diabetes (T1D) patients is used as an application to demonstrate the method’s effectiveness. However, the method focuses on structural sparsification rather than parameter estimation and does not explicitly address patient-specific personalization. Ref. [
37] focuses on glucose simulation rather than pure prediction, proposing physiologically constrained neural network digital twins. Their approach constructs a neural state-space model aligned with a system of biological ODEs, where each equation is approximated by a dedicated neural network restricted to the same inputs as the original equation. The model is trained to both accurately simulate glucose dynamics and maintain physiologically consistent internal states. Rather than estimating explicit physiological parameters, the neural network weights implicitly encode the system’s dynamics. Personalization is achieved by augmenting the population model with an additional individual-specific network that learns residual dynamics from each person’s data.
A physiology-informed glucose–insulin neural network (PIGNN) is proposed in [
38], a method for 30 min ahead glucose prediction that integrates a five-state ODE into an LSTM via physiology-inspired input windows and an ODE-consistency loss penalty. The model incorporates a physiological ODE as prior knowledge but does not explicitly estimate patient-specific physiological parameters; personalization is achieved primarily through data-driven training rather than system identification. The work in [
39] proposes a blood glucose prediction method targeting T1D management by constructing a dual-channel physiology-informed neural network (PINN) architecture constrained by the Bergman Minimal Model. The framework combines a data-driven LSTM/GRU module with a physiology-informed module, integrating physiological knowledge both through architectural design and by incorporating a differential-equation residual penalty into the loss function. The approach shows consistent improvements in prediction accuracy. It is among the few reviewed works that simultaneously perform glucose prediction while enforcing Bergman-model consistency, although physiological parameters are not explicitly identified or reported and are only indirectly constrained during training.
Collectively, these works suggest that physiology-informed hybrid approaches can improve performance, particularly in data-scarce settings, while enabling more physiologically consistent and interpretable outputs and supporting richer functionalities such as prediction, simulation, control-oriented state reconstruction, and parameter inference. While Ref. [
33] shows that joint parameter and hidden state recovery is theoretically possible from minimal observations, subsequent applied works do not fully realize this potential. Most either treat parameters as fixed constants in loss function and focus solely on prediction or estimate parameters only under controlled synthetic conditions. Even where latent physiological states are predicted, as in [
39], individual parameter estimation remains unrealized. The present work addresses these limitations and extends the state of the art in three meaningful directions. First, it moves beyond existing approaches that either embed physiological parameters as fixed constraints to enforce physiological plausibility in loss function or use them merely as static input features by making key parameters trainable and subject-specific, enabling individual-level parameter estimation from real-time data. Second, the proposed Neural ODE (NODE)-based approximation of insulin and meal kinetics offers greater flexibility than fixed-parameter ODE solvers, enabling adaptation to inter-individual variability through a two-stage transfer learning strategy that pre-trains on population-level data to capture general physiological dynamics and subsequently fine-tunes on individual patient BGL data. Third, the sensitivity and counterfactual analyses provide a structured framework for clinical interpretability that most competing methods, including MTL-LSTM, SAN, and GAN-based data-driven approaches, entirely lack. Together, these contributions address the gap that no existing framework simultaneously achieves: interpretable parameter estimation from free-living CGM data, robust individual-level personalization, and counterfactual analysis.
To overcome the problems with state-of-the-art methods, we propose a novel methodology combining physiological and data-driven models to leverage their complementary strengths. In particular, we leverage NODE [
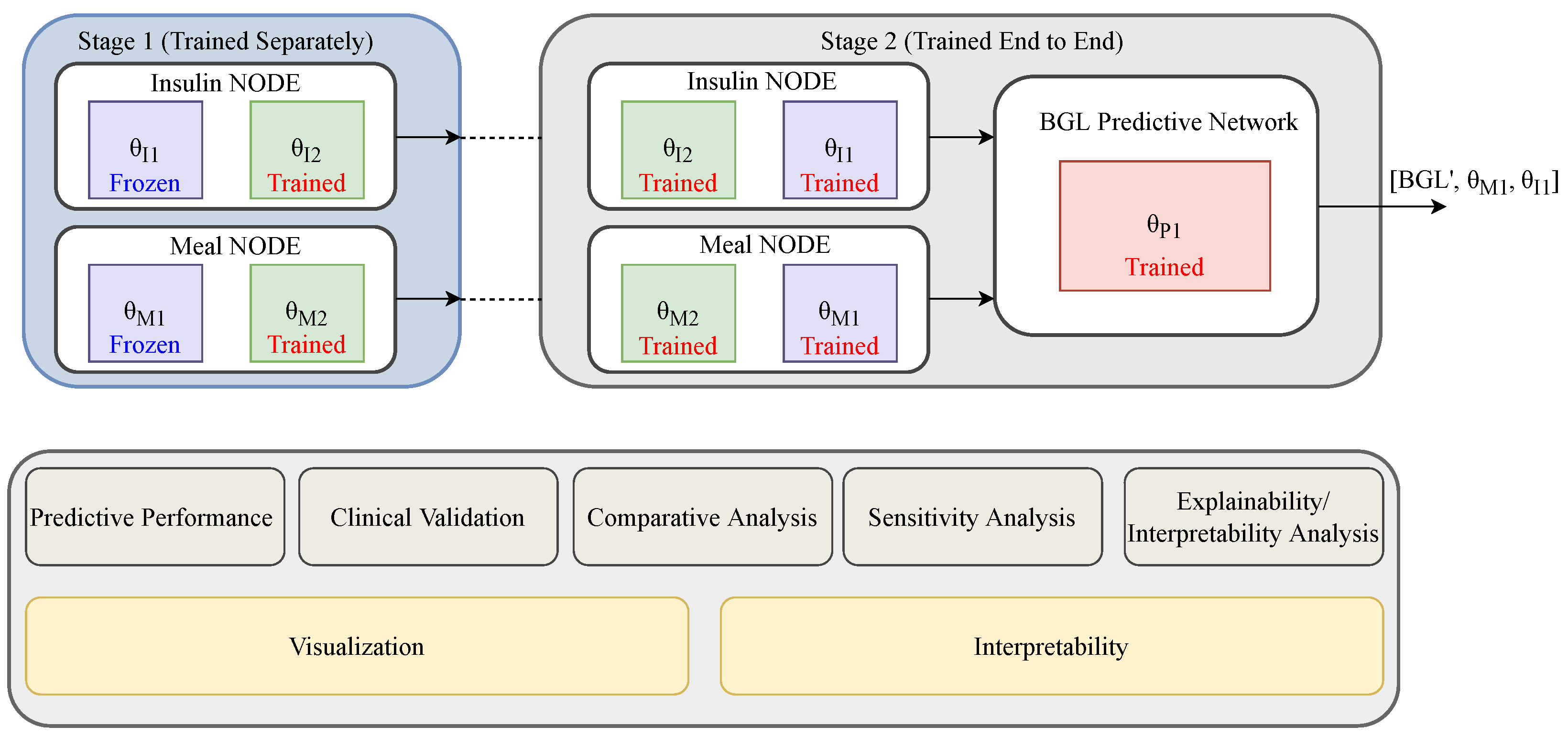
40] to approximate the evolution of insulin and glucose dynamics. These dynamics play a crucial role in blood glucose regulation and are thus integrated into the predictive model. The result is a synergistic framework that harmonizes data and physiological principles, enabling learning from both empirical observations and established physiological models. The proposed physiology-informed blood glucose prediction network (PIBGN) simultaneously predicts BGLs and identifies individualized physiological parameters using data-driven insights, facilitating the development of personalized predictive models. The system follows a structured two-stage transfer learning approach. In the first stage, it approximates the evolution of insulin and meal dynamics. The second stage refines this by learning to identify or infer the optimal physiological parameters that best correspond to the predicted glucose levels, ensuring an accurate and meaningful representation of the observed data. The main contributions of this work are summarized as follows:
Unlike prior methods that treat physiological parameters as static inputs or restrict physiological modeling to fixed training constraints, the proposed approach embeds trainable, subject-specific parameters and physiology into the network architecture, enabling interpretable estimation of physiological parameters from free-living CGM data.
A NODE-based approximation of insulin and meal kinetics is proposed that combines the structural knowledge of physiological models with the flexibility of data-driven learning. A two-stage transfer learning strategy is introduced, first pre-training on population-level data to capture general physiological dynamics, then fine-tuning on individual patient data by integrating the resulting representations into a BGL predictive network, allowing the model to adapt to inter-individual variability.
The framework incorporates sensitivity and counterfactual analyses, demonstrating how changes in insulin dosing and carbohydrate intake affect predicted glucose trajectories, offering both predictive accuracy and physiological explainability.
The remainder of this article is organized as follows. The proposed physiology-informed BGL prediction method is described in
Section 2. We discuss the experimental results and performance analysis in
Section 3. The limitations of the proposed model are demonstrated in
Section 4. Finally, this article is concluded in
Section 5.