Frequency and Spectrum of Radiation-Induced Mutations Revealed by Whole-Genome Sequencing Analyses of Plants
Abstract
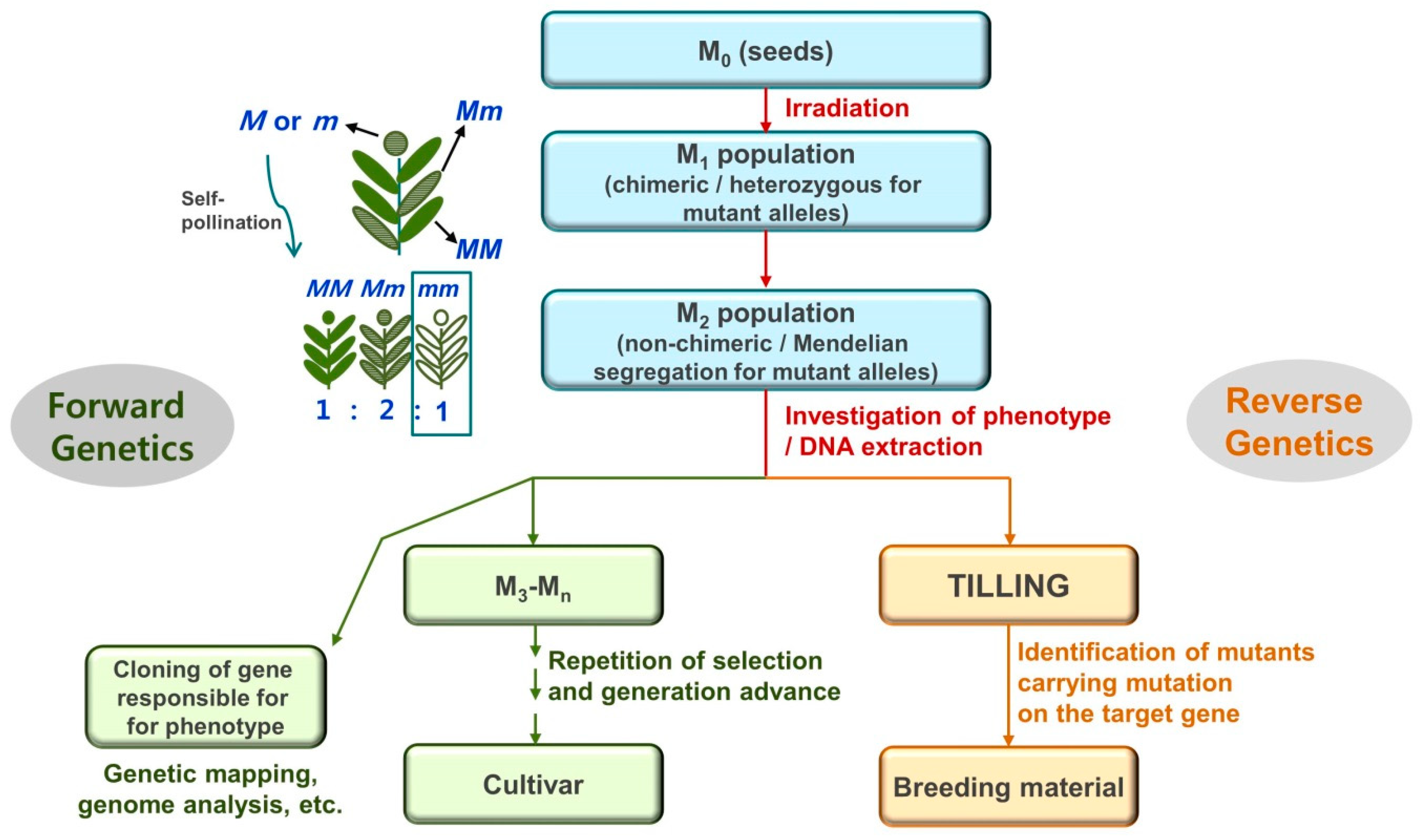
1. Introduction
2. Factors Affecting the Mutation Frequency and Spectrum
2.1. Linear Energy Transfer
2.2. Irradiation Dose
2.3. Effects of Plant Tissue Type and Condition
3. Characteristics of DNA Mutations Revealed by Whole-Genome Sequencing
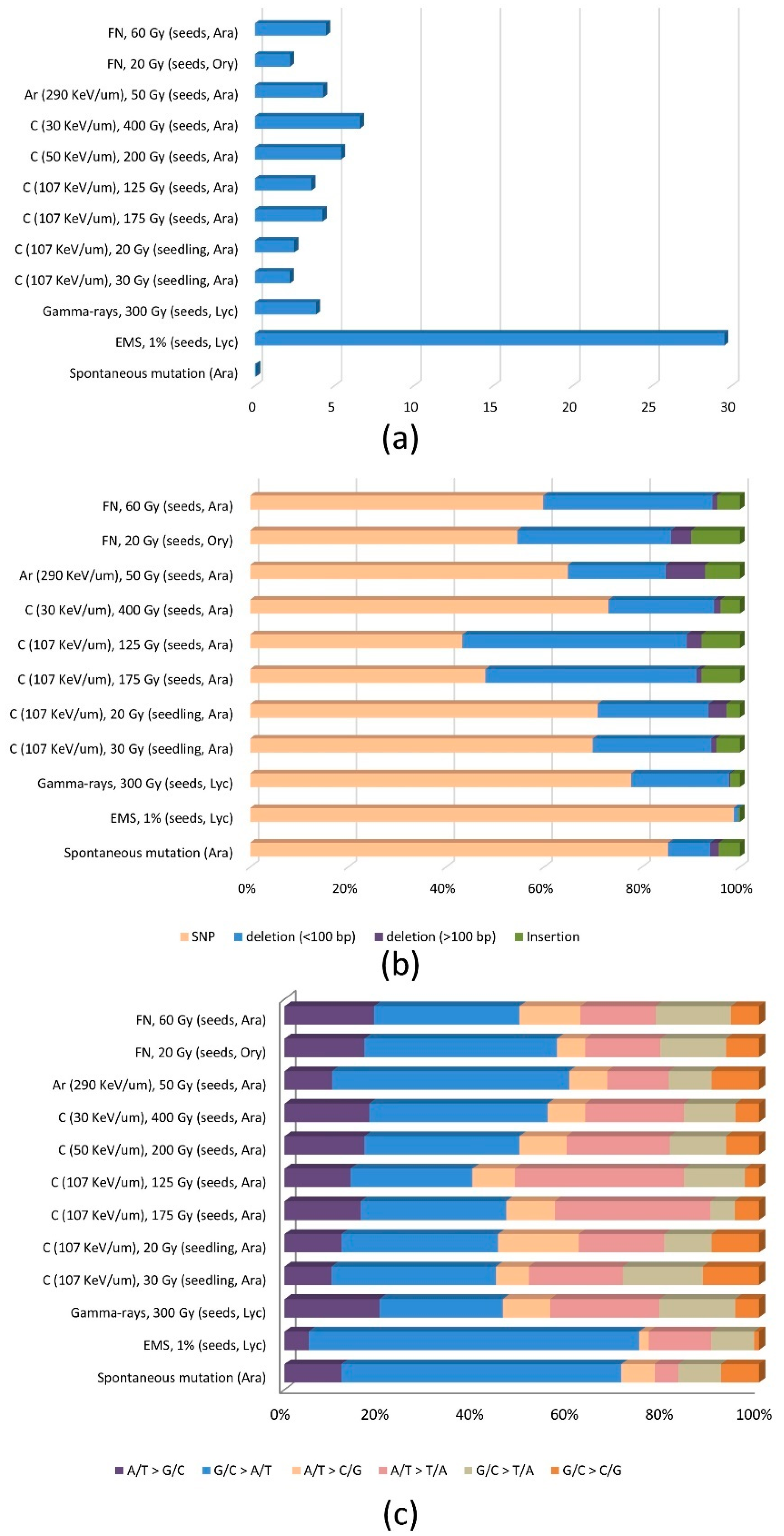
3.1. Frequency and Distribution of Mutations
3.2. Mutation Spectrum and Mechanism
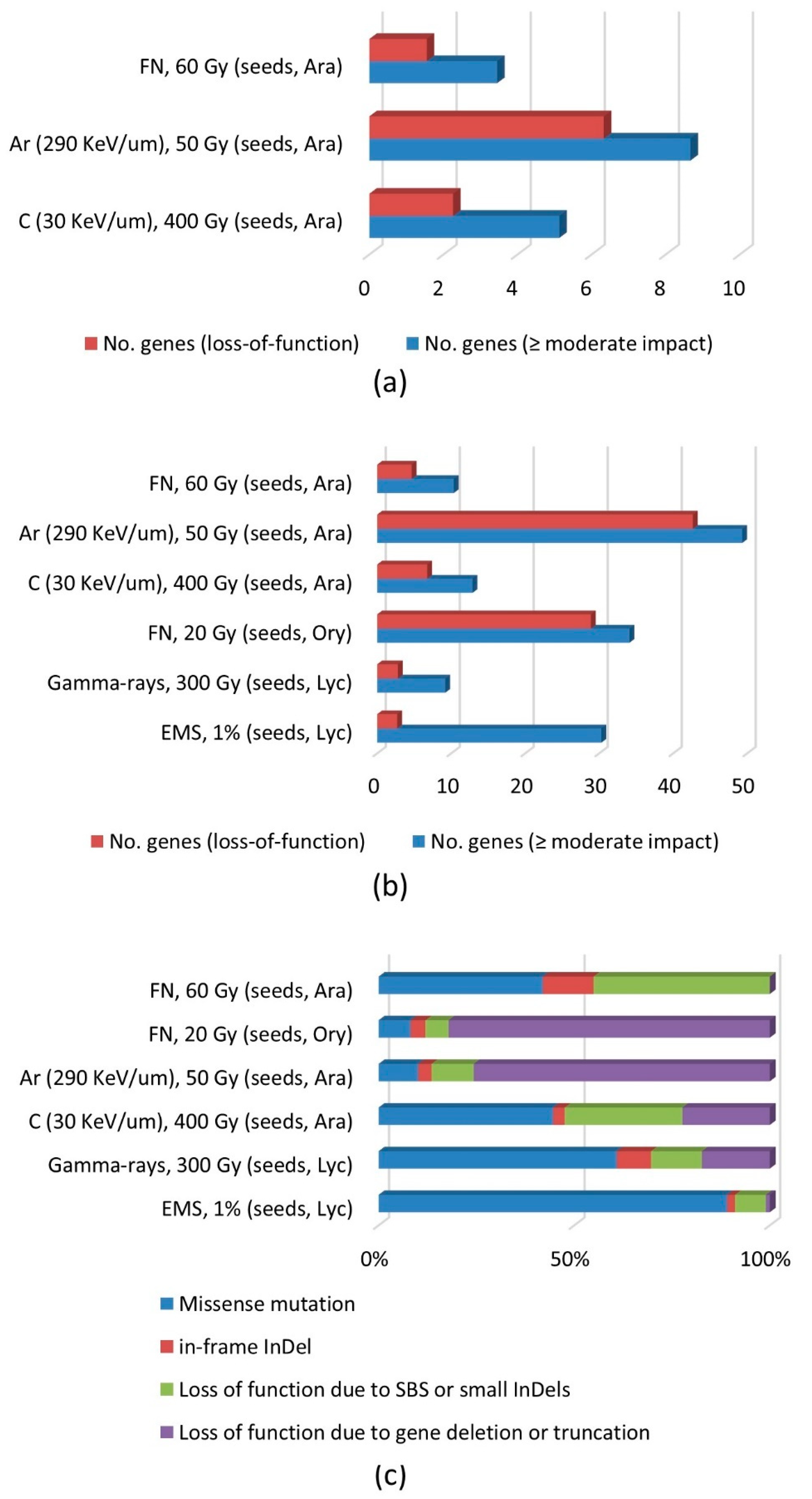
3.3. Effects on Gene Function
4. Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Stadler, L.J. Mutations in barley induced by X-rays and radium. Science 1928, 68, 186–187. [Google Scholar] [CrossRef] [PubMed]
- Stadler, L.J. Genetic effects of X-rays in maize. Proc. Natl. Acad. Sci. USA 1928, 14, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Shu, Q.Y.; Forster, B.P.; Nakagawa, H. Plant Mutation Breeding and Biotechnology; CAB International: Wallingford, UK, 2012; ISBN 9789251070222. [Google Scholar]
- Tanaka, A.; Shikazono, N.; Hase, Y. Studies on Biological Effects of Ion Beams on Lethality, Molecular Nature of Mutation, Mutation Rate, and Spectrum of Mutation Phenotype for Mutation Breeding in Higher Plants. J. Radiat. Res. 2010, 233, 223–233. [Google Scholar] [CrossRef]
- Kazama, Y.; Hirano, T.; Saito, H.; Liu, Y.; Ohbu, S.; Hayashi, Y.; Abe, T. Characterization of highly efficient heavy-ion mutagenesis in Arabidopsis thaliana. BMC Plant Biol. 2011, 11, 161. [Google Scholar] [CrossRef] [PubMed]
- Koornneef, M.; Rolff, E.; Spruit, C.J.P. Genetic control of light-inhibited hypocotyl elongation in Arabidopsis thaliana (L) Heynh. Z. Pflanzenphysiol. 1980, 100, 147–160. [Google Scholar] [CrossRef]
- Shirasawa, K.; Hirakawa, H.; Nunome, T.; Tabata, S.; Isobe, S. Genome-wide survey of artificial mutations induced by ethyl methanesulfonate and gamma rays in tomato. Plant Biotechnol. J. 2016, 14, 51–60. [Google Scholar] [CrossRef]
- IAEA Mutant Variety Database. Available online: http://mvd.iaea.org (accessed on 13 March 2019).
- Hase, Y.; Satoh, K.; Kitamu, S.; Oono, Y. Physiological status of plant tissue affects the frequency and types of mutations induced by carbon-ion irradiation in Arabidopsis. Sci. Rep. 2018, 8, 1394. [Google Scholar] [CrossRef]
- Yamaguchi, H.; Hase, Y.; Tanaka, A.; Shikazono, N.; Degi, K.; Shimizu, A.; Morishita, T. Mutagenic effects of ion beam irradiation on rice. Breed. Sci. 2009, 177, 169–177. [Google Scholar] [CrossRef]
- Hirano, T.; Kazama, Y.; Ishii, K.; Ohbu, S.; Shirakawa, Y.; Abe, T. Comprehensive identification of mutations induced by heavy-ion beam irradiation in Arabidopsis thaliana. Plant J. 2015, 82, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Kazama, Y.; Ishii, K.; Hirano, T.; Wakana, T.; Yamada, M.; Ohbu, S.; Abe, T. Different mutational function of low- and high-linear energy transfer heavy-ion irradiation demonstrated by whole-genome resequencing of Arabidopsis mutants. Plant J. 2017, 92, 1020–1030. [Google Scholar] [CrossRef] [PubMed]
- Taheri, S.; Abdullah, T.L. TILLING, high-resolution melting (HRM), and next-generation sequencing (NGS) techniques in plant mutation breeding. Mol. Breed. 2017, 37, 40. [Google Scholar] [CrossRef]
- Li, X.; Song, Y.; Century, K.; Straight, S.; Ronald, P.; Dong, X.; Lassner, M. A fast neutron deletion mutagenesis-based reverse genetics system for plants. Plant J. 2001, 27, 235–242. [Google Scholar] [CrossRef]
- Rodgers, C.; Wen, J.; Chen, R.; Oldroyd, G. Deletion-based reverse genetics in Medicago truncatula. Plant Physiol. 2009, 151, 1077–1086. [Google Scholar] [CrossRef]
- Arase, S.; Hase, Y.; Abe, J.; Kasai, M.; Yamada, T.; Kitamura, K.; Narumi, I.; Tanaka, A.; Kanazawa, A. Optimization of ion-beam irradiation for mutagenesis in soybean: Effects on plant growth and production of visibly altered mutants. Plant Biotechnol. 2011, 329, 323–329. [Google Scholar] [CrossRef]
- Jo, Y.D.; Kim, S.H.; Hwang, J.E.; Kim, Y.S.; Kang, H.S.; Kim, S.W.; Kwon, S.J.; Ryu, J.; Kim, J.B.; Kang, S.Y. Construction of mutation populations by gamma-ray and carbon beam irradiation in chili pepper (Capsicum annuum L.). Hortic. Environ. Biotechnol. 2016, 57, 606–614. [Google Scholar] [CrossRef]
- Yamaguchi, H.; Shimizu, A.; Degi, K.; Morishita, T. Effects of dose and dose rate of gamma ray irradiation on mutation induction and nuclear DNA content in chrysanthemum. Breed. Sci. 2008, 335, 331–335. [Google Scholar] [CrossRef]
- Hirano, T.; Kazama, Y.; Ohbu, S.; Shirakawa, Y.; Liu, Y.; Kambara, T. Fundamental and molecular mechanisms of mutagenesis molecular nature of mutations induced by high-LET irradiation with argon and carbon ions in Arabidopsis thaliana. Mutat. Res./Fundam. Mol. Mech. Mutagen. 2012, 735, 19–31. [Google Scholar] [CrossRef]
- Shikazono, N.; Tanaka, A.; Watanabe, H.; Tano, S. Rearrangements of the DNA in Carbon Ion-Induced Mutants of Arabidopsis thaliana. Genetics 2001, 157, 379–387. [Google Scholar]
- Shikazono, N.; Yokota, Y.; Kitamura, S.; Suzuki, C.; Watanabe, H.; Tano, S.; Tanaka, A. Mutation Rate and Novel tt Mutants of Arabidopsis thaliana Induced by Carbon Ions. Genetics 2003, 1455, 1449–1455. [Google Scholar]
- Shikazono, N.; Suzuki, C.; Kitamura, S.; Watanabe, H.; Tano, S. Analysis of mutations induced by carbon ions in Arabidopsis thaliana. J. Exp. Bot. 2005, 56, 587–596. [Google Scholar] [CrossRef] [PubMed]
- Belfield, E.J.; Gan, X.; Mithani, A.; Brown, C.; Jiang, C.; Franklin, K.; Alvey, E.; Wibowo, A.; Jung, M.; Bailey, K.; et al. Genome-wide analysis of mutations in mutant lineages selected following fast-neutron irradiation mutagenesis of Arabidopsis thaliana. Genome Res. 2012, 22, 1306–1315. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Luo, S.; Li, X.; Yang, J.; Cui, T.; Li, W.; Yu, L. Identification of Substitutions and Small Insertion-Deletions Induced by Carbon-Ion Beam Irradiation in Arabidopsis thaliana. Front. Plant Sci. 2017, 8, 1851. [Google Scholar] [CrossRef]
- Li, G.; Chern, M.; Jain, R.; Martin, J.A.; Schackwitz, W.S.; Jiang, L.; Vega-Sánchez, M.E.; Lipzen, A.M.; Barry, K.W.; Schmutz, J.; et al. Genome-wide sequencing of 41 rice (Oryza sativa L.) mutated lines reveals diverse mutations induced by fast neutron irradiation. Mol. Plant 2016, 9, 1078–1081. [Google Scholar] [CrossRef]
- Li, G.; Jain, R.; Chern, M.; Pham, N.T.; Martin, J.A.; Wei, T. The sequences of 1504 mutants in the model rice variety Kitaake facilitate rapid functional genomic studies. Plant Cell 2017, 29, 1218–1231. [Google Scholar]
- Hase, Y.; Yoshihara, R.; Nozawa, S.; Narumi, I. Fundamental and Molecular Mechanisms of Mutagenesis Mutagenic effects of carbon ions near the range end in plants. Mutat. Res./Fundam. Mol. Mech. Mutagen. 2012, 731, 41–47. [Google Scholar] [CrossRef]
- Kazama, Y.; Saito, H.; Yamamoto, Y.Y.; Hayashi, Y.; Ichida, H.; Ryuto, H.; Fukunishi, N.; Abe, T. LET-dependent effects of heavy-ion beam irradiation in Arabidopsis thaliana. Plant Biotechnol. 2008, 117, 113–117. [Google Scholar] [CrossRef]
- Yoshihara, R.; Nozawa, S.; Hase, Y.; Narumi, I.; Hidema, J.; Sakamoto, A.N. Mutational effects of γ-rays and carbon ion beams on Arabidopsis seedlings. J. Radiat. Res. 2013, 54, 1050–1056. [Google Scholar] [CrossRef] [PubMed]
- Ballarini, F.; Alloni, D.; Facoetti, A.; Ottolenghi, A. Heavy-ion effects: From track structure to DNA and chromosome damage. New J. Phys. 2008, 10, 075008. [Google Scholar] [CrossRef]
- Hada, M.; Georgakilas, A.G. Formation of clustered DNA damage after high-LET irradiation: A Review. J. Radiat. Res. 2008, 49, 203–210. [Google Scholar] [CrossRef]
- Roots, R.; Okada, S. Estimation of Life times and diffusion distances of radicals DNA strand breaks involved in X-ray-induced or killing of mammalian cells. Radiat. Res. 1975, 320, 306–320. [Google Scholar] [CrossRef]
- Wallace, S.S. Enzymatic processing of radiation-induced free radical damage in DNA1. Radiat. Res. 1998, 150, S60–S79. [Google Scholar] [CrossRef]
- Kikuchi, S.; Saito, Y.; Ryuto, H.; Fukunishi, N.; Abe, T.; Tanaka, H.; Tsujimoto, H. Mechanisms of mutagenesis effects of heavy-ion beams on chromosomes of common wheat, Triticum aestivum. Mutagenes. Res. 2009, 669, 63–66. [Google Scholar] [CrossRef]
- Okamura, M. Wide variety of flower-color and -shape mutants regenerated from leaf cultures irradiated with ion beams. Nucl. Instrum. Methods Phys. Res. 2003, 206, 574–578. [Google Scholar] [CrossRef]
- Tanaka, A. (Kansai Photon Science Institute Kyoto, Japan). Personal communication, 2016.
- Vochita, G.; Focea-ghioc, R.; Creanga, D. Direct versus indirect radiation action in irradiated vegetal embryos. Cent. Eur. J. Biol. 2014, 9, 993–1003. [Google Scholar] [CrossRef][Green Version]
- Kowyama, Y.; Saba, T.; Tsuji, T.; Kawase, T. Specific developmental stages of gametogenesis for radiosensitivity and mutagenesis in rice. Euphytica 1994, 80, 27–38. [Google Scholar] [CrossRef]
- Hase, Y.; Okamura, M.; Takeshita, D.; Narumi, I. Efficient induction of flower-color mutants by ion beam irradiation in petunia seedlings treated with high sucrose. Plant Biotechnol. 2010, 103, 99–103. [Google Scholar] [CrossRef]
- Hase, Y.; Akita, Y.; Kitamura, S.; Narumi, I.; Tanaka, A. Development of an efficient mutagenesis technique using ion beams: Toward more controlled mutation breeding. Plant Biotechnol. 2012, 200, 193–200. [Google Scholar] [CrossRef][Green Version]
- Naito, K.; Kusaba, M.; Shikazono, N.; Takano, T.; Tanaka, A.; Tanisaka, T.; Nishimura, M. Transmissible and nontransmissible mutations induced by irradiating Arabidopsis thaliana pollen With λ-rays and carbon ions. Genetics 2005, 169, 881–889. [Google Scholar] [CrossRef]
- Song, Q.; Cannistraro, V.J.; Taylor, J. Synergistic modulation of cyclobutane pyrimidine dimer photoproduct formation and deamination at a TmCG site over a full helical DNA turn in a nucleosome core particle. Nucleic Acids Res. 2014, 42, 13122–13133. [Google Scholar] [CrossRef]
- Decottignies, A. Alternative end-joining mechanisms: A historical perspective. Front. Genet. 2013, 4, 48. [Google Scholar] [CrossRef]
- Ishii, K.; Kazama, Y.; Morita, R.; Hirano, T.; Ikeda, T. Linear energy transfer-dependent change in rice gene expression profile after heavy-ion beam irradiation. PLoS ONE 2016, 11, e0160061. [Google Scholar] [CrossRef]
- Henry, I.M.; Nagalakshm, U.; Lieberman, M.C.; Ngo, K.J.; Krasileva, K.V.; Vasquez-Gross, H.; Akhunova, A.; Akhunov, E.; Dubcovsky, J.; Tai, T.H.; et al. Efficient genome-wide detection and cataloging of EMS-induced mutations using exome capture and next-generation sequencing. Plant Cell 2014, 26, 1382–1397. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.J.; Ryu, J.; Im, J.; Kim, S.H.; Ikeda, T. Molecular characterization of proton beam-induced mutations in soybean using genotyping-by-sequencing. Mol. Genet. Genom. 2018, 293, 1169–1180. [Google Scholar] [CrossRef]
| Mutagen | Dosage | Species and Tissue | Biological Effect in M1 | Generation of the Sequenced Samples | Number of Sequenced Samples | Selection of Samples by Phenotype w | Reference | |
|---|---|---|---|---|---|---|---|---|
| Fast neutrons | 60 Gy | Arabidopsis thaliana (Ler), seeds | - z | M3 | 6 | Yes | [23] | |
| 20 Gy | Oryza sativa (cv. X. Kitaake), seeds | 81% fertility | M2 | 1370 | No | [27] | ||
| M3 | 134 | |||||||
| Ion beams | Argon ion beams (290 KeV μm−1) | 50 Gy | Arabidopsis thaliana (Col-0), seeds | 95% survival | M2 (pooled M3) y | 3 | Yes | [11] |
| 8 | ||||||||
| Carbon ion beams (30 KeV μm−1) | 400 Gy | 95% survival | M2 (pooled M3) | 8 | Yes | [12] | ||
| Carbon ion beams (50 KeV μm−1) | 200 Gy | - | M3 | 11 | Yes/No v | [24] | ||
| Carbon ion beams (107 KeV μm−1) | 125 Gy | 50% Dq x | M2 | 6 | No | [27] | ||
| 175 Gy | 75% Dq | 6 | ||||||
| 20 Gy | Arabidopsis thaliana (Col-0), 7-day-old seedlings | 50% Dq | 6 | |||||
| 30 Gy | 75% Dq | 6 | ||||||
| Gamma rays | 300 Gy | Solanum lycopersicum (cv. Micro- Tom), seeds | - | M3 | 3 | No | [7] | |
| EMS | 1.0% | - | 4 | |||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jo, Y.D.; Kim, J.-B. Frequency and Spectrum of Radiation-Induced Mutations Revealed by Whole-Genome Sequencing Analyses of Plants. Quantum Beam Sci. 2019, 3, 7. https://doi.org/10.3390/qubs3020007
Jo YD, Kim J-B. Frequency and Spectrum of Radiation-Induced Mutations Revealed by Whole-Genome Sequencing Analyses of Plants. Quantum Beam Science. 2019; 3(2):7. https://doi.org/10.3390/qubs3020007
Chicago/Turabian StyleJo, Yeong Deuk, and Jin-Baek Kim. 2019. "Frequency and Spectrum of Radiation-Induced Mutations Revealed by Whole-Genome Sequencing Analyses of Plants" Quantum Beam Science 3, no. 2: 7. https://doi.org/10.3390/qubs3020007
APA StyleJo, Y. D., & Kim, J.-B. (2019). Frequency and Spectrum of Radiation-Induced Mutations Revealed by Whole-Genome Sequencing Analyses of Plants. Quantum Beam Science, 3(2), 7. https://doi.org/10.3390/qubs3020007






