Enhanced Heart Disease Prediction Based on Machine Learning and χ2 Statistical Optimal Feature Selection Model
Abstract
1. Introduction
- To develop a new heart disease (HD) classification model based on the ML (SVM) algorithm to improve the detection of heart disease;
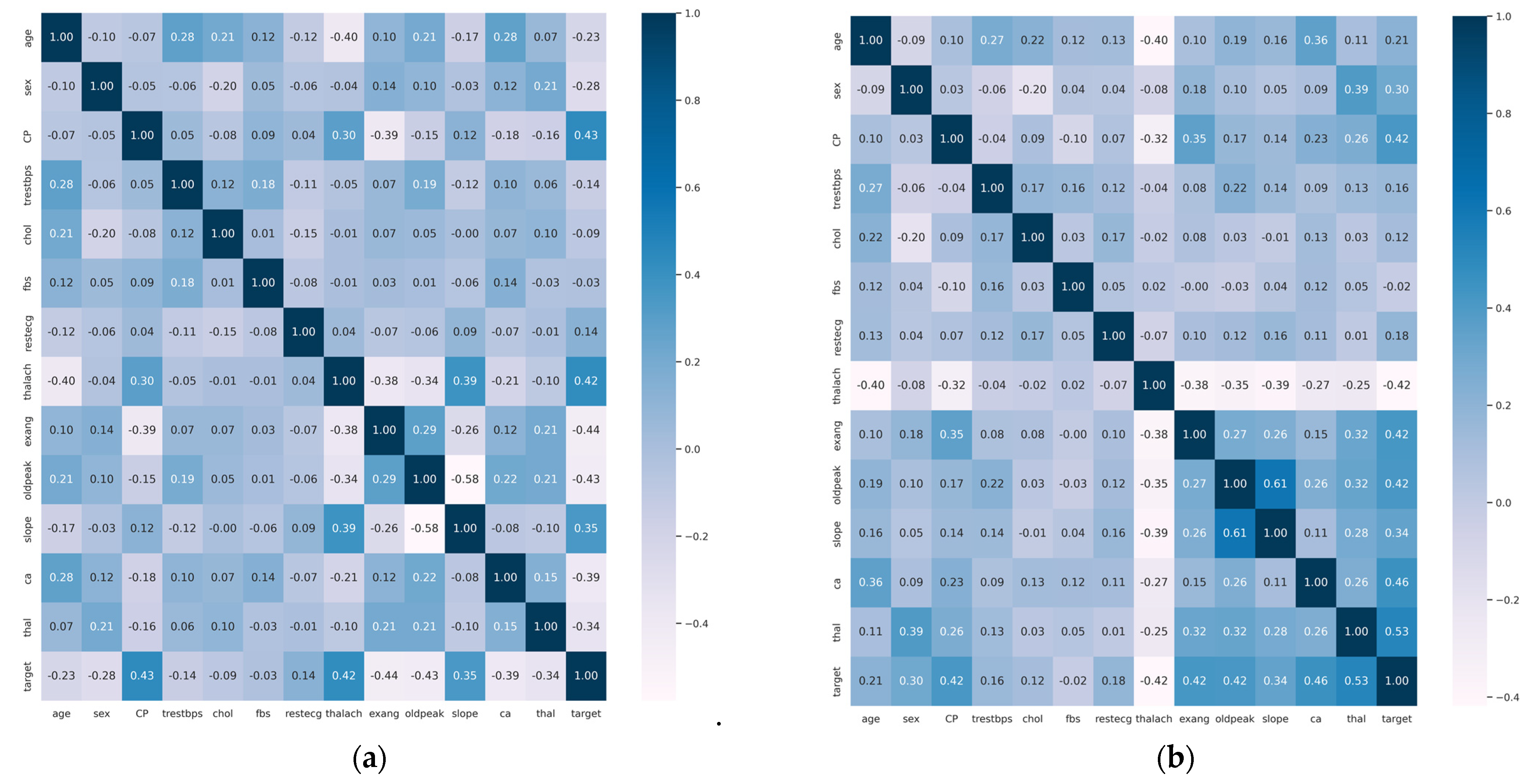
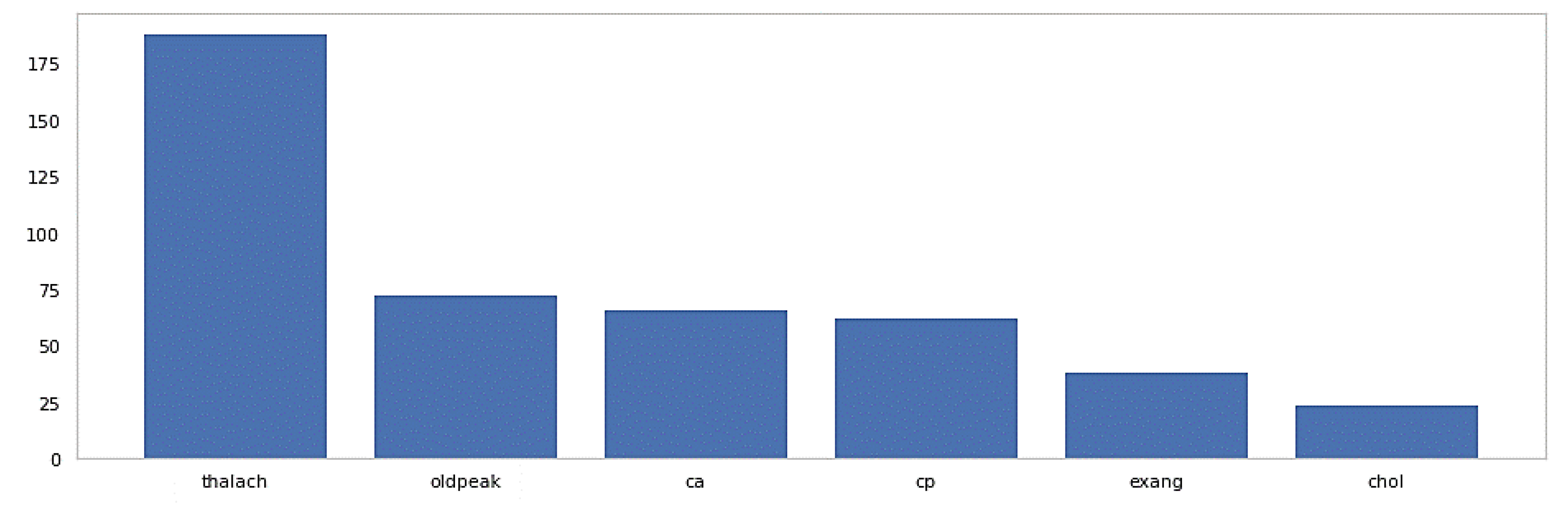
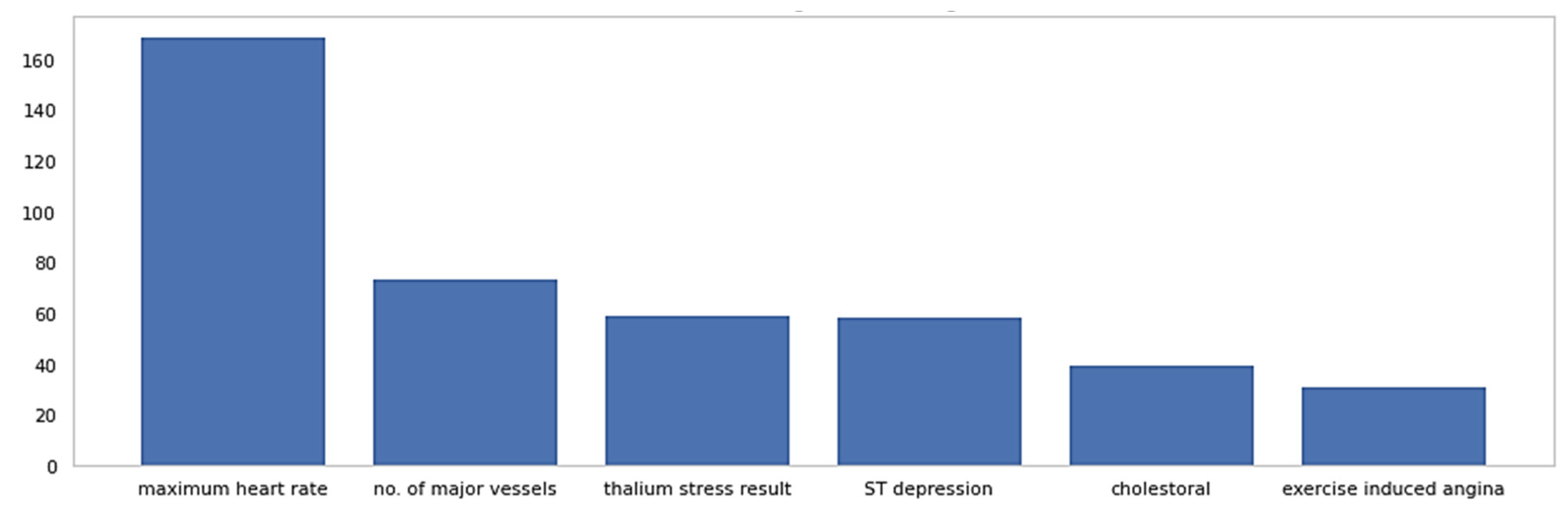
- To implement an optimal feature selection model using the χ2 statistical method for the extraction of the most informative attributes to improve prediction accuracy;
- To validate the proposed heart disease diagnosis model’s accuracy by comparing it with traditional models through the analysis of several performance metrics.
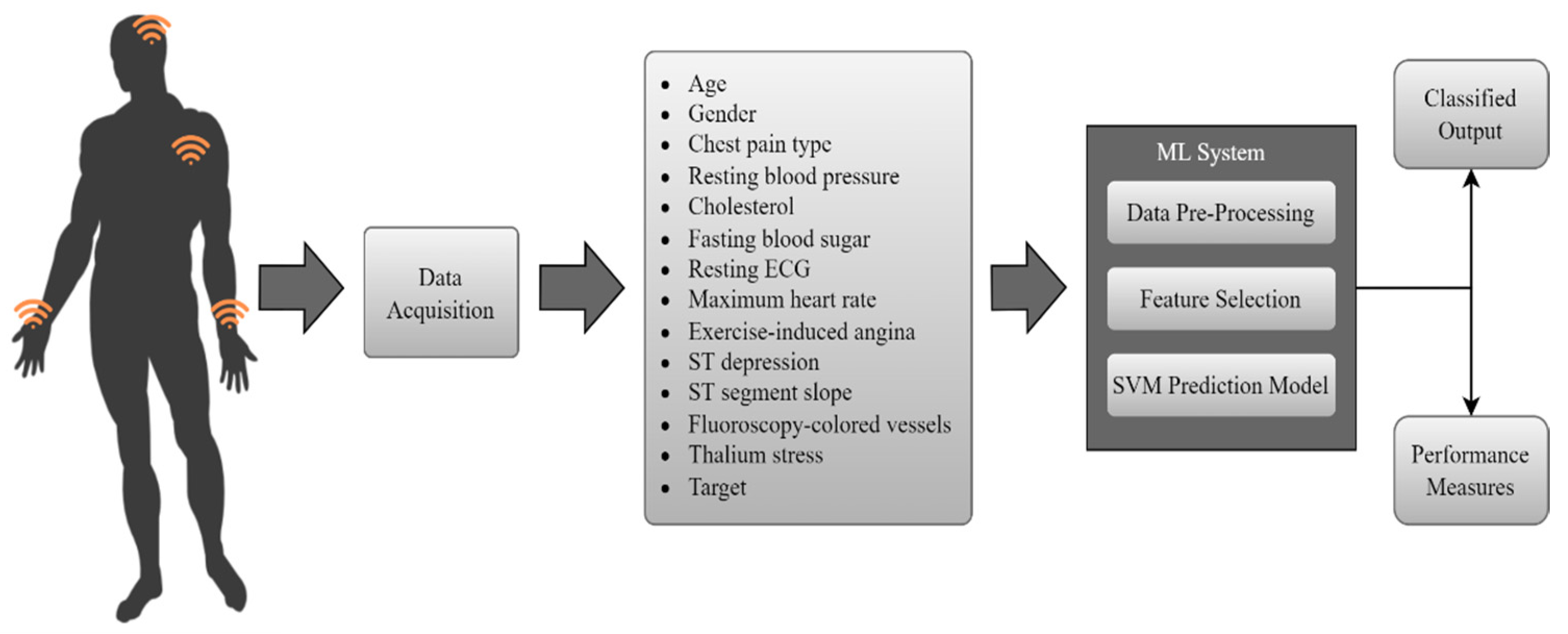
2. Methodology
2.1. Support Vector Machine (SVM) Model
2.1.1. Heart Disease Dataset Features
2.1.2. Data Pre-Processing
2.2. Enhanced SVM Model with Feature Selection
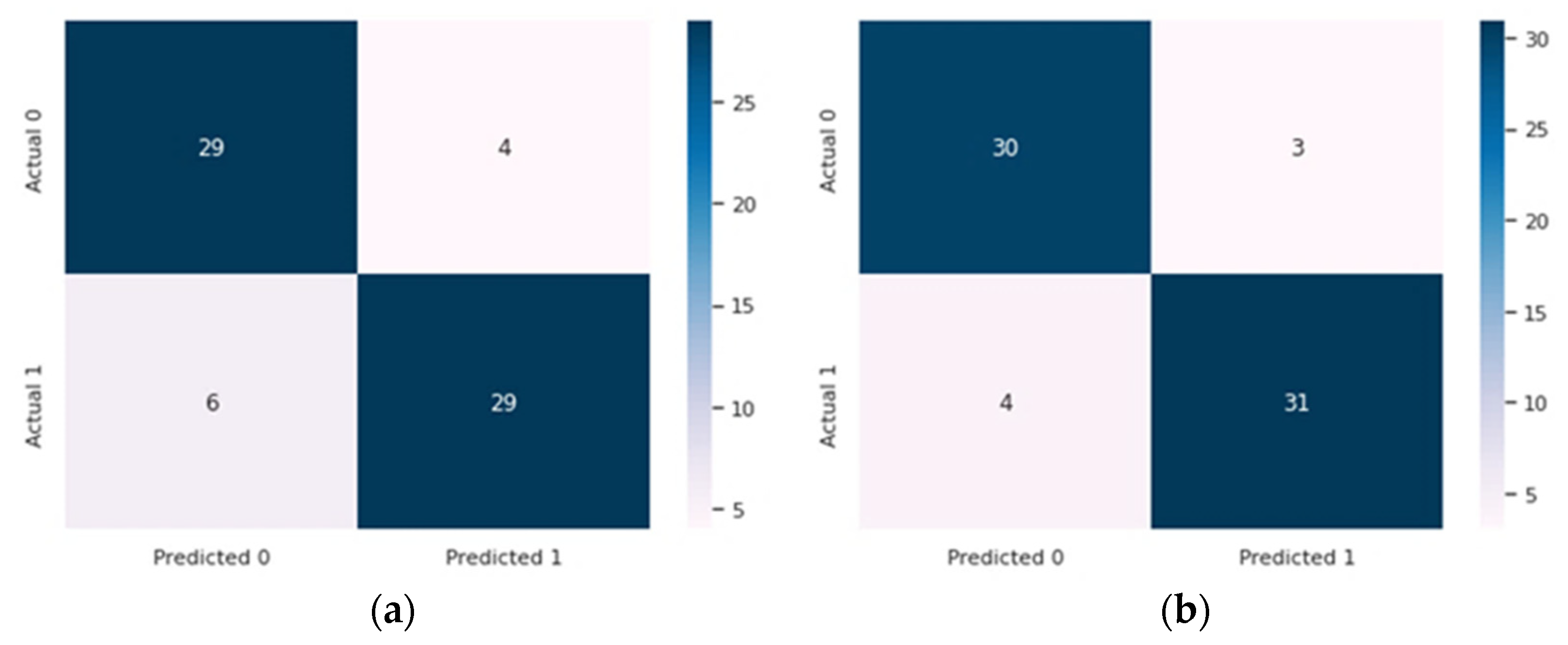
3. Results and Discussion
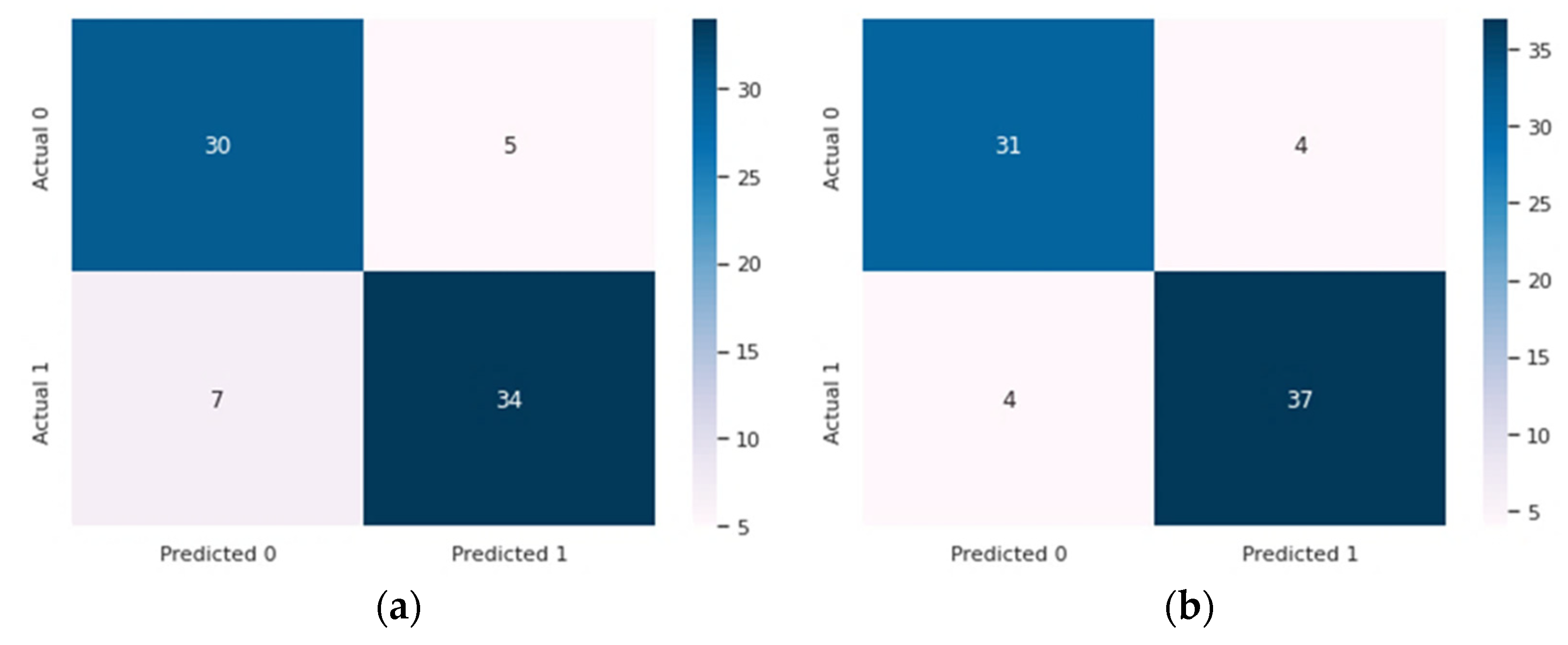
- Accuracy (Acc): defined as the proportion of total positive instances of the model to the total number of instances, as shown in Equation (6).
- Specificity (Spe): the percentage of true negatives out of all healthy individuals, calculated by Equation (7). It was used to determine the degree of the attribute to appropriately classify the individuals without diseases.
- Sensitivity (Sen): used to determine the degree of the attribute in order to appropriately classify the individuals who have diseases, as illustrated in Equation (8);
- F1-score: defined as the harmonic mean of the specificity and sensitivity. It can be computed as shown in Equation (9);
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Das, S.; Sharma, R.; Gourisaria, M.; Rautaray, S.; Pandey, M. Heart disease detection using core machine learning and deep learning techniques: A comparative study. Int. J. Emerg. Technol. 2020, 11, 531–538. [Google Scholar]
- Hasan, T.T.; Jasim, M.H.; Hashim, I.A. FPGA Design and Hardware Implementation of Heart Disease Diagnosis System Based on NVG-RAM Classifier. In Proceedings of the 2018 Third Scientific Conference of Electrical Engineering (SCEE), Baghdad, Iraq, 19–20 December 2018; pp. 33–38. [Google Scholar] [CrossRef]
- Rahman, A.U.; Saeed, M.; Mohammed, M.A.; Jaber, M.M.; Garcia-Zapirain, B. A novel fuzzy parameterized fuzzy hypersoft set and riesz summability approach based decision support system for diagnosis of heart diseases. Diagnostics 2022, 12, 1546. [Google Scholar] [CrossRef] [PubMed]
- Javid, I.; Khalaf, A.; Ghazali, R. Enhanced accuracy of heart disease prediction using machine learning and recurrent neural networks ensemble majority voting method. Int. J. Adv. Comput. Sci. Appl. 2020, 11, 540–551. [Google Scholar] [CrossRef]
- Muhsen, D.K.; Khairi, T.W.A.; Alhamza, N.I.A. Machine Learning System Using Modified Random Forest Algorithm. In Intelligent Systems and Networks, Singapore; Tran, D.-T., Jeon, G., Nguyen, T.D.L., Lu, J., Xuan, T.-D., Eds.; Springer: Singapore, 2021; pp. 508–515. [Google Scholar]
- Wah, T.Y.; Mohammed, M.A.; Iqbal, U.; Kadry, S.; Majumdar, A.; Thinnukool, O. Novel DERMA fusion technique for ECG heartbeat classification. Life 2022, 12, 842. [Google Scholar]
- Mohammed, M.A.; Abdulkareem, K.H.; Al-Waisy, A.S.; Mostafa, S.A.; Al-Fahdawi, S.; Dinar, A.M.; Alhakami, W.; Abdullah, B.A.Z.; Al-Mhiqani, M.N.; Alhakami, H.; et al. Benchmarking methodology for selection of optimal COVID-19 diagnostic model based on entropy and TOPSIS methods. IEEE Access 2020, 8, 99115–99131. [Google Scholar] [CrossRef]
- Dinar, A.M.; Zain, A.M.; Salehuddin, F. Utilizing of CMOS ISFET sensors in DNA applications detection: A systematic review. J. Adv. Res. Dyn. Control Syst. 2018, 10, 569–583. [Google Scholar]
- Soni, M.; Gomathi, S.; Kumar, P.; Churi, P.P.; Mohammed, M.A.; Salman, A.O. Hybridizing Convolutional Neural Network for Classification of Lung Diseases. Int. J. Swarm Intell. Res. (IJSIR) 2022, 13, 1–15. [Google Scholar] [CrossRef]
- Nasser, A.R.; Hasan, A.M.; Humaidi, A.J.; Alkhayyat, A.; Alzubaidi, L.; Fadhel, M.A.; Santamaría, J.; Duan, Y. IoT and Cloud Computing in Health-Care: A New Wearable Device and Cloud-Based Deep Learning Algorithm for Monitoring of Diabetes. Electronics 2021, 10, 2719. Available online: https://www.mdpi.com/2079-9292/10/21/2719 (accessed on 1 May 2022).
- Diwakar, M.; Tripathi, A.; Joshi, K.; Memoria, M.; Singh, P. Latest trends on heart disease prediction using machine learning and image fusion. Mater. Today Proc. 2021, 37, 3213–3218. [Google Scholar] [CrossRef]
- Rahman, A.U.; Saeed, M.; Mohammed, M.A.; Krishnamoorthy, S.; Kadry, S.; Eid, F. An Integrated Algorithmic MADM Approach for Heart Diseases’ Diagnosis Based on Neutrosophic Hypersoft Set with Possibility Degree-Based Setting. Life 2022, 12, 729. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Root, M.M. Building prediction models for coronary heart disease by synthesizing multiple longitudinal research findings. Eur. J. Prev. Cardiol. 2005, 12, 459–464. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Deo, R.C. Machine learning in medicine. Circulation 2015, 132, 1920–1930. [Google Scholar] [CrossRef] [PubMed]
- Mythili, T.; Mukherji, D.; Padalia, N.; Naidu, A. A heart disease prediction model using SVM-decision trees-logistic regression (SDL). Int. J. Comput. Appl. 2013, 68, 0975–8887. [Google Scholar]
- Elhoseny, M.; Mohammed, M.A.; Mostafa, S.A.; Abdulkareem, K.H.; Maashi, M.S.; Garcia-Zapirain, B.; Mutlag, A.A.; Maashi, M.S. A new multi-agent feature wrapper machine learning approach for heart disease diagnosis. Comput. Mater. Contin 2021, 67, 51–71. [Google Scholar] [CrossRef]
- Detrano, R.; Janosi, A.; Steinbrunn, W.; Pfisterer, M.; Schmid, J.J.; Sandhu, S.; Guppy, K.H.; Lee, S.; Froelicher, V. International application of a new probability algorithm for the diagnosis of coronary artery disease. Am. J. Cardiol. 1989, 64, 304–310. [Google Scholar] [CrossRef]
- Gennari, J.H.; Langley, P.; Fisher, D. Models of incremental concept formation. Artif. Intell. 1989, 40, 11–61. [Google Scholar] [CrossRef]
- Janosi, A.; Steinbrunn, W.; Pfisterer, M.; Detrano, R. UCI Machine Learning Repository: Heart Disease Dataset [Online]. Available online: https://archive-beta.ics.uci.edu/ml/datasets/heart+disease (accessed on 1 March 2022).
- Machine Learning Repository: Statlog (Heart) [Online]. Available online: http://archive.ics.uci.edu/ml/datasets/Statlog+%28Heart%29 (accessed on 1 March 2022).
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Sajja, T.K.; Kalluri, H.K. A Deep Learning Method for Prediction of Cardiovascular Disease Using Convolutional Neural Network. Rev. D’intelligence Artif. 2020, 34, 601–606. [Google Scholar] [CrossRef]
- Guo, C.; Zhang, J.; Liu, Y.; Xie, Y.; Han, Z.; Yu, J. Recursion enhanced random forest with an improved linear model (RERF-ILM) for heart disease detection on the internet of medical things platform. IEEE Access 2020, 8, 59247–59256. [Google Scholar] [CrossRef]
- Ali, S.A.; Raza, B.; Malik, A.K.; Shahid, A.R.; Faheem, M.; Alquhayz, H.; Kumar, Y.J. An optimally configured and improved deep belief network (OCI-DBN) approach for heart disease prediction based on Ruzzo–Tompa and stacked genetic algorithm. IEEE Access 2020, 8, 65947–65958. [Google Scholar] [CrossRef]
- Vijayashree, J.; Parveen Sultana, H. Heart disease classification using hybridized Ruzzo-Tompa memetic based deep trained Neocognitron neural network. Health Technol. 2020, 10, 207–216. [Google Scholar] [CrossRef]
- Bharti, R.; Khamparia, A.; Shabaz, M.; Dhiman, G.; Pande, S.; Singh, P. Prediction of Heart Disease Using a Combination of Machine Learning and Deep Learning. Comput. Intell. Neurosci. 2021, 2021, 8387680. [Google Scholar] [CrossRef] [PubMed]
- Ali, L.; Rahman, A.; Khan, A.; Zhou, M.; Javeed, A.; Khan, J.A. An Automated Diagnostic System for Heart Disease Prediction Based on χ2 Statistical Model and Optimally Configured Deep Neural Network. IEEE Access 2019, 7, 34938–34945. [Google Scholar] [CrossRef]
- Aliyar Vellameeran, F.; Brindha, T. A new variant of deep belief network assisted with optimal feature selection for heart disease diagnosis using IoT wearable medical devices. Comput. Methods Biomech. Biomed. Eng. 2021, 25, 387–411. [Google Scholar] [CrossRef]
- Yue, W.; Wang, Z.; Chen, H.; Payne, A.; Liu, X. Machine Learning with Applications in Breast Cancer Diagnosis and Prognosis. Designs 2018, 2, 13. [Google Scholar] [CrossRef]
- Ali, L.; Zhu, C.; Zhou, M.; Liu, Y. Early diagnosis of Parkinson’s disease from multiple voice recordings by simultaneous sample and feature selection. Expert Syst. Appl. 2019, 137, 22–28. [Google Scholar] [CrossRef]
- Liu, H.; Setiono, R. Chi2: Feature selection and discretization of numeric attributes. In Proceedings of the 7th IEEE International Conference on Tools with Artificial Intelligence, Herndon, VA, USA, 5–8 November 1995; pp. 388–391. [Google Scholar]
- Maldonado, S.; Pérez, J.; Weber, R.; Labbé, M. Feature selection for support vector machines via mixed integer linear programming. Inf. Sci. 2014, 279, 163–175. [Google Scholar] [CrossRef]
- Shorewala, V. Early detection of coronary heart disease using ensemble techniques. Inform. Med. Unlocked 2021, 26, 100655. [Google Scholar] [CrossRef]
- Ali, F.; El-Sappagh, S.; Islam, S.R.; Kwak, D.; Ali, A.; Imran, M.; Kwak, K.S. A smart healthcare monitoring system for heart disease prediction based on ensemble deep learning and feature fusion. Inf. Fusion 2020, 63, 208–222. [Google Scholar] [CrossRef]
- Latha, C.B.C.; Jeeva, S.C. Improving the accuracy of prediction of heart disease risk based on ensemble classification techniques. Inform. Med. Unlocked 2019, 16, 100203. [Google Scholar] [CrossRef]
- Haq, A.U.; Li, J.P.; Memon, M.H.; Nazir, S.; Sun, R. A Hybrid Intelligent System Framework for the Prediction of Heart Disease Using Machine Learning Algorithms. Mob. Inf. Syst. 2018, 2018, 3860146. [Google Scholar] [CrossRef]
- Vijayashree, J.; Sultana, H.P. A Machine Learning Framework for Feature Selection in Heart Disease Classification Using Improved Particle Swarm Optimization with Support Vector Machine Classifier. Program. Comput. Softw. 2019, 44, 388–397. [Google Scholar] [CrossRef]
- Tuli, S.; Basumatary, N.; Gill, S.S.; Kahani, M.; Arya, R.C.; Wander, G.S.; Buyya, R. HealthFog: An ensemble deep learning based Smart Healthcare System for Automatic Diagnosis of Heart Diseases in integrated IoT and fog computing environments. Future Gener. Comput. Syst. 2020, 104, 187–200. [Google Scholar] [CrossRef]

| Name | Type | Description |
|---|---|---|
| Age | Numeric | Age in years |
| Sex | Categorial | 0 = Female or 1 = male |
| Cp | Categorial | Type of Chest pain (1 = typical angina, 2 = atypical angina, 3 = non anginal pain, 4 = asymptomatic) |
| Trestbps | Numeric | Resting blood pressure (mm hg) |
| Chol | Numeric | Serum cholesterol (mg/dL) |
| Fbs | Categorial | Fasting blood sugar > 120 mg/dL (0 = false, 1 = true) |
| Restecg | Categorial | Resting electrocardiography results (0 = normal, 1 = ST-T wave abnormality, 2 = probable or definite left ventricular hypertrophy) |
| Thalach | Numeric | Maximum heart rate achieved during thalium stress test |
| Exang | Categorial | Exercise-induced angina (1 = yes, 0 = no) |
| Oldpeak | Numeric | St depression induced by exercise relative to rest |
| Slope | Categorial | Slope of peak exercise ST segment (1 = upsloping, 2 = flat, 3 = downsloping) |
| Ca | Categorial | Number of significant vessels colored by fluoroscopy |
| Thal | Categorial | Thalium stress test result (3 = normal, 6 = fixed, 7 = reversible defect) |
| Num | Categorial | Heart disease status (0 = < 50% diameter narrowing, 1 = > 50% diameter narrowing) |
| Positive Class | Negative Class | Total | |
|---|---|---|---|
| Feature Xi occurs | α | b | α + b = m |
| Feature Xi does not occur | λ | y | λ + y = t − m |
| Total | α + λ = p | b + y = t − p | t |
| Dataset | Total Records | Total Features | Acc (%) | Spe (%) | Sen (%) | F1 (%) |
|---|---|---|---|---|---|---|
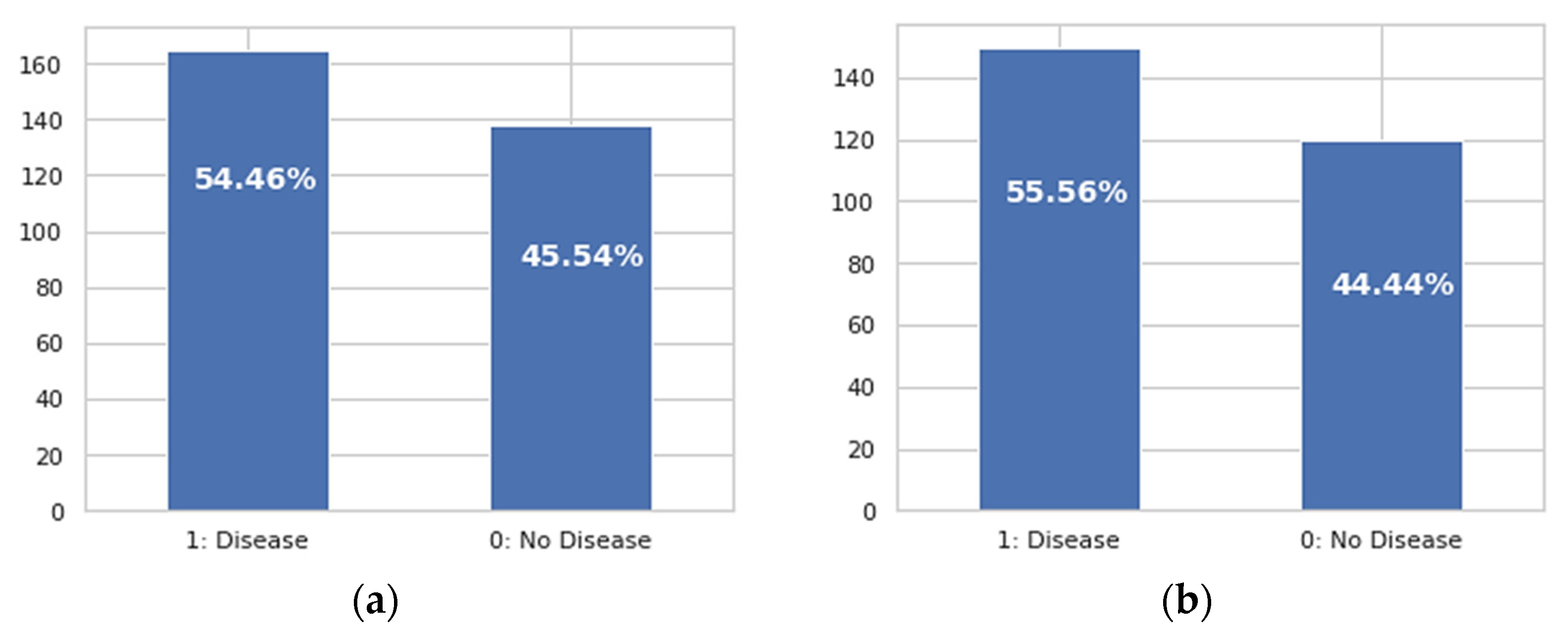
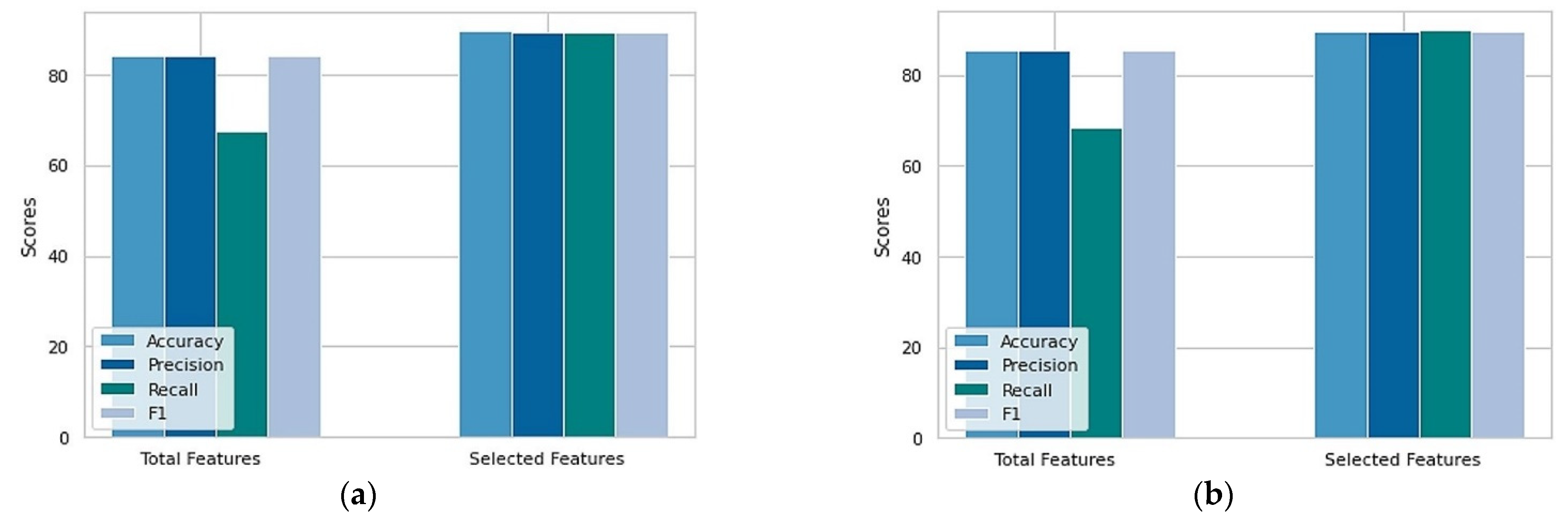
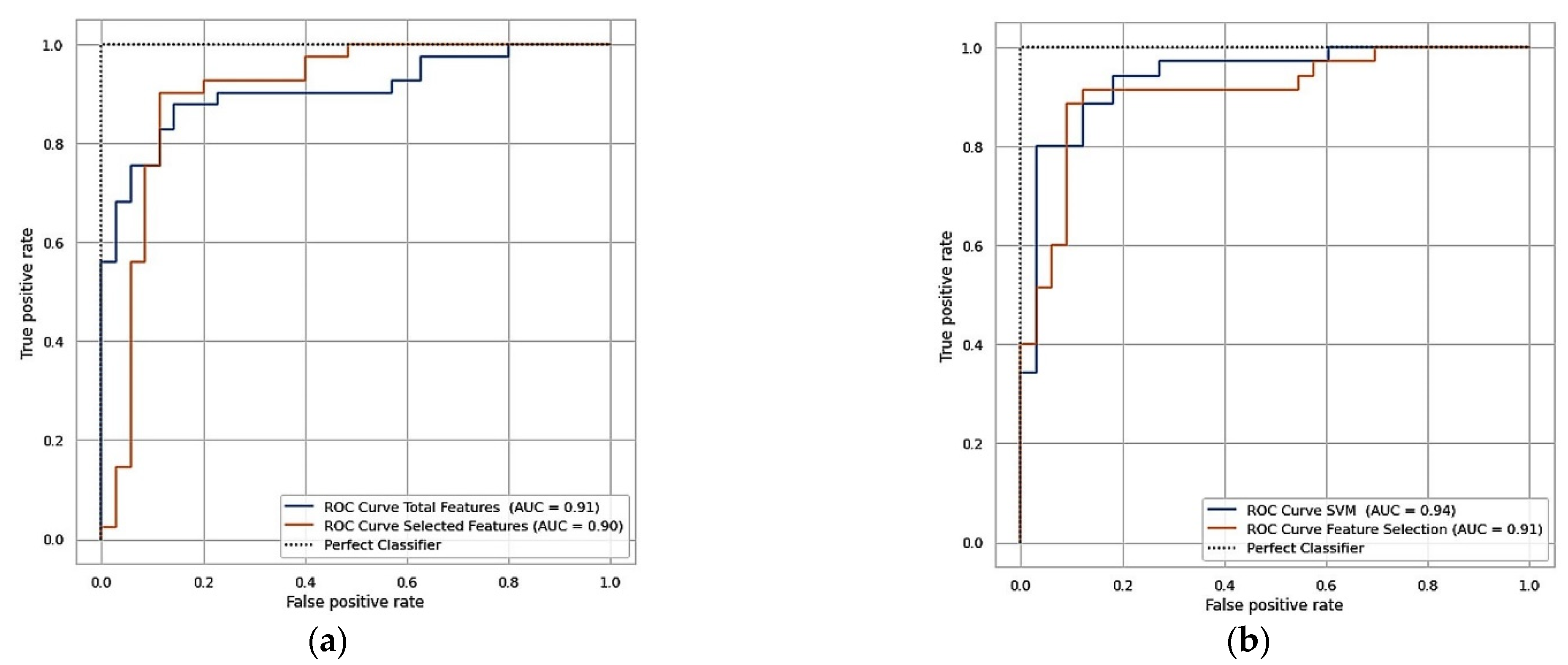
| Cleveland | 303 | 14 | 84.21 | 84.13 | 67.45 | 84.16 |
| Statlog | 270 | 14 | 85.29 | 85.36 | 68.29 | 85.29 |
| Dataset | Total Records | Total Features | Acc (%) | Spe (%) | Sen (%) | F1 (%) |
|---|---|---|---|---|---|---|
| Cleveland | 303 | 6 | 89.47 | 89.40 | 89.40 | 89.40 |
| Statlog | 270 | 6 | 89.70 | 89.70 | 89.74 | 89.70 |
| Study | Feature Selection Method | Dataset | Classifier | Total Features | Acc (%) | Selected Features | Acc (%) |
|---|---|---|---|---|---|---|---|
| [33] | Lasso 1 | Cleveland | Ensemble | 13 | - | 8 | 75.1 |
| [34] | Information Gain | Cleveland | Ensemble | 27 | 72.2 | 16 | 83.5 |
| [26] | Lasso | Cleveland | SVM | 13 | 84.09 | - | 84.26 |
| [35] | Randomly Generated Feature Set | Cleveland | Ensemble | 13 | - | 9 | 85.48 |
| [24] | Ruzzo-Tompa | Cleveland | ANN 2 | 13 | - | 7 | 86.20 |
| [36] | Lasso | Cleveland | SVM (RBF) | 13 | 86 | 6 | 88 |
| [37] | PSO-SVM 3 | Cleveland | SVM | 13 | 79.35 | 6 | 88.22 |
| [28] | PS-GWO | Statlog and Cleveland | DBN 4 | 13 | - | - | 88.8 |
| [38] | PCA 5 | Cleveland | DL 6 | 13 | - | - | 89 |
| Proposed | Chi-squared | SVM | 13 | 85.29 | 6 | 89.47 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sarra, R.R.; Dinar, A.M.; Mohammed, M.A.; Abdulkareem, K.H. Enhanced Heart Disease Prediction Based on Machine Learning and χ2 Statistical Optimal Feature Selection Model. Designs 2022, 6, 87. https://doi.org/10.3390/designs6050087
Sarra RR, Dinar AM, Mohammed MA, Abdulkareem KH. Enhanced Heart Disease Prediction Based on Machine Learning and χ2 Statistical Optimal Feature Selection Model. Designs. 2022; 6(5):87. https://doi.org/10.3390/designs6050087
Chicago/Turabian StyleSarra, Raniya R., Ahmed M. Dinar, Mazin Abed Mohammed, and Karrar Hameed Abdulkareem. 2022. "Enhanced Heart Disease Prediction Based on Machine Learning and χ2 Statistical Optimal Feature Selection Model" Designs 6, no. 5: 87. https://doi.org/10.3390/designs6050087
APA StyleSarra, R. R., Dinar, A. M., Mohammed, M. A., & Abdulkareem, K. H. (2022). Enhanced Heart Disease Prediction Based on Machine Learning and χ2 Statistical Optimal Feature Selection Model. Designs, 6(5), 87. https://doi.org/10.3390/designs6050087










