Identification of the Metallocarboxypeptidase M14 Gene Family in Lysiosquillina maculata, Odontodactylus scyllarus and Oratosquilla oratoria Based on the Full-Length Transcriptome
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection, RNA Extraction, and FL Transcriptome Sequencing
2.2. FL Transcriptome Data Processing
2.3. The Identification MCP M14 Gene Family of L. maculata, O. scyllarus, and O. oratoria
2.4. Characterization of the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
3. Results
3.1. FL Transcriptome Information of L. maculata and O. scyllarus
3.2. Physicochemical Properties of Proteins Belonging to the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
3.3. Phylogenetic Analysis of the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
3.4. Conservation Motif Analysis of the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
3.5. Multiple Sequence Alignment of the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
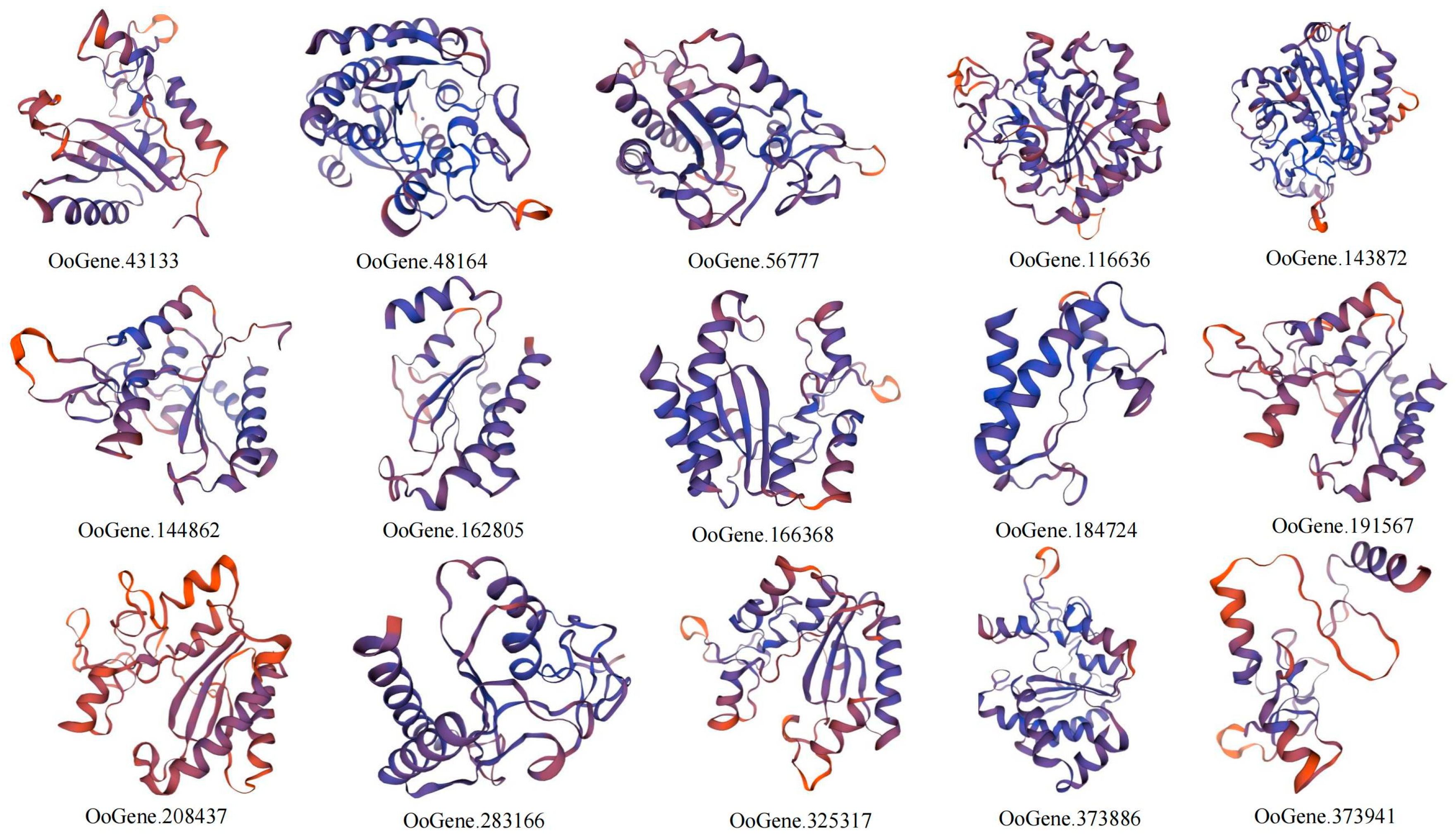
3.6. Prediction of Secondary and Tertiary Protein Structures of the MCP M14 Gene Family in L. maculata, O. scyllarus, and O. oratoria
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahyong, S.T. Raysquilla Manningi, a New Genus and Species of Stomatopod from the Australian Northwest Shelf. J. Crustac. Biol. 2000, 20, 37–41. [Google Scholar] [CrossRef]
- Sha, Z.; Wang, Y.; Cheng, J. Application of mitochondrial COI-based DNA barcoding for the identification of stomatopod species (Crustacea, Stomatopoda) in the China seas. J. Fish. Sci. China 2018, 25, 858. [Google Scholar] [CrossRef]
- Cheng, J.; Wang, Y.L.; Sha, Z.L. Progress on the systematics of Stomatopod (Crustacea: Malacostraca). Mar. Sci. 2015, 39, 173–177. [Google Scholar] [CrossRef]
- Manning, R.B. Stomatopod crustacea of Vietnam: The legacy of Raoul Serene. Crustac. Res. 1995, 4, 1–339. [Google Scholar] [CrossRef]
- Ahyong, S.T.; Harling, C. The phylogeny of the stomatopod Crustacea. Aust. J. Zool. 2000, 48, 607–642. [Google Scholar] [CrossRef]
- Ahyong, S.T. Phylogenetic Analysis of the Stomatopoda (Malacostraca). J. Crustac. Biol. 1997, 17, 695–715. [Google Scholar] [CrossRef][Green Version]
- Marshall, N.J.; Land, M.F.; King, C.A.; Cronin, T.W. The compound eyes of mantis shrimps (Crustacea, Hoplocarida, Stomatopoda). II. Colour pigments in the eyes of stomatopod crustaceans: Polychromatic vision by serial and lateral filtering. Philos. Trans. R. Soc. B Biol. Sci. 1991, 334, 57–84. [Google Scholar] [CrossRef]
- Yaraghi, N.A.; Guarín-Zapata, N.; Grunenfelder, L.K.; Hintsala, E.; Bhowmick, S.; Hiller, J.M.; Betts, M.; Principe, E.L.; Jung, J.Y.; Sheppard, L.; et al. A Sinusoidally Architected Helicoidal Biocomposite. Adv. Mater. 2016, 28, 6835–6844. [Google Scholar] [CrossRef]
- Reaka, M.L. Molting in stomatopod crustaceans. I. Stages of the molt cycle, setagenesis, and morphology. J. Morphol. 1975, 146, 55–80. [Google Scholar] [CrossRef]
- Amini, S.; Masic, A.; Bertinetti, L.; Teguh, J.S.; Herrin, J.S.; Zhu, X.; Su, H.; Miserez, A. Textured fluorapatite bonded to calcium sulphate strengthen stomatopod raptorial appendages. Nat. Commun. 2014, 5, 3187. [Google Scholar] [CrossRef]
- Yaraghi, N.A.; Trikanad, A.A.; Restrepo, D.; Huang, W.; Rivera, J.; Herrera, S.; Zhernenkov, M.; Parkinson, D.Y.; Caldwell, R.L.; Zavattieri, P.D.; et al. The Stomatopod Telson: Convergent Evolution in the Development of a Biological Shield. Adv. Funct. Mater. 2019, 29, 13. [Google Scholar] [CrossRef]
- Patek, S.N.; Rosario, M.V.; Taylor, J.R.A. Comparative spring mechanics in mantis shrimp. J. Exp. Biol. 2012, 216, 1317–1329. [Google Scholar] [CrossRef]
- Devries, M.S.; Murphy, E.A.K.; Patek, S.N. Strike mechanics of an ambush predator: The spearing mantis shrimp. J. Exp. Biol. 2012, 215, 4374–4384. [Google Scholar] [CrossRef]
- Kim, S.; Cho, Y.S.; Kim, H.-M.; Chung, O.; Kim, H.; Jho, S.; Seomun, H.; Kim, J.; Bang, W.Y.; Kim, C.; et al. Comparison of carnivore, omnivore, and herbivore mammalian genomes with a new leopard assembly. Genome Biol. 2016, 17, 211. [Google Scholar] [CrossRef]
- DeVries, M.S. The role of feeding morphology and competition in governing the diet breadth of sympatric stomatopod crustaceans. Biol. Lett. 2017, 13, 20170055. [Google Scholar] [CrossRef]
- Jiao, H.W. Molecular Evolution of Taste and Trehalase Genes in Bats and Its Significance on the Dietary Adaptation; Wuhan University: Wuhan, China, 2019. [Google Scholar]
- Wu, J.W. Molecular Evolution of Lineage-Specifie Traits, Dietary Shift and Visual Function in Bats; Wuhan University: Wuhan, China, 2018. [Google Scholar]
- Chang, Y. Research on the Factors Influencing Dietary Niche Expansion and the Adaptive Evolution of Skull in Piscivorous Bats; Northeast Normal University: Changchun, China, 2020. [Google Scholar] [CrossRef]
- Youngblut, N.D.; Reischer, G.H.; Walters, W.; Schuster, N.; Farnleitner, A.H. Host diet and evolutionary history explain different aspects of gut microbiome diversity among vertebrate clades. Nat. Commun. 2019, 10, 2200. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X. Identification of M14 Family Metal Carboxypeptidases in Antheraea pernyi and Transcriptional Response of Digestive Enzymes Triggered by Starvation; Nanyang Normal University: Nanyang, China, 2022. [Google Scholar] [CrossRef]
- Rossano, R.; Larocca, M.; Lamaina, A.; Viggiani, S.; Riccio, P. The hepatopancreas enzymes of the crustaceans Munida and their potential application in cheese biotechnology. LWT-Food Sci. Technol. 2011, 44, 173–180. [Google Scholar] [CrossRef]
- Fang, L.S.; Lee, B.N. Ontogenic change of digestive enzymes in Penaeus monodon. Comp. Biochem. Physiol. Part B Comp. Biochem. 1992, 103, 1033–1037. [Google Scholar] [CrossRef]
- Tsai, I.H.; Chuano, K.L.; Chuang, J.L. Chymotrypsins in digestive tracts of crustacean decapods (Shrimps). Comp. Biochem. Physiol. B Comp. Biochem. 1986, 85, 235–239. [Google Scholar] [CrossRef]
- Serrano, A.E. Ontogenetic Changes in the Activity of Chymotrypsin and Carboxypeptidases A and B in Mud Crab, Scylla serrata. Isr. J. Aquac. Bamidgeh 2013, 65, 1–6. [Google Scholar]
- Andrés, M.; Gisbert, E.; Díaz, M.; Moyano, F.J.; Rotllant, G. Ontogenetic changes in digestive enzymatic capacities of the spider crab, Maja brachydactyla (Decapoda: Majidae). J. Exp. Mar. Biol. Ecol. 2010, 389, 75–84. [Google Scholar] [CrossRef]
- Vogt, G. Synthesis of digestive enzymes, food processing, and nutrient absorption in decapod crustaceans: A comparison to the mammalian model of digestion. Zoology 2021, 147, 125945. [Google Scholar] [CrossRef]
- Le Moullac, G.; Roy, P.; Van Wormhoudt, A. Effects of trophic prophylatic factors on some digestive enzymatic activities of Penaeus vannamei larvae. In Memorias del Primer Congreso Ecuatoriano de Acuicultura, San Pedro de Manglaralto, Ecuador; CENAIM: Guayaquil, Ecuador, 1992; pp. 81–86. [Google Scholar]
- Long, Q.; Liu, J.; Sun, Y.; Yang, Z.; Tang, B.; Cheng, Y. The Effect of Food Deprivation on Foraging Behavior and Digestive and Metabolic Capacities of the Chinese Mitten Crab, Eriocheir sinensis. Fishes 2023, 8, 47. [Google Scholar] [CrossRef]
- Navarrete-Del-Toro, M.A.; García-Carreño, F.L.; Hernández-Cortés, P.; Molnár, T.; Gráf, L. Biochemical characterisation of chymotrypsin from the midgut gland of yellowleg shrimp, Penaeus californiensis. Food Chem. 2015, 173, 147–155. [Google Scholar] [CrossRef]
- Jones, D.; Kumlu, M.; Le Vay, L.; Fletcher, D. The digestive physiology of herbivorous, omnivorous and carnivorous crustacean larvae: A review. Aquaculture 1997, 155, 285–295. [Google Scholar] [CrossRef]
- Rivera-Pérez, C.; García-Carreño, F.L.; Saborowski, R. Purification and Biochemical Characterization of Digestive Lipase in Whiteleg Shrimp. Mar. Biotechnol. 2010, 13, 284–295. [Google Scholar] [CrossRef]
- Rodríguez-Viera, L.; Alpízar-Pedraza, D.; Mancera, J.M.; Perera, E. Toward a More Comprehensive View of α-Amylase across Decapods Crustaceans. Biology 2021, 10, 947. [Google Scholar] [CrossRef]
- Tsuji, A.; Sato, S.; Kondo, A.; Tominaga, K.; Yuasa, K. Purification and characterization of cellulase from North Pacific krill (Euphausia pacifica). Analysis of cleavage specificity of the enzyme. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2012, 163, 324–333. [Google Scholar] [CrossRef]
- Proespraiwong, P.; Tassanakajon, A.; Rimphanitchayakit, V. Chitinases from the black tiger shrimp Penaeus monodon: Phylogenetics, expression and activities. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2010, 156, 86–96. [Google Scholar] [CrossRef]
- Maugle, P.D.; Deshimaru, O.; Katayama, T.; Simpson, K.L. Digestive enzymes of the shrimp Penaeus japonicus-I. Characteristics of amylase and protease of the shrimp Penaeus japonicus. Nippon. Suisan Gakkaishi 1982, 48, 1753–1757. [Google Scholar] [CrossRef][Green Version]
- Pan, L.Q.; Liu, H.Y.; Xiao, G.Q. A review on digestive enzyme of crustacean larvae. J. Fish. Sci. China 2006, 13, 492–501. [Google Scholar] [CrossRef]
- Skidgel, R.A.; Erdös, E.G. Cellular carboxypeptidases. Immunol. Rev. 1998, 161, 129–141. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Sharma, A.; Sharma, R.; Gakhar, S. Identification, characterization and analysis of expression of gene encoding carboxypeptidase A in Anopheles culicifacies A (Diptera: Culicidae). Acta Trop. 2014, 139, 123–130. [Google Scholar] [CrossRef] [PubMed]
- Fricker, L.D. Activation and membrane binding of carboxypeptidase E. J. Cell. Biochem. 1988, 38, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Isoe, J.; Zamora, J.; Miesfeld, R.L. Molecular analysis of the Aedes aegypti carboxypeptidase gene family. Insect Biochem. Mol. Biol. 2009, 39, 68–73. [Google Scholar] [CrossRef]
- Zhu, Y.P.; Han, M.J.; Zha, X.L.; Chen, Y.; Shen, Y.H. Identification and Expression Analysis of Metallocarboxypeptidase Gene Family in Silkworm, Bombyx mori. Acta Sericologica Sin. 2016, 42, 393–403. [Google Scholar] [CrossRef]
- Ferreira, C.; Rebola, K.G.O.; Cardoso, C.; Bragatto, I.; Ribeiro, A.F.; Terra, W.R. Insect midgut carboxypeptidases with emphasis on S10 hemipteran and M14 lepidopteran carboxypeptidases. Insect Mol. Biol. 2014, 24, 222–239. [Google Scholar] [CrossRef]
- Rodriguez de la Vega, M.; Sevilla, R.G.; Hermoso, A.; Lorenzo, J.; Tanco, S.; Diez, A.; Fricker, L.D.; Bautista, J.M.; Avilés, F.X. Nna1-like proteins are active metallocarboxypeptidases of a new and diverse M14 subfamily. FASEB J. 2007, 21, 851–865. [Google Scholar] [CrossRef]
- Rawlings, N.D.; Barrett, A.J. Evolutionary families of metallopeptidases. Methods Enzymol. 1995, 248, 183–228. [Google Scholar] [CrossRef]
- Bown, D.P.; Gatehouse, J.A. Characterization of a digestive carboxypeptidase from the insect pest corn earworm (Helicoverpa armigera) with novel specificity towards C-terminal glutamate residues. Eur. J. Biochem. 2004, 271, 2000–2011. [Google Scholar] [CrossRef]
- Edwards, M.J.; Lemos, F.J.A.; Donnelly-Doman, M.; Jacobs-Lorena, M. Rapid Induction by a Blood Meal of a Carboxypeptidase Gene in the Gut of the Mosquito Anopheles gambiae. Insect Biochem. Mol. Biol. 1997, 27, 1063–1072. [Google Scholar] [CrossRef] [PubMed]
- Ramos, A.; Mahowald, A.; Jacobs-Lorena, M. Gut-specific genes from the black fly Simulium vittatum encoding trypsin-like and carboxypeptidase-like proteins. Insect Mol. Biol. 1993, 1, 149–163. [Google Scholar] [CrossRef]
- Wang, P.; Li, G.; Kain, W. Characterization and cDNA cloning of midgut carboxypeptidases from Trichoplusia ni. Insect Biochem. Mol. Biol. 2004, 34, 831–843. [Google Scholar] [CrossRef]
- Edwards, M.J.; Moskalyk, L.A.; Donelly-Doman, M.; Vlaskova, M.; Noriega, F.G.; Walker, V.K.; Jacobs-Lorena, M. Characterization of a carboxypeptidase A gene from the mosquito, Aedes aegypti. Insect Mol. Biol. 2000, 9, 33–38. [Google Scholar] [CrossRef]
- Zhang, X.; Zhao, P.; Li, S.; Ma, S.; Du, J.; Liang, S.; Yang, X.; Yao, L.; Duan, J. Genome-Wide Identification of M14 Family Metal Carboxypeptidases in Antheraea pernyi (Lepidoptera: Saturniidae). J. Econ. Èntomol. 2022, 115, 1285–1293. [Google Scholar] [CrossRef] [PubMed]
- Shin, S.C.; Ahn, D.H.; Kim, S.J.; Lee, H.; Oh, T.J.; Lee, J.E.; Park, H. Advantages of Single-Molecule Real-Time Sequencing in High-GC Content Genomes. PLoS ONE 2013, 8, e68824. [Google Scholar] [CrossRef] [PubMed]
- Lou, F.R.; Zhou, J.L.; Schunter, C.; Wang, L.; Tang, Y.Z.; Han, Z.Q.; Kang, B. How Oratosquilla oratoria compound eye response to the polarization of light: In the perspective of vision genes and related proteins. Int. J. Biol. Macromol. 2024, 259, 129053. [Google Scholar] [CrossRef]
- Gordon, S.P.; Tseng, E.; Salamov, A.; Zhang, J.; Meng, X.; Zhao, Z.; Kang, D.; Underwood, J.; Grigoriev, I.V.; Figueroa, M.; et al. Widespread Polycistronic Transcripts in Fungi Revealed by Single-Molecule mRNA Sequencing. PLoS ONE 2015, 10, e0132628. [Google Scholar] [CrossRef]
- Guizard, S.; Miedzinska, K.; Smith, J.; Smith, J.; I Kuo, R.; Davey, M.; Archibald, A.; Watson, M. nf-core/isoseq: Simple gene and isoform annotation with PacBio Iso-Seq long-read sequencing. Bioinformatics 2023, 39, btad150. [Google Scholar] [CrossRef]
- Li, W.Z.; Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 2006, 22, 1658–1659. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, T.J.; Eddy, S.R. nhmmer: DNA homology search with profile HMMs. Bioinformatics 2013, 29, 2487–2489. [Google Scholar] [CrossRef] [PubMed]
- Ambrose, D.P.; Lenin, E.A.; Kiruba, D.A. Intrageneric phylogenetics based on mitochondrial DNA variation among fifteen harpactorine assassin bugs with four ecotypes and three morphs (Hemiptera: Reduviidae: Harpactorinae). Zootaxa 2014, 3779, 540–550. [Google Scholar] [CrossRef]
- Goud, T.S.; Upadhyay, R.C.; Onteru, S.K.; Pichili, V.B.R.; Chadipiralla, K. Identification and sequence characterization of melanocortin 1 receptor gene (MC1R) in Bos indicus versus (Bos taurus × Bos indicus). Anim. Biotechnol. 2019, 31, 283–294. [Google Scholar] [CrossRef]
- Patnaik, B.B.; Hwang, H.J.; Baliarsingh, S.; Chung, J.M.; Sang, M.K.; Min, H.R.; Park, J.E.; Cho, H.C.; Kang, S.W.; Park, S.Y.; et al. In silico characterization of single and tandem-repeat galectin from terrestrial slug, Incilaria fruhstorferi. J. Environ. Biol. 2019, 40, 940–947. [Google Scholar] [CrossRef]
- Cool, D.R.; Loh, Y.P. Carboxypeptidase E is a sorting receptor for prohormones: Binding and kinetic studies. Mol. Cell. Endocrinol. 1998, 139, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Hourdou, M.L.; Guinand, M.; Vacheron, M.J.; Michel, G.; Denoroy, L.; Duez, C.; Englebert, S.; Joris, B.; Weber, G.; Ghuysen, J.M. Characterization of the sporulation-related gamma-D-glutamyl-(L) meso-diaminopimelic-acid-hydrolysing peptidase I of Bacillus sphaericus NCTC 9602 as a member of the metallo(zinc) carboxypeptidase A family. Modular design of the protein. Biochem. J. 1993, 292, 563–570. [Google Scholar] [CrossRef]
- Cheng, J.; Zhang, Z.X.; Li, Y.; Zhang, L.W.; Hui, M.; Sha, Z.L. Rolling with the punches: Organism-environment interactions shape spatial pattern of adaptive differentiation in the widespread mantis shrimp Oratosquilla oratoria. Sci. Total Environ. 2024, 917, 170244. [Google Scholar] [CrossRef]
- Segundo, B.S.; Martínez, M.C.; Vilanova, M.; Cuchillo, C.M.; Avilés, F.X. The severed activation segment of porcine pancreatic procarboxypeptidase A is a powerful inhibitor of the active enzyme isolation and characterisation of the activation peptide. Biochim. Et Biophys. Acta 1982, 707, 74–80. [Google Scholar] [CrossRef]
- Sui, Y.P.; Liu, X.B.; Chai, L.Q.; Wang, J.X.; Zhao, X.F. Characterization and influences of classical insect hormones on the expression profiles of a molting carboxypeptidase A from the cotton bollworm (Helicoverpa armigera). Insect Mol. Biol. 2009, 18, 353–363. [Google Scholar] [CrossRef]
- Ote, M.; Mita, K.; Kawasaki, H.; Daimon, T.; Kobayashi, M.; Shimada, T. Identification of molting fluid carboxypeptidase A (MF-CPA) in Bombyx mori. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2005, 141, 314–322. [Google Scholar] [CrossRef]
- Tham, H.W.; Balasubramaniam, V.R.; Tejo, B.A.; Ahmad, H.; Hassan, S.S. CPB1 of Aedes aegypti interacts with DENV2 E protein and regulates intracellular viral accumulation and release from midgut cells. Viruses 2014, 6, 5028–5046. [Google Scholar] [CrossRef] [PubMed]
- Settle, S.J.; Green, M.M.; Burtis, K.C. The silver gene of Drosophila melanogaster encodes multiple carboxypeptidases similar to mammalian prohormone-processing enzymes. Proc. Natl. Acad. Sci. USA 1995, 92, 9470–9474. [Google Scholar] [CrossRef] [PubMed]
- Fernández, D.; Pallarès, I.; Vendrell, J.; Avilés, F.X. Progress in metallocarboxypeptidases and their small molecular weight inhibitors. Biochimie 2010, 92, 1484–1500. [Google Scholar] [CrossRef] [PubMed]





| Stomatopod | Unigene_ID | CDS (bp) | Amino Acid (bp) | Molecular Weight (kDa) | Isoelectric Point | Instability Index | Aliphatic Index | Grand Average of Hydropathicity |
|---|---|---|---|---|---|---|---|---|
| L. maculata | LmPB.12809.1 | 879 | 292 | 32.74 | 4.99 | 17.64 | 66.47 | −0.395 |
| LmPB.2470.2 | 1296 | 431 | 48.20 | 6.01 | 24.71 | 80.60 | −0.243 | |
| LmPB.14809.1 | 1314 | 437 | 48.66 | 5.33 | 29.51 | 89.92 | −0.206 | |
| LmPB.2470.3 | 816 | 271 | 30.39 | 5.26 | 16.53 | 65.50 | −0.448 | |
| LmPB.10623.2 | 1311 | 436 | 48.42 | 5.02 | 21.49 | 81.24 | −0.181 | |
| LmPB.10623.5 | 1314 | 437 | 48.56 | 5.07 | 21.46 | 81.05 | −0.188 | |
| LmPB.10623.6 | 1071 | 356 | 39.54 | 6.83 | 32.19 | 72.81 | −0.404 | |
| LmPB.4677.1 | 2892 | 963 | 109.30 | 5.35 | 28.60 | 73.64 | −0.419 | |
| LmPB.641.2 | 1065 | 354 | 39.53 | 5.03 | 39.40 | 80.65 | −0.405 | |
| LmPB.13939.1 | 1539 | 512 | 58.43 | 5.67 | 40.02 | 77.89 | −0.608 | |
| LmPB.1608.3 | 3726 | 1241 | 134.73 | 9.81 | 54.01 | 65.50 | −0.528 | |
| LmPB.1608.2 | 3954 | 1317 | 142.96 | 9.30 | 53.57 | 66.83 | −0.534 | |
| LmPB.1608.1 | 3624 | 1207 | 131.36 | 9.39 | 55.89 | 69.53 | −0.528 | |
| O. scyllarus | OsPB.10290.1 | 1767 | 588 | 67.86 | 10.18 | 52.44 | 69.13 | −0.800 |
| OsPB.14360.2 | 1293 | 430 | 48.10 | 5.78 | 26.82 | 81.16 | −0.239 | |
| OsPB.15333.2 | 1062 | 353 | 39.77 | 5.04 | 30.61 | 70.48 | −0.419 | |
| OsPB.15333.1 | 1302 | 433 | 48.33 | 5.20 | 32.76 | 78.18 | −0.285 | |
| OsPB.13536.1 | 873 | 290 | 32.06 | 4.69 | 26.17 | 75.72 | −0.265 | |
| OsPB.13536.2 | 1308 | 435 | 48.44 | 4.91 | 25.02 | 82.99 | −0.198 | |
| OsPB.14654.2 | 1311 | 436 | 48.51 | 4.91 | 24.99 | 83.03 | −0.194 | |
| OsPB.11247.2 | 1233 | 410 | 46.26 | 5.59 | 24.44 | 81.83 | −0.148 | |
| OsPB.11247.1 | 807 | 268 | 30.74 | 5.21 | 22.87 | 72.39 | −0.229 | |
| OsPB.13536.3 | 648 | 215 | 23.84 | 4.57 | 27.15 | 73.12 | −0.262 | |
| OsPB.14524.1 | 1239 | 412 | 46.52 | 4.98 | 50.86 | 74.83 | −0.587 | |
| OsPB.14524.2 | 1548 | 515 | 58.11 | 4.95 | 46.90 | 78.78 | −0.476 | |
| OsPB.10382.1 | 621 | 206 | 23.01 | 5.45 | 35.69 | 70.53 | −0.422 | |
| OsPB.2551.1 | 1464 | 487 | 54.67 | 5.47 | 34.46 | 73.51 | −0.333 | |
| OsPB.2940.1 | 1602 | 533 | 61.51 | 9.09 | 44.82 | 74.03 | −0.479 | |
| OsPB.2812.1 | 3297 | 1098 | 121.85 | 8.26 | 44.30 | 79.54 | −0.307 | |
| OsPB.6250.1 | 3948 | 1315 | 142.52 | 9.32 | 53.37 | 66.51 | −0.540 | |
| O. oratoria | OoGene.48164 | 867 | 288 | 31.77 | 5.27 | 22.42 | 75.28 | −0.267 |
| OoGene.143872 | 1050 | 349 | 38.98 | 5.58 | 28.58 | 76.07 | −0.298 | |
| OoGene.56777 | 684 | 227 | 25.46 | 4.99 | 17.72 | 61.85 | −0.456 | |
| OoGene.373886 | 699 | 232 | 25.72 | 4.97 | 27.00 | 69.91 | −0.285 | |
| OoGene.116636 | 939 | 312 | 34.94 | 9.42 | 45.08 | 70.74 | −0.560 | |
| OoGene.144862 | 627 | 208 | 23.01 | 5.35 | 30.38 | 70.96 | −0.270 | |
| OoGene.166368 | 600 | 199 | 21.97 | 5.40 | 29.10 | 72.16 | −0.255 | |
| OoGene.325317 | 618 | 205 | 22.69 | 5.67 | 35.74 | 62.44 | −0.471 | |
| OoGene.283166 | 573 | 190 | 20.96 | 4.64 | 20.80 | 69.26 | −0.349 | |
| OoGene.43133 | 585 | 194 | 21.85 | 6.23 | 29.27 | 67.42 | −0.454 | |
| OoGene.191567 | 729 | 242 | 25.26 | 4.69 | 38.19 | 78.26 | −0.045 | |
| OoGene.373941 | 426 | 141 | 15.72 | 6.95 | 51.37 | 51.21 | −0.648 | |
| OoGene.162805 | 411 | 136 | 15.05 | 5.04 | 16.53 | 78.90 | −0.213 | |
| OoGene.208437 | 627 | 208 | 23.33 | 10.23 | 80.84 | 51.73 | −0.754 | |
| OoGene.184724 | 312 | 103 | 11.04 | 4.63 | 23.61 | 86.41 | −0.071 |
| Stomatopod | Unigene ID | Alpha Helix (%) | Extended Strand (%) | Beta Turn (%) | Random Coil (%) |
|---|---|---|---|---|---|
| L. maculata | LmPB.12809.1 | 32.19 | 19.86 | 6.51 | 41.44 |
| LmPB.2470.2 | 36.66 | 17.4 | 5.34 | 40.6 | |
| LmPB.14809.1 | 37.53 | 17.62 | 6.64 | 38.22 | |
| LmPB.2470.3 | 29.89 | 21.03 | 7.01 | 42.07 | |
| LmPB.10623.2 | 35.55 | 20.64 | 5.5 | 38.3 | |
| LmPB.10623.5 | 35.24 | 19.91 | 5.26 | 39.59 | |
| LmPB.10623.6 | 21.91 | 24.44 | 5.9 | 47.75 | |
| LmPB.4677.1 | 24.71 | 20.77 | 8.31 | 46.21 | |
| LmPB.641.2 | 28.53 | 19.49 | 9.04 | 42.94 | |
| LmPB.13939.1 | 24.8 | 18.16 | 5.27 | 51.76 | |
| LmPB.1608.3 | 16.92 | 21.43 | 8.14 | 53.51 | |
| LmPB.1608.2 | 17.77 | 20.43 | 8.28 | 53.53 | |
| LmPB.1608.1 | 19.06 | 19.22 | 7.37 | 54.35 | |
| O. scyllarus | OsPB.10290.1 | 28.23 | 18.54 | 8.5 | 44.73 |
| OsPB.14360.2 | 36.74 | 18.84 | 6.05 | 38.37 | |
| OsPB.15333.2 | 37.68 | 17.28 | 6.52 | 38.53 | |
| OsPB.15333.1 | 37.41 | 17.78 | 5.77 | 39.03 | |
| OsPB.13536.1 | 30.69 | 20.69 | 6.55 | 42.07 | |
| OsPB.13536.2 | 39.54 | 18.62 | 5.75 | 36.09 | |
| OsPB.14654.2 | 37.16 | 19.5 | 5.96 | 37.39 | |
| OsPB.11247.2 | 36.1 | 19.51 | 5.12 | 39.27 | |
| OsPB.11247.1 | 33.58 | 20.15 | 5.97 | 40.3 | |
| OsPB.13536.3 | 26.51 | 22.79 | 7.44 | 43.26 | |
| OsPB.14524.1 | 27.18 | 17.72 | 6.55 | 48.54 | |
| OsPB.14524.2 | 21.17 | 20.58 | 6.6 | 51.65 | |
| OsPB.10382.1 | 18.45 | 26.7 | 7.77 | 47.09 | |
| OsPB.2551.1 | 23.61 | 21.97 | 6.98 | 47.43 | |
| OsPB.2940.1 | 25.14 | 21.76 | 3.38 | 49.72 | |
| OsPB.2812.1 | 36.7 | 15.66 | 4.46 | 43.71 | |
| OsPB.6250.1 | 19.47 | 19.24 | 7.45 | 53.84 | |
| O. oratoria | OoGene.48164 | 31.94 | 21.88 | 6.6 | 39.58 |
| OoGene.143872 | 34.96 | 19.77 | 6.02 | 39.26 | |
| OoGene.56777 | 30.4 | 19.38 | 6.17 | 44.05 | |
| OoGene.373886 | 31.9 | 20.69 | 5.17 | 42.24 | |
| OoGene.116636 | 28.53 | 16.35 | 6.09 | 49.04 | |
| OoGene.144862 | 25.48 | 21.63 | 6.25 | 46.63 | |
| OoGene.166368 | 25.63 | 21.61 | 5.53 | 47.24 | |
| OoGene.325317 | 18.05 | 21.95 | 5.85 | 54.15 | |
| OoGene.283166 | 28.42 | 20 | 10 | 41.58 | |
| OoGene.43133 | 19.07 | 24.74 | 5.15 | 51.03 | |
| OoGene.191567 | 21.9 | 21.9 | 7.85 | 48.35 | |
| OoGene.373941 | 18.44 | 17.02 | 1.42 | 63.12 | |
| OoGene.162805 | 37.5 | 20.59 | 10.29 | 31.62 | |
| OoGene.208437 | 12.02 | 15.38 | 3.37 | 69.23 | |
| OoGene.184724 | 33.98 | 20.39 | 8.74 | 36.89 |
| Stomatopod | Unigene ID | Template | Seq Identity (%) | Oligo-State | GMQE | QMEAND |
|---|---|---|---|---|---|---|
| L. maculata | LmPB.12809.1 | 7eqx.1.B | 44.44 | Monomer | 0.82 | 0.77 ± 0.05 |
| LmPB.2470.2 | 1aye.1.A | 28.27 | Monomer | 0.72 | 0.73 ± 0.05 | |
| LmPB.14809.1 | 2boa.1.A | 39.75 | Monomer | 0.71 | 0.72 ± 0.05 | |
| LmPB.2470.3 | 7eqx.1.B | 46.24 | Monomer | 0.83 | 0.78 ± 0.05 | |
| LmPB.10623.2 | 1aye.1.A | 43.37 | Monomer | 0.75 | 0.77 ± 0.05 | |
| LmPB.10623.5 | 1aye.1.A | 43.37 | Monomer | 0.75 | 0.77 ± 0.05 | |
| LmPB.10623.6 | 1aye.1.A | 41.67 | Monomer | 0.63 | 0.75 ± 0.05 | |
| LmPB.4677.1 | 3mn8.1.A | 47.12 | Monomer | 0.33 | 0.72 ± 0.05 | |
| LmPB.641.2 | 2nsm.1A | 47.92 | Monomer | 0.3 | 0.58 ± 0.07 | |
| LmPB.13939.1 | 2nsm.1A | 49.59 | Monomer | 0.56 | 0.66 ± 0.05 | |
| LmPB.1608.3 | 4a37.1.A | 28.93 | Monomer | 0.1 | 0.42 ± 0.05 | |
| LmPB.1608.2 | 312n.1.A | 30.65 | Monomer | 0.12 | 0.55 ± 0.05 | |
| LmPB.1608.1 | 312n.1.A | 30.48 | Monomer | 0.1 | 0.53 ± 0.05 | |
| O. scyllarus | OsPB.10290.1 | 5om9.1.A | 29.67 | Monomer | 0.46 | 0.61 ± 0.05 |
| OsPB.14360.2 | 1aye.1.A | 37.50 | Monomer | 0.73 | 0.73 ± 0.05 | |
| OsPB.15333.2 | 2boa.1.A | 41.76 | Monomer | 0.77 | 0.74 ± 0.05 | |
| OsPB.15333.1 | 1aye.1.A | 39.03 | Monomer | 0.71 | 0.72 ± 0.05 | |
| OsPB.13536.1 | 1dtd.1A | 52.3 | Monomer | 0.85 | 0.81 ± 0.05 | |
| OsPB.13536.2 | 1aye.1.A | 45.92 | Monomer | 0.75 | 0.77 ± 0.05 | |
| OsPB.14654.2 | 1aye.1.A | 45.92 | Monomer | 0.75 | 0.77 ± 0.05 | |
| OsPB.11247.2 | 5om9.1.A | 33.33 | Monomer | 0.65 | 0.68 ± 0.05 | |
| OsPB.11247.1 | 7eqx.1.B | 42.15 | Monomer | 0.74 | 0.74 ± 0.05 | |
| OsPB.13536.3 | 1aye.1.A | 50.48 | Monomer | 0.81 | 0.75 ± 0.06 | |
| OsPB.14524.1 | 2nsm.1.A | 46.65 | Monomer | 0.63 | 0.71 ± 0.05 | |
| OsPB.14524.2 | 2nsm.1.A | 46.15 | Monomer | 0.61 | 0.73 ± 0.05 | |
| OsPB.10382.1 | 1uwy.1.A | 40.65 | Monomer | 0.56 | 0.66 ± 0.07 | |
| OsPB.2551.1 | 1qmu.1.A | 45.01 | Monomer | 0.64 | 0.78 ± 0.05 | |
| OsPB.2940.1 | 312n.1.A | 31.37 | Monomer | 0.31 | 0.47 ± 0.05 | |
| OsPB.2812.1 | 312n.1.A | 25 | Monomer | 0.22 | 0.56 ± 0.05 | |
| OsPB.6250.1 | 312n.1.A | 30.65 | Monomer | 0.12 | 0.55 ± 0.05 | |
| O. oratoria | OoGene.48164 | 1aye.1.A | 51.96 | Monomer | 0.84 | 0.79 ± 0.05 |
| OoGene.143872 | 1aye.1.A | 46.38 | Monomer | 0.82 | 0.78 ± 0.05 | |
| OoGene.56777 | 7eqx.1.B | 45.54 | Monomer | 0.79 | 0.74 ± 0.06 | |
| OoGene.373886 | 1aye.1.A | 50 | Monomer | 0.82 | 0.76 ± 0.06 | |
| OoGene.116636 | 1aye.1.A | 42.18 | Monomer | 0.68 | 0.70 ± 0.05 | |
| OoGene.144862 | 1aye.1.A | 51.71 | Monomer | 0.8 | 0.74 ± 0.06 | |
| OoGene.166368 | 1aye.1.A | 52.04 | Monomer | 0.81 | 0.74 ± 0.06 | |
| OoGene.325317 | 1aye.1.A | 49.49 | Monomer | 0.75 | 0.70 ± 0.06 | |
| OoGene.283166 | 1kwm.2.A | 45.98 | Monomer | 0.77 | 0.77 ± 0.07 | |
| OoGene.43133 | 1aye.1.A | 48.95 | Monomer | 0.76 | 0.69 ± 0.06 | |
| OoGene.191567 | 1aye.1.A | 42.56 | Monomer | 0.64 | 0.69 ± 0.06 | |
| OoGene.373941 | 1aye.1.A | 48.76 | Monomer | 0.62 | 0.65 ± 0.07 | |
| OoGene.162805 | 1kwm.2.A | 44.03 | Monomer | 0.79 | 0.74 ± 0.07 | |
| OoGene.208437 | 3osl.1.A | 28.21 | Monomer | 0.6 | 0.56 ± 0.06 | |
| OoGene.184724 | 5mrv.1.A | 43.14 | Monomer | 0.83 | 0.78 ± 0.08 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Dong, X.; Hu, X.; Duan, X.; Xu, B.; Lou, F. Identification of the Metallocarboxypeptidase M14 Gene Family in Lysiosquillina maculata, Odontodactylus scyllarus and Oratosquilla oratoria Based on the Full-Length Transcriptome. Fishes 2025, 10, 515. https://doi.org/10.3390/fishes10100515
Zhang J, Dong X, Hu X, Duan X, Xu B, Lou F. Identification of the Metallocarboxypeptidase M14 Gene Family in Lysiosquillina maculata, Odontodactylus scyllarus and Oratosquilla oratoria Based on the Full-Length Transcriptome. Fishes. 2025; 10(10):515. https://doi.org/10.3390/fishes10100515
Chicago/Turabian StyleZhang, Jiantong, Xiuqiang Dong, Xinyan Hu, Xiaowen Duan, Bin Xu, and Fangrui Lou. 2025. "Identification of the Metallocarboxypeptidase M14 Gene Family in Lysiosquillina maculata, Odontodactylus scyllarus and Oratosquilla oratoria Based on the Full-Length Transcriptome" Fishes 10, no. 10: 515. https://doi.org/10.3390/fishes10100515
APA StyleZhang, J., Dong, X., Hu, X., Duan, X., Xu, B., & Lou, F. (2025). Identification of the Metallocarboxypeptidase M14 Gene Family in Lysiosquillina maculata, Odontodactylus scyllarus and Oratosquilla oratoria Based on the Full-Length Transcriptome. Fishes, 10(10), 515. https://doi.org/10.3390/fishes10100515







