Cell-Free Synthetic Biology Platform for Engineering Synthetic Biological Circuits and Systems
Abstract
:1. Introduction
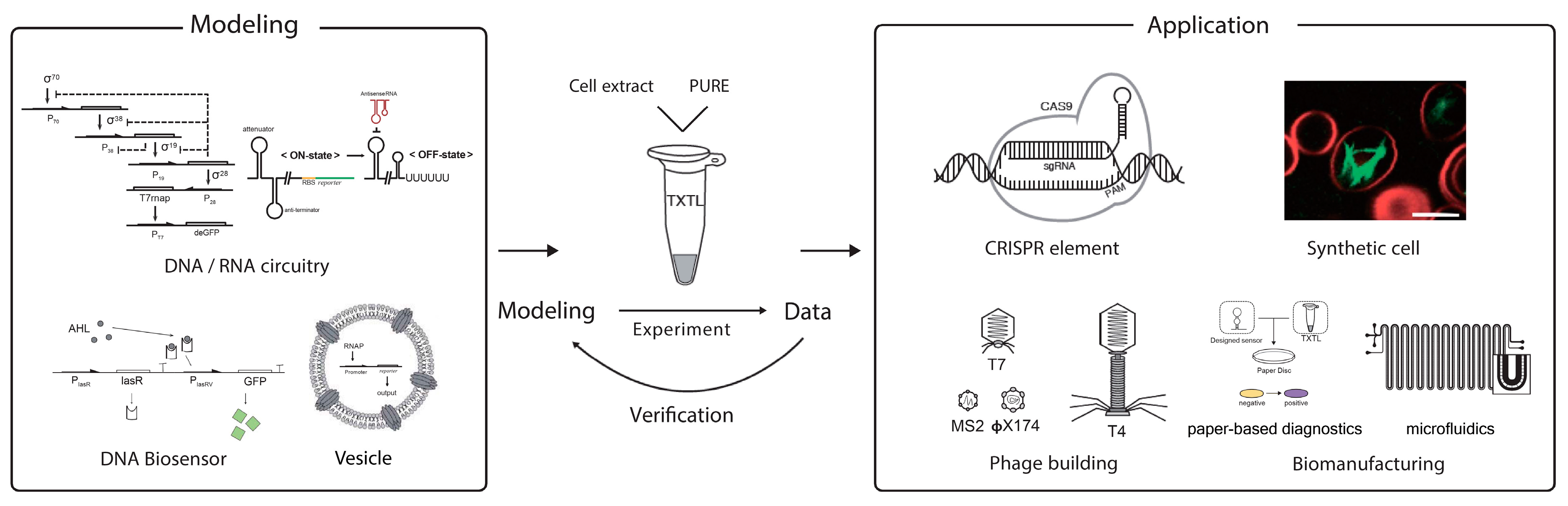
2. Cell-Free Transcription–Translation Platform for Synthetic Biology
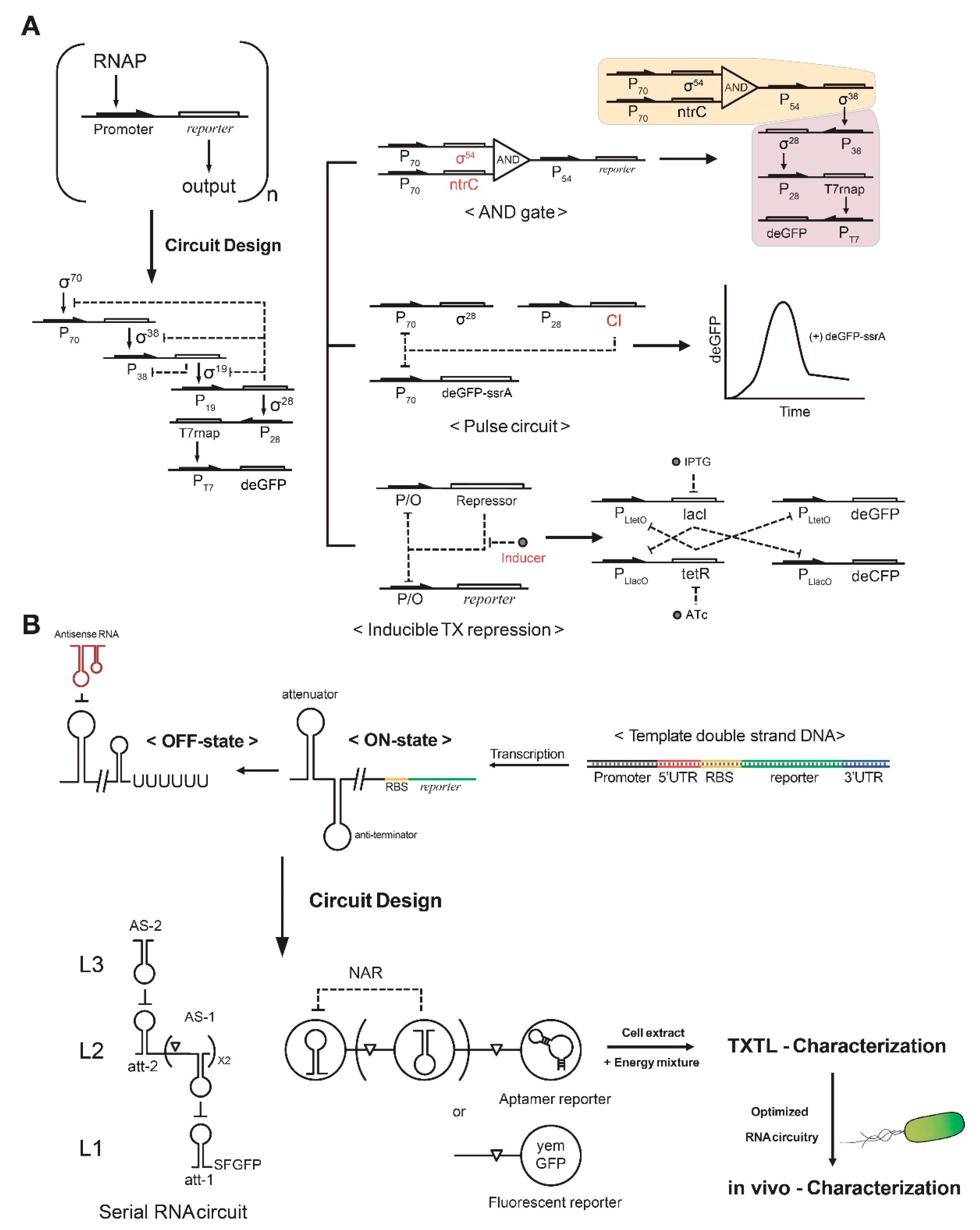
3. In Vitro Synthetic Gene Circuits
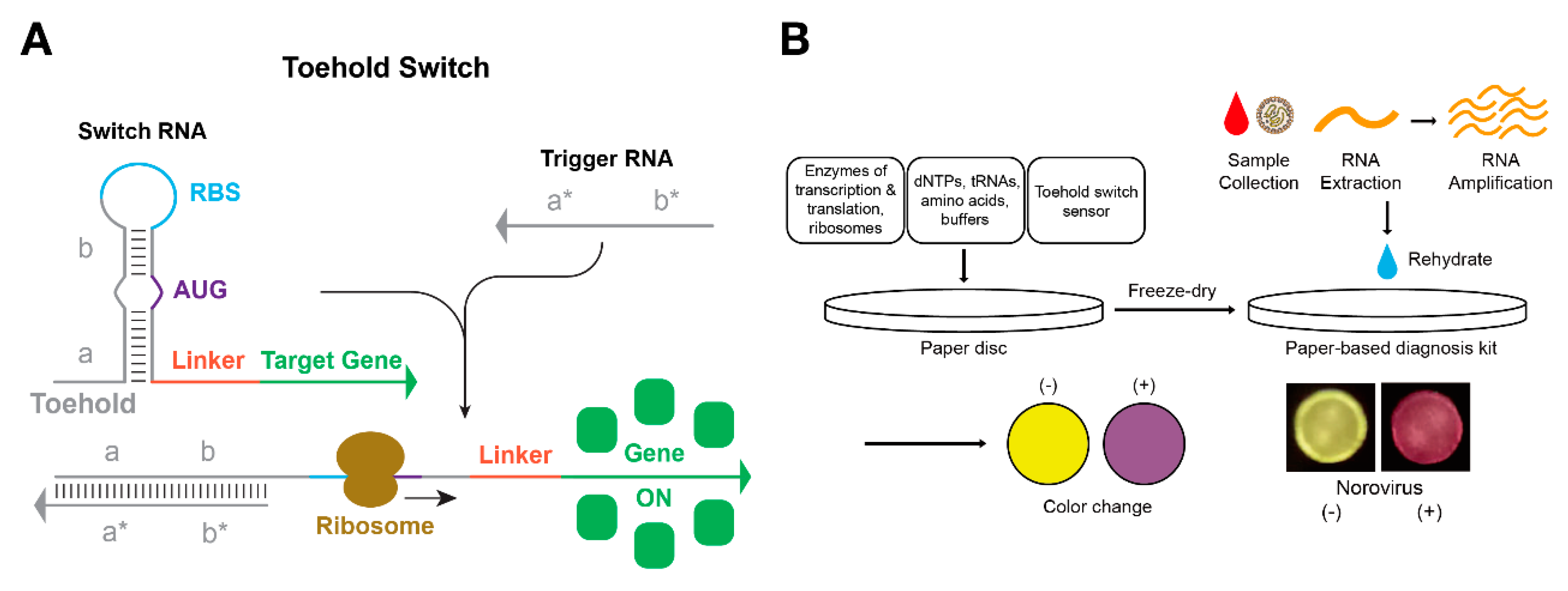
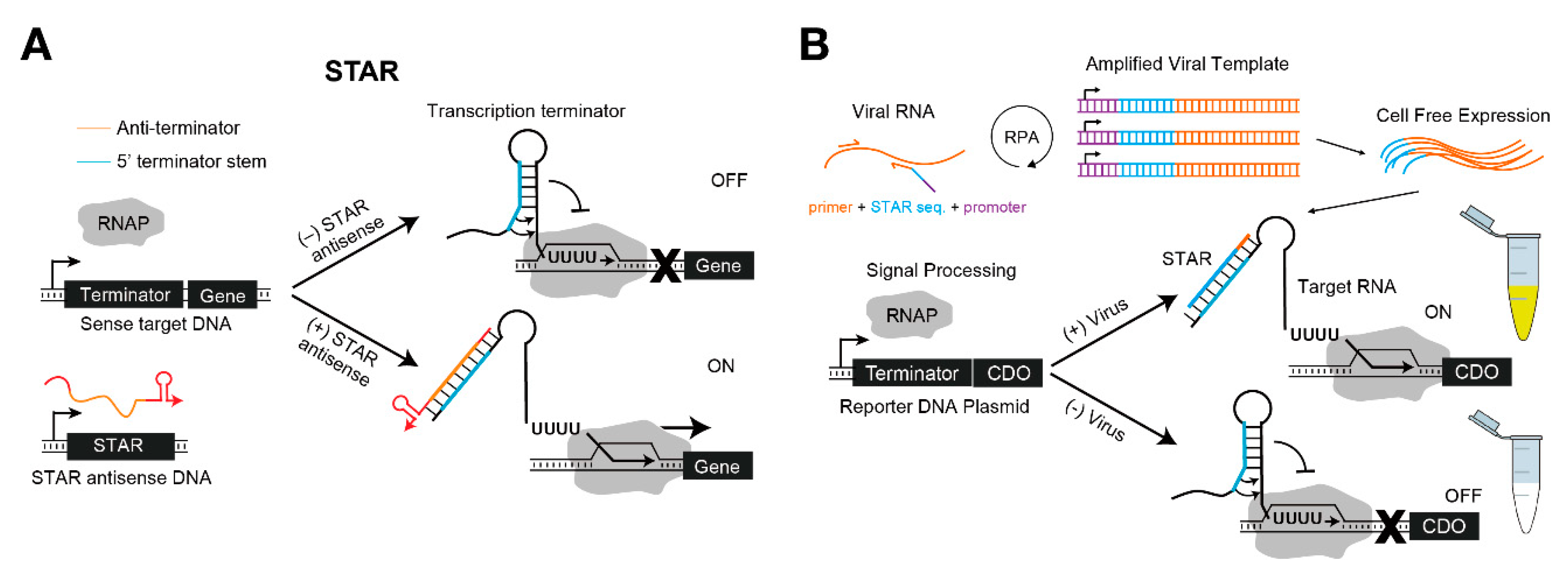
4. RNA Regulatory Circuits for Cell-Free Synthetic Biology
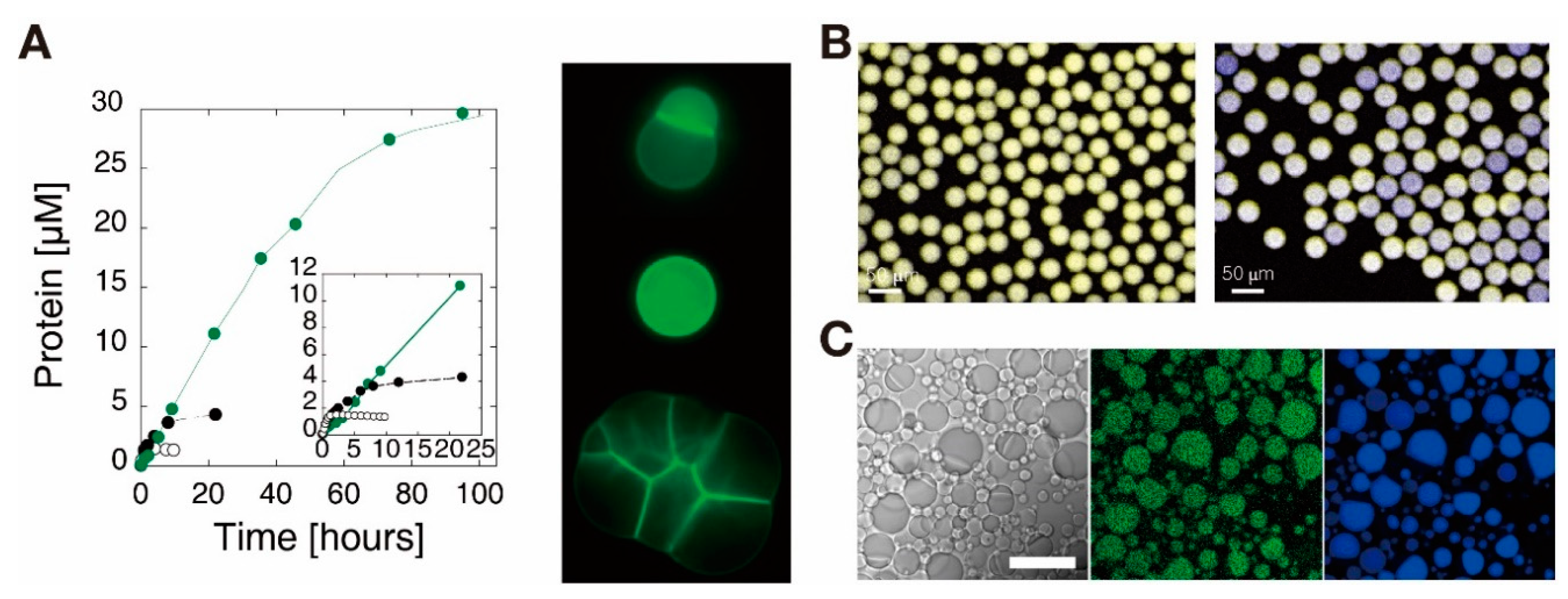
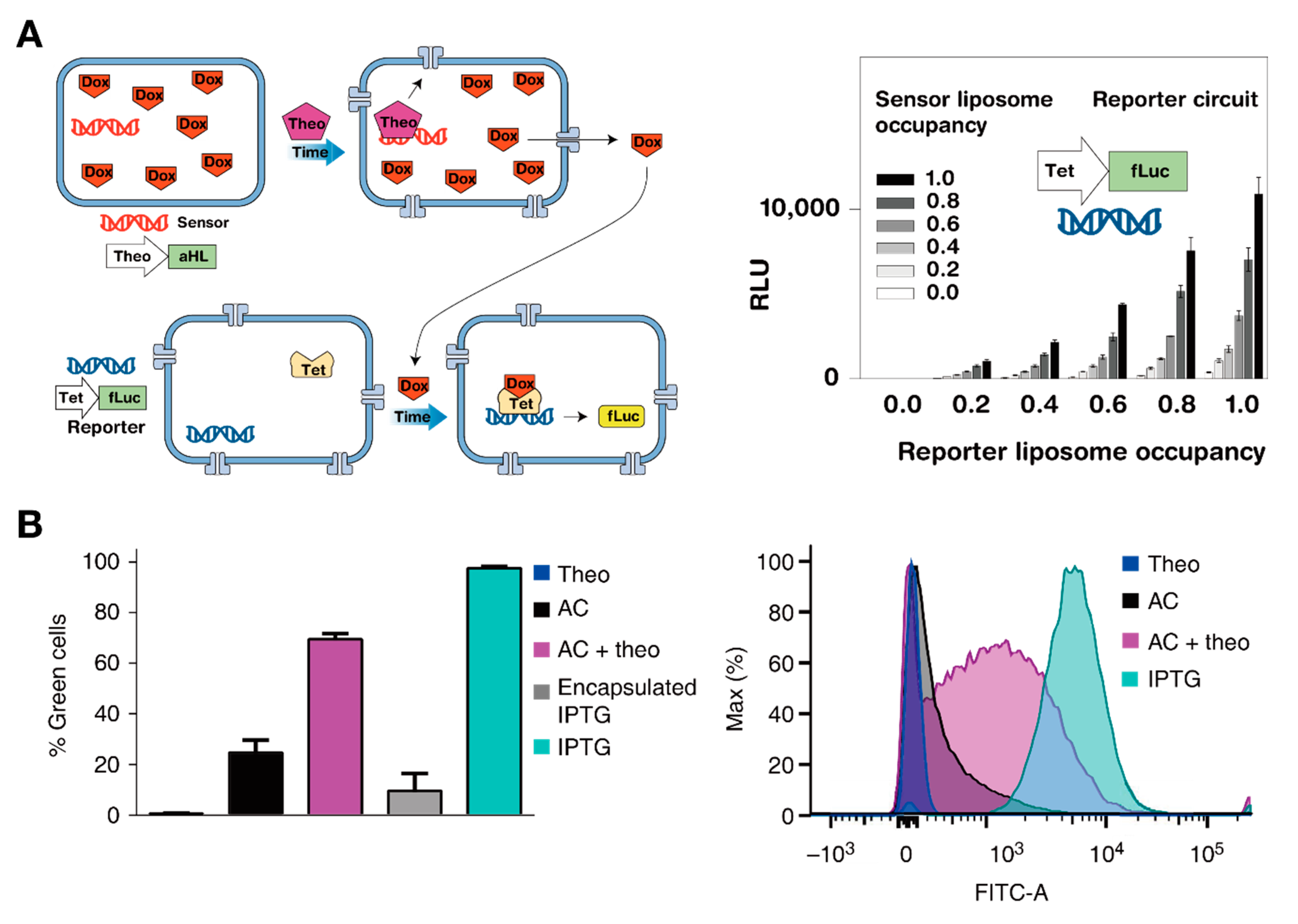
5. Encapsulation of in Vitro Circuits toward the Synthesis of Artificial Cells
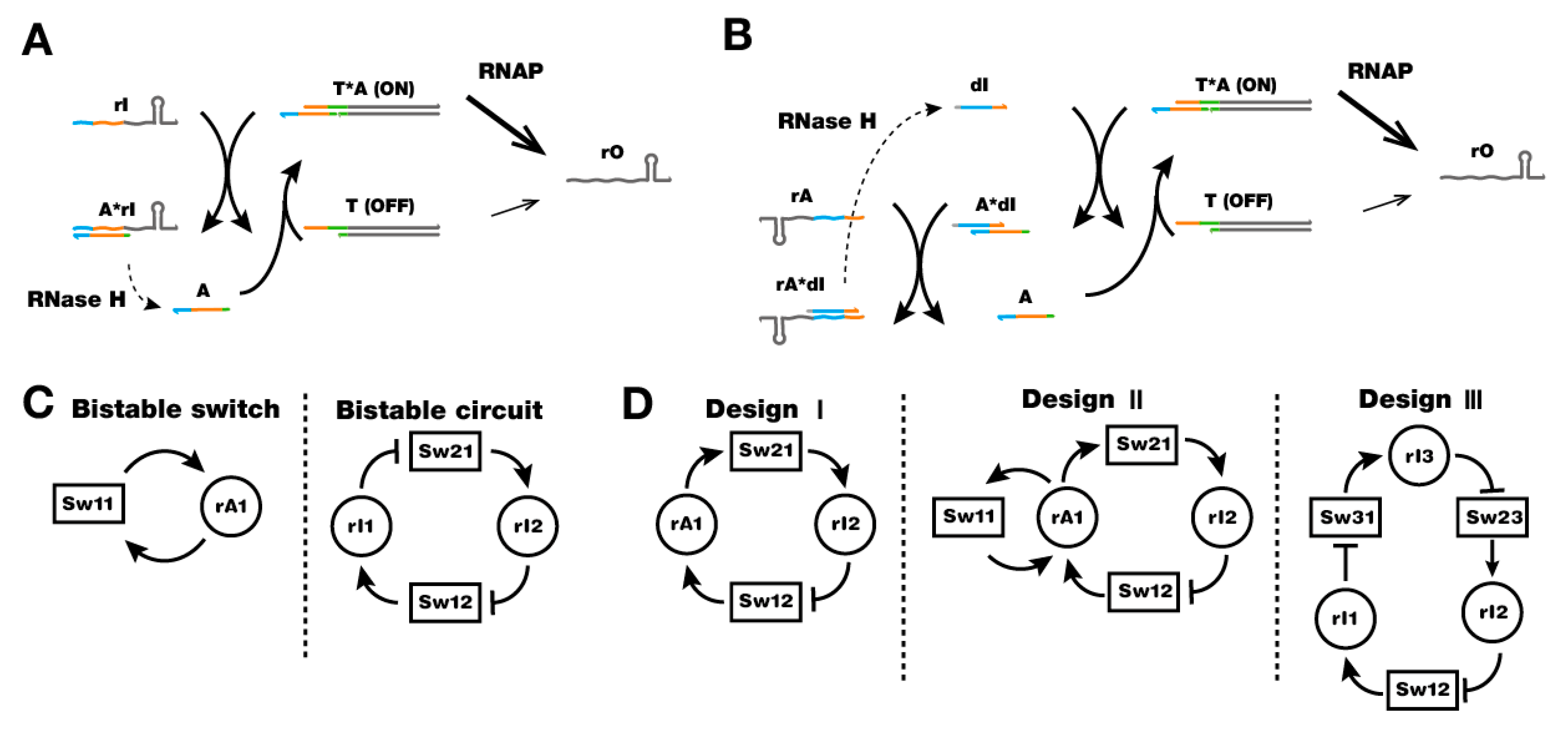
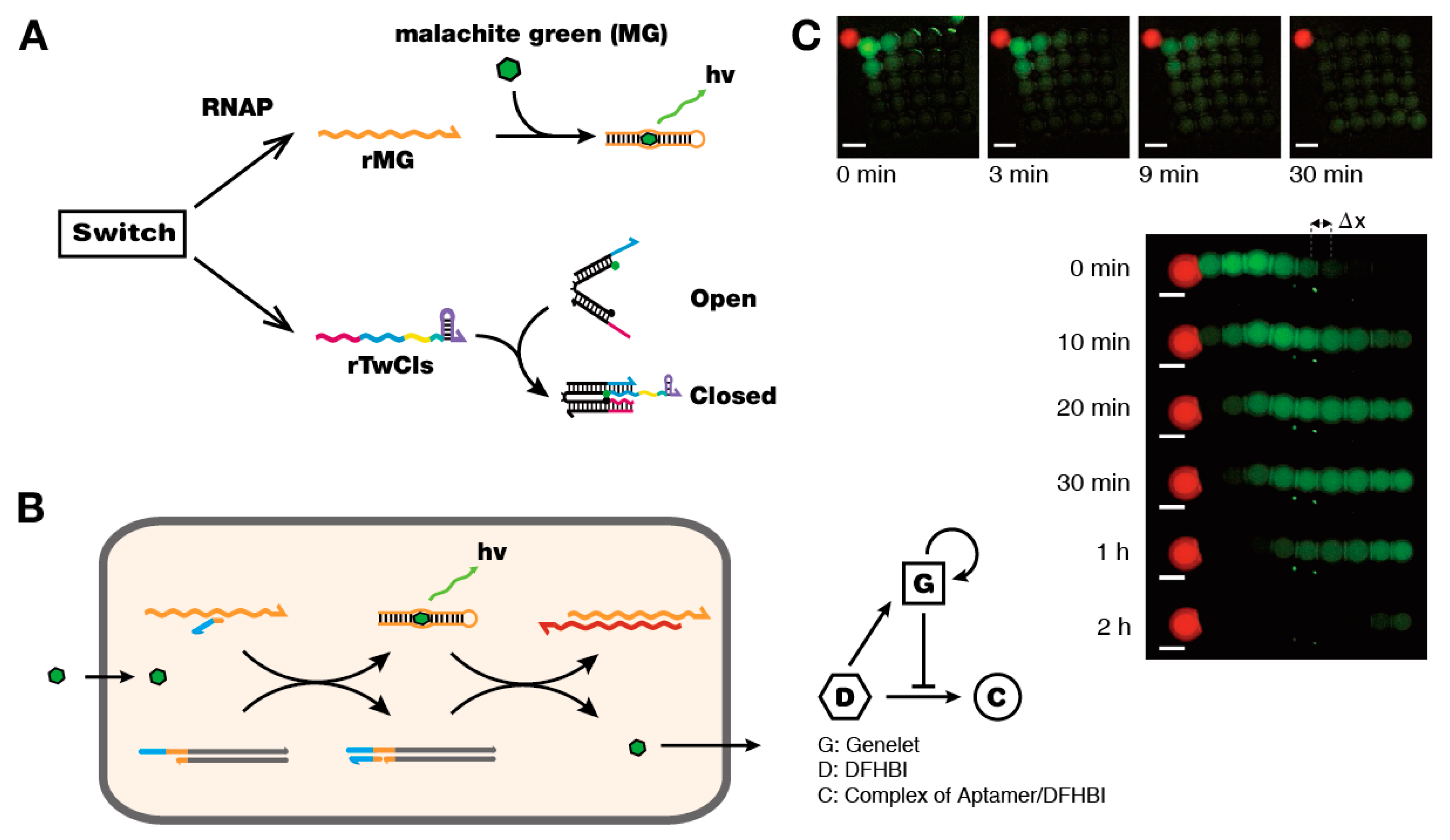
6. Artificial DNA Structures and Systems for in Vitro Synthetic Biology
7. Mathematical Modeling Supports the Development and Analysis of in Vitro Systems
8. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Elowitz, M.B.; Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature. 2000, 403, 335–338. [Google Scholar] [CrossRef] [PubMed]
- Gardner, T.S.; Cantor, C.R.; Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 2000, 403, 339–342. [Google Scholar] [CrossRef] [PubMed]
- Cameron, D.E.; Bashor, C.J.; Collins, J.J. A brief history of synthetic biology. Nat. Rev. Microbiol. 2014, 12, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, A.A.; Der, B.S.; Shin, J.; Vaidyanathan, P.; Paralanov, V.; Strychalski, E.A.; Ross, D.; Densmore, D.; Voigt, C.A. Genetic circuit design automation. Science 2016, 352, aac7341. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, S.P.; Smanski, M.J.; Voigt, C.A.; Densmore, D.M. Genetic Design via Combinatorial Constraint Specification. ACS Synth. Biol. 2017, 6, 2130–2135. [Google Scholar] [CrossRef] [PubMed]
- Garenne, D.; Noireaux, V. Cell-free transcription-translation: Engineering biology from the nanometer to the millimeter scale. Curr. Opin. Biotechnol. 2019, 58, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Dudley, Q.M.; Karim, A.S.; Jewett, M.C. Cell-free metabolic engineering: Biomanufacturing beyond the cell. Biotechnol. J. 2015, 10, 69–82. [Google Scholar] [CrossRef]
- Guo, W.; Sheng, J.; Feng, X. Mini-review: In vitro Metabolic Engineering for Biomanufacturing of High-value Products. Comput. Struct. Biotechnol. J. 2017, 15, 161–167. [Google Scholar] [CrossRef]
- Carlson, E.D.; Gan, R.; Hodgman, C.E.; Jewett, M.C. Cell-free protein synthesis: Applications come of age. Biotechnol. Adv. 2012, 30, 1185–1194. [Google Scholar] [CrossRef]
- Shin, J.; Noireaux, V. An E. coli Cell-Free Expression Toolbox: Application to Synthetic Gene Circuits and Artificial Cells. ACS Synth. Biol. 2012, 1, 29–41. [Google Scholar] [CrossRef]
- Noireaux, V.; Bar-Ziv, R.; Libchaber, A. Principles of cell-free genetic circuit assembly. Proc. Natl. Acad. Sci. USA 2003, 100, 12672–12677. [Google Scholar] [CrossRef]
- Marshall, R.; Maxwell, C.S.; Collins, S.P.; Beisel, C.L.; Noireaux, V. Short DNA containing chi sites enhances DNA stability and gene expression in E. coli cell-free transcription-translation systems. Biotechnol. Bioeng. 2017, 114, 2137–2141. [Google Scholar] [CrossRef] [PubMed]
- Stiege, W.; Erdmann, V.A. The potentials of the in vitro protein biosynthesis system. J. Biotechnol. 1995, 41, 81–90. [Google Scholar] [CrossRef]
- Jewett, M.C.; Noireaux, V. Synthetic biology: Tailor-made genetic codes. Nat. Chem. 2016, 8, 291–292. [Google Scholar] [CrossRef] [PubMed]
- Timm, A.C.; Shankles, P.G.; Foster, C.M.; Doktycz, M.J.; Retterer, S.T. Toward Microfluidic Reactors for Cell-Free Protein Synthesis at the Point-of-Care. Small 2016, 12, 810–817. [Google Scholar] [CrossRef] [PubMed]
- Noireaux, V.; Maeda, Y.T.; Libchaber, A. Development of an artificial cell, from self-organization to computation and self-reproduction. Proc. Natl. Acad. Sci. USA 2011, 108, 3473–3480. [Google Scholar] [CrossRef] [PubMed]
- Matveev, S.V.; Vinokurov, L.M.; Shaloiko, L.A.; Davies, C.; Matveeva, E.A.; Alakhov, Y.B. Effect of the ATP level on the overall protein biosynthesis rate in a wheat germ cell-free system. Biochim. Biophys. Acta 1996, 1293, 207–212. [Google Scholar] [CrossRef]
- Ge, X.; Luo, D.; Xu, J. Cell-Free Protein Expression under Macromolecular Crowding Conditions. PLoS ONE 2011, 6, e28707. [Google Scholar] [CrossRef]
- Siegal-Gaskins, D.; Tuza, Z.A.; Kim, J.; Noireaux, V.; Murray, R.M. Gene Circuit Performance Characterization and Resource Usage in a Cell-Free “Breadboard”. ACS Synth. Biol. 2014, 3, 416–425. [Google Scholar] [CrossRef]
- Shimizu, Y.; Inoue, A.; Tomari, Y.; Suzuki, T.; Yokogawa, T.; Nishikawa, K.; Ueda, T. Cell-free translation reconstituted with purified components. Nat. Biotechnol. 2001, 19, 751–755. [Google Scholar] [CrossRef]
- Pardee, K.; Green, A.A.; Ferrante, T.; Cameron, D.E.; DaleyKeyser, A.; Yin, P.; Collins, J.J. Paper-based Synthetic Gene Networks. Cell 2014, 159, 940–954. [Google Scholar] [CrossRef] [PubMed]
- Garamella, J.; Marshall, R.; Rustad, M.; Noireaux, V. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. ACS Synth. Biol. 2016, 5, 344–355. [Google Scholar] [CrossRef]
- Marshall, R.; Maxwell, C.S.; Collins, S.P.; Jacobsen, T.; Luo, M.L.; Begemann, M.B.; Gray, B.N.; January, E.; Singer, A.; He, Y.; et al. Rapid and Scalable Characterization of CRISPR Technologies Using an E. coli Cell-Free Transcription-Translation System. Mol. Cell 2018, 69, 146–157.e3. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Jensen, K.; Freemont, P.S. Validation of an entirely in vitro approach for rapid prototyping of DNA regulatory elements for synthetic biology. Nucleic Acids Res. 2013, 41, 3471–3481. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.Z.; Yeung, E.; Hayes, C.A.; Noireaux, V.; Murray, R.M. Linear DNA for rapid prototyping of synthetic biological circuits in an Escherichia coli based TX-TL cell-free system. ACS Synth. Biol. 2014, 3, 387–397. [Google Scholar] [CrossRef]
- Takahashi, M.K.; Chappell, J.; Hayes, C.A.; Sun, Z.Z.; Kim, J.; Singhal, V.; Spring, K.J.; Al-Khabouri, S.; Fall, C.P.; Noireaux, V.; et al. Rapidly characterizing the fast dynamics of RNA genetic circuitry with cell-free transcription-translation (TX-TL) systems. ACS Synth. Biol. 2015, 4, 503–515. [Google Scholar] [CrossRef]
- Niederholtmeyer, H.; Sun, Z.Z.; Hori, Y.; Yeung, E.; Verpoorte, A.; Murray, R.M.; Maerkl, S.J. Rapid cell-free forward engineering of novel genetic ring oscillators. eLife 2015, 4, e09771. [Google Scholar] [CrossRef]
- Lucks, J.B.; Qi, L.; Mutalik, V.K.; Wang, D.; Arkin, A.P. Versatile RNA-sensing transcriptional regulators for engineering genetic networks. Proc. Natl. Acad. Sci. USA 2011, 108, 8617–8622. [Google Scholar] [CrossRef]
- Chappell, J.; Takahashi, M.K.; Meyer, S.; Loughrey, D.; Watters, K.E.; Lucks, J. The centrality of RNA for engineering gene expression. Biotechnol. J. 2013, 8, 1379–1395. [Google Scholar] [CrossRef]
- Rosenfeld, N.; Alon, U. Response Delays and the Structure of Transcription Networks. J. Mol. Biol. 2003, 329, 645–654. [Google Scholar] [CrossRef]
- Shen-Orr, S.S.; Milo, R.; Mangan, S.; Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 2002, 31, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.Y.; Takahashi, M.K.; Zhang, Y.; Lucks, J.B. Engineering a Functional Small RNA Negative Autoregulation Network with Model-Guided Design. ACS Synth. Biol. 2018, 7, 1507–1518. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Jardine, P.; Noireaux, V. Genome Replication, Synthesis, and Assembly of the Bacteriophage T7 in a Single Cell-Free Reaction. ACS Synth. Biol. 2012, 1, 408–413. [Google Scholar] [CrossRef] [PubMed]
- Rustad, M.; Eastlund, A.; Jardine, P.; Noireaux, V. Cell-free TXTL synthesis of infectious bacteriophage T4 in a single test tube reaction. Synth. Biol. 2018, 3, ysy002. [Google Scholar] [CrossRef]
- Pardee, K.; Green, A.A.; Takahashi, M.K.; Braff, D.; Lambert, G.; Lee, J.W.; Ferrante, T.; Ma, D.; Donghia, N.; Fan, M.; et al. Rapid, Low-Cost Detection of Zika Virus Using Programmable Biomolecular Components. Cell 2016, 165, 1255–1266. [Google Scholar] [CrossRef] [PubMed]
- Pardee, K.; Slomovic, S.; Nguyen, P.Q.; Lee, J.W.; Donghia, N.; Burrill, D.; Ferrante, T.; McSorley, F.R.; Furuta, Y.; Vernet, A.; et al. Portable, On-Demand Biomolecular Manufacturing. Cell 2016, 167, 248–259.e12. [Google Scholar] [CrossRef] [PubMed]
- Schwarz-Schilling, M.; Kim, J.; Cuba, C.; Weitz, M.; Franco, E.; Simmel, F.C. Building a Synthetic Transcriptional Oscillator. Methods Mol. Biol. 2016, 1342, 185–199. [Google Scholar] [PubMed]
- Kim, J.; Hopfield, J.J.; Winfree, E. Neural network computation by in vitro transcriptional circuits. In Proceedings of the 17th International Conference on Neural Information Processing Systems, Vancouver, BC, Canada, 5–8 December 2005; pp. 681–688. [Google Scholar]
- Kim, J.; White, K.S.; Winfree, E. Construction of an in vitro bistable circuit from synthetic transcriptional switches. Mol. Syst. Biol. 2006, 2, 68. [Google Scholar] [CrossRef]
- Franco, E.; Friedrichs, E.; Kim, J.; Jungmann, R.; Murray, R.; Winfree, E.; Simmel, F.C. Timing molecular motion and production with a synthetic transcriptional clock. Proc. Natl. Acad. Sci. USA 2011, 108, E784–E793. [Google Scholar] [CrossRef]
- Martin, C.T.; Coleman, J.E. Kinetic Analysis of T7 RNA Polymerase-Promoter Interactions with Small Synthetic Promoters. Biochemistry 1987, 26, 2690–2696. [Google Scholar] [CrossRef]
- Subsoontorn, P.; Kim, J.; Winfree, E. Ensemble Bayesian Analysis of Bistability in a Synthetic Transcriptional Switch. ACS Synth. Biol. 2012, 1, 299–316. [Google Scholar] [CrossRef] [PubMed]
- Yurke, B.; Mills, A.P., Jr. Using DNA to power nanostructures. Genet. Program. Evolvable Mach. 2003, 4, 111–122. [Google Scholar] [CrossRef]
- Markevich, N.I.; Hoek, J.B.; Kholodenko, B.N. Signaling switches and bistability arising from multisite phosphorylation in protein kinase cascades. J. Cell Biol. 2004, 164, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Ghaffarizadeh, A.; Flann, N.S.; Podgorski, G.J. Multistable switches and their role in cellular differentiation networks. BMC Bioinform. 2014, 15, S7. [Google Scholar] [CrossRef] [PubMed]
- Proctor, C.J.; Gray, D.A. Explaining oscillations and variability in the p53-Mdm2 system. BMC Syst. Biol. 2008, 2, 75. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.E.; Ihekwaba, A.E.C.; Elliott, M.; Johnson, J.R.; Gibney, C.A.; Foreman, B.E.; Nelson, G.; See, V.; Horton, C.A.; Spiller, D.G.; et al. Oscillations in NF-κB Signaling Control the Dynamics of Gene Expression. Science 2004, 306, 704–708. [Google Scholar] [CrossRef]
- Kim, J.; Winfree, E. Synthetic in vitro transcriptional oscillators. Mol. Syst. Biol. 2011, 7, 465. [Google Scholar] [CrossRef]
- Cho, E.J.; Lee, J.-W.; Ellington, A.D. Applications of Aptamers as Sensors. Annu. Rev. Anal. Chem. 2009, 2, 241–264. [Google Scholar] [CrossRef]
- Dupin, A.; Simmel, F.C. Signalling and differentiation in emulsion-based multi-compartmentalized in vitro gene circuits. Nat. Chem. 2019, 11, 32–39. [Google Scholar] [CrossRef]
- Lloyd, J.; Tran, C.H.; Wadhwani, K.; Cuba Samaniego, C.; Subramanian, H.K.K.; Franco, E. Dynamic Control of Aptamer-Ligand Activity Using Strand Displacement Reactions. ACS Synth. Biol. 2018, 7, 30–37. [Google Scholar] [CrossRef]
- Kim, J.; Quijano, J.F.; Yeung, E.; Murray, R.M. Synthetic logic circuits using RNA aptamer against T7 RNA polymerase. bioRxiv 2014. [Google Scholar] [CrossRef]
- Cuba Samaniego, C.; Franco, E. A Robust Molecular Network Motif for Period-Doubling Devices. ACS Synth. Biol. 2018, 7, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Franco, E.; Giordano, G.; Forsberg, P.-O.; Murray, R.M. Negative Autoregulation Matches Production and Demand in Synthetic Transcriptional Networks. ACS Synth. Biol. 2014, 3, 589–599. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Khetarpal, I.; Sen, S.; Murray, R.M. Synthetic circuit for exact adaptation and fold-change detection. Nucleic Acids Res. 2014, 42, 6078–6089. [Google Scholar] [CrossRef] [PubMed]
- Cuba Samaniego, C.; Giordano, G.; Kim, J.; Blanchini, F.; Franco, E. Molecular Titration Promotes Oscillations and Bistability in Minimal Network Models with Monomeric Regulators. ACS Synth. Biol. 2016, 5, 321–333. [Google Scholar] [CrossRef] [PubMed]
- Ayukawa, S.; Takinoue, M.; Kiga, D. RTRACS: A Modularized RNA-Dependent RNA Transcription System with High Programmability. Acc. Chem. Res. 2011, 44, 1369–1379. [Google Scholar] [CrossRef] [PubMed]
- Takinoue, M.; Kiga, D.; Shohda, K.-I.; Suyama, A. Experiments and simulation models of a basic computation element of an autonomous molecular computing system. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2008, 78, 041921. [Google Scholar] [CrossRef]
- Montagne, K.; Plasson, R.; Sakai, Y.; Fujii, T.; Rondelez, Y. Programming an in vitro DNA oscillator using a molecular networking strategy. Mol. Syst. Biol. 2011, 7, 466. [Google Scholar] [CrossRef]
- Baccouche, A.; Montagne, K.; Padirac, A.; Fujii, T.; Rondelez, Y. Dynamic DNA-toolbox reaction circuits: A walkthrough. Methods 2014, 67, 234–249. [Google Scholar] [CrossRef]
- Padirac, A.; Fujii, T.; Rondelez, Y. Bottom-up construction of in vitro switchable memories. Proc. Natl. Acad. Sci. USA 2012, 109, E3212–E3220. [Google Scholar] [CrossRef]
- Fujii, T.; Rondelez, Y. Predator-prey molecular ecosystems. ACS Nano 2013, 7, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Zadorin, A.S.; Rondelez, Y.; Gines, G.; Dilhas, V.; Urtel, G.; Zambrano, A.; Galas, J.-C.; Estevez-Torres, A. Synthesis and materialization of a reaction–diffusion French flag pattern. Nat. Chem. 2017, 9, 990–996. [Google Scholar] [CrossRef] [PubMed]
- Padirac, A.; Fujii, T.; Rondelez, Y. Quencher-free multiplexed monitoring of DNA reaction circuits. Nucleic Acids Res. 2012, 40, e118. [Google Scholar] [CrossRef] [PubMed]
- Soloveichik, D.; Seelig, G.; Winfree, E. DNA as a universal substrate for chemical kinetics. Proc. Natl. Acad. Sci. USA 2010, 107, 5393–5398. [Google Scholar] [CrossRef] [PubMed]
- Seelig, G.; Soloveichik, D.; Zhang, D.Y.; Winfree, E. Enzyme-Free Nucleic Acid Logic Circuits. Science 2006, 314, 1585–1588. [Google Scholar] [CrossRef]
- Qian, L.; Winfree, E. Scaling Up Digital Circuit Computation with DNA Strand Displacement Cascades. Science 2011, 332, 1196–1201. [Google Scholar] [CrossRef] [PubMed]
- Qian, L.; Winfree, E.; Bruck, J. Neural network computation with DNA strand displacement cascades. Nature 2011, 475, 368–372. [Google Scholar] [CrossRef]
- Cherry, K.M.; Qian, L. Scaling up molecular pattern recognition with DNA-based winner-take-all neural networks. Nature 2018, 559, 370–376. [Google Scholar] [CrossRef]
- Srinivas, N.; Parkin, J.; Seelig, G.; Winfree, E.; Soloveichik, D. Enzyme-free nucleic acid dynamical systems. Science 2017, 358, eaal2052. [Google Scholar] [CrossRef]
- Douglas, S.M.; Bachelet, I.; Church, G.M. A Logic-Gated Nanorobot for Targeted Transport of Molecular Payloads. Science 2012, 335, 831–834. [Google Scholar] [CrossRef]
- Isaacs, F.J.; Dwyer, D.J.; Collins, J.J. RNA synthetic biology. Nat Biotechnol. 2006, 24, 545–554. [Google Scholar] [CrossRef] [PubMed]
- Lease, R.A.; Belfort, M. A trans-acting RNA as a control switch in Escherichia coli: DsrA modulates function by forming alternative structures. Proc. Natl. Acad. Sci. USA 2000, 97, 9919–9924. [Google Scholar] [CrossRef] [PubMed]
- Isaacs, F.J.; Dwyer, D.J.; Ding, C.; Pervouchine, D.D.; Cantor, C.R.; Collins, J.J. Engineered riboregulators enable post-transcriptional control of gene expression. Nat. Biotechnol. 2004, 22, 841–847. [Google Scholar] [CrossRef] [PubMed]
- Green, A.A.; Silver, P.A.; Collins, J.J.; Yin, P. Toehold Switches: De-Novo-Designed Regulators of Gene Expression. Cell 2014, 159, 925–939. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Shen, L.; Wu, K.; Diehnelt, C.W.; Green, A.A. Low-cost detection of norovirus using paper-based cell-free systems and synbody-based viral enrichment. Synth. Biol. 2018, 3, ysy018. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, M.K.; Tan, X.; Dy, A.J.; Braff, D.; Akana, R.T.; Furuta, Y.; Donghia, N.; Ananthakrishnan, A.; Collins, J.J. A low-cost paper-based synthetic biology platform for analyzing gut microbiota and host biomarkers. Nat. Commun. 2018, 9, 3347. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Yin, P.; Green, A.A. Ribocomputing: Cellular Logic Computation Using RNA Devices. Biochemistry 2018, 57, 883–885. [Google Scholar] [CrossRef] [PubMed]
- Green, A.A.; Kim, J.; Ma, D.; Silver, P.A.; Collins, J.J.; Yin, P. Complex cellular logic computation using ribocomputing devices. Nature 2017, 548, 117–121. [Google Scholar] [CrossRef]
- Kim, J.; Zhou, Y.; Carlson, P.; Teichmann, M.; Simmel, F.C.; Silver, P.A.; Collins, J.J.; Lucks, J.B.; Yin, P.; Green, A.A. De-Novo-Designed Translational Repressors for Multi-Input Cellular Logic. bioRxiv 2018. [Google Scholar] [CrossRef]
- Carlson, P.D.; Glasscock, C.J.; Lucks, J.B. De novo Design of Translational RNA Repressors. bioRxiv 2018. [Google Scholar] [CrossRef]
- Takahashi, M.K.; Lucks, J.B. A modular strategy for engineering orthogonal chimeric RNA transcription regulators. Nucleic Acids Res. 2013, 41, 7577–7588. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Takahashi, M.K.; Lucks, J.B. Creating small transcription activating RNAs. Nat. Chem. Biol. 2015, 11, 214–220. [Google Scholar] [CrossRef] [PubMed]
- Chappell, J.; Westbrook, A.; Verosloff, M.; Lucks, J.B. Computational design of small transcription activating RNAs for versatile and dynamic gene regulation. Nat. Commun. 2017, 8, 1051. [Google Scholar] [CrossRef]
- Verosloff, M.; Chappell, J.; Perry, K.L.; Thompson, J.R.; Lucks, J.B. PLANT-Dx: A Molecular Diagnostic for Point of Use Detection of Plant Pathogens. ACS Synth. Biol. 2019, 8, 902–905. [Google Scholar] [CrossRef] [PubMed]
- Hoynes-O’Connor, A.; Hinman, K.; Kirchner, L.; Moon, T.S. De novo design of heat-repressible RNA thermosensors in E. coli. Nucleic Acids Res. 2015, 43, 6166–6179. [Google Scholar] [CrossRef]
- Lee, Y.J.; Moon, T.S. Design rules of synthetic non-coding RNAs in bacteria. Methods 2018, 143, 58–69. [Google Scholar] [CrossRef] [PubMed]
- Rodrigo, G.; Jaramillo, A. AutoBioCAD: Full biodesign automation of genetic circuits. ACS Synth. Biol. 2013, 2, 230–236. [Google Scholar] [CrossRef]
- Rodrigo, G.; Landrain, T.E.; Jaramillo, A. De novo automated design of small RNA circuits for engineering synthetic riboregulation in living cells. Proc. Natl. Acad. Sci. USA 2012, 109, 15271–15276. [Google Scholar] [CrossRef]
- Noireaux, V.; Libchaber, A. A vesicle bioreactor as a step toward an artificial cell assembly. Proc. Natl. Acad. Sci. USA 2004, 101, 17669–17674. [Google Scholar] [CrossRef]
- Minton, A.P. The Influence of Macromolecular Crowding and Macromolecular Confinement on Biochemical Reactions in Physiological Media. J. Biol. Chem. 2001, 276, 10577–10580. [Google Scholar] [CrossRef]
- Holtze, C.; Rowat, A.C.; Agresti, J.J.; Hutchison, J.B.; Angilè, F.E.; Schmitz, C.H.J.; Köster, S.; Duan, H.; Humphry, K.J.; Scanga, R.A.; et al. Biocompatible surfactants for water-in-fluorocarbon emulsions. Lab Chip 2008, 8, 1632. [Google Scholar] [CrossRef] [PubMed]
- Walter, A.; Gutknecht, J. Permeability of small nonelectrolytes through lipid bilayer membranes. J. Membr. Biol. 1986, 90, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, A. Water and nonelectrolyte permeability of lipid bilayer membranes. J. Gen. Physiol. 1976, 68, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Wei, C.; Pohorille, A. Permeation of Membranes by Ribose and Its Diastereomers. J. Am. Chem. Soc. 2009, 131, 10237–10245. [Google Scholar] [CrossRef] [PubMed]
- Abkarian, M.; Loiseau, E.; Massiera, G. Continuous droplet interface crossing encapsulation (cDICE) for high throughput monodisperse vesicle design. Soft Matter 2011, 7, 4610. [Google Scholar] [CrossRef]
- Ishikawa, K.; Sato, K.; Shima, Y.; Urabe, I.; Yomo, T. Expression of a cascading genetic network within liposomes. FEBS Lett. 2004, 576, 387–390. [Google Scholar] [CrossRef]
- Ota, S.; Yoshizawa, S.; Takeuchi, S. Microfluidic Formation of Monodisperse, Cell-Sized, and Unilamellar Vesicles. Angew. Chem. 2009, 121, 6655–6659. [Google Scholar] [CrossRef]
- Weiss, M.; Frohnmayer, J.P.; Benk, L.T.; Haller, B.; Janiesch, J.W.; Heitkamp, T.; Börsch, M.; Lira, R.B.; Dimova, R.; Lipowsky, R.; et al. Sequential bottom-up assembly of mechanically stabilized synthetic cells by microfluidics. Nat. Mater. 2018, 17, 89–96. [Google Scholar] [CrossRef]
- Matosevic, S.; Paegel, B.M. Stepwise Synthesis of Giant Unilamellar Vesicles on a Microfluidic Assembly Line. J. Am. Chem. Soc. 2011, 133, 2798–2800. [Google Scholar] [CrossRef]
- Tan, C.; Saurabh, S.; Bruchez, M.P.; Schwartz, R.; LeDuc, P. Molecular crowding shapes gene expression in synthetic cellular nanosystems. Nat. Nanotechnol. 2013, 8, 602–608. [Google Scholar] [CrossRef]
- Hansen, M.M.; Meijer, L.H.; Spruijt, E.; Maas, R.J.; Rosquelles, M.V.; Groen, J.; Heus, H.A.; Huck, W.T.S. Macromolecular crowding creates heterogeneous environments of gene expression in picolitre droplets. Nat. Nanotechnol. 2016, 11, 191–197. [Google Scholar] [CrossRef] [PubMed]
- Torre, P.; Keating, C.D.; Mansy, S.S. Multiphase Water-in-Oil Emulsion Droplets for Cell-Free Transcription–Translation. Langmuir 2014, 30, 5695–5699. [Google Scholar] [CrossRef]
- Weitz, M.; Kim, J.; Kapsner, K.; Winfree, E.; Franco, E.; Simmel, F.C. Diversity in the dynamical behaviour of a compartmentalized programmable biochemical oscillator. Nat. Chem. 2014, 6, 295–302. [Google Scholar] [CrossRef] [PubMed]
- Adamala, K.P.; Martin-Alarcon, D.A.; Guthrie-Honea, K.R.; Boyden, E.S. Engineering genetic circuit interactions within and between synthetic minimal cells. Nat. Chem. 2017, 9, 431–439. [Google Scholar] [CrossRef] [PubMed]
- de Souza, T.P.; Fahr, A.; Luisi, P.L.; Stano, P. Spontaneous encapsulation and concentration of biological macromolecules in liposomes: An intriguing phenomenon and its relevance in origins of life. J. Mol. Evol. 2014, 79, 179–192. [Google Scholar] [CrossRef] [PubMed]
- Klumpp, S.; Scott, M.; Pedersen, S.; Hwa, T. Molecular crowding limits translation and cell growth. Proc. Natl. Acad. Sci. USA 2013, 110, 16754–16759. [Google Scholar] [CrossRef]
- Albertsson, P.A. Partition of cell particles and macromolecules in polymer two-phase systems. Adv. Protein Chem. 1970, 24, 309–341. [Google Scholar]
- Aumiller, W.M., Jr.; Keating, C.D. Experimental models for dynamic compartmentalization of biomolecules in liquid organelles: Reversible formation and partitioning in aqueous biphasic systems. Adv. Colloid Interface Sci. 2017, 239, 75–87. [Google Scholar] [CrossRef]
- Hyman, A.A.; Weber, C.A.; Jülicher, F. Liquid-Liquid Phase Separation in Biology. Annu. Rev. Cell Dev. Biol. 2014, 30, 39–58. [Google Scholar] [CrossRef]
- Lentini, R.; Martín, N.Y.; Forlin, M.; Belmonte, L.; Fontana, J.; Cornella, M.; Martini, L.; Tamburini, S.; Bentley, W.E.; Jousson, O.; et al. Two-Way Chemical Communication between Artificial and Natural Cells. ACS Cent. Sci. 2017, 3, 117–123. [Google Scholar] [CrossRef]
- Bayoumi, M.; Bayley, H.; Maglia, G.; Sapra, K.T. Multi-compartment encapsulation of communicating droplets and droplet networks in hydrogel as a model for artificial cells. Sci. Rep. 2017, 7, 45167. [Google Scholar] [CrossRef] [PubMed]
- Elani, Y.; Trantidou, T.; Wylie, D.; Dekker, L.; Polizzi, K.; Law, R.V.; Ces, O. Constructing vesicle-based artificial cells with embedded living cells as organelle-like modules. Sci. Rep. 2018, 8, 4564. [Google Scholar] [CrossRef] [PubMed]
- Rampioni, G.; Leoni, L.; Mavelli, F.; Damiano, L.; Stano, P. Interfacing Synthetic Cells with Biological Cells: An Application of the Synthetic Method. In Proceedings of the 2018 Conference on Artificial Life: A Hybrid of the European Conference on Artificial Life (ECAL) and the International Conference on the Synthesis and Simulation of Living Systems (ALIFE), Tokyo, Japan, 23–27 July 2018; pp. 145–146. [Google Scholar]
- Lentini, R.; Santero, S.P.; Chizzolini, F.; Cecchi, D.; Fontana, J.; Marchioretto, M.; Del Bianco, C.; Terrell, J.L.; Spencer, A.C.; Martini, L.; et al. Integrating artificial with natural cells to translate chemical messages that direct E. coli behaviour. Nat. Commun. 2014, 5, 4012. [Google Scholar] [CrossRef] [PubMed]
- Bloomfield, V.A.; Crothers, D.M.; Tinoco, I.; Hearst, J.E.; Wemmer, D.E.; Killman, P.A.; Turner, D.H. Nucleic Acids: Structures, Properties, and Functions, 1st ed.; University Science Books: Sausalito, CA, USA, 2000; 627p. [Google Scholar]
- SantaLucia, J.; Hicks, D. The Thermodynamics of DNA Structural Motifs. Annu. Rev. Biophys. Biomol. Struct. 2004, 33, 415–440. [Google Scholar] [CrossRef] [PubMed]
- Rothemund, P.W.K.; Ekani-Nkodo, A.; Papadakis, N.; Kumar, A.; Fygenson, D.K.; Winfree, E. Design and Characterization of Programmable DNA Nanotubes. J. Am. Chem. Soc. 2004, 126, 16344–16352. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Bell, N.A.W.; Hernández-Ainsa, S.; Thacker, V.V.; Thackray, A.M.; Bujdoso, R.; Keyser, U.F. Single Protein Molecule Detection by Glass Nanopores. ACS Nano 2013, 7, 4129–4134. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.Y.; Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 2011, 3, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Yurke, B.; Turberfield, A.J.; Mills, A.P., Jr.; Simmel, F.C.; Neumann, J.L. A DNA-fuelled molecular machine made of DNA. Nature 2000, 406, 605–608. [Google Scholar] [CrossRef]
- Dirks, R.M.; Pierce, N.A. Triggered amplification by hybridization chain reaction. Proc. Natl. Acad. Sci. USA 2004, 101, 15275–15278. [Google Scholar] [CrossRef]
- Keum, J.W.; Bermudez, H. Enhanced resistance of DNA nanostructures to enzymatic digestion. Chem. Commun. 2009, 7036–7038. [Google Scholar] [CrossRef]
- Castro, C.E.; Kilchherr, F.; Kim, D.N.; Shiao, E.L.; Wauer, T.; Wortmann, P.; Bathe, M.; Dietz, H. A primer to scaffolded DNA origami. Nat. Methods 2011, 8, 221–229. [Google Scholar] [CrossRef] [PubMed]
- Mei, Q.; Wei, X.; Su, F.; Liu, Y.; Youngbull, C.; Johnson, R.; Lindsay, S.; Yan, H.; Meldrum, D. Stability of DNA Origami Nanoarrays in Cell Lysate. Nano Lett. 2011, 11, 1477–1482. [Google Scholar] [CrossRef] [PubMed]
- Klocke, M.A.; Garamella, J.; Subramanian, H.K.K.; Noireaux, V.; Franco, E. Engineering DNA nanotubes for resilience in an E. coli TXTL system. Synth. Biol. 2018, 3, ysy001. [Google Scholar] [CrossRef]
- Bramsen, J.B.; Laursen, M.B.; Nielsen, A.F.; Hansen, T.B.; Bus, C.; Langkjær, N.; Babu, B.R.; Højland, T.; Abramov, M.; Van Aerschot, A.; et al. A large-scale chemical modification screen identifies design rules to generate siRNAs with high activity, high stability and low toxicity. Nucleic Acids Res. 2009, 37, 2867–2881. [Google Scholar] [CrossRef] [PubMed]
- Conway, J.W.; McLaughlin, C.K.; Castor, K.J.; Sleiman, H. DNA nanostructure serum stability: Greater than the sum of its parts. Chem. Commun. 2013, 49, 1172. [Google Scholar] [CrossRef]
- Hahn, J.; Wickham, S.F.J.; Shih, W.M.; Perrault, S.D. Addressing the Instability of DNA Nanostructures in Tissue Culture. ACS Nano 2014, 8, 8765–8775. [Google Scholar] [CrossRef] [PubMed]
- Goltry, S.; Hallstrom, N.; Clark, T.; Kuang, W.; Lee, J.; Jorcyk, C.; Knowlton, W.B.; Yurke, B.; Hughes, W.L.; Graugnard, E. DNA topology influences molecular machine lifetime in human serum. Nanoscale 2015, 7, 10382–10390. [Google Scholar] [CrossRef]
- Zhang, D.Y.; Turberfield, A.J.; Yurke, B.; Winfree, E. Engineering Entropy-Driven Reactions and Networks Catalyzed by DNA. Science 2007, 318, 1121–1125. [Google Scholar] [CrossRef]
- Graugnard, E.; Cox, A.; Lee, J.; Jorcyk, C.; Yurke, B.; Hughes, W.L. Operation of a DNA-Based Autocatalytic Network in Serum. In Proceedings of the 16th International Conference on DNA Computing and Molecular Programming, Hong Kong, China, 14–17 June 2010; pp. 83–88. [Google Scholar]
- Fern, J.; Schulman, R. Design and Characterization of DNA Strand-Displacement Circuits in Serum-Supplemented Cell Medium. ACS Synth. Biol. 2017, 6, 1774–1783. [Google Scholar] [CrossRef]
- Schaffter, S.W.; Green, L.N.; Schneider, J.; Subramanian, H.K.K.; Schulman, R.; Franco, E. T7 RNA polymerase non-specifically transcribes and induces disassembly of DNA nanostructures. Nucleic Acids Res. 2018, 46, 5332–5343. [Google Scholar] [CrossRef]
- Montagne, K.; Gines, G.; Fujii, T.; Rondelez, Y. Boosting functionality of synthetic DNA circuits with tailored deactivation. Nat. Commun. 2016, 7, 13474. [Google Scholar] [CrossRef] [PubMed]
- Westbrook, A.; Tang, X.; Marshall, R.; Maxwell, C.S.; Chappell, J.; Agrawal, D.K.; Dunlop, M.J.; Noireaux, V.; Beisel, C.L.; Lucks, J.; et al. Distinct timescales of RNA regulators enable the construction of a genetic pulse generator. Biotechnol. Bioeng. 2019, 116, 1139–1151. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.Y.; Varner, J.D.; Lucks, J.B. Generating Effective Models and Parameters for RNA Genetic Circuits. ACS Synth. Biol. 2015, 4, 914–926. [Google Scholar] [CrossRef] [PubMed]
- Joesaar, A.; Yang, S.; Bogels, B.; van der Linden, A.; Pieters, P.; Kumar, B.V.V.S.P.; Dalchau, N.; Phillips, A.; Mann, S.; de Greef, T.F.A. Distributed DNA-based Communication in Populations of Synthetic Protocells. Nat. Nanotechnol. 2019, 14, 369–378. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jeong, D.; Klocke, M.; Agarwal, S.; Kim, J.; Choi, S.; Franco, E.; Kim, J. Cell-Free Synthetic Biology Platform for Engineering Synthetic Biological Circuits and Systems. Methods Protoc. 2019, 2, 39. https://doi.org/10.3390/mps2020039
Jeong D, Klocke M, Agarwal S, Kim J, Choi S, Franco E, Kim J. Cell-Free Synthetic Biology Platform for Engineering Synthetic Biological Circuits and Systems. Methods and Protocols. 2019; 2(2):39. https://doi.org/10.3390/mps2020039
Chicago/Turabian StyleJeong, Dohyun, Melissa Klocke, Siddharth Agarwal, Jeongwon Kim, Seungdo Choi, Elisa Franco, and Jongmin Kim. 2019. "Cell-Free Synthetic Biology Platform for Engineering Synthetic Biological Circuits and Systems" Methods and Protocols 2, no. 2: 39. https://doi.org/10.3390/mps2020039
APA StyleJeong, D., Klocke, M., Agarwal, S., Kim, J., Choi, S., Franco, E., & Kim, J. (2019). Cell-Free Synthetic Biology Platform for Engineering Synthetic Biological Circuits and Systems. Methods and Protocols, 2(2), 39. https://doi.org/10.3390/mps2020039




