Fermentation and Microbial Community of Maize Silage Inoculated with Lentilactobacillus buchneri NCIMB 40788 and Contaminated with Bacillus and Clostridium Spore Formers
Abstract
:1. Introduction
2. Materials and Methods
2.1. Silage Preparation and Sampling
2.2. DNA Extraction and 16S rRNA Gene Amplicon Sequencing
2.3. Water-Soluble Chemical Compound Quantification via NMR
2.4. Microbial Analysis of Spore Former Survival
2.5. Bioinformatics
2.6. Statistical Analysis
3. Results
3.1. Physical and Chemical Properties of the Silage
3.2. Diversity of the Bacterial and Fungal Communities of Silage
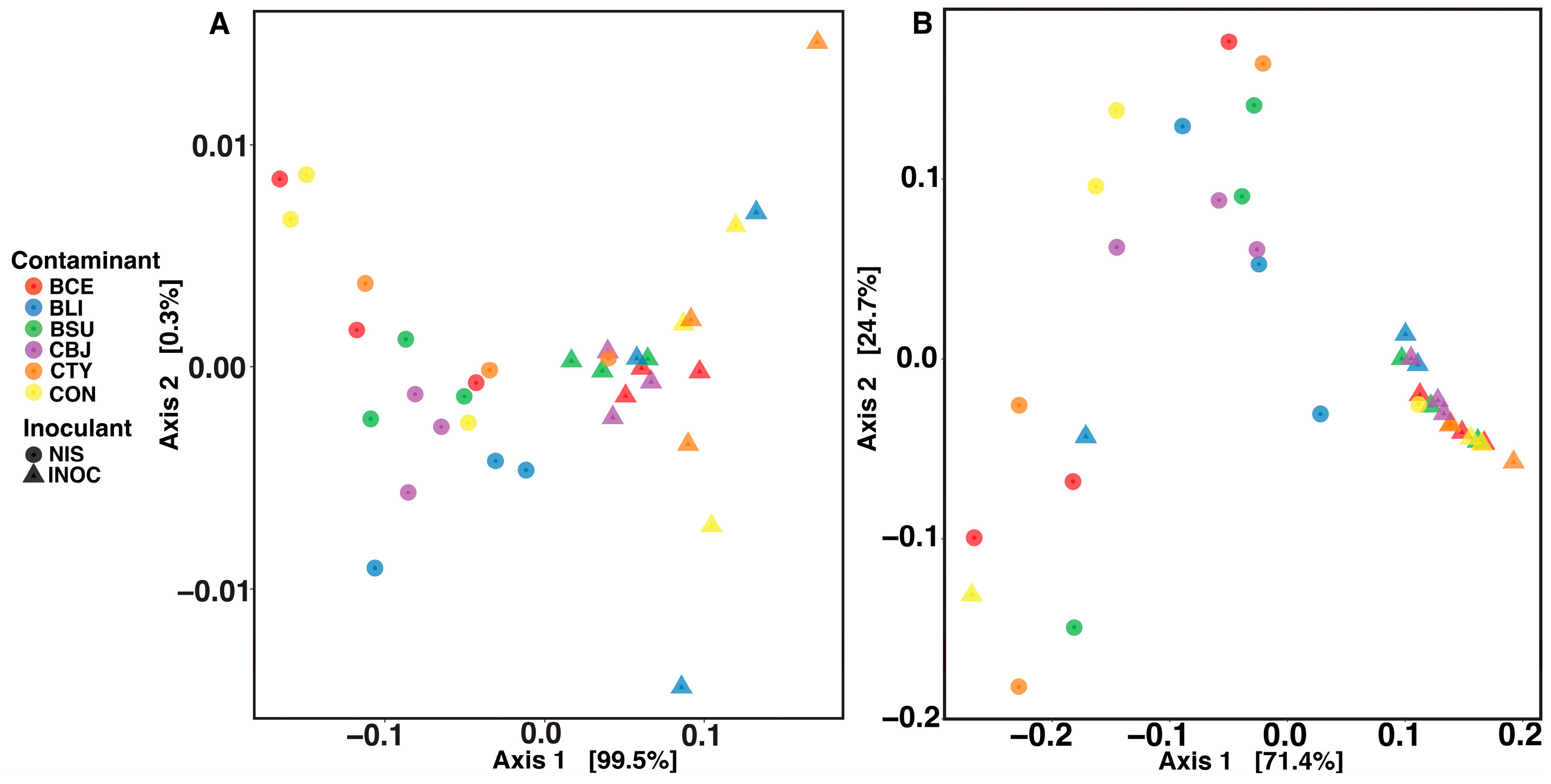
3.3. Impact of Inoculant and Spore Contaminants on the Microbial Communities of Silage
3.4. Changes in Cell and Spore Counts following Inoculation and Contamination
3.5. Correlations between Microbial and Chemical Variables
4. Discussion
4.1. Effects of Inoculation on Silage Quality
4.2. Effects of Bacillus and Clostridium Contaminants on Silage Quality
4.3. Effects of B. cereus on Silage Quality
4.4. Effects of B. licheniformis on Silage Quality
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Vissers, M.M.M.; Te Giffel, M.C.T.; Driehuis, F.; Jong, P.D.; Lankveld, J.M.G. Minimizing the level of Bacillus cereus spores in farm tank milk. J. Dairy Sci. 2007, 90, 3286–3293. [Google Scholar] [CrossRef] [PubMed]
- Queiroz, O.C.M.; Ogunade, I.M.; Weinberg, Z.; Adesogan, A.T. Silage Review: Foodborne pathogens in silage and their mitigation by silage additives. J. Dairy Sci. 2018, 101, 4132–4142. [Google Scholar] [CrossRef] [PubMed]
- McDonald, P.; Henderson, N.; Heron, S. The Biochemistry of Silage, 2nd ed.; Chalcombe: Marlow, England, 1991. [Google Scholar]
- Li, M.M.; White, R.R.; Guan, L.L.; Harthan, L.; Hanigan, M.D. Metatranscriptomic analyses reveal ruminal pH regulates fiber degradation and fermentation by shifting the microbial community and gene expression of carbohydrate-active enzymes. Anim. Microbiome 2021, 3, 32. [Google Scholar] [CrossRef] [PubMed]
- D’Incecco, P.; Pellegrino, L.; Hogenboom, J.A.; Cocconcelli, P.S.; Bassi, D. The late blowing defect of hard cheeses: Behaviour of cells and spores of Clostridium tyrobutyricum throughout the cheese manufacturing and ripening. LWT 2018, 87, 134–141. [Google Scholar] [CrossRef]
- Klijn, N.; Nieuwenhof, F.F.; Hoolwerf, J.D.; van der Waals, C.B.; Weerkamp, A.H. Identification of Clostridium tyrobutyricum as the causative agent of late blowing in cheese by species-specific PCR amplification. Appl. Environ. Microbiol. 1995, 61, 2919–2924. [Google Scholar] [CrossRef]
- Buehner, K.P.; Anand, S.; Djira, G.D.; Garcia, A. Corrigendum to “Prevalence of thermoduric bacteria and spores on 10 midwest dairy farms”. J. Dairy Sci. 2014, 97, 8009–8016. [Google Scholar] [CrossRef]
- Te Giffel, M.C.; Wagendorp, A.; Herrewegh, A.; Driehuis, F. Bacterial spores in silage and raw milk. Antonie Van Leeuwenhoek 2002, 81, 625–630. [Google Scholar] [CrossRef]
- Heyndrickx, M.; Scheldeman, P. Bacilli associated with spoilage in dairy products and other food. In Applications and Systematics of Bacillus and Relatives; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2002; pp. 64–82. ISBN 978-0-470-69674-3. [Google Scholar]
- Beecher, D.J.; Schoeni, J.L.; Wong, A.C. Enterotoxic activity of Hemolysin BL from Bacillus cereus. Infect. Immun. 1995, 63, 4423–4428. [Google Scholar] [CrossRef]
- Ehling-Schulz, M.; Svensson, B.; Guinebretiere, M.-H.; Lindbäck, T.; Andersson, M.; Schulz, A.; Fricker, M.; Christiansson, A.; Granum, P.E.; Märtlbauer, E.; et al. Emetic toxin formation of Bacillus cereus is restricted to a single evolutionary lineage of closely related strains. Microbiol. Read. Engl. 2005, 151, 183–197. [Google Scholar] [CrossRef]
- Griffiths, M. Improving the Safety and Quality of Milk: Milk Production and Processing, 1st ed.; Woodhead Publishing: Thorston, UK, 2010; Volume 1, ISBN 978-1-84569-942-0. [Google Scholar]
- Vidic, J.; Chaix, C.; Manzano, M.; Heyndrickx, M. Food sensing: Detection of Bacillus cereus spores in dairy products. Biosensors 2020, 10, 15. [Google Scholar] [CrossRef]
- Kumari, S.; Sarkar, P.K. Bacillus cereus hazard and control in industrial dairy processing environment. Food Control 2016, 69, 20–29. [Google Scholar] [CrossRef]
- Scheldeman, P.; Herman, L.; Foster, S.; Heyndrickx, M. Bacillus sporothermodurans and other highly heat-resistant spore formers in milk. J. Appl. Microbiol. 2006, 101, 542–555. [Google Scholar] [CrossRef] [PubMed]
- Reginensi, S.M.; González, M.J.; Olivera, J.A.; Sosa, M.; Juliano, P.; Bermúdez, J. RAPD-based screening for spore-forming bacterial populations in Uruguayan commercial powdered milk. Int. J. Food Microbiol. 2011, 148, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Gopal, N.; Hill, C.; Ross, P.R.; Beresford, T.P.; Fenelon, M.A.; Cotter, P.D. The prevalence and control of Bacillus and related spore-forming bacteria in the dairy industry. Front. Microbiol. 2015, 6, 1418. [Google Scholar] [CrossRef]
- Parkinson, T.J. Specific infectious diseases causing infertility and subfertility in cattle. In Veterinary Reproduction and Obstetrics, 10th ed.; Noakes, D.E., Parkinson, T.J., England, G.C.W., Eds.; W.B. Saunders: St. Louis, MO, USA, 2019; pp. 434–466. ISBN 978-0-7020-7233-8. [Google Scholar]
- Scheldeman, P.; Pil, A.; Herman, L.; De Vos, P.; Heyndrickx, M. Incidence and diversity of potentially highly heat-resistant spores isolated at dairy farms. Appl. Environ. Microbiol. 2005, 71, 1480–1494. [Google Scholar] [CrossRef]
- Vaerewijck, M.J.; De Vos, P.; Lebbe, L.; Scheldeman, P.; Hoste, B.; Heyndrickx, M. Occurrence of Bacillus sporothermodurans and other aerobic spore-forming species in feed concentrate for dairy cattle. J. Appl. Microbiol. 2001, 91, 1074–1084. [Google Scholar] [CrossRef]
- Driehuis, F.; Wilkinson, J.M.; Jiang, Y.; Ogunade, I.; Adesogan, A.T. Silage review: Animal and human health risks from silage. J. Dairy Sci. 2018, 101, 4093–4110. [Google Scholar] [CrossRef]
- Gleeson, D.; O’Connell, A.; Jordan, K. Review of potential sources and control of thermoduric bacteria in bulk-tank milk. Ir. J. Agric. Food Res. 2013, 52, 217–227. Available online: http://www.jstor.org/stable/23631033 (accessed on 1 May 2023).
- Heyndrickx, M. The importance of endospore-forming bacteria originating from soil for contamination of industrial food processing. Appl. Environ. Soil Sci. 2011, 2011, e561975. [Google Scholar] [CrossRef]
- Oliveira, A.S.; Weinberg, Z.G.; Ogunade, I.M.; Cervantes, A.A.P.; Arriola, K.G.; Jiang, Y.; Kim, D.; Li, X.; Gonçalves, M.C.M.; Vyas, D.; et al. Meta-analysis of effects of inoculation with homofermentative and facultative heterofermentative lactic acid bacteria on silage fermentation, aerobic stability, and the performance of dairy cows. J. Dairy Sci. 2017, 100, 4587–4603. [Google Scholar] [CrossRef]
- Kleinschmit, D.H.; Kung, L. A Meta-analysis of the effects of Lactobacillus buchneri on the fermentation and aerobic stability of corn and grass and small-grain silages. J. Dairy Sci. 2006, 89, 4005–4013. [Google Scholar] [CrossRef]
- Drouin, P.; Tremblay, J.; Renaud, J.; Apper, E. Microbiota succession during aerobic stability of maize silage inoculated with Lentilactobacillus buchneri NCIMB 40788 and Lentilactobacillus hilgardii CNCM-I-4785. MicrobiologyOpen 2021, 10, e1153. [Google Scholar] [CrossRef]
- Nair, J.; Huaxin, N.; Andrada, E.; Yang, H.-E.; Chevaux, E.; Drouin, P.; McAllister, T.A.; Wang, Y. Effects of inoculation of corn silage with Lactobacillus hilgardii and Lactobacillus buchneri on silage quality, aerobic stability, nutrient digestibility, and growth performance of growing beef cattle. J. Anim. Sci. 2020, 98, skaa267. [Google Scholar] [CrossRef]
- Arriola, K.G.; Oliveira, A.S.; Jiang, Y.; Kim, D.; Silva, H.M.; Kim, S.C.; Amaro, F.X.; Ogunade, I.M.; Sultana, H.; Pech Cervantes, A.A.; et al. Meta-analysis of effects of inoculation with Lactobacillus buchneri, with or without other bacteria, on silage fermentation, aerobic stability, and performance of dairy cows. J. Dairy Sci. 2021, 104, 7653–7670. [Google Scholar] [CrossRef] [PubMed]
- Huisden, C.M.; Adesogan, A.T.; Kim, S.C.; Ososanya, T. Effect of applying molasses or inoculants containing homofermentative or heterofermentative bacteria at two rates on the fermentation and aerobic stability of corn silage. J. Dairy Sci. 2009, 92, 690–697. [Google Scholar] [CrossRef]
- Rosenquist, H.; Hansen, Å. The antimicrobial effect of organic acids, sour dough and nisin against Bacillus subtilis and B. licheniformis isolated from wheat bread. J. Appl. Microbiol. 1998, 85, 621–631. [Google Scholar] [CrossRef]
- Adesogan, A.T.; Krueger, N.; Salawu, M.B.; Dean, D.B.; Staples, C.R. The influence of treatment with dual purpose bacterial inoculants or soluble carbohydrates on the fermentation and aerobic stability of bermudagrass. J. Dairy Sci. 2004, 87, 3407–3416. [Google Scholar] [CrossRef] [PubMed]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Bellemain, E.; Carlsen, T.; Brochmann, C.; Coissac, E.; Taberlet, P.; Kauserud, H. ITS as an environmental DNA barcode for fungi: An in silico approach reveals potential PCR biases. BMC Microbiol. 2010, 10, 189. [Google Scholar] [CrossRef]
- Deshpande, V.; Wang, Q.; Greenfield, P.; Charleston, M.; Porras-Alfaro, A.; Kuske, C.R.; Cole, J.R.; Midgley, D.J.; Tran-Dinh, N. Fungal identification using a bayesian classifier and the Warcup training set of internal transcribed spacer sequences. Mycologia 2016, 108, 1–5. [Google Scholar] [CrossRef]
- Løvdal, I.S.; Hovda, M.B.; Granum, P.E.; Rosnes, J.T. Promoting Bacillus cereus spore germination for subsequent inactivation by mild heat treatment. J. Food Prot. 2011, 74, 2079–2089. [Google Scholar] [CrossRef]
- R Core Team. A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. Available online: https://www.r-project.org/ (accessed on 1 May 2023).
- Santos, A. de O. dos; Ávila, C.L. da S.; Soares, C.; Carvalho, B.F.; Schwan, R.F.; Lima, N. Lactic acid bacteria diversity in corn silage produced in Minas Gerais (Brazil). Ann. Microbiol. 2019, 69, 1445–1459. [Google Scholar] [CrossRef]
- Feng, Q.; Shi, W.; Chen, S.; Degen, A.A.; Qi, Y.; Yang, F.; Zhou, J. Addition of organic acids and Lactobacillus acidophilus to the leguminous forage Chamaecrista rotundifolia improved the quality and decreased harmful bacteria of the silage. Animals 2022, 12, 2260. [Google Scholar] [CrossRef] [PubMed]
- Gilliland, S.E.; Speck, M.L. Antagonistic action of Lactobacillus acidophilus toward intestinal and foodborne pathogens in associative cultures. J. Food Prot. 1977, 40, 820–823. [Google Scholar] [CrossRef] [PubMed]
- Borreani, G.; Tabacco, E.; Schmidt, R.J.; Holmes, B.J.; Muck, R.E. Silage review: Factors affecting dry matter and quality losses in silages. J. Dairy Sci. 2018, 101, 3952–3979. [Google Scholar] [CrossRef] [PubMed]
- Danner, H.; Holzer, M.; Mayrhuber, E.; Braun, R. Acetic acid increases stability of silage under aerobic conditions. Appl. Environ. Microbiol. 2003, 69, 562–567. [Google Scholar] [CrossRef]
- Guan, H.; Ran, Q.; Li, H.; Zhang, X. Succession of microbial communities of corn silage inoculated with heterofermentative lactic acid bacteria from ensiling to aerobic exposure. Fermentation 2021, 7, 258. [Google Scholar] [CrossRef]
- Kung, L.; Shaver, R.D.; Grant, R.J.; Schmidt, R.J. Silage review: Interpretation of chemical, microbial, and organoleptic components of silages. J. Dairy Sci. 2018, 101, 4020–4033. [Google Scholar] [CrossRef]
- Vissers, M.M.M.; Driehuis, F.; Te Giffel, M.C.; De Jong, P.; Lankveld, J.M.G. Concentrations of butyric acid bacteria spores in silage and relationships with aerobic deterioration. J. Dairy Sci. 2007, 90, 928–936. [Google Scholar] [CrossRef]
- Aksu, T.; Baytok, E.; Bolat, D. Effects of a bacterial silage inoculant on corn silage fermentation and nutrient digestibility. Small Rumin. Res. 2004, 55, 249–252. [Google Scholar] [CrossRef]
- Dürre, P. Physiology and sporulation in Clostridium. Microbiol. Spectr. 2014, 2, 315–329. [Google Scholar] [CrossRef]
- Fenner, H.; Barnes, H.D. Improved method for determining dry matter in silage. J. Dairy Sci. 1965, 48, 1324–1328. [Google Scholar] [CrossRef]
- Li, R.; Jiang, D.; Zheng, M.; Tian, P.; Zheng, M.; Xu, C. Microbial community dynamics during alfalfa silage with or without clostridial fermentation. Sci. Rep. 2020, 10, 17782. [Google Scholar] [CrossRef]
- He, J.; Chu, Y.; Li, J.; Meng, Q.; Liu, Y.; Jin, J.; Wang, Y.; Wang, J.; Huang, B.; Shi, L.; et al. Intestinal butyrate-metabolizing species contribute to autoantibody production and bone erosion in rheumatoid arthritis. Sci. Adv. 2022, 8, eabm1511. [Google Scholar] [CrossRef] [PubMed]
- Simpson, K.M.; Callan, R.J.; Van Metre, D.C. Clostridial abomasitis and enteritis in ruminants. Vet. Clin. North Am. Food Anim. Pract. 2018, 34, 155–184. [Google Scholar] [CrossRef] [PubMed]
- Shaver, R.D.; Erdman, R.A.; O’connor, A.M.; Vandersall, J.H. Effects of silage pH on voluntary intake of corn silage and alfalfa haylage. J. Dairy Sci. 1985, 68, 338–346. [Google Scholar] [CrossRef]
- Guo, X.S.; Ke, W.C.; Ding, W.R.; Ding, L.M.; Xu, D.M.; Wang, W.W.; Zhang, P.; Yang, F.Y. Profiling of metabolome and bacterial community dynamics in ensiled Medicago sativa inoculated without or with Lactobacillus plantarum or Lactobacillus buchneri. Sci. Rep. 2018, 8, 357. [Google Scholar] [CrossRef] [PubMed]
- Borreani, G.; Ferrero, F.; Nucera, D.; Casale, M.; Piano, S.; Tabacco, E. Dairy farm management practices and the risk of contamination of tank milk from Clostridium spp. and Paenibacillus spp. spores in silage, total mixed ration, dairy cow feces, and raw milk. J. Dairy Sci. 2019, 102, 8273–8289. [Google Scholar] [CrossRef]
- Canadian Dairy Information Centre. Acts, Regulations, Codes and Standards. Available online: https://agriculture.canada.ca/en/sector/animal-industry/canadian-dairy-information-centre/acts-regulations-codes-standards. (accessed on 1 May 2023).
- Health Canada. Microbial guidelines for ready-to-eat foods–A guide for the conveyance industry and environmental health officers (EHO). Health Canada 2010. Available online: https://publications.gc.ca/site/archivee-archived.html?url=https://publications.gc.ca/collections/collection_2014/sc-hc/H164-167-2013-eng.pdf (accessed on 1 May 2023).
- Borreani, G.; Tabacco, E. Improving corn silage quality in the top layer of farm bunker silos through the use of a next-generation barrier film with high impermeability to oxygen. J. Dairy Sci. 2014, 97, 2415–2426. [Google Scholar] [CrossRef]
- Muck, R.E.; Nadeau, E.M.G.; McAllister, T.A.; Contreras-Govea, F.E.; Santos, M.C.; Kung, L. Silage review: Recent advances and future uses of silage additives. J. Dairy Sci. 2018, 101, 3980–4000. [Google Scholar] [CrossRef]
- Borreani, G.; Tabacco, E. The relationship of silage temperature with the microbiological status of the face of corn silage bunkers. J. Dairy Sci. 2010, 93, 2620–2629. [Google Scholar] [CrossRef]
- Nielsen, P.; Sørensen, J. Multi-target and medium-independent fungal antagonism by hydrolytic enzymes in Paenibacillus polymyxa and Bacillus pumilus strains from barley rhizosphere. FEMS Microbiol. Ecol. 1997, 22, 183–192. [Google Scholar] [CrossRef]
- Kim, Y.-K.; Lee, S.-C.; Cho, Y.-Y.; Oh, H.-J.; Ko, Y.H. Isolation of cellulolytic Bacillus subtilis strains from agricultural environments. ISRN Microbiol. 2012, 2012, 650563. [Google Scholar] [CrossRef]
- Stokes, M.R. Effects of an enzyme mixture, an inoculant, and their interaction on silage fermentation and dairy production. J. Dairy Sci. 1992, 75, 764–773. [Google Scholar] [CrossRef]
- Bai, J.; Franco, M.; Ding, Z.; Hao, L.; Ke, W.; Wang, M.; Xie, D.; Li, Z.; Zhang, Y.; Ai, L.; et al. Effect of Bacillus amyloliquefaciens and Bacillus subtilis on fermentation, dynamics of bacterial community and their functional shifts of whole-plant corn silage. J. Anim. Sci. Biotechnol. 2022, 13, 7. [Google Scholar] [CrossRef]
- Zhang, F.; Wang, X.; Lu, W.; Li, F.; Ma, C. Improved quality of corn silage when combining cellulose-decomposing bacteria and Lactobacillus buchneri during silage fermentation. BioMed. Res. Int. 2019, 2019, 4361358. [Google Scholar] [CrossRef]
- Zhu, Y.; Xiong, H.; Wen, Z.; Tian, H.; Chen, Y.; Wu, L.; Guo, Y.; Sun, B. Effects of different concentrations of Lactobacillus plantarum and Bacillus licheniformis on silage quality, in vitro fermentation and microbial community of hybrid Pennisetum. Animals 2022, 12, 1752. [Google Scholar] [CrossRef]
- Bonaldi, D.S.; Carvalho, B.F.; Ávila, C.L.D.S.; Silva, C.F. Effects of Bacillus subtilis and its metabolites on corn silage quality. Lett. Appl. Microbiol. 2021, 73, 46–53. [Google Scholar] [CrossRef]
- Wang, W.; Fan, G.; Li, X.; Fu, Z.; Liang, X.; Sun, B. Application of Wickerhamomyces anomalus in simulated solid-state fermentation for baijiu production: Changes of microbial community structure and flavor metabolism. Front. Microbiol. 2020, 11, 598758. [Google Scholar] [CrossRef]
- Phuengjayaem, S.; Kuncharoen, N.; Booncharoen, A.; Ongpipattanakul, B.; Tanasupawat, S. Genome analysis and optimization of γ-aminobutyric acid (GABA) production by lactic acid bacteria from plant materials. J. Gen. Appl. Microbiol. 2021, 67, 150–161. [Google Scholar] [CrossRef]
- Os, M.V.; Jailler, M.; Dulphy, J.P. The Influence of ammonia, biogenic amines and γ-aminobutyric acid on grass silage intake in sheep. Br. J. Nutr. 1996, 76, 347–358. [Google Scholar] [CrossRef]
- Tanizawa, Y.; Kobayashi, H.; Nomura, M.; Sakamoto, M.; Arita, M.; Nakamura, Y.; Ohkuma, M.; Tohno, M. Lactobacillus buchneri subsp. silagei subsp. nov., isolated from rice grain silage. Int. J. Syst. Evol. Microbiol. 2020, 70, 3111–3116. [Google Scholar] [CrossRef]
- Cho, Y.R.; Chang, J.Y.; Chang, H.C. Production of gamma-aminobutyric acid (GABA) bSy Lactobacillus buchneri isolated from kimchi and its neuroprotective effect on neuronal cells. J. Microbiol. Biotechnol. 2007, 17, 104–109. [Google Scholar]
- Renchinkhand, G.; Magsar, U.; Bae, H.C.; Choi, S.-H.; Nam, M.S. Identification of β-glucosidase activity of Lentilactobacillus buchneri URN103L and its potential to convert ginsenoside Rb1 from panax ginseng. Foods 2022, 11, 529. [Google Scholar] [CrossRef]
- Dhakal, R.; Bajpai, V.K.; Baek, K.-H. Production of gaba (γ–aminobutyric acid) by microorganisms: A review. Braz. J. Microbiol. 2012, 43, 1230–1241. [Google Scholar] [CrossRef]
- Bach, B.; Meudec, E.; Lepoutre, J.-P.; Rossignol, T.; Blondin, B.; Dequin, S.; Camarasa, C. New insights into γ-aminobutyric acid catabolism: Evidence for γ-hydroxybutyric acid and polyhydroxybutyrate synthesis in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2009, 75, 4231–4239. [Google Scholar] [CrossRef]


| Parameter | Inoculation | Contaminant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NIS | INOC | BCE | BSU | BLI | CTY | CBJ | CON | P (I) | P (C) | P (I × C) | SEM | |
| FM losses (%) | 38.34 1 | 50.45 | 44.55 | 43.40 | 44.79 | 43.94 | 44.30 | 45.41 | <0.001 2 | 0.950 3 | 0.671 4 | 0.97 5 |
| DM (g/kg) | 310.4 | 304.3 | 308.9 | 307.8 | 308.9 | 306.1 | 305.4 | 306.9 | <0.001 | 0.524 | 0.978 | 0.07 |
| DM losses (%) | 6.40 | 8.49 | 6.97 | 7.29 | 6.97 | 7.79 | 8.03 | 7.59 | <0.001 | 0.536 | 0.975 | 0.23 |
| AS10+2 (hours) 6 | 69.55 | 159.47 | 120.10 | 119.74 | 103.38 | 122.31 | 109.18 | 107.10 | <0.001 | 0.761 | 0.409 | 7.21 |
| AS10+3 (hours) 7 | 73.77 | 170.84 | 129.59 | 126.83 | 109.60 | 130.36 | 115.10 | 116.97 | <0.001 | 0.694 | 0.317 | 7.72 |
| Feature | Inoculation | Contaminant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NIS | INOC | BCE | BSU | BLI | CTY | CBJ | CON | P (I) | P (C) | P (I × C) | SEM | |
| At opening (100 d storage) | ||||||||||||
| pH | 3.79 1 | 3.95 | 3.83 cd | 3.82 d | 3.85 cd | 3.93 a | 3.88 bc | 3.91 ab | <0.001 3 | <0.001 4 | 0.003 5 | 0.02 6 |
| Lactic acid | 39.61 | 25.84 | 37.03 a | 32.58 ab | 32.14 ab | 28.96 ab | 32.08 ab | 33.55 b | <0.001 | 0.032 | 0.186 | 1.37 |
| Acetic acid | 20.47 | 31.35 | 27.99 a | 25.10 ab | 24.53 ab | 27.04 ab | 24.59 ab | 26.22 b | <0.001 | 0.022 | 0.104 | 0.99 |
| Propionic acid | 0.09 | 1.37 | 0.67 | 0.58 | 0.76 | 1.10 | 0.46 | 0.80 | <0.001 | 0.614 | 0.589 | 0.15 |
| Butyric acid | 0.22 | 2.30 | 1.33 | 1.29 | 1.14 | 1.44 | 1.01 | 1.35 | <0.001 | 0.306 | 0.275 | 0.19 |
| Malonic acid | 0.97 | 0.77 | 1.60 a | 1.48 a | 0.81 b | 0.48 c | 0.41 c | 0.43 c | 0.002 | <0.001 | <0.001 | 0.09 |
| Methylamine | 0.82 | 1.03 | 1.12 a | 1.23 a | 1.22 a | 1.17 a | 0.22 b | 0.57 b | 0.013 | <0.001 | 0.097 | 0.08 |
| γ-aminobutyric acid | 0.89 | 1.20 | 1.13 | 0.82 | 0.69 | 1.15 | 1.22 | 1.28 | 0.094 | 0.362 | 0.323 | 0.10 |
| Amino acids | 14.81 | 17.08 | 16.31 | 15.92 | 16.07 | 16.19 | 15.09 | 16.10 | <0.001 | 0.574 | 0.291 | 0.28 |
| AGGX 2 | 4.20 | 0.89 | 2.59 | 2.82 | 2.44 | 1.81 | 2.77 | 2.87 | <0.001 | 0.070 | 0.306 | 0.30 |
| Aerobic exposure (3 d) | ||||||||||||
| pH | 3.65 | 3.71 | 3.69 | 3.73 | 3.67 | 3.70 | 3.68 | 3.73 | <0.001 | 0.560 | 0.792 | 0.01 |
| Lactic acid | 30.91 | 24.35 | 24.07 | 27.61 | 29.80 | 25.38 | 29.21 | 29.72 | 0.015 | 0.688 | 0.820 | 1.28 |
| Acetic acid | 18.48 | 25.16 | 23.69 | 21.04 | 20.11 | 23.47 | 21.15 | 21.46 | <0.001 | 0.338 | 0.393 | 0.78 |
| Butyric acid | 0.10 | 0.49 | 0.26 | 0.24 | 0.42 | 0.34 | 0.22 | 0.30 | 0.025 | 0.980 | 0.999 | 0.08 |
| Propionic acid | 0.13 | 1.00 | 0.59 | 0.56 | 0.40 | 0.81 | 0.38 | 0.66 | <0.001 | 0.274 | 0.402 | 0.11 |
| Malonic acid | 1.22 | 1.21 | 1.21 | 1.18 | 1.23 | 1.23 | 1.15 | 1.32 | 0.571 | 0.833 | 0.687 | 0.02 |
| Methylamine | 0.43 | 0.60 | 0.92 | 0.58 | 0.62 | 0.32 | 0.41 | 0.26 | 0.089 | 0.210 | 0.514 | 0.07 |
| γ-aminobutyric acid | 1.02 | 0.90 | 0.40 c | 0.80 bc | 1.59 a | 1.18 ab | 1.45 abc | 0.63 bc | 0.418 | <0.001 | <0.001 | 0.13 |
| Amino acids | 12.41 | 12.97 | 13.26 | 12.47 | 12.78 | 12.66 | 12.23 | 12.71 | 0.763 | 0.172 | 0.805 | 0.18 |
| AGGX | 4.61 | 1.60 | 3.13 | 2.81 | 4.12 | 3.21 | 2.79 | 2.55 | <0.001 | 0.642 | 0.083 | 0.39 |
| Genus | Inoculation | Contaminant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NIS | INOC | BCE | BSU | BLI | CTY | CBJ | CON | P (I) | P (C) | P (I × C) | SEM | |
| At opening (100 d storage) | ||||||||||||
| Lactobacillus | 69.82 1 | 33.64 | 55.44 | 56.24 | 49.54 | 41.95 | 54.77 | 52.43 | <0.001 2 | 0.138 3 | 0.365 4 | 3.50 5 |
| Lentilactobacillus | 29.87 | 65.97 | 44.15 | 43.49 | 49.99 | 57.79 | 44.91 | 47.13 | <0.001 | 0.126 | 0.360 | 3.48 |
| Levilactobacillus | 0.17 | <0.01 | 0.07 | 0.11 | 0.12 | <0.01 | 0.10 | 0.14 | 0.004 | 0.765 | 0.765 | 0.03 |
| Loigolactobacillus | 0.04 | 0.10 | 0.07 | 0.06 | 0.11 | 0.08 | 0.04 | 0.06 | 0.010 | 0.361 | 0.031 | 0.01 |
| Acetobacter | 0.02 | 0.01 | <0.01 | 0.03 | 0.03 | <0.01 | 0.02 | <0.01 | 0.667 | 0.689 | 0.356 | 0.01 |
| Aerobic exposure (3 d) | ||||||||||||
| Lactobacillus | 50.76 | 27.97 | 37.33 | 38.37 | 41.60 | 35.31 | 42.50 | 41.08 | <0.001 | 0.954 | 0.827 | 2.92 |
| Lentilactobacillus | 39.31 | 69.54 | 52.29 | 57.10 | 53.40 | 57.36 | 54.36 | 52.04 | <0.001 | 0.975 | 0.296 | 3.40 |
| Levilactobacillus | 0.73 | 0.02 | 1.74 | 0.05 | 0.26 | 0.11 | 0.06 | 0.06 | 0.224 | 0.474 | 0.452 | 0.28 |
| Loigolactobacillus | 0.37 | 0.01 | 1.03 | <0.01 | 0.01 | 0.01 | <0.01 | <0.01 | 0.292 | 0.432 | 0.433 | 0.17 |
| Acetobacter | 8.27 | 2.22 | 6.45 | 4.20 | 4.34 | 7.03 | 2.89 | 6.60 | 0.012 | 0.862 | 0.200 | 1.21 |
| Genus | Inoculation | Contaminant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NIS | INOC | BCE | BSU | BLI | CTY | CBJ | CON | P (I) | P (C) | P (I × C) | SEM | |
| At opening (100 d storage) | ||||||||||||
| Kazachstania | 5.37 1 | 5.11 | 3.31 | 6.07 | 6.19 | 6.11 | 6.82 | 2.95 | 0.858 2 | 0.470 3 | 0.868 4 | 0.65 5 |
| Mucor | 33.50 | 37.24 | 25.82 | 35.81 | 50.29 | 31.47 | 40.60 | 28.21 | 0.539 | 0.223 | 0.194 | 3.21 |
| Nakaseomyces | 0.27 | 0.21 | 0.10 | 0.22 | 0.37 | 0.28 | 0.35 | 0.10 | 0.586 | 0.655 | 0.458 | 0.06 |
| Torulaspora | 0.10 | 0.11 | 0.09 | 0.13 | 0.04 | 0.18 | 0.09 | 0.16 | 0.807 | 0.739 | 0.696 | 0.03 |
| Wickerhamomyces | 13.62 | 7.43 | 14.19 | 16.11 | 7.78 | 7.83 | 3.00 | 14.24 | 0.346 | 0.831 | 0.477 | 3.07 |
| Saccharomyces | 0.56 | 0.24 | 0.27 | 1.27 | 0.06 | 0.50 | 0.23 | 0.09 | 0.435 | 0.534 | 0.434 | 0.20 |
| Aerobic exposure (3 d) | ||||||||||||
| Kazachstania | 2.04 | 5.04 | 0.38 | 5.39 | 3.99 | 4.62 | 5.61 | 3.66 | 0.010 | 0.298 | 0.087 | 0.81 |
| Mucor | 18.01 | 19.89 | 10.63 | 16.16 | 7.48 | 25.61 | 28.44 | 25.40 | 0.740 | 0.186 | 0.438 | 2.88 |
| Nakaseomyces | 0.42 | 0.28 | <0.01 | <0.01 | 1.64 | 0.12 | <0.01 | 0.34 | 0.768 | 0.276 | 0.732 | 0.22 |
| Torulaspora | 4.13 | 0.30 | <0.01 | 10.90 | 0.34 | 0.84 | <0.01 | 1.20 | 0.291 | 0.455 | 0.507 | 1.76 |
| Wickerhamomyces | 33.86 | 19.78 | 75.99 a | 18.12 b | 27.49 ab | 11.28 b | 1.61 b | 26.43 ab | 0.083 | <0.001 | 0.622 | 5.42 |
| Saccharomyces | 12.85 | 5.93 | 1.99 | 7.84 | 24.09 | 14.91 | 5.77 | 1.77 | 0.148 | 0.074 | 0.189 | 2.66 |
| Cell Count | Inoculation | Contaminant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NIS | INOC | BCE | BSU | BLI | CTY | CBJ | CON | P (I) | P (C) | P (I×C) | SEM | |
| At opening (100 d storage) | ||||||||||||
| Aerobic spore counts | 3.49 1 | 3.68 | 3.56 abc | 3.68 ab | 3.83 a | 3.51 bc | 3.67 ab | 3.26 c | 0.003 2 | < 0.001 3 | 0.002 4 | 0.04 5 |
| Anaerobic spore counts | 2.74 | 3.19 | 2.42 c | 3.35 a | 3.30 a | 2.84 b | 3.13 ab | 2.89 b | <0.001 | <0.001 | <0.001 | 0.10 |
| LAB | 8.39 | 8.76 | 8.59 | 8.52 | 8.63 | 8.58 | 8.54 | 8.59 | <0.001 | 0.887 | 0.306 | 0.04 |
| Yeasts | 2.34 | 1.65 | 2.46 | 1.56 | 1.97 | 1.72 | 2.19 | 2.10 | 0.001 | 0.152 | 0.073 | 0.12 |
| Aerobic exposure (3 d) | ||||||||||||
| Aerobic spore counts | 3.49 | 3.53 | 3.96 a | 3.67 b | 3.64 b | 3.28 c | 3.14 c | 3.36 c | 0.513 | <0.001 | <0.001 | 0.05 |
| Anaerobic spore counts | 3.40 | 3.24 | 3.37 ab | 3.29 ab | 3.47 a | 3.36 ab | 3.33 ab | 3.13 b | 0.002 | 0.007 | 0.035 | 0.03 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huffman, J.; Drouin, P.; Dunière, L.; LaPointe, G. Fermentation and Microbial Community of Maize Silage Inoculated with Lentilactobacillus buchneri NCIMB 40788 and Contaminated with Bacillus and Clostridium Spore Formers. Fermentation 2023, 9, 837. https://doi.org/10.3390/fermentation9090837
Huffman J, Drouin P, Dunière L, LaPointe G. Fermentation and Microbial Community of Maize Silage Inoculated with Lentilactobacillus buchneri NCIMB 40788 and Contaminated with Bacillus and Clostridium Spore Formers. Fermentation. 2023; 9(9):837. https://doi.org/10.3390/fermentation9090837
Chicago/Turabian StyleHuffman, Jesse, Pascal Drouin, Lysiane Dunière, and Gisèle LaPointe. 2023. "Fermentation and Microbial Community of Maize Silage Inoculated with Lentilactobacillus buchneri NCIMB 40788 and Contaminated with Bacillus and Clostridium Spore Formers" Fermentation 9, no. 9: 837. https://doi.org/10.3390/fermentation9090837
APA StyleHuffman, J., Drouin, P., Dunière, L., & LaPointe, G. (2023). Fermentation and Microbial Community of Maize Silage Inoculated with Lentilactobacillus buchneri NCIMB 40788 and Contaminated with Bacillus and Clostridium Spore Formers. Fermentation, 9(9), 837. https://doi.org/10.3390/fermentation9090837







