Abstract
The present study was based on bacterial isolation with probiotic potential from artisanal fermented pickles. A total of 36 bacterial strains were isolated from 50 different artisanal fermented pickle samples. Nine isolates with promising probiotic potential (PCR99, PCR100, PCR118, PCR119, PCR121, PCR125, PCR137, PCR140 and PCR141) were selected. The strains showed varied protease, amylase, lipase and cellulase patterns. The isolated strains displayed varied responses towards various antibiotic classes, i.e., PCR140 showed resistance to penicillin G, polymyxin B, Metronidazole and Streptomycin. PCR140 showed highest resistance to bile salt concentrations (0.3% and 0.5%) and acidic conditions (pH 3 and pH 4) when exposed to mimicked gastrointestinal conditions. The cell viability against enzymes produced in stomach and intestines showed different patterns as pepsin was in the range of 94.32–91.22%, pancreatic resistance 97.32–93.11% and lysozyme resistance was detected at 99.12–92.55%. Furthermore, the auto-aggregation capability of isolated strains was in the range of 46.11–33.33% and cell surface hydrophobicity was in the range of 36.55–31.33%. PCR 140 showed maximum antioxidant activity in lyophilized cells as well as probiotic potential. A phylogenetic analysis based on 16S rRNA gene sequencing confirmed that PCR140 (NMCC91) with higher in vitro probiotic and antioxidant potential belongs to the genus Lactobacillus with 97% similarity with Lacticaseibacillus paracasei. This work demonstrated that the isolate PCR 140 (NMCC91) is suitable for use in food and medical industries.
1. Introduction
Food fermentation has traditionally been associated with a myriad of cultures since antiquity. The outcomes of this centuries-old practice are better food storage, improved quality, and an enhanced consistency of final products through pickling or fermentation processes. Since the beginnings of human civilization, fermentation has been one of most important components in our diet in the form of wine, beer, yogurt, pickles, cheese, etc. Pickling can be defined as “a procedure of converting sugar into acids by lactic acid bacteria” [1]. These acids inhibit pathogens and spoilage-causing non-acid tolerant microorganisms [1]. Pickling is a common practice for food preservation based on the fermentation of lactic acid, alcohol and acetic acid [2]. Globally, pickles are consumed as appetizers that improve the digestion of grains and vegetables. Furthermore, pickles are stored at room temperature [3,4].
Lactic Acid bacteria involved in the pickling process can be defined as “Gram positive, organotrophic, non-sporulating, acid and air tolerant, catalase-negative, non-motile cocci or rod and carbohydrate fermentative and lactic acid forming bacteria” [5,6]. These pickle-associated bacteria produce unique flavors and also promote overall health and fitness [7]. Pickles improve human health by providing vitamins, carbohydrates, minerals, and certain pigments such as anthocyanin, glucosinolates, lycopene, 𝛽-carotene and flavonoids [8,9]. Pickling serves to preserve the nutritional value of fruits and vegetables, prevents food spoilage, and increases food safety. Thus, fermentation acts as an alternative for food additives [1,10,11].
Lactic Acid Bacteria [LAB] can be used as a starter culture for the processing and preservation of meats, fruits, vegetables and dairy items [12]. The final products must meet the standards of consumer acceptability and consistent quality [13]. Lactiplantibacillus plantarum and Limosilactobacillus fermentum are well known microorganisms that are suitable for use as starter cultures [12,13]. The safety and production of antimicrobial compounds makes LAB a befitting choice for the bio-preservation of food commodities. The antimicrobials produced by LAB are di-acetyl, ethanol, organic acid, hydrogen peroxide and bactericidal peptides [14,15,16].
Probiotics are known to combat the putrefactive impacts of the gut metabolism which could cause aging and illness. Probiotics were discovered by Elie Metchnikoff, who is known as the “father of probiotics”. Fermented foods prepared using probiotics are functional foods and have beneficial impacts on the human gut system [17].
Food and agricultural industries are continuously improving through innovational strategies, leading to the generation of constantly emerging research technologies. The change in the acceptance, needs and preferences of consumers is a dynamic process, but food quality maintenance through technology innovations is obvious. The habits of consumers, sustainability factors and their cultural heritage affect technological innovations which are applied in our food industry. Recently, consumers became increasingly health conscious and now prefer products with higher beneficial values. Manufacturers are incentivized to produce functional foods due to these demands. Hence, the acceptance of novel products and successful marketing requires added food value functionalities. Novel food products can be processed or naturally enriched with active compounds secreted by biological agents. These compounds provide health benefits apart from those provided by nutrients when administered in a required quantity.
In the context of previous studies, LABs are recognized as the most prominent probiotic strains as few of the species inhabit the small intestine by resisting harsh environmental conditions, i.e., a low pH, various bile salts and interact with other pathogenic strains and natural inhibitors through their antagonistic activities and antibiotic susceptibility [18]. Besides, their probiotic capabilities such as production of enzymes and hydrophobicity, auto-aggregation and antioxidative activity against DPPH are well demonstrated by L. plantarum, P. pentosaceus, P. acidilactici, E. lactis and E. hira which were isolated from fermented fish Shidal [18], whereas Lactiplantibacillus plantarum F1 and Levilactobacillus brevis OG1 showed antagonistic activity against pathogens [19], and E. faecalis and E. faecium demonstrated high responses towards cell-surface auto-aggregation characteristics [18].
This is the first study based on microflora of traditional homemade pickles in this region of Pakistan. Therefore, this research was performed with the aim to isolate and identify the potential probiotic LAB from traditional homemade pickles. The strains were evaluated based on FAO/WHO parameters on the classification of probiotics. These parameters include the ability to survive simulated GIT conditions, bile salt tolerance, assessment of antibiotic susceptibility, antagonistic activity, screening of bile salt hydrolase (BSH) and lipase activity, antioxidative activity against DPPH free radicals and cell surface characteristics, so that they can be employed for the formation of other beneficial products for living beings.
2. Materials and Methods
2.1. Isolation and Characterization of Lactic Acid Bacteria
Fifty samples of homemade pickles made in brine were collected randomly from Islamabad, Pakistan. Carrot pickles [Daucus carota subsp. Sativus], garlic pickles (Allium sativumi), radish pickles (Raphanus raphanistrum subsp. Sativus) and green chili pickles (Capsicum annuum) were collected in sterile sample bottles. All the samples were transferred to the Probiotic Laboratory in the National Institute for Genomics and Advanced Biotechnology (NIGAB) NARC and were stored at 4 °C. Furthermore, samples were serially diluted and 100 μL of a 100-10-5 serial dilution was poured on MRS agar plates and incubated at 37 °C for 48 h. Colonies were randomly selected and purified on MRS agar by repeated streaking about 4–5 times [4,20,21]. Purified cultures were stored in 30% glycerol at −80 °C. The preliminary identification of isolates included the study of cell morphology, Gram staining, as well as oxidase and catalase testing [22].
2.2. Molecular Identification of Bacterial Isolates
The molecular identification of selected purified strains was performed by adopting the procedure described in [23]. The bacterial DNA was isolated through a DNA extraction process in which colonies were suspended in 1× TE buffer and placed in a PCR machine for 2–3 min at 6000 rpm after heating at 95 °C for 10 min. The 16S rRNA gene of 9 isolated strains was amplified by PCR [Polymerase Chain Reaction] using the reaction mixture containing PCR water, Taq buffer, DNTPs, MgCl2, Universal primers, =, and Taq polymerase. After the formation of the reaction mixture, a second set of tubes was placed in PCR for 2 h and 10 min with the following PCR conditions: initial denaturation at 94 °C for 2 min then 30 cycles of denaturation at 94 °C for 1 min. Initial denaturation was followed by an annealing process for 1 min at 50 °C. After the annealing process, extension was performed at 72 °C for 1.5 min. Lastly, the final extension of the PCR product was performed at 72 °C for almost 5 min. The PCR product was electrophorized by using 2% agarose gel and visualized with ethidium bromide staining. Selected strains were sent for 16S rRNA sequencing at the commercial sequencing facility of Macrogen Inc. (Seoul, South Korea) Sequences were identified by using the BLAST system (https://blast.ncbi.nlm.nih.gov/Blast.cgi, accessed on 10 June 2019) and were submitted to NCBI on 26 June 2019 (http://www.blast.ncbi.nlm.nih.gov). Phylogenetic trees were developed by aligning the selected sequences into MEGA version X with the Neighbor-Joining method.
2.3. Probiotic Characterization
2.3.1. Acid Tolerance
Bacterial cultures were grown in MRS broth with different pH levels of 2, 3, 4 and 5 at 37 °C for 24 h in a shaking incubator followed by a measurement of Optical density at 600 nm. The experimentation was performed in triplicate for validity [24].
2.3.2. Bile Tolerance
Bile tolerance was evaluated with a previously used method as described by Kılıç and Karahan (2010) [25]. Fresh cultures of lactic acid bacterial strains were incubated in MRS broth of 0.3% and 0.5% bile acids for 24 h followed by the monitoring of bacterial growth at 600 nm with a spectrophotometer. All the experiments were performed in triplicate.
2.3.3. Cell Auto-Aggregation
Cell auto-aggregation of bacterial isolates was tested using the procedure outlined by Yadav et al. (2016) [26]. Pure bacterial cultures grown in MRS broth for 18 h were centrifuged at 6000 rpm for 5 min followed by washing and re-suspension in PBS. Absorbance was measured at 600 nm. This suspension was then incubated for 2 h at 37 °C. After 2 h, 1 mL of upper surface supernatant was removed from this suspension and absorbance was measured at 600 nm. Cell auto-aggregation was determined by the decline in absorbance capability and measured with the given formula. All the experiments were performed in triplicate.
Auto-aggregation = (Initial OD − final OD/Initial OD) × 100
2.4. Cell Surface Hydrophobicity
Cell surface hydrophobicity was carried out following the previously described protocol by Gharbi et al. (2019) [27] with slight modifications. Bacterial cultures were incubated in MRS broth for 24 h at 37 °C followed by centrifugation at 6000 rpm for 5 min. Pellets were washed with normal saline, Phosphate Saline Buffer (PSB), suspended in autoclaved distilled water and incubated for 20–30 min after adding 600 μL of Xylene. The aqueous phase was separated carefully, and its OD was recorded at 600 nm. Calculations of percentage of hydrophobicity were conducted according to the following formula:
where Ao = OD before Xylene addition and A1 = OD of aqueous layer.
Hydrophobicity Percentage (%) = [(Ao − A1)/Ao] × 100
2.5. Determination of Pepsin and Pancreatin Resistance
Lactobacillus strains were tested for their resistance to pepsin and pancreatin by following the method given by Jamaly (2011) [28]. Bacterial cultures were centrifuged at 10,000× g for 5 min followed by washing twice with Phosphate-buffered saline of neutral pH. Afterwards these cultures were resuspended in PBS solution of pH 2.0 containing 3 mg/mL pepsin and in PBS solution of pH 8 containing 1 mg/mL pancreatin for 24 h. Resistance to pepsin and pancreatin was determined by calculating viable colony counts.
2.6. Lysozyme Resistance
The lysozyme resistance of selected bacterial isolates was observed using the procedure suggested by Yadav et al. (2016) [26]. Pure Lactobacilli strains were centrifuged for 10 min at 7000 rpm. The isolates were washed with PBS twice and suspended in Ringer’s solution. A total of 10 µL from this suspension was incubated at 37 °C in a sterilized electrolyte solution containing 100 mg/L lysozyme, NaHCO3 1.2 g/L, NaCl 6.2 g/L, CaCl2 0.22 g/L and KCl 2.2 g/L. A control sample was also prepared by the inoculation of strains in an electrolyte solution without lysozyme. After 2 h, the plate count method was used to determine the cell count viability.
2.7. Amylolytic Activity
Amylolytic activity was assessed using a previously recommended procedure described by Ghazanfar (2016) [29]. Media was prepared by adding nutrients agar and starch in distilled water. Media was inoculated with these strains and incubated at 37 °C for 24 h. After that, iodine was sprinkled over it and the presence or absence of luminous zones was observed.
2.8. Cellulolytic Activity
Cellulolytic activity was checked using the method described by Ghazanfar (2016) with slight modifications applied [29]. CMC media was prepared followed by inoculation and incubation for 37 °C for 24 h. The plates were first stained with Congo red dye for 15 min followed by staining with NaCl for 15 min. The presence or absence of a luminous zone was recorded.
2.9. Proteolytic Activity
Proteolytic activity of LAB strains was measured by streaking these strains on skim milk agar plates using previously described method as per Monika et al. (2017) [4]. Luminous zones were observed.
2.10. Lipolytic Activity
For the determination of the lipolytic activity of concerned strains, TSA was prepared with 1 mL Tween 80 in 100 mL distilled water and phenol red. Colonies were spread on the solid surface of agar and incubated for 37 °C for 24 h. Change in color was recorded.
2.11. Assessment of Antibiotic Susceptibility
Antibiotic susceptibility was assessed using the disc diffusion method described by Monika et al. (2017) [4]. Bacterial cultures which were 24 h old and grown at 37 °C were tested against 15 antibiotics including Penicillin G (10 iu), Polymyxin B (300 iu), Chloramphenicol (30 mcg), Ampicillin (10 mcg), Bacitracin (10 iu), Kanamycin (30 mcg), Cephalexin (30 mcg), Tetracycline (30 mcg), Amoxycillin (30 mcg), Metronidazole (5 mcg), Vancomycin (30 mcg), Streptomycin (10 mcg), Gentamycin (10 mcg) and Nalidixic Acid (30 mcg). Bacterial suspensions were swabbed on Mueller Hinton Agar (MHA) agar plates followed by the placement of antibiotics discs on the MHA agar plates, which were then incubated for 24 h at 37 °C. Results were recorded afterwards. The measurement of inhibition zones was evaluated on the basis of VETLAB standards. All the experiments were performed in triplicate.
2.12. Antagonistic Activity
The antimicrobial activity test of concerned strains was carried out using the agar-well diffusion method [4]. The pathogenic strains were Escherichia coli (ATCC8739), Listeria monocytogenes (ATCC13932), Staphylococcus aureus (ATCC6538), and Bacillus cereus (ATCC11778). Swabbed plates with a pathogen suspension were used to make wells filled by 50 μL suspension Lactobacillus strains and zones were measured after incubation at 37 °C for 24 h. The experiments were performed in triplicate.
2.13. Antioxidant Activity
The antioxidant activity of bacterial isolates was monitored against (2, 2-diphenylpicrylhydrazyl) DPPH free radicals using a protocol described by Nadri et al. (2014) [30]. To perform this assay, 0.03 g of DPPH was added to 100 mL of methanol. An 18 h fresh bacterial culture was prepared and added (200 mL) to 800 mL of DPPH and kept for 30 min at 37 °C. Ascorbic acid was used as the standard. Scavenging or the inhibition of free radicals was measured at 517 nm with a UV–visible spectrophotometer. The value was calculated using the following formula:
Inhibition (%) = [(Absorbance of control Absorbance of tests)/Absorbance of control] × 100
2.14. Statistical Analysis
The data were expressed as mean ± standard error mean (SEM) calculated over individual experiments which were performed in triplicate. For the inference statistics, a one-way ANOVA was used to determine the significant differences among the isolated strains on quantitative parameters (p < 0.05).
3. Results
3.1. Purification and Biochemical Characterization of Bacterial Isolates
In this experimental study, nine bacterial isolates were isolated as LAB strains based on their morphological, biochemical, and molecular characteristics. All the isolated strains were Gram positive rods. All bacterial isolates were catalase and oxidase negative.
3.2. Enzymatic Potential of Lactobacillus Strains
Enzymatic activity was evaluated to observe the capability of Lactobacillus strains for producing industrially important enzymes. Bacterial strains did not show any amylolytic activity except strain PCR125. Clear zones were formed only in the “125” strain’s plate which indicated that this bacterial strain has potential to hydrolyze the starch. Five strains, i.e., PCR99, PCR100, PCR118, PCR119 and PCR141 including L. plantarum and L. paracasei were able to show positive results and to utilize Tween 80 as a lipid source. Two strains PCR137 and PCR119) showed proteolytic activity (Table 1).

Table 1.
Determination of Enzymatic Potential of Selected Lactobacillus Strains.
3.3. Antibiotic Suscenptibility
The antibiotic susceptibility results of nine bacterial strains against 15 different antimicrobial agents are presented in Table 2. Results demonstrated that almost all isolated strains are sensitive to protein synthesis inhibitors, i.e., Chloramohenicol, Gentamycin and Streptomycin. Strains showed varied susceptibility responses towards cell wall and DNA synthesis inhibitors, i.e., Penicilin G, Plymyxin B, Ampicillin, Bacitracin, Vancomycin, etc. Furthermore, strains were observed to be resistant to Niroimidazoles, i.e., Metronidazoles.

Table 2.
Antibiotic Susceptibility Determination of Isolated Lactobacillus strains according to VETLAB standards.
3.4. Determination of Cell Viability under Different Probiotic Parameters
3.4.1. pH Tolerance Assay
The isolated bacteria demonstrated diverse results when exposed to different acidic conditions, i.e., pH, as mentioned in Figure 1. The graph depicts that isolated strains are able to survive all pH conditions, specifically pH 5 and pH 7 while at pH 2, PCR118 displayed lowest and PCR140 showed highest tolerance, and PCR100 showed highest tolerance and PCR137 showed the lowest tolerance at pH 3. At pH 4, a tolerance range was observed between PCR121 and PCR100. On the whole, significant results were observed.

Figure 1.
Detection of pH tolerance in bacterial strains. Values are means of replicate experiments; ± indicates standard deviation from the mean. Values differs significantly (p < 0.05).
3.4.2. Bile Salt Tolerance Assay
While observing bacterial tolerance at different bile salt concentrations, bacterial isolates demonstrated that at a 0.3% concentration, strains were highly tolerant as compared to 0.5% concentration (Figure 2). It was observed that PCR99, PCR100, PCR140 showed the highest growth at a concentration of 0.3% and PCR121, PCR118 and PCR119 showed highest growth among other strains at 0.5%.
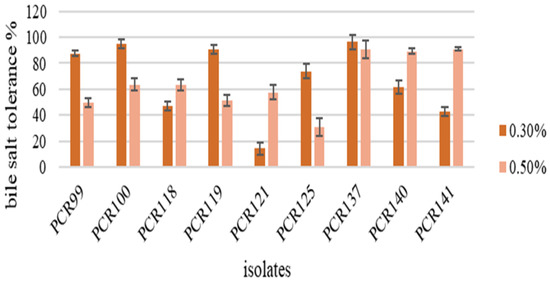
Figure 2.
Detection of Bile Salt tolerance in bacterial strains. Values are means of replicate experiments; ±indicates standard deviation from the mean. Values differs significantly (p < 0.05).
3.4.3. Auto-Aggregation and Cell Surface Hydrophobicity Determination
Among the tested strains, all bacterial isolates showed good auto-aggregation in the range of 46.11–33.33%. Among them, PCR141 was found to be the best auto-aggregating strain and showed 46.11% aggregation over the 24 h incubation period which means this strain is capable of aggregating on epithelial layers of intestine (Figure 3). In the present study, six strains showed highest hydrophobicity, i.e., PCR99, PCR100, PCR118, PCR121 and PCR140, in the range of 36.65–31.33% while the capacity of other strains was also satisfactory, thereby depicting that the strains are able to survive and adhere themselves with the epithelial intestinal layers (Figure 4).
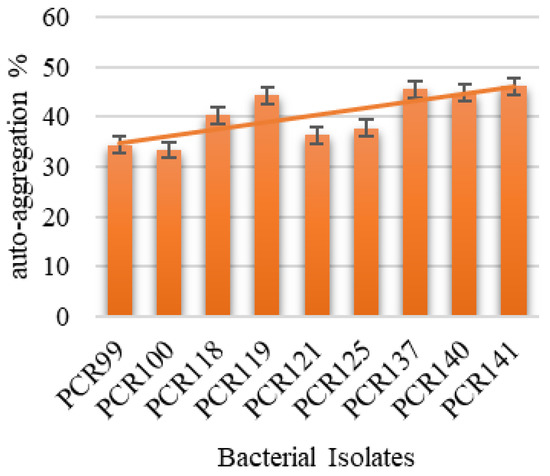
Figure 3.
Determination of Auto-aggregation % of isolated bacterial strains.
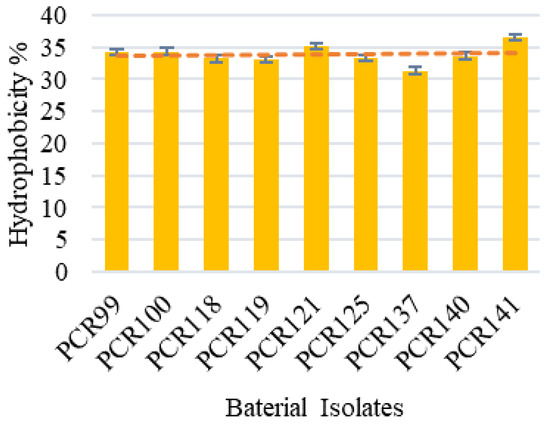
Figure 4.
Determination of Hydrophobicity % of bacterial isolates. Values are means of replicate experiments; ±indicates standard deviation from the mean. Values differs significantly (p < 0.05).
3.4.4. Effect of Lysozyme on Cell Viability
Among the tested bacterial isolates against the bactericidal activity of lysozyme, all strains showed resistance in the range of 99.12–92.55%. PCR141 was observed to be highly resistant to lysozyme, showing viability of up to 99.12%, while the trend showed that the property is strain-specific as all isolates displayed different patterns (Figure 5).
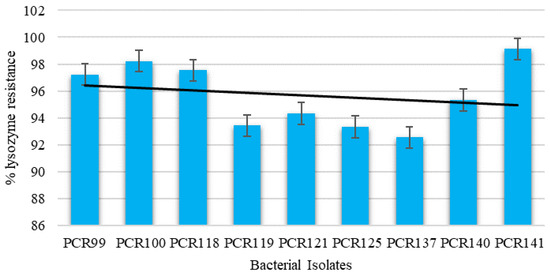
Figure 5.
Determination of cell viability of bacterial isolates against Lysozyme resistance. Values are means of replicate experiments; ± indicates standard deviation from the mean. Values differ significantly (p < 0.05).
3.4.5. Pepsin and Pancreatin Resistance Assay
The pepsin and pancreatin resistance test revealed that the selected bacterial isolates are capable of surviving under intestinal and gastric enzymes as their viability rates were in the range of 94.34–90.34% for pepsin resistance (Figure 6a) and 97.32–93.11% for pancreatin resistance (Figure 6b). The results show that PCR140 and PCR137 demonstrated the highest pancreatin resistance while PCR125 showed lowest resistance. Furthermore, PCR118 and PCR140 possessed the highest pepsin resistance in our study and PCR119 showed lowest resistance.
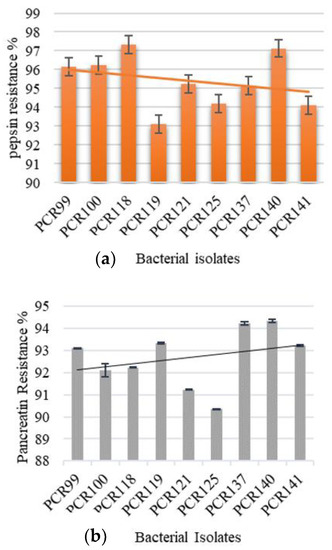
Figure 6.
Determination of bacterial viability in the presence of; (a) pepsin conditions and (b) pancreatin conditions.
3.4.6. Antioxidant Activity against DPPH
Table 3 displays the antioxidant capability of PCR140. The results show that PCR140 displayed the highest DPPH free radical scavenging property. The probiotic strains were observed with almost the same inhibition ability as displayed with ascorbic acid.

Table 3.
In vitro antioxidant of the PCR140.
3.5. Phylogenetic Analysis
The bacterial isolates were identified on a molecular level by using the BLAST tool provided on the National Center for Biotechnology Information [NCBI] website. Four of the sequences were submitted to the NCBI database and the taxonomy of these bacterial isolates is illustrated in Table 4.

Table 4.
Provisionally Taxonomy of Bacterial Isolates.
Provisionally molecular identification of strain PCR140 was followed by a phylogenetic analysis of Lacticaseibacillus paracasei to study their relationship to the nearest species, as mentioned in Figure 7.
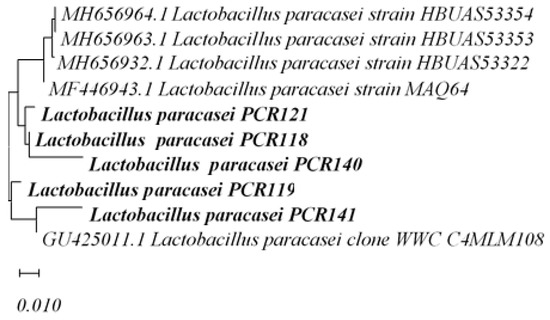

Figure 7.
Phylogenetic Analysis of Lacticaseibacillus paracasei PCR121, PCR118, PCR140, PCR141, PCR119.
4. Discussion
In the present study, nine bacterial isolates were isolated and characterized from fifty different pickled samples, which displayed potent probiotic potential. Despite the fact that the isolates of this genus are already being consumed in the market for the production of many nutraceutical products, no reports have been published reported about their isolation from pickles made in brine solution. Furthermore, the present research is novel due to its sampling area, i.e., Islamabad, where no reports have as of yet been published. The samples selected for the isolation of LAB strains form an indigenous part of the Pakistani diet and are consumed unknowingly as probiotic drinks/foods by people. The presence of LAB strains with potential probiotic characteristics has been reported in many studies. In a study by Kumar et al. (2011), many Lactic Acid Bacteria were isolated from Persian traditional vegetable pickles in which Lactobacillus was the predominant species along with L. Paracasei, L. casei, L. Pentosus, L. brevis and L. mesenteroides [31]. Monika et al. (2017) isolated Lactiplantibacillus plantarum from pickles made in Himachal Pradesh, India [4].
In terms of the enzymatic activity of functional foods in industries, Lactiplantibacillus plantarum [PCR125] strains showed amylase activity while the other eight strains did not exhibit this activity, as they are not involved in α-amylase production, which is in agreement with the results of our research [32]. The literature shows that some strains of Lactiplantibacillus plantarum are capable of α-amylase production [33,34]. In contrast to the literature, Lacticaseibacillus paracasei did not show amylolytic activity [35]. The amylase enzyme has its applications in food, textile, paper, pharmaceutical and detergent manufacturing industries as it can be used for corn syrup, glucose syrups, alcohol fermentation, maltose syrups and in detergents [36,37]. In the present study, the absence of cellulolytic activity revealed that these strains are incapable of metabolizing carboxymethyl cellulose. Therefore, it was concluded that cellulase activity is strain dependent. Cellulolytic activity was observed in the present study, contrary to the study by Singhvi et al. (2010), which declared mutant LAB strains (Lactobacillus lactis mutant RM2-24) were used for cellulose production [38]. In accordance with a study conducted by Dinçer et al. (2018), L. plantarum has the highest lipolysis activity as this activity has been observed in strains isolated from Pastirma in Turkey [39]. The development and production of cheese flavors such as cheddar cheese can be achieved with lipase enzymes [40], which reveals that they are able to make peptide populations of a medium size as a result of proteolytic processes [41]. Protease enzymes can be used for bread quality improvement, brewing, meat tenderization and for the coagulation of milk [40,42].
In this study, antibacterial tests were performed in which bacterial strains were tested against pathogenic strains of E. coli (ATCC8739), Listeria monocytogenes (ATCC13932), Staphylococcus aureus (ATCC6538), and Bacillus cereus. No bacterial strain was detected to be resistant against pathogenic strains. The results revealed that most strains are resistant to Penicillin G and metronidazole, which is in accordance with previous research [43,44,45]. Multiple drug resistance was observed in strains PCR100 and PCR141 while most strains were observed to be sensitive to these antimicrobial agents. The resistance observed against some antibiotics tested suggests that our strains would not be affected by therapies using these antibiotics and might help maintain the natural balance of intestinal microflora during antibiotic treatments.
Microorganisms must be able to resist uncongenial Gastrointestinal Tract (GIT) conditions to be classified as probiotics and to exert beneficial effects. The stomach pH can be reduced to 1.0 in the presence of pepsin but in vitro experimentations were performed with a pH of 1.5–4.5 due to the buffering mechanisms of the food matrix, which produces shielding effects on gut microbiota [16,23]. Resistance to low pH is essential not only for use as a probiotic for humans, but also to produce various food products because low pH resistance helps them to survive in acidic conditions such as in yogurt, etc. Moreover, researchers also discussed that low acidic conditions affect the survival of lactic acid bacterial strains, especially at pH 2 [23,46,47]. The ability to resist bile salts is crucial for colonization and metabolic activities of bacterial isolates in intestines. In intestines, bile salt, at a concentration of 0.3%, is secreted; therefore, bacterial tolerance was observed against bile salt concentrations of 0.3% and 0.5%. In the present study, the results demonstrated that isolated strains are significantly tolerant to pH 1–pH 4 and 0.3% bile salt concentrations. BSH activity is a relevant property for probiotic strains to survive the toxicity of conjugated bile salts in the duodenum [25]. Pancreatin enzymes help in the digestion of fats, proteins and carbohydrate and are released in intestines through the pancreatin duct, while pepsin is secreted by gastric chief cells to digest proteins in an inactive form, i.e., zymogens, which become active when HCl lowers the stomach pH. The tolerance of bacterial strains against pancreatin and pepsin is considered as another criterion to predict their survival in harsh GIT conditions [48]. The nine isolated strains displayed 90–95% resistance against pancreatin.
The auto-aggregation property of bacterial isolates allows them to adhere to the epithelial lining of intestines, which is considered as a beneficial property because it prevents the flush out of bacterial strains from the body through peristalsis. The auto-aggregation percentages of isolated strains increased with time which is consistent with previous findings [49,50]. In a study conducted by Abid et al., it was revealed that Lactobacilli strains, i.e., NMCC-14, exhibit the highest auto-aggregation properties, i.e., 47.55 ± 0.08.
A hydrophobicity test was conducted to detect bacterial attachment capability to intestinal epithelial cells in the presence of hydrocarbons. Bacterial potential colonization is considered an important property for the selection of strains in probiotics. Researchers have discussed hydrophobicity values of some Lactobacillus strains in the range of 82.41–97.96% but values of 15–60% were also observed [51,52]. The interactive forces involved in the adhesion process of Lactobacilli strains include electrostatic interaction, passive forces, and steric and hydrophobic forces. The adherence property of probiotic strains inhibits or prevents the attachment of pathogenic strains to the epithelial layer of the gut which, in turn, prevents pathogenic activity in the GIT tract of organisms [53,54]. Our results are in accordance with the literature [53,55] which demonstrated that Lactobacilli strains isolated in Islamabad are capable of growing under lysozyme conditions in the range of 96.69–53.45%. The results of gastric conditions are in accordance with the literature; however, according to Tokatlı et al., pancreatin has a negative impact on strain-survival rates which is in contrast to our results [54]. Furthermore, these results are in accordance with the data published by Adnan et al. (2017) [56].
In this study, the antioxidant activity of strain PCR140 was observed using the DPPH radical method. The intracellular LAB extracts are reported to have a chelating ability due to metal ions. Besides, intact LAB cells are known to have antioxidant capabilities; therefore, they are present in the GIT tract. [55,57,58]. While passing through GIT tract, LAB released an antioxidant constituent which was reported as a healthy mechanism. Our experimental results showed higher antioxidant capability which is in accordance with the previous literature [59,60,61]. Therefore, strain PCR 140 can be classified as probiotic. The phylogenetic analysis of isolated strains demonstrated the evolutionary relationship and showed that isolates possess > 95% similarity with Lacticaseibacillus paracasei strains.
5. Conclusions
The need to isolate Lactobacillus strains with potent probiotic potential was satisfied with the isolation and characterization of nine Lactobacillus strains from fifty traditional pickles from Islamabad, Pakistan. This was the first study on Lactobacillus isolates conducted in Islamabad on fermented products prepared in brine solution. All the nine isolates were examined on the basis of FAO/WHO guidelines and possessed the minimum criteria to be classified as probiotics. Lactobacillus strains demonstrated satisfactory resistance against gastrointestinal conditions such as an acidic environment, bile salt, bile salt hydrolase, pepsin and pancreatin conditions. Isolated bacterial strains displayed remarkable cell surface characteristics, i.e., hydrophobicity and auto-aggregation properties. The antibiotic susceptibility assay concluded that isolated Lactobacilli strains can be classified as safe. Moreover, Lactobacilli strains with an enzymatic potential of varied significant performances were observed. Lacticaseibacillus paracasei (PCR140) displayed higher antioxidant potential as well. The molecular identification of isolated strains showed that three strains were of Lactiplantibacillus plantarum, five of Lacticaseibacillus paracasei and one was identified as Levilactobacillus brevis. The phylogenetic analysis of these bacterial isolates was performed to analyze the evolutionary relationship. After all these experiments, their significance results demonstrated that these isolates are suitable for use in food and medical industries for the benefits of humans and animals. Furthermore, assays such as the hemolysis assay and gelatinase assay, and the co-aggregation of bacterial isolates with pathogenic strains need to be studied for a further safety evaluation of isolated strains. Moreover, in vivo studies evaluating their safety need to be conducted.
Author Contributions
Conceptualization, U.A. and I.G.; methodology, S.G.; software, A.F. and M.M.; validation, S.G., I.G. and U.A.; formal analysis, A.M.E.; investigation, S.G. and M.A.E.N.; resources, S.M.B.A.; data curation, S.G.; writing—original draft preparation, S.G. and A.S.A.; writing—review and editing, S.G., A.S.A. and S.M.B.A.; visualization, M.A.E.N.; supervision, I.G.; project administration, S.G.; funding acquisition, B.A., A.S.A. and S.M.B.A. All authors have read and agreed to the published version of the manuscript.
Funding
Abdulhakeem S. Alamri would like to acknowledge Taif University for support No. TURSP (2020/288). Syed Mohammed Basheeruddin Asdaq wishes to express his gratitude to AlMaarefa University in Riyadh, Saudi Arabia, for providing support (TUMA-2021-1) to do this study. The authors also acknowledge PARC laboratory for providing financial support for this research project.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All data generated and analyzed during this study are included in this published article.
Acknowledgments
The authors extend their appreciation to the Deputyship for Research and Innovation, Ministry of Education in Saudi Arabia for funding this work through grant no. 375213500. Abdulhakeem S. Alamri would like to acknowledge Taif university for support No. TURSP (2020/288).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Sayın, F.K. The effect of pickling on total phenolic contents and antioxidant activity of 10 vegetables. J. Food Sci. 2015, 1, 135–141. [Google Scholar] [CrossRef]
- Lee, E.; Jung, S.-R.; Lee, S.-Y.; Lee, N.-K.; Paik, H.-D.; Lim, S.-I. Lactiplantibacillus plantarum strain LN4 attenuates diet-induced obesity, insulin resistance, and changes in hepatic mrna levels associated with glucose and lipid metabolism. Nutrients 2018, 10, 643. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Khaskheli, G.B.; Zuo, F.L.; Yu, R.; Chen, S.W. Overexpression of small heat shock protein enhances heat- and salt-stress tolerance of Bifidobacterium Longum NCC2705. Curr. Microbiol. 2015, 71, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Monika, S.; Kumar, V.; Kumari, A.; Angmo, K.; Bhalla, T.C. Isolation and characterization of lactic acid bacteria from traditional pickles of Himachal Pradesh, India. J. Food Sci. Technol. 2017, 54, 1945–1952. [Google Scholar] [CrossRef]
- Perczak, A.; Goliński, P.; Bryła, M.; Waśkiewicz, A. The efficiency of lactic acid bacteria against pathogenic fungi and mycotoxins. Arch. Ind. Hyg. Toxicol. 2018, 69, 32–45. [Google Scholar] [CrossRef] [Green Version]
- Alhashem, Y.N.; Farid, A.; Al Mohaini, M.; Muzammal, M.; Khan, M.H.; Dadrasnia, A.; Alsalman, A.J.; Al Hawaj, M.A.; Ghazanfar, S.; Almusalami, E.M.; et al. Protein Isolation and Separation Techniques of Pasteurella multocidavia One-and Two-Dimensional Gel Electrophoresis. Int. J. Cur. Res. Rev. 2022, 14, 1–8. [Google Scholar] [CrossRef]
- Choi, I.H.; Noh, J.S.; Han, J.-S.; Kim, H.J.; Han, E.-S.; Song, Y.O. Kimchi, a fermented vegetable, improves serum lipid profiles in healthy young adults: Randomized clinical trial. J. Med. Food 2013, 16, 223–229. [Google Scholar] [CrossRef] [Green Version]
- Isabelle, M.; Lee, B.L.; Lim, M.T.; Koh, W.-P.; Huang, D.; Ong, C.N. Antioxidant activity and profiles of common vegetables in Singapore. Food Chem. 2010, 120, 993–1003. [Google Scholar] [CrossRef]
- Takebayashi, J.; Oki, T.; Watanabe, J.; Yamasaki, K.; Chen, J.; Sato-Furukawa, M.; Tsubota-Utsugi, M.; Taku, K.; Goto, K.; Matsumoto, T.; et al. Hydrophilic antioxidant capacities of vegetables and fruits commonly consumed in Japan and estimated average daily intake of hydrophilic antioxidants from these foods. J. Food Compos. Anal. 2013, 29, 25–31. [Google Scholar] [CrossRef]
- Klein, M.; Sanders, M.E.; Duong, T.; Young, H.A. Probiotics: From bench to market. Ann. N. Y. Acad. Sci. 2010, 1212, E1–E14. [Google Scholar] [CrossRef]
- Zuo, F.; Yu, R.; Feng, X.; Chen, L.; Zeng, Z.; Khaskheli, G.B.; Ma, H.; Chen, S. Characterization and in vitro properties of potential probiotic bifidobacterium strains isolated from breast-fed infant feces. Ann. Microbiol. 2015, 66, 1027–1037. [Google Scholar] [CrossRef]
- Olaoye, O.A.; Ndife, J.; Raymond, V.I. Use of Lactiplantibacillus plantarum as starter culture and its influence on physicochemical, microbiological, and sensory characteristics of kunnu-aya produced from Sorghum and tigernut. J. Food Qual. 2017, 2017, 6738137. [Google Scholar] [CrossRef] [Green Version]
- Akabanda, F.; Owusu-Kwarteng, J.; Tano-Debrah, K.; Parkouda, C.; Jespersen, L. The use of lactic acid bacteria starter culture in the production of Nunu, a spontaneously fermented milk product in Ghana. Int. J. Food Sci. 2014, 2014, 721067. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yaseen, M.; Kamran, M.; Farid, A.; Ismail, S.; Muzammal, M.; Amir, K.A.; Rashid, S.A. Antibacterial, Hemagglutination, and Insecticidal Activity Studies on the Solvent Extracts of the Roots of Olea ferruginea. Makara J. Sci. 2022, 26, 8. [Google Scholar]
- Salomskiene, J.; Jonkuviene, D.; Macioniene, I.; Abraitiene, A.; Zeime, J.; Repeckiene, J.; Vaiciulyte-Funk, L. Differences in the occurence and efficiency of antimicrobial compounds produced by lactic acid bacteria. Eur. Food Res. Technol. 2019, 245, 569–579. [Google Scholar] [CrossRef]
- Kanwal, H.; Di Cerbo, A.; Zulfiqar, F.; Sabia, C.; Nawaz, A.; Siddiqui, F.M.; Aqeel, M.; Ghazanfar, S. Probiotic characterization and population diversity analysis of gut-associated Pediococcus acidilactici for its potential use in the dairy industry. Appl. Sci. 2021, 11, 9586. [Google Scholar] [CrossRef]
- Daliri, E.B.-M.; Lee, B.H. New Perspectives on probiotics in health and disease. Food Sci. Hum. Wellness 2015, 4, 56–65. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.; Mohanty, U.; Majumdar, R.K. Isolation and characterization of lactic acid bacteria from traditional fermented fish product Shidal of India with reference to their probiotic potential. LWT 2021, 146, 111641. [Google Scholar] [CrossRef]
- Ogunbanwo, S.T.; Sanni, A.I.; Onilude, A.A. Characterization of bacteriocin produced by Lactiplantibacillus plantarum F1 and Lactobacillus brevis OG1. Afr. J. Biotechnol. 2003, 2, 219–227. [Google Scholar]
- Mohammed, S.; Çon, A.H. Isolation and characterization of potential probiotic lactic acid bacteria from traditional cheese. LWT 2021, 152, 112319. [Google Scholar] [CrossRef]
- Prawan, K.; Bhima, B. Isolation and characterization of lactic acid bacteria for probiotic application from plant sources. Int. J. Adv. Res. 2017, 5, 869–876. [Google Scholar] [CrossRef] [Green Version]
- Breed, R.S.; Bergey, D.H. Bergey’s Manual of Determinative Bacteriology, 7th ed.; Williams & Wilkins Co.: Baltimore, MD, USA, 1957. [Google Scholar]
- Guo, X.-H.; Kim, J.-M.; Nam, H.-M.; Park, S.-Y.; Kim, J.-M. Screening lactic acid bacteria from swine origins for multistrain probiotics based on in vitro functional properties. Anaerobe 2010, 16, 321–326. [Google Scholar] [CrossRef] [PubMed]
- Singhal, N.; Singh, N.S.; Mohanty, S.; Kumar, M.; Virdi, J.S. Rhizospheric lactobacillus plantarum (Lactiplantibacillus plantarum) strains exhibit bile salt hydrolysis, hypocholestrolemic and probiotic capabilities in vitro. Sci. Rep. 2021, 11, 15288. [Google Scholar] [CrossRef] [PubMed]
- Kılıç, G.B.; Karahan, A.G. Identification of lactic acid bacteria isolated from the fecal samples of healthy humans and patients with dyspepsia, and determination of their ph, bile, and antibiotic tolerance properties. J. Mol. Microbiol. Biotechnol. 2010, 18, 220–229. [Google Scholar] [CrossRef]
- Yadav, R.; Puniya, A.K.; Shukla, P. Probiotic properties of lactobacillus plantarum rypr1 from an indigenous fermented beverage Raabadi. Front. Microbiol. 2016, 7, 1683. [Google Scholar] [CrossRef] [Green Version]
- Gharbi, Y.; Fhoula, I.; Ruas-Madiedo, P.; Afef, N.; Boudabous, A.; Gueimonde, M.; Ouzari, H.-I. In-vitro characterization of potentially probiotic lactobacillus strains isolated from human microbiota: Interaction with pathogenic bacteria and the enteric cell line HT29. Ann. Microbiol. 2018, 69, 61–72. [Google Scholar] [CrossRef]
- Jamaly, N. Probiotic potential of lactobacillus strains isolated from known popular traditional Moroccan dairy products. Br. Microbiol. Res. J. 2011, 1, 79–94. [Google Scholar] [CrossRef]
- Ghazanfar, S. Study on the Effect of Dietary Supplementation of Saccharomyces cerevisiae on Performance of Dairy Cattle and Heifers. Ph.D. Thesis, Department of Microbiology, Quaid-i-Azam University, Islamabad, Pakistan, 2016. [Google Scholar]
- Nadri, M.H.; Salim, Y.; Basar, N.; Yahya, A.; Zulkifli, R.M. Antioxidant activities and tyrosinase inhibition effects of phaleria macrocarpa extracts. Afr. J. Tradit. Complement. Altern. Med. 2014, 11, 107–111. [Google Scholar] [CrossRef] [Green Version]
- Al Mohaini, M.; Farid, A.; Muzammal, M.; Dadrasnia, A.; Alsalman, A.J.; Al Hawaj, M.A.; Alhashem, Y.N.; Ismail, S. Pathological study of Pasteurella Multocida Recombinant Clone ABA392. Pak. J. Med. Health Sci. 2022, 16, 1112–1116. [Google Scholar] [CrossRef]
- Adesulu-Dahunsi, A.T.; Jeyaram, K.; Sanni, A.I. Probiotic and technological properties of exopolysaccharide producing lactic acid bacteria isolated from cereal-based Nigerian fermented food products. Food Control 2018, 92, 225–231. [Google Scholar] [CrossRef] [Green Version]
- Hattingh, M.; Alexander, A.; Meijering, I.; Van, R.C.; Dicks, L.M. Amylolytic strains of lactobacillus plantarum isolated from Barley. Afr. J. Biotechnol. 2015, 14, 310–318. [Google Scholar] [CrossRef] [Green Version]
- Khalil, T.; Okla, M.K.; Al-Qahtani, W.H.; Ali, F.; Zahra, M.; Shakeela, Q.; Ahmed, S.; Akhtar, N.; AbdElgawad, H.; Asif, R.; et al. Tracing probiotic producing bacterial species from gut of buffalo (bubalus bubalis), south-east-Asia. Braz. J. Biol. 2022, 84, e259094. [Google Scholar] [CrossRef]
- Atanassova, M. Isolation and partial biochemical characterization of a proteinaceous anti-bacteria and anti-yeast compound produced by Lacticaseibacillus paracasei subsp. paracasei strain M3. Int. J. Food Microbiol. 2003, 87, 63–73. [Google Scholar] [CrossRef]
- Khan, A.N.; Yasmin, H.; Ghazanfar, S.; Hassan, M.N.; Keyani, R.; Khan, I.; Gohar, M.; Shahzad, A.; Hashim, M.J.; Ahmad, A. Antagonistic, anti-oxidant, anti-inflammatory and anti-diabetic probiotic potential of lactobacillus agilis isolated from the rhizosphere of the medicinal plants. Saudi J. Biol. Sci. 2021, 28, 6069–6076. [Google Scholar] [CrossRef] [PubMed]
- Singhvi, M.; Joshi, D.; Adsul, M.; Varma, A.; Gokhale, D. D-(−)-lactic acid production from cellobiose and cellulose by lactobacillus lactis mutant RM2-24. Curr. Green Chem. 2010, 12, 1106–1109. [Google Scholar] [CrossRef]
- Dinçer, E.; Kıvanç, M. Lipolytic activity of lactic acid bacteria isolated from Turkish pastırma. Anadolu Univ. J. Sci. Technol.–C Life Sci. Biotechnol. 2018, 7, 12–19. [Google Scholar] [CrossRef]
- Farid, A.; Shah, A.H.; Ayaz, M.; Amin, A.; Yaseen, M.; Ullah, H.; Haq, F. Comparative study of biological activity of glutathione, sodium tungstate and glutathione-tungstate mixture. Afr. J. Biotechnol. 2012, 11, 10431–10437. [Google Scholar]
- Abid, R.; Ghazanfar, S.; Farid, A.; Sulaman, S.M.; Idrees, M.; Amen, R.A.; Muzammal, M.; Shahzad, M.K.; Mohamed, M.O.; Khaled, A.A.; et al. Pharmacological Properties of 4′, 5,7-Trihydroxyflavone (Apigenin) and Its Impact on Cell Signaling Pathways. Molecules 2022, 27, 4304. [Google Scholar] [CrossRef]
- Patel, M.; Siddiqui, A.J.; Hamadou, W.S.; Surti, M.; Awadelkareem, A.M.; Ashraf, S.A.; Alreshidi, M.; Snoussi, M.; Rizvi, S.M.; Bardakci, F.; et al. Inhibition of bacterial adhesion and antibiofilm activities of a glycolipid biosurfactant from lactobacillus rhamnosus with its physicochemical and functional properties. Antibiotics 2021, 10, 1546. [Google Scholar] [CrossRef]
- Abriouel, H.; Casado Muñoz, M.d.C.; Lavilla Lerma, L.; Pérez Montoro, B.; Bockelmann, W.; Pichner, R.; Kabisch, J.; Cho, G.-S.; Franz, C.M.A.P.; Gálvez, A.; et al. New insights in antibiotic resistance of lactobacillus species from fermented foods. Int. Food Res. J. 2015, 78, 465–481. [Google Scholar] [CrossRef]
- Alsalman, A.J.; Farid, A.; Al Mohaini, M.; Al Hawaj, M.A.; Muzammal, M.; Khan, M.H.; Dadrasnia, A.; Alhashem, Y.N.; Ghazanfar, S.; Almusalami, E.M.; et al. Analysis and Characterization of Chitinase in Bacillus salmalaya Strain 139SI. Int. J. Curr. Res. Rev. 2022, 14, 31–36. [Google Scholar] [CrossRef]
- Al Hawaj, M.A.; Farid, A.; Al Mohaini, M.; Alsalman, A.J.; Muzammal, M.; Khan, M.H.; Dadrasnia, A.; Alhashem, Y.N.; Ghazanfar, S.; Almusalami, E.M.; et al. Biosurfactant Screening and Antibiotic Analysis of Bacillus salmalaya. Int. J. Cur. Res. Rev. 2022, 14, 56–64. [Google Scholar] [CrossRef]
- Al Mohaini, M.; Farid, A.; Alsalman, A.J.; Al Hawaj, M.A.; Alhashem, Y.N.; Ghazanfar, S.; Muzammal, M.; Khan, M.H.; Dadrasnia, A.; Ismail, S. Screening of Anticancer and Immunomodulatory Properties of Recombinant pQE-HAS113 Clone Derived from Streptococcus Equi. Pak. J. Med. Health Sci. 2022, 16, 1100. [Google Scholar] [CrossRef]
- Angmo, K.; Kumari, A.; Savitri; Bhalla, T.C. Probiotic characterization of lactic acid bacteria isolated from fermented foods and beverage of Ladakh. LWT—Food Sci. Tech. 2016, 66, 428–435. [Google Scholar] [CrossRef]
- FAO/WHO. Probiotics in food. In Health and Nutritional Properties and Guidelines for Evaluation; FAO Food and Nutrition Paper 85; FAO: Rome, Italy, 2006. [Google Scholar]
- Saadullah, M.; Asif, M.; Farid, A.; Naseem, F.; Rashid, S.A.; Ghazanfar, S.; Muzammal, M.; Ahmad, S.; Bin Jardan, Y.A.; Alshaya, H.; et al. A Novel Distachionate from Breynia distachia Treats Inflammations by Modulating COX-2 and Inflammatory Cytokines in Rat Liver Tissue. Molecules 2022, 27, 2596. [Google Scholar] [CrossRef]
- Borah, D.; Gogoi, O.; Adhikari, C.; Kakoti, B.B. Isolation and characterization of the new indigenous Staphylococcus sp. DBOCP06 as a probiotic bacterium from traditionally fermented fish and meat products of Assam State. Egypt. J. Basic Appl. Sci. 2016, 3, 232–240. [Google Scholar] [CrossRef] [Green Version]
- Al Mohaini, M.; Farid, A.; Muzammal, M.; Gazanffar, S.; Dadrasnia, A.; Alsalman, A.J.; Al Hawaj, M.A.; Alhashem, Y.N.; Ismail, S. Enhancing Lipase Production of Bacillus salmalay Strain 139SI Using Different Carbon Sources and Surfactants. Appl. Microbiol. 2022, 2, 237–247. [Google Scholar] [CrossRef]
- Ghalouni, E.; Hassaine, O.; Karam, N.-E. Phenotypic identification and technological characterization of lactic acid bacteria isolated from l’ben, an Algerian traditional fermented cow milk. J. Pure Appl. Microbiol. 2018, 12, 521–532. [Google Scholar] [CrossRef]
- Alsalman, A.J.; Farid, A.; Al Mohaini, M.; Muzammal, M.; Khan, M.H.; Dadrasnia, A.; Ismail, S. Chitinase Activity by Chitin Degrading Strain (Bacillus Salmalaya) in Shrimp Waste. Int. J. Curr. Res. Rev. 2022, 14, 11–17. [Google Scholar] [CrossRef]
- Tokatlı, M.; Gülgör, G.; Bağder Elmacı, S.; Arslankoz İşleyen, N.; Özçelik, F. In vitro properties of potential probiotic indigenous lactic acid bacteria originating from traditional pickles. Biomed Res. Int. 2015, 2015, 315819. [Google Scholar] [CrossRef] [Green Version]
- Adnan, M.; Patel, M.; Hadi, S. Functional and health promoting inherent attributes ofenterococcus hiraeF2 as a novel probiotic isolated from the digestive tract of the freshwater fishcatla catla. PeerJ 2017, 5, e3085. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Abid, S.; Farid, A.; Abid, R.; Rehman, M.U.; Alsanie, W.F.; Alhomrani, M.; Alamri, A.S.; Asdaq, S.M.B.; Hefft, D.I.; Saqib, S.; et al. Identification, Biochemical Characterization, and Safety Attributes of Locally Isolated Lactobacillus fermentum from Bubalus bubalis (Buffalo) Milk as a Probiotic. Microorganisms 2022, 10, 954. [Google Scholar] [CrossRef]
- Shivangi, S.; Devi, P.B.; Ragul, K.; Shetty, P.H. Probiotic Potential of Bacillus Strains Isolated from an Acidic Fermented Food Idli. Probiotics Antimicrob. Proteins 2020, 12, 1502–1513. [Google Scholar]
- Fatima, S.; Muzammal, M.; Khan, M.A.; Farid, A.; Kamran, M.; Qayum, J.; Qureshi, M.; Khan, M.N.; Khan, M.A. CRISPR/Cas9 endonucleases: A new era of genetic engineering. Abasyn J. life Sci. 2021, 4, 29–39. [Google Scholar] [CrossRef]
- Farid, A.; Javed, R.; Hayat, M.; Muzammal, M.; Khan, M.H.; Ismail, S.; Rashid, S.A. Screening of Strobilanthes urticifolia wall.ex kuntze for Antitermite and insecticidal activities. Abasyn J. life Sci. 2021, 4, 40–45. [Google Scholar] [CrossRef]
- Amaretti, A.; Di Nunzio, M.; Pompei, A.; Raimondi, S.; Rossi, M.; Bordoni, A. Antioxidant properties of potentially probiotic bacteria: In vitro and in vivo activities. Appl. Microbiol. Biotechnol. 2013, 97, 809–817. [Google Scholar] [CrossRef]
- Hameed, A.; Condò, C.; Tauseef, I.; Idrees, M.; Ghazanfar, S.; Farid, A.; Muzammal, M.; Al Mohaini, M.; Alsalman, A.J.; Al Hawaj, M.A.; et al. Isolation and Characterization of a Cholesterol-Lowering Bacteria from Bubalus bubalis Raw Milk. Fermentation 2022, 8, 163. [Google Scholar] [CrossRef]
- Ghazanfar, S. Understanding the Mechanism of Action of Indigenous Target Probiotic Yeast: Linking the Manipulation of Gut Microbiota and Performance in Animals. In Saccharomyces; Intech Open: London, UK, 2021. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).