Yeast Nanometric Scale Oscillations Highlights Fibronectin Induced Changes in C. albicans
Abstract
1. Introduction
2. Materials and Methods
2.1. Yeast Strains
2.2. Experimental Procedures
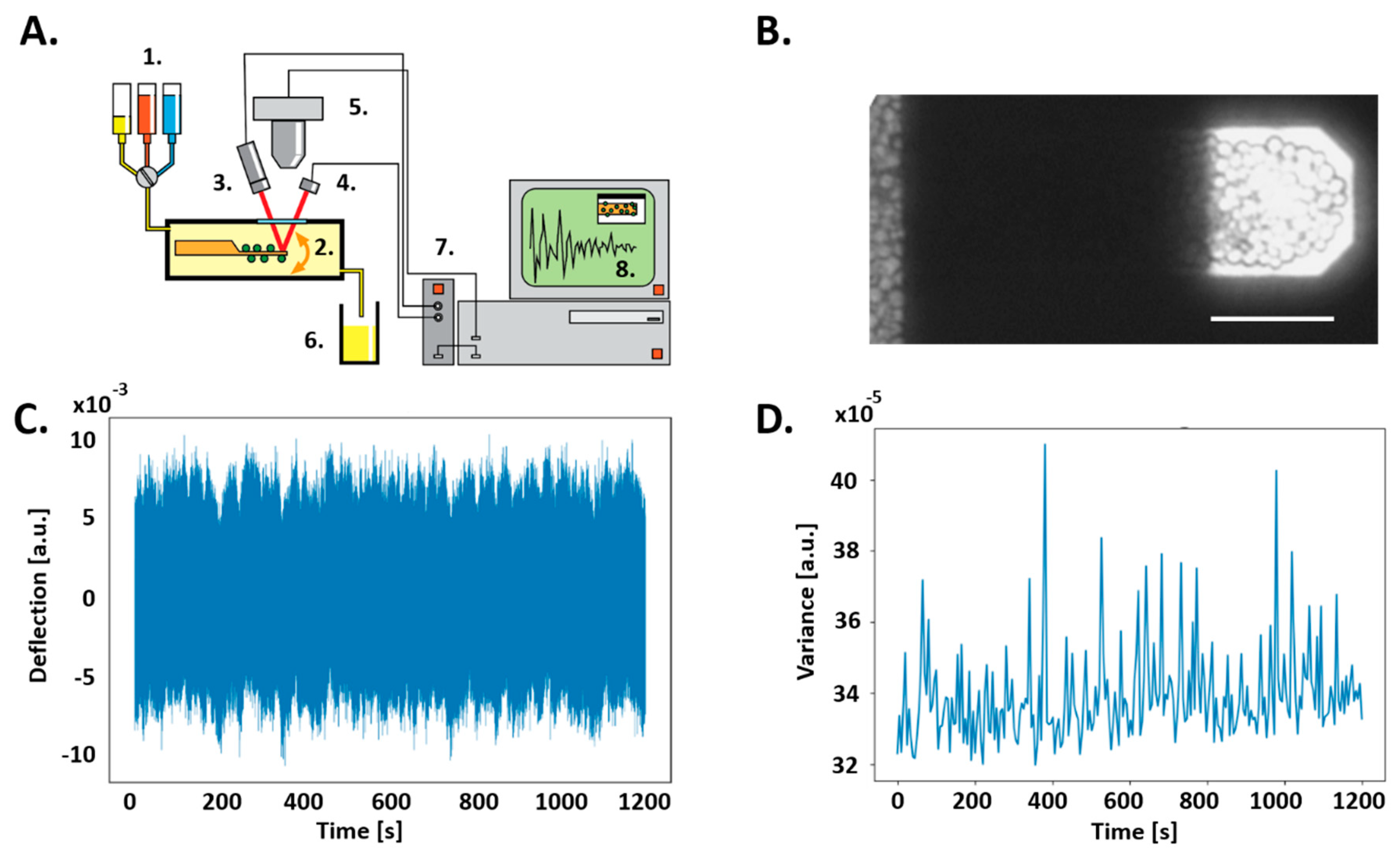
2.3. Nanomotion Detector
2.4. Software and Nanomotion Analysis
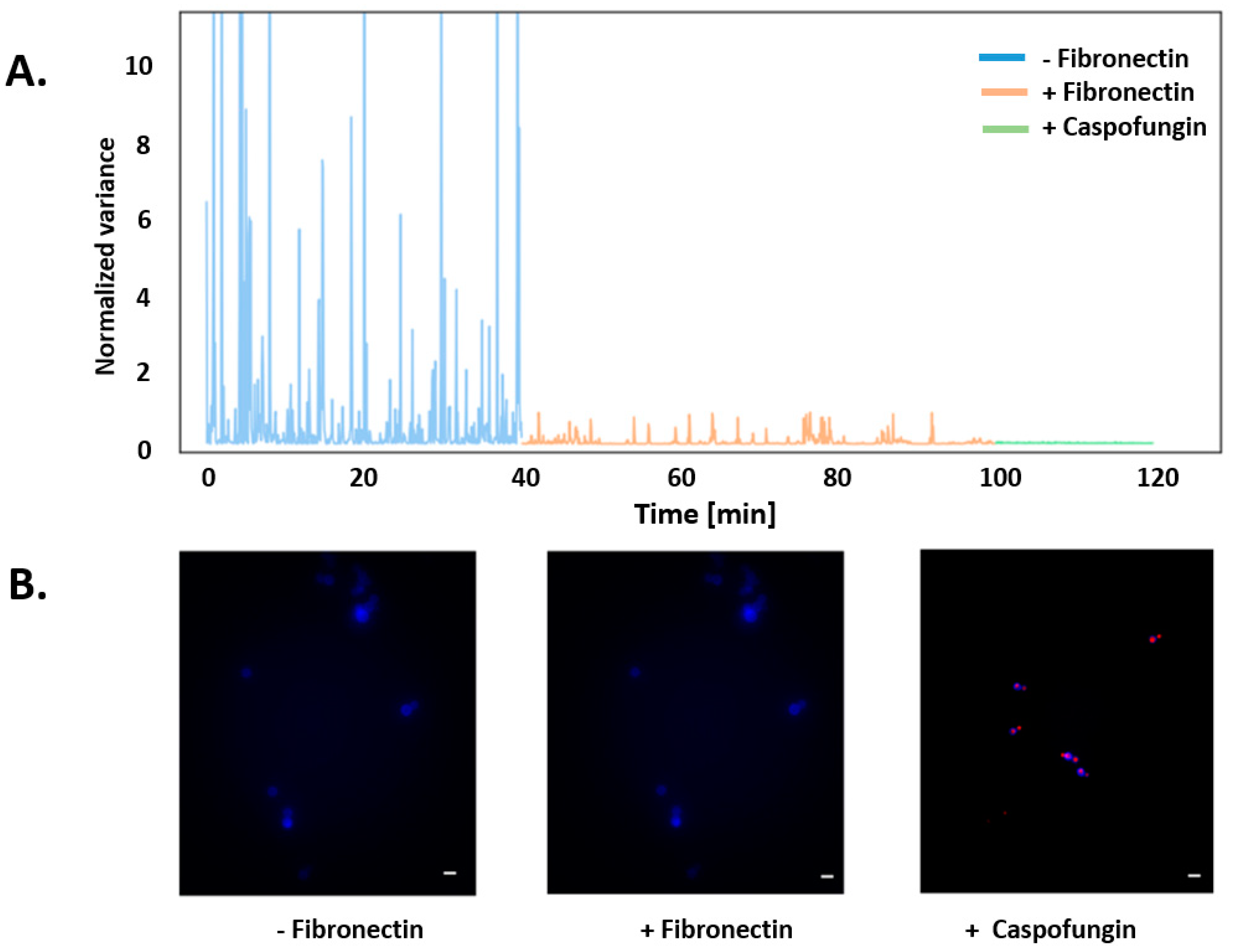
2.5. Viability Assay
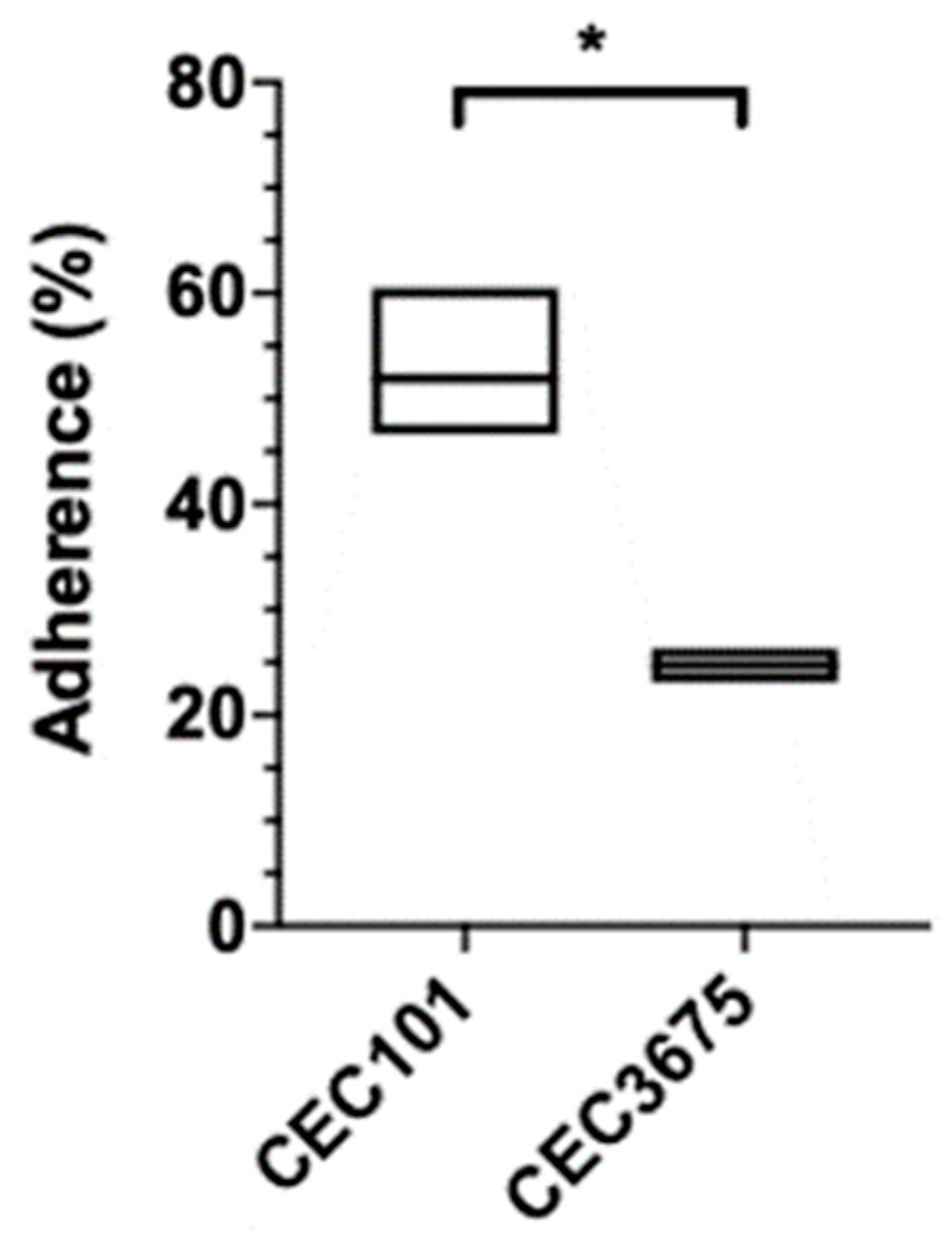
2.6. Adhesion Assay
2.7. Statistical Analysis
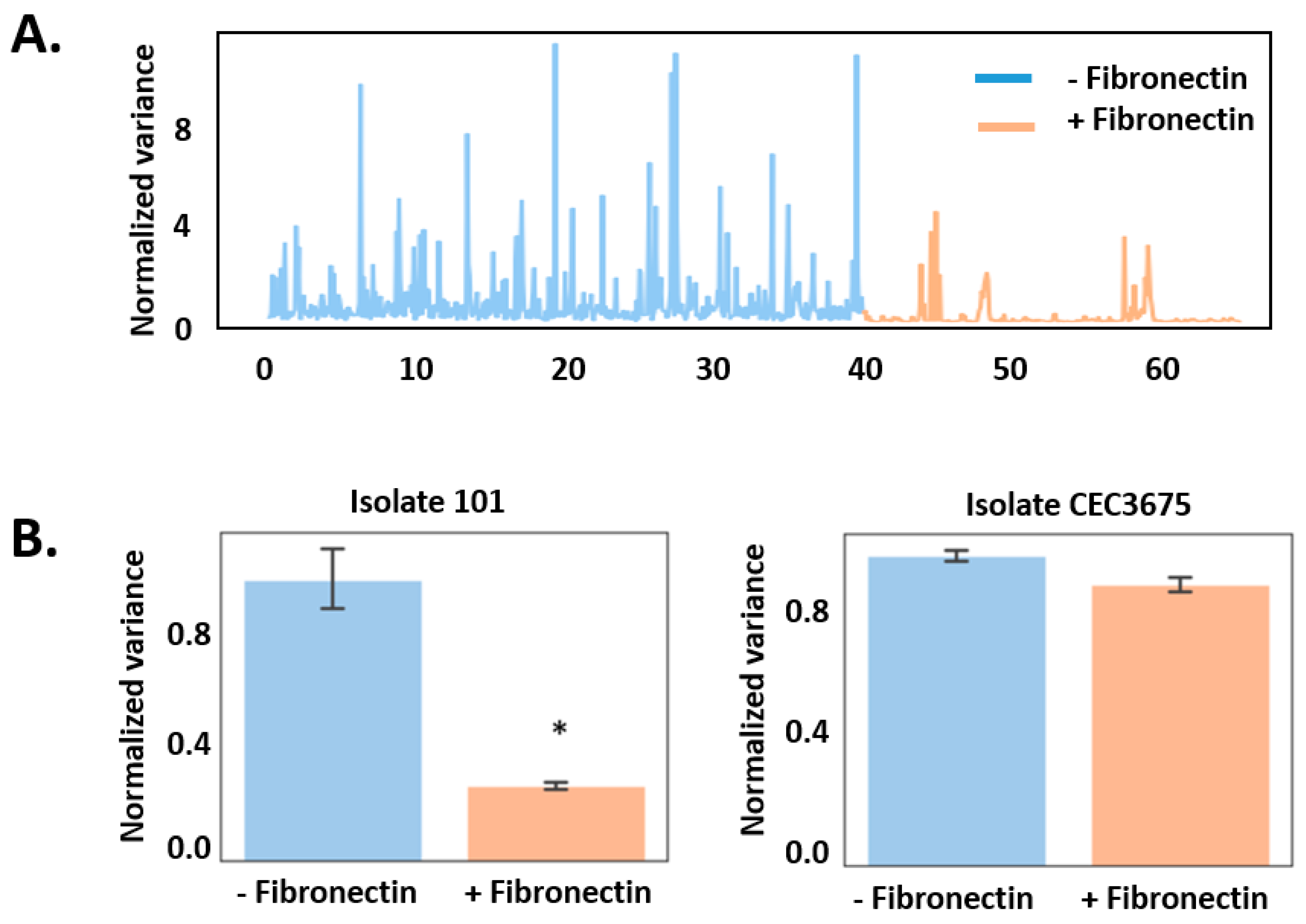
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Willaert, R.; Kasas, S.; Devreese, B.; Dietler, G. Yeast Nanobiotechnology. Fermentation 2016, 2, 18. [Google Scholar] [CrossRef]
- Formosa-Dague, C.; Duval, R.E.; Dague, E. Cell biology of microbes and pharmacology of antimicrobial drugs explored by Atomic Force Microscopy. Semin. Cell Dev. Biol. 2018, 73, 165–176. [Google Scholar] [CrossRef]
- Alsteens, D.; Müller, D.J.; Dufrêne, Y.F. Multiparametric Atomic Force Microscopy Imaging of Biomolecular and Cellular Systems. Acc. Chem. Res. 2017, 50, 924–931. [Google Scholar] [CrossRef] [PubMed]
- Kasas, S.; Stupar, P.; Dietler, G. AFM contribution to unveil pro- and eukaryotic cell mechanical properties. Semin. Cell Dev. Biol. 2018, 73, 177–187. [Google Scholar] [CrossRef] [PubMed]
- Longo, G.; Alonso-Sarduy, L.; Rio, L.M.; Bizzini, A.; Trampuz, A.; Notz, J.; Dietler, G.; Kasas, S. Rapid detection of bacterial resistance to antibiotics using AFM cantilevers as nanomechanical sensors. Nat. Nanotechnol. 2013, 8, 522–526. [Google Scholar] [CrossRef]
- Kohler, A.C.; Venturelli, L.; Longo, G.; Dietler, G.; Kasas, S. Nanomotion detection based on atomic force microscopy cantilevers. Cell Surf. 2019, 5, 100021. [Google Scholar] [CrossRef]
- Villalba, M.I.; Stupar, P.; Chomicki, W.; Bertacchi, M.; Dietler, G.; Arnal, L.; Vela, M.E.; Yantorno, O.; Kasas, S. Nanomotion Detection Method for Testing Antibiotic Resistance and Susceptibility of Slow-Growing Bacteria. Small 2018, 14, 1702671. [Google Scholar] [CrossRef]
- Stupar, P.; Opota, O.; Longo, G.; Prod’hom, G.; Dietler, G.; Greub, G.; Kasas, S. Nanomechanical sensor applied to blood culture pellets: A fast approach to determine the antibiotic susceptibility against agents of bloodstream infections. Clin. Microbiol. Infect. 2017, 23, 400–405. [Google Scholar] [CrossRef]
- Mustazzolu, A.; Venturelli, L.; Dinarelli, S.; Brown, K.; Floto, R.A.; Dietler, G.; Fattorini, L.; Kasas, S.; Girasole, M.; Longo, G. A rapid unravelling of mycobacterial activity and of their susceptibility to antibiotics. Antimicrob. Agents Chemother. 2019, 63, 02194-18. [Google Scholar] [CrossRef]
- Kasas, S.; Ruggeri, F.S.; Benadiba, C.; Maillard, C.; Stupar, P.; Tournu, H.; Dietler, G.; Longo, G. Detecting nanoscale vibrations as signature of life. Proc. Natl. Acad. Sci. USA 2015, 112, 378–381. [Google Scholar] [CrossRef]
- Brown, G.D.; Denning, D.W.; Levitz, S.M. Tackling human fungal infections. Science 2012, 336, 647. [Google Scholar] [CrossRef] [PubMed]
- Ghannoum, M.A.; Jurevic, R.J.; Mukherjee, P.K.; Cui, F.; Sikaroodi, M.; Naqvi, A.; Gillevet, P.M. Characterization of the Oral Fungal Microbiome (Mycobiome) in Healthy Individuals. PLoS Pathog. 2010, 6, e1000713. [Google Scholar] [CrossRef] [PubMed]
- Rupp, S. Interactions of the fungal pathogen Candida albicans with the host. Future Microbiol. 2007, 2, 141–151. [Google Scholar] [CrossRef] [PubMed]
- Filler, S.G.; Sheppard, D.C. Fungal Invasion of Normally Non-Phagocytic Host Cells. PLoS Pathog. 2006, 2, e129. [Google Scholar] [CrossRef] [PubMed]
- Nobile, C.J.; Johnson, A.D. Candida albicans Biofilms and Human Disease. Annu. Rev. Microbiol. 2015, 69, 71–92. [Google Scholar] [CrossRef] [PubMed]
- Chaffin, W.L.; López-Ribot, J.L.; Casanova, M.; Gozalbo, D.; Martínez, J.P. Cell Wall and Secreted Proteins ofCandida albicans: Identification, Function, and Expression. Microbiol. Mol. Biol. Rev. 1998, 62, 130–180. [Google Scholar] [CrossRef]
- Sundstrom, P. Adhesion in Candida spp. Cell. Microbiol. 2002, 4, 461–469. [Google Scholar] [CrossRef]
- Filler, S.G. Candida-host cell receptor-ligand interactions. Curr. Opin. Microbiol. 2006, 9, 333–339. [Google Scholar] [CrossRef]
- Ghannoum, M.A.; Burns, G.R.; Abu Elteen, K.; Radwan, S.S. Experimental evidence for the role of lipids in adherence of Candida spp. to human buccal epithelial cells. Infect. Immun. 1986, 54, 189–193. [Google Scholar] [CrossRef]
- Jimenez-Lucho, V.; Ginsburg, V.; Krivan, H.C. Cryptococcus neoformans, Candida albicans, and other fungi bind specifically to the glycosphingolipid lactosylceramide (Gal beta 1-4Glc beta 1-1Cer), a possible adhesion receptor for yeasts. Infect. Immun. 1990, 58, 2085–2090. [Google Scholar] [CrossRef]
- Yu, L.; Lee, K.K.; Sheth, H.B.; Lane-Bell, P.; Srivastava, G.; Hindsgaul, O.; Paranchych, W.; Hodges, R.S.; Irvin, R.T. Fimbria-mediated adherence of Candida albicans to glycosphingolipid receptors on human buccal epithelial cells. Infect. Immun. 1994, 62, 2843–2848. [Google Scholar] [CrossRef] [PubMed]
- Cameron, B.J.; Douglas, L.J. Blood group glycolipids as epithelial cell receptors for Candida albicans. Infect. Immun. 1996, 64, 891–896. [Google Scholar] [CrossRef] [PubMed]
- Sandin, R.L.; Rogers, A.L.; Patterson, R.J.; Beneke, E.S. Evidence for mannose-mediated adherence of Candida albicans to human buccal cells in vitro. Infect. Immun. 1982, 35, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Critchley, I.A.; Douglas, L.J. Role of glycosides as epithelial cell receptors for Candida albicans. J. Gen. Microbiol. 1987, 133, 637–643. [Google Scholar] [CrossRef]
- Macura, A.B.; Tondyra, E. Influence of some carbohydrates and concanavalin A on the adherence of Candida albicans in vitro to buccal epithelial cells. Zentralbl. Bakteriol. 1989, 272, 196–201. [Google Scholar] [CrossRef]
- Brassart, D.; Woltz, A.; Golliard, M.; Neeser, J.R. In vitro inhibition of adhesion of Candida albicans clinical isolates to human buccal epithelial cells by Fuc alpha 1----2Gal beta-bearing complex carbohydrates. Infect. Immun. 1991, 59, 1605–1613. [Google Scholar] [CrossRef]
- Donohue, D.S.; Ielasi, F.S.; Goossens, K.V.Y.; Willaert, R.G. The N-terminal part of Als1 protein from Candida albicans specifically binds fucose-containing glycans. Mol. Microbiol. 2011, 80, 1667–1679. [Google Scholar] [CrossRef]
- Ielasi, F.S.; Alioscha-Perez, M.; Donohue, D.; Claes, S.; Sahli, H.; Schols, D.; Willaert, R.G. Lectin-glycan interaction network-based identification of host receptors of microbial pathogenic adhesins. MBio 2016, 7. [Google Scholar] [CrossRef]
- Everest-Dass, A.V.; Kolarich, D.; Pascovici, D.; Packer, N.H. Blood group antigen expression is involved in C. albicans interaction with buccal epithelial cells. Glycoconj. J. 2017, 34, 31–50. [Google Scholar] [CrossRef] [PubMed]
- Calderone, R.A.; Scheld, W.M. Role of fibronectin in the pathogenesis of candidal infections. Rev. Infect. Dis. 1987, 9, S400–S403. [Google Scholar] [CrossRef]
- Nett, J.E.; Cabezas-Olcoz, J.; Marchillo, K.; Mosher, D.F.; Andes, D.R. Targeting fibronectin to disrupt in vivo Candida albicans biofilms. Antimicrob. Agents Chemother. 2016, 60, 3152–3155. [Google Scholar] [CrossRef] [PubMed]
- Ropars, J.; Maufrais, C.; Diogo, D.; Marcet-Houben, M.; Perin, A.; Sertour, N.; Mosca, K.; Permal, E.; Laval, G.; Bouchier, C.; et al. Gene flow contributes to diversification of the major fungal pathogen Candida albicans. Nat. Commun. 2018, 9, 2253. [Google Scholar] [CrossRef] [PubMed]
- Murciano, C.; Moyes, D.L.; Runglall, M.; Tobouti, P.; Islam, A.; Hoyer, L.L.; Naglik, J.R. Evaluation of the Role of Candida albicans Agglutinin-Like Sequence (Als) Proteins in Human Oral Epithelial Cell Interactions. PLoS ONE 2012, 7, e33362. [Google Scholar] [CrossRef] [PubMed]
- Schönherr, F.A.; Sparber, F.; Kirchner, F.R.; Guiducci, E.; Trautwein-Weidner, K.; Gladiator, A.; Sertour, N.; Hetzel, U.; Le, G.T.T.; Pavelka, N.; et al. The intraspecies diversity of C. albicans triggers qualitatively and temporally distinct host responses that determine the balance between commensalism and pathogenicity. Mucosal Immunol. 2017, 10, 1335–1350. [Google Scholar] [CrossRef] [PubMed]
- Skerl, K.G.; Calderone, R.A.; Segal, E.; Sreevalsan, T.; Scheld, W.M. In vitro binding of Candida albicans yeast cells to human fibronectin. Can. J. Microbiol. 1984, 30, 221–227. [Google Scholar] [CrossRef] [PubMed]
- Douglas, L.J. Adhesin—Receptor interactions in the attachment of Candida albicans to host epithelial cells. Can. J. Bot. 1995, 73, 1147–1153. [Google Scholar] [CrossRef]
- Alsteens, D.; Beaussart, A.; Derclaye, S.; El-Kirat-Chatel, S.; Park, H.R.; Lipke, P.N.; Dufrêne, Y.F. Single-cell force spectroscopy of Als-mediated fungal adhesion. Anal. Methods 2013, 5, 3657–3662. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Palecek, S.P. EAP1, a Candida albicans Gene Involved in Binding Human Epithelial Cells. Eukaryot. Cell 2003, 2, 1266–1273. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Svarovsky, M.J.; Karlsson, A.J.; Wagner, J.P.; Marchillo, K.; Oshel, P.; Andes, D.; Palecek, S.P. Eap1p, an adhesin that mediates Candida albicans biofilm formation in vitro and in vivo. Eukaryot. Cell 2007, 6, 931–939. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Palecek, S.P. Distinct domains of the Candida albicans adhesin EAP1 p mediate cell-cell and cell-substrate interactions. Microbiology 2008, 154, 1193–1203. [Google Scholar] [CrossRef]
- Hooshdaran, M.Z.; Barker, K.S.; Hilliard, G.M.; Kusch, H.; Morschhäuser, J.; Rogers, P.D. Proteomic analysis of azole resistance in Candida albicans clinical isolates. Antimicrob. Agents Chemother. 2004, 48, 2733–2735. [Google Scholar] [CrossRef] [PubMed]
- Singleton, D.R.; Fidel, P.L.; Wozniak, K.L.; Hazen, K.C. Contribution of cell surface hydrophobicity protein 1 (Csh1p) to virulence of hydrophobic Candida albicans serotype A cells. FEMS Microbiol. Lett. 2005, 244, 373–377. [Google Scholar] [CrossRef] [PubMed]
- de Groot, P.W.J.; Hellingwerf, K.J.; Klis, F.M. Genome-wide identification of fungal GPI proteins. Yeast 2003, 20, 781–796. [Google Scholar] [CrossRef] [PubMed]
- McCall, A.D.; Pathirana, R.U.; Prabhakar, A.; Cullen, P.J.; Edgerton, M. Candida albicans biofilm development is governed by cooperative attachment and adhesion maintenance proteins. NPJ Biofilms Microbiomes 2019, 5, 21. [Google Scholar] [CrossRef]
- Watts, H.; Cheah, F.S.; Hube, B.; Sanglard, D.; Gow, N.A. Altered adherence in strains of Candida albicans harbouring null mutations in secreted aspartic proteinase genes. FEMS Microbiol. Lett. 1998, 159, 129–135. [Google Scholar] [CrossRef]
- Naglik, J.R.; Challacombe, S.J.; Hube, B. Candida albicans Secreted Aspartyl Proteinases in Virulence and Pathogenesis. Microbiol. Mol. Biol. Rev. 2003, 67, 400–428. [Google Scholar] [CrossRef]
- Kumar, R.; Breindel, C.; Saraswat, D.; Cullen, P.J.; Edgerton, M. Candida albicans Sap6 amyloid regions function in cellular aggregation and zinc binding, and contribute to zinc acquisition. Sci. Rep. 2017, 7, 1–15. [Google Scholar] [CrossRef]
- Klotz, S.A.; Gaur, N.K.; Lake, D.F.; Chan, V.; Rauceo, J.; Lipke, P.N. Degenerate peptide recognition by Candida albicans adhesins Als5p and Als1p. Infect. Immun. 2004, 72, 2029–2034. [Google Scholar] [CrossRef]
- Sheppard, D.C.; Yeaman, M.R.; Welch, W.H.; Phan, Q.T.; Fu, Y.; Ibrahim, A.S.; Filler, S.G.; Zhang, M.; Waring, A.J.; Edwards, J.E. Functional and structural diversity in the Als protein family of Candida albicans. J. Biol. Chem. 2004, 279, 30480–30489. [Google Scholar] [CrossRef]
- Argimón, S.; Wishart, J.A.; Leng, R.; Macaskill, S.; Mavor, A.; Alexandris, T.; Nicholls, S.; Knight, A.W.; Enjalbert, B.; Walmsley, R.; et al. Developmental regulation of an adhesin gene during cellular morphogenesis in the fungal pathogen Candida albicans. Eukaryot. Cell 2007, 6, 682–692. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kohler, A.-C.; Venturelli, L.; Kannan, A.; Sanglard, D.; Dietler, G.; Willaert, R.; Kasas, S. Yeast Nanometric Scale Oscillations Highlights Fibronectin Induced Changes in C. albicans. Fermentation 2020, 6, 28. https://doi.org/10.3390/fermentation6010028
Kohler A-C, Venturelli L, Kannan A, Sanglard D, Dietler G, Willaert R, Kasas S. Yeast Nanometric Scale Oscillations Highlights Fibronectin Induced Changes in C. albicans. Fermentation. 2020; 6(1):28. https://doi.org/10.3390/fermentation6010028
Chicago/Turabian StyleKohler, Anne-Céline, Leonardo Venturelli, Abhilash Kannan, Dominique Sanglard, Giovanni Dietler, Ronnie Willaert, and Sandor Kasas. 2020. "Yeast Nanometric Scale Oscillations Highlights Fibronectin Induced Changes in C. albicans" Fermentation 6, no. 1: 28. https://doi.org/10.3390/fermentation6010028
APA StyleKohler, A.-C., Venturelli, L., Kannan, A., Sanglard, D., Dietler, G., Willaert, R., & Kasas, S. (2020). Yeast Nanometric Scale Oscillations Highlights Fibronectin Induced Changes in C. albicans. Fermentation, 6(1), 28. https://doi.org/10.3390/fermentation6010028






