Effects of Kuding Tea on the Succession and Assembly of the Fungal Community During Fermentation of Daqu
Abstract
1. Introduction
2. Materials and Methods
2.1. Daqu Block Preparation and Sample Collection
2.2. Analysis of Endogenous Factors
2.3. Total DNA Extraction, PCR Amplification, and Sequencing
2.4. Sequence Data Processing
2.5. Statistical Analysis
3. Results
3.1. Community Composition and Its Diversity
3.2. Assembly Mechanisms and Temporal Dynamics

3.3. Influential Fungal Drivers of Community Dissimilarity and Assembly

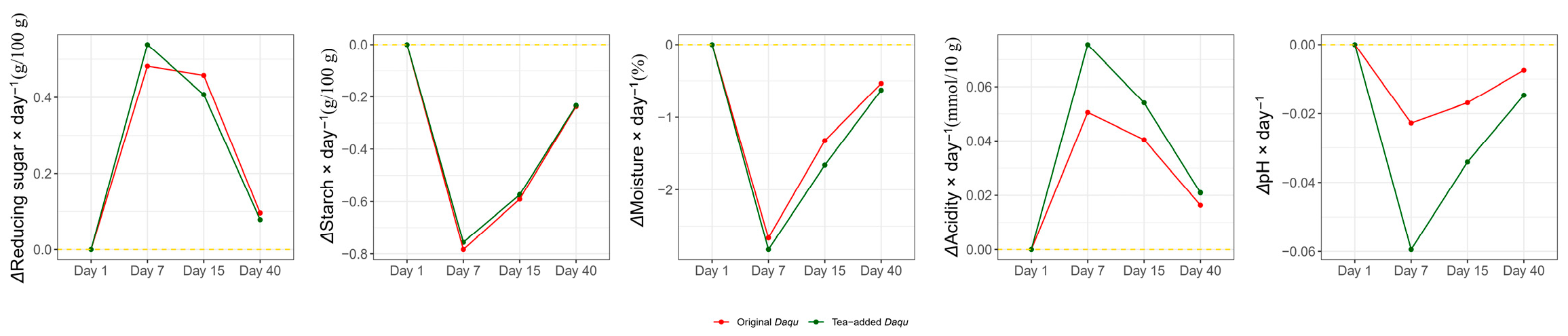
3.4. Changes in Endogenous Factors and Their Associations with Community Succession

4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Li, W.; Fan, G.; Fu, Z.; Wang, W.; Xu, Y.; Teng, C.; Zhang, C.; Yang, R.; Sun, B.; Li, X. Effects of fortification of Daqu with various yeasts on microbial community structure and flavor metabolism. Food Res. Int. 2020, 129, 108837. [Google Scholar] [CrossRef]
- Sakandar, H.A.; Hussain, R.; Khan, Q.F.; Zhang, H. Functional microbiota in Chinese traditional Baijiu and Mijiu Qu (starters): A review. Food Res. Int. 2020, 138, 109830. [Google Scholar] [CrossRef]
- Du, H.; Wang, X.S.; Zhang, Y.H.; Xu, Y. Exploring the impacts of raw materials and environments on the microbiota in Chinese Daqu starter. Int. J. Food Microbiol. 2019, 297, 32–40. [Google Scholar] [CrossRef]
- Wei, J.; Jie, W.; Baosheng, L.; Jianhui, D.; Xiaojun, W.; Zhihua, S.; Xinglin, H.; Jie, H.; Chunguang, L.; Jianqin, H.; et al. Analysis of characteristic flavor compounds in single-grain chinese baijiu brewed from different raw. Food Sci. 2020, 41, 234–238. (In Chinese) [Google Scholar] [CrossRef]
- Zhao, L.; Li, F.F.; Xiao, H.; Zhao, T.F.; Zhong, Y.X.; Hu, Z.H.; Jiang, L.; Wang, X.Y.; Wang, X.Y. Green Tea Modulates Temporal Dynamics and Environmental Adaptation of Microbial Communities in Daqu Fermentation. Fermentation 2025, 11, 511. [Google Scholar] [CrossRef]
- Wüpper, S.; Lüersen, K.; Rimbach, G. Chemical Composition, Bioactivity and Safety Aspects of Kuding Tea—From Beverage to Herbal Extract. Nutrients 2020, 12, 2796. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Zu, M.; Chen, K.; Gao, L.; Min, H.; Zhuo, W.; Chen, W.; Liu, A. Screening of neuraminidase inhibitory activities of some medicinal plants traditionally used in Lingnan Chinese medicines. BMC Complement. Altern. Med. 2018, 18, 102. [Google Scholar] [CrossRef]
- Alaoui, A.; Sahri, N.; Mahdi, I.; Fahsi, N.; El Herradi, E.H.; Sobeh, M. Argan: Phytochemical profiling and evaluation of the antioxidant, hypoglycemic, and antibacterial properties of its fruit pulp extracts. Heliyon 2024, 10, e23612. [Google Scholar] [CrossRef]
- Ahmed, M.; Sajid, A.R.; Javeed, A.; Aslam, M.; Ahsan, T.; Hussain, D.; Mateen, A.; Li, X.; Qin, P.; Ji, M. Antioxidant, antifungal, and aphicidal activity of the triterpenoids spinasterol and 22,23-dihydrospinasterol from leaves of Citrullus colocynthis L. Sci. Rep. 2022, 12, 4910. [Google Scholar] [CrossRef]
- Zhao, L.; Wang, X.; Luo, Z.; Jiang, L.; Wu, F.; Zhong, Y.; Wang, X. Effect of adding green tea on physicochemical characteristics and bacterial diversity of daqu during fermentation. Food Sci. 2024, 45, 59–68. (In Chinese) [Google Scholar] [CrossRef]
- Zhao, L.; Wang, X.; Luo, Z.; Gan, G.; Wu, F.; Zhong, Y.; Wang, X. Optimization of the fermentation process of Moutai-flavored Daqu with the addition of green tea: Impacts on bacterial succession and endogenous factors. J. Food Sci. Technol. 2025, 62, 1957–1967. [Google Scholar] [CrossRef]
- Bai, X. A Technical Study on the Tea-Flower Daqu Preparation. PhD Thesis, Anhui Agricultural University, Hefei, China, 2016. [Google Scholar]
- Nekola, J.C.; White, P.S. The distance decay of similarity in biogeography and ecology. J. Biogeogr. 1999, 26, 867–878. [Google Scholar] [CrossRef]
- Hatosy, S.M.; Martiny, J.B.H.; Sachdeva, R.; Steele, J.; Fuhrman, J.A.; Martiny, A.C. Beta diversity of marine bacteria depends on temporal scale. Ecology 2013, 94, 1898–1904. [Google Scholar] [CrossRef] [PubMed]
- MacArthur, R.; Wilson, E. Theory of Island Biogeography (MPB-1); Princeton University Press: Princeton, NJ, USA, 2015. [Google Scholar]
- Jiao, S.; Zhang, Z.; Yang, F.; Lin, Y.; Chen, W.; Wei, G. Temporal dynamics of microbial communities in microcosms in response to pollutants. Mol. Ecol. 2017, 26, 923–936. [Google Scholar] [CrossRef]
- Preston, F.W. Time and Space and the Variation of Species. Ecology 1960, 41, 611–627. [Google Scholar] [CrossRef]
- Vermeij, G.J. Species-Diversity in-Space and Time; Rosenzweig, M.L., Ed.; Cambridge University Press: New York, NY, USA, 1995; Volume 270, pp. 503–504. [Google Scholar]
- Ahluwalia, S.; Holmes, I.; von May, R.; Rabosky, D.L.; Rabosky, A.R.D. Assembling microbial communities: A genomic analysis of a natural experiment in neotropical bamboo internodes. PeerJ 2022, 10, e13958. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, Q.; Chen, Q.; Hou, S.; Jiao, N.; Zheng, Q. Deciphering community assembly processes of the microbial community in subtropical coastal-estuarine seawater over a 6-year exploration. ISME Commun. 2025, 5, ycaf091. [Google Scholar] [CrossRef]
- Zuo, Q.C.; Huang, Y.G.; Guo, M. Evaluation of bacterial diversity during fermentation process: A comparison between handmade and machine-made high-temperature Daqu of Maotai-flavor liquor. Ann. Microbiol. 2020, 70, 57. [Google Scholar] [CrossRef]
- QB/T 4257-2011; General Methods of Analysis for Daqu. China Light Industry Press: Beijing, China, 2012.
- GB 5009.7-2016; Determination of Reducing Sugar in Foods. China Standard Press: Beijing, China, 2016.
- Huang, T.; Wei, J.; Zhou, S.; Hao, W.; Liu, X.; Yu, Z.; Zhuang, W.; Zeng, Z. Fungal community structure and succession of light-flavor Baijiu during fermentation process. LWT-Food Sci. Technol. 2025, 224, 117825. [Google Scholar] [CrossRef]
- Dai, Y.; Li, Z.; Tian, Z. Analysis of bacterial diversity in daqu and fermented grains for maotai-flavor liquor. Food Sci. 2019, 40, 152–159. (In Chinese) [Google Scholar] [CrossRef]
- Větrovský, T.; Morais, D.; Kohout, P.; Lepinay, C.; Algora, C.; Awokunle Hollá, S.; Bahnmann, B.D.; Bílohnědá, K.; Brabcová, V.; D’Alò, F.; et al. GlobalFungi, a global database of fungal occurrences from high-throughput-sequencing metabarcoding studies. Sci. Data 2020, 7, 228. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Kessy, A.; Allan, Z.; Timo, P.; Raivo, P.; Filipp, I.; Henrik, N.R.; Urmas, K. UNITE General FASTA Release for Fungi. Version 19.02.2025. Available online: https://doi.org/10.15156/BIO/3301229 (accessed on 19 February 2025).
- White, E.P.; Adler, P.B.; Lauenroth, W.K.; Gill, R.A.; Greenberg, D.; Kaufman, D.M.; Rassweiler, A.; Rusak, J.A.; Smith, M.D.; Steinbeck, J.R.; et al. A comparison of the species-time relationship across ecosystems and taxonomic groups. Oikos 2006, 112, 185–195. [Google Scholar] [CrossRef]
- Dengler, J. Which function describes the species-area relationship best? A review and empirical evaluation. J. Biogeogr. 2009, 36, 728–744. [Google Scholar] [CrossRef]
- Hu, A.; Nie, Y.X.; Yu, G.R.; Han, C.H.; He, J.H.; He, N.P.; Liu, S.R.; Deng, J.; Shen, W.H.; Zhang, G.X. Diurnal Temperature Variation and Plants Drive Latitudinal Patterns in Seasonal Dynamics of Soil Microbial Community. Front. Microbiol. 2019, 10, 674. [Google Scholar] [CrossRef] [PubMed]
- Ning, D.L.; Yuan, M.T.; Wu, L.W.; Zhang, Y.; Guo, X.; Zhou, X.S.; Yang, Y.F.; Arkin, A.P.; Firestone, M.K.; Zhou, J.Z. A quantitative framework reveals ecological drivers of grassland microbial community assembly in response to warming. Nat. Commun. 2020, 11, 4717. [Google Scholar] [CrossRef]
- Stegen, J.C.; Lin, X.J.; Fredrickson, J.K.; Konopka, A.E. Estimating and mapping ecological processes influencing microbial community assembly. Front. Microbiol. 2015, 6, 370. [Google Scholar] [CrossRef] [PubMed]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Archer, E. rfPermute: Estimate Permutation P-Values for Random Forest Importance Metrics. Available online: https://CRAN.R-project.org/package=rfPermute (accessed on 10 June 2013).
- Zhao, L.; Yuan, H.; Zheng, K.; Zhuo, M.; Wu, F.; Zhang, Y.; Jiang, L.; Guangdong, G.; Wang, X. Functional predictions of bacterial communities associated with sorghum cultivars across different rhizocompartments. Emir. J. Food Agric. 2025, 37, 1–12. [Google Scholar] [CrossRef]
- Legendre, P.; Legendre, L. Numerical Ecology, 3rd ed.; Elsevier: Amsterdam, The Netherlands, 2012; Volume 24. [Google Scholar]
- Oksanen, J.; Simpson, G.; Blanchet, F.; Kindt, R.; Legendre, P.; Minchin, P.; O’Hara, R.; Solymos, P.; Stevens, M.; Szoecs, E.; et al. Vegan: Community Ecology Package (R Package Version 2.7-3). Available online: https://CRAN.R-project.org/package=vegan (accessed on 4 March 2025).
- Braak, C.J.F.T. Canonical correspondence analysis: A new eigenvector technique for multivariate direct gradient analysis. Ecology 1986, 67, 1167–1179. [Google Scholar] [CrossRef]
- Yun, Y.H.; Deng, B.C.; Cao, D.S.; Wang, W.T.; Liang, Y.Z. Variable importance analysis based on rank aggregation with applications in metabolomics for biomarker discovery. Anal. Chim. Acta 2016, 911, 27–34. [Google Scholar] [CrossRef]
- Zhao, C.; Su, W.; Mu, C.; Zheng, P.; Wang, H. Effects of different raw materials on microbial community structure and diversity in Jiang-flavor Daqu. Food Sci. 2022, 43, 133–141. (In Chinese) [Google Scholar] [CrossRef]
- Zeng, Q.; Lv, S.; Li, X.; Hu, X.; Liang, Y.; Fan, C.; Duan, J.; Deng, T. Bioactive ingredients and microbial diversity of fuzhuan tea produced from different raw materials. Food Sci. 2020, 41, 69–77. (In Chinese) [Google Scholar] [CrossRef]
- Ren, H.; Cai, Z.; Du, C.; Li, Z.; Guo, X.; Wang, Y.; Ma, D.; Zhang, B.; Zheng, Y. Interrelated spatiotemporal variations between bacterial community and physicochemical factors in pit mud of Chinese strong-flavor Baijiu. LWT-Food Sci. Technol. 2024, 192, 115630. [Google Scholar] [CrossRef]
- Shi, G.; Fang, C.; Xing, S.; Guo, Y.; Li, X.; Han, X.; Lin, L.; Zhang, C. Heterogenetic mechanism in high-temperature Daqu fermentation by traditional craft and mechanical craft: From microbial assembly patterns to metabolism phenotypes. Food Res. Int. 2024, 187, 114327. [Google Scholar] [CrossRef] [PubMed]
- Gong, L.; Qin, S.; Zheng, X.; Zhao, J.; Liu, M.; Zhao, M. Traceability between microbial community and environmental microbial Community in Maotai-flavor Daqu. Food Chem. X 2025, 27, 102321. [Google Scholar] [CrossRef]
- Tang, J.; Wang, L.; Zhao, Y.; Wu, C.; Qiao, L.; Xiao, X.; Guo, M.; Yang, L. Multi-omics analysis of the spatial heterogeneity of Moutai-flavored Daqu. Food Sci. 2025, 46, 126–134. (In Chinese) [Google Scholar] [CrossRef]
- Ropars, J.; Rodríguez de la Vega, R.C.; López-Villavicencio, M.; Gouzy, J.; Dupont, J.; Swennen, D.; Dumas, E.; Giraud, T.; Branca, A. Diversity and Mechanisms of Genomic Adaptation in Penicillium. In Aspergillus and Penicillium in the Post-Genomic Era; de Vries, R.P., Benoit Gelber, I., Andersen, M.R., Eds.; Caister Academic Press: Norfolk, UK, 2016; pp. 27–42. [Google Scholar]
- Zareb, A.; Banachewicz, P.; Havrysh, P.; Blaszczyk, L.; Hammad, T.; Meftah, C.; Salamon, S. Endophytic fungal communities of Calicotome spinosa-an important medicinal plant of Tizi-Ouzou (Algeria). J. Appl. Genet. 2025, 66, 763–769. [Google Scholar] [CrossRef]
- Germain, R.M.; Mayfield, M.M.; Gilbert, B. The ‘filtering’ metaphor revisited: Competition and environment jointly structure invasibility and coexistence. Biol. Lett. 2018, 14, 20180460. [Google Scholar] [CrossRef] [PubMed]
- De Wit, R. Does the Environment “Filter” or “Select” Species? Bridging the Ecologies of Microbes and Macro-Organisms for a Common Niche Assembly Theory. Environments 2025, 10, 350. [Google Scholar] [CrossRef]
- Caporaso, J.G.; Paszkiewicz, K.; Field, D.; Knight, R.; Gilbert, J.A. The Western English Channel contains a persistent microbial seed bank. ISME J. 2012, 6, 1089–1093. [Google Scholar] [CrossRef]
- Ashley, S.; Gregory, C.J.; Jo, H.; Rob, K.; Noah, F. A meta-analysis of changes in bacterial and archaeal communities with time. ISME J. 2013, 7, 1493–1506. [Google Scholar] [CrossRef] [PubMed]
- Jiao, S.; Chen, W.; Wang, E.; Wang, J.; Liu, Z.; Li, Y.; Wei, G. Microbial succession in response to pollutants in batch-enrichment culture. Sci. Rep. 2016, 6, 21791. [Google Scholar] [CrossRef] [PubMed]
- Waill, A.E.; Abdu, G.A.K.; Azza, E.; Ghoson, M.D. Allergen, pathogen, or biotechnological tool? The dematiaceous fungi Alternaria what’s for it and what’s on it? J. Pharm. Pharmacol. Res. 2021, 4, 1–6. [Google Scholar] [CrossRef]
- Zhao, S.; Li, J.; Liu, J.; Xiao, S.; Yang, S.; Mei, J.; Ren, M.; Wu, S.; Zhang, H.; Yang, X. Secondary metabolites of Alternaria: A comprehensive review of chemical diversity and pharmacological properties. Front. Microbiol. 2023, 13, 1085666. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Peng, Y.; Fu, L.; Lou, J. Metabolites from Alternaria Fungi and Their Bioactivities. Molecules 2013, 18, 5891–5935. [Google Scholar] [CrossRef]
- Zhao, L.; Lu, J.J.; Jiang, L.; Wang, X.Y.; Wu, F.Y.; Luo, Z.B.; Yang, L.; Wang, X.Y. Assembly, diversity and coexistence of bacteria communities in various rhizocompartment niches in Sorghum Cultivars. Rhizosphere 2023, 27, 100779. [Google Scholar] [CrossRef]
- Zhang, X.; Li, H.; Li, B.; Song, K.; Sha, Y.; Liu, Y.; Dong, S.; Wang, D.; Yang, L. Microbial Community Shifts in Tea Plant Rhizosphere under Seawater Stress: Enrichment of Beneficial Taxa. Microorganisms 2024, 12, 1287. [Google Scholar] [CrossRef]
- Wang, B.-T.; Hu, S.; Yu, X.-Y.; Jin, L.; Zhu, Y.-J.; Jin, F.-J. Studies of Cellulose and Starch Utilization and the Regulatory Mechanisms of Related Enzymes in Fungi. Polymers 2020, 12, 530. [Google Scholar] [CrossRef]
- Dong, W.; Yu, X.; Wang, L.; Zou, M.; Ma, J.; Liu, J.; Feng, Y.; Zhao, S.; Yang, Q.; Hu, Y.; et al. Unveiling the microbiota of sauce-flavor Daqu and its relationships with flavors and color during maturation. Front. Microbiol. 2024, 15, 1345772. [Google Scholar] [CrossRef]
- Yang, J.; Xue, F.; Li, D.; Chen, J.; Shi, G.; Song, G.; Li, Y. Oxygen regulation of microbial communities and chemical compounds in cigar tobacco curing. Front. Microbiol. 2024, 15, 1425553. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Wang, B.T.; Li, T.; Bu, S.; Jin, C.Z.; Jin, L.; Ruan, H.H.; Shin, K.S.; Jin, F.J. Successional patterns of microbial communities across various stages of leaf litter decomposition in poplar plantations. Front. Microbiol. 2025, 16, 1628355. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhao, L.; Liu, J.; Zhang, L.; Luo, Z.; Tang, Q.; Zhao, J.; Ji, Q.; Wang, X. Effects of Kuding Tea on the Succession and Assembly of the Fungal Community During Fermentation of Daqu. Fermentation 2026, 12, 136. https://doi.org/10.3390/fermentation12030136
Zhao L, Liu J, Zhang L, Luo Z, Tang Q, Zhao J, Ji Q, Wang X. Effects of Kuding Tea on the Succession and Assembly of the Fungal Community During Fermentation of Daqu. Fermentation. 2026; 12(3):136. https://doi.org/10.3390/fermentation12030136
Chicago/Turabian StyleZhao, Liang, Jialin Liu, Liang Zhang, Zhenbiao Luo, Qulai Tang, Jingjing Zhao, Qing Ji, and Xinye Wang. 2026. "Effects of Kuding Tea on the Succession and Assembly of the Fungal Community During Fermentation of Daqu" Fermentation 12, no. 3: 136. https://doi.org/10.3390/fermentation12030136
APA StyleZhao, L., Liu, J., Zhang, L., Luo, Z., Tang, Q., Zhao, J., Ji, Q., & Wang, X. (2026). Effects of Kuding Tea on the Succession and Assembly of the Fungal Community During Fermentation of Daqu. Fermentation, 12(3), 136. https://doi.org/10.3390/fermentation12030136



