The Mechanistic Roles of ncRNAs in Promoting and Supporting Chemoresistance of Colorectal Cancer
Abstract
1. Introduction
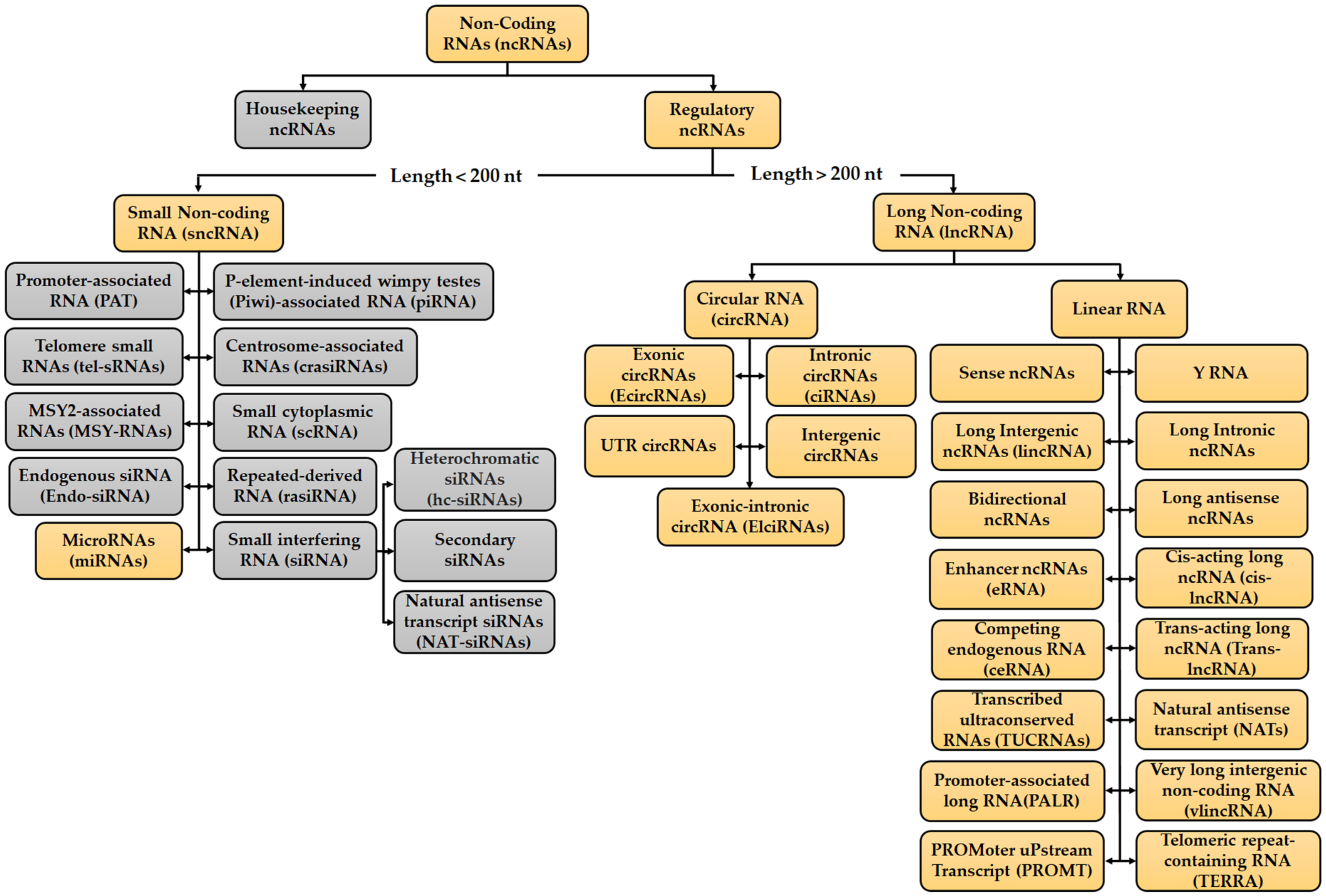
2. Overview of ncRNAs
3. Drug Resistance in CRC
3.1. ncRNAs in Chemotherapeutic Resistance
3.2. Drug Resistance in CRC Due to miRNAs, lncRNAs and circRNAs
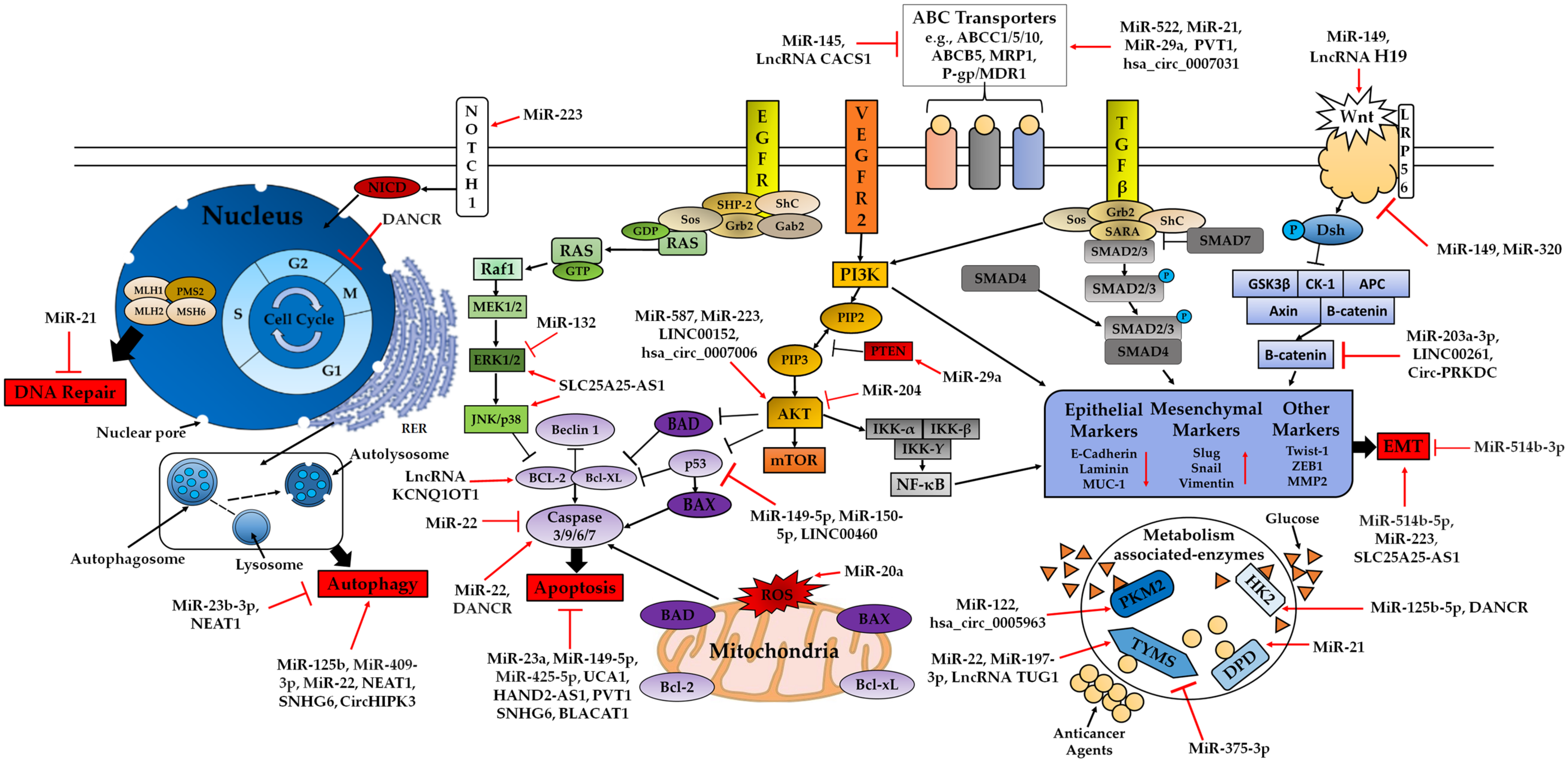
3.2.1. miRNAs, lncRNAs, circRNAs in 5-FU Resistance
ABC Transporters Family
PI3K/AKT Signalling Pathway
MAPK/ERK Signalling Pathway and EMT
DNA Repair Pathways
Apoptosis-Related Pathway
Autophagy
Hippo Signalling Pathway
Wnt/β-catenin Signalling Pathway
Other Chemoresistance Related miRNAs, lncRNAs or circRNAs
3.2.2. miRNAs, lncRNAs, circRNAs in OXA Resistance
ABC Transporters Family
PI3K/AKT Signalling Pathway
Apoptosis-Related Pathway
Autophagy Related Pathway
Wnt/β-catenin Signalling Pathway
TNF-α Pathway
Glycolysis
Other Chemoresistance Related miRNAs, lncRNAs or circRNAs
3.2.3. miRNAs, lncRNAs, circRNAs in Cisplatin Resistance
ABC Transporters Family
Apoptosis-Related Pathway
Autophagy
Wnt/β-catenin Signaling Pathway
EMT
Other Chemoresistance Related miRNAs or lncRNAs
3.2.4. miRNAs, lncRNAs, circRNAs in DOX Resistance
ABC Transporters Family
PI3K/AKT, MAPK/ERK Signalling Pathways and EMT
Apoptosis-Related Pathway
Hippo Signaling Pathway
Other Chemoresistance Related miRNAs or lncRNAs
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Imran, A.; Qamar, H.Y.; Ali, Q.; Naeem, H.; Riaz, M.; Amin, S.; Kanwal, N.; Ali, F.; Sabar, M.F.; Nasir, I.A. Role of Molecular Biology in Cancer Treatment: A Review Article. Iran. J. Public Health 2017, 46, 1475–1485. [Google Scholar]
- Senore, C.; Segnan, N.; Gunter, K. Chapter 5.5 Colorectal cancer. Decreasing disparities and promoting prevention are policy priorities. In World Cancer Report, 1st ed.; Wild, C.P., Weiderpass, E., Stewart, B.W., Eds.; International Agency for Research on Cancer: Lyon, France, 2020; pp. 344–354. [Google Scholar]
- Simon, K. Colorectal Cancer Development and Advances in Screening. Clin. Interv. Aging 2016, 11, 967–976. [Google Scholar]
- Nguyen, H.T.; Duong, H.Q. The molecular characteristics of colorectal cancer: Implications for diagnosis and therapy. Oncol. Lett. 2018, 16, 9–18. [Google Scholar] [CrossRef]
- Centelles, J.J. General aspects of colorectal cancer. Int. Sch. Res. Not. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Brenner, H.; Kloor, M.; Pox, C.P. Colorectal cancer. Lancet 2014, 383, 1490–1502. [Google Scholar] [CrossRef]
- Pan, J.J.; Xie, X.J.; Li, X.; Chen, W. Long Non-Coding RNAs and Drug Resistance. Asian Pac. J. Cancer Prev. 2015, 16, 8067–8073. [Google Scholar] [CrossRef] [PubMed]
- Gherman, A.; Balacescu, L.; Gheorghe-Cetean, S.; Vlad, C.; Balacescu, O.; Irimie, A.; Lisencu, C. Current and New Predictors for Treatment Response in Metastatic Colorectal Cancer. The Role of Circulating MiRNAs as Biomarkers. Int. J. Mol. Sci. 2020, 21, 2089. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Yan, Y.; Chen, M.; Luo, G.; Hao, J.; Pan, J.; Hu, S.; Guo, P.; Li, W.; Wang, R.; et al. Aspirin Enhances the Sensitivity of Colon Cancer Cells to Cisplatin by Abrogating the Binding of NF-ΚB to the COX-2 Promoter. Aging 2020, 12, 611–627. [Google Scholar] [CrossRef]
- Xiong, S.; Xiao, G.W. Reverting Doxorubicin Resistance in Colon Cancer by Targeting a Key Signaling Protein, Steroid Receptor Coactivator. Exp. Ther. Med. 2018, 15, 3751–3758. [Google Scholar] [CrossRef]
- Chortkoff, B.; Stenehjem, D. Chemotherapy, Immunosuppression, and Anesthesia. In Pharmacology and Physiology for Anesthesia; Hemmings, H.C., Egan, T.D., Eds.; Elsevier: Amsterdam, The Netherlands, 2019; pp. 753–768. [Google Scholar]
- Briffa, R.; Langdon, S.P.; Grech, G.; Harrison, D.J. Acquired and Intrinsic Resistance to Colorectal Cancer Treatment. In Colorectal Cancer—Diagnosis, Screening and Management; Chen, J., Ed.; InTech: London, UK, 2018. [Google Scholar]
- Blondy, S.; David, V.; Verdier, M.; Mathonnet, M.; Perraud, A.; Christou, N. 5-Fluorouracil Resistance Mechanisms in Colorectal Cancer: From Classical Pathways to Promising Processes. Cancer Sci. 2020, 111, 3142–3154. [Google Scholar] [CrossRef]
- Martinez-Balibrea, E.; Martínez-Cardús, A.; Ginés, A.; Ruiz de Porras, V.; Moutinho, C.; Layos, L.; Manzano, J.L.; Bugés, C.; Bystrup, S.; Esteller, M.; et al. Tumor-Related Molecular Mechanisms of Oxaliplatin Resistance. Mol. Cancer Ther. 2015, 14, 1767–1776. [Google Scholar] [CrossRef]
- Dasari, S.; Tchounwou, P.B. Cisplatin in Cancer Therapy: Molecular Mechanisms of Action. Eur. J. Pharmacol. 2014, 740, 364–378. [Google Scholar] [CrossRef] [PubMed]
- Micallef, I.; Baron, B. Doxorubicin: An Overview of the Anti-Cancer and Chemoresistance Mechanisms. Ann. Clin. Toxicol. 2020, 3, 1031. [Google Scholar]
- Hammond, W.A.; Swaika, A.; Mody, K. Pharmacologic Resistance in Colorectal Cancer: A Review. Ther. Adv. Med. Oncol. 2016, 8, 57–84. [Google Scholar] [CrossRef] [PubMed]
- Hu, T.; Li, Z.; Gao, C.Y.; Cho, C.H. Mechanisms of Drug Resistance in Colon Cancer and Its Therapeutic Strategies. World J. Gastroenterol. 2016, 22, 6876. [Google Scholar] [CrossRef]
- Xie, Y.H.; Chen, Y.X.; Fang, J.Y. Comprehensive Review of Targeted Therapy for Colorectal Cancer. Signal Transduct. Target. Ther. 2020, 5, 22. [Google Scholar] [CrossRef]
- Vaghari-Tabari, M.; Majidinia, M.; Moein, S.; Qujeq, D.; Asemi, Z.; Alemi, F.; Mohamadzadeh, R.; Targhazeh, N.; Safa, A.; Yousefi, B. MicroRNAs and Colorectal Cancer Chemoresistance: New Solution for Old Problem. Life Sci. 2020, 259, 118255. [Google Scholar] [CrossRef]
- Skarkova, V.; Kralova, V.; Vitovcova, B.; Rudolf, E. Selected Aspects of Chemoresistance Mechanisms in Colorectal Carcinoma-A Focus on Epithelial-to-Mesenchymal Transition, Autophagy, and Apoptosis. Cells 2019, 8, 234. [Google Scholar] [CrossRef] [PubMed]
- Corrà, F.; Agnoletto, C.; Minotti, L.; Baldassari, F.; Volinia, S. The Network of Non-Coding RNAs in Cancer Drug Resistance. Front. Oncol. 2018, 8, 327. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Wang, X.; Lv, L.; Zheng, Y.; Zhang, N.; Yang, M. The Emerging Role of Noncoding RNAs in Colorectal Cancer Chemoresistance. Cell. Oncol. 2019, 42, 757–768. [Google Scholar] [CrossRef]
- Liu, K.; Gao, L.; Ma, X.; Huang, J.J.; Chen, J.; Zeng, L.; Ashby, C.R., Jr.; Zou, C.; Chen, Z.S. Long Non-Coding RNAs Regulate Drug Resistance in Cancer. Mol. Cancer 2020, 19, 54. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Yang, J.; Li, X.; Liu, D.; Fu, L.; Wang, X. Functions and Mechanisms of Circular RNAs in Cancer Radiotherapy and Chemotherapy Resistance. Mol. Cancer 2020, 19, 58. [Google Scholar] [CrossRef]
- He, Q.; Long, J.; Yin, Y.; Li, Y.; Lei, X.; Li, Z.; Zhu, W. Emerging Roles of LncRNAs in the Formation and Progression of Colorectal Cancer. Front. Oncol. 2020, 9, 1542. [Google Scholar] [CrossRef]
- Qu, Y.; Tan, H.Y.; Chan, Y.T.; Jiang, H.; Wang, N.; Wang, D. The Functional Role of Long Noncoding RNA in Resistance to Anticancer Treatment. Ther. Adv. Med. Oncol. 2020, 12, 1758835920927850. [Google Scholar] [CrossRef] [PubMed]
- Djebali, S.; Davis, C.A.; Merkel, A.; Dobin, A.; Lassmann, T.; Mortazavi, A.; Tanzer, A.; Lagarde, J.; Lin, W.; Schlesinger, F.; et al. Landscape of Transcription in Human Cells. Nature 2012, 489, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Pavet, V.; Portal, M.M.; Moulin, J.C.; Herbrecht, R.; Gronemeyer, H. Towards Novel Paradigms for Cancer Therapy. Oncogene 2011, 30, 1–20. [Google Scholar] [CrossRef]
- Esteller, M. Non-Coding RNAs in Human Disease. Nat. Rev. Genet. 2011, 12, 861–874. [Google Scholar] [CrossRef]
- Diamantopoulos, M.A.; Tsiakanikas, P.; Scorilas, A. Non-Coding RNAs: The Riddle of the Transcriptome and Their Perspectives in Cancer. Ann. Transl. Med. 2018, 6, 241. [Google Scholar] [CrossRef]
- Geng, X.; Jia, Y.; Zhang, Y.; Shi, L.; Li, Q.; Zang, A.; Wang, H. Circular RNA: Biogenesis, Degradation, Functions and Potential Roles in Mediating Resistance to Anticarcinogens. Epigenomics 2020, 12, 267–283. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Song, X.; Glass, C.K.; Rosenfeld, M.G. The Long Arm of Long Noncoding RNAs: Roles as Sensors Regulating Gene Transcriptional Programs. Cold Spring Harb. Perspect. Biol. 2011, 3, a003756. [Google Scholar] [CrossRef]
- Rinn, J.L.; Chang, H.Y. Genome Regulation by Long Noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Froberg, J.E.; Lee, J.T. Long Noncoding RNAs: Fresh Perspectives into the RNA World. Trends Biochem. Sci. 2014, 39, 35–43. [Google Scholar] [CrossRef]
- Batista, P.J.; Chang, H.Y. Long Noncoding RNAs: Cellular Address Codes in Development and Disease. Cell 2013, 152, 1298–1307. [Google Scholar] [CrossRef]
- Fatica, A.; Bozzoni, I. Long Non-Coding RNAs: New Players in Cell Differentiation and Development. Nat. Rev. Genet. 2014, 15, 7–21. [Google Scholar] [CrossRef]
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and Their Integrated Networks. J. Integr. Bioinform. 2019, 16. [Google Scholar] [CrossRef]
- Chen, Q.; Meng, X.; Liao, Q.; Chen, M. Versatile Interactions and Bioinformatics Analysis of Noncoding RNAs. Brief. Bioinform. 2019, 20, 1781–1794. [Google Scholar] [CrossRef] [PubMed]
- Anastasiadou, E.; Faggioni, A.; Trivedi, P.; Slack, F. The Nefarious Nexus of Noncoding RNAs in Cancer. Int. J. Mol. Sci. 2018, 19, 2072. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, J.M. Revisiting LncRNAs: How Do You Know Yours Is Not an ERNA? Mol. Cell 2016, 62, 1–2. [Google Scholar] [CrossRef] [PubMed]
- Cech, T.R.; Steitz, J.A. The Noncoding RNA Revolution-Trashing Old Rules to Forge New Ones. Cell 2014, 157, 77–94. [Google Scholar] [CrossRef] [PubMed]
- Peschansky, V.J.; Wahlestedt, C. Non-Coding RNAs as Direct and Indirect Modulators of Epigenetic Regulation. Epigenetics 2014, 9, 3–12. [Google Scholar] [CrossRef]
- Ponjavic, J.; Ponting, C.P.; Lunter, G. Functionality or Transcriptional Noise? Evidence for Selection within Long Noncoding RNAs. Genome Res. 2007, 17, 556–565. [Google Scholar] [CrossRef] [PubMed]
- Klinge, C.M. Non-Coding RNAs in Breast Cancer: Intracellular and Intercellular Communication. Noncoding RNA 2018, 4, 40. [Google Scholar] [CrossRef] [PubMed]
- Hombach, S.; Kretz, M. Non-Coding RNAs: Classification, Biology and Functioning. Adv. Exp. Med. Biol. 2016, 937, 3–17. [Google Scholar] [PubMed]
- Fu, X.D. Non-Coding RNA: A New Frontier in Regulatory Biology. Natl. Sci. Rev. 2014, 1, 190–204. [Google Scholar] [CrossRef]
- Kowalski, M.P.; Krude, T. Functional Roles of Non-Coding Y RNAs. Int. J. Biochem. Cell Biol. 2015, 66, 20–29. [Google Scholar] [CrossRef]
- Taft, R.J.; Pang, K.C.; Mercer, T.R.; Dinger, M.; Mattick, J.S. Non-Coding RNAs: Regulators of Disease. J. Pathol. 2010, 220, 126–139. [Google Scholar] [CrossRef] [PubMed]
- Piatek, M.J.; Werner, A. Endogenous SiRNAs: Regulators of Internal Affairs. Biochem. Soc. Trans. 2014, 42, 1174–1179. [Google Scholar] [CrossRef] [PubMed]
- Han Li, C.; Chen, Y. Small and Long Non-Coding RNAs: Novel Targets in Perspective Cancer Therapy. Curr. Genom. 2015, 16, 319–326. [Google Scholar]
- Kaikkonen, M.U.; Lam, M.T.Y.; Glass, C.K. Non-Coding RNAs as Regulators of Gene Expression and Epigenetics. Cardiovasc. Res. 2011, 90, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Sana, J.; Faltejskova, P.; Svoboda, M.; Slaby, O. Novel classes of non-coding RNAs and cancer. J. Transl. Med. 2012, 10, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Mewalal, R.; Hu, R.; Tuskan, G.A.; Yang, X. New technologies accelerate the exploration of non-coding RNAs in horticultural plants. Hortic. Res. 2017, 4, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Dahariya, S.; Paddibhatla, I.; Kumar, S.; Raghuwanshi, S.; Pallepati, A.; Gutti, R.K. Long non-coding RNA: Classification, biogenesis and functions in blood cells. Mol. Immunol. 2019, 112, 82–92. [Google Scholar] [CrossRef] [PubMed]
- Srijyothi, L.; Ponne, S.; Prathama, T.; Ashok, C.; Baluchamy, S. Roles of non-coding RNAs in transcriptional regulation. In Transcriptional and Post-Transcriptional Regulation, 1st ed.; Ghedira, K., Ed.; IntechOpen: London, UK, 2018. [Google Scholar]
- Laurent, G.S.; Wahlestedt, C.; Kapranov, P. The Landscape of long noncoding RNA classification. Trends Genet. 2015, 31, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Amin, N.; McGrath, A.; Chen, Y.P.P. Evaluation of deep learning in non-coding RNA classification. Nat. Mach. Intell. 2019, 1, 246–256. [Google Scholar]
- Anastasiadou, E.; Jacob, L.S.; Slack, F.J. Non-Coding RNA Networks in Cancer. Nat. Rev. Cancer 2018, 18, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Wahid, F.; Shehzad, A.; Khan, T.; Kim, Y.Y. MicroRNAs: Synthesis, Mechanism, Function, and Recent Clinical Trials. Biochim. Biophys. Acta 2010, 1803, 1231–1243. [Google Scholar] [CrossRef] [PubMed]
- Yamamura, S.; Imai-Sumida, M.; Tanaka, Y.; Dahiya, R. Interaction and Cross-Talk between Non-Coding RNAs. Cell. Mol. Life Sci. 2018, 75, 467–484. [Google Scholar] [CrossRef] [PubMed]
- Di Leva, G.; Garofalo, M.; Croce, C.M. MicroRNAs in Cancer. Annu. Rev. Pathol. 2014, 9, 287–314. [Google Scholar] [CrossRef]
- Pedroza-Torres, A.; Romero-Córdoba, S.L.; Justo-Garrido, M.; Salido-Guadarrama, I.; Rodríguez-Bautista, R.; Montaño, S.; Muñiz-Mendoza, R.; Arriaga-Canon, C.; Fragoso-Ontiveros, V.; Álvarez-Gómez, R.M.; et al. MicroRNAs in Tumor Cell Metabolism: Roles and Therapeutic Opportunities. Front. Oncol. 2019, 9, 1404. [Google Scholar] [CrossRef]
- Giovannetti, E.; Erozenci, A.; Smit, J.; Danesi, R.; Peters, G.J. Molecular Mechanisms Underlying the Role of MicroRNAs (MiRNAs) in Anticancer Drug Resistance and Implications for Clinical Practice. Crit. Rev. Oncol. Hematol. 2012, 81, 103–122. [Google Scholar] [CrossRef]
- Geisler, S.; Coller, J. RNA in Unexpected Places: Long Non-Coding RNA Functions in Diverse Cellular Contexts. Nat. Rev. Mol. Cell Biol. 2013, 14, 699–712. [Google Scholar] [CrossRef]
- Wei, M.M.; Zhou, G.B. Long Non-Coding RNAs and Their Roles in Non-Small-Cell Lung Cancer. Genom. Proteom. Bioinform. 2016, 14, 280–288. [Google Scholar] [CrossRef]
- Rutenberg-Schoenberg, M.; Sexton, A.N.; Simon, M.D. The Properties of Long Noncoding RNAs That Regulate Chromatin. Annu. Rev. Genomics Hum. Genet. 2016, 17, 69–94. [Google Scholar] [CrossRef]
- Balas, M.M.; Johnson, A.M. Exploring the Mechanisms behind Long Noncoding RNAs and Cancer. Noncoding RNA Res. 2018, 3, 108–117. [Google Scholar] [CrossRef]
- Li, Z.; Zhao, W.; Wang, M.; Zhou, X. The Role of Long Noncoding RNAs in Gene Expression Regulation. In Gene Expression Profiling in Cancer; Vlachakis, D., Ed.; IntechOpen: London, UK, 2019. [Google Scholar]
- Bermúdez, M.; Aguilar-Medina, M.; Lizárraga-Verdugo, E.; Avendaño-Félix, M.; Silva-Benítez, E.; López-Camarillo, C.; Ramos-Payán, R. LncRNAs as Regulators of Autophagy and Drug Resistance in Colorectal Cancer. Front. Oncol. 2019, 9, 1008. [Google Scholar] [CrossRef]
- Bonasio, R.; Shiekhattar, R. Regulation of Transcription by Long Noncoding RNAs. Annu. Rev. Genet. 2014, 48, 433–455. [Google Scholar] [CrossRef]
- Wang, C.; Wang, L.; Ding, Y.; Lu, X.; Zhang, G.; Yang, J.; Zheng, H.; Wang, H.; Jiang, Y.; Xu, L. LncRNA Structural Characteristics in Epigenetic Regulation. Int. J. Mol. Sci. 2017, 18, 2659. [Google Scholar] [CrossRef]
- Ginn, L.; Shi, L.; Montagna, M.L.; Garofalo, M. LncRNAs in Non-Small-Cell Lung Cancer. Noncoding RNA 2020, 6, 25. [Google Scholar]
- Noh, J.H.; Kim, K.M.; McClusky, W.G.; Abdelmohsen, K.; Gorospe, M. Cytoplasmic Functions of Long Noncoding RNAs. Wiley Interdiscip. Rev. RNA 2018, 9, e1471. [Google Scholar] [CrossRef]
- Kung, J.T.Y.; Colognori, D.; Lee, J.T. Long Noncoding RNAs: Past, Present, and Future. Genetics 2013, 193, 651–669. [Google Scholar] [CrossRef]
- Yoon, J.H.; Abdelmohsen, K.; Gorospe, M. Posttranscriptional Gene Regulation by Long Noncoding RNA. J. Mol. Biol. 2013, 425, 3723–3730. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Feng, J.; Lei, L.; Hu, J.; Xia, L.; Wang, J.; Xiang, Y.; Liu, L.; Zhong, S.; Han, L.; et al. Comprehensive Characterization of Tissue-Specific Circular RNAs in the Human and Mouse Genomes. Brief. Bioinform. 2017, 18, 984–992. [Google Scholar] [CrossRef]
- Su, M.; Xiao, Y.; Ma, J.; Tang, Y.; Tian, B.; Zhang, Y.; Li, X.; Wu, Z.; Yang, D.; Zhou, Y.; et al. Circular RNAs in Cancer: Emerging Functions in Hallmarks, Stemness, Resistance and Roles as Potential Biomarkers. Mol. Cancer 2019, 18, 90. [Google Scholar] [CrossRef]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs Are a Large Class of Animal RNAs with Regulatory Potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef]
- Noto, J.J.; Schmidt, C.A.; Matera, A.G. Engineering and Expressing Circular RNAs via TRNA Splicing. RNA Biol. 2017, 14, 978–984. [Google Scholar] [CrossRef]
- Chen, X.; Fan, S.; Song, E. Noncoding RNAs: New Players in Cancers. Adv. Exp. Med. Biol. 2016, 927, 1–47. [Google Scholar]
- Huang, A.; Zheng, H.; Wu, Z.; Chen, M.; Huang, Y. Circular RNA-Protein Interactions: Functions, Mechanisms, and Identification. Theranostics 2020, 10, 3503–3517. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Qi, X.; Liu, L.; Hu, X.; Liu, J.; Yang, J.; Yang, J.; Lu, L.; Zhang, Z.; Ma, S.; et al. Emerging Epigenetic Regulation of Circular RNAs in Human Cancer. Mol. Ther. Nucleic Acids 2019, 16, 589–596. [Google Scholar] [CrossRef]
- Salzman, J. Circular RNA Expression: Its Potential Regulation and Function. Trends Genet. 2016, 32, 309–316. [Google Scholar] [CrossRef] [PubMed]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. CircRNA Biogenesis Competes with Pre-MRNA Splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, A.M.; Chang, H.Y. Long Noncoding RNAs in Cancer Pathways. Cancer Cell 2016, 29, 452–463. [Google Scholar] [CrossRef]
- Panda, A.C. Circular RNAs Act as MiRNA Sponges. Adv. Exp. Med. Biol. 2018, 1087, 67–79. [Google Scholar] [PubMed]
- Ma, D.; Zhao, Y.; Yu, S.; Zhang, H.; Cheng, M.; Cao, H.; Li, Q.; Min, L. CircRNA as CeRNA Mediated by MicroRNA May Be Involved in Goat Lactation. Small Rumin. Res. 2019, 171, 63–72. [Google Scholar] [CrossRef]
- Sen, R.; Ghosal, S.; Das, S.; Balti, S.; Chakrabarti, J. Competing endogenous RNA: The key to posttranscriptional regulation. Sci. World J. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Fabbri, M.; Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 2013, 12, 847–865. [Google Scholar] [CrossRef]
- Lekka, E.; Hall, J. Noncoding RNAs in Disease. FEBS Lett. 2018, 592, 2884–2900. [Google Scholar] [CrossRef] [PubMed]
- Gutschner, T.; Diederichs, S. The Hallmarks of Cancer: A Long Non-Coding RNA Point of View: A Long Non-Coding RNA Point of View. RNA Biol. 2012, 9, 703–719. [Google Scholar] [CrossRef]
- Zhang, X.; Xie, K.; Zhou, H.; Wu, Y.; Li, C.; Liu, Y.; Liu, Z.; Xu, Q.; Liu, S.; Xiao, D.; et al. Role of Non-Coding RNAs and RNA Modifiers in Cancer Therapy Resistance. Mol. Cancer 2020, 19, 47. [Google Scholar] [CrossRef] [PubMed]
- Agostini, M.; Ganini, C.; Candi, E.; Melino, G. The Role of Noncoding RNAs in Epithelial Cancer. Cell Death Discov. 2020, 6, 13. [Google Scholar] [CrossRef]
- Strubberg, A.M.; Madison, B.B. MicroRNAs in the Etiology of Colorectal Cancer: Pathways and Clinical Implications. Dis. Model. Mech. 2017, 10, 197–214. [Google Scholar] [CrossRef]
- Ibrahim, H.; Lim, Y.C. KRAS-Associated MicroRNAs in Colorectal Cancer. Oncol. Rev. 2020, 14, 454. [Google Scholar] [CrossRef]
- Saberinia, A.; Alinezhad, A.; Jafari, F.; Soltany, S.; Akhavan Sigari, R. Oncogenic MiRNAs and Target Therapies in Colorectal Cancer. Clin. Chim. Acta 2020, 508, 77–91. [Google Scholar] [CrossRef] [PubMed]
- Ahadi, A. The Significance of MicroRNA Deregulation in Colorectal Cancer Development and the Clinical Uses as a Diagnostic and Prognostic Biomarker and Therapeutic Agent. Noncoding RNA Res. 2020, 5, 125–134. [Google Scholar] [CrossRef]
- Thomas, J.; Ohtsuka, M.; Pichler, M.; Ling, H. MicroRNAs: Clinical Relevance in Colorectal Cancer. Int. J. Mol. Sci. 2015, 16, 28063–28076. [Google Scholar] [CrossRef]
- Jiang, D.; Xie, X.; Lu, Z.; Liu, L.; Qu, Y.; Wu, S.; Li, Y.; Li, G.; Wang, H.; Xu, G. Establishment of a Colorectal Cancer-Related MicroRNA-MRNA Regulatory Network by Microarray and Bioinformatics. Front. Genet. 2020, 11, 560186. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Duanmu, J.; Fu, X.; Li, T.; Jiang, Q. Identifying a New MicroRNA Signature as a Prognostic Biomarker in Colon Cancer. PLoS ONE 2020, 15, e0228575. [Google Scholar] [CrossRef]
- Xiao, Z.; Chen, S.; Feng, S.; Li, Y.; Zou, J.; Ling, H.; Zeng, Y.; Zeng, X. Function and Mechanisms of MicroRNA-20a in Colorectal Cancer. Exp. Ther. Med. 2020, 19, 1605–1616. [Google Scholar] [CrossRef]
- Garajová, I.; Ferracin, M.; Porcellini, E.; Palloni, A.; Abbati, F.; Biasco, G.; Brandi, G. Non-Coding RNAs as Predictive Biomarkers to Current Treatment in Metastatic Colorectal Cancer. Int. J. Mol. Sci. 2017, 18, 1547. [Google Scholar] [CrossRef] [PubMed]
- Neve, B.; Jonckheere, N.; Vincent, A.; Van Seuningen, I. Epigenetic Regulation by LncRNAs: An Overview Focused on UCA1 in Colorectal Cancer. Cancers 2018, 10, 440. [Google Scholar] [CrossRef] [PubMed]
- Bolha, L.; Ravnik-Glavač, M.; Glavač, D. Long Noncoding RNAs as Biomarkers in Cancer. Dis. Markers 2017, 2017, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Tang, B.; Xiao, Y.F.; Xie, R.; Li, B.S.; Dong, H.; Zhou, J.Y.; Yang, S.M. Long Non-Coding RNAs in Colorectal Cancer. Oncotarget 2016, 7, 5226–5239. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Qu, J.; Wu, D.K.; Lu, Z.L.; Sun, Y.S.; Qu, Q. Long Non-Coding RNAs: A Rising Biotarget in Colorectal Cancer. Oncotarget 2017, 8, 22187–22202. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Song, Y.X.; Ma, B.; Wang, J.J.; Sun, J.X.; Chen, X.W.; Zhao, J.H.; Yang, Y.C.; Wang, Z.N. Regulatory Roles of Non-Coding RNAs in Colorectal Cancer. Int. J. Mol. Sci. 2015, 16, 19886–19919. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.D.; Qi, P.; Du, X. Long Non-Coding RNAs in Colorectal Cancer: Implications for Pathogenesis and Clinical Application. Mod. Pathol. 2014, 27, 1310–1320. [Google Scholar] [CrossRef]
- Poursheikhani, A.; Abbaszadegan, M.R.; Kerachian, M.A. Mechanisms of Long Non-Coding RNA Function in Colorectal Cancer Tumorigenesis. Asia Pac. J. Clin. Oncol. 2020. [Google Scholar] [CrossRef]
- Bachmayr-Heyda, A.; Reiner, A.T.; Auer, K.; Sukhbaatar, N.; Aust, S.; Bachleitner-Hofmann, T.; Mesteri, I.; Grunt, T.W.; Zeillinger, R.; Pils, D. Correlation of Circular RNA Abundance with Proliferation—Exemplified with Colorectal and Ovarian Cancer, Idiopathic Lung Fibrosis, and Normal Human Tissues. Sci. Rep. 2015, 5, 8057. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhang, L.; Su, Y.; Zhang, X. Screening Potential Biomarkers for Colorectal Cancer Based on Circular RNA Chips. Oncol. Rep. 2018, 39, 2499–2512. [Google Scholar] [CrossRef]
- Jiang, W.; Zhang, X.; Chu, Q.; Lu, S.; Zhou, L.; Lu, X.; Liu, C.; Mao, L.; Ye, C.; Timko, M.P.; et al. The Circular RNA Profiles of Colorectal Tumor Metastatic Cells. Front. Genet. 2018, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Mármol, I.; Sánchez-de-Diego, C.; Pradilla Dieste, A.; Cerrada, E.; Rodriguez Yoldi, M.J. Colorectal Carcinoma: A General Overview and Future Perspectives in Colorectal Cancer. Int. J. Mol. Sci. 2017, 18, 197. [Google Scholar] [CrossRef]
- Svoronos, A.A.; Engelman, D.M.; Slack, F.J. OncomiR or Tumor Suppressor? The Duplicity of MicroRNAs in Cancer. Cancer Res. 2016, 76, 3666–3670. [Google Scholar] [CrossRef] [PubMed]
- Si, W.; Shen, J.; Zheng, H.; Fan, W. The Role and Mechanisms of Action of MicroRNAs in Cancer Drug Resistance. Clin. Epigenet. 2019, 11, 25. [Google Scholar] [CrossRef]
- Iorio, M.V.; Croce, C.M. MicroRNAs in Cancer: Small Molecules with a Huge Impact. J. Clin. Oncol. 2009, 27, 5848–5856. [Google Scholar] [CrossRef] [PubMed]
- Gaur, A.; Jewell, D.A.; Liang, Y.; Ridzon, D.; Moore, J.H.; Chen, C.; Ambros, V.R.; Israel, M.A. Characterization of MicroRNA Expression Levels and Their Biological Correlates in Human Cancer Cell Lines. Cancer Res. 2007, 67, 2456–2468. [Google Scholar] [CrossRef]
- Calin, G.A.; Croce, C.M. MicroRNA Signatures in Human Cancers. Nat. Rev. Cancer 2006, 6, 857–866. [Google Scholar] [CrossRef] [PubMed]
- Bhan, A.; Soleimani, M.; Mandal, S.S. Long Noncoding RNA and Cancer: A New Paradigm. Cancer Res. 2017, 77, 3965–3981. [Google Scholar] [CrossRef] [PubMed]
- Nie, L.; Wu, H.J.; Hsu, J.M.; Chang, S.S.; Labaff, A.M.; Li, C.W.; Wang, Y.; Hsu, J.L.; Hung, M.C. Long Non-Coding RNAs: Versatile Master Regulators of Gene Expression and Crucial Players in Cancer. Am. J. Transl. Res. 2012, 4, 127–150. [Google Scholar] [PubMed]
- Artemaki, P.I.; Scorilas, A.; Kontos, C.K. Circular RNAs: A New Piece in the Colorectal Cancer Puzzle. Cancers 2020, 12, 2464. [Google Scholar] [CrossRef]
- Ahmed, F.E.; Ahmed, N.C.; Vos, P.W.; Bonnerup, C.; Atkins, J.N.; Casey, M.; Nuovo, G.J.; Naziri, W.; Wiley, J.E.; Mota, H.; et al. Diagnostic MicroRNA Markers to Screen for Sporadic Human Colon Cancer in Stool: I. Proof of Principle. Cancer Genom. Proteom. 2013, 10, 93–113. [Google Scholar]
- Goswami, C.P.; Nakshatri, H. PROGmiR: A Tool for Identifying Prognostic MiRNA Biomarkers in Multiple Cancers Using Publicly Available Data. J. Clin. Bioinforma. 2012, 2, 23. [Google Scholar] [CrossRef] [PubMed]
- Ayers, D.; Vandesompele, J. Influence of MicroRNAs and Long Non-Coding RNAs in Cancer Chemoresistance. Genes 2017, 8, 95. [Google Scholar] [CrossRef] [PubMed]
- Dai, F.; Dai, L.; Zheng, X.; Guo, Y.; Zhang, Y.; Niu, M.; Lu, Y.; Li, H.; Hou, R.; Zhang, Y.; et al. Non-Coding RNAs in Drug Resistance of Head and Neck Cancers: A Review. Biomed. Pharmacother. 2020, 127, 110231. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Zhang, L.; Li, Z.; Li, Y.; Huang, C.; Lu, X. MicroRNAs in DNA Damage Response, Carcinogenesis, and Chemoresistance. Int. Rev. Cell Mol. Biol. 2017, 333, 1–49. [Google Scholar]
- Liu, H.; Lei, C.; He, Q.; Pan, Z.; Xiao, D.; Tao, Y. Nuclear Functions of Mammalian MicroRNAs in Gene Regulation, Immunity and Cancer. Mol. Cancer 2018, 17, 1–14. [Google Scholar] [CrossRef]
- Wu, G.; Wilson, G.; George, J.; Liddle, C.; Hebbard, L.; Qiao, L. Overcoming Treatment Resistance in Cancer: Current Understanding and Tactics. Cancer Lett. 2017, 387, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Haider, T.; Pandey, V.; Banjare, N.; Gupta, P.N.; Soni, V. Drug Resistance in Cancer: Mechanisms and Tackling Strategies. Pharmacol. Rep. 2020, 72, 1125–1151. [Google Scholar] [CrossRef] [PubMed]
- Housman, G.; Byler, S.; Heerboth, S.; Lapinska, K.; Longacre, M.; Snyder, N.; Sarkar, S. Drug Resistance in Cancer: An Overview. Cancers 2014, 6, 1769–1792. [Google Scholar] [CrossRef]
- Zahreddine, H.; Borden, K.L.B. Mechanisms and Insights into Drug Resistance in Cancer. Front. Pharmacol. 2013, 4, 28. [Google Scholar] [CrossRef]
- Mansoori, B.; Mohammadi, A.; Davudian, S.; Shirjang, S.; Baradaran, B. The Different Mechanisms of Cancer Drug Resistance: A Brief Review. Adv. Pharm. Bull. 2017, 7, 339–348. [Google Scholar] [CrossRef] [PubMed]
- Wilkens, S. Structure and mechanism of ABC transporters. F1000prime Rep. 2015, 7. [Google Scholar] [CrossRef] [PubMed]
- Kathawala, R.J.; Gupta, P.; Ashby, C.R., Jr.; Chen, Z.S. The modulation of ABC transporter-mediated multidrug resistance in cancer: A review of the past decade. Drug Resist. Updates 2015, 18, 1–17. [Google Scholar] [CrossRef]
- Wang, T.; Chen, Z.; Zhu, Y.; Pan, Q.; Liu, Y.; Qi, X.; Jin, L.; Jin, J.; Ma, X.; Hua, D. Inhibition of transient receptor potential channel 5 reverses 5-Fluorouracil resistance in human colorectal cancer cells. J. Biol. Chem. 2015, 290, 448–456. [Google Scholar] [CrossRef] [PubMed]
- Herraez, E.; Gonzalez-Sanchez, E.; Vaquero, J.; Romero, M.R.; Serrano, M.A.; Marin, J.J.; Briz, O. Cisplatin-induced chemoresistance in colon cancer cells involves FXR-dependent and FXR-independent up-regulation of ABC proteins. Mol. Pharm. 2012, 9, 2565–2576. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Qiu, M.; Tang, Q.; Liu, M.; Lang, N.; Bi, F. Establishment and biological characteristics of oxaliplatin-resistant human colon cancer cell lines. Chin. J. Cancer 2010, 29, 661–667. [Google Scholar] [CrossRef]
- Fulda, S. Tumor resistance to apoptosis. Int. J. Cancer 2009, 124, 511–515. [Google Scholar] [CrossRef]
- Hu, T.; Wang, L.; Zhang, L.; Lu, L.; Shen, J.; Chan, R.L.; Li, M.; Wu, W.K.; To, K.K.; Cho, C.H. Sensitivity of apoptosis-resistant colon cancer cells to tanshinones is mediated by autophagic cell death and p53-independent cytotoxicity. Phytomedicine 2015, 22, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhao, H.; Zhou, X.; Song, L. Inhibition of lactate dehydrogenase A by microRNA-34a resensitizes colon cancer cells to 5-fluorouracil. Mol. Med. Rep. 2015, 11, 577–582. [Google Scholar] [CrossRef]
- Yuan, S.; Tao, F.; Zhang, X.; Zhang, Y.; Sun, X.; Wu, D. Role of Wnt/β-catenin signaling in the chemoresistance modulation of colorectal cancer. BioMed Res. Int. 2020, 2020. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Wang, W.; Cai, H.; Du, B.; Zhang, L.; Ma, W.; Hu, Y.; Feng, S.; Miao, G. MACC1 facilitates chemoresistance and cancer stem cell-like properties of colon cancer cells through the PI3K/AKT signaling pathway. Mol. Med. Rep. 2017, 16, 8747–8754. [Google Scholar] [CrossRef]
- Koveitypour, Z.; Panahi, F.; Vakilian, M.; Peymani, M.; Forootan, F.S.; Esfahani, M.H.N.; Ghaedi, K. Signaling pathways involved in colorectal cancer progression. Cell Biosci. 2019, 9, 1–14. [Google Scholar] [CrossRef]
- Soleimani, A.; Rahmani, F.; Ferns, G.A.; Ryzhikov, M.; Avan, A.; Hassanian, S.M. Role of the NF-κB signaling pathway in the pathogenesis of colorectal cancer. Gene 2019, 726, 144132. [Google Scholar] [CrossRef] [PubMed]
- Suman, S.; Das, T.P.; Ankem, M.K.; Damodaran, C. Targeting Notch signaling in colorectal cancer. Curr. Colorectal Cancer Rep. 2014, 10, 411–416. [Google Scholar] [CrossRef] [PubMed]
- Duan, L.; Yang, W.; Feng, W.; Cao, L.; Wang, X.; Niu, L.; Li, Y.; Zhou, W.; Zhang, Y.; Liu, J.; et al. Molecular Mechanisms and Clinical Implications of MiRNAs in Drug Resistance of Colorectal Cancer. Ther. Adv. Med. Oncol. 2020, 12, 175883592094734. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, L.; Ni, Z.; Sun, J.; Gao, H.; Cheng, Z.; Xu, J.; Yin, P. Resveratrol induces AMPK-dependent MDR1 inhibition in colorectal cancer HCT116/L-OHP cells by preventing activation of NF-κB signaling and suppressing cAMP-responsive element transcriptional activity. Tumor Biol. 2015, 36, 9499–9510. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.J.; Kang, G.J.; Kang, J.I.; Boo, H.J.; Hyun, J.W.; Koh, Y.S.; Chang, W.Y.; Kim, Y.R.; Kwon, J.M.; Maeng, Y.H.; et al. Over-activation of AKT signaling leading to 5-Fluorouracil resistance in SNU-C5/5-FU cells. Oncotarget 2018, 9, 19911–19928. [Google Scholar] [CrossRef]
- Kukcinaviciute, E.; Jonusiene, V.; Sasnauskiene, A.; Dabkeviciene, D.; Eidenaite, E.; Laurinavicius, A. Significance of Notch and Wnt signaling for chemoresistance of colorectal cancer cells HCT116. J. Cell. Biochem. 2018, 119, 5913–5920. [Google Scholar] [CrossRef]
- Almendro, V.; Ametller, E.; García-Recio, S.; Collazo, O.; Casas, I.; Augé, J.M.; Maurel, J.; Gascón, P. The role of MMP7 and its cross-talk with the FAS/FASL system during the acquisition of chemoresistance to oxaliplatin. PLoS ONE 2009, 4, e4728. [Google Scholar] [CrossRef]
- Suman, S.; Kurisetty, V.; Das, T.P.; Vadodkar, A.; Ramos, G.; Lakshmanaswamy, R.; Damodaran, C. Activation of AKT signaling promotes epithelial–mesenchymal transition and tumor growth in colorectal cancer cells. Mol. Carcinog. 2014, 53, E151–E160. [Google Scholar] [CrossRef]
- Li, J.; Liu, H.; Yu, J.; Yu, H. Chemoresistance to doxorubicin induces epithelial-mesenchymal transition via upregulation of transforming growth factor β signaling in HCT116 colon cancer cells. Mol. Med. Rep. 2015, 12, 192–198. [Google Scholar] [CrossRef]
- Kim, A.Y.; Kwak, J.H.; Je, N.K.; Lee, Y.H.; Jung, Y.S. Epithelial-mesenchymal transition is associated with acquired resistance to 5-fluorocuracil in HT-29 colon cancer cells. Toxicol. Res. 2015, 31, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.M.; Lv, J.F.; Gong, L.; Yu, L.Y.; Chen, X.P.; Zhou, H.H.; Fan, L. Role of deficient mismatch repair in the personalized management of colorectal cancer. Int. J. Environ. Res. Public Health 2016, 13, 892. [Google Scholar] [CrossRef] [PubMed]
- Arnold, C.N.; Goel, A.; Boland, C.R. Role of hMLH1 promoter hypermethylation in drug resistance to 5-fluorouracil in colorectal cancer cell lines. Int. J. Cancer 2003, 106, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Yao, X.; Wang, Y.; Hu, F.; Wang, F.; Jiang, L.; Liu, Y.; Wang, D.; Sun, G.; Zhao, Y. MLH1 promoter methylation frequency in colorectal cancer patients and related clinicopathological and molecular features. PLoS ONE 2013, 8, e59064. [Google Scholar] [CrossRef] [PubMed]
- Tajima, A.; Hess, M.T.; Cabrera, B.L.; Kolodner, R.D.; Carethers, J.M. The mismatch repair complex hMutSα recognizes 5-fluorouracil-modified DNA: Implications for chemosensitivity and resistance. Gastroenterology 2004, 127, 1678–1684. [Google Scholar] [CrossRef] [PubMed]
- Martin, L.P.; Hamilton, T.C.; Schilder, R.J. Platinum resistance: The role of DNA repair pathways. Clin. Cancer Res. 2008, 14, 1291–1295. [Google Scholar] [CrossRef]
- Fedier, A.; Schwarz, V.A.; Walt, H.; Carpini, R.D.; Haller, U.; Fink, D. Resistance to topoisomerase poisons due to loss of DNA mismatch repair. Int. J. Cancer 2001, 93, 571–576. [Google Scholar] [CrossRef]
- Ahmad, S. Platinum–DNA interactions and subsequent cellular processes controlling sensitivity to anticancer platinum complexes. Chem. Biodivers. 2010, 7, 543–566. [Google Scholar] [CrossRef] [PubMed]
- Cassidy, J. Review of oxaliplatin: An active platinum agent in colorectal cancer. Int. J. Clin. Pract. 2000, 54, 399–402. [Google Scholar]
- Bohanes, P.; LaBonte, M.J.; Lenz, H.J. A review of excision repair cross-complementation group 1 in colorectal cancer. Clin. Colorectal Cancer 2011, 10, 157–164. [Google Scholar] [CrossRef]
- Wigle, T.J.; Tsvetkova, E.V.; Welch, S.A.; Kim, R.B. DPYD and fluorouracil-based chemotherapy: Mini review and case report. Pharmaceutics 2019, 11, 199. [Google Scholar] [CrossRef]
- Muhale, F.A.; Wetmore, B.A.; Thomas, R.S.; McLeod, H.L. Systems pharmacology assessment of the 5-fluorouracil pathway. Pharmacogenomics 2011, 12, 341–350. [Google Scholar] [CrossRef]
- Chen, Q.N.; Wei, C.C.; Wang, Z.X.; Sun, M. Long Non-Coding RNAs in Anti-Cancer Drug Resistance. Oncotarget 2017, 8, 1925–1936. [Google Scholar] [CrossRef]
- Majidinia, M.; Yousefi, B. Long Non-Coding RNAs in Cancer Drug Resistance Development. DNA Repair 2016, 45, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Rynkeviciene, R.; Simiene, J.; Strainiene, E.; Stankevicius, V.; Usinskiene, J.; Miseikyte Kaubriene, E.; Meskinyte, I.; Cicenas, J.; Suziedelis, K. Non-Coding RNAs in Glioma. Cancers 2018, 11, 17. [Google Scholar] [CrossRef] [PubMed]
- Wei, L.; Sun, J.; Zhang, N.; Zheng, Y.; Wang, X.; Lv, L.; Liu, J.; Xu, Y.; Shen, Y.; Yang, M. Noncoding RNAs in Gastric Cancer: Implications for Drug Resistance. Mol. Cancer 2020, 19, 62. [Google Scholar] [CrossRef]
- Luo, Y.J.; Huang, Q.M.; Ren, Y.; Liu, Z.L.; Xu, C.F.; Wang, H.; Xiao, J.W. Non-Coding RNA in Drug Resistance of Gastric Cancer. World J. Gastrointest. Oncol. 2019, 11, 957–970. [Google Scholar] [CrossRef] [PubMed]
- Lin, Z.; Lu, S.; Xie, X.; Yi, X.; Huang, H. Noncoding RNAs in Drug-Resistant Pancreatic Cancer: A Review. Biomed. Pharmacother. 2020, 131, 110768. [Google Scholar] [CrossRef]
- Lv, Y.; Huang, S. Role of Non-Coding RNA in Pancreatic Cancer. Oncol. Lett. 2019, 18, 3963–3973. [Google Scholar] [CrossRef] [PubMed]
- Du, T.; Shi, Y.; Xu, S.; Wan, X.; Sun, H.; Liu, B. Long Non-Coding RNAs in Drug Resistance of Breast Cancer. Onco. Targets. Ther. 2020, 13, 7075–7087. [Google Scholar] [CrossRef] [PubMed]
- Crudele, F.; Bianchi, N.; Reali, E.; Galasso, M.; Agnoletto, C.; Volinia, S. The Network of Non-Coding RNAs and Their Molecular Targets in Breast Cancer. Mol. Cancer 2020, 19, 61. [Google Scholar] [CrossRef] [PubMed]
- Latowska, J.; Grabowska, A.; Zarębska, Ż.; Kuczyński, K.; Kuczyńska, B.; Rolle, K. Non-Coding RNAs in Brain Tumors, the Contribution of LncRNAs, CircRNAs, and SnoRNAs to Cancer Development-Their Diagnostic and Therapeutic Potential. Int. J. Mol. Sci. 2020, 21, 7001. [Google Scholar] [CrossRef] [PubMed]
- Zeng, T.; Li, L.; Zhou, Y.; Gao, L. Exploring Long Noncoding RNAs in Glioblastoma: Regulatory Mechanisms and Clinical Potentials. Int. J. Genom. 2018, 2018, 1–13. [Google Scholar] [CrossRef]
- Ding, B.; Lou, W.; Xu, L.; Fan, W. Non-Coding RNA in Drug Resistance of Hepatocellular Carcinoma. Biosci. Rep. 2018, 38, BSR20180915. [Google Scholar] [CrossRef]
- Huang, Z.; Zhou, J.-K.; Peng, Y.; He, W.; Huang, C. The Role of Long Noncoding RNAs in Hepatocellular Carcinoma. Mol. Cancer 2020, 19, 77. [Google Scholar] [CrossRef] [PubMed]
- Mondal, P.; Natesh, J.; Kamal, M.A.; Meeran, S.M. Non-Coding RNAs in Lung Cancer Chemoresistance. Curr. Drug Metab. 2019, 20, 1023–1032. [Google Scholar] [CrossRef] [PubMed]
- Ye, R.; Tang, R.; Gan, S.; Li, R.; Cheng, Y.; Guo, L.; Zeng, C.; Sun, Y. New Insights into Long Non-Coding RNAs in Non-Small Cell Lung Cancer. Biomed. Pharmacother. 2020, 131, 110775. [Google Scholar] [CrossRef]
- Bhat, A.A.; Younes, S.N.; Raza, S.S.; Zarif, L.; Nisar, S.; Ahmed, I.; Mir, R.; Kumar, S.; Sharawat, S.K.; Hashem, S.; et al. Role of Non-Coding RNA Networks in Leukemia Progression, Metastasis and Drug Resistance. Mol. Cancer 2020, 19, 57. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Cheng, Z.; Pang, Y.; Cui, L.; Qian, T.; Quan, L.; Zhao, H.; Shi, J.; Ke, X.; Fu, L. Role of MicroRNAs, CircRNAs and Long Noncoding RNAs in Acute Myeloid Leukemia. J. Hematol. Oncol. 2019, 12, 51. [Google Scholar] [CrossRef]
- Abildgaard, C.; Do Canto, L.M.; Steffensen, K.D.; Rogatto, S.R. Long Non-Coding RNAs Involved in Resistance to Chemotherapy in Ovarian Cancer. Front. Oncol. 2019, 9, 1549. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Li, H.; Du, R.; Meng, J.; Yang, G. Involvement of Non-Coding RNAs in Chemotherapy Resistance of Ovarian Cancer. J. Cancer 2018, 9, 1966–1972. [Google Scholar] [CrossRef]
- Wu, Q.B.; Sheng, X.; Zhang, N.; Yang, M.W.; Wang, F. Role of MicroRNAs in the Resistance of Colorectal cancer to Chemoradiotherapy (Review). Mol. Clin. Oncol. 2018, 8, 523–527. [Google Scholar]
- Magee, P.; Shi, L.; Garofalo, M. Role of MicroRNAs in Chemoresistance. Ann. Transl. Med. 2015, 3, 332. [Google Scholar] [PubMed]
- An, X.; Sarmiento, C.; Tan, T.; Zhu, H. Regulation of Multidrug Resistance by MicroRNAs in Anti-Cancer Therapy. Acta Pharm. Sin. B 2017, 7, 38–51. [Google Scholar] [CrossRef] [PubMed]
- Hua, X.; Sun, Y.; Chen, J.; Wu, Y.; Sha, J.; Han, S.; Zhu, X. Circular RNAs in Drug Resistant Tumors. Biomed. Pharmacother. 2019, 118, 109233. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Kong, S.; Wang, F.; Ju, S. CircRNAs: Biogenesis, Functions, and Role in Drug-Resistant Tumours. Mol. Cancer 2020, 19, 119. [Google Scholar] [CrossRef]
- Xu, T.; Wang, M.; Jiang, L.; Ma, L.; Wan, L.; Chen, Q.; Wei, C.; Wang, Z. CircRNAs in Anticancer Drug Resistance: Recent Advances and Future Potential. Mol. Cancer 2020, 19, 127. [Google Scholar] [CrossRef] [PubMed]
- Qi, F.F.; Yang, Y.; Zhang, H.; Chen, H. Long Non-Coding RNAs: Key Regulators in Oxaliplatin Resistance of Colorectal Cancer. Biomed. Pharmacother. 2020, 128, 110329. [Google Scholar] [CrossRef]
- Ashrafizadeh, M.; Zarrabi, A.; Hushmandi, K.; Hashemi, F.; Hashemi, F.; Samarghandian, S.; Najafi, M. MicroRNAs in cancer therapy: Their involvement in oxaliplatin sensitivity/resistance of cancer cells with a focus on colorectal cancer. Life Sci. 2020, 256, 117973. [Google Scholar] [CrossRef] [PubMed]
- Raziq, K.; Cai, M.; Dong, K.; Wang, P.; Afrifa, J.; Fu, S. Competitive endogenous network of lncRNA, miRNA, and mRNA in the chemoresistance of gastrointestinal tract adenocarcinomas. Biomed. Pharmacother. 2020, 130, 110570. [Google Scholar] [CrossRef]
- Xie, P.; Mo, J.L.; Liu, J.H.; Li, X.; Tan, L.M.; Zhang, W.; Zhou, H.H.; Liu, Z.Q. Pharmacogenomics of 5-fluorouracil in colorectal cancer: Review and update. Cell. Oncol. 2020, 43, 989–1001. [Google Scholar]
- Wu, L.; Li, S.; Peng, R.; Gong, S.; Xu, L.; Zou, F. Drug resistance of colon cancer cells to 5-fluorouracil mediated by microRNA-21. Zhonghua Yi Xue Yi Chuan Xue Za Zhi Zhonghua Yixue Yichuanxue Zazhi Chin. J. Med. Genet. 2015, 32, 620–624. [Google Scholar]
- Sun, L.H.; Tian, D.; Yang, Z.C.; Li, J.L. Exosomal MiR-21 Promotes Proliferation, Invasion and Therapy Resistance of Colon Adenocarcinoma Cells through Its Target PDCD4. Sci. Rep. 2020, 10, 8271. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Li, B.; Zhang, B.; Zhang, H.; Suo, J. MiR-361 Enhances Sensitivity to 5-Fluorouracil by Targeting the FOXM1-ABCC5/10 Signaling Pathway in Colorectal Cancer. Oncol. Lett. 2019, 18, 4064–4073. [Google Scholar] [CrossRef]
- Xie, T.; Geng, J.; Wang, Y.; Wang, L.; Huang, M.; Chen, J.; Zhang, K.; Xue, L.; Liu, X.; Mao, X.; et al. FOXM1 evokes 5-fluorouracil resistance in colorectal cancer depending on ABCC10. Oncotarget 2017, 8, 8574. [Google Scholar] [CrossRef]
- Fan, H.; Zhu, J.; Yao, X. Knockdown of Long Non-coding RNA PVT1 Reverses Multidrug Resistance in Colorectal Cancer Cells. Mol. Med. Rep. 2018, 17, 8309–8315. [Google Scholar] [CrossRef]
- He, X.; Ma, J.; Zhang, M.; Cui, J.; Yang, H. Circ_0007031 Enhances Tumor Progression and Promotes 5-Fluorouracil Resistance in Colorectal Cancer through Regulating MiR-133b/ABCC5 Axis. Cancer Biomark. 2020, 29, 531–542. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Ai, Y.Q.; Li, Y.F.; Ye, Q.; Chen, Z.T.; Qin, J.Y.; Liu, Q.Y.; Wang, H.; Ju, Y.H.; Li, W.H.; et al. Microarray Analysis of Circular RNA Expression Profile Associated with 5-Fluorouracil-Based Chemoradiation Resistance in Colorectal Cancer Cells. Biomed Res. Int. 2017, 2017, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Talmon, G.; Wang, J. Erratum: MicroRNA-587 Antagonizes 5-FU-Induced Apoptosis and Confers Drug Resistance by Regulating PPP2R1B Expression in Colorectal Cancer. Cell Death Dis. 2016, 7, e2525. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Liang, Y.; Shen, L.; Shen, L. MicroRNA-204 Modulates Colorectal Cancer Cell Sensitivity in Response to 5-Fluorouracil-Based Treatment by Targeting High Mobility Group Protein A2. Biol. Open 2016, 5, 563–570. [Google Scholar] [CrossRef]
- Li, Y.; Huang, S.; Li, Y.; Zhang, W.; He, K.; Zhao, M.; Lin, H.; Li, D.; Zhang, H.; Zheng, Z.; et al. Decreased Expression of LncRNA SLC25A25-AS1 Promotes Proliferation, Chemoresistance, and EMT in Colorectal Cancer Cells. Tumour Biol. 2016, 37, 14205–14215. [Google Scholar] [CrossRef] [PubMed]
- Valeri, N.; Gasparini, P.; Braconi, C.; Paone, A.; Lovat, F.; Fabbri, M.; Sumani, K.M.; Alder, H.; Amadori, D.; Patel, T.; et al. MicroRNA-21 Induces Resistance to 5-Fluorouracil by down-Regulating Human DNA MutS Homolog 2 (HMSH2). Proc. Natl. Acad. Sci. USA 2010, 107, 21098–21103. [Google Scholar] [CrossRef]
- Deng, J.; Lei, W.; Fu, J.C.; Zhang, L.; Li, J.H.; Xiong, J.P. Targeting MiR-21 Enhances the Sensitivity of Human Colon Cancer HT-29 Cells to Chemoradiotherapy in Vitro. Biochem. Biophys. Res. Commun. 2014, 443, 789–795. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Yang, F.; Wang, Y.; Wang, Y.; Xue, G.; Mei, Q.; Wang, F.; Sun, S. MicroRNA-23a Antisense Enhances 5-Fluorouracil Chemosensitivity through APAF-1/Caspase-9 Apoptotic Pathway in Colorectal Cancer Cells: Mir-23a Downregulated 5-Fu-Induced Apoptosis. J. Cell. Biochem. 2014, 115, 772–784. [Google Scholar] [CrossRef] [PubMed]
- Chai, H.; Liu, M.; Tian, R.; Li, X.; Tang, H. MiR-20a Targets BNIP2 and Contributes Chemotherapeutic Resistance in Colorectal Adenocarcinoma SW480 and SW620 Cell Lines. Acta Biochim. Biophys. Sin. (Shanghai) 2011, 43, 217–225. [Google Scholar] [CrossRef]
- Zheng, H.; Yan, B.; Wu, Q.; Zhang, J. MicroRNA-9-5p increases the sensitivity of colorectal cancer cells to 5-fluorouracil by downregulating high mobility group A2 expression. Oncol. Lett. 2021, 21. [Google Scholar] [CrossRef]
- Bian, Z.; Jin, L.; Zhang, J.; Yin, Y.; Quan, C.; Hu, Y.; Feng, Y.; Liu, H.; Fei, B.; Mao, Y.; et al. LncRNA-UCA1 Enhances Cell Proliferation and 5-Fluorouracil Resistance in Colorectal Cancer by Inhibiting MiR-204-5p. Sci. Rep. 2016, 6, 23892. [Google Scholar] [CrossRef]
- Xian, Z.; Hu, B.; Wang, T.; Zeng, J.; Cai, J.; Zou, Q.; Zhu, P. LncRNA UCA1 Contributes to 5-Fluorouracil Resistance of Colorectal Cancer Cells through MiR-23b-3p/ZNF281 Axis. OncoTargets Ther. 2020, 13, 7571–7583. [Google Scholar] [CrossRef]
- Jiang, Z.; Li, L.; Hou, Z.; Liu, W.; Wang, H.; Zhou, T.; Li, Y.; Chen, S. LncRNA HAND2-AS1 Inhibits 5-Fluorouracil Resistance by Modulating MiR-20a/PDCD4 Axis in Colorectal Cancer. Cell. Signal. 2020, 66, 109483. [Google Scholar] [CrossRef]
- Qu, L.; Chen, Y.; Zhang, F.; He, L. Long non-coding RNA DLGAP1-AS1/miR-149-5p/TGFB2 axis contributes to colorectal cancer progression and 5-FU resistance by regulating smad2 pathway. Mol. Ther. Oncolytics 2021, 20, 607–624. [Google Scholar] [CrossRef]
- Ren, T.J.; Liu, C.; Hou, J.F.; Shan, F.X. CircDDX17 Reduces 5-Fluorouracil Resistance and Hinders Tumorigenesis in Colorectal Cancer by Regulating MiR-31-5p/KANK1 Axis. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 1743–1754. [Google Scholar] [PubMed]
- Yu, X.; Shi, W.; Zhang, Y.; Wang, X.; Sun, S.; Song, Z.; Liu, M.; Zeng, Q.; Cui, S.; Qu, X. CXCL12/CXCR4 Axis Induced MiR-125b Promotes Invasion and Confers 5-Fluorouracil Resistance through Enhancing Autophagy in Colorectal Cancer. Sci. Rep. 2017, 7, 42226. [Google Scholar] [CrossRef]
- Zhang, H.; Tang, J.; Li, C.; Kong, J.; Wang, J.; Wu, Y.; Xu, E.; Lai, M. MiR-22 Regulates 5-FU Sensitivity by Inhibiting Autophagy and Promoting Apoptosis in Colorectal Cancer Cells. Cancer Lett. 2015, 356 Pt B, 781–790. [Google Scholar] [CrossRef]
- Liu, F.; Ai, F.Y.; Zhang, D.C.; Tian, L.; Yang, Z.Y.; Liu, S.J. LncRNA NEAT1 knockdown attenuates autophagy to elevate 5-FU sensitivity in colorectal cancer via targeting miR-34a. Cancer Med. 2019, 9, 1079–1091. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Lan, Z.; He, J.; Lai, Q.; Yao, X.; Li, Q.; Liu, Y.; Lai, H.; Gu, C.; Yan, Q.; et al. LncRNA SNHG6 Promotes Chemoresistance through ULK1-Induced Autophagy by Sponging MiR-26a-5p in Colorectal Cancer Cells. Cancer Cell Int. 2019, 19, 234. [Google Scholar] [CrossRef]
- Xu, X.; Chen, X.; Xu, M.; Liu, X.; Pan, B.; Qin, J.; Xu, T.; Zeng, K.; Pan, Y.; He, B.; et al. MiR-375-3p Suppresses Tumorigenesis and Partially Reverses Chemoresistance by Targeting YAP1 and SP1 in Colorectal Cancer Cells. Aging 2019, 11, 7357–7385. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Xie, T.; Mao, X.; Xue, L.; Chu, X.; Chen, L. MicroRNA-149 increases the sensitivity of colorectal cancer cells to 5-fluorouracil by targeting forkhead box transcription factor FOXM1. Cell. Physiol. Biochem. 2016, 39, 617–629. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.Y.; Deng, J.; Xiang, X.J.; Zhang, L.; Yu, F.; Chen, J.; Sun, Z.; Feng, M.; Xiong, J.P. miR-320 enhances the sensitivity of human colon cancer cells to chemoradiotherapy in vitro by targeting FOXM1. Biochem. Biophys. Res. Commun. 2015, 457, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Pei, L.; Xie, P.; Guo, G. Circ-PRKDC Contributes to 5-Fluorouracil Resistance of Colorectal Cancer Cells by Regulating MiR-375/FOXM1 Axis and Wnt/β-Catenin Pathway. OncoTargets Ther. 2020, 13, 5939–5953. [Google Scholar] [CrossRef]
- Xu, F.; Ye, M.L.; Zhang, Y.P.; Li, W.J.; Li, M.T.; Wang, H.Z.; Qiu, X.; Xu, Y.; Yin, J.W.; Hu, Q.; et al. MicroRNA-375-3p Enhances Chemosensitivity to 5-Fluorouracil by Targeting Thymidylate Synthase in Colorectal Cancer. Cancer Sci. 2020, 111, 1528–1541. [Google Scholar] [CrossRef]
- Wang, X.; Jiang, G.; Ren, W.; Wang, B.; Yang, C.; Li, M. LncRNA NEAT1 Regulates 5-Fu Sensitivity, Apoptosis and Invasion in Colorectal Cancer through the MiR-150-5p/CPSF4 Axis. Onco. Targets. Ther. 2020, 13, 6373–6383. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Hu, H.; Yuan, Z.; Zhang, Q.; Xiong, H.; Hu, Z.; Wu, H.; Huang, R.; Wang, G.; Tang, Q. LncRNA NEAT1 Remodels Chromatin to Promote the 5-Fu Resistance by Maintaining Colorectal Cancer Stemness. Cell Death Dis. 2019, 11, 962. [Google Scholar] [CrossRef]
- Jiang, X.; Li, Q.; Zhang, S.; Song, C.; Zheng, P. Long Noncoding RNA GIHCG Induces Cancer Progression and Chemoresistance and Indicates Poor Prognosis in Colorectal Cancer. Onco. Targets Ther. 2019, 12, 1059–1070. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Hu, H.; Wang, Y.; Huang, Q.; Huang, R.; Chen, Y.; Ma, T.; Qiao, T.; Zhang, Q.; Wu, H.; et al. Long Non-Coding RNA TUG1 Mediates 5-Fluorouracil Resistance by Acting as a CeRNA of MiR-197-3p in Colorectal Cancer. J. Cancer 2019, 10, 4603–4613. [Google Scholar] [CrossRef] [PubMed]
- Qiao, L.; Liu, X.; Tang, Y.; Zhao, Z.; Zhang, J.; Liu, H. Knockdown of Long Non-Coding RNA Prostate Cancer-Associated NcRNA Transcript 1 Inhibits Multidrug Resistance and c-Myc-Dependent Aggressiveness in Colorectal Cancer Caco-2 and HT-29 Cells. Mol. Cell. Biochem. 2018, 441, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Ragusa, M.; Majorana, A.; Statello, L.; Maugeri, M.; Salito, L.; Barbagallo, D.; Guglielmino, M.R.; Duro, L.R.; Angelica, R.; Caltabiano, R.; et al. Specific alterations of microRNA transcriptome and global network structure in colorectal carcinoma after cetuximab treatment. Mol. Cancer Ther. 2010, 9, 3396–3409. [Google Scholar] [CrossRef]
- Abu, N.; Hon, K.W.; Jeyaraman, S.; Yahaya, A.; Abdullah, N.M.A.; Mustangin, M.; Sulaiman, S.A.; Jamal, R.; Ab-Mutalib, N.S. Identification of Differentially Expressed Circular RNAs in Chemoresistant Colorectal Cancer. Epigenomics 2019, 11, 875–884. [Google Scholar] [CrossRef]
- Li, S.; Zheng, S. Down-Regulation of Circ_0032833 Sensitizes Colorectal Cancer to 5-Fluorouracil and Oxaliplatin Partly Depending on the Regulation of MiR-125-5p and MSI1. Cancer Manag. Res. 2020, 12, 11257–11269. [Google Scholar] [CrossRef]
- Panczyk, M. Pharmacogenetics research on chemotherapy resistance in colorectal cancer over the last 20 years. World J. Gastroenterol. WJG 2014, 20, 9775. [Google Scholar] [CrossRef]
- Mao, L.; Li, Y.; Zhao, J.; Li, Q.; Yang, B.; Wang, Y.; Zhu, Z.; Sun, H.; Zhai, Z. Transforming growth factor-β1 contributes to oxaliplatin resistance in colorectal cancer via epithelial to mesenchymal transition. Oncol. Lett. 2017, 14, 647–654. [Google Scholar] [CrossRef]
- Martinez-Cardús, A.; Martinez-Balibrea, E.; Bandrés, E.; Malumbres, R.; Ginés, A.; Manzano, J.L.; Taron, M.; Garcia-Foncillas, J.; Abad, A. Pharmacogenomic approach for the identification of novel determinants of acquired resistance to oxaliplatin in colorectal cancer. Mol. Cancer Ther. 2009, 8, 194–202. [Google Scholar] [CrossRef]
- Huang, W.S.; Hsieh, M.C.; Huang, C.Y.; Kuo, Y.H.; Tung, S.Y.; Shen, C.H.; Hsieh, Y.Y.; Teng, C.C.; Lee, K.F.; Chen, T.C.; et al. The association of CXC receptor 4 mediated signaling pathway with oxaliplatin-resistant human colorectal cancer cells. PLoS ONE 2016, 11, e0159927. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Deng, G.; Fu, Y.; Han, Y.; Guo, C.; Yin, L.; Cai, C.; Shen, H.; Wu, S.; Zeng, S. FOXC2 promotes oxaliplatin resistance by inducing epithelial-mesenchymal transition via MAPK/ERK signaling in colorectal cancer. OncoTargets Ther. 2020, 13, 1625. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xu, Z.; Sun, Y.; Chi, P.; Lu, X. Knockdown of KLK11 reverses oxaliplatin resistance by inhibiting proliferation and activating apoptosis via suppressing the PI3K/AKT signal pathway in colorectal cancer cell. OncoTargets Ther. 2018, 11, 809. [Google Scholar] [CrossRef]
- Gao, R.; Fang, C.; Xu, J.; Tan, H.; Li, P.; Ma, L. LncRNA CACS15 Contributes to Oxaliplatin Resistance in Colorectal Cancer by Positively Regulating ABCC1 through Sponging MiR-145. Arch. Biochem. Biophys. 2019, 663, 183–191. [Google Scholar] [CrossRef]
- Yue, B.; Cai, D.; Liu, C.; Fang, C.; Yan, D. Linc00152 Functions as a Competing Endogenous RNA to Confer Oxaliplatin Resistance and Holds Prognostic Values in Colon Cancer. Mol. Ther. 2016, 24, 2064–2077. [Google Scholar] [CrossRef] [PubMed]
- Lin, Y.C.; Yu, Y.S.; Lin, H.H.; Hsiao, K.Y. Oxaliplatin-Induced DHX9 Phosphorylation Promotes Oncogenic Circular RNA CCDC66 Expression and Development of Chemoresistance. Cancers 2020, 12, 697. [Google Scholar] [CrossRef]
- Zhang, L.; Pickard, K.; Jenei, V.; Bullock, M.D.; Bruce, A.; Mitter, R.; Kelly, G.; Paraskeva, C.; Strefford, J.; Primrose, J.; et al. MiR-153 Supports Colorectal Cancer Progression via Pleiotropic Effects That Enhance Invasion and Chemotherapeutic Resistance. Cancer Res. 2013, 73, 6435–6447. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Hu, X.; Miao, X.; Zhu, K.; Cui, S.; Meng, Q.; Sun, J.; Wang, T. MicroRNA-425-5p Regulates Chemoresistance in Colorectal Cancer Cells via Regulation of Programmed Cell Death 10. J. Cell. Mol. Med. 2016, 20, 360–369. [Google Scholar] [CrossRef]
- Qin, Y.; Li, L.; Wang, F.; Zhou, X.; Liu, Y.; Yin, Y.; Qi, X. Knockdown of Mir-135b sensitizes colorectal cancer cells to oxaliplatin-induced apoptosis through increase of FOXO1. Cell. Physiol. Biochem. 2018, 48, 1628–1637. [Google Scholar]
- Chen, R.; Zhou, S.; Chen, J.; Lin, S.; Ye, F.; Jiang, P. LncRNA BLACAT1/MiR-519d-3p/CREB1 Axis Mediates Proliferation, Apoptosis, Migration, Invasion, and Drug-Resistance in Colorectal Cancer Progression. Cancer Manag. Res. 2020, 12, 13137–13148. [Google Scholar] [CrossRef]
- Li, L.; Shang, J.; Zhang, Y.; Liu, S.; Peng, Y.; Zhou, Z.; Pan, H.; Wang, X.; Chen, L.; Zhao, Q. MEG3 Is a Prognostic Factor for CRC and Promotes Chemosensitivity by Enhancing Oxaliplatin-Induced Cell Apoptosis. Oncol. Rep. 2017, 38, 1383–1392. [Google Scholar] [CrossRef]
- Wang, H.; Li, H.; Zhang, L.; Yang, D. Overexpression of MEG3 Sensitizes Colorectal Cancer Cells to Oxaliplatin through Regulation of MiR-141/PDCD4 Axis. Biomed. Pharmacother. 2018, 106, 1607–1615. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.; Shi, H.; Ba, M.; Lin, S.; Tang, H.; Zeng, X.; Zhang, X. MiR-409-3p Sensitizes Colon Cancer Cells to Oxaliplatin by Inhibiting Beclin-1-Mediated Autophagy. Int. J. Mol. Med. 2016, 37, 1030–1038. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, C.; Liu, X.; Wang, Y.; Zhao, R.; Yang, Y.; Zheng, X.; Zhang, Y.; Zhang, X. CircHIPK3 Promotes Oxaliplatin-Resistance in Colorectal Cancer through Autophagy by Sponging MiR-637. EBioMedicine 2019, 48, 277–288. [Google Scholar] [CrossRef]
- Ren, J.; Ding, L.; Zhang, D.; Shi, G.; Xu, Q.; Shen, S.; Wang, Y.; Wang, T.; Hou, Y. Carcinoma-Associated Fibroblasts Promote the Stemness and Chemoresistance of Colorectal Cancer by Transferring Exosomal LncRNA H19. Theranostics 2018, 8, 3932–3948. [Google Scholar] [CrossRef]
- Lai, M.; Liu, G.; Li, R.; Bai, H.; Zhao, J.; Xiao, P.; Mei, J. Hsa_circ_0079662 Induces the Resistance Mechanism of the Chemotherapy Drug Oxaliplatin through the TNF-α Pathway in Human Colon Cancer. J. Cell. Mol. Med. 2020, 24, 5021–5027. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, H.; Yang, H.; Bai, M.; Ning, T.; Deng, T.; Liu, R.; Fan, Q.; Zhu, K.; Li, J.; et al. Exosome-Delivered CircRNA Promotes Glycolysis to Induce Chemoresistance through the MiR-122-PKM2 Axis in Colorectal Cancer. Mol. Oncol. 2020, 14, 539–555. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Wan, G.; Spizzo, R.; Ivan, C.; Mathur, R.; Hu, X.; Ye, X.; Lu, J.; Fan, F.; Xia, L.; et al. MiR-203 Induces Oxaliplatin Resistance in Colorectal Cancer Cells by Negatively Regulating ATM Kinase. Mol. Oncol. 2014, 8, 83–92. [Google Scholar] [CrossRef]
- Liang, H.; Xu, Y.; Zhang, Q.; Yang, Y.; Mou, Y.; Gao, Y.; Chen, R.; Chen, C.; Dai, P. MiR-483-3p Regulates Oxaliplatin Resistance by Targeting FAM171B in Human Colorectal Cancer Cells. Artif. Cells Nanomed. Biotechnol. 2019, 47, 725–736. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Zhu, H.; Wang, J.; Sui, H.; Zhang, H.; Jin, C.; Li, L.; Xu, T.; Miao, R. MiR-492 Is Functionally Involved in Oxaliplatin Resistance in Colon Cancer Cells LS174T via Its Regulating the Expression of CD147. Mol. Cell. Biochem. 2015, 405, 73–79. [Google Scholar] [CrossRef]
- Wu, Y.Z.; Lin, H.Y.; Zhang, Y.; Chen, W.F. miR-200b-3p mitigates oxaliplatin resistance via targeting TUBB3 in colorectal cancer. J. Gene Med. 2020, 22, e3178. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Sun, W.; Yu, J.; Zhou, Y.; Gu, Y.; Han, J.; Zhou, L.; Jiang, X.; Wang, C. LINC00460-MiR-149-5p/MiR-150-5p-Mutant P53 Feedback Loop Promotes Oxaliplatin Resistance in Colorectal Cancer. Mol. Ther. Nucleic Acids 2020, 22, 1004–1015. [Google Scholar] [CrossRef]
- Xie, L.; Cui, G.; Li, T. Long Non-Coding RNA CBR3-AS1 Promotes Stem-like Properties and Oxaliplatin Resistance of Colorectal Cancer by Sponging MiR-145-5p. Res. Sq. 2020. [Google Scholar] [CrossRef]
- Fan, C.; Yuan, Q.; Liu, G.; Zhang, Y.; Yan, M.; Sun, Q.; Zhu, C. Long Non-Coding RNA MALAT1 Regulates Oxaliplatin-Resistance via MiR-324-3p/ADAM17 Axis in Colorectal Cancer Cells. Cancer Cell Int. 2020, 20, 473. [Google Scholar] [CrossRef] [PubMed]
- Ekblad, L.; Kjellström, J.; Johnsson, A. Reduced drug accumulation is more important in acquired resistance against oxaliplatin than against cisplatin in isogenic colon cancer cells. Anti-Cancer Drugs 2010, 21, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Kang, Y.; Chen, L.; Wang, H.; Liu, J.; Zeng, S.; Yu, L. The drug-resistance mechanisms of five platinum-based antitumor agents. Front. Pharmacol. 2020, 11, 343. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhang, Y.; Qin, X.; Wang, Y.; Fu, J. FGF9 promotes cisplatin resistance in colorectal cancer via regulation of Wnt/β-catenin signaling pathway. Exp. Ther. Med. 2020, 19, 1711–1718. [Google Scholar] [CrossRef] [PubMed]
- Ping, G.; Xiong, W.; Zhang, L.; Li, Y.; Zhang, Y.; Zhao, Y. Silencing Long Noncoding RNA PVT1 Inhibits Tumorigenesis and Cisplatin Resistance of Colorectal Cancer. Am. J. Transl. Res. 2018, 10, 138–149. [Google Scholar]
- Zheng, Z.H.; You, H.Y.; Feng, Y.J.; Zhang, Z.T. LncRNA KCNQ1OT1 Is a Key Factor in the Reversal Effect of Curcumin on Cisplatin Resistance in the Colorectal Cancer Cells. Res. Sq. 2020. [Google Scholar] [CrossRef]
- Han, Y.; Zhou, S.; Wang, X.; Mao, E.; Huang, L. SNHG14 Stimulates Cell Autophagy to Facilitate Cisplatin Resistance of Colorectal Cancer by Regulating MiR-186/ATG14 Axis. Biomed. Pharmacother. 2020, 121, 109580. [Google Scholar] [CrossRef]
- Xiao, Z.; Qu, Z.; Chen, Z.; Fang, Z.; Zhou, K.; Huang, Z.; Guo, X.; Zhang, Y. LncRNA HOTAIR Is a Prognostic Biomarker for the Proliferation and Chemoresistance of Colorectal Cancer via MiR-203a-3p-Mediated Wnt/ß-Catenin Signaling Pathway. Cell. Physiol. Biochem. 2018, 46, 1275–1285. [Google Scholar] [CrossRef]
- Wang, Z.K.; Yang, L.; Wu, L.L.; Mao, H.; Zhou, Y.H.; Zhang, P.F.; Dai, G.H. Long non-coding RNA LINC00261 sensitizes human colon cancer cells to cisplatin therapy. Braz. J. Med. Biol. Res. 2018, 51. [Google Scholar] [CrossRef] [PubMed]
- Ren, L.L.; Yan, T.T.; Shen, C.Q.; Tang, J.Y.; Kong, X.; Wang, Y.C.; Chen, J.; Liu, Q.; He, J.; Zhong, M.; et al. The Distinct Role of Strand-Specific MiR-514b-3p and MiR-514b-5p in Colorectal Cancer Metastasis. Cell Death Dis. 2018, 9, 687. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; He, L.; Zhang, H.; Chen, Y. Knockdown of MiR-20a Enhances Sensitivity of Colorectal Cancer Cells to Cisplatin by Increasing ASK1 Expression. Cell. Physiol. Biochem. 2018, 47, 1432–1441. [Google Scholar] [CrossRef]
- Shi, H.; Li, K.; Feng, J.; Liu, G.; Feng, Y.; Zhang, X. LncRNA-DANCR Interferes with MiR-125b-5p/HK2 Axis to Desensitize Colon Cancer Cells to Cisplatin Vis Activating Anaerobic Glycolysis. Front. Oncol. 2020, 10, 1034. [Google Scholar] [CrossRef]
- Luo, P.; Wu, S.; Ji, K.; Yuan, X.; Li, H.; Chen, J.; Tian, Y.; Qiu, Y.; Zhong, X. LncRNA MIR4435-2HG Mediates Cisplatin Resistance in HCT116 Cells by Regulating Nrf2 and HO-1. PLoS ONE 2020, 15, e0223035. [Google Scholar] [CrossRef]
- Sonowal, H.; Pal, P.B.; Wen, J.J.; Awasthi, S.; Ramana, K.V.; Srivastava, S.K. Aldose reductase inhibitor increases doxorubicin-sensitivity of colon cancer cells and decreases cardiotoxicity. Sci. Rep. 2017, 7, 3182. [Google Scholar] [CrossRef]
- Khameneh, Z.R.; Mohammadian, M.; Rasouli, M.A.; Moradi, Z.; Ahmadi, Z.; Khiyavi, A.A. Effects of Curcumin in Combination with Doxorubicin in Human Colorectal Cancer Cell Line. Asian Pac. J. Cancer Biol. 2018, 3, 89–92. [Google Scholar] [CrossRef]
- Du, J.; He, Y.; Li, P.; Wu, W.; Chen, Y.; Ruan, H. IL-8 regulates the doxorubicin resistance of colorectal cancer cells via modulation of multidrug resistance 1 (MDR1). Cancer Chemother. Pharmacol. 2018, 81, 1111–1119. [Google Scholar] [CrossRef]
- Yang, G.E.; Jiang, O.U.; Ling, D.; Jiang, X.; Yuan, P.; Zeng, G.; Zhu, J.; Tian, J.; Weng, Y.; Wu, D. MicroRNA-522 Reverses Drug Resistance of Doxorubicin-Induced HT29 Colon Cancer Cell by Targeting ABCB5. Mol. Med. Rep. 2015, 12, 3930–3936. [Google Scholar] [CrossRef]
- Shi, X.; Valizadeh, A.; Mir, S.M.; Asemi, Z.; Karimian, A.; Majidina, M.; Safa, A.; Yosefi, B. MiRNA-29a Reverses P-Glycoprotein-Mediated Drug Resistance and Inhibits Proliferation via up-Regulation of PTEN in Colon Cancer Cells. Eur. J. Pharmacol. 2020, 880, 173138. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Ma, T.; Duan, J.; Liu, X.; Liu, L. MicroRNA-223-induced inhibition of the FBXW7 gene affects the proliferation and apoptosis of colorectal cancer cells via the Notch and Akt/mTOR pathways. Mol. Med. Rep. 2021, 23, 154. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, M. MiR-132 Regulates Adriamycin Resistance in Colorectal Cancer Cells through Targeting Extracellular Signal-Regulated Kinase 1. Cancer Biother. Radiopharm. 2019, 34, 398–404. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Zhao, Z.; Song, J.; Luo, B.; Huang, L. MiR-223 promotes the doxorubicin resistance of colorectal cancer cells via regulating epithelial–mesenchymal transition by targeting FBXW7. Acta Biochim. Biophys. Sin. 2018, 50, 597–604. [Google Scholar] [CrossRef]
- Qu, J.; Zhao, L.; Zhang, P.; Wang, J.; Xu, N.; Mi, W.; Jiang, X.; Zhang, C.; Qu, J. MicroRNA-195 Chemosensitizes Colon Cancer Cells to the Chemotherapeutic Drug Doxorubicin by Targeting the First Binding Site of BCL2L2 MRNA: MicroRNA-195 Chemosensitizes Colon Cancer Cells. J. Cell. Physiol. 2015, 230, 535–545. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Zhang, R.; Yang, D.; Li, J.; Yan, X.; Jin, K.; Li, W.; Liu, X.; Zhao, J.; Shang, W.; et al. Knockdown of long non-coding RNA XIST inhibited doxorubicin resistance in colorectal cancer by upregulation of miR-124 and downregulation of SGK1. Cell. Physiol. Biochem. 2018, 51, 113–128. [Google Scholar] [CrossRef]
- Xiong, M.; Wu, M.; Peng, D.; Huang, W.; Chen, Z.; Ke, H.; Chen, Z.; Song, W.; Zhao, Y.; Xiang, A.P.; et al. LncRNA DANCR represses Doxorubicin-induced apoptosis through stabilizing MALAT1 expression in colorectal cancer cells. Cell Death Dis. 2021, 12, 1–17. [Google Scholar] [CrossRef] [PubMed]
- He, Y.; Wang, J.; Wang, J.; Yung, V.Y.W.; Hsu, E.; Li, A.; Kang, Q.; Ma, J.; Han, Q.; Jin, P.; et al. MicroRNA-135b regulates apoptosis and chemoresistance in colorectal cancer by targeting large tumor suppressor kinase 2. Am. J. Cancer Res. 2015, 5, 1382. [Google Scholar]
- Silva-Fisher, J.M.; Dang, H.X.; White, N.M.; Strand, M.S.; Krasnick, B.A.; Rozycki, E.B.; Jeffers, G.G.L.; Grossman, J.G.; Highkin, M.K.; Tang, C.; et al. Long Non-Coding RNA RAMS11 Promotes Metastatic Colorectal Cancer Progression. Nat. Commun. 2020, 11, 2156. [Google Scholar] [CrossRef]
- Zhu, R.; Yang, G.; Cao, Z.; Shen, K.; Zheng, L.; Xiao, J.; You, L.; Zhang, T. The Prospect of Serum and Glucocorticoid-Inducible Kinase 1 (SGK1) in Cancer Therapy: A Rising Star. Ther. Adv. Med. Oncol. 2020, 12, 1758835920940946. [Google Scholar] [CrossRef]
| Mechanism | Type of ncRNA | Reported ncRNA | Alteration in CRC Cells | Validated ncRNA Targets | References |
|---|---|---|---|---|---|
| ABC Transporter Family | miRNA | miR-21 | Upregulated | PDCD4/ABCC5/CD44 | [195,196] |
| miRNA | miR-361 | Downregulated | FOXM1, ABCC5/10 | [197,198] | |
| lncRNA | PVT1 | Upregulated | MRP1, P-gp | [199] | |
| circRNA | hsa_circ_0007031 | Upregulated | miR-133b/ABCC5 | [200,201] | |
| PI3K/AKT signaling pathway | miRNA | miR-587 | Upregulated | PPP2R1B/pAKT/XIAP | [202] |
| miRNA | miR-204 | Downregulated | HMGA2 | [203] | |
| circRNA | hsa_circ_0007031, hsa_circ_0007006, hsa_circ_0000504 | Upregulated | AKT3 | [201] | |
| MAPK/ERK Pathways and EMT | lncRNA | SLC25A25-AS1 | Downregulated | ERK, p38 | [204] |
| DNA repair mechanisms | miRNA | miR-21 | Upregulated | hMSH2 | [205,206] |
| Apoptosis | miRNA | miR-361 | Downregulated | Caspase 3/7 | [197] |
| miRNA | miR-23a | Upregulated | APAF-1 | [207] | |
| miRNA | miR-20a | Upregulated | BNIP2 | [208] | |
| miRNA | miR-9-5p | Downregulated | HMGA2 | [209] | |
| lncRNA | PVT1 | Upregulated | mTOR, BCL-2 | [198] | |
| lncRNA | UCA1 | Upregulated | miR-204-5p | [210] | |
| lncRNA | UCA1 | Upregulated | miR-23b-3p/ZNF281 | [211] | |
| miRNA | miR-20a | Upregulated | PDCD4 | [212] | |
| lncRNA | HAND2-AS1 | Downregulated | miR-20a/PDCD4 | [212] | |
| lncRNA | DLGAP1-AS1 | Upregulated | miR-149-5p/TGFB2 | [213] | |
| circRNA | hsa_circ_0007031, hsa_circ_0000504 | Upregulated | BCL2 | [201] | |
| circRNA | circDDX17 | Downregulated | miR-31-5p/KANK1 | [214] | |
| Autophagy | miRNA | miR-23b-3p | Downregulated | UCA1 | [211] |
| miRNA | miR-125b | Upregulated | CXCL12/CXCR4 | [215] | |
| miRNA | miR-125b | Upregulated | CXCL12/CXCR4 | [215] | |
| miRNA | miR-22 | Downregulated | PARP/ATG5 | [216] | |
| miRNA | miR-22 | Downregulated | BTG1 | [216] | |
| miRNA | miR-34a | Upregulated | HMGB1/ATG9A/ATG4B | [217] | |
| lncRNA | NEAT1 | Upregulated | miR-34a/HMGB1/ATG9A/ATG4B | [217] | |
| lncRNA | SNHG6 | Upregulated | miR-26a-5p/ULK1 | [218] | |
| Hippo signaling pathway | miRNA | miR-375-3p | Downregulated | YAP1, SP1 | [219] |
| Wnt/β-catenin signaling pathway | miRNA | miR-149 | Downregulated | FOXM1 | [220] |
| miRNA | miR-320 | Downregulated | FOXM1 | [221] | |
| miRNA | miR-375 | Downregulated | FOXM1 | [222] | |
| circRNA | circ-PRKDC | Upregulated | miR-375/FOXM1 | [222] | |
| JAK/STAT signalling Pathway | circRNA | hsa_circ_0000504 | Upregulated | hsa-miR-485-5p/STAT3 | [201] |
| Other chemoresistance related ncRNAs | miRNA | miR-375-3p | Downregulated | TYMS | [223] |
| lncRNA | NEAT1 | Upregulated | miR-150-5p/CPSF4 | [224] | |
| lncRNA | NEAT1 | Upregulated | H3K27ac | [225] | |
| lncRNA | GIHCG | Upregulated | / | [226] | |
| lncRNA | TUG1 | Upregulated | miR-197-3p/TYMS | [227] | |
| lncRNA | PCAT-1 | Upregulated | / | [228] | |
| circRNA | hsa_circ_0048234 | Downregulated | miR-671-5p | [201] | |
| circRNA | hsa_circ_0007031 | Upregulated | miR-885-3p | [201] | |
| circRNA | hsa_circ_0007006, hsa_circ_0000504 | Upregulated | / | [201] | |
| circRNA | has_circ_0008509, has_circ_0084021, has_circ_0087862 | Downregulated | / | [201] | |
| circRNA | has_circ_0008509, has_circ_0084021, has_circ_0087862 | Downregulated | / | [201] | |
| circRNA | has_circRNA_103306 | Upregulated | / | [229,230] | |
| circRNA | has_circRNA_406937 | Downregulated | / | [230] | |
| circRNA | circ_0032833 | Upregulated | miR-125-5p/MSI1 | [231] |
| Mechanism | Type of ncRNA | Reported ncRNA | Alteration in CRC Cells | Validated ncRNA Targets | References |
|---|---|---|---|---|---|
| ABC Transporter Family | lncRNA | CACS15 | Upregulated | miR-145/ABCC1 | [238] |
| PI3K/AKT signaling pathway | lncRNA | LINC00152 | Upregulated | miR-193a-3p/ERBB4/AKT | [239] |
| circRNA | circCCDC66 | Upregulated | DHX9 | [240] | |
| Apoptosis | miRNA | miR-20a | Upregulated | BNIP2 | [208] |
| miRNA | miR-153 | Upregulated | FOXO3a | [241] | |
| miRNA | miR-425-5p | Upregulated | PDCD10 | [242] | |
| miRNA | miR-135b | Upregulated | FOXO1 | [243] | |
| lncRNA | BLACAT1 | Upregulated | miR-519d-3p/ CREB1 | [244] | |
| lncRNA | MEG3 | Downregulated | miR-141/PDCD4 | [245,246] | |
| Autophagy | miRNA | miR-409-3p | Downregulated | Beclin 1 | [247] |
| circRNA | circHIPK3 | Upregulated | miR-637/STAT3/BCL-2/Beclin1 | [248] | |
| Wnt/β-catenin signaling pathway | miRNA | miR-320 | Downregulated | FOXM1 | [221] |
| lncRNA | H19 | Upregulated | miR-141 | [249] | |
| TNF-α signalling pathway | circRNA | hsa_circ_0079662 | Upregulated | hsa-mir-324-5p/HOXA6 | [250] |
| Glycolysis | circRNA | hsa_circ_0005963 | Upregulated | miR-122/PKM2 | [251] |
| Other chemoresistance related ncRNA | miRNA | miR-203 | Upregulated | ATM | [252] |
| miRNA | miR-483-3p | Downregulated | FAM171B | [253] | |
| miRNA | miR-492 | Downregulated | CD147 | [254] | |
| miRNA | miR-200b-3p | Downregulated | TUBB3 | [255] | |
| lncRNA | GIHCG | Upregulated | / | [226] | |
| lncRNA | LINC00460 | Upregulated | miR-149-5p/miR-150-5p/ Mut p53 | [256] | |
| lncRNA | CBR3-AS1 | Upregulated | miR-145-5p | [257] | |
| lncRNA | MALAT1 | Upregulated | miR-324-3p/ADAM17 | [258] |
| Mechanism | Type of ncRNA | Reported ncRNA | Alteration in CRC Cells | Validated ncRNA Targets | References |
|---|---|---|---|---|---|
| ABC Transporter Family | lncRNA | PVT1 | Upregulated | MRP1, MDR1 | [262] |
| Apoptosis | miRNA | miR-153 | Upregulated | FOXO3a | [241] |
| lncRNA | PVT1 | Upregulated | BCL-2, BAX, Cleaved Caspase-3 | [262] | |
| lncRNA | KCNQ1OT1 | Upregulated | miR-497/BCL-2 | [263] | |
| Autophagy | lncRNA | SNHG14 | Upregulated | miR-186/ATG14 | [264] |
| Wnt/β-catenin signaling pathway | miRNA | miR-203a-3p | Downregulated | β-catenin, GRG5 | [265] |
| lncRNA | HOTAIR | Upregulated | miR-203a-3p/β-catenin/GRG5 | [265] | |
| lncRNA | LINC00261 | Downregulated | β-catenin | [266] | |
| EMT | miRNA | miR-514b-3p | Downregulated | E-cadherin, CLDN-1 | [267] |
| miRNA | miR-514b-5p | Upregulated | E-cadherin, CLDN-1 | [267] | |
| ROS signalling pathway | miRNA | miR-20a | Upregulated | ASK1/JNK | [268] |
| Glycolysis | lncRNA | DANCR | Upregulated | miR-125b-5p/HK2 | [269] |
| Nrf2/HO-1 pathway | lncRNA | MIR4435-2HG | Upregulated | Nrf2/HO-1 | [270] |
| Mechanism | Type of ncRNA | Reported ncRNA | Alteration in CRC Cells | Validated ncRNA Targets | References |
|---|---|---|---|---|---|
| ABC Transporter Family | miRNA | miR-522 | Downregulated | ABCB5 | [274] |
| miRNA | miR-29a | Downregulated | MDR1/P-gp | [275] | |
| PI3K/AKT signaling pathway | miRNA | miR-29a | Downregulated | PTEN | [275] |
| miRNA | miR-223 | Upregulated | FBXW7 | [276] | |
| MAPK/ERK Pathways and EMT | miRNA | miR-132 | Downregulated | ERK1 | [277] |
| miRNA | miR-223 | Upregulated | FBXW7 | [278] | |
| lncRNA | SLC25A25-AS1 | Downregulated | ERK, p38 | [204] | |
| Apoptosis | miRNA | miR-195 | Downregulated | BCL2L2 | [279] |
| lncRNA | XIST | Upregulated | SGK1/miR-124 | [280] | |
| lncRNA | DANCR | Upregulated | QK/MALAT1 | [281] | |
| Hippo signaling pathway | miRNA | miR-135b | Upregulated | LATS2 | [282] |
| Other chemoresistance related ncRNA | miRNA | miR-137, miR-127, miR-22 | Downregulated | / | [279] |
| miRNA | miR-21, miR-592 | Upregulated | / | [279] | |
| lncRNA | RAMS11 | Upregulated | TOP2α | [283] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Micallef, I.; Baron, B. The Mechanistic Roles of ncRNAs in Promoting and Supporting Chemoresistance of Colorectal Cancer. Non-Coding RNA 2021, 7, 24. https://doi.org/10.3390/ncrna7020024
Micallef I, Baron B. The Mechanistic Roles of ncRNAs in Promoting and Supporting Chemoresistance of Colorectal Cancer. Non-Coding RNA. 2021; 7(2):24. https://doi.org/10.3390/ncrna7020024
Chicago/Turabian StyleMicallef, Isaac, and Byron Baron. 2021. "The Mechanistic Roles of ncRNAs in Promoting and Supporting Chemoresistance of Colorectal Cancer" Non-Coding RNA 7, no. 2: 24. https://doi.org/10.3390/ncrna7020024
APA StyleMicallef, I., & Baron, B. (2021). The Mechanistic Roles of ncRNAs in Promoting and Supporting Chemoresistance of Colorectal Cancer. Non-Coding RNA, 7(2), 24. https://doi.org/10.3390/ncrna7020024







