microRNA Expression Profiles in the Ventral Hippocampus during Pubertal Development and the Impact of Peri-Pubertal Binge Alcohol Exposure
Abstract
1. Introduction
2. Results
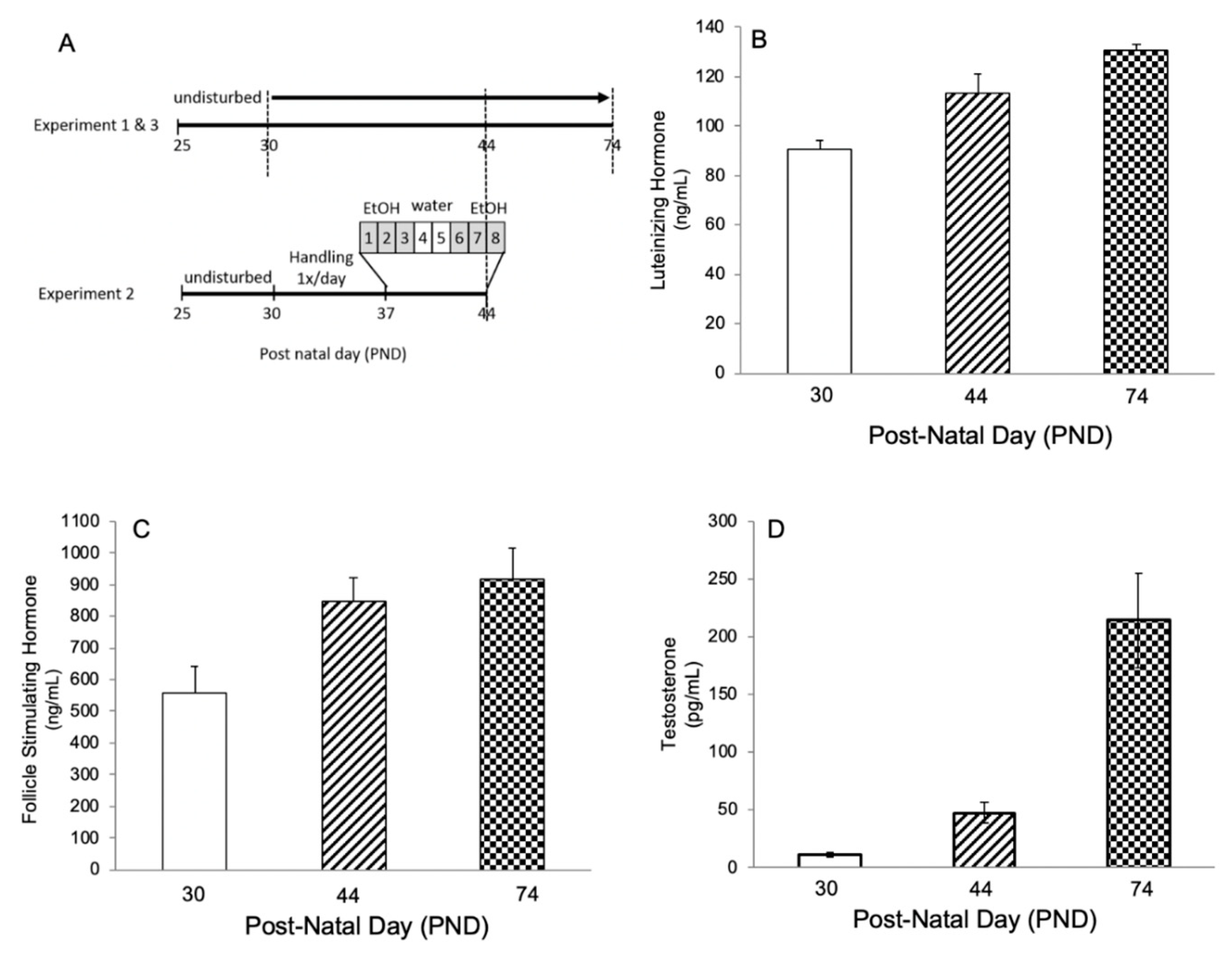
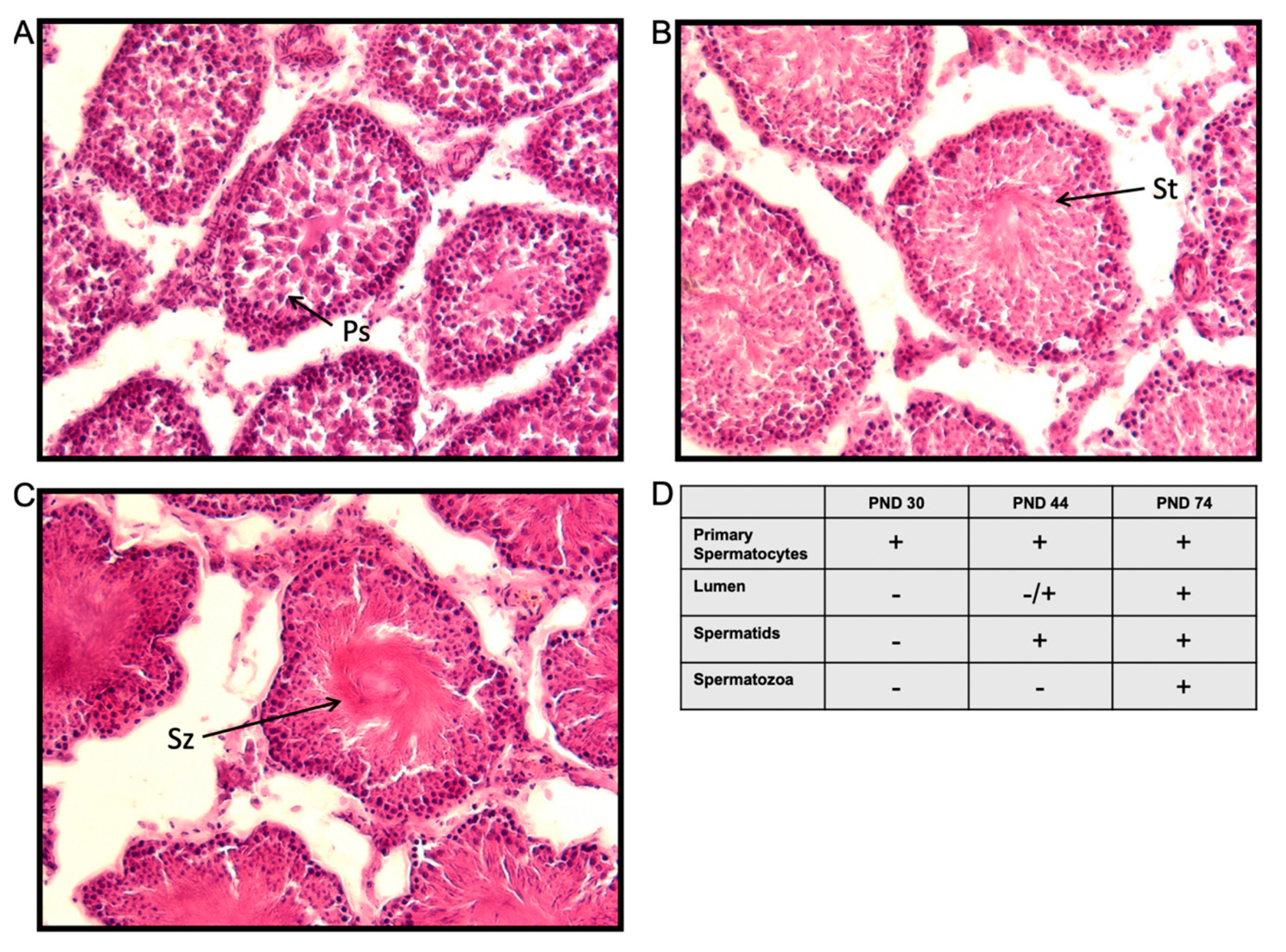
2.1. Experiment 1: Hormone Levels and Spermatogenesis in Untreated Male Wistar Rats at Post-Natal Day (PND) 30, 44, and 74
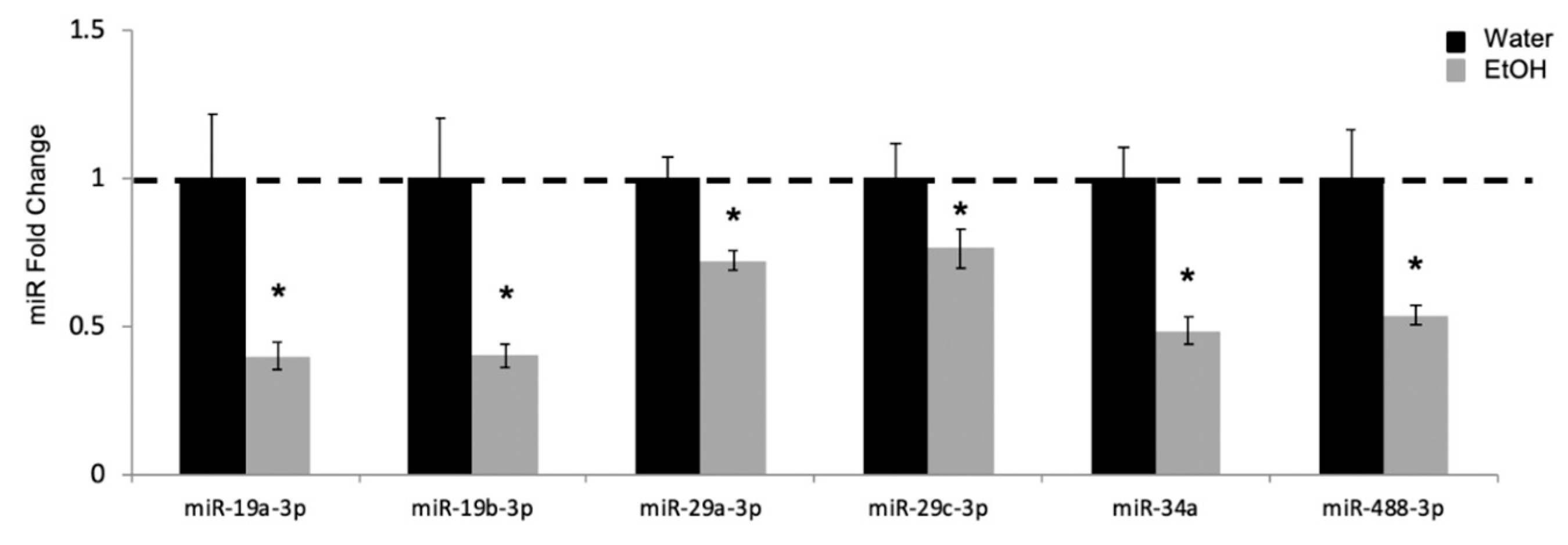
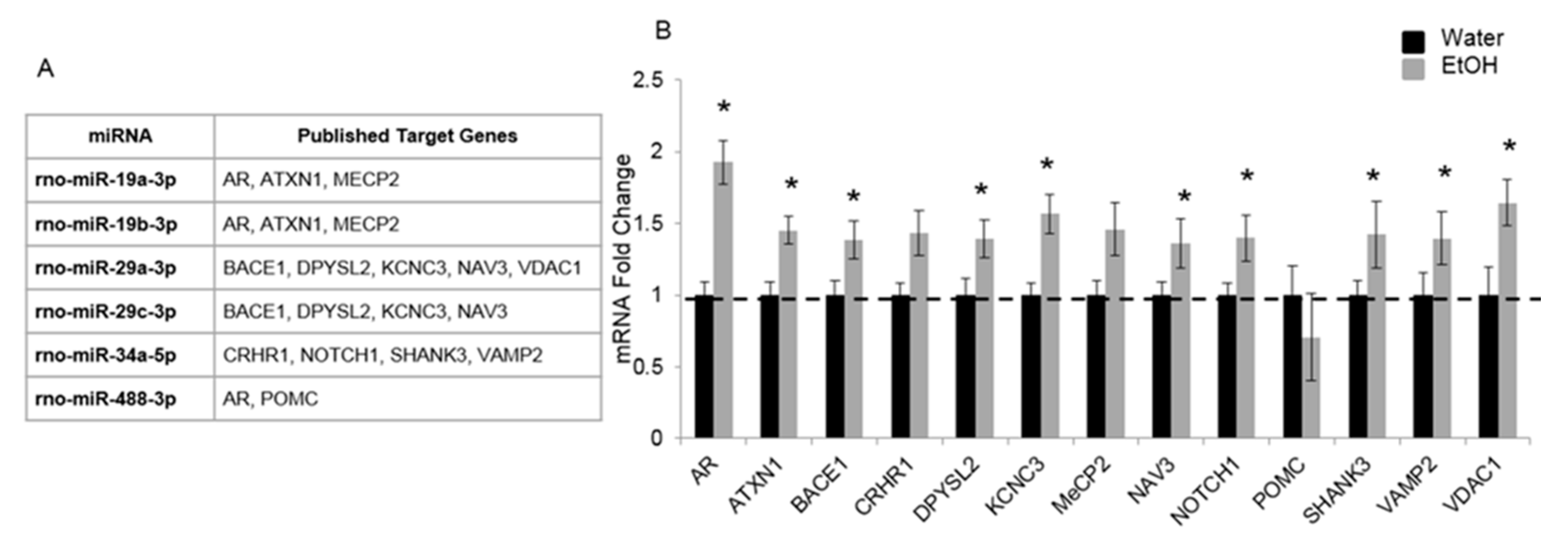
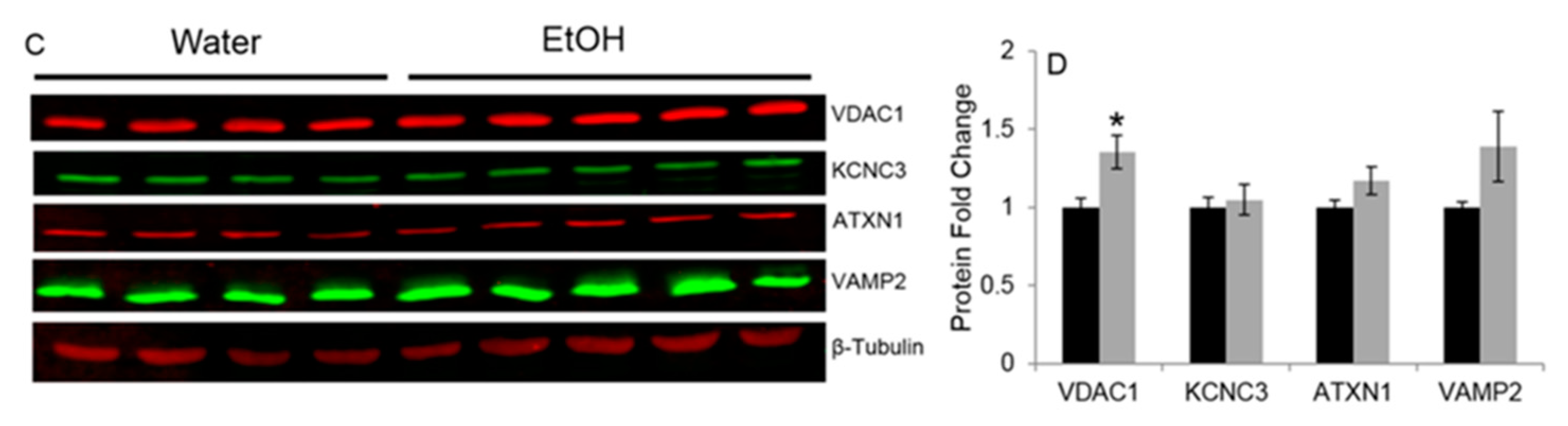
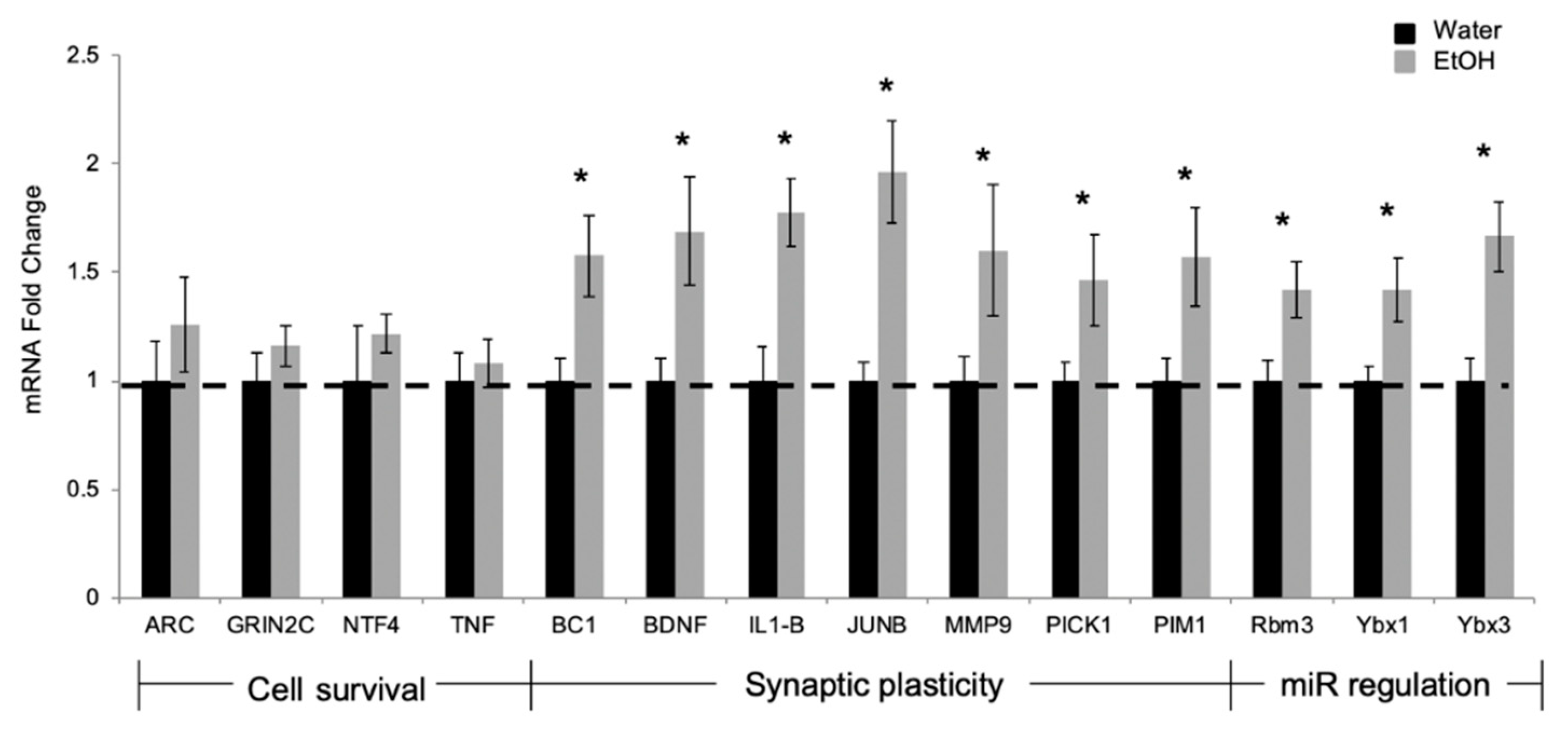
2.2. Experiment 2: Repeated Binge Alcohol during Pubertal Development Alters the Neurodevelopmental Gene Network in the Ventral Hippocampus
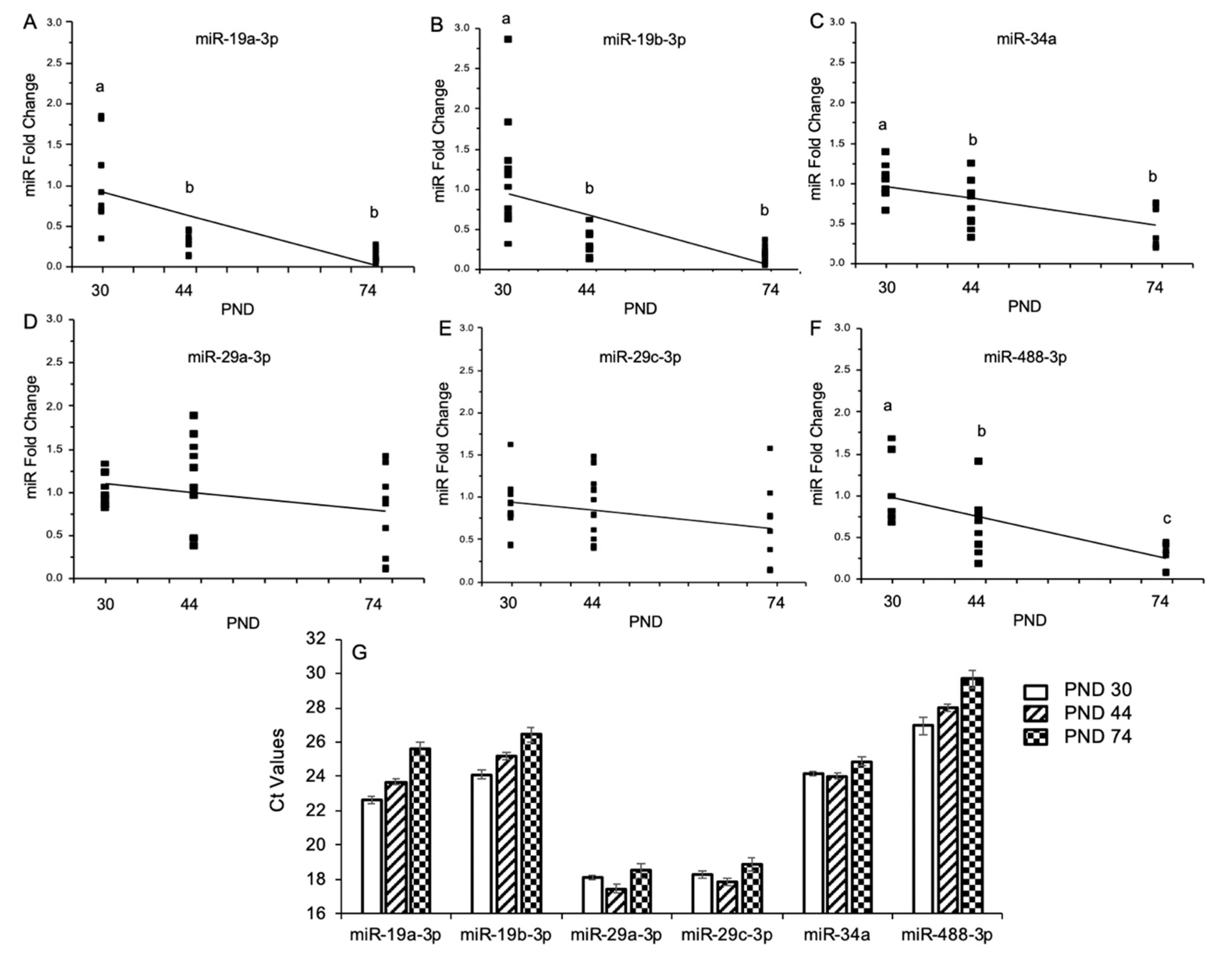
2.3. Experiment 3: Temporal Regulation of Alcohol-Sensitive miRs Through Puberty
3. Discussion
4. Materials and Methods
4.1. Animals
4.1.1. Experiment 1: Normal Pubertal Development (N = 10/age group)
4.1.2. Experiment 2: Repeated Binge-Pattern Alcohol Exposure Paradigm (N = 10/treatment)
4.1.3. Experiment 3: Normal Pubertal miR Expression Patterns (N = 10/age group)
4.2. Tissue Collection
4.3. Hormone Measurements
4.4. Testes Histology and Analysis
4.5. RNA Isolation and Reverse Transcription
4.6. Gene Expression Profiling with Microarray
4.7. Reverse Transcription Quantitative PCR (RT-qPCR)
4.8. Western Blots
4.9. Statistics
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- White, A.; Hingson, R. The Burden of Alcohol Use: Excessive Alcohol Consumption and Related Consequences among College Students. Alcohol Res. 2013, 35, 201–218. [Google Scholar] [PubMed]
- Miller, J.W.; Naimi, T.S.; Brewer, R.D.; Jones, S.E. Binge Drinking and Associated Health Risk Behaviors among High School Students. Pediatrics 2007, 119, 76–85. [Google Scholar] [CrossRef] [PubMed]
- Smyth, A.; Teo, K.K.; Rangarajan, S.; O’Donnell, M.; Zhang, X.; Rana, P.; Leong, D.P.; Dagenais, G.; Seron, P.; Rosengren, A.; et al. Alcohol consumption and cardiovascular disease, cancer, injury, admission to hospital, and mortality: A prospective cohort study. Lancet 2015. [Google Scholar] [CrossRef]
- Song, Y.; Liu, R.; Guo, X.; Park, Y.; Wang, J.; Huang, X.; Hollenbeck, A.; Blair, A.; Chen, H. Alcohol Consumption, Types of Alcohol, and Parkinson’s Disease. PLoS ONE 2013, 8, e66452. [Google Scholar] [CrossRef]
- Vargas, W.M.; Bengston, L.; Gilpin, N.W.; Whitcomb, B.W.; Richardson, H.N. Alcohol Binge Drinking during Adolescence or Dependence during Adulthood Reduces Prefrontal Myelin in Male Rats. J. Neurosci. 2014, 34, 14777–14782. [Google Scholar] [CrossRef] [PubMed]
- Risher, M.-L.; Fleming, R.L.; Risher, W.C.; Miller, K.M.; Klein, R.C.; Wills, T.; Acheson, S.K.; Moore, S.D.; Wilson, W.A.; Eroglu, C.; et al. Adolescent Intermittent Alcohol Exposure: Persistence of Structural and Functional Hippocampal Abnormalities into Adulthood. Alcohol. Clin. Exp. Res. 2015, 39, 989–997. [Google Scholar] [CrossRef] [PubMed]
- Przybycien-Szymanksa, M.M.; Rao, Y.S.; Pak, T.R. Binge-pattern alcohol exposure during puberty induces sexually dimorphic changes in genes regulating the HPA axis. Am. J. Physiol. Endocrinol. Metab. 2010, 298, E320–E328. [Google Scholar] [CrossRef] [PubMed]
- Prins, S.A.; Przybycien-Szymanska, M.M.; Rao, Y.S.; Pak, T.R. Long-Term Effects of Peripubertal Binge EtOH Exposure on Hippocampal microRNA Expression in the Rat. PLoS ONE 2014, 9, e83166. [Google Scholar] [CrossRef] [PubMed]
- Torcaso, A.; Asimes, A.; Meagher, M.; Pak, T.R. Adolescent binge alcohol exposure increases risk assessment behaviors in male Wistar rats after exposure to an acute psychological stressor in adulthood. Psychoneuroendocrinology 2017, 76, 154–161. [Google Scholar] [CrossRef] [PubMed]
- Asimes, A.; Torcaso, A.; Pinceti, E.; Kim, C.K.; Zeleznik-Le, N.J.; Pak, T.R. Adolescent binge-pattern alcohol exposure alters genome-wide DNA methylation patterns in the hypothalamus of alcohol-naïve male offspring. Alcohol 2017, 60, 179–189. [Google Scholar] [CrossRef] [PubMed]
- Asimes, A.; Kim, C.K.; Cuarenta, A.; Auger, A.P.; Pak, T.R. Binge Drinking and Intergenerational Implications: Parental Preconception Alcohol Impacts Offspring Development in Rats. J. Endocr. Soc. 2018, 2, 672–686. [Google Scholar] [CrossRef] [PubMed]
- Przybycien-Szymanska, M.M.; Mott, N.N.; Pak, T.R. Alcohol Dysregulates Corticotropin-Releasing-Hormone (CRH) Promoter Activity by Interfering with the Negative Glucocorticoid Response Element (nGRE). PLoS ONE 2011, 6, e26647. [Google Scholar] [CrossRef] [PubMed]
- Schaefer, A.; O’Carroll, D.; Tan, C.L.; Hillman, D.; Sugimori, M.; Llinas, R.; Greengard, P. Cerebellar neurodegeneration in the absence of microRNAs. J. Exp. Med. 2007, 204, 1553–1558. [Google Scholar] [CrossRef] [PubMed]
- Giraldez, A.J.; Cinalli, R.M.; Glasner, M.E.; Enright, A.J.; Thomson, J.M.; Baskerville, S.; Hammond, S.M.; Bartel, D.P.; Schier, A.F. MicroRNAs regulate brain morphogenesis in zebrafish. Science 2005, 308, 833–838. [Google Scholar] [CrossRef] [PubMed]
- Delaloy, C.; Liu, L.; Lee, J.A.; Su, H.; Shen, F.; Yang, G.Y.; Young, W.L.; Ivey, K.N.; Gao, F.B. MicroRNA-9 coordinates proliferation and migration of human embryonic stem cell-derived neural progenitors. Cell Stem Cell 2010, 6, 323–335. [Google Scholar] [CrossRef] [PubMed]
- Giusti, S.A.; Vogl, A.M.; Brockmann, M.M.; Vercelli, C.A.; Rein, M.L.; Trumbach, D.; Wurst, W.; Cazalla, D.; Stein, V.; Deussing, J.M.; et al. MicroRNA-9 controls dendritic development by targeting REST. Elife 2014, 3, e02755. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Luo, M.; Ni, N.; Den, Y.; Xia, J.; Chen, J.; Ji, J.; Zhou, X.; Fan, X.; Gu, P. Reciprocal actions of microRNA-9 and TLX in the proliferation and differentiation of retinal progenitor cells. Stem Cells Dev. 2014, 23, 2771–2781. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Kurokawa, D.; Nakao, H.; Ohmura, T.; Aizawa, S. MicroRNA-9 modulates Cajal-Retzius cell differentiation by suppressing Foxg1 expression in mouse medial pallium. J. Neurosci. 2008, 28, 10415–10421. [Google Scholar] [CrossRef] [PubMed]
- Shibata, M.; Nakao, H.; Kiyonari, H.; Abe, T.; Aizawa, S. MicroRNA-9 regulates neurogenesis in mouse telencephalon by targeting multiple transcription factors. J. Neurosci. 2011, 31, 3407–3422. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Sun, G.; Li, S.; Shi, Y. A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nat. Struct. Mol. Biol. 2009, 16, 365–371. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Sun, G.; Ye, P.; Li, S.; Shi, Y. MicroRNA let-7d regulates the TLX/microRNA-9 cascade to control neural cell fate and neurogenesis. Sci. Rep. 2013, 3, 1329. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Lin, Q.; Kim, K.J.; Dardashti, F.D.; Kim, J.; He, F.; Sun, Y. Ngn1 inhibits astrogliogenesis through induction of miR-9 during neuronal fate specification. Elife 2015, 4, e06885. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J.E.; Lee, P.R.; Chen, S.; Li, W.; Fields, R.D. MicroRNA regulation of homeostatic synaptic plasticity. Proc. Natl. Acad. Sci. USA 2011, 108, 11650–11655. [Google Scholar] [CrossRef] [PubMed]
- Edbauer, D.; Neilson, J.R.; Foster, K.A.; Wang, C.F.; Seeburg, D.P.; Batterton, M.N.; Tada, T.; Dolan, B.M.; Sharp, P.A.; Sheng, M. Regulation of synaptic structure and function by FMRP-associated microRNAs miR-125b and miR-132. Neuron 2010, 65, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Fiore, R.; Rajman, M.; Schwale, C.; Bicker, S.; Antoniou, A.; Bruehl, C.; Draguhn, A.; Schratt, G. MiR-134-dependent regulation of Pumilio-2 is necessary for homeostatic synaptic depression. Embo J. 2014, 33, 2231–2246. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Wang, W.Y.; Mao, Y.W.; Graff, J.; Guan, J.S.; Pan, L.; Mak, G.; Kim, D.; Su, S.C.; Tsai, L.H. A novel pathway regulates memory and plasticity via SIRT1 and miR-134. Nature 2010, 466, 1105–1109. [Google Scholar] [CrossRef] [PubMed]
- Lambert, T.J.; Storm, D.R.; Sullivan, J.M. MicroRNA132 modulates short-term synaptic plasticity but not basal release probability in hippocampal neurons. PLoS ONE 2010, 5, e15182. [Google Scholar] [CrossRef] [PubMed]
- Luikart, B.W.; Bensen, A.L.; Washburn, E.K.; Perederiy, J.V.; Su, K.G.; Li, Y.; Kernie, S.G.; Parada, L.F.; Westbrook, G.L. miR-132 mediates the integration of newborn neurons into the adult dentate gyrus. PLoS ONE 2011, 6, e19077. [Google Scholar] [CrossRef] [PubMed]
- Manakov, S.A.; Grant, S.G.; Enright, A.J. Reciprocal regulation of microRNA and mRNA profiles in neuronal development and synapse formation. BMC Genom. 2009, 10, 419. [Google Scholar] [CrossRef] [PubMed]
- Remenyi, J.; van den Bosch, M.W.; Palygin, O.; Mistry, R.B.; McKenzie, C.; Macdonald, A.; Hutvagner, G.; Arthur, J.S.; Frenguelli, B.G.; Pankratov, Y. miR-132/212 knockout mice reveal roles for these miRNAs in regulating cortical synaptic transmission and plasticity. PLoS ONE 2013, 8, e62509. [Google Scholar] [CrossRef] [PubMed]
- Schratt, G.M.; Tuebing, F.; Nigh, E.A.; Kane, C.G.; Sabatini, M.E.; Kiebler, M.; Greenberg, M.E. A brain-specific microRNA regulates dendritic spine development. Nature 2006, 439, 283–289. [Google Scholar] [CrossRef] [PubMed]
- Scott, H.L.; Tamagnini, F.; Narduzzo, K.E.; Howarth, J.L.; Lee, Y.B.; Wong, L.F.; Brown, M.W.; Warburton, E.C.; Bashir, Z.I.; Uney, J.B. MicroRNA-132 regulates recognition memory and synaptic plasticity in the perirhinal cortex. Eur. J. Neurosci. 2012, 36, 2941–2948. [Google Scholar] [CrossRef] [PubMed]
- Dohler, K.; Wuttke, W. Changes with Age in Levels of Serum Gonadotropins, Prolactin, and Gonadal Steroid in Prepubertal Male and Female Rats. Endocrinology 1974, 97, 898–907. [Google Scholar] [CrossRef] [PubMed]
- Dresios, J.; Aschrafi, A.; Owens, G.C.; Vanderklish, P.W.; Edelman, G.M.; Mauro, V.P. Cold stress-induced protein Rbm3 binds 60S ribosomal subunits, alters microRNA levels, and enhances global protein synthesis. Proc. Natl. Acad. Sci. USA 2005, 102, 1865–1870. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.L.; Fu, X.; Huang, J.; Jia, T.T.; Zong, F.Y.; Mu, S.R.; Zhu, H.; Yan, Y.; Qiu, S.; Wu, Q.; et al. Genome-wide analysis of YB-1-RNA interactions reveals a novel role of YB-1 in miRNA processing in glioblastoma multiforme. Nucleic Acids Res. 2015, 43, 8516–8528. [Google Scholar] [CrossRef] [PubMed]
- Messina, A.; Langlet, F.; Chachlaki, K.; Roa, J.; Rasika, S.; Jouy, N.; Gallet, S.; Gaytan, F.; Parkash, J.; Tena-Sempere, M.; et al. A microRNA switch regulates the rise in hypothalamic GnRH production before puberty. Nat. Neurosci. 2016, 19, 835. [Google Scholar] [CrossRef] [PubMed]
- Zanato, V.; Martins, M.; Anselmo-Franci, J.; Petenusci, S.; Lamano-Carvalho, T. Sexual development of male Wistar rats. Braz. J. Med. Biol. Res. 1994, 27, 1273–1280. [Google Scholar] [PubMed]
- Picut, C.; Remick, A.; de Rijk, E.; Simons, M.; Stump, D.; Parker, G. Postnatal development of the testis in the rat: morphologic study and correlation of morphology to neuroendocrine parameters. Toxicol. Pathol. 2015, 43, 326–342. [Google Scholar] [CrossRef] [PubMed]
- Mollinari, C.; Racaniello, M.; Berry, A.; Pieri, M.; de Stefano, M.C.; Cardinale, A.; Zona, C.; Cirulli, F.; Garaci, E.; Merlo, D. miR-34a regulates cell proliferation, morphology and function of newborn neurons resulting in improved behavioural outcomes. Cell Death Dis. 2015, 6, e1622. [Google Scholar] [CrossRef] [PubMed]
- Morgado, A.L.; Xavier, J.M.; Dionisio, P.A.; Ribeiro, M.F.; Dias, R.B.; Sebastiao, A.M.; Sola, S.; Rodrigues, C.M. MicroRNA-34a Modulates Neural Stem Cell Differentiation by Regulating Expression of Synaptic and Autophagic Proteins. Mol. Neurobiol. 2015, 51, 1168–1183. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.; Jakel, L.; Bruinsma, I.B.; Claassen, J.A.; Kuiperij, H.B.; Verbeek, M.M. MicroRNA-29a Is a Candidate Biomarker for Alzheimer’s Disease in Cell-Free Cerebrospinal Fluid. Mol. Neurobiol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulou, A.S.; Serneels, L.; Achsel, T.; Mandemakers, W.; Callaerts-Vegh, Z.; Dooley, J.; Lau, P.; Ayoubi, T.; Radaelli, E.; Spinazzi, M.; et al. Deficiency of the miR-29a/b-1 cluster leads to ataxic features and cerebellar alterations in mice. Neurobiol. Dis. 2015, 73, 275–288. [Google Scholar] [CrossRef] [PubMed]
- Roshan, R.; Shridhar, S.; Sarangdhar, M.A.; Banik, A.; Chawla, M.; Garg, M.; Singh, V.P.; Pillai, B. Brain-specific knockdown of miR-29 results in neuronal cell death and ataxia in mice. Rna 2014, 20, 1287–1297. [Google Scholar] [CrossRef] [PubMed]
- Zong, Y.; Yu, P.; Cheng, H.; Wang, H.; Wang, X.; Liang, C.; Zhu, H.; Qin, Y.; Qin, C. miR-29c regulates NAV3 protein expression in a transgenic mouse model of Alzheimer’s disease. Brain Res. 2015, 1624, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Kim, H.J.; Schafer, S.T.; Paquola, A.; Clemenson, G.D.; Toda, T.; Oh, J.; Pankonin, A.R.; Lee, B.S.; Johnston, S.T.; et al. Functional Implications of miR-19 in the Migration of Newborn Neurons in the Adult Brain. Neuron 2016, 89, 79–89. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Baek, M.; Gusev, Y.; Brackett, D.J.; Nuovo, G.J.; Schmittgen, T.D. Systematic evaluation of microRNA processing patterns in tissues, cell lines, and tumors. Rna 2008, 14, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Labouesse, M.A.; Polesel, M.; Clementi, E.; Müller, F.; Markkanen, E.; Mouttet, F.; Cattaneo, A.; Richetto, J. MicroRNA Expression Profiling in the Prefrontal Cortex: Putative Mechanisms for the Cognitive Effects of Adolescent High Fat Feeding. Sci. Rep. 2018, 8. [Google Scholar] [CrossRef] [PubMed]
- Crews, F.; Vetreno, R.; Broadwater, M.; Robinson, D. Adolescent Alcohol Exposure Persistently Impacts Adult Neurobiology and Behavior. Pharm. Rev 2016, 68, 1074–1109. [Google Scholar] [CrossRef] [PubMed]
- Trantham-Davidson, H.; Centanni, S.; Garr, S.; New, N.; Mulholland, P.; Gass, J.; Glover, E.; Floresco, S.; Crews, F.; Krishnan, H.; et al. Binge-Like Alcohol Exposure During Adolescence Disrupts Dopaminergic Neurotransmission in the Adult Prelimbic Cortex. Neuropsychopharmacology 2017, 42, 1024–1036. [Google Scholar] [CrossRef] [PubMed]
- Crews, F.; He, J.; Hodge, C. Adolescent cortical development: a critical period of vulnerability for addiction. Pharm. Biochem. Behav. 2007, 86, 189–199. [Google Scholar] [CrossRef] [PubMed]
- Curlik, D.M.; DiFeo, G.; Shors, T.J. Preparing for adulthood: thousands upon thousands of new cells are born in the hippocampus during puberty, and most survive with effortful learning. Front. Neurosci. 2014, 8. [Google Scholar] [CrossRef] [PubMed]
- Ketelslegers, J.; Hetzel, W.; Sherins, R.; Catt, K. Developmental changes in testicular gonadotropin receptors: plasma gonadotropins and plasma testosterone in the rat. Endocrinology 1978, 103, 212–222. [Google Scholar] [CrossRef] [PubMed]
- Przybycien-Szymanska, M.M.; Rao, Y.S.; Prins, S.A.; Pak, T.R. Parental binge alcohol abuse alters F1 generation hypothalamic gene expression in the absence of direct fetal alcohol exposure. PLoS ONE 2014, 9, e89320. [Google Scholar] [CrossRef] [PubMed]
- Lauing, K.; Himes, R.; Rachwalski, M.; Strotman, P.; Callaci, J.J. Binge alcohol treatment of adolescent rats followed by alcohol abstinence is associated with site-specific differences in bone loss and incomplete recovery of bone mass and strength. Alcohol 2008, 42, 649–656. [Google Scholar] [CrossRef] [PubMed]
- Paxinos, G.; Watson, C. The Rat Brain in Stereotaxic Coordinates; Academic Press: Cambridge, MA, USA, 2013; p. 472. [Google Scholar]
- Pak, T.R.; Lynch, G.R.; Tsai, P.-S. Estrogen Accelerates Gonadal Recrudescence in Photo-Regressed Male Siberian Hamsters. Endocrinology 2002, 143, 4131–4134. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Asimes, A.; Kim, C.K.; Rao, Y.S.; Bartelt, K.; Pak, T.R. microRNA Expression Profiles in the Ventral Hippocampus during Pubertal Development and the Impact of Peri-Pubertal Binge Alcohol Exposure. Non-Coding RNA 2019, 5, 21. https://doi.org/10.3390/ncrna5010021
Asimes A, Kim CK, Rao YS, Bartelt K, Pak TR. microRNA Expression Profiles in the Ventral Hippocampus during Pubertal Development and the Impact of Peri-Pubertal Binge Alcohol Exposure. Non-Coding RNA. 2019; 5(1):21. https://doi.org/10.3390/ncrna5010021
Chicago/Turabian StyleAsimes, AnnaDorothea, Chun K. Kim, Yathindar S. Rao, Kyle Bartelt, and Toni R. Pak. 2019. "microRNA Expression Profiles in the Ventral Hippocampus during Pubertal Development and the Impact of Peri-Pubertal Binge Alcohol Exposure" Non-Coding RNA 5, no. 1: 21. https://doi.org/10.3390/ncrna5010021
APA StyleAsimes, A., Kim, C. K., Rao, Y. S., Bartelt, K., & Pak, T. R. (2019). microRNA Expression Profiles in the Ventral Hippocampus during Pubertal Development and the Impact of Peri-Pubertal Binge Alcohol Exposure. Non-Coding RNA, 5(1), 21. https://doi.org/10.3390/ncrna5010021



