1. Introduction
Laccases (EC 1.10.3.2) are multicopper oxidases (MCOs) widely distributed in nature (fungi, bacteria and plants) that catalyse the oxidation of a large variety of organic substrates (substituted phenols, aromatic amines, benzenethiols, heterocycles, etc.) coupled to the reduction of O
2 to water. They hold four catalytic copper ions in their active site, one copper type 1 (T1), one type 2 (T2) and two T3 coppers. The reducing substrate is oxidized in the T1 site and four electrons are transferred to the T2/T3 trinuclear cluster (TNC), where one molecule of oxygen is reduced to two molecules of water. The oxidation of the substrate at the T1 site and the electron transfer from the T1 site to the TNC is assisted by highly conserved residues coordinating the catalytic coppers [
1]. The T1 site is coordinated by two His and one Cys residues, while a total of eight His residues coordinate the TNC coppers. A fourth Met ligand typically binds axially the T1 copper in plant and bacterial laccases, resulting in a tetrahedral geometry, while a non-coordinating Phe or Leu occupies this position in most fungal laccases, resulting in a T1 site with trigonal geometry. The redox potential of the T1 site classifies laccases as low-redox potential (E
° < 500 mV), in bacteria and plants, and medium (E
° = 500–710 mV) or high-redox potential (E
° = 720–800 mV) in fungi [
2,
3]. Pieces of evidence support the importance of the geometry of the T1 site in tuning the redox potential of laccases [
4,
5] although this is not the only determinant [
6,
7,
8].
The high redox potential of certain laccases secreted by white-rot fungi like PM1 basidiomycete, or different species of
Trametes or
Pycnoporus [
9,
10,
11,
12,
13], expand their oxidation capabilities to a wider substrate range than their medium- and low-redox potential counterparts [
6]. For instance, oxidation of some high-redox potential mediator compounds such as 4-hydroxybenzotriazole (HBT), violuric acid or p-coumaric acid is restricted to high-redox potential laccases (HRPLs) [
14,
15]. Once oxidized by the enzyme, the redox mediators expand the laccase substrate portfolio by acting as diffusible electron shuttles between the enzyme and the oxidizable compound in the so-called laccase-mediator systems. Therefore, HRPLs are particularly relevant in biotechnology, with applications in organic chemistry, pulp & paper, food and textile industries, bioremediation, and in biosensors and biofuel cells [
16,
17]. The possibility of adjusting the intrinsic enzyme properties to the industrial requirements through protein engineering promotes the biotechnological potential of these laccases by developing tailor-made biocatalysts for specific applications [
18,
19,
20,
21].
Typically, the majority of well-characterised laccases belong to the order Polyporales. This is because Basidiomycota (Agaricomycetes) laccases have been thoroughly studied due to their participation in lignin biodegradation during wood decay, a process that is mainly carried out by white-rot Polyporales species [
22]. However, the order Agaricales contains the largest number of known saprotrophic fungal species that have diversified during evolution to colonise a variety of lignocellulosic substrates. Their unparalleled diversity of lifestyles (comprising white-rot, brown-rot, leaf-litter, grass-litter and decayed wood decomposers) have been recently correlated with changes in their enzymatic toolkits of lignocellulolytic oxidoreductases [
23]. The high evolutionary rates of class-II peroxidase, laccase, glucose–methanol–choline oxidase, unspecific peroxygenase and lytic polysaccharide monooxygenase gene families, paralleling the ecological diversification in Agaricales, support the relevance of oxidative enzymatic machinery in the evolution of saprotrophic lifestyles in this order. In particular, Agaricales species growing on forest litter, decayed wood and grass litter constitute remarkable sources of laccases, both in number and in the diversity of laccase-like types. One of these 52 Agaricomycetes genomes analysed in that study was
Agrocybe pediades, a representative Agaricales species growing on pastures and meadows (grass litter lifestyle). The fungus possesses a significant set of MCO genes, most of which are laccases [
23].
The objective of this work is to characterise in detail a novel laccase from
A. pediades specifically selected from the genome of this fungus because it was secreted under ligninolytic conditions during solid-state fermentation on wheat straw. This makes the enzyme an interesting subject of study to evaluate if it might show promising properties as a biocatalyst like other laccases secreted by white-rot Polyporales. The laccase gene was synthesized de novo and expressed in
S. cerevisiae, and the enzyme was subjected to directed evolution to improve the laccase activity detected in the culture broth. Enzyme directed evolution is a powerful protein engineering approach that mimics the main processes of natural evolution (mutagenesis and selection of the fittest variants) to obtain improved enzymes for specific purposes [
24]. The high homologous recombination frequency of
S. cerevisiae provides important advantages for enzyme directed evolution. Moreover, the yeast is a preferred host for expression and directed evolution of fungal enzymes due to its capability to perform the post-translational modifications required to secret active eukaryotic enzymes [
25,
26,
27]. Glycosylation is one of the major requirements of fungal laccases. Although glycosylation is supposed to play a crucial biological role in enzyme protection against proteolysis and in protein folding [
28,
29,
30], some of its effects are not yet fully understood and contradictory results can be found in the literature. This prompted us to study the influence of the linked sugars on the secretion, activity and stability of the recombinant
A. pediades laccase.
2. Materials and Methods
2.1. Reagents and Strains
Yeast Transformation Kit, 2,6-dimethoxyphenol (DMP), N,N-dimethyl-1,4-phenylenediamine (DMPD), 5-Hydroxyimino-2,4,6(1H,3H,5H)-pyrimidinetrione (violuric acid), Evans Blue (EB), Reactive Black 5 (RB5), aniline, p-phenylenediamine (PPD) and HBT were purchased from Sigma-Aldrich (St. Louis, MI, USA). High Pure Plasmid Isolation Kit and 2,2′azinobis (3 ethylbenzothiazoline-6 sulphonic acid) (ABTS) were obtained from ROCHE (Basel, Switzerland). Detergents: polyoxyethylene (10) tridecyl (PET), TWEEN 20 and CHAPS were purchased from Sigma-Aldrich (St. Louis, MI, USA). Phusion High-Fidelity DNA polymerase and Restriction enzymes were purchased from New England Biolabs (Ipswich, MA, USA). QIAquick gel extraction kit from Qiagen (Hilden, Germany). ZymoprepTM Yeast Plasmid Miniprep II was purchased from Zymo Research (Irvine, CA, USA). S. cerevisiae BJ5465 strain was purchased from LGC Promochem (Teddington, UK) and Mutazyme II DNA polymerase was from Agilent (Santa Clara, CA, USA). Agrocybe pediades AH40210 was obtained from the University of Alcalá Herbarium Culture Collection, Alcalá de Henares, Spain.
2.2. Culture and Media
Glucose-ammonium medium [
31], Minimal medium (MM), EB expression medium [
32] and SEM expression medium (without including ethanol) [
33] were prepared as already described. For laccase expression EB and SEM were supplemented with 4 mM and 2 mM CuSO
4, respectively.
2.3. Agrocybe pediades Secretome
The secretome of A. pediades AH40210 was collected from cultures in glucose-ammonium medium and on wheat straw as follows. The fungus was grown in 250 mL containing 50 mL of glucose-ammonium medium or 4 g of chopped wheat (Triticum aestivum) straw (particle size ~5–20 mm long × 1–3 mm wide) soaked with 10 mL of distilled water. Inoculum for both culture media consisted of 4 mL of homogenized actively growing mycelium from glucose-ammonium cultures (at 180 rpm and 28 °C) washed and resuspended in sterile distilled water. Both liquid and solid-state fermentation cultures were grown at 28 °C under static conditions in the dark. Samples (entire flasks in triplicate) were collected after 6, 14 and 43 days of incubation. Samples from fungal cultures on lignocellulose were treated with 80 mL distilled water at 180 rpm and 24 °C for 100 min. These and the samples from fungal cultures grown on glucose-ammonium medium were filtered under vacuum and the filtrates were used for proteomic analyses.
Total extracellular proteins in the above filtrates were freeze-dried, resuspended in 20 mM sodium tartrate (pH 5), and the impurities were removed by a short SDS-PAGE (10% polyacrylamide) stained with Coomassie. The protein bands were cut, destained using 50 mM ammonium bicarbonate in 50% acetonitrile and subjected to tryptic digestion [
34]. Tryptic peptides were analysed in an LTQ-Orbitrap Velos mass spectrometer coupled to an Easy-nLC 1000 HPLC system (Thermo Scientific, Waltham, MA, USA). Peptides were first loaded into a precolumn Acclaim PepMap 100 (Thermo Scientific, Waltham, MA, USA), and then eluted onto an Acclaim PepMap C18 colum (25 cm long, 75 µm inner diameter and 3 µm particle size) (Thermo Scientific, Waltham, MA, USA) using a 120 min gradient set as follows: 0–35% solvent B for 90 min, 35–45% solvent B for 10 min, 45–95% solvent B for 5 min, 95% solvent B for 10 min, 95–100% solvent B for 1 min and 100% solvent B for 4 min, at a flow rate of 250 nL/min (solvent A: 0.1% formic acid in 2% acetonitrile; solvent B: 0.1% formic acid in pure acetonitrile). Mass spectrometry (MS) analysis was performed in the Orbitrap at 30,000 (at
m/z 400) resolution using a 200–1600
m/z mass range. After the survey scan, the 15 most intense precursor ions were selected for collision-induced dissociation fragmentation in the ionic trap. Fragmentation was performed with a normalized collision energy of 35%. Charge state screening was enabled to reject unassigned and singly charged protonated ions. A dynamic exclusion time of 45 s was used to discriminate against previously selected ions.
The MS data were analysed with Proteome Discoverer (version1.4.1.14) (Thermo Scientific, Waltham, MA, USA) using standardized workflows. Acquired spectra were searched against the catalog of predicted proteins from the
A. pediades AH40210 genome, available at the JGI fungal genome portal MycoCosm (
https://mycocosm.jgi.doe.gov/Agrped1 accessed on 3 May 2021), using the SEQUEST search engine. Precursor and fragment mass tolerance were set to 10 ppm and 0.5 Da, respectively, allowing a maximum of two missed cleavages, carbamidomethylation of cysteines as a fixed modification, and methionine oxidation as a variable modification. Identified peptides were validated using a Percolator algorithm [
35] with a q-value threshold of 0.01.
2.4. Predictions and Modelling
NetNGlyc 1.0 Server at
http://www.cbs.dtu.dk/services/NetNGlyc/ accessed on 3 May 2021 was used for prediction of
N-glycosylation sites. The 3D molecular structure model of ApL was built with
Trametes trogii laccase (PDB 2HRG) as template using the Swiss-model server [
36]. Analysis of mutations and representation of the 3D protein structures were performed with PyMol. Graphical representation of amino acid frequencies was done with WebLogo [
37] and the server ESBRI [
38] was used for the evaluation of salt bridges.
2.5. Enzyme Engineering in S. cerevisiae
The coding sequence (CDS) of
A. pediades laccase (ID 823363, JGI) fused to α
9H2 signal peptide and cloned in the episomic pJRoC30 vector was obtained in a previous work [
39].
Error prone PCR (epPCR) was carried out with Mutazyme II DNA polymerase following seller recommendations and using the ExtFw sense primer and ExtRv antisense primer (
Table S1). PCR products were purified using QIAquick gel extraction kit and mutated genes were co-cloned with linearized pJRoC30 vector (BamHI/NotI) in
S. cerevisiae by IVOE [
40].
Site-directed mutagenesis were carried out using customised mutagenic primers (
Table S1) to introduce single point mutations. For each mutated site, two fragments were obtained: one with the ExtFw sense and specific mutation antisense primers and the second with the specific mutation sense and ExtRv antisense primers (
Table S1). Products were co-cloned in
S. cerevisiae as mentioned above.
Saturation Mutagenesis (SM) and Combinatorial Saturation Mutagenesis (CSM): two complementary degenerated mutagenic primers were designed for obtaining CSM libraries over 453rd and 454th or 459th and 460th positions, and SM over the fourth axial ligand position of the DM variant (
Table S1). PCR and co-cloning methodology were carried out as described above. GLUE-IT was used to determine the number of clones to be screened in order to cover all possible amino acid combinations (coverage at 95% of confidence) [
41].
2.6. Laccase Expression in Yeast Microfermentations and High-Throughput Screening of Mutant Libraries
Individual colonies from the mutagenic library were picked and transferred to 50 µL of MM in sterile 96-well plates. H1 position was not inoculated (negative control) and column 6 was inoculated with the parent type for comparison. The plates were incubated at 28 °C, 70% humidity, 200 rpm in a humidity shaker (Minitron-INFORS). After 24 h, we added 160 µL of SEM and plates were incubated for another 48 h. Then, plates were centrifugated at 1000× g, 4 °C, 10 min and 20 µL aliquots of supernatants were transferred to replica plates. Addition of 180 µL 3 mM ABTS in Citrate-phosphate (CP) buffer at pH 3 or acetate phosphate buffer pH 6 started the enzymatic reactions that were monitored in a plate reader SpectraMax M2 (Molecular Devices, Sunnyvale, CA, USA), in kinetic mode at 25 °C by the increment in absorption at 418 nm (εABTS = 36,000 M−1 cm−1). The activities of the clones were normalized to the activity of the parent type in each plate. One activity unit (U) is defined as the amount of enzyme needed to transform 1 μmol substrate/minute at room temperature.
First re-screening. Aliquots (5 µL) from selected clone cultures were inoculated in a new sterile 96-well plate together with 50 µL MM in columns 2 and 7. Well D7 was inoculated with the parent type. Columns 1 and 12 and rows A and H were not used. Plates were incubated at 28 °C, 70% humidity, 200 rpm. After 24 h, 5 µL of growth medium (columns 2 and 7) was transferred to the four adjacent wells and another 24 h incubation was performed. 160 µL of SEM medium was added and the plates were incubated again under same conditions for 48 h. Laccase activity was measured as described previously.
Second rescreening. Selected clones from first rescreening were incubated in 3 mL MM (Supplemented with glucose) at 28 °C, 200 rpm, 24 h. DH5α E. coli was transformed with plasmids isolated from MM cultures and E. coli cells grown overnight in LB + Ampicillin plates at 37 °C. Insolate plasmid from a single colony was obtained and sent to sequencing. After sequence confirmation, S. cerevisiae cells were transformed with confirmed plasmid. Five single colonies were picked and screened for each clone, as described above.
Microfermentation of
S. cerevisiae clones expressing the selected laccase variants for comparison studies were performed as described previously [
39].
2.7. Flask Scale Production of Laccase Variants
Saccharomyces cerevisiae cells transformed with selected laccase genes were inoculated in 3 mL MM and incubated for 48 h, 200 rpm, 28 °C. An aliquot of the culture was used to inoculate 10 mL MM in 100 mL flasks (final optical density OD600 = 0.3) and incubated until OD600 around 1 was reached. Then, cells were diluted to OD600 = 0.1 in 30 mL EB medium in 100 mL flasks and incubated at 28 °C or 20 °C, 200 rpm. After maximum activity was reached, cells were centrifuged (5000 rpm, 4 °C) and supernatants concentrated with Amicon Ultra Centrifugal filters (30 KDa) at 5000 rpm, 10 min.
2.8. Laccase Characterization
Enzymatic assays were carried out in 96-well plates with 100 mU/mL of non-purified laccases (activity measured with ABTS at pH 3), in a SpectraMax M2 plate reader. The pH activity profiles, enzyme thermotolerance (T50, 10 min) and pH stability assays were, as previously described, measuring laccase activity in triplicate to determine mean values and standard deviations [
39]. T50 (10 min) was defined as the temperature at which 50% initial activity is kept after 10 min incubation. Enzyme stability to the presence of solvents and halides was performed in triplicate as was described for pH stability assay [
39], but using purified enzyme incubated in 20 mM Tris-HCl pH 7 with either Tween 20, CHAPS and PET detergents at 34 mM, NaCl at 2M or 0.1% SDS (
v/v).
Twenty μL aliquots of purified enzyme solution (at 100 mU/mL activity) were added to 170 μL 50 mM CP buffer (pH 3) in the presence of a gradient of 10% to 90% (v/v) of organic solvents (acetone, ethanol and DMSO) and different concentrations of halides (0.01–0.15 mM for NaF, 10–200 mM for NaCl, 0.1–15 mM for SDS and 5–200 mM for EDTA) (96-well plates). Ten μL ABTS 60 mM were added to the mixture (three replicates per sample) and laccase activity was immediately measured for 2 min. Solvent inhibition was fitted to a sigmoidal function and halides to a biexponential decay function to calculate IC50 values.
The activities of the crude enzymes were tested by quintuplicate in 96-well plates. The assays were performed by adding 20 μL of enzyme solution with 1 U/mL laccase activity (measured with ABTS at pH 3) to 180 μL of 3 mM DMP or 9 mM guaiacol in 100 mM acetate phosphate buffer (pH 5), to 180 μL of 5 mM DMPD or PPD in 100 mM tartrate buffer (pH 4) or to 180 μL of 100 µM EB in 100 mM tartrate buffer (pH 4). Respective activities were calculated by the increments in absorbance at 469 nm (εDMP = 27,500 M
−1 cm
−1), 470 nm (εguaiacol = 26,600 M
−1 cm
−1), 550 nm (εDMPD = 4134 M
−1 cm
−1) and 450 nm (εPPD = 14,685 M
−1 cm
−1). The specific activities (U/mg) of purified enzymes were calculated measuring the increment of absorbance at 409 nm (εHBT = 321 M
−1 cm
−1) in 100 mM acetate phosphate pH 5, and 410 nm (εaniline = 1167 M
−1 cm
−1), 515 nm (εvioluric acid = 113 M
−1 cm
−1,
Figure S1) in 100 mM tartrate buffer pH 4. For EB and RB5, the activities were measured by the decrement in absorbance at 505 nm (εEB = 55,500 M
−1 cm
−1) and 495 nm RB5 (εRB5 = 22,500 M
−1 cm
−1) in 100 mM tartrate buffer pH 4.
Kinetic constants were determined as previously described [
42] using a range from 0.00125 mM to 0.1 mM for ABTS and DMP and 0.00125 mM to 2 mM for DMPD, and the following buffers: 100 mM CP buffer for assays at pH 3, 100 mM tartrate for pH 4 and 100 mM acetate phosphate for pH 5. To calculate
KM and
kcat values the average Vmax was represented versus substrate concentration and fitted to a single rectangular hyperbola function in SigmaPlot (version 14.0) software for DMP and DMPD, and fitted to a Hill-sigmoidal for ABTS, where parameter a was equal to
kcat and parameter b was equal to
KM. In all assays three replicates of each laccase variant were used.
2.9. Purification of Laccase Variants
Crude enzymes were filtered with 0.22 µm pore size membrane and concentrated and ultra-diafiltrated using Pellicon cassettes (Merck Millipore, Darmstadt, Germany) and Amicon stirred cells (Merck Millipore, Darmstadt, Germany), both with a 10 kDa cutoff. The concentrated solution was dialyzed against 20 mM TrisHCl buffer pH 7 and immediately concentrated by Amicon Ultra (10 KDa) tubes at 5000 rpm, 10 min. Laccases were purified by FPLC (AKTA purifier system, GE Healthcare) in three steps: (i) anion exchange HiPrep QFF 16/10 column (GE Healthcare) using a salt gradient 0–40% (20 mM Tris-HCl, 1M NaCl, pH 7) to elute the enzyme; (ii) anion exchange Mono Q 5/50 GL column (GE Healthcare), using a 0–25% salt gradient (20 mM Tris-HCl, 1M NaCl, pH 7) to elute the laccase; (iii) molecular exclusion column HiLoad 16/600 Superdex 75pg (GE Healthcare). Between each purification step, fractions with laccase activity were collected, dialyzed and concentrated using Amicon® Ultra (10 KDa) tubes. The purity of the enzyme was estimated in SDS-PAGE and N-deglycosylation was performed with Endoglycosidase H following manufacturer’s instructions. Additionally, a chromatographic affinity step with HiTrap Con A 4B column (GE Healthcare) was added for evaluating NGly laccases variants. The eluent buffer was 20 mM Tris-HCl, 0.5 M NaCl, 1M Glucose, pH 7.
2.10. Protein Quantification
Protein quantification was calculated by the Qubit 3.0 fluorometer of Sigma-Aldrich (St. Louis, MI, USA).
4. Discussion
Agrocybe pediades is a representative Agaricales species that grows on pastures and meadows (grass-litter lifestyle). Although its genome encodes 11 different laccases, all classified as sensu stricto [
23], only the laccase named ApL in this study was secreted by the fungus under ligninolytic conditions (solid-state fermentation of wheat straw). We addressed the functional expression, engineering and characterization of this enzyme using
S. cerevisiae as a heterologous expression system. The yeast was chosen since it is the preferred platform for the directed evolution of fungal laccases and has provided successful secretion of active enzymes with interesting properties as biocatalysts [
20,
32,
47].
In a previous work, we demonstrated the capability of the evolved α
9H2 leader [
20] to improve the secretion by
S. cerevisiae of several fungal laccases (ApL included), compared to different signal peptides derived from the α-factor preproleader [
39]. Here, the use of α
9H2 leader allowed us to achieve the functional expression of ApL in the yeast, although the levels obtained were insufficient for a deep characterization of the enzyme. To increase laccase production, and simultaneously to give rise to an improved version of the enzyme as a biocatalyst, the laccase was first subjected to a round of directed evolution through random mutagenesis over the α
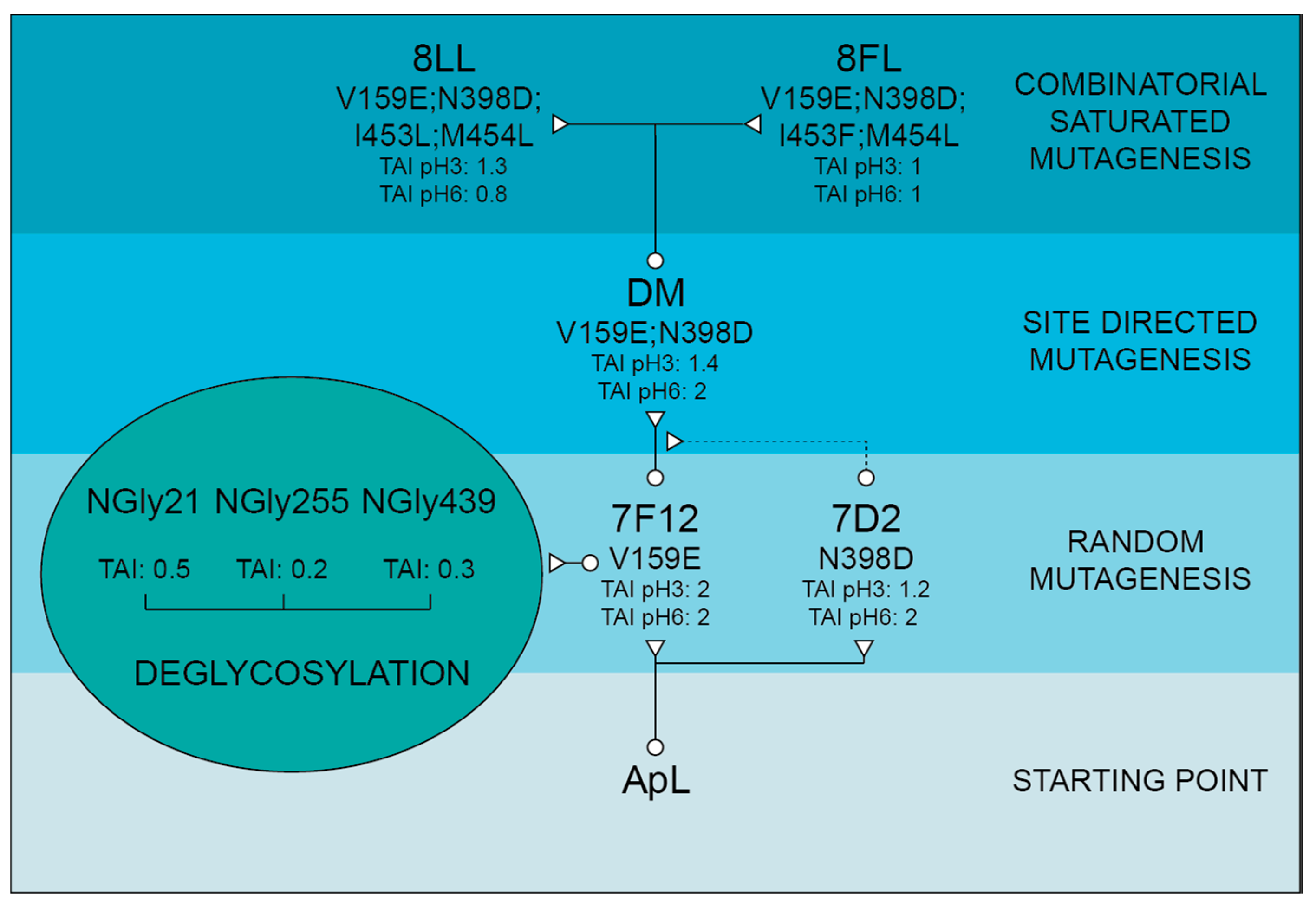
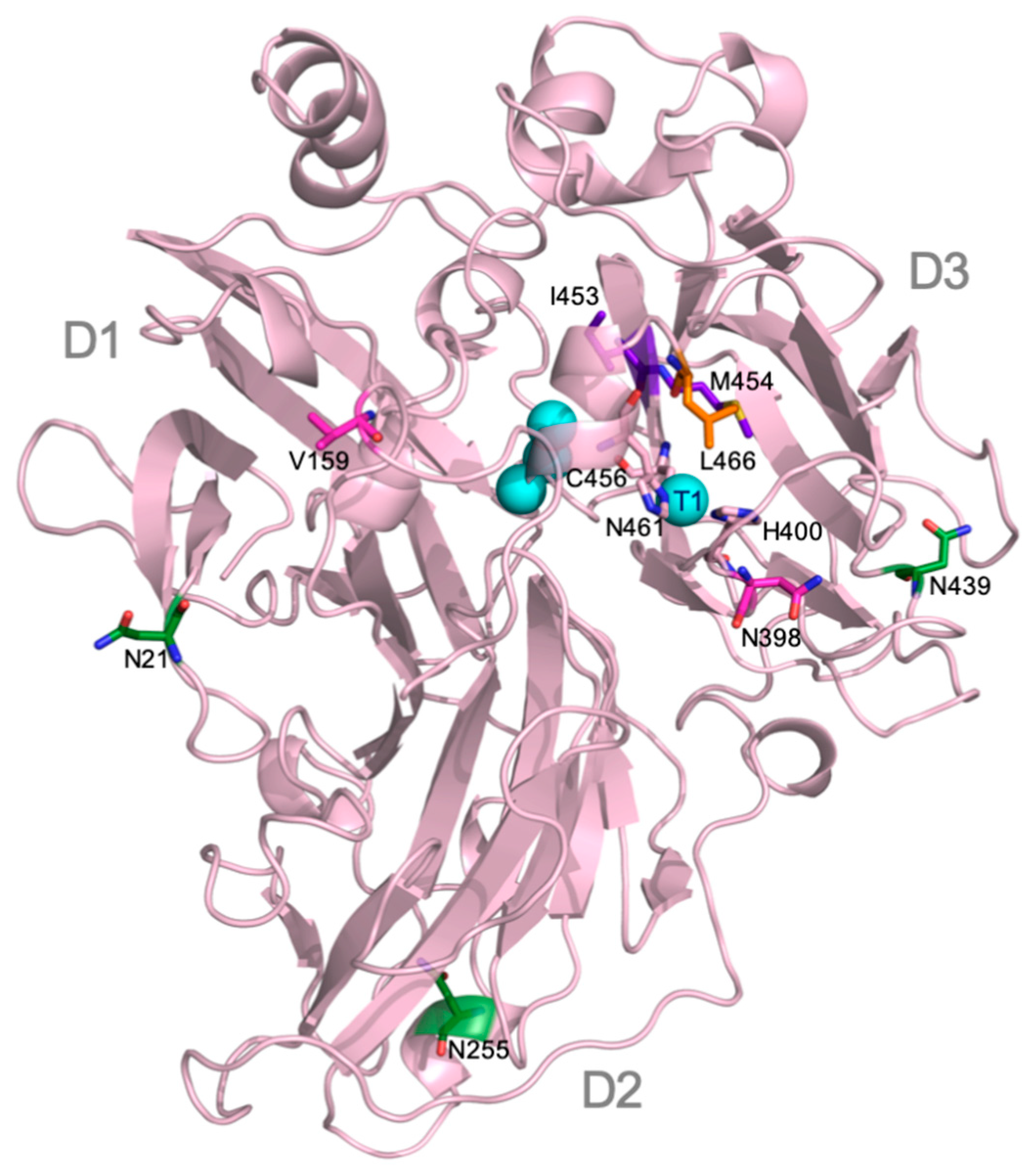
9H2-ApL construction. Mutation V159E selected in 7F12 variant produced two-fold TAI in microfermentation, without affecting the optimum pH of the enzyme. The location of V159E mutation at the protein surface, far away from the T1 site (
Figure 2), together with the similar activities for different substrates of native ApL and 7F12 variant, evidenced an enhancement of enzyme production by this mutation. Analysis of the structure model of ApL suggested that V159E substitution caused no changes in H bonding or electrostatic interactions (using the evaluating salt bridge server ESBRI [
38]). Although it is not possible to conclude what the exact contribution of this mutation is, it might facilitate the polypeptide maturation during expression and secretion by the yeast, in agreement with reported single-point mutations enhancing protein expression by aiding in protein flexibility during folding [
32] or stabilizing buried regions [
48].
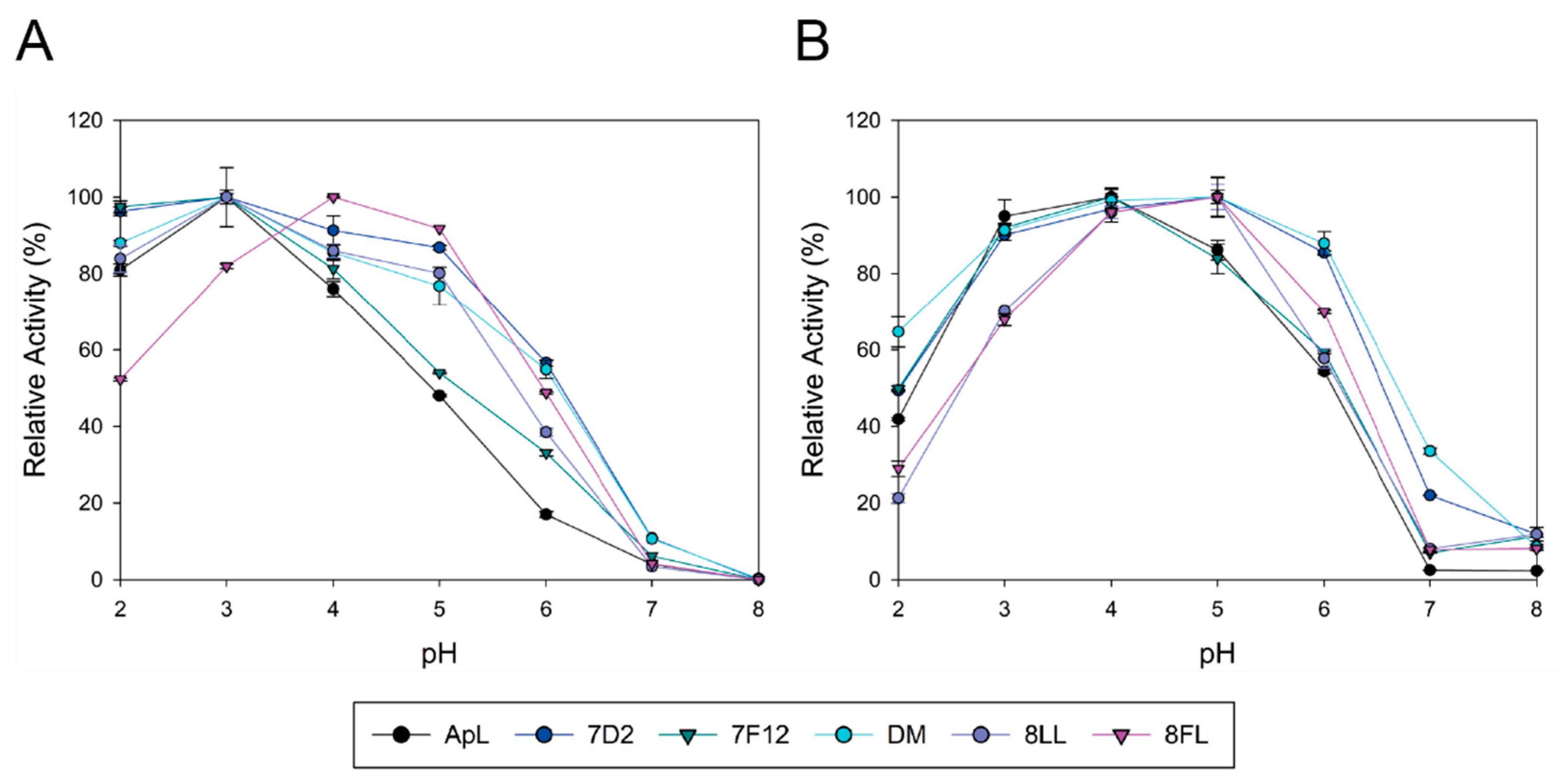
Basidiomycete laccases generally exhibit acidic pH activity profiles [
44,
49]. By contrast, ApL shows significant activity at pH 6. This constitutes an interesting property that can be enhanced in the laboratory to facilitate the applicability of the enzyme since alkaline conditions are required for industrial applications such as lignin valorization [
50,
51]. Thus, we included an activity assay at pH 6 during the screening of the mutant library to facilitate the selection of mutations enhancing the activity of the enzyme at neutral pH. As a result, 7D2 variant, holding mutation N398D, was selected with improved activity at pH 6. Based on these results and on the distance between V159E and N398D, we evaluated their possible joint effect in the DM variant (V159E; N398D). DM exhibited properties from their parents: it showed a wider pH profile towards neutral pH and notably higher laccase activity with respect to native ApL. Then, we studied the amino acids adjacent to the conserved tripeptide His-Cys-His. These residues coordinate T3 and T1 coppers, and the Cys-T1 bond has a strong influence on laccase activity [
1]. The alignment of 482 laccase sequences of 33 fungal Agaricales genomes revealed differences in conservation among the residues upstream and downstream the tripeptide. Therefore, the variable positions were randomised in the DM variant in two independent mutagenesis libraries, CSM 453–454 and CSM 459–460, using customised primers to substitute the residues of ApL by the most frequent amino acids found in the alignment. Noticeably, in the CSM 459–460 library, the pair Asp 459, Trp 460 of DM laccase was the only combination that ensured detectable laccase activity, suggesting a strong restriction in these positions for ApL (even amino acid changes of same nature are not allowed). In CSM 453–454 only two new amino acid combinations provided parental-like activities: I453L, M454L (8LL variant) and I453F, M454L (8FL variant). Mutations selected in 453 position were conservative (Ile was replaced by Leu or Phe), suggesting that a hydrophobic residue is required in this position, whereas in both mutants Met 454 was replaced by Leu. Important changes in the pH activity profiles (significant narrower and more neutral profiles with ABTS and DMP for variant 8FL) and in T50 (significantly diminished in 8LL and 8FL) indicated that the amino acids placed in these positions not only affect the pH dependence of laccase activity, but also the thermotolerance of the enzyme. Contribution of the second-shell amino acids of T1 copper to modulate laccase activity and restriction to acid pH in fungal HRPLs has been previously reported [
52,
53].
Substitution of Asn 398 by Asp stood out as the most relevant mutation for the catalytic activity of the enzyme. Asn 398 is located in a loop delimiting the entrance of the substrate binding pocket, nearby His 400 (T1 Cu ligand). Comparison of 7F12 and DM catalytic constants for the oxidation of different model compounds showed that N398D mutation is responsible for a notable superior catalytic efficiency of DM and a significant improvement of activity at neutral pH values. This is in agreement with previous results found in Ascomycete [
52] and Basidiomycete [
32,
42] laccases where substitution in positions equivalent or contiguous to ApL 398th produced an increment in laccase activity and a shifted profile to more neutral pH values. Actually, P394H substitution (equivalent to 399 ApL) increased the activity at neutral pHs of a swap-domain laccase [
42], shifted the optimal pH from 3 to 5 (with DMP) and improved the catalytic activity of an evolved
Pycnoporus cinnabarinus laccase [
54]. In the latter enzyme a new substrate binding mode increases the turnover rate associated to an enhanced stabilization of the oxidized form of certain substrates [
55].
On the other hand, the kinetic parameters of DM variant for the oxidation of model compounds are remarkable. Its catalytic efficiencies for the oxidation of ABTS and DMP are notably superior than those of other wild laccases from Agaricales like
Agaricus blazei laccase [
56] or HRPLs from Polyporales such as PM1L [
13] or
Trametes trogii laccase [
10]. In addition, the kinetic constants of DM with these two substrates are also notably superior to those of laccases from
Coprinopsis cinerea [
57] and
Pleurotus sajor-caju [
58] or
Trametes versicolor [
53] expressed in yeast. Moreover, the specific activities of the developed ApL for the oxidation of recalcitrant organic dyes or high-redox potential mediators is similar or better than those of PM1L, suggesting the enzyme possesses a high redox potential as well, although it holds a Leu as fourth “non-coordinating” axial ligand instead of a Phe typical of Polyporales HRPLs. In addition, the activity of the DM variant was not severely affected by the presence of the different inhibitors tested here. In fact, the enzyme showed high tolerance to the presence of EDTA, SDS and halides, with IC50 superior to those of other fungal laccases [
59]. The enzyme exhibited a striking activity under the presence of EDTA and organic solvents by contrast to other laccases, which in general hardly tolerate high concentration of these inhibitors [
12,
60,
61,
62]. It also exhibited outstanding stability to NaCl, whereas the tolerance to NaF was remarkably poorer. These results are in agreement with the reported strong potential of fluoride to obstruct the electron transfer in the TNC site, due to the influence that the diameter of the anion has on the inhibitory potential of the halide (Fl- > Cl- > Br-) [
19,
60,
63].
A Phe residue is fully conserved as the fourth non-coordinating axial ligand of T1 in the majority of the HRPLs crystalized so far (most of them from Polyporales). Some examples with Phe in this position are those from
Trametes versicolor [
7],
Trametes trogii [
64],
Coriolopsis caperata [
65],
P. cinnabarinus, PcL [
66] or PM1 basidiomycete [
13,
18]. Instead, ApL has a leucine as non-coordinating residue in the axial position. The correlation between the high redox potential of the T1 site and the presence of Phe at this position has been called into question because other factors like the charge distribution near T1 site [
67] or the length of the bonding between T1 and its ligands [
7] seem to exert an effect on the redox potential of laccases. In this line, a laccase from
Rigidosporus lignosus exhibited a 730 mV with a Leu [
68] or, more noticeable, an Ascomycete laccase from
Botrytis aclada with a Leu at this position also showed a considerable redox potential (720 mV) [
69]. Here we proved the oxidation of several recalcitrant substrates by DM, suggesting that this ApL variant possesses a high-redox potential comparable to that of the HRPL from PM1 basidiomycete. DM showed even superior specific activity than PM1L with violuric acid and RB5, although these differences could be attributed to substrate affinity that is determined by the size, shape or polarity of the substrate binding pocket [
18,
70]. The oxidation of the high-redox mediator violuric acid and the organic dye RB5 are of relevance for the respectively application of the enzyme in laccase-mediator systems and in the degradation of industrial dyes [
15,
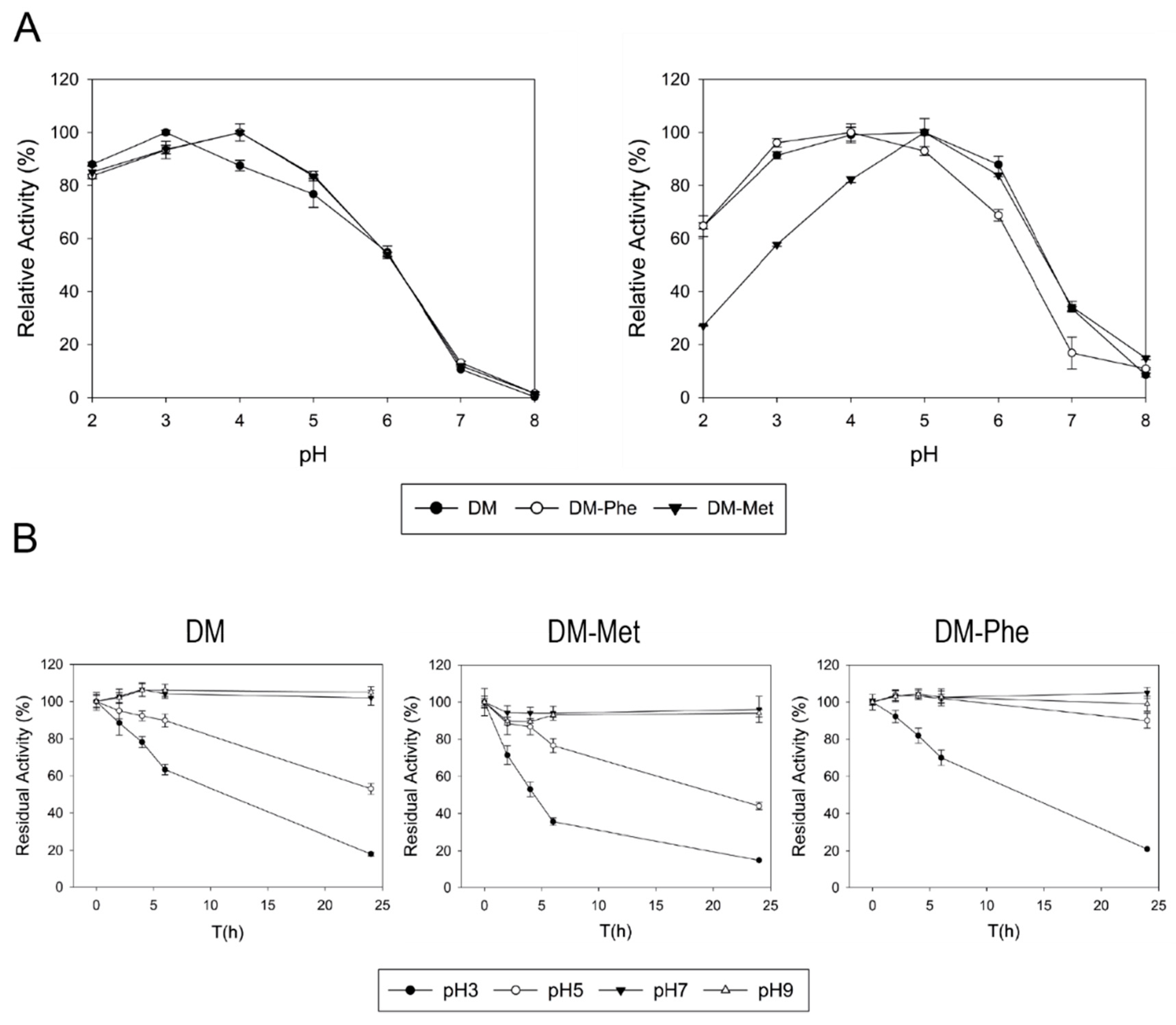
17]. Saturation mutagenesis of this position in DM variant confirmed the preference of Leu in ApL scaffold. Substitution by Met barely allowed the oxidation of guaiacol or EB dye by DM-Met variant, probably related to a decrease in the redox potential accordingly to other reports [
69]. On the other hand, while DM-Phe variant exhibited better activity with ABTS than DM, it also showed poorer oxidation for EB dye or guaiacol. Even though these differences could be attributed to the strong dependence of pH profiles on the axial ligand shown here and in other studies [
52], this does not seem to determine the oxidation of EB dye, guaiacol or PPD because the assays were carried out at pH values where both enzymes work efficiently.
Glycosylation is a major post-translational modification described to facilitate protein folding and structural stabilization or prevent from protein proteolysis [
28,
29,
71]. Most fungal laccases are glycoproteins with glycan moieties contributing to up to 25–50% of the enzyme molecular weight [
25,
32,
42,
72].
N-glycosylation is known as the predominant type of sugar anchoring in fungal laccases obtained by either homologous [
7,
65,
73] or heterologous [
42,
74,
75] expression. The
N-glycosylation patterns of Basidiomycete [
74,
76,
77] and Ascomycete [
75] laccases are clearly different. Most of these studies provide a reliable position of the sugar residues because they are based on crystallized structures, but the number of sampled laccases is still reduced, in particular of Agaricales laccases (only one crystallized [
78]). Taking all this into account, we compared the three
N-glycosylation sites of ApL (N21, N255 and N439), each respectively placed in one of the three cupredoxin domains of laccase (D1, D2 and D3) (
Figure 2), with those predicted by the NetNGlyc 1.0 Server in laccases from Polypolares (82 laccases with PcL as query sequence) and from Agaricales (482 laccases with ApL as query sequence). In Polyporales laccases, two sites, N54 and N434 (PcL numbering), located in D1 and D3, respectively, were almost fully conserved [
74,
76,
77]. On the contrary, Agaricales laccases exhibited more variability of
N-sites among the three laccase domains, although N434 (N439 in ApL numbering) stood out as the most conserved
N-glycosylation site (92% frequency). The three putative
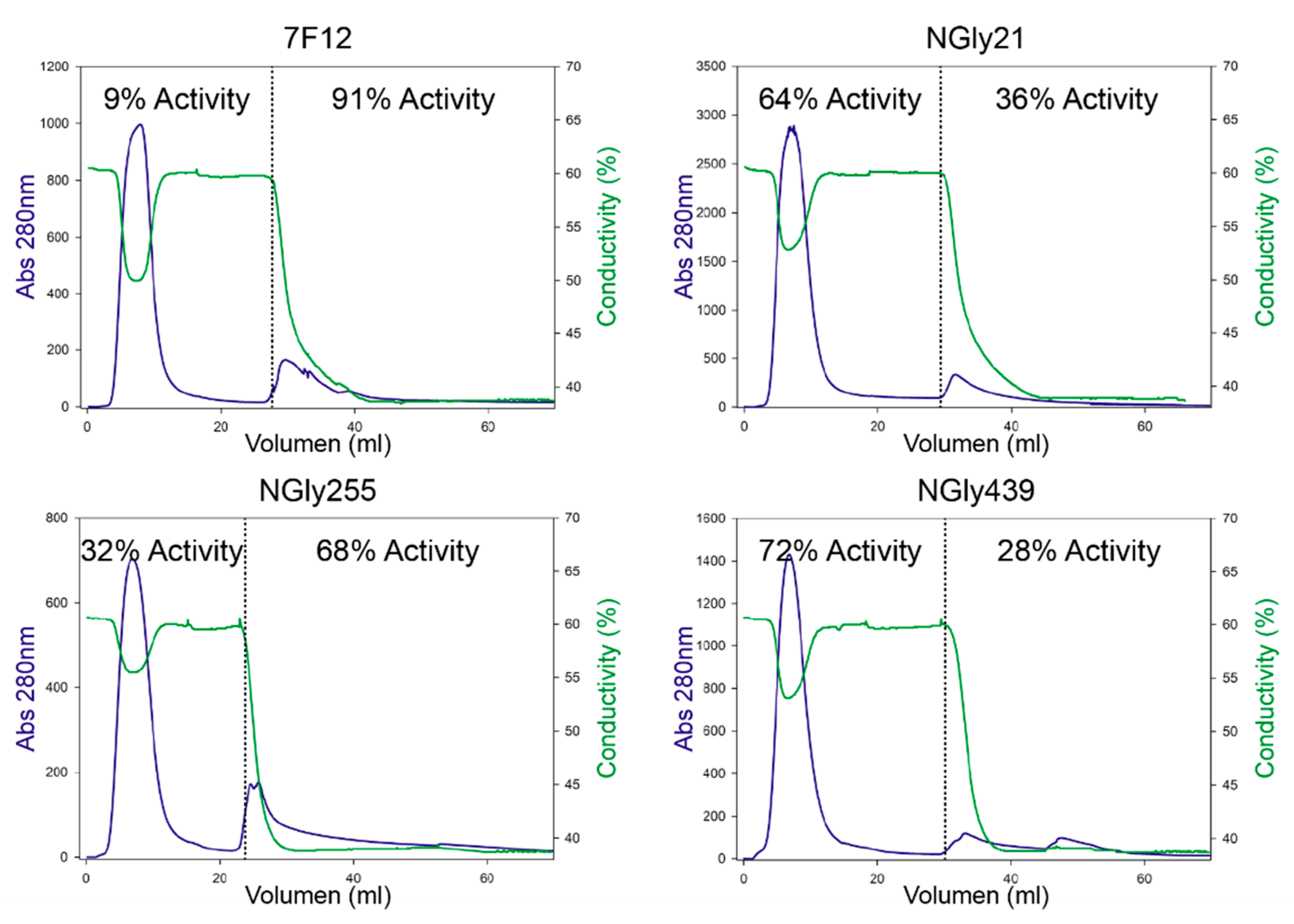
N-glycosylation sites of ApL were later proved as real sugar anchoring sites after we individually removed the three sites and the corresponding N-Gly laccase variants were subjected to concanavalin A chromatography. This allowed us to detect the partial deglycosylation of the three NGly variants, that showed dissimilar glycosylation patterns among them and different from the full glycosylated enzyme (7F12) holding the three sugar-anchoring sites.
To study if
N-glycosylation in these sites is critical for the heterologous expression and activity of ApL, the three
N-Gly variants were produced in
S. cerevisiae flask cultures and characterised. All
N-Gly variants showed diminished laccase activities in the crude extracts, especially those removing N255 and N439 sites, which barely reached a 12% and 5% of the detected parental activity. These results confirm the crucial role of both sites. The essential role of N439 had been suggested by the strict conservation of this
N-glycosylation site in Agaricales and Polyporales laccases. Besides, our results agree with those obtained with
Lentinus sp laccase where almost no activity was found when N238 and N458 sites (coinciding with N255 and N439 in ApL) were removed [
74]. The role of sugar anchoring at these positions was proposed as a mechanism to maintain the intrinsic laccase activity by stabilizing a large loop connecting the two cupredoxin domains D2–D3 [
49,
74]. Some authors reported that the lack of sugar could derived into conformational changes for substrate binding during the catalytic reaction, which is supported by the correlations found between deglycosylation of laccase and loss of activity [
73,
79,
80]. However, these studies were based on the kinetic characterization of the recombinant enzymes once secreted by the yeast and after enzymatic deglycosylation, without considering the effect that
N-glycosylation may have in early protein processing (post-translational modification, folding, secretion, etc.). Aiming at evaluating the biological role of glycosylation on the heterologous expression of ApL and/or on its catalytic activity, we purified the three
N-Gly variants specifically obtained from the removal of each site N21, N255 and N439. Comparison of their kinetic constants showed, in general, poorer activities than the parent laccase (7F12 used as reference for native glycosylated laccase) due to lower turnover rates for the oxidation of ABTS and DMP. However, this does not fully explain the outstanding decrease of activity detected in the corresponding
S. cerevisiae liquid cultures, specially at 28 °C. In fact, reduction of the temperature to 20 °C (to maximize the synthesis of correctly folded heterologous protein) raised the laccase activities detected in the culture broths and shortened the differences among
N-Gly variants and parent laccase. The latter could be attributed to an increment on enzyme secretion by the yeast [
81], avoiding harmful protein aggregation [
82]. All these pieces of information pointed out to a possible double function of
N-glycosylation in ApL, having an effect on enzyme production and activity. To evaluate both contributions, for each NGly variant and respecting the parent laccase, we compared the decrease in activity detected during 20 °C yeast fermentation and the decrease in
kcat for ABTS oxidation (under saturated substrate conditions the catalytic activity depends on this parameter). In the case of NGly21 variant both decrements were similar (around two-fold), indicating the contribution of glycosylation on N21 site only to enzyme activity. Conversely, the effect of sugars linked on N255 and N439 sites seemed to exert an effect on the catalytic activity (impaired two-fold and 1.3-fold, respectively) and, particularly, on the production of the enzyme by the yeast (the activities secreted in the culture broths were reduced by six- fold and 3.1-fold, respectively).
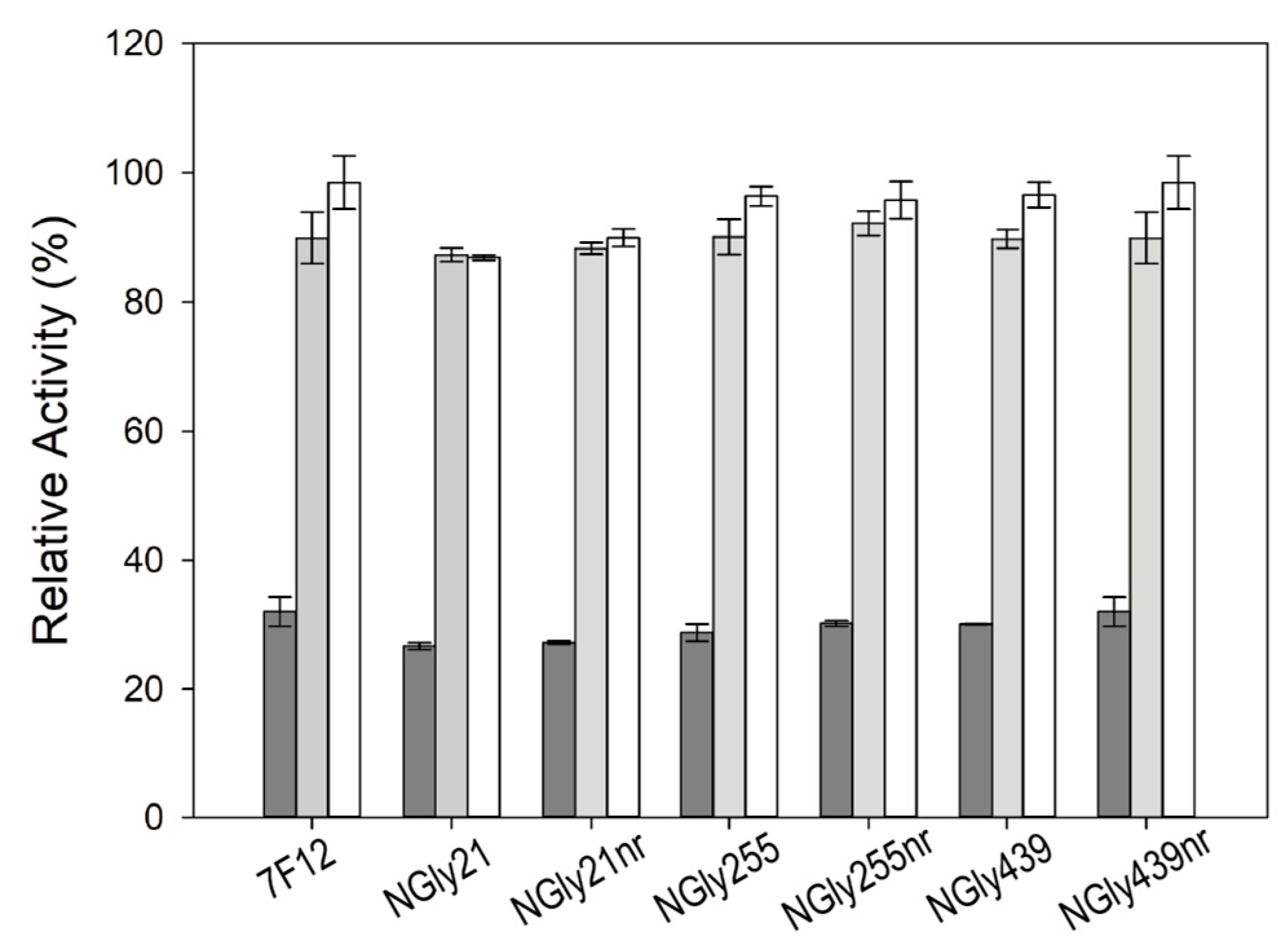
Finally, on the basis that glycoproteins with a similar degree of glycosylation could have different stability depending on where the sugar are linked [
28], we studied the stability to pH and temperature of the different (de)glycosylated fractions of the NGly variants. No major differences were found among the stability of NGly variants and parent laccase, showing no direct correlation between the glycosylation degree and the thermotolerance or stability to pHs of ApL. These data disagree with the commonly accepted role of glycosylation to enhance enzyme thermostability [
83] but are in agreement with other studies on fungal laccases where deglycosylation of the enzyme did not alter thermostability [
42,
74].