Characterization of a Candida albicans Mutant Defective in All MAPKs Highlights the Major Role of Hog1 in the MAPK Signaling Network
Abstract
1. Introduction
2. Materials and Methods
2.1. Strain Construction and Growth Conditions
2.2. Interaction Assays with Murine Macrophages
2.3. Morphogenesis Assays
3. Results
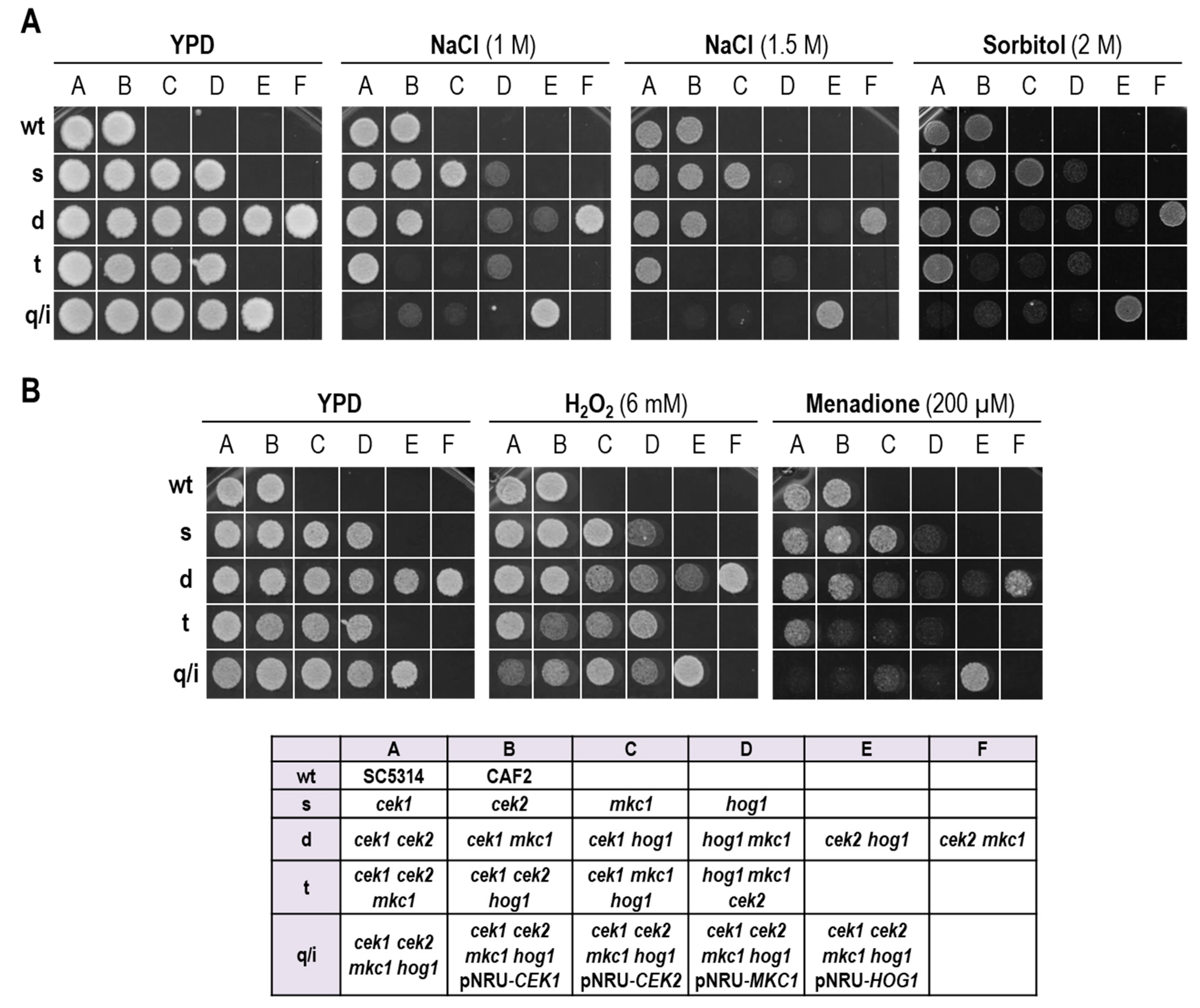
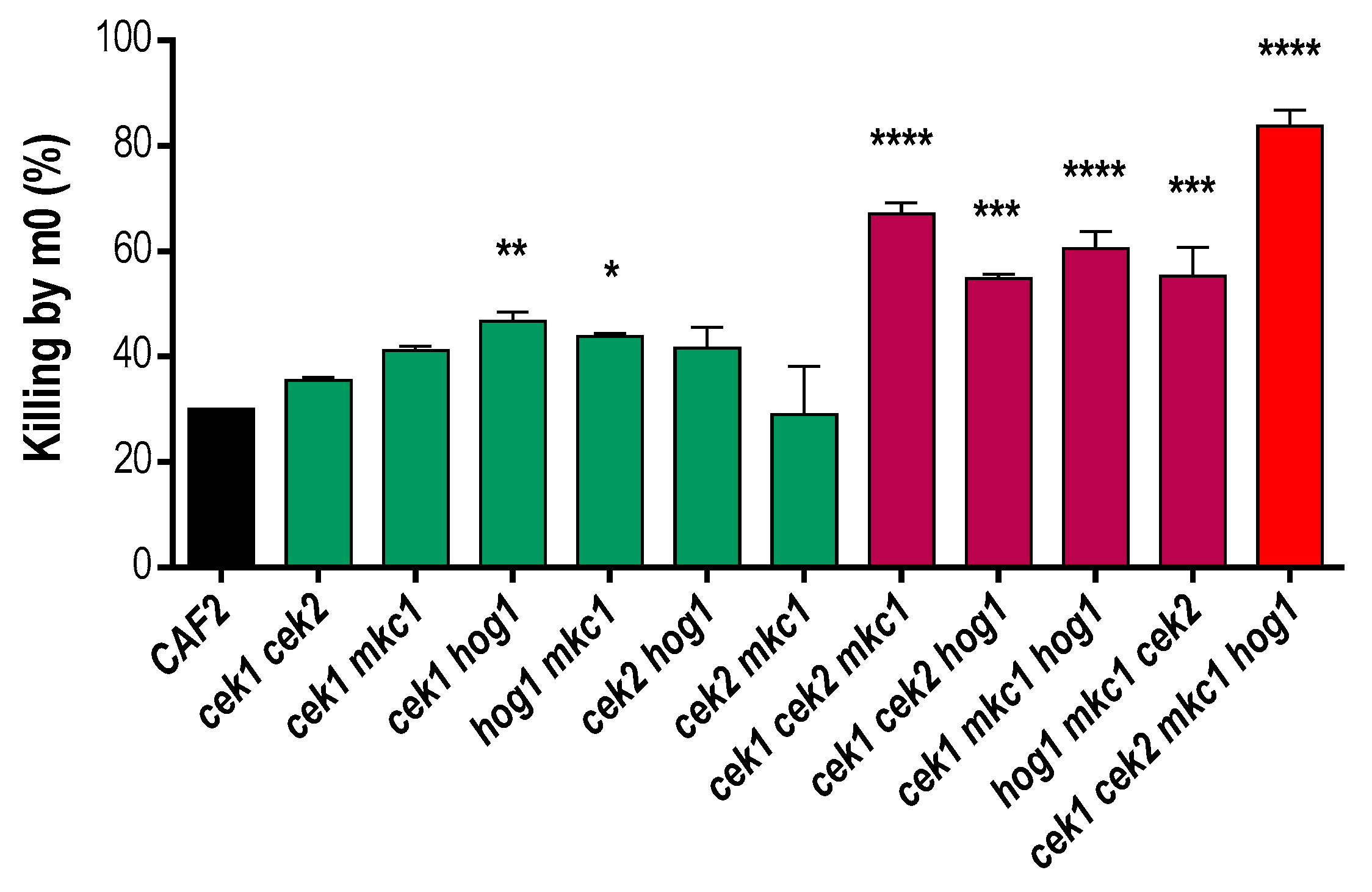
3.1. A Strain Lacking CEK1, CEK2, MKC1 and HOG1 Is Viable but Highly Sensitive to Osmotic and Oxidative Stress
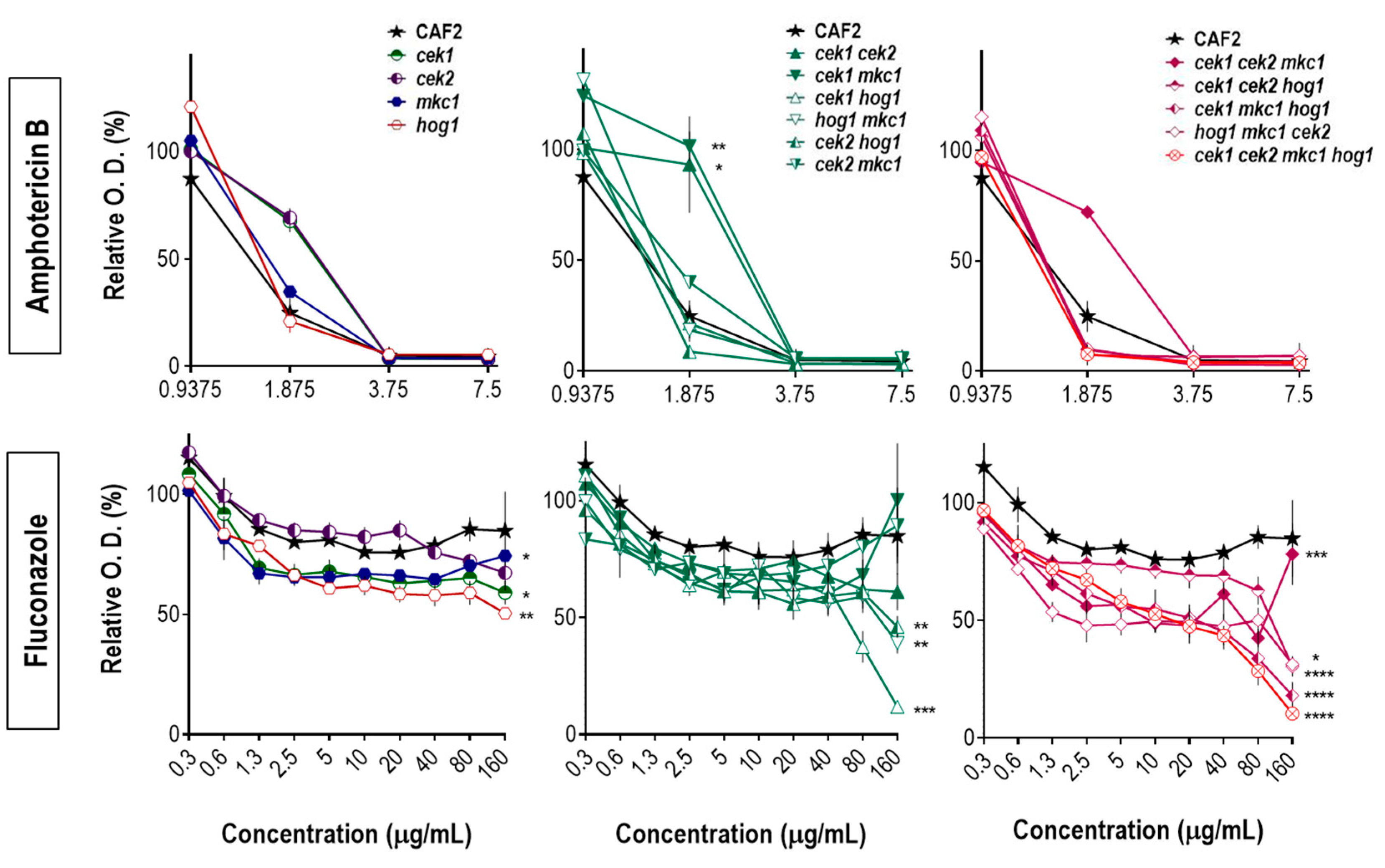
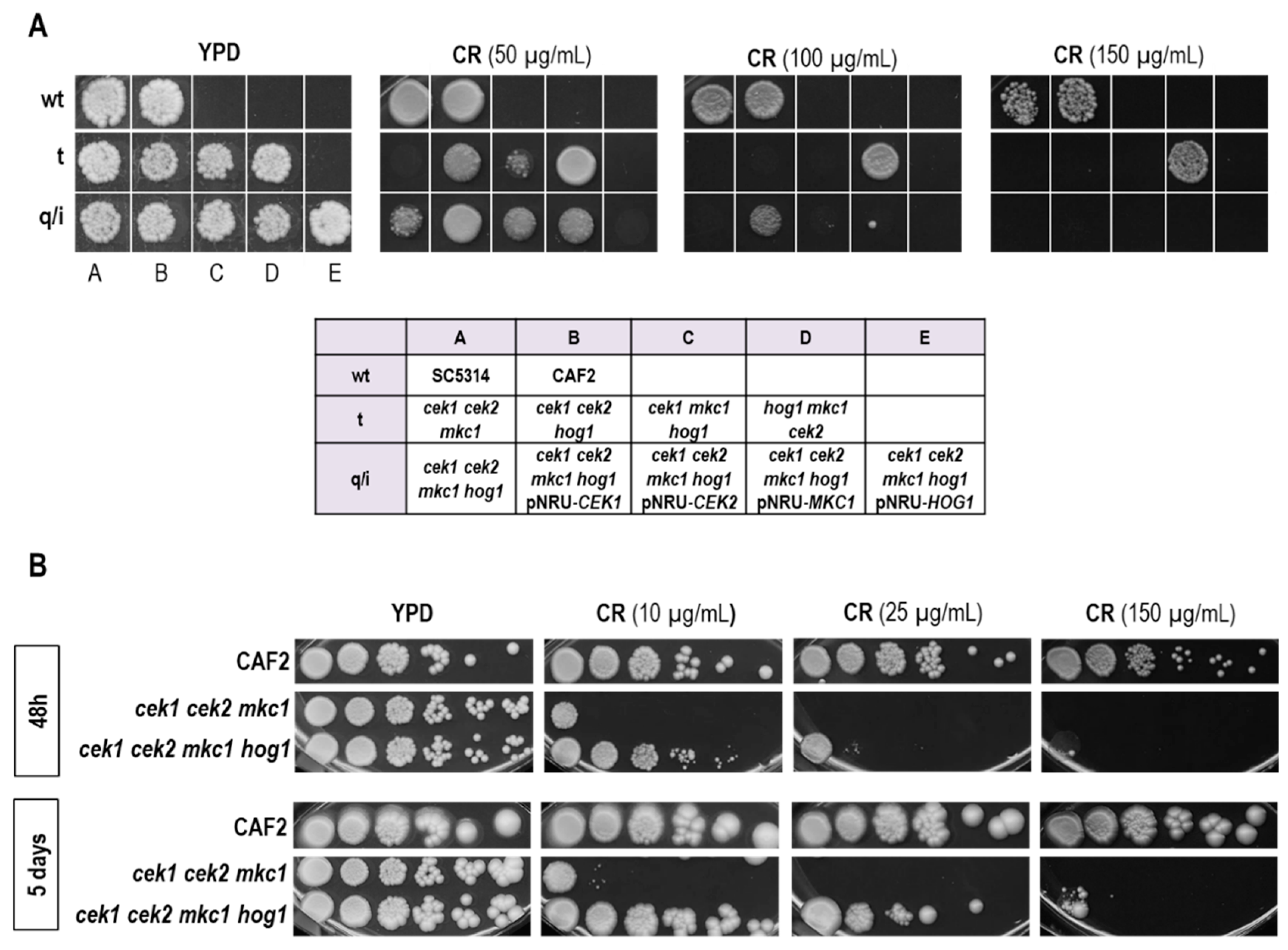
3.2. Hog1 Resistance to Cell Wall Interfering Drugs Is Not Exclusively Due to Cek1 Hyperactivation
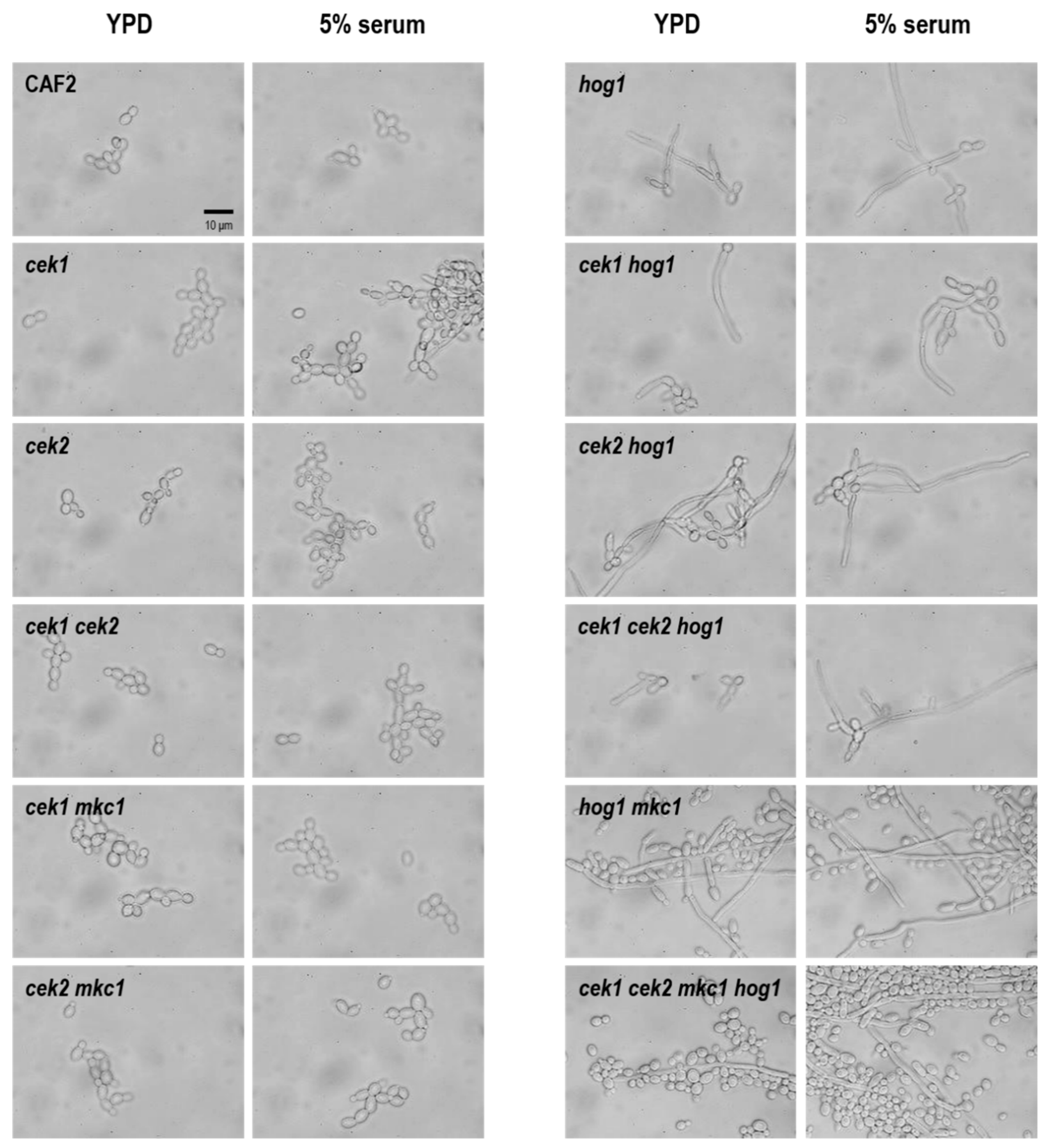
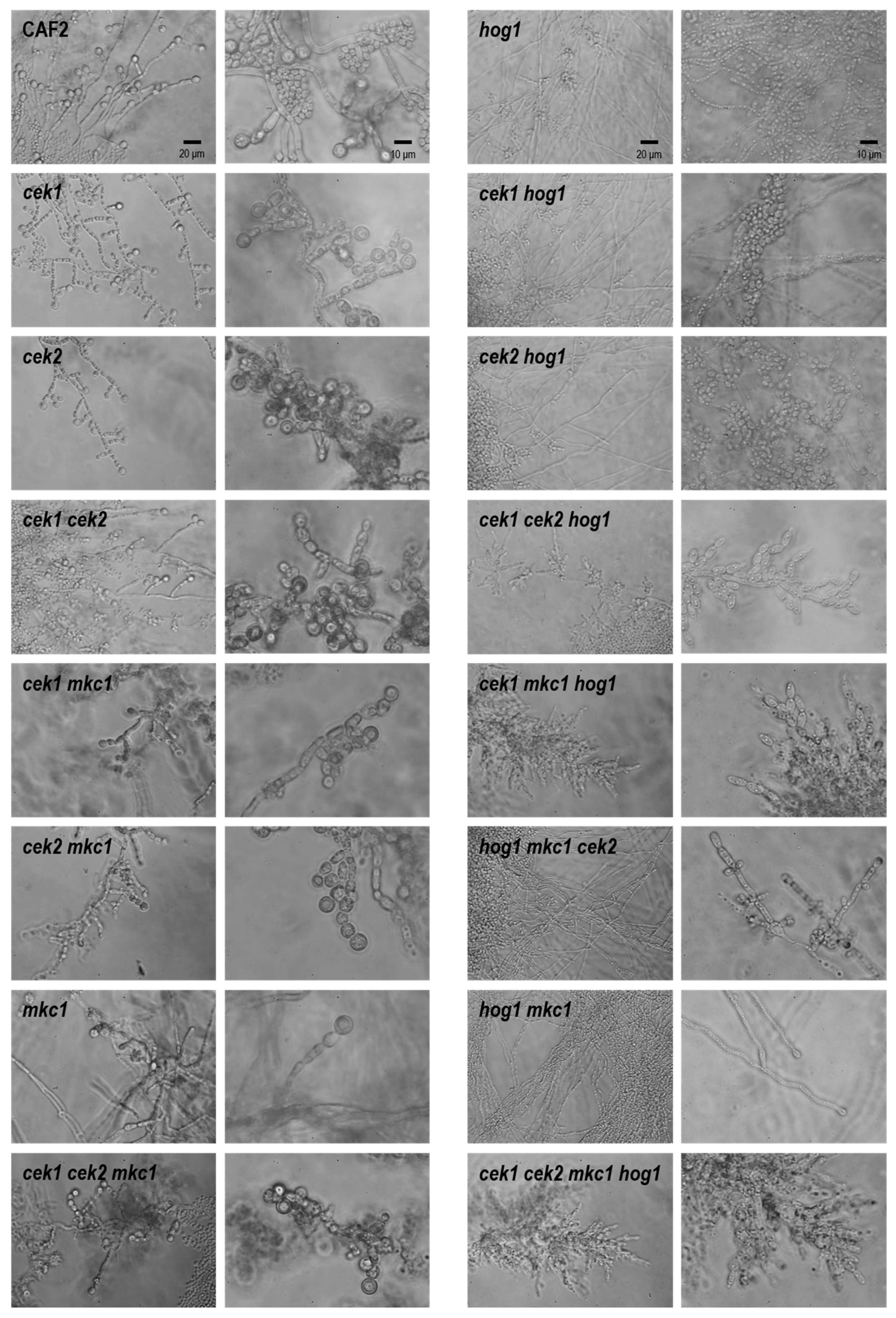
3.3. Hog1 Is the Key MAPK for C. albicans Morphological Transitions
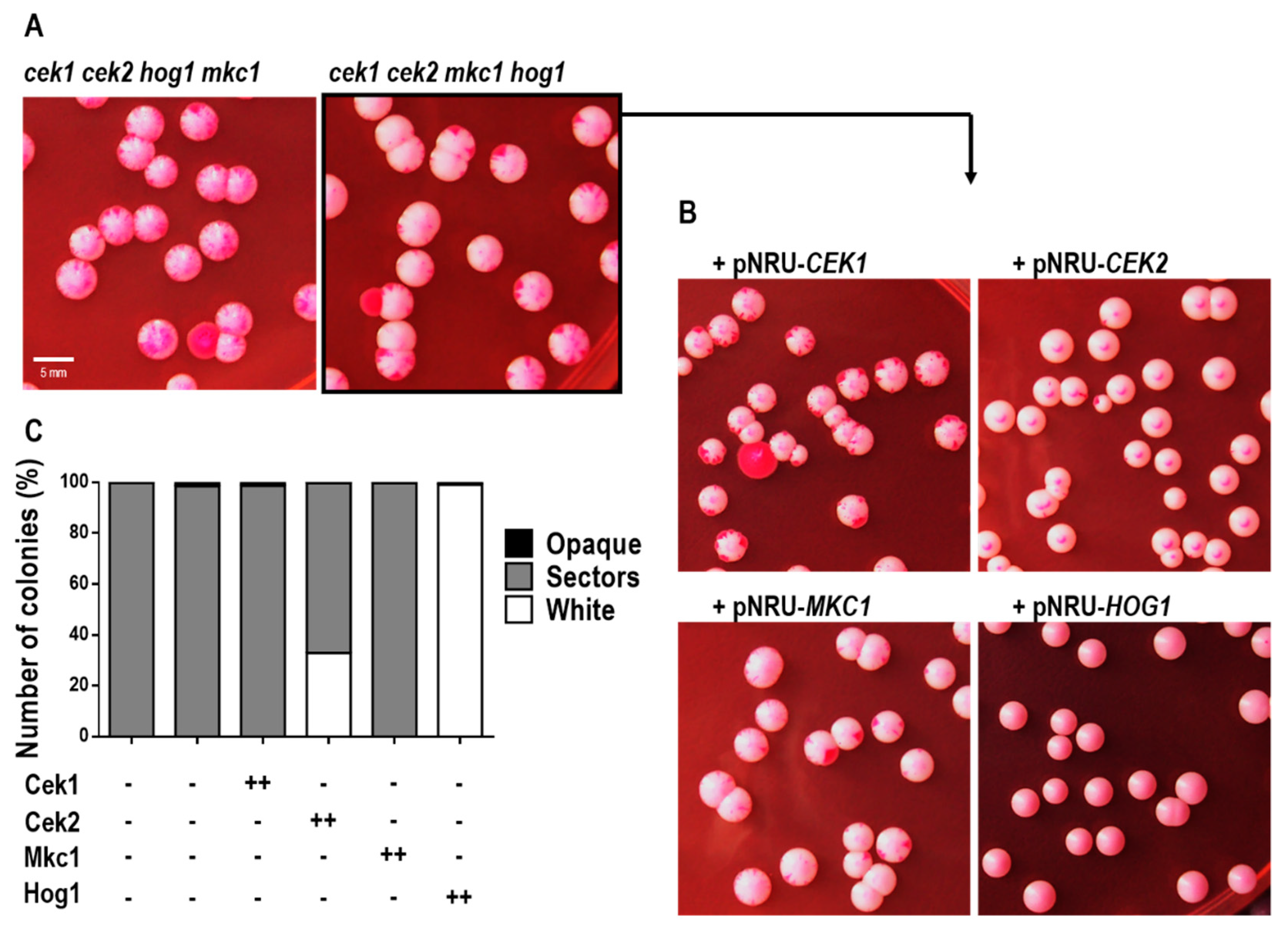
3.4. The Absence of MAPKs Triggers White-Opaque Switching in Rich Medium in a Hog1-Dependent Manner
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Pfaller, M.A.; Diekema, D.J.; Turnidge, J.D.; Castanheira, M.; Jones, R.N. Twenty Years of the SENTRY Antifungal Surveillance Program: Results for Candida Species From 1997-2016. Open forum Infect. Dis. 2019, 6, S79–S94. [Google Scholar] [CrossRef] [PubMed]
- Pfaller, M.A.; Diekema, D.J. Epidemiology of invasive candidiasis: A persistent public health problem. Clin. Microbiol. Rev. 2007, 20, 133–163. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Denning, D.W.; Gow, N.A.R.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden killers: Human fungal infections. Sci. Transl. Med. 2012, 4, 165rv113. [Google Scholar] [CrossRef]
- Lo, H.J.; Kohler, J.R.; DiDomenico, B.; Loebenberg, D.; Cacciapuoti, A.; Fink, G.R. Nonfilamentous Candida albicans mutants are avirulent. Cell 1997, 90, 939–949. [Google Scholar] [CrossRef]
- Saville, S.P.; Lazzell, A.L.; Monteagudo, C.; Lopez-Ribot, J.L. Engineered control of cell morphology in vivo reveals distinct roles for yeast and filamentous forms of Candida albicans during infection. Eukaryot. Cell 2003, 2, 1053–1060. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Navarathna, D.H.; Roberts, D.D.; Cooper, J.T.; Atkin, A.L.; Petro, T.M.; Nickerson, K.W. Arginine-induced germ tube formation inCandida albicans is essential for escape from murine macrophage line RAW 264.7. Infect. Immun. 2009, 77, 1596–1605. [Google Scholar] [CrossRef]
- Gantner, B.N.; Simmons, R.M.; Underhill, D.M. Dectin-1 mediates macrophage recognition of Candida albicans yeast but not filaments. EMBO J. 2005, 24, 1277–1286. [Google Scholar] [CrossRef]
- Saville, S.P.; Lazzell, A.L.; Chaturvedi, A.K.; Monteagudo, C.; Lopez-Ribot, J.L. Use of a genetically engineered strain to evaluate the pathogenic potential of yeast cell and filamentous forms during Candida albicans systemic infection in immunodeficient mice. Infect. Immun. 2008, 76, 97–102. [Google Scholar] [CrossRef]
- Dujardin, L.; Walbaum, S.; Biguet, J. Effect of glucose and nitrogen concentrations on the morphology of Candida albicans and the formation of chlamydospores in synthetic culture media. Mycopathologia 1980, 71, 113–118. [Google Scholar] [CrossRef]
- Montazeri, M.; Hedrick, H.G. Factors affecting spore formation in a Candida albicans strain. Appl. Environ. Microbiol. 1984, 47, 1341–1342. [Google Scholar] [CrossRef]
- Jansons, V.K.; Nickerson, W.J. Induction, morphogenesis, and germination of the chlamydospore of Candida albicans. J. Bacteriol. 1970, 104, 910–921. [Google Scholar] [CrossRef] [PubMed]
- Odds, F.C. Candida and candidosis; Baillière Tindall: London, UK, 1988; Vol. 2. [Google Scholar]
- Citiulo, F.; Moran, G.P.; Coleman, D.C.; Sullivan, D.J. Purification and germination of Candida albicans and Candida dubliniensis chlamydospores cultured in liquid media. FEMS Yeast Res. 2009, 9, 1051–1060. [Google Scholar] [CrossRef]
- Sonneborn, A.; Bockmuhl, D.P.; Ernst, J.F. Chlamydospore formation in Candida albicans requires the Efg1p morphogenetic regulator. Infect. Immun. 1999, 67, 5514–5517. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Monge, R.; Navarro-Garcia, F.; Román, E.; Negredo, A.I.; Eisman, B.; Nombela, C.; Pla, J. The Hog1 mitogen-activated protein kinase is essential in the oxidative stress response and chlamydospore formation in Candida albicans. Eukaryot. Cell 2003, 2, 351–361. [Google Scholar] [CrossRef]
- Palige, K.; Linde, J.; Martin, R.; Bottcher, B.; Citiulo, F.; Sullivan, D.J.; Weber, J.; Staib, C.; Rupp, S.; Hube, B.; et al. Global transcriptome sequencing identifies chlamydospore specific markers in Candida albicans and Candida dubliniensis. PLoS ONE 2013, 8, e61940. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.G.; Johnson, A.D. White-opaque switching in Candida albicans is controlled by mating-type locus homeodomain proteins and allows efficient mating. Cell 2002, 110, 293–302. [Google Scholar] [CrossRef]
- Lockhart, S.R.; Daniels, K.J.; Zhao, R.; Wessels, D.; Soll, D.R. Cell biology of mating in Candida albicans. Eukaryot. Cell 2003, 2, 49–61. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, S.R.; Pujol, C.; Daniels, K.J.; Miller, M.G.; Johnson, A.D.; Pfaller, M.A.; Soll, D.R. In Candida albicans, white-opaque switchers are homozygous for mating type. Genetics 2002, 162, 737–745. [Google Scholar] [PubMed]
- Soll, D.R. The role of phenotypic switching in the basic biology and pathogenesis of Candida albicans. J. Oral Microbiol. 2014, 6. [Google Scholar] [CrossRef]
- Ene, I.V.; Bennett, R.J.; Anderson, M.Z. Mechanisms of genome evolution in Candida albicans. Curr. Opin. Microbiol. 2019, 52, 47–54. [Google Scholar] [CrossRef]
- Forche, A.; Abbey, D.; Pisithkul, T.; Weinzierl, M.A.; Ringstrom, T.; Bruck, D.; Petersen, K.; Berman, J. Stress alters rates and types of loss of heterozygosity in Candida albicans. MBio 2011, 2. [Google Scholar] [CrossRef]
- Forche, A.; Magee, P.T.; Selmecki, A.; Berman, J.; May, G. Evolution in Candida albicans populations during a single passage through a mouse host. Genetics 2009, 182, 799–811. [Google Scholar] [CrossRef]
- Ropars, J.; Maufrais, C.; Diogo, D.; Marcet-Houben, M.; Perin, A.; Sertour, N.; Mosca, K.; Permal, E.; Laval, G.; Bouchier, C.; et al. Gene flow contributes to diversification of the major fungal pathogen Candida albicans. Nat. Commun. 2018, 9, 2253. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Monge, R.; Navarro-García, F.; Molero, G.; Díez-Orejas, R.; Gustin, M.; Pla, J.; Sánchez, M.; Nombela, C. Role of the mitogen-activated protein kinase Hog1p in morphogenesis and virulence of Candida albicans. J. Bacteriol. 1999, 181, 3058–3068. [Google Scholar] [CrossRef]
- San José, C.; Alonso-Monge, R.; Pérez-Díaz, R.M.; Pla, J.; Nombela, C. The mitogen-activated protein kinase homolog HOG1 gene controls glycerol accumulation in the pathogenic fungus Candida albicans. J. Bacteriol. 1996, 178, 5850–5852. [Google Scholar] [CrossRef]
- Navarro-García, F.; Sanchez, M.; Pla, J.; Nombela, C. Functional characterization of the MKC1 gene of Candida albicans, which encodes a mitogen-activated protein kinase homolog related to cell integrity. Mol. Cell. Biol. 1995, 15, 2197–2206. [Google Scholar] [CrossRef]
- Kumamoto, C.A. A contact-activated kinase signals Candida albicans invasive growth and biofilm development. Proc. Natl. Acad. Sci. USA 2005, 102, 5576–5581. [Google Scholar] [CrossRef]
- Román, E.; Nombela, C.; Pla, J. The Sho1 adaptor protein links oxidative stress to morphogenesis and cell wall biosynthesis in the fungal pathogen Candida albicans. Mol. Cell. Biol. 2005, 25, 10611–10627. [Google Scholar] [CrossRef]
- Román, E.; Alonso-Monge, R.; Gong, Q.; Li, D.; Calderone, R.; Pla, J. The Cek1 MAPK is a short-lived protein regulated by quorum sensing in the fungal pathogen Candida albicans. FEMS Yeast Res. 2009, 9, 942–955. [Google Scholar] [CrossRef] [PubMed]
- Román, E.; Correia, I.; Salazin, A.; Fradin, C.; Jouault, T.; Poulain, D.; Liu, F.T.; Pla, J. The Cek1-mediated MAP kinase pathway regulates exposure of α−(1,2) and β−(1,2)-mannosides in the cell wall of Candida albicans modulating immune recognition. Virulence 2016, 7, 558–577. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Chen, J.; Lane, S.; Liu, H. A conserved mitogen-activated protein kinase pathway is required for mating in Candida albicans. Mol. Microbiol. 2002, 46, 1335–1344. [Google Scholar] [CrossRef]
- Román, E.; Correia, I.; Prieto, D.; Alonso, R.; Pla, J. The HOG MAPK pathway in Candida albicans: More than an osmosensing pathway. Int. Microbiol. 2019. [Google Scholar] [CrossRef]
- Correia, I.; Prieto, D.; Román, E.; Wilson, D.; Hube, B.; Alonso-Monge, R.; Pla, J. Cooperative Role of MAPK Pathways in the Interaction of Candida albicans with the Host Epithelium. Microorganisms 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Román, E.; Arana, D.M.; Nombela, C.; Alonso-Monge, R.; Pla, J. MAP kinase pathways as regulators of fungal virulence. Trends Microbiol. 2007, 15, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Navarro-García, F.; Eisman, B.; Fiuza, S.M.; Nombela, C.; Pla, J. The MAP kinase Mkc1p is activated under different stress conditions in Candida albicans. Microbiology 2005, 151, 2737–2749. [Google Scholar] [CrossRef] [PubMed]
- Correia, I.; Román, E.; Prieto, D.; Eisman, B.; Pla, J. Complementary roles of the Cek1 and Cek2 MAP kinases in Candida albicans cell-wall biogenesis. Future Microbiol. 2016, 11, 51–67. [Google Scholar] [CrossRef] [PubMed]
- Eisman, B.; Alonso-Monge, R.; Román, E.; Arana, D.M.; Nombela, C.; Pla, J. The Cek1 and Hog1 mitogen-activated protein kinases play complementary roles in cell wall biogenesis and chlamydospore formation in the fungal pathogen Candida albicans. Eukaryot. Cell 2006, 5, 347–358. [Google Scholar] [CrossRef]
- Fonzi, W.A.; Irwin, M.Y. Isogenic strain construction and gene mapping in Candida albicans. Genetics 1993, 134, 717–728. [Google Scholar]
- Gillum, A.M.; Tsay, E.Y.H.; Kirsch, D.R. Isolation of the Candida albicans gene for orotidine-5’- phosphate decarboxylase by complementation of S. cerevisiae ura3 and E. coli pyrF mutations. Mol. Gen. Genet. 1984, 198, 179–182. [Google Scholar] [CrossRef] [PubMed]
- Csank, C.; Schröppel, K.; Leberer, E.; Harcus, D.; Mohamed, O.; Meloche, S.; Thomas, D.Y.; Whiteway, M. Roles of the Candida albicans mitogen-activated protein kinase homolog, Cek1p, in hyphal development and systemic candidiasis. Infect. Immun. 1998, 66, 2713–2721. [Google Scholar] [CrossRef] [PubMed]
- Prieto, A.D.; Román, E.; Correia, I.; Pla, J. The HOG pathway is critical for the colonization of the mouse gastrointestinal tract by Candida albicans. PLoS ONE 2014, 9, e87128. [Google Scholar] [CrossRef] [PubMed]
- Arana, D.M.; Alonso-Monge, R.; Du, C.; Calderone, R.; Pla, J. Differential susceptibility of mitogen-activated protein kinase pathway mutants to oxidative-mediated killing by phagocytes in the fungal pathogen Candida albicans. Cell. Microbiol. 2007, 9, 1647–1659. [Google Scholar] [CrossRef]
- Herrero-de-Dios, C.; Alonso-Monge, R.; Pla, J. The lack of upstream elements of the Cek1 and Hog1 mediated pathways leads to a synthetic lethal phenotype upon osmotic stress in Candida albicans. Fungal Genet. Biol. 2014, 69, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, D.; Kondo, K.; Uehara, N.; Otokozawa, S.; Tsuji, N.; Yagihashi, A.; Watanabe, N. Endogenous reactive oxygen species is an important mediator of miconazole antifungal effect. Antimicrob. Agents Chemother. 2002, 46, 3113–3117. [Google Scholar] [CrossRef]
- Mesa-Arango, A.C.; Trevijano-Contador, N.; Román, E.; Sanchez-Fresneda, R.; Casas, C.; Herrero, E.; Arguelles, J.C.; Pla, J.; Cuenca-Estrella, M.; Zaragoza, O. The production of reactive oxygen species is a universal action mechanism of Amphotericin B against pathogenic yeasts and contributes to the fungicidal effect of this drug. Antimicrob. Agents Chemother. 2014, 58, 6627–6638. [Google Scholar] [CrossRef] [PubMed]
- Román, E.; Prieto, D.; Martin, R.; Correia, I.; Mesa Arango, A.C.; Alonso-Monge, R.; Zaragoza, O.; Pla, J. Role of catalase overproduction in drug resistance and virulence in Candida albicans. Future Microbiol. 2016, 11, 1279–1297. [Google Scholar] [CrossRef]
- Correia, I. Role of the Cek1 and Cek2 kinases within the MAPK signalling network of the fungal pathogen Candida albicans. Ph.D. Thesis, Universidad Complutense de Madrid, Facultad de Farmacia, Madrid, Spain, 2014. [Google Scholar]
- Cabib, E. Differential inhibition of chitin synthetases 1 and 2 from Saccharomyces cerevisiae by polyoxin D and nikkomycins. Antimicrob. Agents Chemother. 1991, 35, 170–173. [Google Scholar] [CrossRef] [PubMed]
- Navarro-García, F.; Alonso-Monge, R.; Rico, H.; Pla, J.; Sentandreu, R.; Nombela, C. A role for the MAP kinase gene MKC1 in cell wall construction and morphological transitions in Candida albicans. Microbiology 1998, 144, 411–424. [Google Scholar] [CrossRef]
- Slutsky, B.; Staebell, M.; Anderson, J.; Risen, L.; Pfaller, M.; Soll, D.R. “White-opaque transition”: A second high-frequency switching system in Candida albicans. J. Bacteriol. 1987, 169, 189–197. [Google Scholar] [CrossRef]
- Rikkerink, E.H.; Magee, B.B.; Magee, P.T. Opaque-white phenotype transition: A programmed morphological transition in Candida albicans. J. Bacteriol. 1988, 170, 895–899. [Google Scholar] [CrossRef]
- Soll, D.R.; Morrow, B.; Srikantha, T. High-frequency phenotypic switching in Candida albicans. Trends Genet. 1993, 9, 61–65. [Google Scholar] [CrossRef]
- Rosenberg, S.M. Stress-induced loss of heterozygosity in Candida: A possible missing link in the ability to evolve. MBio 2011, 2. [Google Scholar] [CrossRef]
- Day, A.M.; Quinn, J. Stress-Activated Protein Kinases in Human Fungal Pathogens. Front. Cell Infect. Microbiol. 2019, 9, 261. [Google Scholar] [CrossRef]
- Pradhan, A.; Avelar, G.M.; Bain, J.M.; Childers, D.; Pelletier, C.; Larcombe, D.E.; Shekhova, E.; Netea, M.G.; Brown, G.D.; Erwig, L.; et al. Non-canonical signalling mediates changes in fungal cell wall PAMPs that drive immune evasion. Nat. Commun. 2019, 10, 5315. [Google Scholar] [CrossRef]
- Sokol-Anderson, M.L.; Brajtburg, J.; Medoff, G. Amphotericin B-induced oxidative damage and killing of Candida albicans. J. Infect. Dis. 1986, 154, 76–83. [Google Scholar] [CrossRef] [PubMed]
- Guirao-Abad, J.P.; Sanchez-Fresneda, R.; Roman, E.; Pla, J.; Arguelles, J.C.; Alonso-Monge, R. The MAPK Hog1 mediates the response to amphotericin B in Candida albicans. Fungal Genet. Biol. 2020, 136, 103302. [Google Scholar] [CrossRef] [PubMed]
- Galan-Diez, M.; Arana, D.M.; Serrano-Gomez, D.; Kremer, L.; Casasnovas, J.M.; Ortega, M.; Cuesta-Dominguez, A.; Corbi, A.L.; Pla, J.; Fernandez-Ruiz, E. Candida albicans beta-glucan exposure is controlled by the fungal CEK1-mediated mitogen-activated protein kinase pathway that modulates immune responses triggered through dectin-1. Infect. Immun. 2010, 78, 1426–1436. [Google Scholar] [CrossRef] [PubMed]
- Munro, C.A.; Selvaggini, S.; de, B.I.; Walker, L.; Lenardon, M.D.; Gerssen, B.; Milne, S.; Brown, A.J.; Gow, N.A. The PKC, HOG and Ca2+ signalling pathways co-ordinately regulate chitin synthesis in Candida albicans. Mol. Microbiol. 2007, 63, 1399–1413. [Google Scholar] [CrossRef] [PubMed]
- Arana, D.M.; Nombela, C.; Alonso-Monge, R.; Pla, J. The Pbs2 MAP kinase kinase is essential for the oxidative-stress response in the fungal pathogen Candida albicans. Microbiology 2005, 151, 1033–1049. [Google Scholar] [CrossRef]
- Su, C.; Lu, Y.; Liu, H. Reduced TOR signaling sustains hyphal development in Candida albicans by lowering Hog1 basal activity. Mol. Biol. Cell 2013, 24, 385–397. [Google Scholar] [CrossRef]
- Biswas, S.; Van Dijck, P.; Datta, A. Environmental sensing and signal transduction pathways regulating morphopathogenic determinants of Candida albicans. Microbiol. Mol. Biol. Rev. 2007, 71, 348–376. [Google Scholar] [CrossRef] [PubMed]
- Srikantha, T.; Soll, D.R. A white-specific gene in the white-opaque switching system of Candida albicans. Gene 1993, 131, 53–60. [Google Scholar] [CrossRef]
- Ramirez-Zavala, B.; Reuss, O.; Park, Y.N.; Ohlsen, K.; Morschhauser, J. Environmental induction of white-opaque switching in Candida albicans. PLoS Path. 2008, 4, e1000089. [Google Scholar] [CrossRef]
- Huang, G.; Srikantha, T.; Sahni, N.; Yi, S.; Soll, D.R. CO2 regulates white-to-opaque switching in Candida albicans. Curr. Biol. 2009, 19, 330–334. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Yi, S.; Sahni, N.; Daniels, K.J.; Srikantha, T.; Soll, D.R. N-acetylglucosamine induces white to opaque switching, a mating prerequisite in Candida albicans. PLoS Path. 2010, 6, e1000806. [Google Scholar] [CrossRef]
- Alby, K.; Schaefer, D.; Bennett, R.J. Homothallic and heterothallic mating in the opportunistic pathogen Candida albicans. Nature 2009, 460, 890–893. [Google Scholar] [CrossRef]
- Kolotila, M.P.; Diamond, R.D. Effects of neutrophils and in vitro oxidants on survival and phenotypic switching of Candida albicans WO-1. Infect. Immun. 1990, 58, 1174–1179. [Google Scholar] [CrossRef]
- Liang, S.H.; Cheng, J.H.; Deng, F.S.; Tsai, P.A.; Lin, C.H. A novel function for Hog1 stress-activated protein kinase in controlling white-opaque switching and mating in Candida albicans. Eukaryot. Cell 2014, 13, 1557–1566. [Google Scholar] [CrossRef]
- Chang, W.H.; Liang, S.H.; Deng, F.S.; Lin, C.H. The conserved dual phosphorylation sites of the Candida albicans Hog1 protein are crucial for white-opaque switching, mating, and pheromone-stimulated cell adhesion. Med Mycol. J. 2016. [Google Scholar] [CrossRef]
- Zordan, R.E.; Miller, M.G.; Galgoczy, D.J.; Tuch, B.B.; Johnson, A.D. Interlocking transcriptional feedback loops control white-opaque switching in Candida albicans. PLoS Biol. 2007, 5, e256. [Google Scholar] [CrossRef]
- Srikantha, T.; Borneman, A.R.; Daniels, K.J.; Pujol, C.; Wu, W.; Seringhaus, M.R.; Gerstein, M.; Yi, S.; Snyder, M.; Soll, D.R. TOS9 regulates white-opaque switching in Candida albicans. Eukaryot. Cell 2006, 5, 1674–1687. [Google Scholar] [CrossRef]
- Ramirez-Zavala, B.; Weyler, M.; Gildor, T.; Schmauch, C.; Kornitzer, D.; Arkowitz, R.; Morschhauser, J. Activation of the Cph1-dependent MAP kinase signaling pathway induces white-opaque switching in Candida albicans. PLoS Path. 2013, 9, e1003696. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Tao, L.; Nobile, C.J.; Tong, Y.; Guan, G.; Sun, Y.; Cao, C.; Hernday, A.D.; Johnson, A.D.; Zhang, L.; et al. White-opaque switching in natural MTLa/alpha isolates of Candida albicans: Evolutionary implications for roles in host adaptation, pathogenesis, and sex. PLoS Biol. 2013, 11, e1001525. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Wang, H.; Chou, S.; Nie, X.; Chen, J.; Liu, H. Bistable expression of WOR1, a master regulator of white-opaque switching in Candida albicans. Proc. Natl. Acad. Sci. USA 2006, 103, 12813–12818. [Google Scholar] [CrossRef]
- Rastghalam, G.; Omran, R.P.; Alizadeh, M.; Fulton, D.; Mallick, J.; Whiteway, M. MAP Kinase Regulation of the Candida albicans Pheromone Pathway. mSphere 2019, 4. [Google Scholar] [CrossRef]
- Selmecki, A.M.; Dulmage, K.; Cowen, L.E.; Anderson, J.B.; Berman, J. Acquisition of aneuploidy provides increased fitness during the evolution of antifungal drug resistance. PLoS Genet. 2009, 5, e1000705. [Google Scholar] [CrossRef] [PubMed]
| Strain Name | Background Strain and Genotype | Source |
|---|---|---|
| SC5314 (wt) | Clinical isolate | [40] |
| CAF2 (wt) | ura3Δ::imm434/ura3Δ::imm434-URA3 | [39] |
| CAI4 | ura3Δ::imm434/ura3Δ::imm434 | [39] |
| cek1 | [CAI4] cek1Δ::hisG-URA3-hisG/cek1Δ::hisG | [41] |
| cek2 | [CAI4] cek2::cat-URA3-cat/cek2::cat | [37] |
| mkc1 | [CAI4] mkc1::hisG-URA3-hisG/mkc1::hisG | [27] |
| hog1 | [CAI4] hog1::hisG-URA3-hisG/hog1::hisG | [42] |
| cek1 cek2 | [CAI4] cek1::hisG/cek1::hisG cek2::cat-URA3-cat/cek2::cat | [37] |
| cek1 mkc1 | [CAI4] cek1::hisG/cek1::hisG mkc1::hisG-URA3-hisG/mkc1::hisG | [37] |
| cek1 hog1 | [CAI4] cek1::hisG/cek1::hisG hog1::hisG/hog1::hisG ARD1/ard1::FRT SAP2pr-FLPURA3 | [43] |
| hog1 mkc1 | [CAI4] hog1::hisG/hog1::hisG mkc1::hisG-URA3-hisG/mkc1::hisG | [37] |
| cek2 hog1 | [CAI4] cek2::cat/cek2::cat hog1::hisG-URA3-hisG/hog1::hisG | [37] |
| cek2 mkc1 | [CAI4] cek2::cat/cek2::cat mkc1::hisG-URA3-hisG/mkc1::hisG | [37] |
| cek1 cek2 mkc1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG-URA3-hisG/mkc1::hisG | This work |
| cek1 cek2 hog1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat hog1::hisG-URA3-hisG/hog1::hisG | This work |
| cek1 mkc1 hog1 | [CAI4] cek1::hisG/cek1::hisG mkc1::hisG/mkc1::hisG hog1::hisG-URA3-hisG/hog1::hisG | This work |
| hog1 mkc1 cek2 | [CAI4] hog1::hisG/hog1::hisG mkc1::hisG/mkc1::hisG cek2::cat-URA3-cat/cek2::cat | This work |
| cek1 cek2 hog1 mkc1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat hog1::hisG/hog1::hisG mkc1::hisG-URA3-hisG/mkc1::hisG | This work |
| cek1 cek2 mkc1 hog1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG/mkc1::hisG hog1::hisG-URA3-hisG/hog1::hisG | This work |
| cek1 cek2 mkc1 hog1 pNRU-CEK1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG/mkc1::hisG hog1::hisG/hog1::hisG ADH1/adh1::tTA pTet-CEK1-myc-URA3 | This work |
| cek1 cek2 mkc1 hog1 pNRU-CEK2 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG/mkc1::hisG hog1::hisG/hog1::hisG ADH1/adh1::tTA pTet-CEK2-myc-URA3 | This work |
| cek1 cek2 mkc1 hog1 pNRU-MKC1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG/mkc1::hisG hog1::hisG/hog1::hisG ADH1/adh1::tTA pTet-MKC1-myc-URA3 | This work |
| cek1 cek2 mkc1 hog1 pNRU-HOG1 | [CAI4] cek1::hisG/cek1::hisG cek2::cat/cek2::cat mkc1::hisG/mkc1::hisG hog1::hisG/hog1::hisG ADH1/adh1::tTA pTet-HOG1-myc-URA3 | This work |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Correia, I.; Wilson, D.; Hube, B.; Pla, J. Characterization of a Candida albicans Mutant Defective in All MAPKs Highlights the Major Role of Hog1 in the MAPK Signaling Network. J. Fungi 2020, 6, 230. https://doi.org/10.3390/jof6040230
Correia I, Wilson D, Hube B, Pla J. Characterization of a Candida albicans Mutant Defective in All MAPKs Highlights the Major Role of Hog1 in the MAPK Signaling Network. Journal of Fungi. 2020; 6(4):230. https://doi.org/10.3390/jof6040230
Chicago/Turabian StyleCorreia, Inês, Duncan Wilson, Bernhard Hube, and Jesús Pla. 2020. "Characterization of a Candida albicans Mutant Defective in All MAPKs Highlights the Major Role of Hog1 in the MAPK Signaling Network" Journal of Fungi 6, no. 4: 230. https://doi.org/10.3390/jof6040230
APA StyleCorreia, I., Wilson, D., Hube, B., & Pla, J. (2020). Characterization of a Candida albicans Mutant Defective in All MAPKs Highlights the Major Role of Hog1 in the MAPK Signaling Network. Journal of Fungi, 6(4), 230. https://doi.org/10.3390/jof6040230







