Abstract
Fungal infections are a growing public health concern, and an increasingly important cause of human mortality, with Candida species being amongst the most frequently encountered of these opportunistic fungal pathogens. Several Candida species are polymorphic, and able to transition between distinct morphological states, including yeast, hyphal, and pseudohyphal forms. While not all Candida pathogens are polymorphic, the ability to undergo morphogenesis is linked with the virulence of many of these pathogens. There are also many connections between Candida morphogenesis and antifungal drug treatment and susceptibility. Here, we review how Candida morphogenesis—a key virulence trait—is linked with antifungal drugs and antifungal drug resistance. We highlight how antifungal therapeutics are able to modulate morphogenesis in both sensitive and drug-resistant Candida strains, the shared signaling pathways that mediate both morphogenesis and the cellular response to antifungal drugs and drug resistance, and the connection between Candida morphology, drug resistance, and biofilm growth. We further review the development of anti-virulence drugs, and targeting Candida morphogenesis as a novel therapeutic strategy to target fungal pathogens. Together, this review highlights important connections between fungal morphogenesis, virulence, and susceptibility to antifungals.
1. Introduction to Candida and Candidiasis Infections
The fungal kingdom contains diverse organisms which make significant contributions to supporting life on our planet. Fungi are of vital importance to humanity as industrial manufacturers, play a key role in environmental nutrient recycling, and are model organisms for many eukaryotic processes. Our use of fungi throughout history has expanded in scope, from an edible agricultural resource, to the production of food, beverages and are a valuable source of antimicrobial compounds. However, there are also many fungal species that are serious pathogenic threats to plant, animal, and human health [1]. Currently, approximately 5 million fungal species have been classified, of which an estimated 300 are capable of establishing disease within a mammalian host [2,3]. As a human pathogen, it has recently been reported that, globally, fungal infections currently cause 1.5 million deaths per year [4], a statistic that rivals that of tuberculosis. Invasive fungal infections are a relatively modern problem, associated with newly-emerging populations of aging and immunocompromised individuals, such as HIV/AIDS patients, those undergoing cancer chemotherapy treatment, organ transplantation recipients, and others.
Amongst the most notable human-associated fungal pathogens are members from the Candida genus, including Candida albicans, Candida tropicalis, Candida glabrata, Candida krusei, Candida parapsilosis, and the emerging pathogens Candida haemulonii and Candida auris (Table 1); these fungi all cause a range of infections, which, particularly in immunocompromised patients, can lead to severe, invasive candidiasis including bloodstream infections [5,6,7]. Both acquired and inherited immunodeficiencies and other risk factors may predispose patients to Candida infections. The risk of candidiasis is increased in patients with underlying malignancies and those undergoing chemotherapy, patients undergoing hematopoietic stem cell or solid organ transplantation, those with immunosuppressive diseases (i.e., HIV/AIDS), patients using broad-spectrum antibiotics or corticosteroids and using invasive medical interventions (i.e., central vascular catheters), and those with certain genetic risk factors (i.e., impaired IL-17 immunity) [8,9,10,11]. In most cases, Candida spp. are opportunistic pathogens that reside as commensal organisms present in the body’s endogenous microbiome [12]. However, factors such as an impaired immune system, or implanted medical devices, can create an environment that enables invasive infections to arise [13]. Nosocomial infections of candidiasis are becoming more common, and account for more than 85% of all invasive fungal infections in both Europe and the United States [6]. Not only are these infections becoming more prevalent, they are accompanied by dangerously high mortality rates in both developing and developed countries alike. Globally, 30-day all-cause mortality rates range from 29 to 72% [14]. In 2016, Dio et al. [5] reported crude mortality rates of candidemia of 72.2% in Brazil, and Bongomin et al. [4] reported mortality rates in the UK in excess of 40%.

Table 1.
Candida species: morphogenesis, antifungal responses and virulence processes.
There are currently almost 200 species of yeasts in the genus Candida [31], and the notable pathogens amongst them include C. albicans, C. dubliniensis, C. parapsilosis, C. glabrata, C. tropicalis, C. lusitaniae, and C. krusei. Additionally, C. auris has recently emerged as a highly multidrug-resistant pathogen, and a serious global health threat [32]. Amongst these Candida pathogens, C. albicans is the species mostly common associated with human infection [14]. Each of these Candida species possess critical factors that contribute to these fungi being well adapted as a pathogen to the human host. Processes such as cellular morphogenesis, cell-surface adhesion, phenotypic switching, biofilm formation, antifungal drug resistance, and secretion of hydrolytic enzymes are all well-established virulence mechanisms that aid in the success of these species as pathogens [33,34,35,36,37,38,39,40].
2. Candida Morphogenesis: Utility and Regulation
Certain Candida species possess the ability to undergo complex cellular morphological transitions; this trait is frequently linked to the virulence of these opportunistic pathogens [33,36,39,41]. There are three relevant morphotypes of Candida spp.: yeasts, hyphae and pseudohyphae, with hyphae and pseudohyphae sometimes collectively referred to as filamentous growth states [41,42]. Yeasts are ovoid, single cells that grow and divide via axial and bipolar budding, reminiscent of the the model organism Saccharomyces cerevisiae [43,44]. Hyphal states are indicative by their elongated state with a uniform width and parallel sides. They have pores in their septa that allow for cell to cell communication and tend to grow in a polarized manner [42,45]. Pseudohyphae share the traits of polarized growth and elongation with hyphae; however, they are ellipsoid and their septa are constricted. Pseudohyphae also tend to be phenotypically distinct from hyphae, via their highly branched nature, as their cells are smaller and take less time to divide [46]. While pseudohyphal growth is commonly observed amongst Candida species, the extent and frequency to which different species can undergo pseudohyphal growth, and the length and appearance of the pseudohyphae can vary significantly between species. Amongst Candida spp., only certain species possess the ability to undergo the morphological transition between yeast and filamentous growth states (Table 1). The best studied of these is C. albicans, which is readily able to transition between yeast, pseudohyphal, and hyphal growth states, under diverse environmental conditions [39]. Other Candida species, are similarly able to undergo a yeast to hyphal/pseudohyphal transition, including C. tropicalis [47] and C. dubliniensis [48]. The Candida species C. parapsilosis, C. lusitaniae, C. haemulonii and C. krusei are unable to form true hyphae, but can transition between yeast and pseudohyphal states [24,49,50,51]. Recently, a filamentous growth state was observed in the emerging Candida pathogen, C. auris, upon growth on NaCl-rich YPD, or upon deletion of the molecular chaperone HSP90 [52,53]. Passage through a mammalian host also revealed a hyphal-like phenotype in C. auris [26]. This is an important finding, and one that may further help explain the success of C. auris as a highly virulent, emerging fungal pathogen.
Cellular morphogenesis in Candida species between yeast and filamentous growth is often associated with virulence, with both yeast and filamentous forms contributing to pathogenesis in distinct capacities [54]. For C. albicans, it is suggested that the yeast morphology aids in fungal colonization, biofilm formation, and rapid dissemination into the host bloodstream [55]. On the other hand, the hyphal growth state facilitates deep tissue colonization, enhances biofilm formation, and enables macrophage evasion [45]. C. albicans cells in a hyphal morphology can invade epithelial cells by exerting a mechanical force to force growth between tissues, and similar mechanical stretching can enable hyphal escape from macrophages [56,57]. C. albicans hyphae also secrete candidalysin, a peptide toxin that is critical for virulence, via damage to host epithelial cell membranes, and stimulating host cell signaling pathways [58]. Candidalysin is itself derived from proteolytic cleavage of the Ece1 protein, the product of the hyphal-specific gene ECE1 [58,59]. C. albicans’ thigmotropism also allows the pathogen to find weak points such as grooves in the host tissue to penetrate and cause further host damage [60].
Much experimental evidence has demonstrated that, when polymorphic Candida spp. are locked into one morphotype or the other, its virulence is lowered, or completely abolished [54,61,62,63]. This indicates that the specific functions demonstrated by each morphotype is directly responsible for the overall virulence of these polymorphic pathogens. Conversely, there are other species in the Candida genus that exist mainly in a yeast morphology, yet remain virulent. For example, C. glabrata is typically a non-dimorphic yeast that is capable of being highly virulent [17]. Additionally, while deleting genes involved in morphogenesis often results in loss of virulence, there are factors that are involved in morphogenesis and not virulence, and, conversely, virulence factors that play no apparent role in morphogenesis [54]. This suggests that, while filamentation and morphogenesis is an important virulence strategy for some Candida species, it is not universal, nor is it strictly required for virulence. There is a complex evolutionary relationship between fungal morphogenesis and virulence, which has been subject to in-depth exploration [63].
2.1. Key Signal Transduction Pathways That Govern Morphogenesis
The genes that promote C. albicans filamentation are tightly regulated by a network of key signaling pathways and transcription factors [64]. These signaling processes have been subject to significant research and review [39,64,65,66,67]. Briefly, these filamentous growth-promoting transcription factors are controlled by complex cellular signaling pathways that are ultimately triggered by a wide range of environmental signals, such as serum, nutrient limitation, temperature, pH and CO2 concentration [64]. These environmental factors regulating morphogenesis are controlled via multiple signalling pathways, depending on the physiological cues present. Pathways such as the cAMP-PKA and Ras1-MAPK are both initiated by the guanine nucleotide binding protein Ras1, but diverge their signalling cascades downstream of Ras1 activation. Most components of these pathways are critical for filamentous growth, including the downstream transcriptional regulator Efg1 [39]. Several additional MAPK pathways are also critical regulators of C. albicans morphogenesis; the protein kinase C (PKC) and the high osmolarity glycerol (HOG) MAPK pathways are involved in maintaining cell wall integrity and have components that modulate morphogenesis [68,69,70]. Other pathways such as the pH, TOR, and cell cycle arrest pathways all control filamentous growth and the expression of hyphae-specific genes via different signalling cascades [39]. Additionally, the heat shock factor Hsp90 governs temperature-dependent morphogenesis in C. albicans via the cAMP-PKA pathway [71,72,73], and the heat shock transcription factor Hsf1 also regulates filamentous growth by compromising the Hsp90 function [74].
Hyphal formation in C. albicans can also be negatively regulated by different signalling cues, and, most notably, by the fungal-produced quorum sensing molecule farnesol [75,76,77]. Farnesol initiates repression of filamentous growth by targeting key signaling cascades involved in the yeast-to-hyphal growth transition [78]. Farnesol targets the cAMP-PKA pathway via hydrophobically disrupting the membrane-bound Ras1-Cyr1 complex that affects downstream cAMP production [79]. In addition to the cAMP-PKA pathway, farnesol acts through Cup9 and Ubr1 signaling. When C. albicans cells are released from farnesol inhibition, the ubiquitin protein ligase Ubr1 degrades the transcriptional repressor Cup9, which in turn de-represses expression of SOK1, which is required for degradation of Nrg1, and initiation of filamentous growth [80,81,82]. Farnesol has also been shown to regulate multiple kinases and transcription factors responsible for hyphal formation [83], establishing this molecule as an important modulator of the yeast to hyphal transition. For C. albicans, farnesol is only produced under aerobic growth conditions [84].
Ultimately, activation of filamentous growth pathways and transcription factors reads the expression of ‘hyphal-specific genes’, which are specifically induced or upregulating in filamentous growth forms. These include a number of cell surface adhesin proteins that aid in fungal adhesion, including agglutinin-like sequence (ALS) family proteins, and ‘hyphal wall protein’ HWP1 [85]. Other genes that are upregulated upon yeast-to-filamentous growth transition in C. albicans include secreted aspartic proteases, encoded by SAP genes [86,87] proteins that exhibit extracellular proteolytic activity, aid in the degradation of host proteins, and are tightly linked to fungal virulence [88,89]. ECE1 is also a hyphal-specific gene in C. albicans; the proteolytic cleavage of the Ece1 protein results in production of the peptide toxin candidalysin [59]. Other hyphal-specific genes associated with the filamentous form of C. albicans include HGC1 [90], DDR48, RBT1, and RBT4 [91,92].
Transcriptional control over genes governing morphogenesis is critical, and multiple transcription factors are core modulators of the different C. albicans morphogenetic circuitry. For example, the filament-specific transcriptional regulator UME6 is needed for proper expression of these hyphal-specific genes [93]. In experiments where Ume6 is constitutively expressed, it forms hyphae even in non-filament-inducing cues [94]. EFG1 is also a major transcriptional regulator of filamentation that governs yeast to hyphal morphogenesis downstream of cAMP-PKA signaling [95,96,97]. Many other transcription factors such as FLO8, CPH1, EED1, CZF1 and TEC1 are also positive regulators of filamentous growth [98,99,100,101]. Other transcription factors such as TUP1, NRG1 and RFG1 are negative regulators of filamentation. They act by repressing hyphae specific genes and when null mutants are created, hyphal formation occurs in C. albicans without the need for any inducing cues [102,103,104]. Together, these factors are uniquely expressed or upregulated in C. albicans filamentous cells, and are responsible for the phenotypes associated with filamentous growth.
2.2. Biofilm Formation
Candida spp., are frequently found as part of microbial biofilms in both their yeast and filamentous growth states. Biofilms are structured microbial communities that are comprised of single or multiple microbial species, encased in an extracellular matrix (ECM). Similar to other microbial biofilms, Candida biofilms tend to be more resistant to host defences and antimicrobial agents, particularly azoles, and can be an important component of persistence within the host [12,29,105,106]. Candida biofilms are composed of multiple layers of microbial cells and a matrix of extracellular material that may vary between species, consisting of polysaccharides, including β-1,3-glucan, β-1,6-glucan, mannans, and proteins [107,108]. Biofilm associated C. albicans cells contain cell walls which are twice as thick, and contain more carbohydrates and β-1,3-glucan, than their planktonic cell counterpart [107]. While C. albicans is commonly associated with biofilm growth, many other non-albicans Candida species are also capable of biofilm growth, as well as filamentous growth, such as C. tropicalis, C. parapsilosis, and C. auris [37,109]. Biofilms significantly increase fungal pathogenicity due to their ability to secrete a protective ECM, and anchor robustly to host tissues [12].
The ability of several Candida species, including C. albicans, C. tropicalis, and C. parapsilosis to transition between distinct morphological states, and the large number of surface adhesins associated specifically with the hyphal form, such as ALS3 and HWP1, both play key roles in maintaining biofilm formation [37,110,111]. In many cases, C. albicans mutants unable to undergo morphogenesis are also defective in forming biofilms [112]. Filamentous cells contribute structural integrity and strength to a biofilm and act as a support for adherence of yeast cells [38], while yeast cells may allow for dissemination of biofilm cells within a host [110]. Filamentous cells also enable C. albicans to invade host cells and tissues, and allow biofilms to mature [12]. The primary morphological composition of the biofilm dictates its strength: biofilms that are composed of 50% or more hyphae are more robust and more difficult to disrupt [64,112], due to filamentous cells expressing more adhesin proteins, facilitating growth of more mature, robust biofilms [38]. Biofilm formation is also regulated by a complex transcriptional network that is intrinsically linked to morphogenetic switching in polymorphic Candida spp. [111]. This network is governed by several key transcription factors, including Ndt80, Rob1, Bcr1, Efg1, Tec1, Brg1, Flo8, Gal4, and Rfx2, many of which are also crucial for filamentous growth [113,114].
3. Fungal Treatment: Classes of Antifungals and Drug Resistance
3.1. Antifungal Drugs
Fungal morphogenesis has links with treatment and response to antifungal drugs. Here, we briefly review the major classes of antifungal drugs and mechanisms of resistance. Due to the relatively close evolutionary relationship between eukaryotic fungal pathogens and mammalian cells, there are limited cellular targets which can be exploited for use in antifungal drug therapy, while also exhibiting limited toxicity against human patients. The four major classes of antifungal drugs used for treatment of Candida infections are the azoles, polyenes, echinocandins, and flucytosine.
Azoles target production of ergosterol [115]: a key sterol which serves as the functional equivalent of cholesterol within mammalian cells, in order to regulate the integrity and fluidity of the fungal cell membrane [116]. Azoles exert their fungistatic effect on Candida by targeting ergosterol production. They function by specifically targeting the enzyme lanosterol 14α-demethylase, (that is encoded by the ERG11 gene in Candida) which is a key enzyme in the ergosterol biosynthetic pathway [117]. Azoles contain a free nitrogen atom which binds to the heme group at the active site of Erg11, preventing the demethylation of lanosterol [118,119,120] and effectively blocking ergosterol production.
The echinocandins are a class of antifungal drugs that, unlike azoles, typically act in a fungicidal manner against most Candida species [121]. Echinocandins exhibit low toxicity against the host and target the fungal cell wall specifically by acting as non-competitive inhibitors of the enzyme (1,3)-β-d-glucan synthase [121]. This inhibition results in the disruption of the synthesis of the carbohydrate β-1,3-glucan, a major component of the fungal cell wall [122]. Disruption of (1,3)-β-d-glucan synthase causes severe stress on the fungal cell wall and results in loss of cell wall integrity, followed by lysis [123].
Polyenes are potent, amphipathic compounds, which contain both hydrophobic and hydrophilic regions, giving these drugs a unique molecular structure capable of directly targeting and strongly binding to ergosterol in the fungal cell membrane [124]. Similar to echinocandins, they exert a fungicidal effect on C. albicans, as the binding of a polyene to ergosterol, forms a large drug-lipid complex which spans the fungal cell membrane. This forms a leaky channel, disrupting the membrane and allowing the escape of cellular ions out of the cell which significantly disrupts the proton gradient of the cell [125]. Recently, the production of reactive oxygen species (ROS) has also been proposed to play a role in the fungicidal activity of the polyene amphotericin B [126,127]. Amphotericin B has an impact on fungal redox homeostasis [127], and alters C. albicans cellular metabolism and respiration, ultimately leading to the production of intracellular ROS, which is lethal to the cell [126]. Polyenes have been well known antifungals for over 50 years, but their clinical use has been limited due to both poor solubility and strong nephrotoxicity, caused by structural similarities between ergosterol, and its mammalian analogue cholesterol [128,129].
Finally, the pyrimidine analog flucytosine works by blocking both RNA and DNA biosynthesis pathways in fungi. Flucytosine is rarely, if ever, used in antifungal monotherapy against Candida infections, due to the rapid and frequent development of resistance to this antifungal [130]. Typically, flucytosine is used in combination with other antifungal, such as amphotericin B. Flucytosine (in combination with amphotericin B), is recommended for the treatment of drug-resistant Candida infections, including fluconazole-resistant C. glabrata infections [131].
3.2. Antifungal Drug Resistance
Candida species are able to evolve various drug resistance against all three classes of antifungals, and some have intrinsic resistance to antifungals (Table 1). Resistance to the azoles are most dominant and widely reported in literature, due to their extensive clinical use over the past few decades [129]. Four major mechanisms have been reported to induce azole resistance in Candida, with many of these mechanisms being reported in multiple Candida species, and some azole-resistant isolates demonstrating multiple resistance strategies [132]: 1. Point mutations in the gene ERG11 at different “hotspot” regions which confers poor or insufficient azole binding [117,133,134,135,136,137]; 2. Transcriptional upregulation of the gene ERG11 resulting in upregulated ergosterol production [138,139], or amplification of the ERG11 gene [140,141]. Evidence has also suggested that in Candida species, ERG11 may be upregulated by the zinc-cluster transcriptional regulator Upc2p [142,143,144,145]; 3. Overexpression or amplification of multiple classes of drug efflux pumps, including the ABC transporters CDR1 and CDR2 [117,146,147,148] and the major facilitator superfamilies (MFS) proteins, such as transporter genes MDR1 [149,150,151], which may be overexpressed in azole-resistant Candida isolates (C. krusei, is intrinsically resistant to azoles, which is associated with upregulation of efflux pumps [117]); 4. Regulation and induction of numerous cellular stress response pathways such as the phosphatase protein calcineurin, and the molecular chaperone Hsp90 [39,123].
Echinocandin resistance has been linked to mutations occurring within FKS genes, including the essential gene FKS1 [152] and its paralog FKS2 [153]. These FKS genes encode for the catalytic subunit (1,3)-β-D-glucan synthase, which is the target of echinocandin binding [121]. FKS mutations confer a structural alteration which results in poor echinocandin binding [152] and have been reported to occur within two “hotspot regions” within FKS1, the first of which mutates Ser645 to either Tyr645, Pro645, or Phe645; the second hotspot region has been mapped to point mutations occuring between the amino acids 1345 to 365 [154]. In C. albicans, mutations within FKS1 alone are sufficient enough to confer echinocandin resistance. However, within some other Candida species, such as C. glabrata, mutations within both FKS1 and FKS2 have been linked to echinocandin resistance [153]. Echinocandin resistance has also been linked to stress response pathways, including elevated expression levels of the genes CNB1 and MID1 [154], which play a role in the calcineurin cellular stress response signalling pathway [123]. The calcineurin, PKC and HOG pathways have all also been implicated in regulating the increased expression of chitin synthase genes in response to cell wall stress due to echinocandins [39].
Antifungal resistance to polyenes is rare, but has been reported in diverse Candida species, due primarily to alterations in membrane sterol composition [155,156,157,158,159]. In C. albicans, resistance has been reported due to mutations in the C-5,6-desaturase enzyme, which blocks ergosterol biosynthesis and leads to the accumulation of an alternative sterol in the fungal cell membrane [160]. Some Candida species, such as C. lusitaniae and Candida guilliermondii have been reported to demonstrate intrinsic resistance to the polyene amphotericin B [39], which may occur due to mutations in sterol biosynthesis pathways or other components of their cell membrane [128].
4. Candida Morphogenesis, Antifungal Drugs and Drug Resistance
In this section, we highlight the connections between cellular morphogenesis in Candida species, and treatment with antifungal agents. In particular, we analyze how existing antifungal agents are able to modulate cellular morphogenesis; the conserved signaling pathways involved in morphogenesis and the response to antifungal drugs, including antifungal drug resistance pathways; and how Candida biofilms connect fungal morphology to resistance to antifungal drug treatment. Finally, we will discuss novel ‘anti-virulence’ drugs, that specifically target fungal morphogenesis as a means to limit the pathogenicity of Candida infections, and may serve as innovative new strategies to treat fungal infections.
4.1. Effects of Antifungals on Fungal Morphology
The interactions between fungal morphogenesis and exposure to antifungal agents have been previously studied amongst Candida species upon treatment with antifungal drugs. It has been well documented that the azoles—one of the most widespread antifungals—also have a direct effect on C. albicans morphology, and prevent the formation of hyphae (Figure 1a) [161,162]. The effect of azole-mediated hyphal repression is prominent enough that at subinhibitory azole concentrations hyphal branching is restricted, and at clinically relevant doses, the yeast-to-hyphal transition is prevented entirely [161,162]. It was shown that C. albicans cells treated with azoles produced higher levels of the farnesol [163], a fungal-produced quorum sensing molecule with the ability to block filamentous growth in C. albicans. This occurs because azoles target enzymes involved in sterol biosynthesis, and azole treatment leads to a build up of the sterol biosynthetic intermediate, farnesyl pyrophosphate, which indirectly stimulates the over-production of farnesol [163,164]. As previously described, farnesol is able to inhibit filamentous growth in C. albicans by targeting key pathways, such as the cAMP-PKA cascade and via Ubr1-mediated degradation of Cup9 [80], which are involved in the yeast-to-filamentous growth transition [78]. Under anaerobic growth conditions, C. albicans does not produce farnesol, and is more resistant to antifungals, including azoles and amphotericin B, potentially due to reduced carbon flow through the sterol pathway [84], further highlighting the connection between quorum sensing and antifungal resistance. Together this explains a critical connection between a clinically-relevant class of antifungal agents (the azoles), and the repression of morphogenesis—a critical virulence trait in Candida pathogens. This may also explain the success of the azoles as therapeutics.
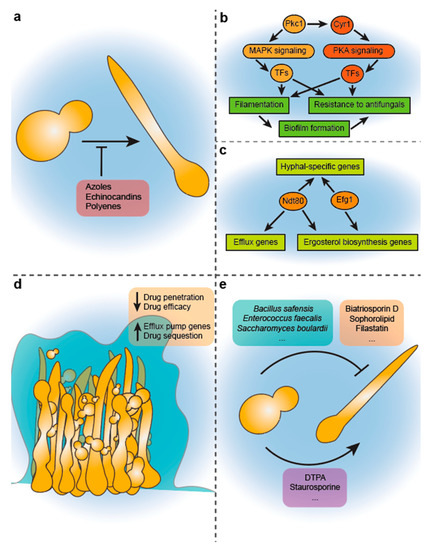
Figure 1.
Linking morphogenesis with antifungal drug treatment and drug resistance in Candida. (a) antifungals and their impact on morphogenesis. Candida morphogenesis can be blocked with treatment with antifungal compounds, including azoles, echinocandins, and polyenes [161,162,165,166,167]. This morphogenetic inhibition is seen even at sub-inhibitory concentrations of all three classes. (b) interactions between signalling pathways that govern morphogenesis and drug resistance. The protein kinase C (PKC)-MAPK pathway and the cAMP-PKA pathway are both involved in drug resistance and morphogenesis in Candida. Depicted are the downstream effects of PKC which regulates MAPK signalling to maintain cell wall integrity when exposed to drugs. Pkc1 has been found to regulate Cyr1 [171], which is a part of the cAMP-PKA signalling pathway, which governs filamentous growth [39]. Pkc1 is implicated in regulation of both drug resistance [172] and hyphal formation [171]. Signaling from the PKA pathway also plays a role in drug resistance via Cyr1 signaling [173,174], as well as morphogenesis [39]. Downstream transcriptional regulators of both MAPK and PKA pathways ultimately regulate morphogenesis and resistance to antifungals. Morphogenesis and filamentous growth also contribute to biofilm formation, which in turn enhances antifungal resistance. (c) transcriptional regulation of antifungal drug resistance and morphogenesis. Transcription factors such as Efg1 and Ndt80 have previously been linked to modulating expression of hyphal specific genes and promoting filamentation [95,96,97,175,176]. Both of these transcriptional regulators are also involved in drug resistance, via regulation of genes involved in drug efflux and/or ergosterol biosynthesis [176,177,178,179,180]; (d) coupling Candida morphogenesis and drug resistance through biofilm formation. Biofilms of polymorphic Candida species are typically composed of diverse cellular morphologies: hyphae, pseudohyphae and yeast [38]. Once established, these polymorphic biofilms secrete an extracellular matrix (ECM) that helps shield Candida from the external environment. Biofilms play a significant role in antifungal drug resistance by preventing antifungal penetration, sequestering antifungals, upregulating antifungal drug efflux, and metabolic regulation that limits antifungal activity [105,181,182,183]; (e) novel compounds for targeting Candida morphogenesis. New compounds are being discovered that can modulate filamentation in Candida species, and may serve as novel therapeutic strategies for treating fungal infections. Examples include drugs (i.e., filastatin), and compounds secreted by competing microbes (i.e., Bacillus safensis) that prevent filamentation, and drugs that enhances filamentation (i.e., diethylenetriaminepentaacetic acid (DTPA)).
Aside from azoles, polyenes and an echinocandin-like antifungal have also been documented to disrupt the yeast-to-hyphal morphogenetic transition in C. albicans (Figure 1a) [165,166,167]. For polyenes, it has been demonstrated that sub-MIC concentrations of amphotericin B can limit the transition between yeast and filamentous growth [165,167]. Similarly, the echinocandin-like antifungal mulundocandin prevents yeast to filamentous growth when administered at sub-MIC levels [165]. In the case of treatment with echinocandins, there are many changes in cellular and cell wall composition that occur in Candida species treated with these antifungals, including increase in cell wall chitin content, decrease in β-glucans, and the upregulation of cell surface proteins [168]. The prevention of hyphal formation by the echinocandin-like drug mulundocandin may be due to these alterations in cell wall composition, resulting in disrupted cellular morphogenesis and cell elongation.
4.2. Fungal Morphogenesis of Drug Resistant Isolates
Aside from antifungal drugs that are able to modulate cellular morphogenesis in C. albicans, there is also a relationship between morphogenesis and resistance to antifungal drugs. It has been observed that there is a correlation between resistance to antifungals and the ability to undergo filamentous growth [161,169]. For azoles, when C. albicans clinical isolates with variable levels of resistance to azole antifungals were grown under filament-inducing conditions, and, in the presence of an azole drug, drug-resistant isolates were able to form hyphae more efficiently than susceptible isolates [161,170]. In C. lusitaniae, there is also an association between resistance to the polyene amphotericin B, and pseudohyphal filamentation: phenotypic switching produces a distinct phenotype (dark brown colony pigmentation on copper sulfate media), which is associated with both pseudohyphal growth as well as an slightly elevated MIC relative to the dominant “light brown” phenotype [50]. Similarly, for echinocandin-resistant C. albicans strains, there is a correlation between resistance and cellular morphogenesis: strains harbouring an fks1 mutation, rendering cells resistant to echinocandins, have defects in filamentous growth [169]. Interestingly, fks1 mutants with impaired filamentation are those with the highest relative levels of cellular chitin in the resistant strains [169], suggesting that echinocandin resistance-mediated over-production of chitin and altered cell-wall composition limits filamentation.
4.3. Shared Cellular Signaling Pathways Mediating Candida Morphogenesis and Drug Resistance
Important links have been discovered between the cellular signalling pathways that modulate morphogenesis in Candida, and those that regulate resistance to antifungal drugs. Indeed, many of the central morphogenetic pathways, such as the cAMP-PKA and MAPK pathways, are additionally involved in antifungal drug resistance (Figure 1b) [39]. For instance, in C. albicans, mutation or deletion of Cyr1 (the adenylate cyclase that synthesizes cAMP) renders cells unable to filament, and hypersensitive to azole antifungals [173,174]. This antifungal sensitivity is conferred, at least in part, due to lack of upregulation of the drug efflux pump Cdr1 in Cyr1-deficient Candida strains [174]. Pkc1, a component of PKC-MAPK signaling in C. albicans is an important regulator of antifungal drug resistance [172]. Recent studies have also discovered functions in the PKC pathway that also regulates filamentation: the protein kinase Pkc1 can function alongside the cAMP-PKA pathway to stimulate activation of Cyr1 to generate cAMP, and induce filamentation [171]. Since Pkc1 is known to interact with actin in S. cerevisiae [184], and Cyr1 also interacts with G-actin [185], it may be that actin is required for Pkc1 to govern morphogenesis in Candida. Other MAPK-related response regulators have similarly been implicated in filamentous growth and drug sensitivity in C. lusitaniae [186].
The essential molecular chaperone Hsp90 is a central regulator of fungal morphogenesis and antifungal drug resistance. Hsp90 is involved in temperature-dependent morphogenesis in C. albicans, such that inhibition of Hsp90 leads to enhanced filamentous growth, via the cAMP-PKA signaling pathway [72]. Inhibition of Hsp90 also blocks the emergence of resistance to the azoles, enhances antifungal sensitivity in azole-resistant strains [187], and reduces tolerance and resistance to echinocandins [123]. Other proteins and pathways involved with Hsp90 signalling are similarly involved in both morphogenesis and antifungal drug resistance, including the Hsp90 client protein and stress response factor calcineurin [123,188,189,190], and the Hsp90 co-chaperone Sgt1 [71]. Hsp90 plays a conserved role in mediating morphogenesis and azole tolerance in C. auris [53]. Additionally, lysine deacetylases (KDACs), including Hos2, Hda1, Rpd3, and Rpd31, are key regulators of both antifungal drug resistance and morphogenesis in C. albicans [191,192], at least in part via their deacetylation and regulation of Hsp90 function [191,192]. The ability of KDACs to mediate drug resistance via Hsp90 is conserved across C. tropicalis and C. guilliermondii [192].
Additional cellular factors have a shared role in the modulation of fungal morphogenesis and susceptibility to antifungal agents. There are several examples of Candida mutant strains that are both defective in filamentous growth, and demonstrate increased susceptibility to antifungal agents. For instance, the C. albicans PMT genes encode a family of protein O-mannosyltransferases that are critical for O-mannosylation of fungal secretory proteins, and are involved in both morphogenesis and antifungal drug resistance. Genetic deletion or depletion of C. albicans PMT genes, including PMT1, PMT2, PMT4, and PMT6 inhibits hyphal formation under diverse environmental growth conditions, and renders cells highly susceptible to cell wall and cell membrane stressors, including the azole antifungal ketoconazole [193,194]. Disruption of the sphingolipid biosynthetic gene IPT1 also renders cells sensitive to azole antifungals, and defective in hyphal formation [195]. This effect is has also been observed with the sphingolipid genes FEN1 and FEN12, although with the polyene amphotericin B instead of azoles [196]. This suggests that several key factors involved in either cell wall modelling or membrane composition, which in turn affects the capacity of the cell to undergo cellular morphogenesis and respond to the presence of antifungal agents.
Factors involved in ergosterol biosynthesis and function are also involved in mediating antifungal drug resistance and morphogenesis. For instance, the squalene epoxidase ERG1, which catalyzes an essential step in the biosynthesis of ergosterol, is involved in resistance to azoles and hyphal formation [197]. C. albicans mutants depleted of ERG1 show abnormal sterol composition, leading to increased sensitivity to both azole and polyene antifungals, along with an inability to form hyphal cells [197]. Similarly, the C. albicans O-acyltransferase GUP1 gene is also involved in mediating drug resistance to the azoles via alterations in membrane ergosterol constitution and distribution, and is strongly implicated in hyphal growth and biofilm formation [198].
Transcription Factors Coupling Morphogenesis and Antifungal Drug Susceptibility
There are also many transcription factors that are recently being understood to control circuitry involved in both Candida morphogenetic programs, as well as antifungal drug resistance (Figure 1c). For instance, in C. albicans, the Zn cluster transcription factors Tac1 and Znc1 are upregulated by farnesol (which blocks C. albicans filamentation), and in turn bind to the promoter of efflux pump genes such as CDR1 and CDR2, leading to their overexpression [199]. This suggests that drug treatment with the azoles, which, as previously described, leads to increased farnesol production [163], limiting filamentous growth, and in turn activating Tac1 and Znc1, can further lead to increase drug efflux and thus resistance. The well-characterized morphogenesis transcription factor Efg1, is also linked to susceptibility to azoles, polyenes and echinocandins in different Candida species (Figure 1c) [177,178]. C. albicans efg1∆ mutant strains downregulate the expression of ERG11, and have decreased levels of ergosterol, resulting in enhanced susceptibility to azoles and polyenes [200]. Other studies have found that in C. parapsilosis, efg1Δ mutants were also susceptible to echinocandins [178].
The transcription factor Ndt80 has also been implicated in resistance to antifungals and morphogenesis in C. albicans (Figure 1c). Ndt80 regulates drug resistance by upregulating expression of the efflux pumps CDR1, CDR2, and MDR1 [176,179] as well as ergosterol biosynthesis genes, such as ERG3 and ERG11 [180]. Ndt80 also plays an essential role in filamentous growth, as this transcriptional regulator interacts with hyphal-specific gene promoters, and upregulation of factors such as HWP1, ECE1, RBT4, and ALS3 [176], and deletion of NDT80 renders C. albicans unable to filament [175].
The transcription factor CAS5 and the coactivator, ADA2 also play a key role in the response to antifungal drugs, and cell morphology. Deletion of these transcription factors renders C. albicans hypersensitive to caspofungin, and both factors are required for expression of numerous caspofungin-responsive genes [201]. Similarly, disruption or deletion of both Ada2 and Cas5 renders C. albicans hypersensitive to fluconazole and enhances azole-mediated killing, via regulation of efflux pumps and ergosterol biosynthesis genes [151,202]. Ada2 and Cas5 are also both important for C. albicans filamentation in vivo, and contribute to fungal virulence [203,204]. Recently, a new role for Cas5 was uncovered in C. albicans, as a key regulator coupling the response to cellular stress, drug resistance, and cell cycle regulation [205], which may help explain the pleiotropic role of this transcription factor.
It is likely that many additional transcriptional regulators are similarly involved in morphogenesis and antifungal susceptibility. For example, Ume6 is a transcription factor that is critical for C. albicans morphogenesis [94,206,207]. Given that Ume6 interacts directly with the cell wall biogenesis protein Sun41 [208], which itself has been implicated in basal caspofungin [209] and fluconazole tolerance [210], it may play a pleiotropic role in morphogenesis as well as susceptibility to antifungal agents. Future research will likely identify additional signaling networks and transcriptional regulators with key functions in coupling antifungal susceptibility with cellular morphogenesis in C. albicans.
4.4. Biofilm Formation and Drug Resistance
One of the key connections between Candida morphogenesis and antifungal drug resistance, is the formation of biofilms (Figure 1d). As previously described, amongst polymorphic Candida species, biofilms are typically comprised of a dense community of yeast and filamentous cell types [37,38,211]. These polymorphic Candida biofilm communities also play a significant role in resistance to antifungal drugs, and confer enhanced drug resistance [12,105,183]. Biofilms provide many mechanisms for drug resistance, including prevention of antifungal penetration, upregulation of antifungal drug efflux pumps, drug sequestration, and the presence of persister cells (Figure 1d) [105,181,182,183].
Biofilms prevent antifungals from penetrating cells due to the chemical composition of the thick extracellular matrix (ECM), allowing cells to grow and propagate within a biofilm without interference [105]. Matrix polymers in the ECM such as polysaccharides, including β-1,3-glucan, β-1,6-glucan, mannans, and proteins, are thought to confer antifungal resistance to biofilms as they prevent deep penetration of antifungals into the biofilm [105,107]. It was recently found that in C. albicans, biofilms produce extracellular vesicles (EVs) that play a central role in biofilm matrix production and resistance to antifungal drugs [210]. EVs, which are membranous compartments released by almost all cells, are commonly found amongst fungi, and carry proteins, lipids, polysaccharides, and RNA [212,213,214]. In C. albicans, EVs produced by planktonic cells versus those in biofilms, have a unique proteomic composition; biofilm-associated EV composition is very similar to the protein and polysaccharide composition of the ECMs, including a significant amount of mannan and glucan [210]. It has recently been shown that C. albicans mutants defective in the ESCRT (endosomal sorting complexes required for transport) pathway, have decreased vesicle production and are hypersusceptible to fluconazole during biofilm growth [210], likely due to decreased vesicle delivery of ECM materials (i.e., mannan and glucan), and thus decreased ECM-associated drug sequestration. This novel finding reinforces the critical link between fungal biofilm ECM and antifungal drug resistance, and suggests ESCRT proteins as putative targets to limit drug resistance in C. albicans biofilms.
If drugs bypass this defense and enter the biofilm-associated cells, upregulated efflux pumps force antifungal drugs out of the cells, to prevent any further damage to Candida [215]. Studies have shown that while efflux pumps have minimal effects on mature biofilms, during the early phases of biofilm formation, efflux pumps play a significant role in antifungal resistance and are significantly upregulated in early biofilms [105]. Drugs that were not pumped out by efflux pumps and remain in the cell undergo drug sequestration in which the drugs are compartmentalized in the cell in less harmful locations [105]. Drug sequestration of azoles, echinocandins, pyrimidines, and polyenes is regulated by β-1,3 glucan, a major carbohydrate component of Candida biofilms [105].
Persister cells are a type of metabolically dormant cell that may arise in biofilms, and which may be more tolerant to antifungal drug treatment. These persister cells may be involved in antifungal tolerance and recalcitrant infections [181]. These metabolically dormant Candida cells are found in biofilms [216,217], and this allows these biofilms to be more tolerant to antifungal drugs, compared to planktonic fungal cells [181,216,217,218]. Typically, fungicidal drugs induce cell death via the production of reactive oxygen species (ROS) [126]; however, persister cells are capable of dealing with larger quantities of ROS, and survive via activation of an oxidative stress response [181]. This response involves an overexpression of superoxide dismutases (SODs), which assist in protecting cells from ROS and allows biofilm-associated cells to survive in the presence of antifungal drugs [181].
While Candida biofilms are, in general, more resistant to antifungals, they are particularly resistant to the azole class of antifungals [106], and are generally more susceptible to the echinocandins and liposomal formulations of the polyene amphotericin B [219,220]. This may be due to several factors, including the fact that Candida biofilms have an altered sterol composition relative to planktonic cells [221], which may disproportionately affect resistance to the sterol biosynthesis-targeting azoles. Biofilm cells also overexpress the target of the azoles, ERG11 [222], which may promote azole-specific resistance. Additionally, Candida cells in biofilms overexpress drug-efflux pumps [106]; while this may increase overall resistance to antifungals, resistance to azoles is uniquely affected by the overexpression of the MFS transporter MDR1 in biofilms [223,224,225,226].
4.5. New Antivirulence Compounds That Target Candida Morphogenesis
We have established that there is a connection between antifungal drugs and Candida morphogenesis: existing antifungal agents can modulate filamentation (and thus virulence), and many signaling pathways are shared between those involved in cellular morphology, and those involved in the response and resistance to antifungal agents. Here, we focus on how anti-morphogenesis compounds can be harnessed as novel therapeutics for the treatment of fungal pathogens (Table 2).

Table 2.
Compounds that alter Candida morphogenesis.
Given the limited number of clinically-available antifungal drugs, increasing rates of antifungal drug resistance, and emergence of multi-drug resistant pathogens, such as C. auris [32], there is a renewed urgency for the development of new strategies to tackle fungal pathogens. As described earlier, existing antifungal drugs create a very strong selection pressure, and Candida species are able to evolve resistance to antifungal drugs via multiple mechanisms [227]. Recently, there has been a research emphasis on identifying and characterizing drugs that target virulence processes, such as fungal morphogenesis, that may limit or inhibit pathogenicity of a microbial organism, while still allowing it to live (Figure 1e) [228,229,230,231]. It is suggested that such anti-virulence compounds will create a weaker selection pressure on the microbes, thus preventing selection for the rapid evolution of drug resistance. Furthermore, by allowing survival of their microbial targets, anti-virulence compounds would theoretically be less disruptive to the ecological balance of the host microbiome.
In the case of C. albicans, the inhibition of morphogenesis is an excellent target for anti-virulence drugs (Table 2), given that cellular morphogenesis is coupled with fungal virulence and inhibiting the process may aid in better containment and clearance by the host’s own immune system [78,232]. As a result, several screens have been conducted to identify new small-molecule inhibitors that target the morphogenetic transition in Candida pathogens [78,233,234,235,236,237,238]. The majority of these experiments have focused on treatment of Candida strains with the small molecules under conditions which promote filamentous growth, followed by either a physical or genetic assessment of hyphal formation. Here, we will review some key findings of novel, anti-morphogenesis inhibitors in Candida pathogens (Figure 1e), with potential applications as innovative therapeutic strategies for limiting the virulence of these pathogens.
Zhang et al. recently discovered that C. albicans treated with the small phenolic compound Biatriosporin D prevents yeast-to-hyphal morphogenesis [239]. It does so by downregulating the expression of the hyphal-specific genes ALS3, HWP1, and ECE1 by directly affecting the cAMP-PKA pathway via the regulation of the yeast’s intracellular cAMP levels. Cells treated with Biatriosporin D showed a dose-dependent decrease in intracellular cAMP levels and exogenous cAMP can partially restore hyphal formation. Biatriosporin D was also found to upregulate the expression of Dpp3, which is involved in the synthesis of the hyphal-inhibiting quorum-sensing molecule farnesol [239,240]. The combination of increased farnesol production, decreased cAMP levels, and low toxicity, renders Biatriosporin D a strong inhibitor of filamentation, and an emerging therapeutic candidate.
Some compounds, such as the glycolipid sophorolipid, has previously shown antimicrobial activity [248]. Haque et al. have investigated the antifungal effects on planktonic Candida spp. and also demonstrated that sophorolipid prevents hyphal formation [241]. Upon treatment with sophorolipid, biofilm formation was significantly disrupted and the expression of hyphal-specific genes (HWP1, ALS1, ALS3, ECE1 and SAP4), were downregulated [241]. This downregulation of hyphal genes abrogated the capacity of C. albicans to form biofilms, as well as increased the efficacy of treatment with other antifungals, implying its potential utility in combination with antifungal treatment.
Recently, Romo et al. identified a novel group of bioactive compounds with a common biaryl amide core structure that were potent inhibitors of filamentation in C. albicans. Genetic analysis suggests that these inhibitors function downstream of multiple signalling pathways and are likely impacting filamentation via the transcription factor Brg1. After confirming the ability of these compounds to inhibit filamentation and biofilm growth in vitro, this group performed in vivo studies using mouse models of disseminated and oropharyngeal candidiasis. The hyphal-inhibiting drugs were able to increase mouse survival, and inhibit filamentation in vivo, as demonstrated by lack of hyphal cells in histological samples of kidneys from infected mice [237]. Importantly, while these drugs blocked hyphal growth, they did not impact fungal survival, as overall fungal burdens remained similar between treated and untreated mice [237].
Another potent inhibitor of Candida filamentation is the compound filastatin, which blocks the yeast-to-hyphal morphological transition in C. albicans, and also inhibits cellular adhesion, biofilm formation, and pathogenesis [238]. Filastatin inhibits filamentation in diverse environmental conditions, in both liquid and solid growth media [238]. Genetic pathway analysis was performed to identify the mechanism of action of this inhibitor, and revealed that it likely acts downstream of two key transcriptional regulators: Efg1 and Cph1 [238]. These transcription factors are the terminal transcriptional regulators of the cAMP-PKA pathway and MAPK pathway, respectively, and both are key regulators of filamentous growth in C. albicans. Another recent report used similar chemical-genetic profiling to identify the function of hyphal inhibitors Niclosamide and TCSA (Tri-Chloro-Salicyanilide) [233]. These small molecule inhibitors, which block hyphal formation in C. albicans and inhibit biofilm growth of C. albicans and C. auris, were similarly found to act on effectors downstream of cAMP-PKA and MAPK signalling pathways [233].
In addition to small molecule inhibitors of filamentation identified through drug screening assays, another relevant reservoir for anti-morphogenesis compounds is other microorganisms which may coexist and compete with Candida species (Figure 1e). Previous work has shown that bacteria such as Bacillus safensis can produce a chitinase compound with antipathogenic capabilities which can specifically target C. albicans’ capacity to form functional hyphae [249]. B. safensis is able to directly bind to C. albicans and secrete a chitinase that can destabilize the cell wall and prevent proper hyphal formation. This leads to the antivirulent effect of halting C. albicans’ morphological transition from yeast to hyphal growth, and also efficiently preventing biofilm formation. Similarly, Enterococcus faecalis bacteria produce the EntV bacteriocin, which is able to inhibit hyphal morphogenesis, biofilm formation, and virulence in C. albicans [243]. Many Lactobacillus species are able to inhibit hyphal formation of C. albicans through the production of organic acids that lower the pH of the surrounding microenvironment, promoting yeast-form growth [250], and Escherichia coli can modulate biofilm formation and hyphal growth of diverse Candida pathogens [251]. Candida species can also regulate hyphal formation of other competing Candida organisms: C. krusei and C. glabrata can prevent hyphal formation in C. albicans, resulting in downregulation of HWP1 expression [252]. The yeast species Saccharomyces boulardii shows similar anti-virulence effects on C. albicans, and can block the yeast to hyphal growth transition via the secretion of capric acid [244,253]. Identifying additional factors secreted by competing microorganisms may provide a practical method to identify new drugs to inhibit Candida morphogenesis and virulence.
4.6. Inhibitors of Morphogenesis and Antifungal Drug Resistance
Interestingly, some small molecules have been identified that are able to both inhibit morphogenesis, as well as block resistance to antifungal drugs, or sensitize drug-resistant fungal lineages. As described above, there are many overlapping cellular pathways in Candida that mediate both morphogenesis, as well as resistance to antifungal drugs. As such, identifying drugs that target these pathways may be a powerful strategy for inhibiting virulence and treating drug-resistant fungal infections. For example, recently, it was found that halogenated salicylanilides compounds elicit overexpression of drug resistance genes and are also capable of inhibiting morphogenesis in C. albicans [233]. For example, the drug Niclosamide was able to inhibit hyphal formation in both wild-type and drug-resistant C. albicans lineages, and transcriptome analysis showed that strains exposed to this drug upregulated genes responsible for drug efflux, such as CDR1, MDR1 and TPO3, while also downregulating genes responsible for filamentous growth.
The natural product beauvericin has been shown to block the morphogenetic transition between yeast and filamentous growth in C. albicans via the transcription factor Brg1, while simultaneously potentiating the activity of the azole antifungals against azole resistant isolates of C. albicans and C. glabrata [245]. This potentiation of azole activity and sensitization of drug-resistant Candida isolates occurs by inhibition of multidrug efflux via the Cdr1 efflux pump [245]. Similarly, the natural product staurosporine enhances the activity of both the azole and echinocandin antifungals [172], and regulates fungal morphogenesis [246]. Interestingly, staurosporine induces, as oppose to represses, C. albicans morphogenesis, via cAMP-PKA signaling (Figure 1e) [246]. This is reminiscent of the effect of inhibiting HSP90 in C. albicans, as described above, which both enhances antifungal activity, and induces filamentous growth [72,123,187,254]. Given that Hsp90 is such a highly-connected protein, with many chaperone client proteins and a large interaction network [255], it is likely that depletion of this interconnected protein affects multiple downstream cellular alterations, resulting in pleiotropic effects. For instance, Hsp90 regulates morphogenesis via its interactions with the cAMP-PKA signaling pathway, the cyclin-dependent kinase Cdc28, and other signaling pathways [72,74,256,257]. Through its interaction with the protein phosphatase calcineurin, Hsp90 is able to modulate antifungal drug resistance in C. albicans [123,187,188]. Hsp90 also interacts with co-chaperones, such as Sgt1 and Cdc37 [71,258], which impact its role in morphogenesis and drug susceptibility. For example, Cdc37, via its interaction with the kinase Crk1, plays a crucial role in C. albicans morphogenesis [259]. Inhibitors which affect both fungal morphogenesis and resistance to antifungal drugs hold great promise as novel therapeutics against fungal infections, or for use in combination with existing antifungal agents to provide more effective treatment of drug-resistant infections.
Finally, the broad-spectrum chelator diethylenetriaminepentaacetic acid (DTPA) was recently discovered to strongly induce filamentation, while also enhancing the susceptibility of C. albicans to antifungal drugs [247]. DTPA causes robust filamentation of C. albicans, even in non-hyphal inducing cues, by depleting zinc. This filamentation occurs in a manner that is dependent upon the cAMP-PKA signalling pathway, as well as additional transcription factors Brg1 and Rob1 [247]. DTPA also increases fungal susceptibility to echinocandin antifungals and is able to enhance the efficacy of caspofungin against echinocandin-resistant C. albicans strains both in vitro and in vivo. DTPA’s role in antifungal susceptibility occurs via chelation of magnesium; it acts to modulate Hog1-MAPK signaling and enhance echinocandin activity [247].
5. Conclusions
It is clear that, amongst Candida pathogens, there are important connections between antifungal drug treatment, cellular morphogenesis, and resistance to antifungal drugs. It may not be surprising that such a connection exists, given that a majority of antifungal agents target the fungal membrane or cell wall, and that cell wall remodelling is an important component of fungal morphogenesis and the yeast-to-hyphal growth transition. Given the important role of Candida filamentation in fungal virulence, and the numerous shared pathways governing filamentation and susceptibility to antifungal agents, targeting these shared signaling pathways may be a powerful strategy to enhance antifungal efficacy while limiting fungal virulence. Indeed, novel anti-virulence compounds that target Candida morphogenesis are promising alternatives or complements to the current antifungal arsenal. Future research on fungal signaling pathways that couple morphogenesis with susceptibility to antifungal drugs may help identify new cellular targets for the treatment of life-threatening invasive fungal infections, and the development of novel clinical solutions.
Author Contributions
J.S., S.R., I.R., and R.S.S. each contributed to researching, writing, and editing this manuscript.
Funding
R.S.S. is supported by a CIHR Priority Grant on Antimicrobial Resistance, and a Banting Research Foundation Discovery Award.
Acknowledgments
We would like to thank all Shapiro lab members for helpful discussion. R.S.S. is supported by a Canadian Institutes of Health Research (CIHR) Priority Grant on Antimicrobial Resistance, and a Banting Research Foundation Discovery Award.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Fisher, M.C.; Henk, D.A.; Briggs, C.J.; Brownstein, J.S.; Madoff, L.C.; McCraw, S.L.; Gurr, S.J. Emerging fungal threats to animal, plant and ecosystem health. Nature 2012, 484, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Taylor, L.H.; Latham, S.M.; Woolhouse, M.E. Risk factors for human disease emergence. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2001, 356, 983–989. [Google Scholar] [CrossRef] [PubMed]
- Köhler, J.R.; Casadevall, A.; Perfect, J. The spectrum of fungi that infects humans. Cold Spring Harb. Perspect. Med. 2014, 5, a019273. [Google Scholar] [CrossRef] [PubMed]
- Bongomin, F.; Gago, S.; Oladele, R.O.; Denning, D.W. Global and multi-national prevalence of fungal diseases-estimate precision. J. Fungi (Basel) 2017, 3, 57. [Google Scholar] [CrossRef] [PubMed]
- Doi, A.M.; Pignatari, A.C.C.; Edmond, M.B.; Marra, A.R.; Camargo, L.F.A.; Siqueira, R.A.; da Mota, V.P.; Colombo, A.L. Epidemiology and microbiologic characterization of nosocomial candidemia from a Brazilian national surveillance program. PLoS ONE 2016, 11, e0146909. [Google Scholar] [CrossRef] [PubMed]
- Chatterjee, S.; Alampalli, S.V.; Nageshan, R.K.; Chettiar, S.T.; Joshi, S.; Tatu, U.S. Draft genome of a commonly misdiagnosed multidrug resistant pathogen Candida auris. BMC Genomics 2015, 16, 686. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.F.; Liu, W.L.; Hsieh, M.H.; Hii, I.M.; Lee, Y.L.; Lin, Y.T.; Ho, M.W.; Liu, C.E.; Chen, Y.H.; Wang, F.D. Epidemiology and antifungal susceptibility of candidemia isolates of non-albicans Candida species from cancer patients. Emerg. Microbes Infect. 2017, 6, e87. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Lionakis, M.S.; Arendrup, M.C.; Ostrosky-Zeichner, L.; Kullberg, B.J. Invasive candidiasis. Nat. Rev. Dis. Primers 2018, 4, 18026. [Google Scholar] [CrossRef] [PubMed]
- Lionakis, M.S.; Levitz, S.M. Host control of fungal infections: lessons from basic studies and human cohorts. Annu. Rev. Immunol. 2018, 36, 157–191. [Google Scholar] [CrossRef] [PubMed]
- Yapar, N. Epidemiology and risk factors for invasive candidiasis. Ther. Clin. Risk Manag. 2014, 10, 95. [Google Scholar] [CrossRef] [PubMed]
- Desai, J.V.; van de Veerdonk, F.L.; Lionakis, M.S. Understanding the role of host immune responses in invasive candidiasis. Intensive Care Med. 2018, 44, 1310–1314. [Google Scholar] [CrossRef] [PubMed]
- Tsui, C.; Kong, E.F.; Jabra-Rizk, M.A. Pathogenesis of Candida albicans biofilm. Pathog. Dis. 2016, 74, ftw018. [Google Scholar] [CrossRef] [PubMed]
- Arendrup, M.C. Candida and candidaemia. Susceptibility and epidemiology. Dan. Med. J. 2013, 60, B4698. [Google Scholar] [PubMed]
- Lamoth, F.; Lockhart, S.R.; Berkow, E.L.; Calandra, T. Changes in the epidemiological landscape of invasive candidiasis. J. Antimicrob. Chemother. 2018, 73, i4–i13. [Google Scholar] [CrossRef] [PubMed]
- Sellam, A.; Whiteway, M. Recent advances on biology and virulence. F1000Res. 2016, 5, 2582. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Wilson, D.; Hube, B. Candida albicans pathogenicity mechanisms. Virulence 2013, 4, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Fidel, P.L., Jr.; Vazquez, J.A.; Sobel, J.D. Candida glabrata: review of epidemiology, pathogenesis, and clinical disease with comparison to C. albicans. Clin. Microbiol. Rev. 1999, 12, 80–96. [Google Scholar] [CrossRef] [PubMed]
- Deorukhkar, S.C.; Saini, S.; Mathew, S. Virulence Factors Contributing to pathogenicity of Candida tropicalis and its antifungal susceptibility profile. Int. J. Microbiol. 2014, 2014, 456878. [Google Scholar] [CrossRef]
- Zuza-Alves, D.L.; Silva-Rocha, W.P.; Chaves, G.M. An update on based on basic and clinical approaches. Front. Microbiol. 2017, 8, 1927. [Google Scholar] [CrossRef]
- Neji, S.; Hadrich, I.; Trabelsi, H.; Abbes, S.; Cheikhrouhou, F.; Sellami, H.; Makni, F.; Ayadi, A. Virulence factors, antifungal susceptibility and molecular mechanisms of azole resistance among Candida parapsilosis complex isolates recovered from clinical specimens. J. Biomed. Sci. 2017, 24, 67. [Google Scholar] [CrossRef]
- Richardson, J.; Ho, J.; Naglik, J. Candida–Epithelial Interactions. J. Fungi 2018, 4, 22. [Google Scholar] [CrossRef] [PubMed]
- Turner, S.A.; Butler, G. The Candida pathogenic species complex. Cold Spring Harb. Perspect. Med. 2014, 4, a019778. [Google Scholar] [CrossRef]
- Priest, S.J.; Lorenz, M.C. Characterization of virulence-related phenotypes in Candida species of the CUG Clade. Eukaryot. Cell 2015, 14, 931–940. [Google Scholar] [CrossRef]
- Samaranayake, Y.H.; Samaranayake, L.P. Candida krusei: biology, epidemiology, pathogenicity and clinical manifestations of an emerging pathogen. J. Med. Microbiol. 1994, 41, 295–310. [Google Scholar] [CrossRef]
- Larkin, E.; Hager, C.; Chandra, J.; Mukherjee, P.K.; Retuerto, M.; Salem, I.; Long, L.; Isham, N.; Kovanda, L.; Borroto-Esoda, K.; et al. The emerging pathogen Candida auris: growth phenotype, virulence factors, activity of antifungals, and effect of SCY-078, a novel glucan synthesis inhibitor, on growth morphology and biofilm formation. Antimicrob. Agents Chemother. 2017, 61, e02396-16. [Google Scholar] [CrossRef]
- Yue, H.; Bing, J.; Zheng, Q.; Zhang, Y.; Hu, T.; Du, H.; Wang, H.; Huang, G. Filamentation in Candida auris, an emerging fungal pathogen of humans: passage through the mammalian body induces a heritable phenotypic switch. Emerg. Microbes Infect. 2018, 7, 188. [Google Scholar] [CrossRef] [PubMed]
- Silva-Dias, A.; Miranda, I.M.; Branco, J.; Monteiro-Soares, M.; Pina-Vaz, C.; Rodrigues, A.G. Adhesion, biofilm formation, cell surface hydrophobicity, and antifungal planktonic susceptibility: Relationship among Candida spp. Front. Microbiol. 2015, 6, 205. [Google Scholar] [CrossRef]
- Sullivan, D.J.; Moran, G.P.; Coleman, D.C. Candida dubliniensis: ten years on. FEMS Microbiol. Lett. 2005, 253, 9–17. [Google Scholar] [CrossRef] [PubMed]
- Jabra-Rizk, M.A.; Falkler, W.A.; Meiller, T.F. Fungal biofilms and drug resistance. Emerg. Infect. Dis. 2004, 10, 14–19. [Google Scholar] [CrossRef]
- Arendrup, M.C.; Patterson, T.F. Multidrug-resistant Candida: epidemiology, molecular mechanisms, and treatment. J. Infect. Dis. 2017, 216, S445–S451. [Google Scholar] [CrossRef] [PubMed]
- Brandt, M.E.; Lockhart, S.R. Recent taxonomic developments with Candida and other opportunistic yeasts. Curr. Fungal Infect. Rep. 2012, 6, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Geddes-McAlister, J.; Shapiro, R.S. New pathogens, new tricks: emerging, drug-resistant fungal pathogens and future prospects for antifungal therapeutics. Ann. N. Y. Acad. Sci. 2018, 1435, 57–78. [Google Scholar] [CrossRef]
- Kumamoto, C.A.; Vinces, M.D. Contributions of hyphae and hypha-co-regulated genes to Candida albicans virulence. Cell. Microbiol. 2005, 7, 1546–1554. [Google Scholar] [CrossRef] [PubMed]
- Munro, C. Candida albicans cell wall mediated virulence. In Pathogenic Yeasts; Springer: Dordrecht, Germany, 2009; pp. 69–95. [Google Scholar]
- Canteri de Souza, P.; Custódio Caloni, C.; Wilson, D.; Sergio Almeida, R. An invertebrate host to study fungal infections, Mycotoxins and Antifungal Drugs. J. Fungi (Basel) 2018, 4, 125. [Google Scholar] [CrossRef] [PubMed]
- Whiteway, M.; Oberholzer, U. Candida morphogenesis and host-pathogen interactions. Curr. Opin. Microbiol. 2004, 7, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.; Negri, M.; Henriques, M.; Oliveira, R.; Williams, D.W.; Azeredo, J. Adherence and biofilm formation of non-Candida albicans Candida species. Trends Microbiol. 2011, 19, 241–247. [Google Scholar] [CrossRef] [PubMed]
- Nobile, C.J.; Johnson, A.D. Candida albicans Biofilms and Human Disease. Annu. Rev. Microbiol. 2015, 69, 71–92. [Google Scholar] [CrossRef]
- Shapiro, R.S.; Robbins, N.; Cowen, L.E. Regulatory circuitry governing fungal development, drug resistance, and disease. Microbiol. Mol. Biol. Rev. 2011, 75, 213–267. [Google Scholar] [CrossRef]
- Cowen, L.E.; Sanglard, D.; Howard, S.J.; Rogers, P.D.; Perlin, D.S. Mechanisms of antifungal drug resistance. Cold Spring Harb. Perspect. Med. 2014, 5, a019752. [Google Scholar] [CrossRef]
- Desai, J. Candida albicans hyphae: from growth initiation to invasion. J. Fungi 2018, 4, 10. [Google Scholar] [CrossRef]
- Sudbery, P.; Gow, N.; Berman, J. The distinct morphogenic states of Candida albicans. Trends Microbiol. 2004, 12, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Slaughter, B.D.; Smith, S.E.; Li, R. Symmetry breaking in the life cycle of the budding yeast. Cold Spring Harb. Perspect. Biol. 2009, 1, a003384. [Google Scholar] [CrossRef] [PubMed]
- Gale, C.; Gerami-Nejad, M.; McClellan, M.; Vandoninck, S.; Longtine, M.S.; Berman, J. Candida albicans Int1p interacts with the septin ring in yeast and hyphal cells. Mol. Biol. Cell 2001, 12, 3538–3549. [Google Scholar] [CrossRef] [PubMed]
- Sudbery, P.E. Growth of Candida albicans hyphae. Nat. Rev. Microbiol. 2011, 9, 737–748. [Google Scholar] [CrossRef]
- Veses, V.; Gow, N.A.R. Pseudohypha budding patterns of Candida albicans. Med. Mycol. 2009, 47, 268–275. [Google Scholar] [CrossRef] [PubMed]
- Porman, A.M.; Hirakawa, M.P.; Jones, S.K.; Wang, N.; Bennett, R.J. MTL-independent phenotypic switching in Candida tropicalis and a dual role for Wor1 in regulating switching and filamentation. PLoS Genet. 2013, 9, e1003369. [Google Scholar] [CrossRef]
- Martins, M.; Henriques, M.; Azeredo, J.; Rocha, S.M.; Coimbra, M.A.; Oliveira, R. Morphogenesis control in Candida albicans and Candida dubliniensis through signaling molecules produced by planktonic and biofilm cells. Eukaryot. Cell 2007, 6, 2429–2436. [Google Scholar] [CrossRef]
- Laffey, S.F.; Butler, G. Phenotype switching affects biofilm formation by Candida parapsilosis. Microbiology 2005, 151, 1073–1081. [Google Scholar] [CrossRef]
- Miller, N.S.; Dick, J.D.; Merz, W.G. Phenotypic switching in Candida lusitaniae on copper sulfate indicator agar: association with amphotericin B resistance and filamentation. J. Clin. Microbiol. 2006, 44, 1536–1539. [Google Scholar] [CrossRef]
- Kumar, A.; Prakash, A.; Singh, A.; Kumar, H.; Hagen, F.; Meis, J.F.; Chowdhary, A. Candida haemulonii species complex: an emerging species in India and its genetic diversity assessed with multilocus sequence and amplified fragment-length polymorphism analyses. Emerg. Microbes Infect. 2016, 5, e49. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Bing, J.; Zheng, Q.; Zhang, F.; Liu, J.; Yue, H.; Tao, L.; Du, H.; Wang, Y.; Wang, H.; et al. The first isolate of Candida auris in China: clinical and biological aspects. Emerg. Microbes Infect. 2018, 7, 93. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.H.; Iyer, K.R.; Pardeshi, L.; Muñoz, J.F.; Robbins, N.; Cuomo, C.A.; Wong, K.H.; Cowen, L.E. Genetic analysis of Candida auris implicates Hsp90 in morphogenesis and azole tolerance and Cdr1 in azole resistance. MBio 2019, 10, e02529-18. [Google Scholar] [CrossRef] [PubMed]
- Noble, S.M.; French, S.; Kohn, L.A.; Chen, V.; Johnson, A.D. Systematic screens of a Candida albicans homozygous deletion library decouple morphogenetic switching and pathogenicity. Nat. Genet. 2010, 42, 590–598. [Google Scholar] [CrossRef] [PubMed]
- Saville, S.P.; Lazzell, A.L.; Monteagudo, C.; Lopez-Ribot, J.L. Engineered control of cell morphology in vivo reveals distinct roles for yeast and filamentous forms of Candida albicans during infection. Eukaryot. Cell 2003, 2, 1053–1060. [Google Scholar] [CrossRef] [PubMed]
- El-Kirat-Chatel, S.; Dufrêne, Y.F. Nanoscale imaging of the Candida-macrophage interaction using correlated fluorescence-atomic force microscopy. ACS Nano 2012, 6, 10792–10799. [Google Scholar] [CrossRef] [PubMed]
- Westman, J.; Moran, G.; Mogavero, S.; Hube, B.; Grinstein, S. Candida albicans hyphal expansion causes phagosomal membrane damage and luminal alkalinization. MBio 2018, 9, e01226-18. [Google Scholar] [CrossRef]
- Moyes, D.L.; Wilson, D.; Richardson, J.P.; Mogavero, S.; Tang, S.X.; Wernecke, J.; Höfs, S.; Gratacap, R.L.; Robbins, J.; Runglall, M.; et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 2016, 532, 64–68. [Google Scholar] [CrossRef]
- Richardson, J.P.; Mogavero, S.; Moyes, D.L.; Blagojevic, M.; Krüger, T.; Verma, A.H.; Coleman, B.M.; De La Cruz Diaz, J.; Schulz, D.; Ponde, N.O.; et al. Processing of Ece1p is critical for candidalysin maturation and fungal virulence. MBio 2018, 9, e02178-17. [Google Scholar] [CrossRef]
- Watts, H.J.; Véry, A.A.; Perera, T.H.; Davies, J.M.; Gow, N.A. Thigmotropism and stretch-activated channels in the pathogenic fungus Candida albicans. Microbiology 1998, 144, 689–695. [Google Scholar] [CrossRef]
- Lo, H.J.; Köhler, J.R.; DiDomenico, B.; Loebenberg, D.; Cacciapuoti, A.; Fink, G.R. Nonfilamentous C. albicans mutants are avirulent. Cell 1997, 90, 939–949. [Google Scholar] [CrossRef]
- Murad, A.M.; Leng, P.; Straffon, M.; Wishart, J.; Macaskill, S.; MacCallum, D.; Schnell, N.; Talibi, D.; Marechal, D.; Tekaia, F.; et al. NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans. EMBO J. 2001, 20, 4742–4752. [Google Scholar] [CrossRef] [PubMed]
- Thompson, D.S.; Carlisle, P.L.; Kadosh, D. Coevolution of morphology and virulence in Candida species. Eukaryot. Cell 2011, 10, 1173–1182. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, R.S.; Ryan, O.; Boone, C.; Cowen, L.E. Regulatory circuitry governing morphogenesis in Saccharomyces cerevisiae and Candida albicans. Cell Cycle 2012, 11, 4294–4295. [Google Scholar] [CrossRef] [PubMed]
- Gow, N.A.R.; van de Veerdonk, F.L.; Brown, A.J.P.; Netea, M.G. Candida albicans morphogenesis and host defence: Discriminating invasion from colonization. Nat. Rev. Microbiol. 2011, 10, 112–122. [Google Scholar] [CrossRef] [PubMed]
- Whiteway, M.; Bachewich, C. Morphogenesis in Candida albicans. Annu. Rev. Microbiol. 2007, 61, 529–553. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.J.; Gow, N.A. Regulatory networks controlling Candida albicans morphogenesis. Trends Microbiol. 1999, 7, 333–338. [Google Scholar] [CrossRef]
- Monge, R.A.; Román, E.; Nombela, C.; Pla, J. The MAP kinase signal transduction network in Candida albicans. Microbiology 2006, 152, 905–912. [Google Scholar] [CrossRef]
- Kumamoto, C.A. A contact-activated kinase signals Candida albicans invasive growth and biofilm development. Proc. Natl. Acad. Sci. USA 2005, 102, 5576–5581. [Google Scholar] [CrossRef]
- Alonso-Monge, R.; Navarro-García, F.; Molero, G.; Diez-Orejas, R.; Gustin, M.; Pla, J.; Sánchez, M.; Nombela, C. Role of the mitogen-activated protein kinase Hog1p in morphogenesis and virulence of Candida albicans. J. Bacteriol. 1999, 181, 3058–3068. [Google Scholar]
- Shapiro, R.S.; Zaas, A.K.; Betancourt-Quiroz, M.; Perfect, J.R.; Cowen, L.E. The Hsp90 co-chaperone Sgt1 governs Candida albicans morphogenesis and drug resistance. PLoS ONE 2012, 7, e44734. [Google Scholar] [CrossRef]
- Shapiro, R.S.; Uppuluri, P.; Zaas, A.K.; Collins, C.; Senn, H.; Perfect, J.R.; Heitman, J.; Cowen, L.E. Hsp90 orchestrates temperature-dependent Candida albicans morphogenesis via Ras1-PKA signaling. Curr. Biol. 2009, 19, 621–629. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, R.S.; Cowen, L. Coupling temperature sensing and development: Hsp90 regulates morphogenetic signalling in Candida albicans. Virulence 2010, 1, 45–48. [Google Scholar] [CrossRef]
- Veri, A.O.; Miao, Z.; Shapiro, R.S.; Tebbji, F.; O’Meara, T.R.; Kim, S.H.; Colazo, J.; Tan, K.; Vyas, V.K.; Whiteway, M.; et al. Tuning Hsf1 levels drives distinct fungal morphogenetic programs with depletion impairing Hsp90 function and overexpression expanding the target space. PLoS Genet. 2018, 14, e1007270. [Google Scholar] [CrossRef]
- Davis-Hanna, A.; Piispanen, A.E.; Stateva, L.I.; Hogan, D.A. Farnesol and dodecanol effects on the Candida albicans Ras1-cAMP signalling pathway and the regulation of morphogenesis. Mol. Microbiol. 2007, 67, 47–62. [Google Scholar] [CrossRef] [PubMed]
- Hornby, J.M.; Jensen, E.C.; Lisec, A.D.; Tasto, J.J.; Jahnke, B.; Shoemaker, R.; Dussault, P.; Nickerson, K.W. Quorum sensing in the dimorphic fungus Candida albicans is mediated by farnesol. Appl. Environ. Microbiol. 2001, 67, 2982–2992. [Google Scholar] [CrossRef] [PubMed]
- Nickerson, K.W.; Atkin, A.L. Deciphering fungal dimorphism: Farnesol’s unanswered questions. Mol. Microbiol. 2017, 103, 567–575. [Google Scholar] [CrossRef] [PubMed]
- Shareck, J.; Belhumeur, P. Modulation of morphogenesis in Candida albicans by various small molecules. Eukaryot. Cell 2011, 10, 1004–1012. [Google Scholar] [CrossRef]
- Piispanen, A.E.; Bonnefoi, O.; Carden, S.; Deveau, A.; Bassilana, M.; Hogan, D.A. Roles of Ras1 membrane localization during Candida albicans hyphal growth and farnesol response. Eukaryot. Cell 2011, 10, 1473–1484. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Su, C.; Unoje, O.; Liu, H. Quorum sensing controls hyphal initiation in Candida albicans through Ubr1-mediated protein degradation. Proc. Natl. Acad. Sci. USA 2014, 111, 1975–1980. [Google Scholar] [CrossRef]
- Lu, Y.; Su, C.; Liu, H. Candida albicans hyphal initiation and elongation. Trends Microbiol. 2014, 22, 707–714. [Google Scholar] [CrossRef]
- Meir, J.; Hartmann, E.; Eckstein, M.T.; Guiducci, E.; Kirchner, F.; Rosenwald, A.; LeibundGut-Landmann, S.; Pérez, J.C. Identification of Candida albicans regulatory genes governing mucosal infection. Cell. Microbiol. 2018, 20, e12841. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.-Y.; Cao, Y.-B.; Xu, Z.; Ying, K.; Li, Y.; Xie, Y.; Zhu, Z.-Y.; Chen, W.-S.; Jiang, Y.-Y. cDNA microarray analysis of differential gene expression in Candida albicans biofilm exposed to farnesol. Antimicrob. Agents Chemother. 2005, 49, 584–589. [Google Scholar] [CrossRef] [PubMed]
- Dumitru, R.; Hornby, J.M.; Nickerson, K.W. Defined anaerobic growth medium for studying Candida albicans basic biology and resistance to eight antifungal drugs. Antimicrob. Agents Chemother. 2004, 48, 2350–2354. [Google Scholar] [CrossRef]
- De Groot, P.W.J.; Bader, O.; de Boer, A.D.; Weig, M.; Chauhan, N. Adhesins in human fungal pathogens: Glue with plenty of stick. Eukaryot. Cell 2013, 12, 470–481. [Google Scholar] [CrossRef] [PubMed]
- Nantel, A.; Dignard, D.; Bachewich, C.; Harcus, D.; Marcil, A.; Bouin, A.-P.; Sensen, C.W.; Hogues, H.; van het Hoog, M.; Gordon, P.; et al. Transcription profiling of Candida albicans cells undergoing the yeast-to-hyphal transition. Mol. Biol. Cell 2002, 13, 3452–3465. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Li, Y.H.; Yu, S.-B.; Li, W.G.; Liu, X.S.; Zhao, L.; Lu, J.X. A Genome-wide transcriptional analysis of yeast-hyphal transition in Candida tropicalis. by RNA-Seq. PLoS ONE 2016, 11, e0166645. [Google Scholar] [CrossRef] [PubMed]
- Felk, A.; Kretschmar, M.; Albrecht, A.; Schaller, M.; Beinhauer, S.; Nichterlein, T.; Sanglard, D.; Korting, H.C.; Schäfer, W.; Hube, B. Candida albicans hyphal formation and the expression of the Efg1-regulated proteinases Sap4 to Sap6 are required for the invasion of parenchymal organs. Infect. Immun. 2002, 70, 3689–3700. [Google Scholar] [CrossRef] [PubMed]
- Naglik, J.R.; Challacombe, S.J.; Hube, B. Candida albicans secreted aspartyl proteinases in virulence and pathogenesis. Microbiol. Mol. Biol. Rev. 2003, 67, 400–428. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, Y.; Wang, Y. Hgc1, a novel hypha-specific G1 cyclin-related protein regulates Candida albicans hyphal morphogenesis. EMBO J. 2004, 23, 1845–1856. [Google Scholar] [CrossRef]
- Lane, S.; Birse, C.; Zhou, S.; Matson, R.; Liu, H. DNA array studies demonstrate convergent regulation of virulence factors by Cph1, Cph2, and Efg1 in Candida albicans. J. Biol. Chem. 2001, 276, 48988–48996. [Google Scholar] [CrossRef]
- Bahn, Y.-S.; Molenda, M.; Staab, J.F.; Lyman, C.A.; Gordon, L.J.; Sundstrom, P. Genome-wide transcriptional profiling of the cyclic AMP-dependent signaling pathway during morphogenic transitions of Candida albicans. Eukaryot. Cell 2007, 6, 2376–2390. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, P.L.; Kadosh, D. Candida albicans Ume6, a filament-specific transcriptional regulator, directs hyphal growth via a pathway involving Hgc1 cyclin-related protein. Eukaryot. Cell 2010, 9, 1320–1328. [Google Scholar] [CrossRef]
- Banerjee, M.; Thompson, D.S.; Lazzell, A.; Carlisle, P.L.; Pierce, C.; Monteagudo, C.; López-Ribot, J.L.; Kadosh, D. UME6, a novel filament-specific regulator of Candida albicans hyphal extension and virulence. Mol. Biol. Cell 2008, 19, 1354–1365. [Google Scholar] [CrossRef] [PubMed]
- Ernst, J.F. Transcription factors in Candida albicans—Environmental control of morphogenesis. Microbiology 2000, 146, 1763–1774. [Google Scholar] [CrossRef] [PubMed]
- Doedt, T.; Krishnamurthy, S.; Bockmühl, D.P.; Tebarth, B.; Stempel, C.; Russell, C.L.; Brown, A.J.P.; Ernst, J.F. APSES proteins regulate morphogenesis and metabolism in Candida albicans. Mol. Biol. Cell 2004, 15, 3167–3180. [Google Scholar] [CrossRef] [PubMed]
- Mulhern, S.M.; Logue, M.E.; Butler, G. Candida albicans transcription factor Ace2 regulates metabolism and is required for filamentation in hypoxic conditions. Eukaryot. Cell 2006, 5, 2001–2013. [Google Scholar] [CrossRef] [PubMed]
- Cao, F.; Lane, S.; Raniga, P.P.; Lu, Y.; Zhou, Z.; Ramon, K.; Chen, J.; Liu, H. The Flo8 transcription factor is essential for hyphal development and virulence in Candida albicans. Mol. Biol. Cell 2006, 17, 295–307. [Google Scholar] [CrossRef]
- Martin, R.; Moran, G.P.; Jacobsen, I.D.; Heyken, A.; Domey, J.; Sullivan, D.J.; Kurzai, O.; Hube, B. The Candida albicans-specific gene EED1 encodes a key regulator of hyphal extension. PLoS ONE 2011, 6, e18394. [Google Scholar] [CrossRef] [PubMed]
- Vinces, M.D.; Haas, C.; Kumamoto, C.A. Expression of the Candida albicans morphogenesis regulator gene CZF1 and its regulation by Efg1p and Czf1p. Eukaryot. Cell 2006, 5, 825–835. [Google Scholar] [CrossRef]
- Maiti, P.; Ghorai, P.; Ghosh, S.; Kamthan, M.; Tyagi, R.K.; Datta, A. Mapping of functional domains and characterization of the transcription factor Cph1 that mediate morphogenesis in Candida albicans. Fungal Genet. Biol. 2015, 83, 45–57. [Google Scholar] [CrossRef]
- Khalaf, R.A.; Zitomer, R.S. The DNA binding protein Rfg1 is a repressor of filamentation in Candida albicans. Genetics 2001, 157, 1503–1512. [Google Scholar] [PubMed]
- Braun, B.R.; Head, W.S.; Wang, M.X.; Johnson, A.D. Identification and characterization of TUP1-regulated genes in Candida albicans. Genetics 2000, 156, 31–44. [Google Scholar] [PubMed]
- Braun, B.R.; Kadosh, D.; Johnson, A.D. NRG1, a repressor of filamentous growth in C. albicans, is down-regulated during filament induction. EMBO J. 2001, 20, 4753–4761. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.; Rodrigues, C.; Araújo, D.; Rodrigues, M.; Henriques, M. Candida species biofilms’ antifungal resistance. J. Fungi 2017, 3, 8. [Google Scholar] [CrossRef] [PubMed]
- Ramage, G.; Rajendran, R.; Sherry, L.; Williams, C. Fungal biofilm resistance. Int. J. Microbiol. 2012, 2012, 528521. [Google Scholar] [CrossRef] [PubMed]
- Baillie, G.S.; Douglas, L.J. Matrix polymers of Candida biofilms and their possible role in biofilm resistance to antifungal agents. J. Antimicrob. Chemother. 2000, 46, 397–403. [Google Scholar] [CrossRef] [PubMed]
- Al-Fattani, M.A.; Douglas, L.J. Biofilm matrix of Candida albicans and Candida tropicalis: chemical composition and role in drug resistance. J. Med. Microbiol. 2006, 55, 999–1008. [Google Scholar] [CrossRef] [PubMed]
- Sherry, L.; Ramage, G.; Kean, R.; Borman, A.; Johnson, E.M.; Richardson, M.D.; Rautemaa-Richardson, R. Biofilm-forming capability of highly virulent, multidrug-resistant Candida auris. Emerg. Infect. Dis. 2017, 23, 328–331. [Google Scholar] [CrossRef] [PubMed]
- Nobile, C.J.; Andes, D.R.; Nett, J.E.; Smith, F.J.; Yue, F.; Phan, Q.-T.; Edwards, J.E.; Filler, S.G.; Mitchell, A.P. Critical role of Bcr1-dependent adhesins in C. albicans biofilm formation in vitro and in vivo. PLoS Pathog. 2006, 2, e63. [Google Scholar] [CrossRef]
- Araújo, D.; Henriques, M.; Silva, S. Portrait of Candida species biofilm regulatory network genes. Trends Microbiol. 2017, 25, 62–75. [Google Scholar] [CrossRef]
- Paramonova, E.; Krom, B.P.; van der Mei, H.C.; Busscher, H.J.; Sharma, P.K. Hyphal content determines the compression strength of Candida albicans biofilms. Microbiology 2009, 155, 1997–2003. [Google Scholar] [CrossRef] [PubMed]
- Nobile, C.J.; Fox, E.P.; Nett, J.E.; Sorrells, T.R.; Mitrovich, Q.M.; Hernday, A.D.; Tuch, B.B.; Andes, D.R.; Johnson, A.D. A recently evolved transcriptional network controls biofilm development in Candida albicans. Cell 2012, 148, 126–138. [Google Scholar] [CrossRef] [PubMed]
- Fox, E.P.; Bui, C.K.; Nett, J.E.; Hartooni, N.; Mui, M.C.; Andes, D.R.; Nobile, C.J.; Johnson, A.D. An expanded regulatory network temporally controls Candida albicans biofilm formation. Mol. Microbiol. 2015, 96, 1226–1239. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Gao, N.; Li, C.; Gao, J.; Ying, C. A newly identified amino acid substitution T123I in the 14α-demethylase (Erg11p) of Candida albicans confers azole resistance. FEMS Yeast Res. 2017, 17, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Lv, Q.-Z.; Yan, L.; Jiang, Y.-Y. The synthesis, regulation, and functions of sterols in Candida albicans: Well-known but still lots to learn. Virulence 2016, 7, 649–659. [Google Scholar] [CrossRef] [PubMed]
- Whaley, S.G.; Berkow, E.L.; Rybak, J.M.; Nishimoto, A.T.; Barker, K.S.; David Rogers, P. Azole antifungal resistance in Candida albicans and emerging non-albicans Candida species. Front. Microbiol. 2017, 7, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Sheehan, D.J.; Hitchcock, C.A.; Sibley, C.M. Current and emerging azole antifungal agents. Clin. Microbiol. Rev. 1999, 12, 40–79. [Google Scholar] [CrossRef]
- White, T.C.; Marr, K.A.; Bowden, R.A. Clinical, cellular, and molecular factors that contribute to antifungal drug resistance. Clin. Microbiol. Rev. 1998, 11, 382–402. [Google Scholar] [CrossRef]
- Hitchcock, C.A. Cytochrome P-450-dependent 14α-sterol demethylase of Candida albicans and its interaction with azole antifungals. Biochem. Soc. Trans. 1991, 19, 782–787. [Google Scholar] [CrossRef]
- Perlin, D.S. Echinocandin resistance in Candida. Clin. Infect. Dis. 2015, 61, S612–S617. [Google Scholar] [CrossRef]
- Lee, K.K.; Maccallum, D.M.; Jacobsen, M.D.; Walker, L.A.; Odds, F.C.; Gow, N.A.R.; Munro, C.A. Elevated cell wall chitin in Candida albicans confers echinocandin resistance in vivo. Antimicrob. Agents Chemother. 2012, 56, 208–217. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.D.; Robbins, N.; Zaas, A.K.; Schell, W.A.; Perfect, J.R.; Cowen, L.E. Hsp90 governs echinocandin resistance in the pathogenic yeast Candida albicans via calcineurin. PLoS Pathog. 2009, 5, e1000532. [Google Scholar] [CrossRef] [PubMed]
- Nett, J.E.; Crawford, K.; Marchillo, K.; Andes, D.R. Role of Fks1p and matrix glucan in Candida albicans biofilm resistance to an echinocandin, pyrimidine, and polyene. Antimicrob. Agents Chemother. 2010, 54, 3505–3508. [Google Scholar] [CrossRef] [PubMed]
- Broughton, M.C.; Christine Broughton, M.; Bard, M.; Lees, N.D. Polyene resistance in ergosterol producing strains of Candida albicans. Mycoses 2009, 34, 75–83. [Google Scholar] [CrossRef]
- Belenky, P.; Camacho, D.; Collins, J.J. Fungicidal drugs induce a common oxidative-damage cellular death pathway. Cell Rep. 2013, 3, 350–358. [Google Scholar] [CrossRef] [PubMed]
- Shekhova, E.; Kniemeyer, O.; Brakhage, A.A. Induction of mitochondrial ROS production by itraconazole, terbinafine and amphotericin B as a mode of action against Aspergillus fumigatus. Antimicrob. Agents Chemother. 2017, 61, e00978-17. [Google Scholar]
- McCarthy, M.; O’Shaughnessy, E.M.; Walsh, T.J. Amphotericin B: Polyene resistance mechanisms. In Antimicrobial Drug Resistance; Springer: Cham, Switzerland, 2017; pp. 387–395. [Google Scholar]
- Perfect, J.R. The antifungal pipeline: a reality check. Nat. Rev. Drug Discov. 2017, 16, 603–616. [Google Scholar] [CrossRef] [PubMed]
- Vermes, A.; Guchelaar, H.J.; Dankert, J. Flucytosine: A review of its pharmacology, clinical indications, pharmacokinetics, toxicity and drug interactions. J. Antimicrob. Chemother. 2000, 46, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Pappas, P.G.; Kauffman, C.A.; Andes, D.R.; Clancy, C.J.; Marr, K.A.; Ostrosky-Zeichner, L.; Reboli, A.C.; Schuster, M.G.; Vazquez, J.A.; Walsh, T.J.; et al. Clinical practice guideline for the management of candidiasis: 2016 update by the Infectious Diseases Society of America. Clin. Infect. Dis. 2016, 62, 409–417. [Google Scholar] [CrossRef] [PubMed]
- Berkow, E.L.; Lockhart, S.R. Fluconazole resistance in Candida species: A current perspective. Infect. Drug Resist. 2017, 10, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Marichal, P.; Koymans, L.; Willemsens, S.; Bellens, D.; Verhasselt, P.; Luyten, W.; Borgers, M.; Ramaekers, F.C.S.; Odds, F.C.; Vanden Bossche, H. Contribution of mutations in the cytochrome P450 14alpha-demethylase (Erg11p, Cyp51p) to azole resistance in Candida albicans. Microbiology 1999, 145, 2701–2713. [Google Scholar] [CrossRef] [PubMed]
- Flowers, S.A.; Colón, B.; Whaley, S.G.; Schuler, M.A.; David Rogers, P. The contribution of clinically derived mutations in ERG11 to azole resistance in Candida albicans. Antimicrob. Agents Chemother. 2014, 59, 450–460. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos Silva, D.B.; Rodrigues, L.M.C.; de Almeida, A.A.; de Oliveira, K.M.P.; Grisolia, A.B. Novel point mutations in the ERG11 gene in clinical isolates of azole resistant Candida species. Mem. Inst. Oswaldo Cruz 2016, 111, 192–199. [Google Scholar] [CrossRef] [PubMed]
- White, T.C.; Holleman, S.; Dy, F.; Mirels, L.F.; Stevens, D.A. Resistance mechanisms in clinical isolates of Candida albicans. Antimicrob. Agents Chemother. 2002, 46, 1704–1713. [Google Scholar] [CrossRef] [PubMed]
- Xiang, M.-J.; Liu, J.-Y.; Ni, P.-H.; Wang, S.; Shi, C.; Wei, B.; Ni, Y.-X.; Ge, H.-L. Erg11 mutations associated with azole resistance in clinical isolates of Candida albicans. FEMS Yeast Res. 2013, 13, 386–393. [Google Scholar] [CrossRef]
- He, X.; Zhao, M.; Chen, J.; Wu, R.; Zhang, J.; Cui, R.; Jiang, Y.; Chen, J.; Cao, X.; Xing, Y.; et al. Overexpression of both ERG11 and ABC2 genes might be responsible for itraconazole resistance in clinical isolates of Candida krusei. PLoS ONE 2015, 10, e0136185. [Google Scholar] [CrossRef]
- Rosana, Y.; Yasmon, A.; Lestari, D.C. Overexpression and mutation as a genetic mechanism of fluconazole resistance in Candida albicans isolated from human immunodeficiency virus patients in Indonesia. J. Med. Microbiol. 2015, 64, 1046–1052. [Google Scholar] [CrossRef]
- Selmecki, A.; Forche, A.; Berman, J. Aneuploidy and isochromosome formation in drug-resistant Candida albicans. Science 2006, 313, 367–370. [Google Scholar] [CrossRef]
- Selmecki, A.; Gerami-Nejad, M.; Paulson, C.; Forche, A.; Berman, J. An isochromosome confers drug resistance in vivo by amplification of two genes, ERG11 and TAC1. Mol. Microbiol. 2008, 68, 624–641. [Google Scholar] [CrossRef]
- Flowers, S.A.; Barker, K.S.; Berkow, E.L.; Toner, G.; Chadwick, S.G.; Gygax, S.E.; Morschhäuser, J.; Rogers, P.D. Gain-of-function mutations in UPC2 are a frequent cause of ERG11 upregulation in azole-resistant clinical isolates of Candida albicans. Eukaryot. Cell 2012, 11, 1289–1299. [Google Scholar] [CrossRef]
- Whaley, S.G.; Caudle, K.E.; Vermitsky, J.-P.; Chadwick, S.G.; Toner, G.; Barker, K.S.; Gygax, S.E.; Rogers, P.D. UPC2A is required for high-level azole antifungal resistance in Candida glabrata. Antimicrob. Agents Chemother. 2014, 58, 4543–4554. [Google Scholar] [CrossRef] [PubMed]
- Hoot, S.J.; Smith, A.R.; Brown, R.P.; White, T.C. An A643V amino acid substitution in Upc2p contributes to azole resistance in well-characterized clinical isolates of Candida albicans. Antimicrob. Agents Chemother. 2011, 55, 940–942. [Google Scholar] [CrossRef]
- Feng, W.; Yang, J.; Wang, Y.; Chen, J.; Xi, Z.; Qiao, Z. ERG11 mutations and upregulation in clinical itraconazole-resistant isolates of Candida krusei. Can. J. Microbiol. 2016, 62, 938–943. [Google Scholar] [CrossRef] [PubMed]
- Prasad, R.; Banerjee, A.; Khandelwal, N.K.; Dhamgaye, S. The ABCs of Candida albicans multidrug transporter Cdr1. Eukaryot. Cell 2015, 14, 1154–1164. [Google Scholar] [CrossRef] [PubMed]
- Holmes, A.R.; Lin, Y.-H.; Niimi, K.; Lamping, E.; Keniya, M.; Niimi, M.; Tanabe, K.; Monk, B.C.; Cannon, R.D. ABC transporter Cdr1p contributes more than Cdr2p does to fluconazole efflux in fluconazole-resistant Candida albicans clinical isolates. Antimicrob. Agents Chemother. 2008, 52, 3851–3862. [Google Scholar] [CrossRef] [PubMed]
- Tsao, S.; Rahkhoodaee, F.; Raymond, M. Relative contributions of the Candida albicans ABC transporters Cdr1p and Cdr2p to clinical azole resistance. Antimicrob. Agents Chemother. 2009, 53, 1344–1352. [Google Scholar] [CrossRef]
- Morschhäuser, J.; Barker, K.S.; Liu, T.T.; BlaB-Warmuth, J.; Homayouni, R.; Rogers, P.D. The transcription factor Mrr1p controls expression of the MDR1 efflux pump and mediates multidrug resistance in Candida albicans. PLoS Pathog. 2007, 3, e164. [Google Scholar] [CrossRef]
- Cavalheiro, M.; Pais, P.; Galocha, M.; Teixeira, M.C. Host-pathogen interactions mediated by MDR transporters in fungi: as pleiotropic as it gets! Genes 2018, 9, 332. [Google Scholar] [CrossRef]
- Ramírez-Zavala, B.; Mogavero, S.; Schöller, E.; Sasse, C.; Rogers, P.D.; Morschhäuser, J. SAGA/ADA complex subunit Ada2 is required for Cap1- but not Mrr1-mediated upregulation of the Candida albicans multidrug efflux pump MDR1. Antimicrob. Agents Chemother. 2014, 58, 5102–5110. [Google Scholar] [CrossRef]
- Balashov, S.V.; Park, S.; Perlin, D.S. Assessing resistance to the echinocandin antifungal drug caspofungin in Candida albicans by profiling mutations in FKS1. Antimicrob. Agents Chemother. 2006, 50, 2058–2063. [Google Scholar] [CrossRef]
- Katiyar, S.K.; Alastruey-Izquierdo, A.; Healey, K.R.; Johnson, M.E.; Perlin, D.S.; Edlind, T.D. Fks1 and Fks2 are functionally redundant but differentially regulated in Candida glabrata: implications for echinocandin resistance. Antimicrob. Agents Chemother. 2012, 56, 6304–6309. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Zhang, L.; Wakabayashi, H.; Myers, J.; Jiang, Y.; Cao, Y.; Jimenez-Ortigosa, C.; Perlin, D.S.; Rustchenko, E. Tolerance to caspofungin in Candida albicans is associated with at least three distinctive mechanisms that govern expression of FKS genes and cell wall remodeling. Antimicrob. Agents Chemother. 2017, 61, e00071-17. [Google Scholar] [CrossRef] [PubMed]
- Jensen, R.H.; Astvad, K.M.T.; Silva, L.V.; Sanglard, D.; Jørgensen, R.; Nielsen, K.F.; Mathiasen, E.G.; Doroudian, G.; Perlin, D.S.; Arendrup, M.C. Stepwise emergence of azole, echinocandin and amphotericin B multidrug resistance in vivo in Candida albicans orchestrated by multiple genetic alterations. J. Antimicrob. Chemother. 2015, 70, 2551–2555. [Google Scholar] [CrossRef] [PubMed]
- Young, L.Y.; Hull, C.M.; Heitman, J. Disruption of ergosterol biosynthesis confers resistance to amphotericin B in Candida lusitaniae. Antimicrob. Agents Chemother. 2003, 47, 2717–2724. [Google Scholar] [CrossRef] [PubMed]
- Woods, R.A.; Bard, M.; Jackson, I.E.; Drutz, D.J. Resistance to polyene antibiotics and correlated sterol changes in two isolates of Candida tropicalis from a patient with an amphotericin B-resistant funguria. J. Infect. Dis. 1974, 129, 53–58. [Google Scholar] [CrossRef] [PubMed]
- Vandeputte, P.; Tronchin, G.; Bergès, T.; Hennequin, C.; Chabasse, D.; Bouchara, J.-P. Reduced susceptibility to polyenes associated with a missense mutation in the ERG6 gene in a clinical isolate of Candida glabrata with pseudohyphal growth. Antimicrob. Agents Chemother. 2007, 51, 982–990. [Google Scholar] [CrossRef] [PubMed]
- Forastiero, A.; Mesa-Arango, A.C.; Alastruey-Izquierdo, A.; Alcazar-Fuoli, L.; Bernal-Martinez, L.; Pelaez, T.; Lopez, J.F.; Grimalt, J.O.; Gomez-Lopez, A.; Cuesta, I.; et al. Candida tropicalis antifungal cross-resistance is related to different azole target (Erg11p) modifications. Antimicrob. Agents Chemother. 2013, 57, 4769–4781. [Google Scholar] [CrossRef]
- Kelly, S.L.; Lamb, D.C.; Kelly, D.E.; Manning, N.J.; Loeffler, J.; Hebart, H.; Schumacher, U.; Einsele, H. Resistance to fluconazole and cross-resistance to amphotericin B in Candida albicans from AIDS patients caused by defective sterol delta5,6-desaturation. FEBS Lett. 1997, 400, 80–82. [Google Scholar] [CrossRef]
- Ha, K.C.; White, T.C. Effects of azole antifungal drugs on the transition from yeast cells to hyphae in susceptible and resistant isolates of the pathogenic yeast Candida albicans. Antimicrob. Agents Chemother. 1999, 43, 763–768. [Google Scholar] [CrossRef]
- Odds, F.C.; Cockayne, A.; Hayward, J.; Abbott, A.B. Effects of imidazole- and triazole-derivative antifungal compounds on the growth and morphological development of Candida albicans hyphae. J. Gen. Microbiol. 1985, 131, 2581–2589. [Google Scholar] [CrossRef]
- Hornby, J.M.; Nickerson, K.W. Enhanced production of farnesol by Candida albicans treated with four azoles. Antimicrob. Agents Chemother. 2004, 48, 2305–2307. [Google Scholar] [CrossRef] [PubMed]
- Bandara, H.M.H.N.; Lam, O.L.T.; Jin, L.J.; Samaranayake, L. Microbial chemical signaling: a current perspective. Crit. Rev. Microbiol. 2012, 38, 217–249. [Google Scholar] [CrossRef] [PubMed]
- Hawser, S.; Islam, K. Comparisons of the effects of fungicidal and fungistatic antifungal agents on the morphogenetic transformation of Candida albicans. J. Antimicrob. Chemother. 1999, 43, 411–413. [Google Scholar] [CrossRef] [PubMed]
- Kabli, S.A. In vitro, susceptibilities of clinical yeast isolates to antifungal drugs of polyene, pyrimidine, and azoles, and their effect in yeast adhesion and mycelial Formation. Saudi J. Biol. Sci. 2008, 15, 189–197. [Google Scholar]
- Hawser, S.; Francolini, M.; Islam, K. The effects of antifungal agents on the morphogenetic transformation by Candida albicans in vitro. J. Antimicrob. Chemother. 1996, 38, 579–587. [Google Scholar] [CrossRef] [PubMed]
- Formosa, C.; Schiavone, M.; Martin-Yken, H.; François, J.M.; Duval, R.E.; Dague, E. Nanoscale effects of caspofungin against two yeast species, Saccharomyces cerevisiae and Candida albicans. Antimicrob. Agents Chemother. 2013, 57, 3498–3506. [Google Scholar] [CrossRef] [PubMed]
- Ben-Ami, R.; Garcia-Effron, G.; Lewis, R.E.; Gamarra, S.; Leventakos, K.; Perlin, D.S.; Kontoyiannis, D.P. Fitness and virulence costs of Candida albicans FKS1 hot spot mutations associated with echinocandin resistance. J. Infect. Dis. 2011, 204, 626–635. [Google Scholar] [CrossRef]
- Costa, C.R.; Hasimoto e Souza, L.K.; Ataídes, F.S.; Ferri, P.H.; da Costa, M.P.; de Fátima Lisboa Fernanades, O.; do Rosário Rodrigues Silva, M. Molecular analysis and dimorphism of azole-susceptible and resistant Candida albicans isolates. Rev. Soc. Bras. Med. Trop. 2011, 44, 740–744. [Google Scholar] [CrossRef]
- Xie, J.L.; Grahl, N.; Sless, T.; Leach, M.D.; Kim, S.H.; Hogan, D.A.; Robbins, N.; Cowen, L.E. Signaling through Lrg1, Rho1 and Pkc1 governs Candida albicans morphogenesis in response to diverse cues. PLoS Genet. 2016, 12, e1006405. [Google Scholar] [CrossRef]
- LaFayette, S.L.; Collins, C.; Zaas, A.K.; Schell, W.A.; Betancourt-Quiroz, M.; Gunatilaka, A.A.L.; Perfect, J.R.; Cowen, L.E. PKC signaling regulates drug resistance of the fungal pathogen Candida albicans via circuitry comprised of Mkc1, calcineurin, and Hsp90. PLoS Pathog. 2010, 6, e1001069. [Google Scholar] [CrossRef]
- Rocha, C.R.; Schröppel, K.; Harcus, D.; Marcil, A.; Dignard, D.; Taylor, B.N.; Thomas, D.Y.; Whiteway, M.; Leberer, E. Signaling through adenylyl cyclase is essential for hyphal growth and virulence in the pathogenic fungus Candida albicans. Mol. Biol. Cell 2001, 12, 3631–3643. [Google Scholar] [CrossRef] [PubMed]
- Jain, P.; Akula, I.; Edlind, T. Cyclic AMP signaling pathway modulates susceptibility of candida species and Saccharomyces cerevisiae to antifungal azoles and other sterol biosynthesis inhibitors. Antimicrob. Agents Chemother. 2003, 47, 3195–3201. [Google Scholar] [CrossRef]
- Min, K.; Biermann, A.; Hogan, D.A.; Konopka, J.B. Genetic analysis of NDT80 family transcription factors in Candida albicans using new CRISPR-Cas9 approaches. mSphere 2018, 3, e00545-18. [Google Scholar] [CrossRef] [PubMed]
- Sellam, A.; Askew, C.; Epp, E.; Tebbji, F.; Mullick, A.; Whiteway, M.; Nantel, A. Role of transcription factor CaNdt80p in cell separation, hyphal growth, and virulence in Candida albicans. Eukaryot. Cell 2010, 9, 634–644. [Google Scholar] [CrossRef] [PubMed]
- Bink, A.; Govaert, G.; Vandenbosch, D.; Kuchariková, S.; Coenye, T.; Nelis, H.; Van Dijck, P.; Cammue, B.P.A.; Thevissen, K. Transcription factor Efg1 contributes to the tolerance of Candida albicans biofilms against antifungal agents in vitro and in vivo. J. Med. Microbiol. 2012, 61, 813–819. [Google Scholar] [CrossRef] [PubMed]
- Connolly, L.A.; Riccombeni, A.; Grózer, Z.; Holland, L.M.; Lynch, D.B.; Andes, D.R.; Gácser, A.; Butler, G. The APSES transcription factor Efg1 is a global regulator that controls morphogenesis and biofilm formation in Candida parapsilosis. Mol. Microbiol. 2013, 90, 36–53. [Google Scholar] [PubMed]
- Sasse, C.; Schillig, R.; Dierolf, F.; Weyler, M.; Schneider, S.; Mogavero, S.; David Rogers, P.; Morschhäuser, J. The transcription factor Ndt80 does not contribute to Mrr1-, Tac1-, and Upc2-mediated fluconazole resistance in Candida albicans. PLoS ONE 2011, 6, e25623. [Google Scholar] [CrossRef]
- Sellam, A.; Tebbji, F.; Nantel, A. Role of Ndt80p in sterol metabolism regulation and azole resistance in Candida albicans. Eukaryot. Cell 2009, 8, 1174–1183. [Google Scholar] [CrossRef]
- Wuyts, J.; Van Dijck, P.; Holtappels, M. Fungal persister cells: The basis for recalcitrant infections? PLoS Pathog. 2018, 14, e1007301. [Google Scholar] [CrossRef]
- Dominguez, E.G.; Andes, D.R. Candida biofilm tolerance: comparison of planktonic and biofilm resistance mechanisms. In Candida albicans: Cellular and Molecular Biology; Springer: Cham, Switzerland, 2017; pp. 77–92. [Google Scholar]
- Taff, H.T.; Mitchell, K.F.; Edward, J.A.; Andes, D.R. Mechanisms of Candida biofilm drug resistance. Future Microbiol. 2013, 8, 1325–1337. [Google Scholar] [CrossRef]
- Hsu, J.-M.; Huang, J.; Meluh, P.B.; Laurent, B.C. The yeast RSC chromatin-remodeling complex is required for kinetochore function in chromosome segregation. Mol. Cell. Biol. 2003, 23, 3202–3215. [Google Scholar] [CrossRef] [PubMed]
- Zou, H.; Fang, H.-M.; Zhu, Y.; Wang, Y. Candida albicans Cyr1, Cap1 and G-actin form a sensor/effector apparatus for activating cAMP synthesis in hyphal growth. Mol. Microbiol. 2009, 75, 579–591. [Google Scholar] [CrossRef] [PubMed]
- Ruprich-Robert, G.; Chapeland-Leclerc, F.; Boisnard, S.; Florent, M.; Bories, G.; Papon, N. Contributions of the response regulators Ssk1p and Skn7p in the pseudohyphal development, stress adaptation, and drug sensitivity of the opportunistic yeast Candida lusitaniae. Eukaryot. Cell 2008, 7, 1071–1074. [Google Scholar] [CrossRef] [PubMed]
- Cowen, L.E.; Lindquist, S. Hsp90 potentiates the rapid evolution of new traits: Drug resistance in diverse fungi. Science 2005, 309, 2185–2189. [Google Scholar] [CrossRef] [PubMed]
- Cowen, L.E.; Carpenter, A.E.; Matangkasombut, O.; Fink, G.R.; Lindquist, S. Genetic architecture of Hsp90-dependent drug resistance. Eukaryot. Cell 2006, 5, 2184–2188. [Google Scholar] [CrossRef] [PubMed]
- Sanglard, D.; Ischer, F.; Marchetti, O.; Entenza, J.; Bille, J. Calcineurin A of Candida albicans: Involvement in antifungal tolerance, cell morphogenesis and virulence: Multiple functions of calcineurin in C. albicans. Mol. Microbiol. 2003, 48, 959–976. [Google Scholar] [CrossRef] [PubMed]
- Bader, T.; Schröppel, K.; Bentink, S.; Agabian, N.; Köhler, G.; Morschhäuser, J. Role of calcineurin in stress resistance, morphogenesis, and virulence of a Candida albicans wild-type strain. Infect. Immun. 2006, 74, 4366–4369. [Google Scholar] [CrossRef]
- Robbins, N.; Leach, M.D.; Cowen, L.E. Lysine deacetylases Hda1 and Rpd3 regulate Hsp90 function thereby governing fungal drug resistance. Cell Rep. 2012, 2, 878–888. [Google Scholar] [CrossRef]
- Li, X.; Robbins, N.; O’Meara, T.R.; Cowen, L.E. Extensive functional redundancy in the regulation of Candida albicans drug resistance and morphogenesis by lysine deacetylases Hos2, Hda1, Rpd3 and Rpd31. Mol. Microbiol. 2017, 103, 635–656. [Google Scholar] [CrossRef]
- Prill, S.K.-H.; Klinkert, B.; Timpel, C.; Gale, C.A.; Schröppel, K.; Ernst, J.F. PMT family of Candida albicans: Five protein mannosyltransferase isoforms affect growth, morphogenesis and antifungal resistance. Mol. Microbiol. 2005, 55, 546–560. [Google Scholar] [CrossRef]
- Timpel, C.; Zink, S.; Strahl-Bolsinger, S.; Schröppel, K.; Ernst, J. Morphogenesis, adhesive properties, and antifungal resistance depend on the Pmt6 protein mannosyltransferase in the fungal pathogen Candida albicans. J. Bacteriol. 2000, 182, 3063–3071. [Google Scholar] [CrossRef] [PubMed]
- Prasad, T.; Saini, P.; Gaur, N.A.; Vishwakarma, R.A.; Khan, L.A.; Haq, Q.M.R.; Prasad, R. Functional analysis of CaIPT1, a sphingolipid biosynthetic gene involved in multidrug resistance and morphogenesis of Candida albicans. Antimicrob. Agents Chemother. 2005, 49, 3442–3452. [Google Scholar] [CrossRef] [PubMed]
- Alfatah, M.; Bari, V.K.; Nahar, A.S.; Bijlani, S.; Ganesan, K. Critical role for CaFEN1 and CaFEN12 of Candida albicans in cell wall integrity and biofilm formation. Sci. Rep. 2017, 7, 40281. [Google Scholar] [CrossRef] [PubMed]
- Pasrija, R.; Krishnamurthy, S.; Prasad, T.; Ernst, J.F.; Prasad, R. Squalene epoxidase encoded by ERG1 affects morphogenesis and drug susceptibilities of Candida albicans. J. Antimicrob. Chemother. 2005, 55, 905–913. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, C.; Silva, S.; Faria-Oliveira, F.; Pinho, E.; Henriques, M.; Lucas, C. Candida albicans virulence and drug-resistance requires the O-acyltransferase Gup1p. BMC Microbiol. 2010, 10, 238. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Rossi, J.M.; Myers, L.C. Candida albicans Zn cluster transcription factors Tac1 and Znc1 are activated by farnesol to upregulate a transcriptional program including the multidrug efflux pump. Antimicrob. Agents Chemother. 2018, 62, e00968-18. [Google Scholar] [CrossRef]
- Prasad, T.; Hameed, S.; Manoharlal, R.; Biswas, S.; Mukhopadhyay, C.K.; Goswami, S.K.; Prasad, R. Morphogenic regulator EFG1 affects the drug susceptibilities of pathogenic Candida albicans. FEMS Yeast Res. 2010, 10, 587–596. [Google Scholar] [CrossRef]
- Bruno, V.M.; Kalachikov, S.; Subaran, R.; Nobile, C.J.; Kyratsous, C.; Mitchell, A.P. Control of the C. albicans cell wall damage response by transcriptional regulator Cas5. PLoS Pathog. 2006, 2, e21. [Google Scholar] [CrossRef]
- Vasicek, E.M.; Berkow, E.L.; Bruno, V.M.; Mitchell, A.P.; Wiederhold, N.P.; Barker, K.S.; Rogers, P.D. Disruption of the transcriptional regulator Cas5 results in enhanced killing of Candida albicans by fluconazole. Antimicrob. Agents Chemother. 2014, 58, 6807–6818. [Google Scholar] [CrossRef]
- Pukkila-Worley, R.; Peleg, A.Y.; Tampakakis, E.; Mylonakis, E. Candida albicans hyphal formation and virulence assessed using a Caenorhabditis elegans infection model. Eukaryot. Cell 2009, 8, 1750–1758. [Google Scholar] [CrossRef]
- Chamilos, G.; Nobile, C.J.; Bruno, V.M.; Lewis, R.E.; Mitchell, A.P.; Kontoyiannis, D.P. Candida albicans Cas5, a regulator of cell wall integrity, is required for virulence in murine and toll mutant fly models. J. Infect. Dis. 2009, 200, 152–157. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.L.; Qin, L.; Miao, Z.; Grys, B.T.; Diaz, J.D.L.C.; Ting, K.; Krieger, J.R.; Tong, J.; Tan, K.; Leach, M.D.; et al. The Candida albicans transcription factor Cas5 couples stress responses, drug resistance and cell cycle regulation. Nat. Commun. 2017, 8, 499. [Google Scholar] [CrossRef] [PubMed]
- Zeidler, U.; Lettner, T.; Lassnig, C.; Müller, M.; Lajko, R.; Hintner, H.; Breitenbach, M.; Bito, A. UME6 is a crucial downstream target of other transcriptional regulators of true hyphal development in Candida albicans. FEMS Yeast Res. 2009, 9, 126–142. [Google Scholar] [CrossRef] [PubMed]
- Carlisle, P.L.; Banerjee, M.; Lazzell, A.; Monteagudo, C.; López-Ribot, J.L.; Kadosh, D. Expression levels of a filament-specific transcriptional regulator are sufficient to determine Candida albicans morphology and virulence. Proc. Natl. Acad. Sci. USA 2009, 106, 599–604. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, M.; Uppuluri, P.; Zhao, X.R.; Carlisle, P.L.; Vipulanandan, G.; Villar, C.C.; López-Ribot, J.L.; Kadosh, D. Expression of UME6, a key regulator of Candida albicans hyphal development, enhances biofilm formation via Hgc1- and Sun41-dependent mechanisms. Eukaryot. Cell 2012, 12, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Norice, C.T.; Smith, F.J., Jr.; Solis, N.; Filler, S.G.; Mitchell, A.P. Requirement for Candida albicans Sun41 in biofilm formation and virulence. Eukaryot. Cell 2007, 6, 2046–2055. [Google Scholar] [CrossRef] [PubMed]
- Zarnowski, R.; Sanchez, H.; Covelli, A.S.; Dominguez, E.; Jaromin, A.; Berhardt, J.; Heiss, C.; Azadi, P.; Mitchell, A.; Andes, D.R. Candida albicans biofilm--induced vesicles confer drug resistance through matrix biogenesis. PLoS Biol. 2018, 16, e2006872. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.; Henriques, M.; Martins, A.; Oliveira, R.; Williams, D.; Azeredo, J. Biofilms of non-Candida albicans Candida species: Quantification, structure and matrix composition. Med. Mycol. 2009, 47, 681–689. [Google Scholar] [CrossRef]
- Vargas, G.; Rocha, J.D.B.; Oliveira, D.L.; Albuquerque, P.C.; Frases, S.; Santos, S.S.; Nosanchuk, J.D.; Gomes, A.M.O.; Medeiros, L.C.A.S.; Miranda, K.; et al. Compositional and immunobiological analyses of extracellular vesicles released by Candida albicans. Cell. Microbiol. 2015, 17, 389–407. [Google Scholar] [CrossRef]
- Joffe, L.S.; Nimrichter, L.; Rodrigues, M.L.; Del Poeta, M. Potential roles of fungal extracellular vesicles during infection. mSphere 2016, 1, e00099-16. [Google Scholar] [CrossRef]
- Rodrigues, M.L.; Godinho, R.M.C.; Zamith-Miranda, D.; Nimrichter, L. Traveling into outer space: unanswered questions about fungal extracellular vesicles. PLoS Pathog. 2015, 11, e1005240. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Cerdeira, C.; Gregorio, M.C.; Molares-Vila, A.; López-Barcenas, A.; Fabbrocini, G.; Bardhi, B.; Sinani, A.; Sánchez-Blanco, E.; Arenas-Guzmán, R.; Hernandez-Castro, R. Biofilms and vulvovaginal candidiasis. Colloids Surf. B Biointerfaces 2018, 174, 110–125. [Google Scholar] [CrossRef] [PubMed]
- LaFleur, M.D.; Kumamoto, C.A.; Lewis, K. Candida albicans biofilms produce antifungal-tolerant persister cells. Antimicrob. Agents Chemother. 2006, 50, 3839–3846. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Seneviratne, C.J.; Alpi, E.; Vizcaino, J.A.; Jin, L. Delicate metabolic control and coordinated stress response critically determine antifungal tolerance of Candida albicans biofilm persisters. Antimicrob. Agents Chemother. 2015, 59, 6101–6112. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Li, Z.; Chu, H.; Guo, J.; Jiang, G.; Qi, Q. Candida albicans amphotericin B-tolerant persister formation is closely related to surface adhesion. Mycopathologia 2016, 181, 41–49. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, D.M.; George, T.; Chandra, J.; Mukherjee, P.K.; Ghannoum, M.A. Antifungal susceptibility of Candida biofilms: unique efficacy of amphotericin B lipid formulations and echinocandins. Antimicrob. Agents Chemother. 2002, 46, 1773–1780. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, S.P.; VandeWalle, K.; Ramage, G.; Patterson, T.F.; Wickes, B.L.; Graybill, J.R.; López-Ribot, J.L. In vitro activity of caspofungin against Candida albicans biofilms. Antimicrob. Agents Chemother. 2002, 46, 3591–3596. [Google Scholar] [CrossRef]
- Mukherjee, P.K.; Chandra, J.; Kuhn, D.M.; Ghannoum, M.A. Mechanism of fluconazole resistance in Candida albicans biofilms: Phase-specific role of efflux pumps and membrane sterols. Infect. Immun. 2003, 71, 4333–4340. [Google Scholar] [CrossRef]
- García-Sánchez, S.; Aubert, S.; Iraqui, I.; Janbon, G.; Ghigo, J.-M.; d’Enfert, C. Candida albicans biofilms: a developmental state associated with specific and stable gene expression patterns. Eukaryot. Cell 2004, 3, 536–545. [Google Scholar] [CrossRef]
- Niimi, K.; Maki, K.; Ikeda, F.; Holmes, A.R.; Lamping, E.; Niimi, M.; Monk, B.C.; Cannon, R.D. Overexpression of Candida albicans CDR1, CDR2, or MDR1 does not produce significant changes in echinocandin susceptibility. Antimicrob. Agents Chemother. 2006, 50, 1148–1155. [Google Scholar] [CrossRef]
- Wirsching, S.; Michel, S.; Köhler, G.; Morschhäuser, J. Activation of the multiple drug resistance gene MDR1 in fluconazole-resistant, clinical Candida albicans strains is caused by mutations in a trans-regulatory factor. J. Bacteriol. 2000, 182, 400–404. [Google Scholar] [CrossRef] [PubMed]
- White, T.C. Increased mRNA levels of ERG16, CDR, and MDR1 correlate with increases in azole resistance in Candida albicans isolates from a patient infected with human immunodeficiency virus. Antimicrob. Agents Chemother. 1997, 41, 1482–1487. [Google Scholar] [CrossRef] [PubMed]
- Ramage, G.; Bachmann, S.; Patterson, T.F.; Wickes, B.L.; López-Ribot, J.L. Investigation of multidrug efflux pumps in relation to fluconazole resistance in Candida albicans biofilms. J. Antimicrob. Chemother. 2002, 49, 973–980. [Google Scholar] [CrossRef] [PubMed]
- Revie, N.M.; Iyer, K.R.; Robbins, N.; Cowen, L.E. Antifungal drug resistance: Evolution, mechanisms and impact. Curr. Opin. Microbiol. 2018, 45, 70–76. [Google Scholar] [CrossRef] [PubMed]
- Dickey, S.W.; Cheung, G.Y.C.; Otto, M. Different drugs for bad bugs: antivirulence strategies in the age of antibiotic resistance. Nat. Rev. Drug Discov. 2017, 16, 457–471. [Google Scholar] [CrossRef] [PubMed]
- Maura, D.; Ballok, A.E.; Rahme, L.G. Considerations and caveats in anti-virulence drug development. Curr. Opin. Microbiol. 2016, 33, 41–46. [Google Scholar] [CrossRef]
- Cegelski, L.; Marshall, G.R.; Eldridge, G.R.; Hultgren, S.J. The biology and future prospects of antivirulence therapies. Nat. Rev. Microbiol. 2008, 6, 17–27. [Google Scholar] [CrossRef]
- Rampioni, G.; Visca, P.; Leoni, L.; Imperi, F. Drug repurposing for antivirulence therapy against opportunistic bacterial pathogens. Emerg. Top. Life Sci. 2017, 1, 13–22. [Google Scholar] [CrossRef]
- Vila, T.; Romo, J.A.; Pierce, C.G.; McHardy, S.F.; Saville, S.P.; Lopez-Ribot, J.L. Targeting Candida albicans filamentation for antifungal drug development. Virulence 2017, 8, 150–158. [Google Scholar] [CrossRef]
- Garcia, C.; Burgain, A.; Chaillot, J.; Pic, É.; Khemiri, I.; Sellam, A. A phenotypic small-molecule screen identifies halogenated salicylanilides as inhibitors of fungal morphogenesis, biofilm formation and host cell invasion. Sci. Rep. 2018, 8, 11559. [Google Scholar] [CrossRef]
- Toenjes, K.A.; Munsee, S.M.; Ibrahim, A.S.; Jeffrey, R.; Edwards, J.E., Jr.; Johnson, D.I. Small-molecule inhibitors of the budded-to-hyphal-form transition in the pathogenic yeast Candida albicans. Antimicrob. Agents Chemother. 2005, 49, 963–972. [Google Scholar] [CrossRef] [PubMed]
- Bar-Yosef, H.; Vivanco Gonzalez, N.; Ben-Aroya, S.; Kron, S.J.; Kornitzer, D. Chemical inhibitors of Candida albicans hyphal morphogenesis target endocytosis. Sci. Rep. 2017, 7, 5692. [Google Scholar] [CrossRef] [PubMed]
- Goffena, J.; Toenjes, K.A.; Butler, D.K. Inhibition of yeast-to-filamentous growth transitions in Candida albicans by a small molecule inducer of mammalian apoptosis. Yeast 2018, 35, 291–298. [Google Scholar] [CrossRef] [PubMed]
- Romo, J.A.; Pierce, C.G.; Chaturvedi, A.K.; Lazzell, A.L.; McHardy, S.F.; Saville, S.P.; Lopez-Ribot, J.L. Development of anti-virulence approaches for candidiasis via a novel series of small-molecule inhibitors of Candida albicans filamentation. MBio 2017, 8, e01991-17. [Google Scholar] [CrossRef] [PubMed]
- Fazly, A.; Jain, C.; Dehner, A.C.; Issi, L.; Lilly, E.A.; Ali, A.; Cao, H.; Fidel, P.L., Jr.; Rao, R.P.; Kaufman, P.D. Chemical screening identifies filastatin, a small molecule inhibitor of Candida albicans adhesion, morphogenesis, and pathogenesis. Proc. Natl. Acad. Sci. USA 2013, 110, 13594–13599. [Google Scholar] [CrossRef]
- Zhang, M.; Chang, W.; Shi, H.; Zhou, Y.; Zheng, S.; Li, Y.; Li, L.; Lou, H. Biatriosporin D displays anti-virulence activity through decreasing the intracellular cAMP levels. Toxicol. Appl. Pharmacol. 2017, 322, 104–112. [Google Scholar] [CrossRef]
- Navarathna, D.H.M.L.P.; Hornby, J.M.; Krishnan, N.; Parkhurst, A.; Duhamel, G.E.; Nickerson, K.W. Effect of farnesol on a mouse model of systemic candidiasis, determined by use of a DPP3 knockout mutant of Candida albicans. Infect. Immun. 2007, 75, 1609–1618. [Google Scholar] [CrossRef]
- Haque, F.; Alfatah, M.; Ganesan, K.; Bhattacharyya, M.S. Inhibitory effect of sophorolipid on Candida albicans biofilm formation and hyphal growth. Sci. Rep. 2016, 6, 23575. [Google Scholar] [CrossRef]
- Lee, J.-H.; Kim, Y.-G.; Choi, P.; Ham, J.; Park, J.G.; Lee, J. Antibiofilm and antivirulence activities of 6-Gingerol and 6-Shogaol against Candida albicans due to hyphal inhibition. Front. Cell. Infect. Microbiol. 2018, 8, 299. [Google Scholar] [CrossRef]
- Graham, C.E.; Cruz, M.R.; Garsin, D.A.; Lorenz, M.C. Enterococcus faecalis bacteriocin EntV inhibits hyphal morphogenesis, biofilm formation, and virulence of Candida albicans. Proc. Natl. Acad. Sci. USA 2017, 114, 4507–4512. [Google Scholar] [CrossRef]
- Murzyn, A.; Krasowska, A.; Stefanowicz, P.; Dziadkowiec, D.; Łukaszewicz, M. Capric acid secreted by S. boulardii inhibits C. albicans filamentous growth, adhesion and biofilm formation. PLoS ONE 2010, 5, e12050. [Google Scholar] [CrossRef] [PubMed]
- Shekhar-Guturja, T.; Tebung, W.A.; Mount, H.; Liu, N.; Köhler, J.R.; Whiteway, M.; Cowen, L.E. Beauvericin potentiates azole activity via inhibition of multidrug efflux, blocks Candida albicans morphogenesis, and is effluxed via Yor1 and circuitry controlled by Zcf29. Antimicrob. Agents Chemother. 2016, 60, 7468–7480. [Google Scholar] [PubMed]
- Xie, J.L.; O’Meara, T.R.; Polvi, E.J.; Robbins, N.; Cowen, L.E. Staurosporine induces filamentation in the human fungal pathogen Candida albicans via signaling through Cyr1 and protein kinase A. mSphere 2017, 2, e00056-17. [Google Scholar] [CrossRef] [PubMed]
- Polvi, E.J.; Averette, A.F.; Lee, S.C.; Kim, T.; Bahn, Y.-S.; Veri, A.O.; Robbins, N.; Heitman, J.; Cowen, L.E. Metal chelation as a powerful strategy to probe cellular circuitry governing fungal drug resistance and morphogenesis. PLoS Genet. 2016, 12, e1006350. [Google Scholar] [CrossRef] [PubMed]
- Van Bogaert, I.N.A.; Saerens, K.; De Muynck, C.; Develter, D.; Soetaert, W.; Vandamme, E.J. Microbial production and application of sophorolipids. Appl. Microbiol. Biotechnol. 2007, 76, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Kronstad, J.W. Disarming fungal pathogens: Bacillus safensis inhibits virulence factor production and biofilm formation by Cryptococcus neoformans and Candida albicans. MBio 2017, 8, e01537-17. [Google Scholar] [CrossRef]
- Matsubara, V.H.; Wang, Y.; Bandara, H.M.H.N.; Mayer, M.P.A.; Samaranayake, L.P. Probiotic lactobacilli inhibit early stages of Candida albicans biofilm development by reducing their growth, cell adhesion, and filamentation. Appl. Microbiol. Biotechnol. 2016, 100, 6415–6426. [Google Scholar] [CrossRef]
- Bandara, H.M.H.N.; Cheung, B.P.K.; Watt, R.M.; Jin, L.J.; Samaranayake, L.P. Secretory products of Escherichia coli biofilm modulate Candida biofilm formation and hyphal development. J. Investig. Clin. Dent. 2013, 4, 186–199. [Google Scholar] [CrossRef]
- De Barros, P.P.; Freire, F.; Rossoni, R.D.; Junqueira, J.C.; Jorge, A.O.C. Candida krusei and Candida glabrata reduce the filamentation of Candida albicans by downregulating expression of HWP1 gene. Folia Microbiol. 2017, 62, 317–323. [Google Scholar] [CrossRef]
- Krasowska, A.; Murzyn, A.; Dyjankiewicz, A.; Łukaszewicz, M.; Dziadkowiec, D. The antagonistic effect of Saccharomyces boulardii on Candida albicans filamentation, adhesion and biofilm formation. FEMS Yeast Res. 2009, 9, 1312–1321. [Google Scholar] [CrossRef]
- Shapiro, R.S.; Cowen, L.E. Uncovering cellular circuitry controlling temperature-dependent fungal morphogenesis. Virulence 2012, 3, 400–404. [Google Scholar] [CrossRef] [PubMed]
- O’Meara, T.R.; Robbins, N.; Cowen, L.E. The Hsp90 chaperone network modulates Candida virulence traits. Trends Microbiol. 2017, 25, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, R.S.; Sellam, A.; Tebbji, F.; Whiteway, M.; Nantel, A.; Cowen, L.E. Pho85, Pcl1, and Hms1 signaling governs Candida albicans morphogenesis induced by high temperature or Hsp90 compromise. Curr. Biol. 2012, 22, 461–470. [Google Scholar] [CrossRef] [PubMed]
- Senn, H.; Shapiro, R.S.; Cowen, L.E. Cdc28 provides a molecular link between Hsp90, morphogenesis, and cell cycle progression in Candida albicans. Mol. Biol. Cell 2012, 23, 268–283. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.; Gao, Y.; Liu, H.; Chen, J. Candida albicans Cdc37 interacts with the Crk1 kinase and is required for Crk1 production. FEBS Lett. 2004, 561, 223–230. [Google Scholar] [CrossRef]
- Chen, J.; Zhou, S.; Wang, Q.; Chen, X.; Pan, T.; Liu, H. Crk1, a novel Cdc2-related protein kinase, is required for hyphal development and virulence in Candida albicans. Mol. Cell. Biol. 2000, 20, 8696–8708. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).