PS, It’s Complicated: The Roles of Phosphatidylserine and Phosphatidylethanolamine in the Pathogenesis of Candida albicans and Other Microbial Pathogens
Abstract
1. Introduction
2. Phosphatidylserine and Phosphatidylethanolamine Synthesis in Microbes
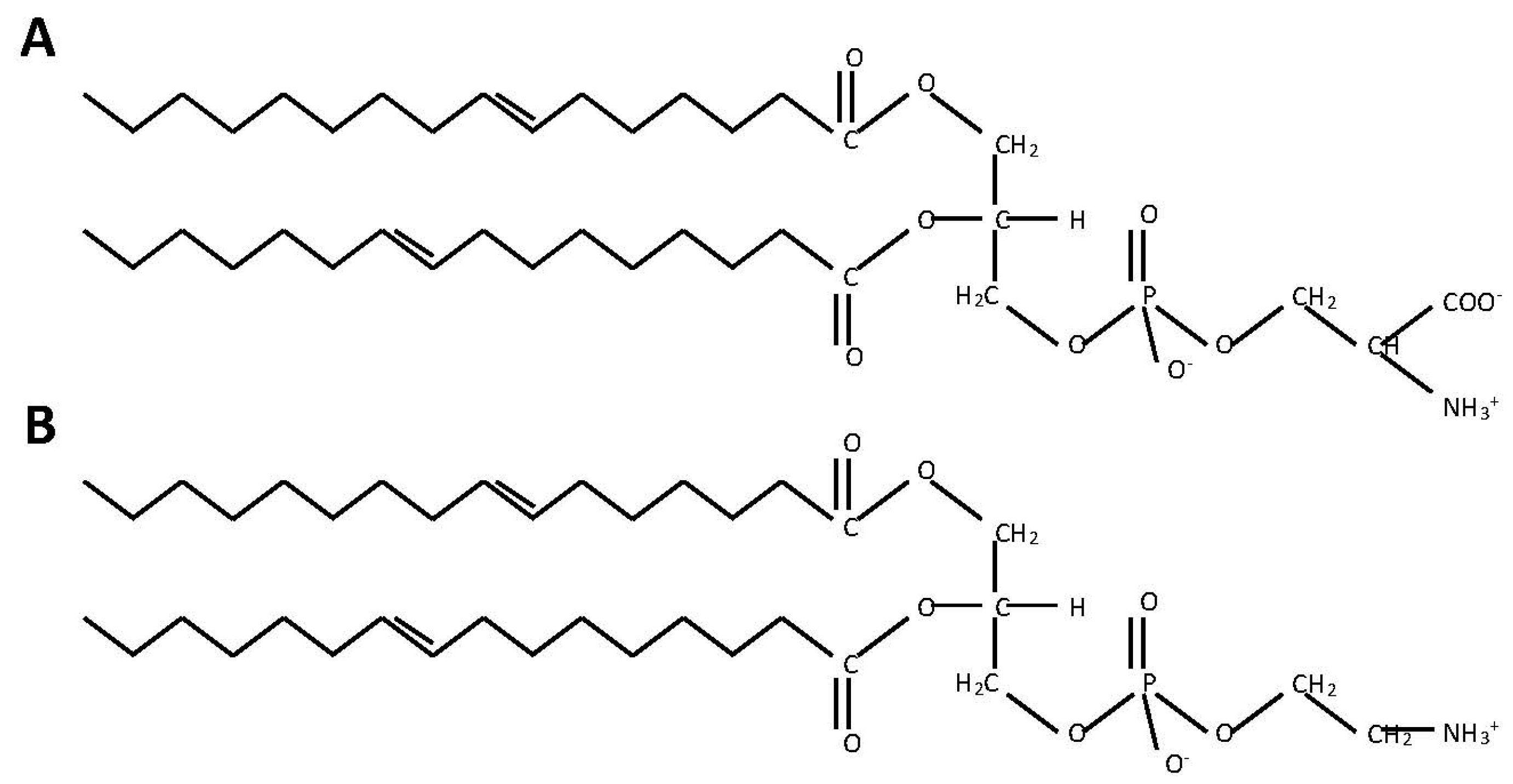
2.1. Phosphatidylserine Synthesis Is Similar between Fungi and Bacteria
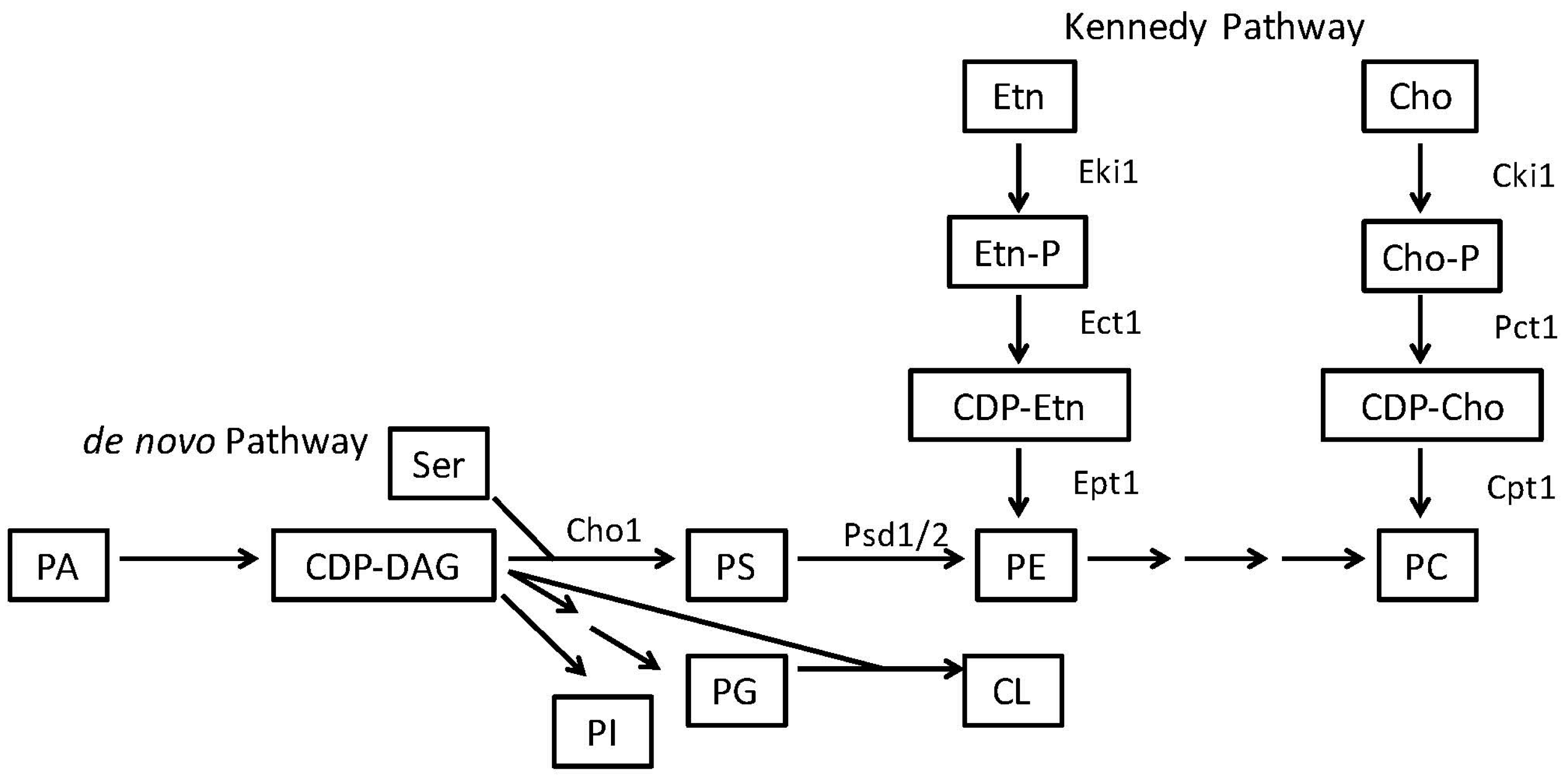
2.2. Phosphatidylethanolamine Is Synthesized by a Variety of Pathways
3. PS and PE Can Act as Modulators of Virulence in Candida, Bacteria and Parasites
3.1. Candida albicans Requires PE Synthesis from PS to Be Virulent
3.2. PS Inhibitors Could Be Effective against Some Bacterial Pathogens
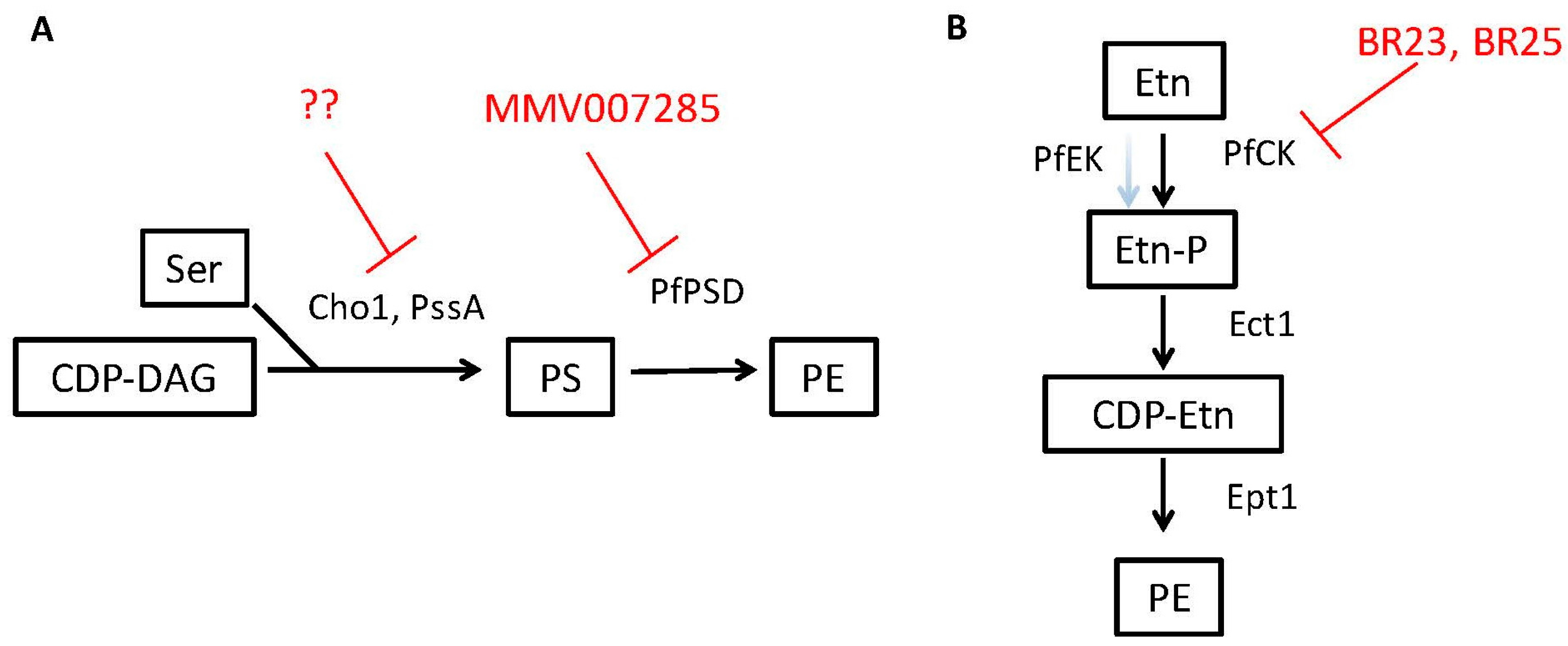
3.3. PE Synthesis Inhibitors Could Be Effective against Eukaryotic Pathogens
4. PS Symmetry in the Membrane Plays a Role in Virulence
4.1. Cryptococcus neoformans Lipid Flippase Impacts Virulence
4.2. PS Exposure in Parasites Facilitates Invasion of Host Cells
5. PS and PE May Play a Role in Extracellular Vesicles in Candida and Other Fungi
6. Perspectus
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Finlay, B.B.; Falkow, S. Common themes in microbial pathogenicity. Microbiol. Rev. 1989, 53, 210–230. [Google Scholar] [PubMed]
- Finlay, B.B.; Falkow, S. Common themes in microbial pathogenicity revisited. Microbiol. Mol. Biol. Rev. 1997, 61, 136–169. [Google Scholar] [PubMed]
- Mahan, M.J.; Heithoff, D.M.; Sinsheimer, R.L.; Low, D.A. Assessment of bacterial pathogenesis by analysis of gene expression in the host. Annu. Rev. Genet. 2000, 34, 139–164. [Google Scholar] [CrossRef] [PubMed]
- Cronan, J.E. Bacterial membrane lipids: Where do we stand? Annu. Rev. Microbiol. 2003, 57, 203–224. [Google Scholar] [CrossRef] [PubMed]
- Dowhan, W. Molecular basis for membrane phospholipid diversity: Why are there so many lipids? Annu. Rev. Biochem. 1997, 66, 199–232. [Google Scholar] [CrossRef] [PubMed]
- Shea, J.M.; Del Poeta, M. Lipid signaling in pathogenic fungi. Curr. Opin. Microbiol. 2006, 9, 352–358. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bogdanov, M.; Dowhan, W. Lipid-assisted protein folding. J. Biol. Chem. 1999, 274, 36827–36830. [Google Scholar] [CrossRef] [PubMed]
- Mirucki, C.S.; Abedi, M.; Jiang, J.; Zhu, Q.; Wang, Y.H.; Safavi, K.E.; Clark, R.B.; Nichols, F.C. Biologic activity of porphyromonas endodontalis complex lipids. J. Endod. 2014, 40, 1342–1348. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jain, M.; Petzold, C.J.; Schelle, M.W.; Leavell, M.D.; Mougous, J.D.; Bertozzi, C.R.; Leary, J.A.; Cox, J.S. Lipidomics reveals control of Mycobacterium tuberculosis virulence lipids via metabolic coupling. Proc. Natl. Acad. Sci. USA 2007, 104, 5133–5138. [Google Scholar] [CrossRef] [PubMed]
- Upreti, H.B.; Rawat, D.S.; Das, S.K. Virulence, capsule size and lipid composition interrelation of Cryptococcus neoformans. Microbiologica 1984, 7, 371–374. [Google Scholar] [PubMed]
- Wessel, M.; Klusener, S.; Godeke, J.; Fritz, C.; Hacker, S.; Narberhaus, F. Virulence of Agrobacterium tumefaciens requires phosphatidylcholine in the bacterial membrane. Mol. Microbiol. 2006, 62, 906–915. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, A.; Molle, V.; Besra, G.S.; Jacobs, W.R., Jr.; Kremer, L. The Mycobacterium tuberculosis FAS-Ⅱ condensing enzymes: Their role in mycolic acid biosynthesis, acid-fastness, pathogenesis and in future drug development. Mol. Microbiol. 2007, 64, 1442–1454. [Google Scholar] [CrossRef] [PubMed]
- Anderes, E.A.; Sandine, W.E.; Elliker, P.R. Lipids of antibiotic-sensitive and -resistant strains of Pseudomonas aeruginosa. Can. J. Microbiol. 1971, 17, 1357–1365. [Google Scholar] [CrossRef] [PubMed]
- Rakotomanga, M.; Saint-Pierre-Chazalet, M.; Loiseau, P.M. Alteration of fatty acid and sterol metabolism in miltefosine-resistant Leishmania donovani promastigotes and consequences for drug-membrane interactions. Antimicrob. Agents Chemother. 2005, 49, 2677–2686. [Google Scholar] [CrossRef] [PubMed]
- Cox, R.A.; Best, G.K. Cell wall composition of two strains of Blastomyces dermatitidis exhibiting differences in virulence for mice. Infect. Immun. 1972, 5, 449–453. [Google Scholar] [PubMed]
- Disalvo, A.F.; Denton, J.F. Lipid content of four strains of Blastomyces dermatitidis of different mouse virulence. J. Bacteriol. 1963, 85, 927–931. [Google Scholar] [PubMed]
- Nielsen, H.S. Variation in lipid content of strains of Histoplasma capsulatum exhibiting different virulence properties for mice. J. Bacteriol. 1965, 91, 273–277. [Google Scholar]
- Rella, A.; Farnoud, A.M.; Del Poeta, M. Plasma membrane lipids and their role in fungal virulence. Prog. Lipid Res. 2016, 61, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Geiger, O. Lipids and Legionella virulence. In Handbook of Hydrocarbon and Lipid Microbiology; Timmis, K.N., Ed.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 3195–3202. [Google Scholar]
- Goren, M.B.; Brokl, O.; Schaefer, W.B. Lipids of putative relevance to virulence in Mycobacterium tuberculosis: Correlation of virulence with elaboration of sulfatides and strongly acidic lipids. Infect. Immun. 1974, 9, 142–149. [Google Scholar] [PubMed]
- Davis, S.E.; Hopke, A.; Minkin, S.C., Jr.; Montedonico, A.E.; Wheeler, R.T.; Reynolds, T.B. Masking of β(1-3)-glucan in the cell wall of Candida albicans from detection by innate immune cells depends on phosphatidylserine. Infect. Immun. 2014, 82, 4405–4413. [Google Scholar] [CrossRef] [PubMed]
- Tilney, L.G.; Harb, O.S.; Connelly, P.S.; Robinson, C.G.; Roy, C.R. How the parasitic bacterium Legionella pneumophila modifies its phagosome and transforms it into rough ER: Implications for conversion of plasma membrane to the ER membrane. J. Cell Sci. 2001, 114, 4637–4650. [Google Scholar] [PubMed]
- Giles, D.K.; Hankins, J.V.; Guan, Z.; Trent, M.S. Remodelling of the Vibrio cholerae membrane by incorporation of exogenous fatty acids from host and aquatic environments. Mol. Microbiol. 2011, 79, 716–728. [Google Scholar] [CrossRef] [PubMed]
- Harp, J.R.; Saito, H.E.; Bourdon, A.K.; Reyes, J.; Arias, C.A.; Campagna, S.R.; Fozo, E.M. Exogenous fatty acids protect Enterococcus faecalis from daptomycin-induced membrane stress independently of the response regulator liar. Appl. Environ. Microbiol. 2016, 82, 4410–4420. [Google Scholar] [CrossRef] [PubMed]
- Saito, H.E.; Harp, J.R.; Fozo, E.M. Incorporation of exogenous fatty acids protects Enterococcus faecalis from membrane-damaging agents. Appl. Environ. Microbiol. 2014, 80, 6527–6538. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.; Rock, C.O. Exogenous fatty acid metabolism in bacteria. Biochimie 2017, 141, 30–39. [Google Scholar] [CrossRef] [PubMed]
- Sant, D.G.; Tupe, S.G.; Ramana, C.V.; Deshpande, M.V. Fungal cell membrane-promising drug target for antifungal therapy. J. Appl. Microbiol. 2016, 121, 1498–1510. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishnan, S.; Serricchio, M.; Striepen, B.; Butikofer, P. Lipid synthesis in protozoan parasites: A comparison between kinetoplastids and apicomplexans. Prog. Lipid Res. 2013, 52, 488–512. [Google Scholar] [CrossRef] [PubMed]
- Mishra, P.; Bolard, J.; Prasad, R. Emerging role of lipids of Candida albicans, a pathogenic dimorphic yeast. Biochim. Biophys. Acta 1992, 1127, 1–14. [Google Scholar] [CrossRef]
- Carman, G.M.; Han, G.S. Regulation of phospholipid synthesis in the yeast Saccharomyces cerevisiae. Annu. Rev. Biochem. 2011, 80, 859–883. [Google Scholar] [CrossRef] [PubMed]
- Carman, G.M.; Zeimetz, G.M. Regulation of phospholipid biosynthesis in the yeast Saccharomyces cerevisiae. J. Biol. Chem. 1996, 271, 13293–13296. [Google Scholar] [CrossRef] [PubMed]
- Raetz, C.R.; Dowhan, W. Biosynthesis and function of phospholipids in Escherichia coli. J. Biol. Chem. 1990, 265, 1235–1238. [Google Scholar] [PubMed]
- Sohlenkamp, C.; de Rudder, K.E.; Geiger, O. Phosphatidylethanolamine is not essential for growth of Sinorhizobium meliloti on complex culture media. J. Bacteriol. 2004, 186, 1667–1677. [Google Scholar] [CrossRef] [PubMed]
- Vance, J.E. Phospholipid synthesis and transport in mammalian cells. Traffic 2015, 16, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Henry, S.A.; Kohlwein, S.D.; Carman, G.M. Metabolism and regulation of glycerolipids in the yeast Saccharomyces cerevisiae. Genetics 2012, 190, 317–349. [Google Scholar] [CrossRef] [PubMed]
- Vance, J.E.; Steenbergen, R. Metabolism and functions of phosphatidylserine. Prog. Lipid Res. 2005, 44, 207–234. [Google Scholar] [CrossRef] [PubMed]
- Kuge, O.; Nishijima, M. Phosphatidylserine synthase I and II of mammalian cells. Biochim. Biophys. Acta 1997, 1348, 151–156. [Google Scholar] [CrossRef]
- Signorell, A.; Rauch, M.; Jelk, J.; Ferguson, M.A.; Butikofer, P. Phosphatidylethanolamine in Trypanosoma brucei is organized in two separate pools and is synthesized exclusively by the kennedy pathway. J. Biol. Chem. 2008, 283, 23636–23644. [Google Scholar] [CrossRef] [PubMed]
- Tasseva, G.; Bai, H.D.; Davidescu, M.; Haromy, A.; Michelakis, E.; Vance, J.E. Phosphatidylethanolamine deficiency in mammalian mitochondria impairs oxidative phosphorylation and alters mitochondrial morphology. J. Biol. Chem. 2013, 288, 4158–4173. [Google Scholar] [CrossRef] [PubMed]
- Kanfer, J.; Kennedy, E.P. Metabolism and function of bacterial lipids II. Biosynthesis of phospholipids in Escherichia coli. J. Biol. Chem. 1964, 239, 1720–1726. [Google Scholar] [PubMed]
- Cassilly, C.D.; Farmer, A.T.; Montedonico, A.E.; Smith, T.K.; Campagna, S.R.; Reynolds, T.B. Role of phosphatidylserine synthase in shaping the phospholipidome of Candida albicans. FEMS Yeast Res. 2017, 17. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.P.; Weiss, S.B. The function of cytidine coenzymes in the biosynthesis of phospholipides. J. Biol Chem. 1956, 222, 193–214. [Google Scholar] [PubMed]
- Gibellini, F.; Smith, T.K. The kennedy pathway—de novo synthesis of phosphatidylethanolamine and phosphatidylcholine. IUBMB Life 2010, 62, 414–428. [Google Scholar] [CrossRef] [PubMed]
- Moser, R.; Aktas, M.; Fritz, C.; Narberhaus, F. Discovery of a bifunctional cardiolipin/phosphatidylethanolamine synthase in bacteria. Mol. Microbiol. 2014, 92, 959–972. [Google Scholar] [CrossRef] [PubMed]
- Pulido, S.A.; Nguyen, V.H.; Alzate, J.F.; Cedeno, D.L.; Makurath, M.A.; Rios-Vasquez, A.; Duque-Benitez, S.M.; Jones, M.A.; Robledo, S.M.; Friesen, J.A. Insights into the phosphatidylcholine and phosphatidylethanolamine biosynthetic pathways in Leishmania parasites and characterization of a choline kinase from Leishmania infantum. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 2017, 213, 45–54. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Pompey, J.M.; Hsu, F.F.; Key, P.; Bandhuvula, P.; Saba, J.D.; Turk, J.; Beverley, S.M. Redirection of sphingolipid metabolism toward de novo synthesis of ethanolamine in Leishmania. EMBO J. 2007, 26, 1094–1104. [Google Scholar] [CrossRef] [PubMed]
- Birner, R.; Burgermeister, M.; Schneiter, R.; Daum, G. Roles of phosphatidylethanolamine and of its several biosynthetic pathways in Saccharomyces cerevisiae. Mol. Biol. Cell. 2001, 12, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Mao, C.; Wadleigh, M.; Jenkins, G.M.; Hannun, Y.A.; Obeid, L.M. Identification and characterization of Saccharomyces cerevisiae dihydrosphingosine-1-phosphate phosphatase. J. Biol. Chem. 1997, 272, 28690–28694. [Google Scholar] [CrossRef] [PubMed]
- Saba, J.D.; Nara, F.; Bielawska, A.; Garrett, S.; Hannun, Y.A. The BST1 gene of Saccharomyces cerevisiae is the sphingosine-1-phosphate lyase. J. Biol. Chem. 1997, 272, 26087–26090. [Google Scholar] [CrossRef] [PubMed]
- Deng, L.; Fukuda, R.; Kakihara, T.; Narita, K.; Ohta, A. Incorporation and remodeling of phosphatidylethanolamine containing short acyl residues in yeast. Biochim. Biophys. Acta 2010, 1801, 635–645. [Google Scholar] [CrossRef] [PubMed]
- Flis, V.V.; Daum, G. Lipid transport between the endoplasmic reticulum and mitochondria. Cold Spring Harb. Perspect. Biol. 2013, 5, a013235. [Google Scholar] [CrossRef] [PubMed]
- Riekhof, W.R.; Voelker, D.R. Uptake and utilization of lyso-phosphatidylethanolamine by Saccharomyces cerevisiae. J. Biol. Chem. 2006, 281, 36588–36596. [Google Scholar] [CrossRef] [PubMed]
- Riekhof, W.R.; Wu, J.; Gijon, M.A.; Zarini, S.; Murphy, R.C.; Voelker, D.R. Lysophosphatidylcholine metabolism in Saccharomyces cerevisiae: The role of P-type ATPases in transport and a broad specificity acyltransferase in acylation. J. Biol. Chem. 2007, 282, 36853–36861. [Google Scholar] [CrossRef] [PubMed]
- Riekhof, W.R.; Wu, J.; Jones, J.L.; Voelker, D.R. Identification and characterization of the major lysophosphatidylethanolamine acyltransferase in Saccharomyces cerevisiae. J. Biol. Chem. 2007, 282, 28344–28352. [Google Scholar] [CrossRef] [PubMed]
- Burgermeister, M.; Birner-Grunberger, R.; Nebauer, R.; Daum, G. Contribution of different pathways to the supply of phosphatidylethanolamine and phosphatidylcholine to mitochondrial membranes of the yeast Saccharomyces cerevisiae. Biochim. Biophys. Acta 2004, 1686, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Elabbadi, N.; Ancelin, M.L.; Vial, H.J. Phospholipid metabolism of serine in plasmodium-infected erythrocytes involves phosphatidylserine and direct serine decarboxylation. Biochem. J. 1997, 324(Pt. 2), 435–445. [Google Scholar] [CrossRef]
- Hartmann, A.; Hellmund, M.; Lucius, R.; Voelker, D.R.; Gupta, N. Phosphatidylethanolamine synthesis in the parasite mitochondrion is required for efficient growth but dispensable for survival of Toxoplasma gondii. J. Biol. Chem. 2014, 289, 6809–6824. [Google Scholar] [CrossRef] [PubMed]
- Tyhach, R.J.; Hawrot, E.; Satre, M.; Kennedy, E.P. Increased synthesis of phosphatidylserine decarboxylase in a strain of Escherichia coli bearing a hybrid plasmid. Altered association of enzyme with the membrane. J. Biol. Chem. 1979, 254, 627–633. [Google Scholar] [PubMed]
- Choi, J.Y.; Kumar, V.; Pachikara, N.; Garg, A.; Lawres, L.; Toh, J.Y.; Voelker, D.R.; Ben Mamoun, C. Characterization of plasmodium phosphatidylserine decarboxylase expressed in yeast and application for inhibitor screening. Mol. Microbiol. 2016, 99, 999–1014. [Google Scholar] [CrossRef] [PubMed]
- Baunaure, F.; Eldin, P.; Cathiard, A.M.; Vial, H. Characterization of a non-mitochondrial type I phosphatidylserine decarboxylase in Plasmodium falciparum. Mol. Microbiol. 2004, 51, 33–46. [Google Scholar] [CrossRef] [PubMed]
- Gupta, N.; Hartmann, A.; Lucius, R.; Voelker, D.R. The obligate intracellular parasite Toxoplasma gondii secretes a soluble phosphatidylserine decarboxylase. J. Biol. Chem. 2012, 287, 22938–22947. [Google Scholar] [CrossRef] [PubMed]
- Bustamante, C.I. Treatment of Candida infection: A view from the trenches! Curr. Opin. Infect. Dis. 2005, 18, 490–495. [Google Scholar] [CrossRef] [PubMed]
- Eggimann, P.; Garbino, J.; Pittet, D. Management of Candida species infections in critically ill patients. Lancet Infect. Dis. 2003, 3, 772–785. [Google Scholar] [CrossRef]
- Cassone, A.; Cauda, R. Candida and candidiasis in hiv-infected patients: Where commensalism, opportunistic behavior and frank pathogenicity lose their borders. Aids 2012, 26, 1457–1472. [Google Scholar] [CrossRef] [PubMed]
- Moyes, D.L.; Wilson, D.; Richardson, J.P.; Mogavero, S.; Tang, S.X.; Wernecke, J.; Hofs, S.; Gratacap, R.L.; Robbins, J.; Runglall, M.; et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 2016, 532, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Kumamoto, C.A.; Vinces, M.D. Contributions of hyphae and hypha-co-regulated genes to Candida albicans virulence. Cell. Microbiol. 2005, 7, 1546–1554. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Gordon, S. Immune recognition. A new receptor for β-glucans. Nature 2001, 413, 36–37. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, R.T.; Fink, G.R. A drug-sensitive genetic network masks fungi from the immune system. PLoS Pathog. 2006, 2, e35. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, R.T.; Kombe, D.; Agarwala, S.D.; Fink, G.R. Dynamic, morphotype-specific Candida albicans β-glucan exposure during infection and drug treatment. PLoS Pathog. 2008, 4, e1000227. [Google Scholar] [CrossRef] [PubMed]
- Hasim, S.; Allison, D.P.; Retterer, S.T.; Hopke, A.; Wheeler, R.T.; Doktycz, M.J.; Reynolds, T.B. β-(1,3)-glucan unmasking in some Candida albicans mutants correlates with increases in cell wall surface roughness and decreases in cell wall elasticity. Infect. Immun. 2017, 85, e00601-16. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.L.; Montedonico, A.E.; Kauffman, S.; Dunlap, J.R.; Menn, F.M.; Reynolds, T.B. Phosphatidylserine synthase and phosphatidylserine decarboxylase are essential for cell wall integrity and virulence in Candida albicans. Mol. Microbiol. 2010, 75, 1112–1132. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.L.; Reynolds, T.B.; University of Tennessee, Knoxville, USA. Unpublished work. 2009.
- Cassilly, C.D.; Maddox, M.M.; Cherian, P.T.; Bowling, J.J.; Hamann, M.T.; Lee, R.E.; Reynolds, T.B. SB-224289 antagonizes the antifungal mechanism of the marine depsipeptide papuamide A. PLoS ONE 2016, 11, e0154932. [Google Scholar] [CrossRef] [PubMed]
- Braun, B.R.; van Het Hoog, M.; d’Enfert, C.; Martchenko, M.; Dungan, J.; Kuo, A.; Inglis, D.O.; Uhl, M.A.; Hogues, H.; Berriman, M.; et al. A human-curated annotation of the Candida albicans genome. PLoS Genet. 2005, 1, 36–57. [Google Scholar] [CrossRef] [PubMed]
- Bukata, L.; Altabe, S.; de Mendoza, D.; Ugalde, R.A.; Comerci, D.J. Phosphatidylethanolamine synthesis is required for optimal virulence of Brucella abortus. J. Bacteriol. 2008, 190, 8197–8203. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Serran-Aguilera, L.; Denton, H.; Rubio-Ruiz, B.; Lopez-Gutierrez, B.; Entrena, A.; Izquierdo, L.; Smith, T.K.; Conejo-Garcia, A.; Hurtado-Guerrero, R. Plasmodium falciparum choline kinase inhibition leads to a major decrease in phosphatidylethanolamine causing parasite death. Sci. Rep. 2016, 6, 33189. [Google Scholar] [CrossRef] [PubMed]
- Comerci, D.J.; Altabe, S.; de Mendoza, D.; Ugalde, R.A. Brucella abortus synthesizes phosphatidylcholine from choline provided by the host. J. Bacteriol. 2006, 188, 1929–1934. [Google Scholar] [CrossRef] [PubMed]
- Conde-Alvarez, R.; Grillo, M.J.; Salcedo, S.P.; de Miguel, M.J.; Fugier, E.; Gorvel, J.P.; Moriyon, I.; Iriarte, M. Synthesis of phosphatidylcholine, a typical eukaryotic phospholipid, is necessary for full virulence of the intracellular bacterial parasite Brucella abortus. Cell. Microbiol. 2006, 8, 1322–1335. [Google Scholar] [CrossRef] [PubMed]
- Thiele, O.W.; Kehr, W. Die “freien” lipids of Brucella abortus bang. FEBS J. 1969, 9, 167–175. [Google Scholar]
- Nakajima, M.; DeChavigny, A.; Johnson, C.E.; Hamada, J.; Stein, C.A.; Nicolson, G.L. Suramin. A potent inhibitor of melanoma heparanase and invasion. J. Biol. Chem. 1991, 266, 9661–9666. [Google Scholar] [PubMed]
- Ben Mamoun, C.; Prigge, S.T.; Vial, H. Targeting the lipid metabolic pathways for the treatment of malaria. Drug Dev. Res. 2010, 71, 44–55. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Denning, D.W.; Gow, N.A.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden killers: Human fungal infections. Sci. Transl. Med. 2012, 4, 165rv113. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.; Liao, G.; Baker, G.M.; Wang, Y.; Lau, R.; Paderu, P.; Perlin, D.S.; Xue, C. Lipid flippase subunit cdc50 mediates drug resistance and virulence in Cryptococcus neoformans. MBio 2016, 7, e00478-16. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, A.; Gupta, C.M. Transbilayer translocation of membrane phosphatidylserine and its role in macrophage invasion in Leishmania promastigotes. Mol. Biochem. Parasitol. 2003, 128, 1–9. [Google Scholar] [CrossRef]
- Wanderley, J.L.; Moreira, M.E.; Benjamin, A.; Bonomo, A.C.; Barcinski, M.A. Mimicry of apoptotic cells by exposing phosphatidylserine participates in the establishment of amastigotes of Leishmania (l) amazonensis in mammalian hosts. J. Immunol. 2006, 176, 1834–1839. [Google Scholar] [CrossRef] [PubMed]
- De Freitas Balanco, J.M.; Moreira, M.E.; Bonomo, A.; Bozza, P.T.; Amarante-Mendes, G.; Pirmez, C.; Barcinski, M.A. Apoptotic mimicry by an obligate intracellular parasite downregulates macrophage microbicidal activity. Curr. Biol. 2001, 11, 1870–1873. [Google Scholar] [CrossRef]
- Van Zandbergen, G.; Solbach, W.; Laskay, T. Apoptosis driven infection. Autoimmunity 2007, 40, 349–352. [Google Scholar] [CrossRef] [PubMed]
- Farias, L.H.; Rodrigues, A.P.; Silveira, F.T.; Seabra, S.H.; DaMatta, R.A.; Saraiva, E.M.; Silva, E.O. Phosphatidylserine exposure and surface sugars in two Leishmania (viannia) braziliensis strains involved in cutaneous and mucocutaneous leishmaniasis. J. Infect. Dis. 2013, 207, 537–543. [Google Scholar] [CrossRef] [PubMed]
- Wanderley, J.L.; Pinto da Silva, L.H.; Deolindo, P.; Soong, L.; Borges, V.M.; Prates, D.B.; de Souza, A.P.; Barral, A.; Balanco, J.M.; do Nascimento, M.T.; et al. Cooperation between apoptotic and viable metacyclics enhances the pathogenesis of leishmaniasis. PLoS ONE 2009, 4, e5733. [Google Scholar] [CrossRef] [PubMed]
- Van Zandbergen, G.; Bollinger, A.; Wenzel, A.; Kamhawi, S.; Voll, R.; Klinger, M.; Muller, A.; Holscher, C.; Herrmann, M.; Sacks, D.; et al. Leishmania disease development depends on the presence of apoptotic promastigotes in the virulent inoculum. Proc. Natl. Acad. Sci. USA 2006, 103, 13837–13842. [Google Scholar] [CrossRef] [PubMed]
- Damatta, R.A.; Seabra, S.H.; Deolindo, P.; Arnholdt, A.C.; Manhaes, L.; Goldenberg, S.; de Souza, W. Trypanosoma cruzi exposes phosphatidylserine as an evasion mechanism. FEMS Microbiol. Lett. 2007, 266, 29–33. [Google Scholar] [CrossRef] [PubMed]
- Freire-de-Lima, C.G.; Nascimento, D.O.; Soares, M.B.; Bozza, P.T.; Castro-Faria-Neto, H.C.; de Mello, F.G.; DosReis, G.A.; Lopes, M.F. Uptake of apoptotic cells drives the growth of a pathogenic trypanosome in macrophages. Nature 2000, 403, 199–203. [Google Scholar] [CrossRef] [PubMed]
- Seabra, S.H.; de Souza, W.; Damatta, R.A. Toxoplasma gondii exposes phosphatidylserine inducing a TGF-β1 autocrine effect orchestrating macrophage evasion. Biochem. Biophys. Res. Commun. 2004, 324, 744–752. [Google Scholar] [CrossRef] [PubMed]
- Moller-Tank, S.; Kondratowicz, A.S.; Davey, R.A.; Rennert, P.D.; Maury, W. Role of the phosphatidylserine receptor TIM-1 in enveloped-virus entry. J. Virol. 2013, 87, 8327–8341. [Google Scholar] [CrossRef] [PubMed]
- Jemielity, S.; Wang, J.J.; Chan, Y.K.; Ahmed, A.A.; Li, W.; Monahan, S.; Bu, X.; Farzan, M.; Freeman, G.J.; Umetsu, D.T.; et al. TIM-family proteins promote infection of multiple enveloped viruses through virion-associated phosphatidylserine. PLoS Pathog. 2013, 9, e1003232. [Google Scholar] [CrossRef] [PubMed]
- Soares, M.M.; King, S.W.; Thorpe, P.E. Targeting inside-out phosphatidylserine as a therapeutic strategy for viral diseases. Nat. Med. 2008, 14, 1357–1362. [Google Scholar] [CrossRef] [PubMed]
- Vanlandschoot, P.; Leroux-Roels, G. Viral apoptotic mimicry: An immune evasion strategy developed by the hepatitis b virus? Trends Immunol. 2003, 24, 144–147. [Google Scholar] [CrossRef]
- Dos Santos, M.G.; Muxel, S.M.; Zampieri, R.A.; Pomorski, T.G.; Floeter-Winter, L.M. Transbilayer dynamics of phospholipids in the plasma membrane of the Leishmania genus. PLoS ONE 2013, 8, e55604. [Google Scholar] [CrossRef] [PubMed]
- Araujo-Santos, J.M.; Gamarro, F.; Castanys, S.; Herrmann, A.; Pomorski, T. Rapid transport of phospholipids across the plasma membrane of Leishmania infantum. Biochem. Biophys. Res. Commun. 2003, 306, 250–255. [Google Scholar] [CrossRef]
- Albuquerque, P.C.; Nakayasu, E.S.; Rodrigues, M.L.; Frases, S.; Casadevall, A.; Zancope-Oliveira, R.M.; Almeida, I.C.; Nosanchuk, J.D. Vesicular transport in Histoplasma capsulatum: An effective mechanism for trans-cell wall transfer of proteins and lipids in ascomycetes. Cell. Microbiol. 2008, 10, 1695–1710. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, M.L.; Nakayasu, E.S.; Oliveira, D.L.; Nimrichter, L.; Nosanchuk, J.D.; Almeida, I.C.; Casadevall, A. Extracellular vesicles produced by Cryptococcus neoformans contain protein components associated with virulence. Eukaryot. Cell 2008, 7, 58–67. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, D.L.; Freire-de-Lima, C.G.; Nosanchuk, J.D.; Casadevall, A.; Rodrigues, M.L.; Nimrichter, L. Extracellular vesicles from Cryptococcus neoformans modulate macrophage functions. Infect. Immun. 2010, 78, 1601–1609. [Google Scholar] [CrossRef] [PubMed]
- Manocha, M.S.; San-Blas, G.; Centeno, S. Lipid composition of Paracoccidioides brasiliensis: Possible correlation with virulence of different strains. J. Gen. Microbiol. 1980, 117, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Schorey, J.S.; Cheng, Y.; Singh, P.P.; Smith, V.L. Exosomes and other extracellular vesicles in host-pathogen interactions. EMBO Rep. 2015, 16, 24–43. [Google Scholar] [CrossRef] [PubMed]
- Vallejo, M.C.; Nakayasu, E.S.; Longo, L.V.; Ganiko, L.; Lopes, F.G.; Matsuo, A.L.; Almeida, I.C.; Puccia, R. Lipidomic analysis of extracellular vesicles from the pathogenic phase of Paracoccidioides brasiliensis. PLoS ONE 2012, 7, e39463. [Google Scholar] [CrossRef]
- Laulagnier, K.; Motta, C.; Hamdi, S.; Roy, S.; Fauvelle, F.; Pageaux, J.F.; Kobayashi, T.; Salles, J.P.; Perret, B.; Bonnerot, C.; et al. Mast cell- and dendritic cell-derived exosomes display a specific lipid composition and an unusual membrane organization. Biochem. J. 2004, 380, 161–171. [Google Scholar] [CrossRef] [PubMed]
- Subra, C.; Laulagnier, K.; Perret, B.; Record, M. Exosome lipidomics unravels lipid sorting at the level of multivesicular bodies. Biochimie 2007, 89, 205–212. [Google Scholar] [CrossRef] [PubMed]
- Wolf, J.M.; Espadas, J.; Luque-Garcia, J.; Reynolds, T.; Casadevall, A. Lipid biosynthetic genes affect Candida albicans extracellular vesicle morphology, cargo, and immunostimulatory properties. Eukaryot. Cell 2015, 14, 745–754. [Google Scholar] [CrossRef] [PubMed]

| Target/Enzyme Activity | Organism | Phospholipid Product | Inhibitor(s) * | Other Potential Pathogens | Ref. |
|---|---|---|---|---|---|
| Cho1/Phosphatidylserine synthase | Candida albicans | phosphatidylserine | NA | Conserved in fungi | [71] |
| PssA/Phosphatidylserine synthase | Brucella abortus | phosphatidylserine | NA | Conserved in gram negative bacteria | [75] |
| PfPSD/Phosphatidylserine decarboxylase | Plasmodium falciparum | Phosphatidyl-ethanolamine | 7-chloro-N-(4-ethoxyphenyl)-4-quinolinamine (MMV007285) | Conserved in parasites, fungi and some bacteria | [59] |
| PfCK/ Ethanolamine kinase activity of choline kinase | Plasmodium falciparum | Phosphatidyl-ethanolamine | BR23, BR25 | unknown | [76] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cassilly, C.D.; Reynolds, T.B. PS, It’s Complicated: The Roles of Phosphatidylserine and Phosphatidylethanolamine in the Pathogenesis of Candida albicans and Other Microbial Pathogens. J. Fungi 2018, 4, 28. https://doi.org/10.3390/jof4010028
Cassilly CD, Reynolds TB. PS, It’s Complicated: The Roles of Phosphatidylserine and Phosphatidylethanolamine in the Pathogenesis of Candida albicans and Other Microbial Pathogens. Journal of Fungi. 2018; 4(1):28. https://doi.org/10.3390/jof4010028
Chicago/Turabian StyleCassilly, Chelsi D., and Todd B. Reynolds. 2018. "PS, It’s Complicated: The Roles of Phosphatidylserine and Phosphatidylethanolamine in the Pathogenesis of Candida albicans and Other Microbial Pathogens" Journal of Fungi 4, no. 1: 28. https://doi.org/10.3390/jof4010028
APA StyleCassilly, C. D., & Reynolds, T. B. (2018). PS, It’s Complicated: The Roles of Phosphatidylserine and Phosphatidylethanolamine in the Pathogenesis of Candida albicans and Other Microbial Pathogens. Journal of Fungi, 4(1), 28. https://doi.org/10.3390/jof4010028





