The Velvet Complex Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum
Abstract
1. Introduction
2. Results
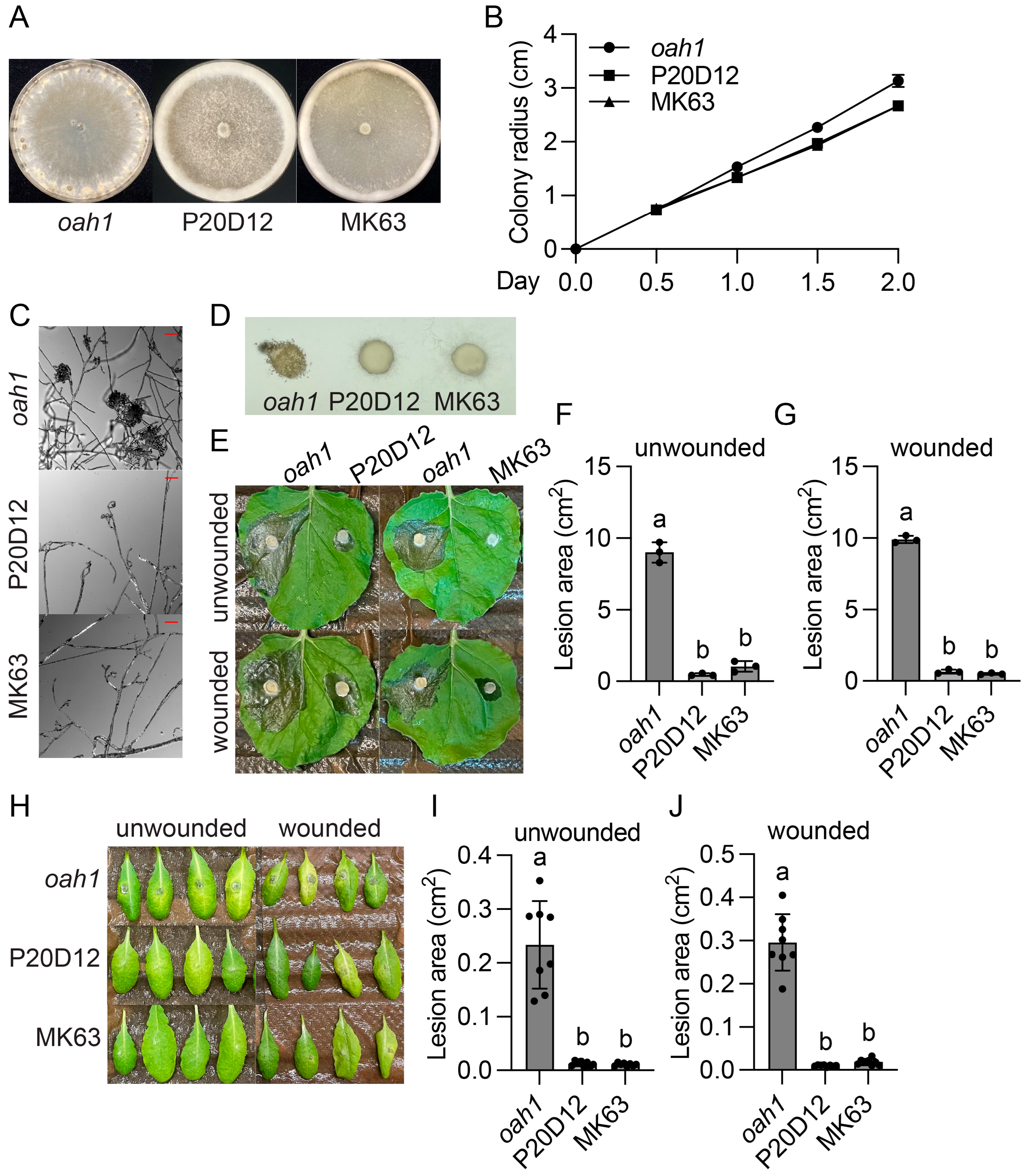
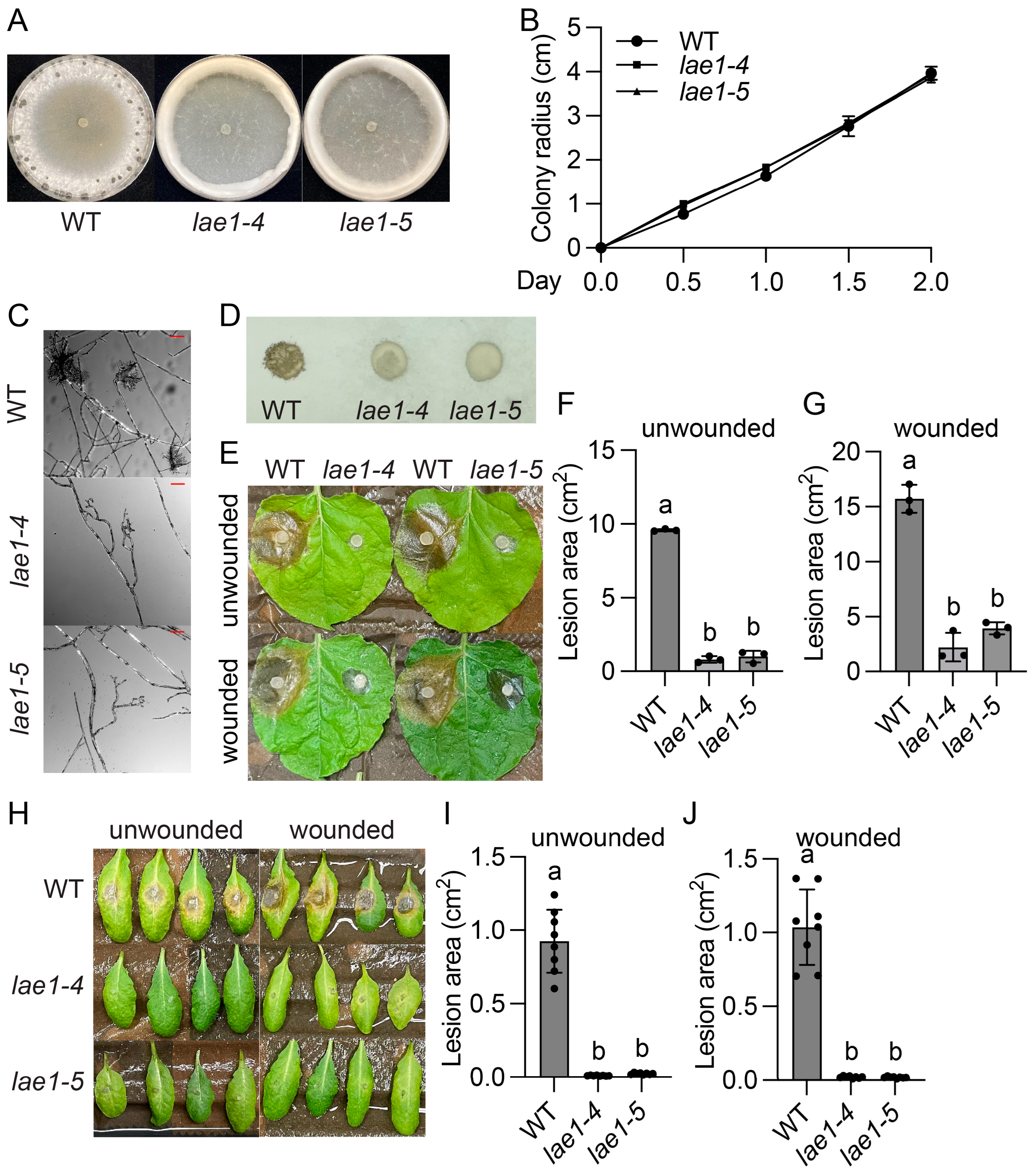
2.1. Identification and Characterization of Three Sclerotinia sclerotiorum UV Mutants That Display Thick Aerial Hyphae, No Sclerotia, and Decreased Virulence
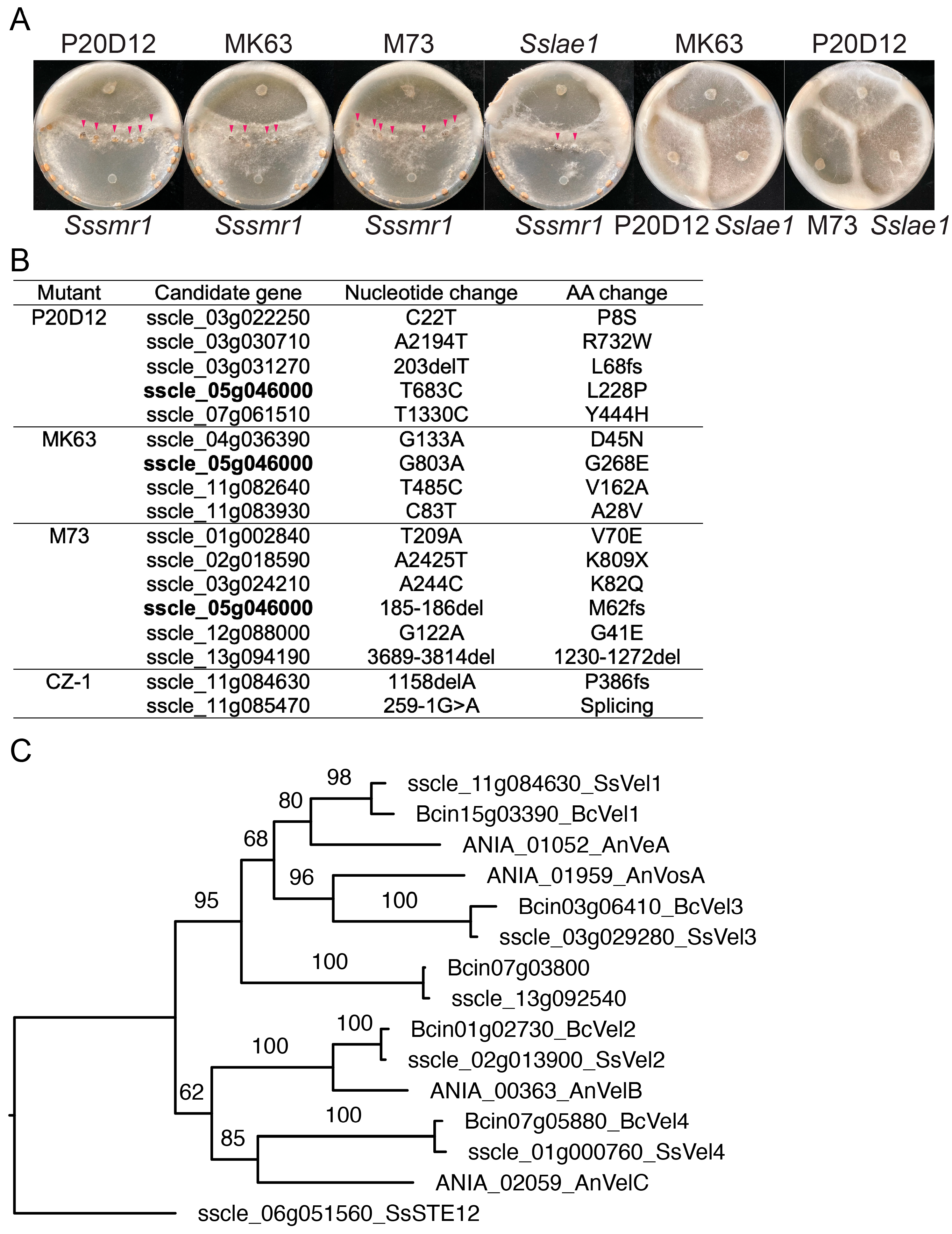
2.2. Molecular Cloning of SsLae1
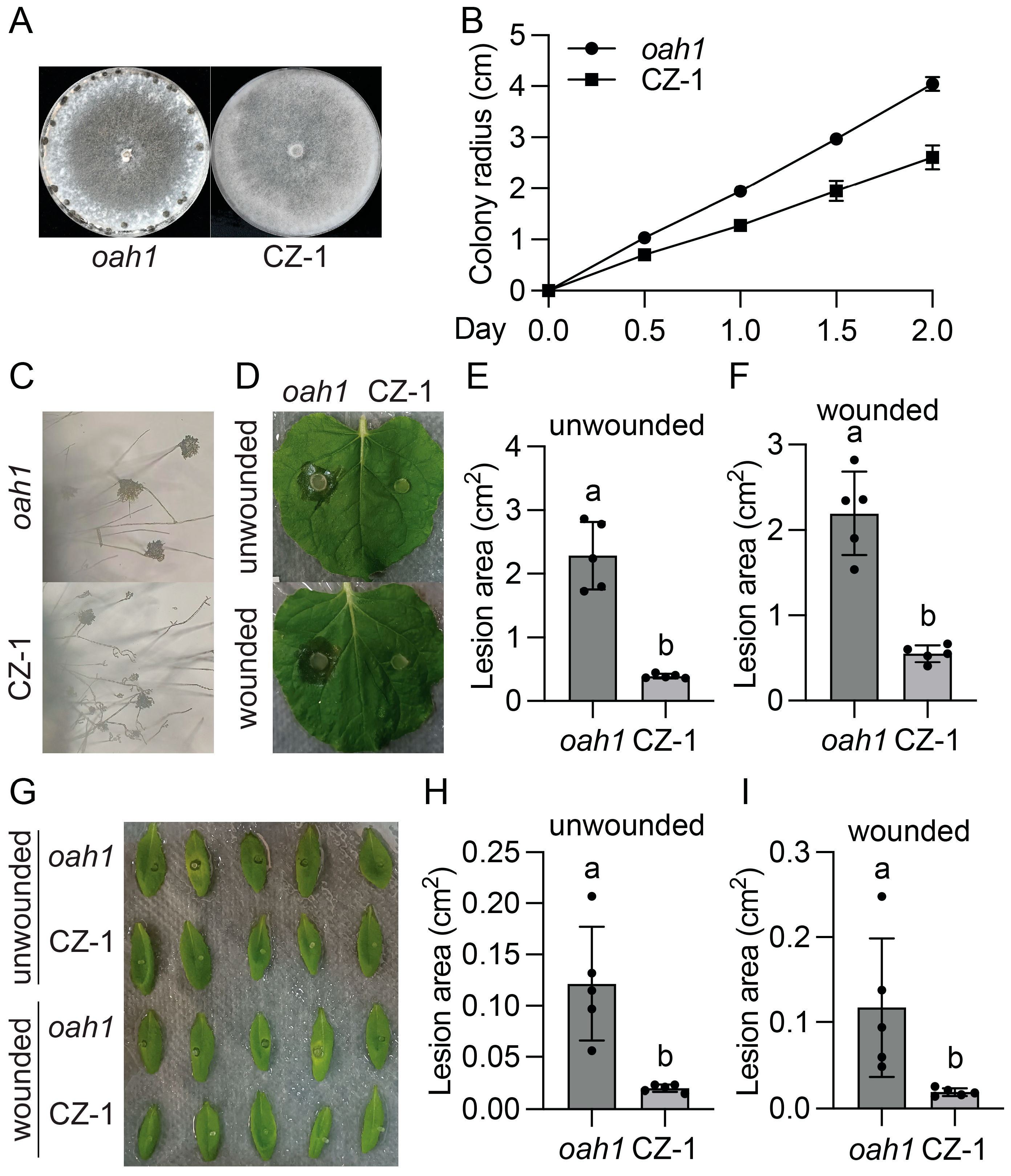
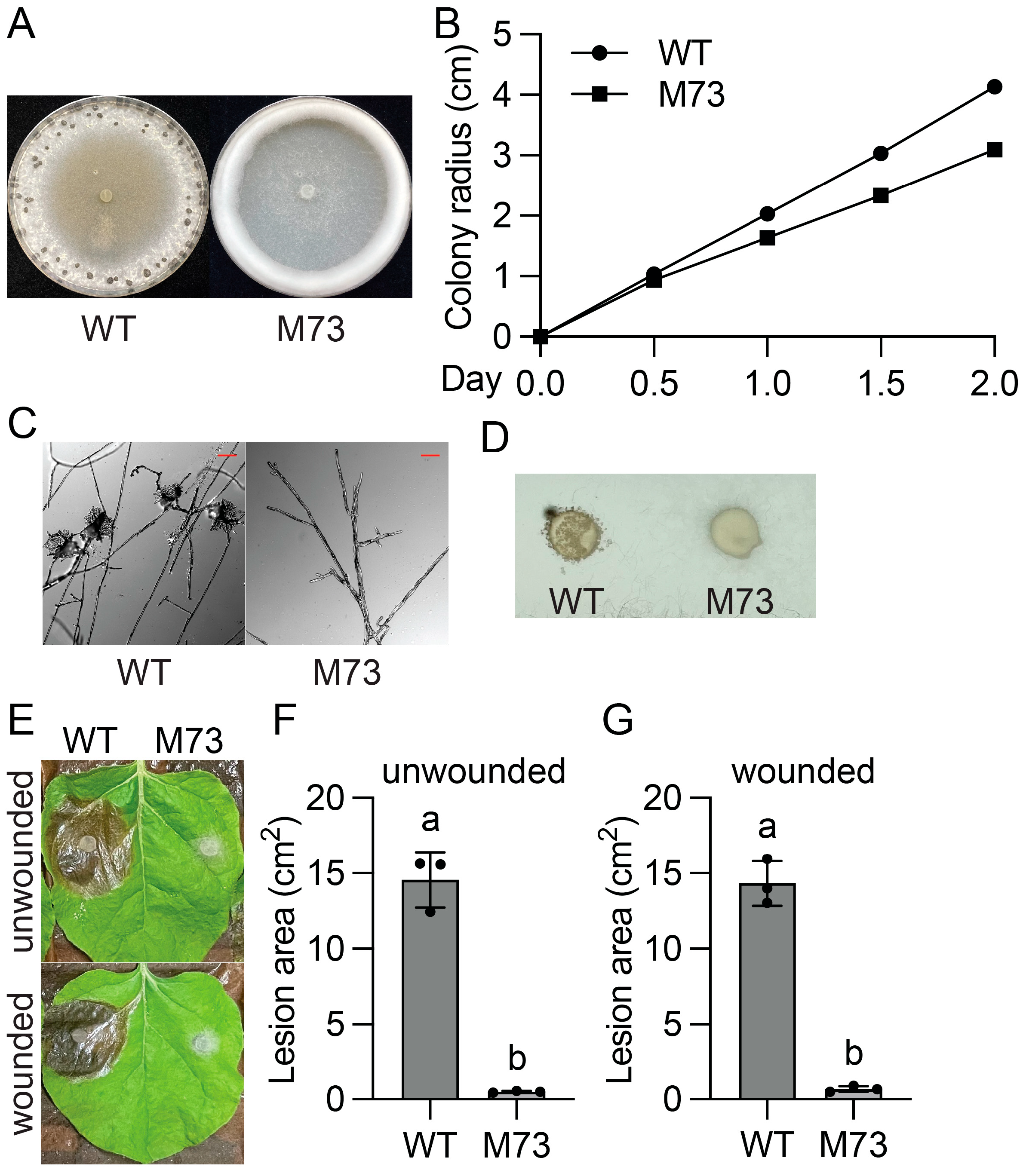
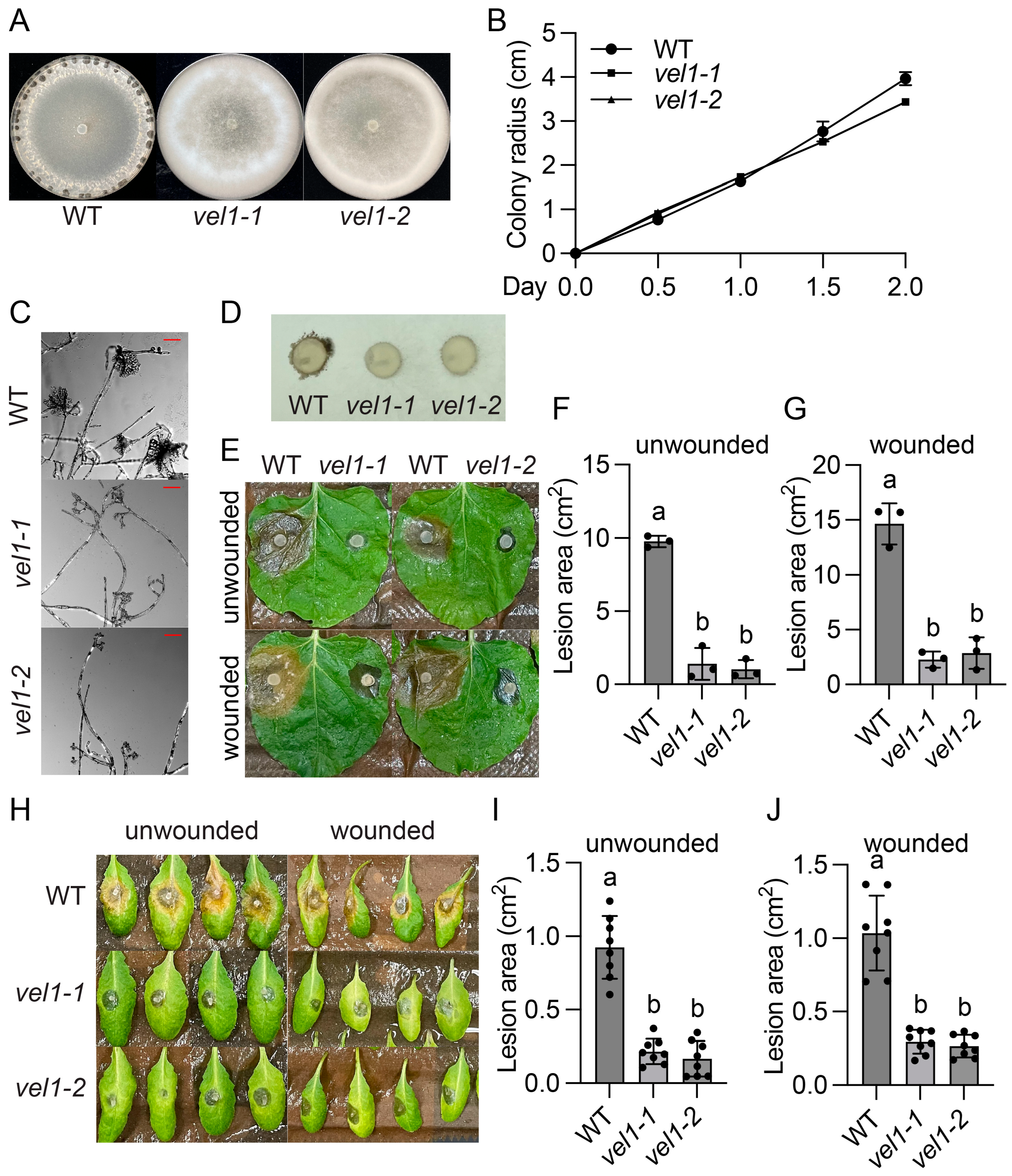
2.3. SsVel1 Is Also Required for Sclerotia Formation, Compound Appressorium Development, and Virulence of S. sclerotiorum
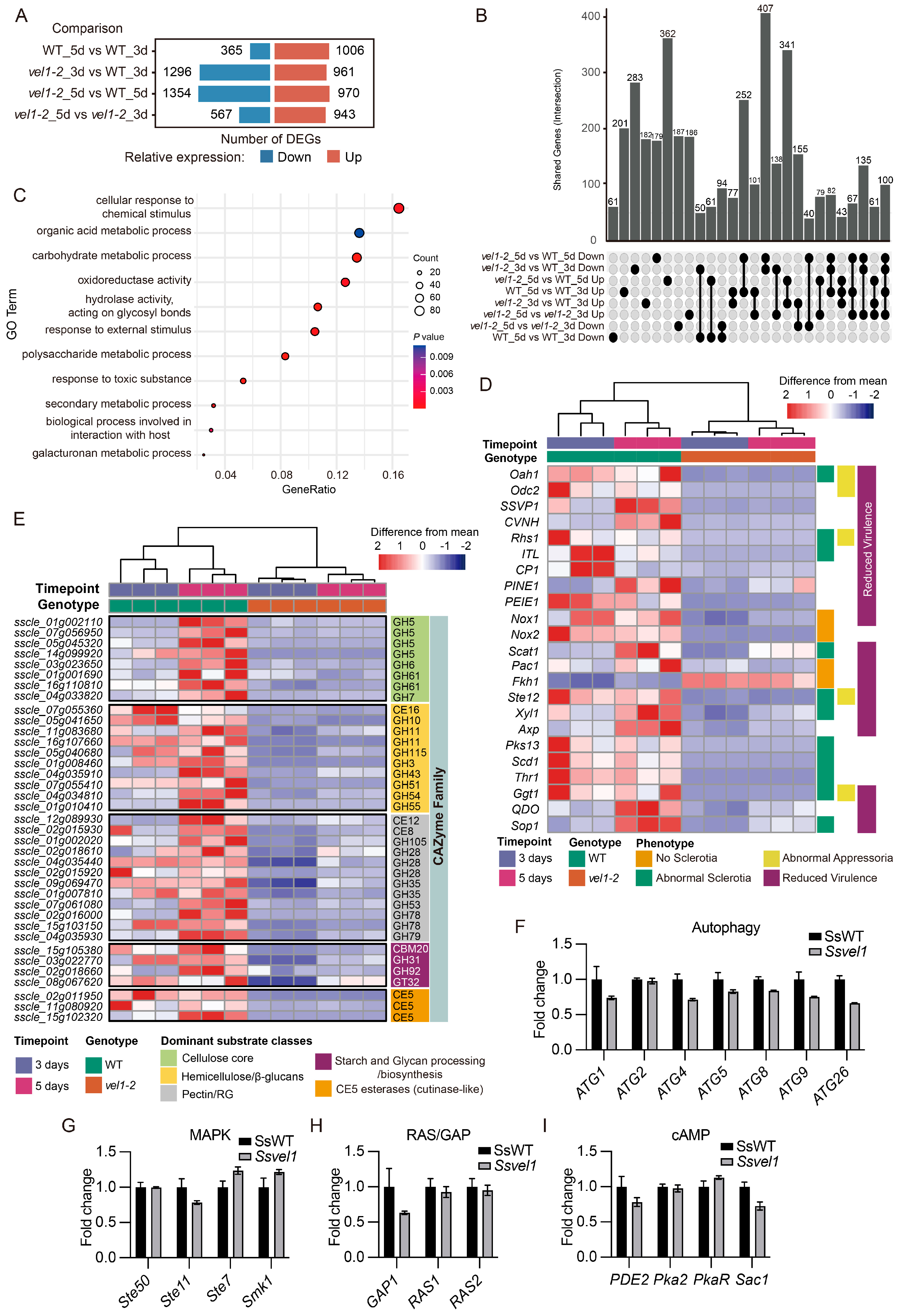
2.4. SsVel1 Broadly Regulates Genes Specific to Infection and Sclerotia Development
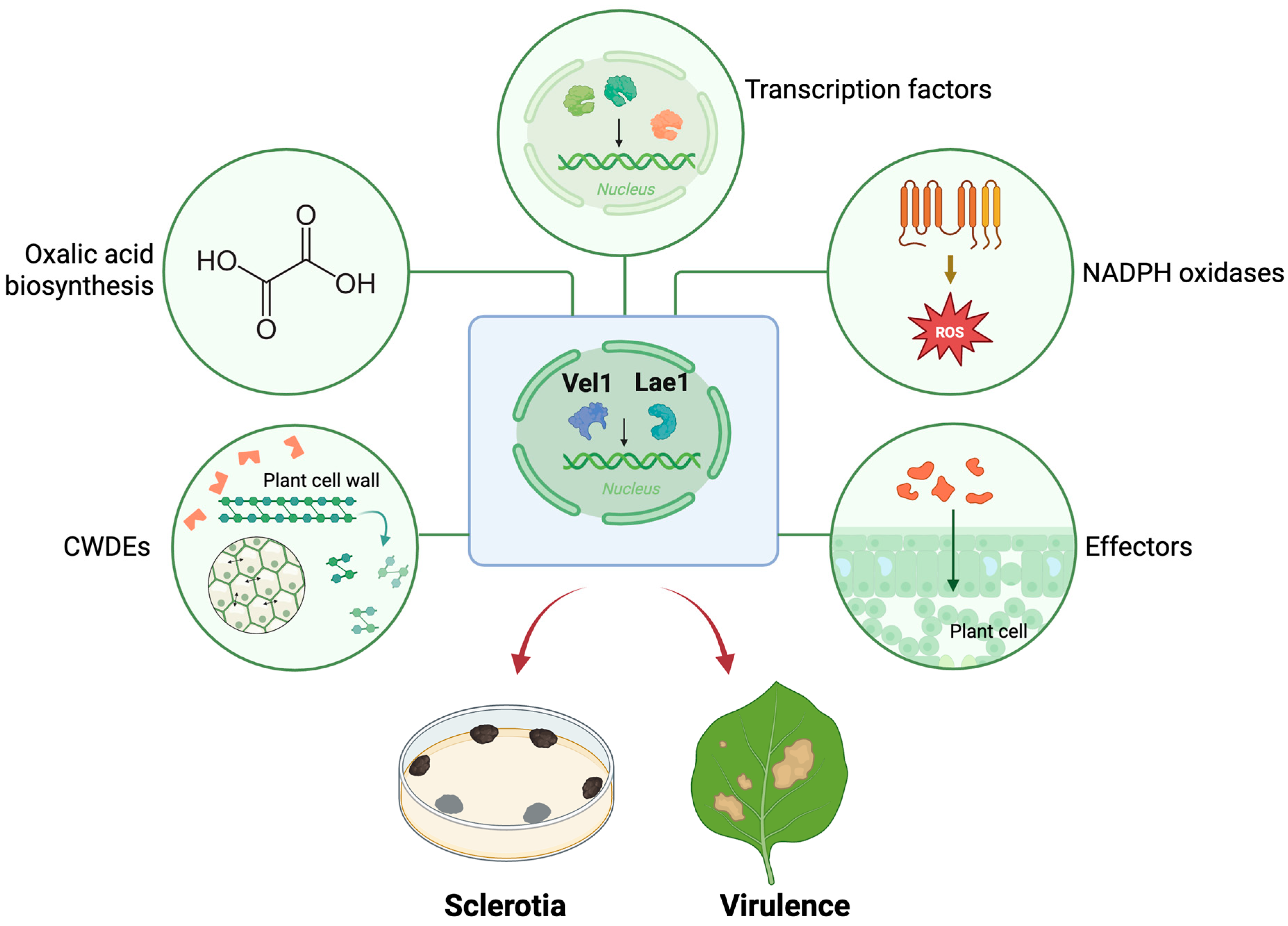
3. Discussion
4. Materials and Methods
4.1. Fungal Strains and Culture Conditions
4.2. UV Mutagenesis
4.3. Genomic DNA Extraction and NGS Analysis
4.4. Target Gene Knockout
4.5. Colony Morphology and Growth Rate Determination
4.6. Plant Infection Assay
4.7. Compound Appressoria Formation
4.8. RNA-seq Analysis
4.9. Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bolton, M.D.; Thomma, B.P.H.J.; Nelson, B.D. Sclerotinia sclerotiorum (Lib.) de Bary: Biology and Molecular Traits of a Cosmopolitan Pathogen. Mol. Plant Pathol. 2006, 7, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Xu, Y.; Hoy, R.; Zhang, J.; Qin, L.; Li, X. The Notorious Soilborne Pathogenic Fungus Sclerotinia sclerotiorum: An Update on Genes Studied with Mutant Analysis. Pathogens 2019, 9, 27. [Google Scholar] [CrossRef]
- Zhu, Y.; Wu, C.; Deng, Y.; Yuan, W.; Zhang, T.; Lu, J. Recent Advances in Virulence of a Broad Host Range Plant Pathogen Sclerotinia sclerotiorum: A Mini-Review. Front. Microbiol. 2024, 15, 1424130. [Google Scholar] [CrossRef] [PubMed]
- Weerasinghe, T.; Li, J.; Chen, X.; Gao, J.; Tian, L.; Xu, Y.; Gong, Y.; Huang, W.; Zhang, Y.; Jiang, L.; et al. Autophagy-Related Proteins (ATGs) Are Differentially Required for Development and Virulence of Sclerotinia sclerotiorum. J. Fungi 2025, 11, 391. [Google Scholar] [CrossRef]
- Zhang, H.; Li, Y.; Lai, W.; Huang, K.; Li, Y.; Wang, Z.; Chen, X.; Wang, A. SsATG8 and SsNBR1 Mediated-Autophagy Is Required for Fungal Development, Proteasomal Stress Response and Virulence in Sclerotinia sclerotiorum. Fungal Genet. Biol. 2021, 157, 103632. [Google Scholar] [CrossRef]
- Xu, Y.; Qiu, Y.; Zhang, Y.; Li, X. A CAMP Phosphodiesterase Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum. Front. Plant Sci. 2023, 14, 1175552. [Google Scholar] [CrossRef]
- Yu, P.-L.; Rollins, J.A. The CAMP-Dependent Protein Kinase A Pathway Perturbs Autophagy and Plays Important Roles in Development and Virulence of Sclerotinia sclerotiorum. Fungal Biol. 2022, 126, 20–34. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Liberti, D.; Li, M.; Kim, Y.T.; Hutchens, A.; Wilson, R.; Rollins, J.A. Oxaloacetate Acetylhydrolase Gene Mutants of Sclerotinia sclerotiorum Do Not Accumulate Oxalic Acid, but Do Produce Limited Lesions on Host Plants. Mol. Plant Pathol. 2015, 16, 559–571. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Xiang, M.; White, D.; Chen, W. PH Dependency of Sclerotial Development and Pathogenicity Revealed by Using Genetically Defined Oxalate-Minus Mutants of Sclerotinia sclerotiorum. Environ. Microbiol. 2015, 17, 2896–2909. [Google Scholar] [CrossRef]
- Tian, L.; Li, J.; Xu, Y.; Qiu, Y.; Zhang, Y.; Li, X. A MAP Kinase Cascade Broadly Regulates the Lifestyle of Sclerotinia sclerotiorum and Can Be Targeted by HIGS for Disease Control. Plant J. 2024, 118, 324–344. [Google Scholar] [CrossRef]
- Kim, H.; Chen, C.; Kabbage, M.; Dickman, M.B. Identification and Characterization of Sclerotinia sclerotiorum NADPH Oxidases. Appl. Environ. Microbiol. 2011, 77, 7721–7729. [Google Scholar] [CrossRef]
- Xu, Y.; Tan, J.; Lu, J.; Zhang, Y.; Li, X. RAS Signalling Genes Can Be Used as Host-Induced Gene Silencing Targets to Control Fungal Diseases Caused by Sclerotinia sclerotiorum and Botrytis cinerea. Plant Biotechnol. J. 2024, 22, 262–277. [Google Scholar] [CrossRef]
- Rollins, J.A. The Sclerotinia Sclerotiorum Pac1 Gene Is Required for Sclerotial Development and Virulence. Mol. Plant-Microbe Interact. 2003, 16, 785–795. [Google Scholar] [CrossRef]
- Xu, Y.; Ao, K.; Tian, L.; Qiu, Y.; Huang, X.; Liu, X.; Hoy, R.; Zhang, Y.; Rashid, K.Y.; Xia, S.; et al. A Forward Genetic Screen in Sclerotinia sclerotiorum Revealed the Transcriptional Regulation of Its Sclerotial Melanization Pathway. Mol. Plant-Microbe Interact. 2022, 35, 244–256. [Google Scholar] [CrossRef]
- Zhang, C.; Xu, Y.; Li, L.; Wu, M.; Fang, Z.; Tan, J.; Rollins, J.A.; Lin, H.; Huang, X.; Mansfield, S.D.; et al. A GDP-Mannose-1-Phosphate Guanylyltransferase as a Potential HIGS Target against Sclerotinia sclerotiorum. PLoS Pathog. 2025, 21, e1013129. [Google Scholar] [CrossRef]
- Bayram, Ö.; Krappmann, S.; Ni, M.; Jin, W.B.; Helmstaedt, K.; Valerius, O.; Braus-Stromeyer, S.; Kwon, N.J.; Keller, N.P.; Yu, J.H.; et al. VelB/VeA/LaeA Complex Coordinates Light Signal with Fungal Development and Secondary Metabolism. Science 2008, 320, 1504–1506. [Google Scholar] [CrossRef] [PubMed]
- Schumacher, J.; Simon, A.; Cohrs, K.C.; Traeger, S.; Porquier, A.; Dalmais, B.; Viaud, M.; Tudzynski, B. The VELVET Complex in the Gray Mold Fungus Botrytis Cinerea: Impact of BcLAE1 on Differentiation, Secondary Metabolism, and Virulence. Mol. Plant-Microbe Interact. 2015, 28, 659–674. [Google Scholar] [CrossRef] [PubMed]
- Bayram, Ö.S.; Dettmann, A.; Karahoda, B.; Moloney, N.M.; Ormsby, T.; McGowan, J.; Cea-Sánchez, S.; Miralles-Durán, A.; Brancini, G.T.P.; Luque, E.M.; et al. Control of Development, Secondary Metabolism and Light-Dependent Carotenoid Biosynthesis by the Velvet Complex of Neurospora Crassa. Genetics 2019, 212, 691–710. [Google Scholar] [CrossRef] [PubMed]
- Lyu, X.; Shen, C.; Fu, Y.; Xie, J.; Jiang, D.; Li, G.; Cheng, J. Comparative Genomic and Transcriptional Analyses of the Carbohydrate-Active Enzymes and Secretomes of Phytopathogenic Fungi Reveal Their Significant Roles during Infection and Development. Sci. Rep. 2015, 5, 15565. [Google Scholar] [CrossRef]
- Liang, Y.; Xiong, W.; Steinkellner, S.; Feng, J. Deficiency of the Melanin Biosynthesis Genes SCD1 and THR1 Affects Sclerotial Development and Vegetative Growth, but Not Pathogenicity, in Sclerotinia sclerotiorum. Mol. Plant Pathol. 2018, 19, 1444–1453. [Google Scholar] [CrossRef]
- Yu, Y.; Xiao, J.; Du, J.; Yang, Y.; Bi, C.; Qing, L. Disruption of the Gene Encoding Endo-β-1, 4-Xylanase Affects the Growth and Virulence of Sclerotinia sclerotiorum. Front. Microbiol. 2016, 7, 1787. [Google Scholar] [CrossRef] [PubMed]
- Yajima, W.; Liang, Y.; Kav, N.N.V. Gene Disruption of an Arabinofuranosidase/β-Xylosidase Precursor Decreases Sclerotinia sclerotiorum Virulence on Canola Tissue. Mol. Plant-Microbe Interact. 2009, 22, 783–789. [Google Scholar] [CrossRef]
- Li, J.; Zhang, Y.; Zhang, Y.; Yu, P.-L.; Pan, H.; Rollins, J.A. Introduction of Large Sequence Inserts by CRISPR-Cas9 To Create Pathogenicity Mutants in the Multinucleate Filamentous Pathogen Sclerotinia sclerotiorum. MBio 2018, 9, e00567-18. [Google Scholar] [CrossRef]
- Yarden, O.; Veluchamy, S.; Dickman, M.B.; Kabbage, M. Sclerotinia Sclerotiorum Catalase SCAT1 Affects Oxidative Stress Tolerance, Regulates Ergosterol Levels and Controls Pathogenic Development. Physiol. Mol. Plant Pathol. 2014, 85, 34–41. [Google Scholar] [CrossRef]
- Chen, J.; Ullah, C.; Reichelt, M.; Gershenzon, J.; Hammerbacher, A. Sclerotinia sclerotiorum Circumvents Flavonoid Defenses by Catabolizing Flavonol Glycosides and Aglycones. Plant Physiol. 2019, 180, 1975–1987. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Liang, X.; Rollins, J.A. Sclerotinia sclerotiorum γ-Glutamyl Transpeptidase (Ss-Ggt1) Is Required for Regulating Glutathione Accumulation and Development of Sclerotia and Compound Appressoria. Mol. Plant-Microbe Interact. 2012, 25, 412–420. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Xiao, J.; Zhu, W.; Yang, Y.; Mei, J.; Bi, C.; Qian, W.; Qing, L.; Tan, W. Ss-Rhs1, a Secretory Rhs Repeat-containing Protein, Is Required for the Virulence of Sclerotinia sclerotiorum. Mol. Plant Pathol. 2017, 18, 1052–1061. [Google Scholar] [CrossRef]
- Lyu, X.; Shen, C.; Fu, Y.; Xie, J.; Jiang, D.; Li, G.; Cheng, J. The Microbial Opsin Homolog Sop1 Is Involved in Sclerotinia sclerotiorum Development and Environmental Stress Response. Front. Microbiol. 2016, 6, 1504. [Google Scholar] [CrossRef]
- Wei, W.; Xu, L.; Peng, H.; Zhu, W.; Tanaka, K.; Cheng, J.; Sanguinet, K.A.; Vandemark, G.; Chen, W. A Fungal Extracellular Effector Inactivates Plant Polygalacturonase-Inhibiting Protein. Nat. Commun. 2022, 13, 2213. [Google Scholar] [CrossRef]
- Liu, X.; Zhao, H.; Yuan, M.; Li, P.; Xie, J.; Fu, Y.; Li, B.; Yu, X.; Chen, T.; Lin, Y.; et al. An Effector Essential for Virulence of Necrotrophic Fungi Targets Plant HIRs to Inhibit Host Immunity. Nat. Commun. 2024, 15, 9391. [Google Scholar] [CrossRef]
- Liang, X.; Moomaw, E.W.; Rollins, J.A. Fungal Oxalate Decarboxylase Activity Contributes to Sclerotinia sclerotiorum Early Infection by Affecting Both Compound Appressoria Development and Function. Mol. Plant Pathol. 2015, 16, 825–836. [Google Scholar] [CrossRef] [PubMed]
- Yang, G.; Tang, L.; Gong, Y.; Xie, J.; Fu, Y.; Jiang, D.; Li, G.; Collinge, D.B.; Chen, W.; Cheng, J. A Cerato-Platanin Protein SsCP1 Targets Plant PR1 and Contributes to Virulence of Sclerotinia sclerotiorum. New Phytol. 2018, 217, 739–755. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Wei, W.; Fu, Y.; Cheng, J.; Xie, J.; Li, G.; Yi, X.; Kang, Z.; Dickman, M.B.; Jiang, D. A Secretory Protein of Necrotrophic Fungus Sclerotinia sclerotiorum That Suppresses Host Resistance. PLoS ONE 2013, 8, e53901. [Google Scholar] [CrossRef]
- Lyu, X.; Shen, C.; Fu, Y.; Xie, J.; Jiang, D.; Li, G.; Cheng, J. A Small Secreted Virulence-Related Protein Is Essential for the Necrotrophic Interactions of Sclerotinia sclerotiorum with Its Host Plants. PLOS Pathog. 2016, 12, e1005435. [Google Scholar] [CrossRef]
- Kim, Y.T.; Prusky, D.; Rollins, J.A. An Activating Mutation of the Sclerotinia sclerotiorum Pac1 Gene Increases Oxalic Acid Production at Low PH but Decreases Virulence. Mol. Plant Pathol. 2007, 8, 611–622. [Google Scholar] [CrossRef]
- Fan, H.; Yu, G.; Liu, Y.; Zhang, X.; Liu, J.; Zhang, Y.; Rollins, J.A.; Sun, F.; Pan, H. An Atypical Forkhead-containing Transcription Factor SsFKH1 Is Involved in Sclerotial Formation and Is Essential for Pathogenicity in Sclerotinia sclerotiorum. Mol. Plant Pathol. 2017, 18, 963–975. [Google Scholar] [CrossRef] [PubMed]
- Ma, M.; Tang, L.; Sun, R.; Lyu, X.; Xie, J.; Fu, Y.; Li, B.; Chen, T.; Lin, Y.; Yu, X.; et al. An Effector SsCVNH Promotes the Virulence of Sclerotinia sclerotiorum through Targeting Class III Peroxidase AtPRX71. Mol. Plant Pathol. 2024, 25, e13464. [Google Scholar] [CrossRef]
- Amselem, J.; Cuomo, C.A.; van Kan, J.A.L.; Viaud, M.; Benito, E.P.; Couloux, A.; Coutinho, P.M.; de Vries, R.P.; Dyer, P.S.; Fillinger, S.; et al. Genomic Analysis of the Necrotrophic Fungal Pathogens Sclerotinia sclerotiorum and Botrytis Cinerea. PLoS Genet. 2011, 7, e1002230. [Google Scholar] [CrossRef]
- Bayram, Ö.; Braus, G.H. Coordination of Secondary Metabolism and Development in Fungi: The Velvet Family of Regulatory Proteins. FEMS Microbiol. Rev. 2012, 36, 1–24. [Google Scholar] [CrossRef]
- Wiemann, P.; Brown, D.W.; Kleigrewe, K.; Bok, J.W.; Keller, N.P.; Humpf, H.; Tudzynski, B. FfVel1 and FfLae1, Components of a Velvet-like Complex in Fusarium fujikuroi, Affect Differentiation, Secondary Metabolism and Virulence. Mol. Microbiol. 2010, 77, 972–994. [Google Scholar] [CrossRef]
- Park, H.S.; Nam, T.Y.; Han, K.H.; Kim, S.C.; Yu, J.H. VelC Positively Controls Sexual Development in Aspergillus nidulans. PLoS ONE 2014, 9, e89883. [Google Scholar] [CrossRef] [PubMed]
- Bayram, Ö.S.; Bayram, Ö.; Valerius, O.; Park, H.S.; Irniger, S.; Gerke, J.; Ni, M.; Han, K.H.; Yu, J.H.; Braus, G.H. Laea Control of Velvet Family Regulatory Proteins for Light-Dependent Development and Fungal Cell-Type Specificity. PLoS Genet. 2010, 6, e1001226. [Google Scholar] [CrossRef]
- Schumacher, J.; Pradier, J.M.; Simon, A.; Traeger, S.; Moraga, J.; Collado, I.G.; Viaud, M.; Tudzynski, B. Natural Variation in the VELVET Gene Bcvel1 Affects Virulence and Light-Dependent Differentiation in Botrytis Cinerea. PLoS ONE 2012, 7, e47840. [Google Scholar] [CrossRef]
- Müller, N.; Leroch, M.; Schumacher, J.; Zimmer, D.; Könnel, A.; Klug, K.; Leisen, T.; Scheuring, D.; Sommer, F.; Mühlhaus, T.; et al. Investigations on VELVET Regulatory Mutants Confirm the Role of Host Tissue Acidification and Secretion of Proteins in the Pathogenesis of Botrytis cinerea. New Phytol. 2018, 219, 1062–1074. [Google Scholar] [CrossRef]
- Cea-Sánchez, S.; Corrochano-Luque, M.; Gutiérrez, G.; Glass, N.L.; Cánovas, D.; Corrochano, L.M. Transcriptional Regulation by the Velvet Protein VE-1 during Asexual Development in the Fungus Neurospora crassa. MBio 2022, 13, e0150522. [Google Scholar] [CrossRef]
- Yang, Q.; Chen, Y.; Ma, Z. Involvement of BcVeA and BcVelB in Regulating Conidiation, Pigmentation and Virulence in Botrytis cinerea. Fungal Genet. Biol. 2013, 50, 63–71. [Google Scholar] [CrossRef] [PubMed]
- López-Berges, M.S.; Hera, C.; Sulyok, M.; Schäfer, K.; Capilla, J.; Guarro, J.; Di Pietro, A. The Velvet Complex Governs Mycotoxin Production and Virulence of Fusarium oxysporum on Plant and Mammalian Hosts. Mol. Microbiol. 2013, 87, 49–65. [Google Scholar] [CrossRef]
- Hoff, B.; Kamerewerd, J.; Sigl, C.; Mitterbauer, R.; Zadra, I.; Kürnsteiner, H.; Kück, U. Two Components of a Velvet-Like Complex Control Hyphal Morphogenesis, Conidiophore Development, and Penicillin Biosynthesis in Penicillium chrysogenum. Eukaryot. Cell 2010, 9, 1236–1250. [Google Scholar] [CrossRef] [PubMed]
- Athukorala, S.N.P.; Fernando, W.G.D.; Rashid, K.Y.; de Kievit, T. The Role of Volatile and Non-Volatile Antibiotics Produced by Pseudomonas chlororaphis Strain PA23 in Its Root Colonization and Control of Sclerotinia sclerotiorum. Biocontrol Sci. Technol. 2010, 20, 875–890. [Google Scholar] [CrossRef]
- Derbyshire, M.; Denton-Giles, M.; Hegedus, D.; Seifbarghi, S.; Rollins, J.; Van Kan, J.; Seidl, M.F.; Faino, L.; Mbengue, M.; Navaud, O.; et al. The Complete Genome Sequence of the Phytopathogenic Fungus Sclerotinia sclerotiorum Reveals Insights into the Genome Architecture of Broad Host Range Pathogens. Genome Biol. Evol. 2017, 9, 593–618. [Google Scholar] [CrossRef]
- Li, H. Aligning Sequence Reads, Clone Sequences and Assembly Contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Van der Auwera, G.A.; Carneiro, M.O.; Hartl, C.; Poplin, R.; del Angel, G.; Levy-Moonshine, A.; Jordan, T.; Shakir, K.; Roazen, D.; Thibault, J.; et al. From FastQ Data to High-Confidence Variant Calls: The Genome Analysis Toolkit Best Practices Pipeline. Curr. Protoc. Bioinforma. 2013, 43, 11.10.1–11.10.33. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.H.; Hamari, Z.; Han, K.H.; Seo, J.A.; Reyes-Domínguez, Y.; Scazzocchio, C. Double-Joint PCR: A PCR-Based Molecular Tool for Gene Manipulations in Filamentous fungi. Fungal Genet. Biol. 2004, 41, 973–981. [Google Scholar] [CrossRef]
- Kim, D.; Paggi, J.M.; Park, C.; Bennett, C.; Salzberg, S.L. Graph-Based Genome Alignment and Genotyping with HISAT2 and HISAT-Genotype. Nat. Biotechnol. 2019, 37, 907–915. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. FeatureCounts: An Efficient General Purpose Program for Assigning Sequence Reads to Genomic Features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-Throughput Functional Annotation and Data Mining with the Blast2GO Suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Villanueva, R.A.M.; Chen, Z.J. Ggplot2: Elegant Graphics for Data Analysis (2nd Ed.). Meas. Interdiscip. Res. Perspect. 2019, 17, 160–167. [Google Scholar] [CrossRef]
- Gu, Z.; Eils, R.; Schlesner, M. Complex Heatmaps Reveal Patterns and Correlations in Multidimensional Genomic Data. Bioinformatics 2016, 32, 2847–2849. [Google Scholar] [CrossRef]
- Conway, J.R.; Lex, A.; Gehlenborg, N. UpSetR: An R Package for the Visualization of Intersecting Sets and Their Properties. Bioinformatics 2017, 33, 2938–2940. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, W.; Chen, Z.; Cui, N.; Wijaya, J.; Xu, Y.; Wu, M.; Zhang, Y.; Li, X. The Velvet Complex Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum. J. Fungi 2025, 11, 786. https://doi.org/10.3390/jof11110786
Huang W, Chen Z, Cui N, Wijaya J, Xu Y, Wu M, Zhang Y, Li X. The Velvet Complex Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum. Journal of Fungi. 2025; 11(11):786. https://doi.org/10.3390/jof11110786
Chicago/Turabian StyleHuang, Weijie, Zhuo Chen, Ning Cui, Jessica Wijaya, Yan Xu, Mingsong Wu, Yuelin Zhang, and Xin Li. 2025. "The Velvet Complex Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum" Journal of Fungi 11, no. 11: 786. https://doi.org/10.3390/jof11110786
APA StyleHuang, W., Chen, Z., Cui, N., Wijaya, J., Xu, Y., Wu, M., Zhang, Y., & Li, X. (2025). The Velvet Complex Is Essential for Sclerotia Formation and Virulence in Sclerotinia sclerotiorum. Journal of Fungi, 11(11), 786. https://doi.org/10.3390/jof11110786






