Development of a Multiplex RT-PCR Assay for Simultaneous Detection of Four Potential Zoonotic Swine RNA Viruses
Abstract
:1. Introduction
2. Materials and Methods
2.1. Nucleic Acid Extraction and Reverse Transcription
2.2. Primer Design and Construction of SaV, EMCV, RVA and AstV Plasmids
2.3. Single RT-PCR and mRT-PCR Reaction Optimization
2.4. Assay Sensitivity, Specificity, and Reproducibility
2.5. Detecting Target Viruses from Field Samples Using mRT-PCR
2.6. Phylogenetic Analysis of the Detected Viruses
3. Results
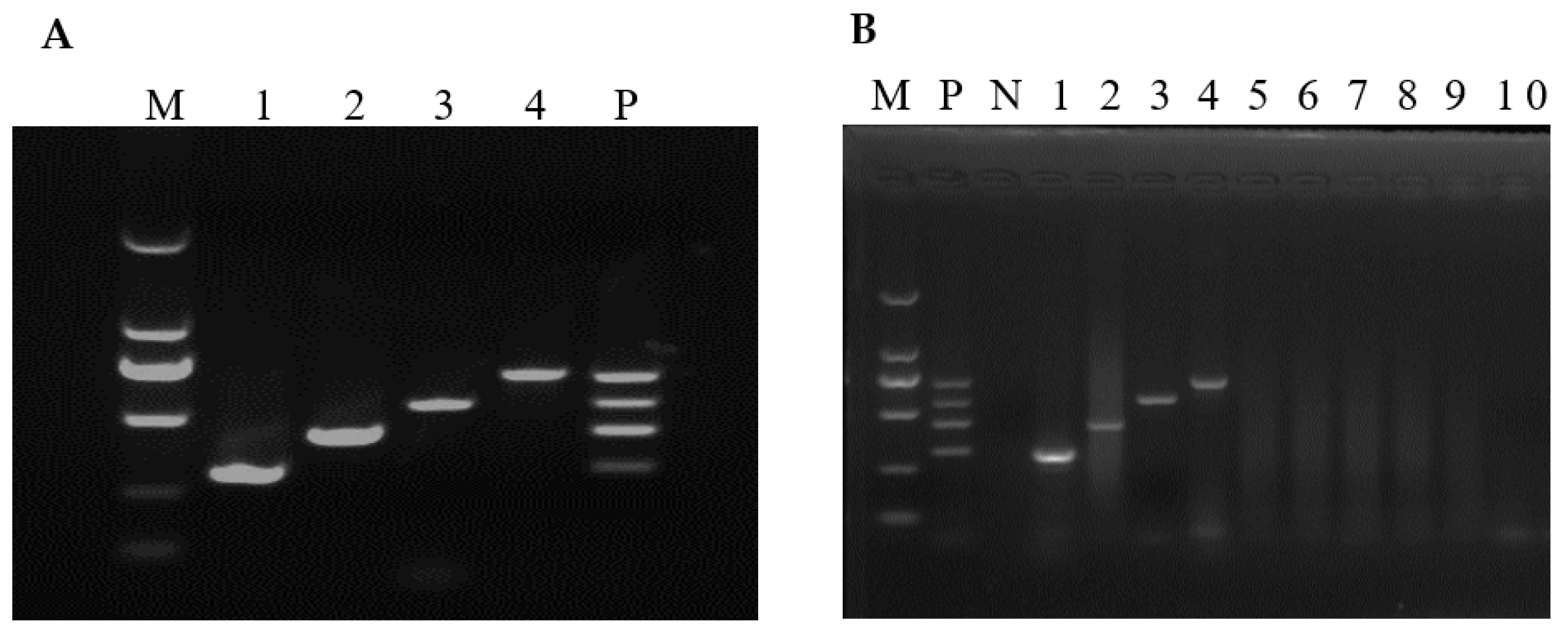
3.1. Optimized Conditions of the mRT-PCR
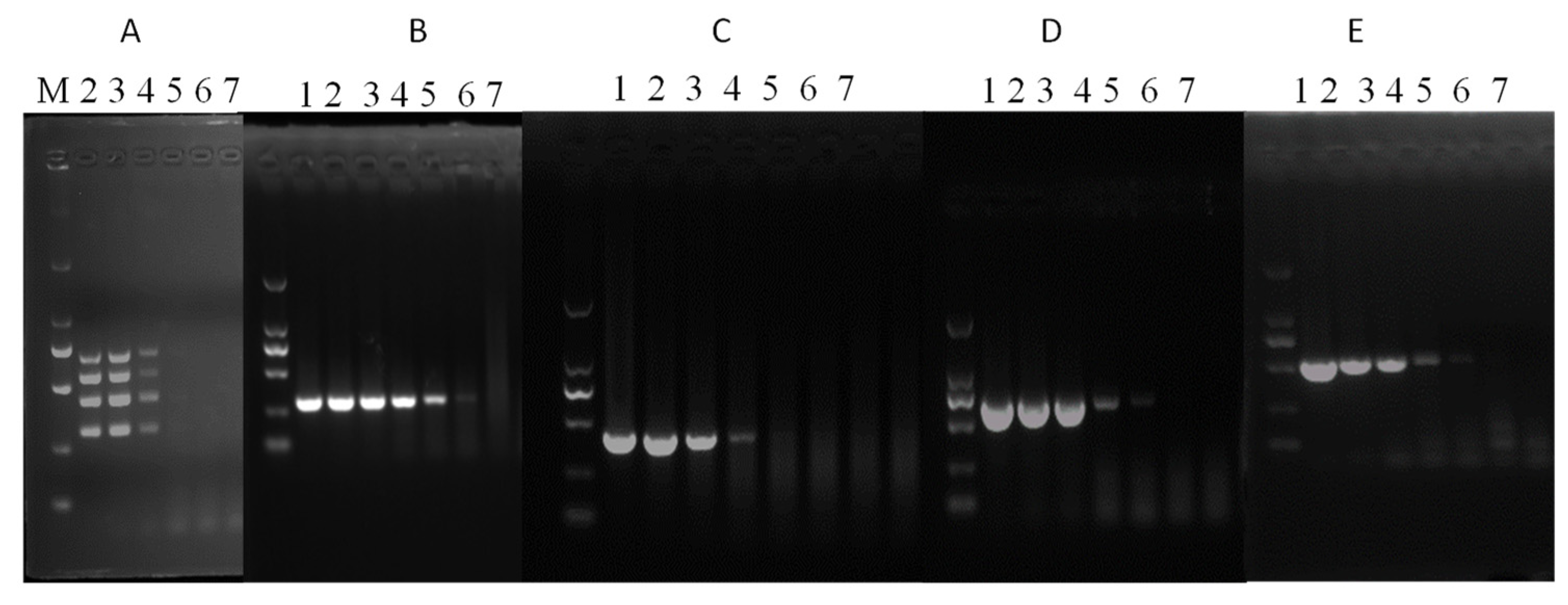
3.2. Assay Sensitivity
3.3. Assay Specificity
3.4. Assay Reproducibility
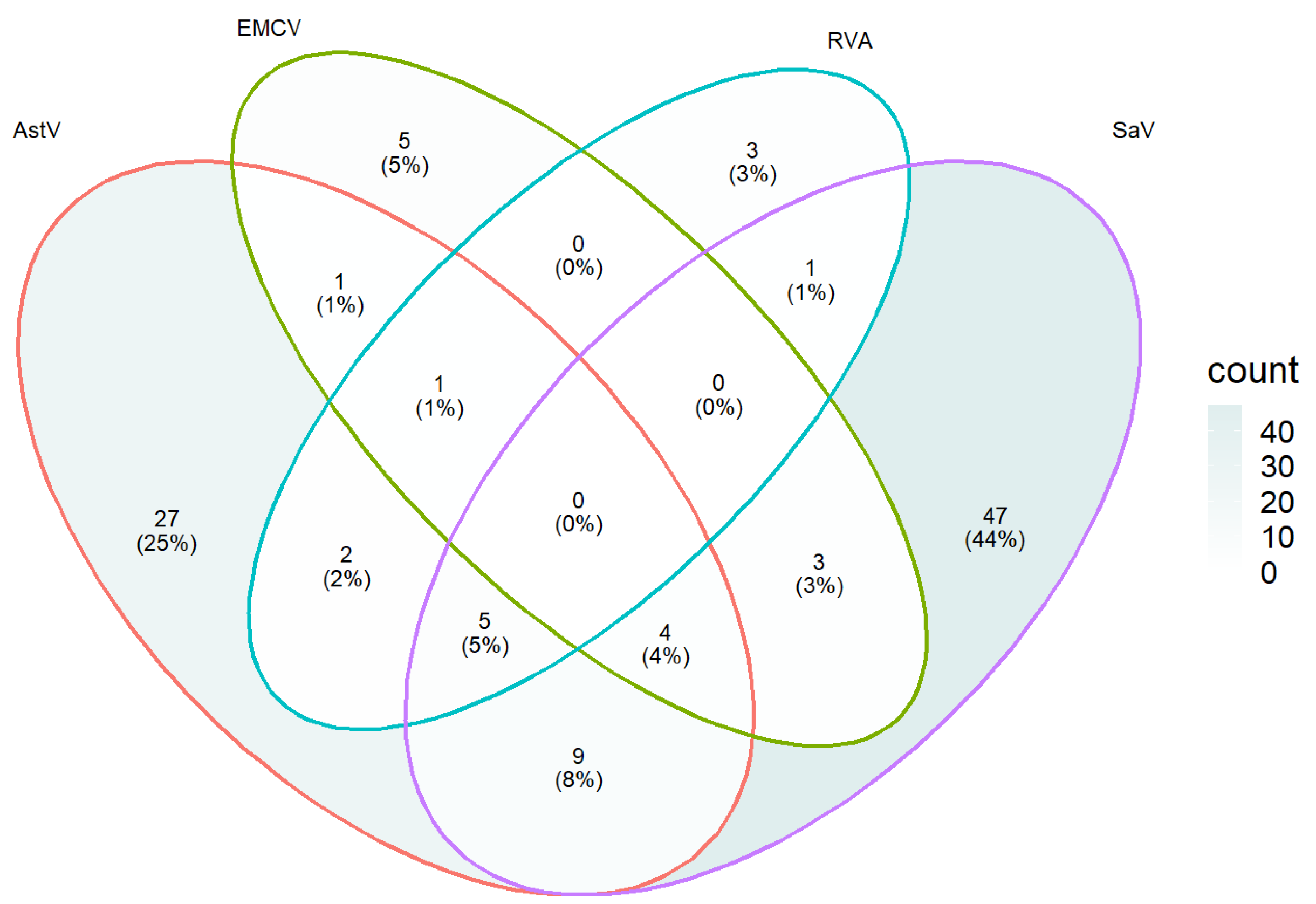
3.5. Detection of Field Samples
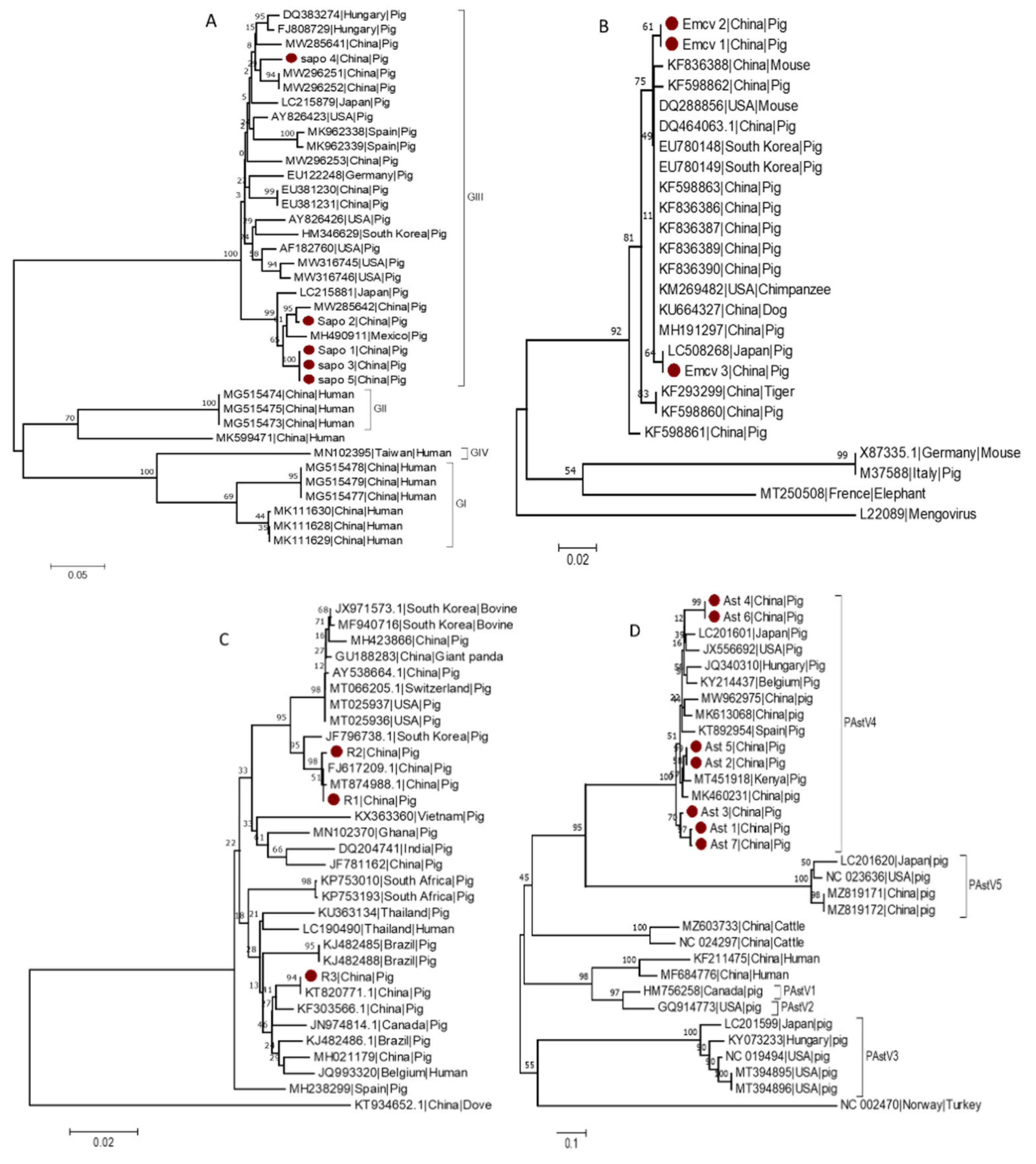
3.6. Phylogenetic Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Zhou, P.; Fan, H.; Lan, T.; Yang, X.-L.; Shi, W.-F.; Zhang, W.; Zhu, Y.; Zhang, Y.-W.; Xie, Q.-M.; Mani, S. Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 2018, 556, 255–258. [Google Scholar] [CrossRef] [PubMed]
- Ge, F.-F.; Yang, D.-Q.; Ju, H.-B.; Wang, J.; Liu, J.; Liu, P.-H.; Zhou, J.-P. Epidemiological survey of porcine epidemic diarrhea virus in swine farms in Shanghai, China. Arch. Virol. 2013, 158, 2227–2231. [Google Scholar] [CrossRef] [PubMed]
- Mandelik, R.; Sarvas, M.; Jackova, A.; Salamunova, S.; Novotny, J.; Vilcek, S. First outbreak with chimeric swine enteric coronavirus (SeCoV) on pig farms in Slovakia–lessons to learn. Acta Vet. Hung. 2018, 66, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Bank-Wolf, B.R.; König, M.; Thiel, H.-J. Zoonotic aspects of infections with noroviruses and sapoviruses. Vet. Microbiol. 2010, 140, 204–212. [Google Scholar] [CrossRef] [Green Version]
- Midgley, S.E.; Bányai, K.; Buesa, J.; Halaihel, N.; Hjulsager, C.K.; Jakab, F.; Kaplon, J.; Larsen, L.E.; Monini, M.; Poljšak-Prijatelj, M. Diversity and zoonotic potential of rotaviruses in swine and cattle across Europe. Vet. Microbiol. 2012, 156, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Saif, L.J. Comparative pathogenesis of enteric viral infections of swine. In Mechanisms in the Pathogenesis of Enteric Diseases 2; Paul, P.S., Francis, D.H., Eds.; Springer: Boston, MA, USA, 1999; Volume 473, pp. 47–59. [Google Scholar] [CrossRef]
- Zhou, W.; Ullman, K.; Chowdry, V.; Reining, M.; Benyeda, Z.; Baule, C.; Juremalm, M.; Wallgren, P.; Schwarz, L.; Zhou, E. Molecular investigations on the prevalence and viral load of enteric viruses in pigs from five European countries. Vet. Microbiol. 2016, 182, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Salamunova, S.; Jackova, A.; Mandelik, R.; Novotny, J.; Vlasakova, M.; Vilcek, S. Molecular detection of enteric viruses and the genetic characterization of porcine astroviruses and sapoviruses in domestic pigs from Slovakian farms. BMC Vet. Res. 2018, 14, 313. [Google Scholar] [CrossRef] [PubMed]
- Pan, Y.; Tian, X.; Qin, P.; Wang, B.; Zhao, P.; Yang, Y.-L.; Wang, L.; Wang, D.; Song, Y.; Zhang, X. Discovery of a novel swine enteric alphacoronavirus (SeACoV) in southern China. Vet. Microbiol. 2017, 211, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Theuns, S.; Conceicao-Neto, N.; Zeller, M.; Heylen, E.; Roukaerts, I.D.; Desmarets, L.M.; Van Ranst, M.; Nauwynck, H.J.; Matthijnssens, J. Characterization of a genetically heterogeneous porcine rotavirus C, and other viruses present in the fecal virome of a non-diarrheic Belgian piglet. Infect. Genet. Evol. 2016, 43, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Chao, D.Y.; Wei, J.Y.; Chang, W.F.; Wang, J.; Wang, L.C. Detection of multiple genotypes of calicivirus infection in asymptomatic swine in Taiwan. Zoonoses Public Health 2012, 59, 434–444. [Google Scholar] [CrossRef] [PubMed]
- Machnowska, P.; Ellerbroek, L.; Johne, R. Detection and characterization of potentially zoonotic viruses in faeces of pigs at slaughter in Germany. Vet. Microbiol. 2014, 168, 60–68. [Google Scholar] [CrossRef]
- Monini, M.; Di Bartolo, I.; Ianiro, G.; Angeloni, G.; Magistrali, C.F.; Ostanello, F.; Ruggeri, F.M. Detection and molecular characterization of zoonotic viruses in swine fecal samples in Italian pig herds. Arch. Virol. 2015, 160, 2547–2556. [Google Scholar] [CrossRef]
- Dufkova, L.; Scigalkova, I.; Moutelikova, R.; Malenovska, H.; Prodelalova, J. Genetic diversity of porcine sapoviruses, kobuviruses, and astroviruses in asymptomatic pigs: An emerging new sapovirus GIII genotype. Arch. Virol. 2013, 158, 549–558. [Google Scholar] [CrossRef] [PubMed]
- Mendenhall, I.H.; Smith, G.J.; Vijaykrishna, D. Ecological drivers of virus evolution: Astrovirus as a case study. J. Virol. 2015, 89, 6978–6981. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Czechowicz, J.; Huaman, J.L.; Forshey, B.M.; Morrison, A.C.; Castillo, R.; Huaman, A.; Caceda, R.; Eza, D.; Rocha, C.; Blair, P.J. Prevalence and risk factors for encephalomyocarditis virus infection in Peru. Vector-Borne Zoonotic Dis. 2011, 11, 367–374. [Google Scholar] [CrossRef] [PubMed]
- Martella, V.; Lorusso, E.; Banyai, K.; Decaro, N.; Corrente, M.; Elia, G.; Cavalli, A.; Radogna, A.; Costantini, V.; Saif, L. Identification of a porcine calicivirus related genetically to human sapoviruses. J. Clin. Microbiol. 2008, 46, 1907–1913. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vlasova, A.N.; Amimo, J.O.; Saif, L.J. Porcine Rotaviruses: Epidemiology, Immune Responses and Control Strategies. Viruses 2017, 9, 48. [Google Scholar] [CrossRef] [PubMed]
- Lipkin, W.I.; Firth, C. Viral surveillance and discovery. Curr. Opin. Virol. 2013, 3, 199–204. [Google Scholar] [CrossRef] [Green Version]
- Akdag, A.I.; Gupta, S.; Khan, N.; Upadhayay, A.; Ray, P. Epidemiology and clinical features of rotavirus, adenovirus, and astrovirus infections and coinfections in children with acute gastroenteritis prior to rotavirus vaccine introduction in Meerut, North India. J. Med. Virol. 2020, 92, 1102–1109. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Hu, R.; Tang, X.; Wu, C.; He, Q.; Zhao, Z.; Chen, H.; Wu, B. Occurrence and investigation of enteric viral infections in pigs with diarrhea in China. Arch. Virol. 2013, 158, 1631–1636. [Google Scholar] [CrossRef] [PubMed]
- Valko, A.; Marosi, A.; Csagola, A.; Farkas, R.; Ronai, Z.; Dan, A. Frequency of diarrhoea-associated viruses in swine of various ages in Hungary. Acta Vet. Hung. 2019, 67, 140–150. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Shi, B.J.; Huang, X.G.; Peng, M.Y.; Zhang, X.M.; He, D.N.; Pang, R.; Zhou, B.; Chen, P.Y. A multiplex RT-PCR assay for rapid and differential diagnosis of four porcine diarrhea associated viruses in field samples from pig farms in East China from 2010 to 2012. J. Virol. Methods 2013, 194, 107–112. [Google Scholar] [CrossRef]
- Yue, F.; Cui, S.; Zhang, C.; Yoon, K.J. A multiplex PCR for rapid and simultaneous detection of porcine circovirus type 2, porcine parvovirus, porcine pseudorabies virus, and porcine reproductive and respiratory syndrome virus in clinical specimens. Virus Genes 2009, 38, 392–397. [Google Scholar] [CrossRef]
- Ding, G.; Fu, Y.; Li, B.; Chen, J.; Wang, J.; Yin, B.; Sha, W.; Liu, G. Development of a multiplex RT-PCR for the detection of major diarrhoeal viruses in pig herds in China. Transbound. Emerg. Dis. 2020, 67, 678–685. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Zhao, Y.; Hu, Q.; Lv, C.; Zhang, C.; Zhao, R.; Hu, F.; Lin, W.; Cui, S. A multiplex RT-PCR for rapid and simultaneous detection of porcine teschovirus, classical swine fever virus, and porcine reproductive and respiratory syndrome virus in clinical specimens. J. Virol. Methods 2011, 172, 88–92. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3—New capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547. [Google Scholar] [CrossRef]
- Oka, T.; Lu, Z.; Phan, T.; Delwart, E.L.; Saif, L.J.; Wang, Q. Genetic characterization and classification of human and animal sapoviruses. PLoS ONE 2016, 11, e0156373. [Google Scholar] [CrossRef] [PubMed]
- Scheuer, K.A.; Oka, T.; Hoet, A.E.; Gebreyes, W.A.; Molla, B.Z.; Saif, L.J.; Wang, Q. Prevalence of porcine noroviruses, molecular characterization of emerging porcine sapoviruses from finisher swine in the United States, and unified classification scheme for sapoviruses. J. Clin. Microbiol. 2013, 51, 2344–2353. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yinda, C.K.; Conceição-Neto, N.; Zeller, M.; Heylen, E.; Maes, P.; Ghogomu, S.M.; Van Ranst, M.; Matthijnssens, J. Novel highly divergent sapoviruses detected by metagenomics analysis in straw-colored fruit bats in Cameroon: Divergent bat sapoviruses. Emerg. Microbes Infect. 2017, 6, 1–7. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Marthaler, D.; Fredrickson, R.; Gauger, P.C.; Zhang, J.; Burrough, E.R.; Petznick, T.; Li, G. Genetically divergent porcine sapovirus identified in pigs, United States. Transbound. Emerg. Dis. 2020, 67, 18–28. [Google Scholar] [CrossRef]
- Carocci, M.; Bakkali-Kassimi, L. The encephalomyocarditis virus. Virulence 2012, 3, 351–367. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, H.; Li, Y.; Zhang, G.; Sang, S.; Wang, C.; Chang, H. Complete genome sequences and phylogenetic analysis of encephalomyocarditis virus strains isolated from pigs and rats origin. Infect. Genet. Evol. 2017, 55, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Templeton, K.E.; Scheltinga, S.A.; Graffelman, A.W.; Van Schie, J.M.; Crielaard, J.W.; Sillekens, P.; Van Den Broek, P.J.; Goossens, H.; Beersma, M.F.; Claas, E.C. Comparison and evaluation of real-time PCR, real-time nucleic acid sequence-based amplification, conventional PCR, and serology for diagnosis of Mycoplasma pneumoniae. J. Clin. Microbiol. 2003, 41, 4366–4371. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ogawa, H.; Taira, O.; Hirai, T.; Takeuchi, H.; Nagao, A.; Ishikawa, Y.; Tuchiya, K.; Nunoya, T.; Ueda, S. Multiplex PCR and multiplex RT-PCR for inclusive detection of major swine DNA and RNA viruses in pigs with multiple infections. J. Virol. Methods 2009, 160, 210–214. [Google Scholar] [CrossRef]
- Xu, X.G.; Chen, G.D.; Huang, Y.; Ding, L.; Li, Z.C.; Chang, C.D.; Wang, C.Y.; Tong, D.W.; Liu, H.J. Development of multiplex PCR for simultaneous detection of six swine DNA and RNA viruses. J. Virol. Methods 2012, 183, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Shi, K.; Sun, W.; Zhao, J.; Yin, Y.; Si, H.; Qu, S.; Lu, W. Development a multiplex RT-PCR assay for simultaneous detection of African swine fever virus, classical swine fever virus and atypical porcine pestivirus. J. Virol. Methods 2021, 287, 114006. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Lin, X.Y.; Yang, Z.X.; Yao, X.P.; Li, G.L.; Peng, S.Z.; Wang, Y. A multiplex PCR for simultaneous detection of classical swine fever virus, African swine fever virus, highly pathogenic porcine reproductive and respiratory syndrome virus, porcine reproductive and respiratory syndrome virus and pseudorabies in swines. Pol. J. Vet. Sci. 2015, 18, 715–723. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakhaie, M.; Soleimanjahi, H.; Mollaie, H.R.; Arabzadeh, S.M.A. Development of multiplex reverse transcription-polymerase chain reaction for simultaneous detection of influenza A, B and adenoviruses. Iran. J. Pathol. 2018, 13, 54. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.; Choi, C.; Kim, B.; Chae, C. Detection and differentiation of porcine epidemic diarrhoea virus and transmissible gastroenteritis virus in clinical samples by multiplex RT-PCR. Vet. Rec. 2000, 146, 637–640. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Hung, J.J.; Wu, C.Y.; Chien, M.S. Multiplex PCR for rapid detection of pseudorabies virus, porcine parvovirus and porcine circoviruses. Vet. Microbiol. 2004, 101, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Day, J.M.; Spackman, E.; Pantin-Jackwood, M. A multiplex RT-PCR test for the differential identification of turkey astrovirus type 1, turkey astrovirus type 2, chicken astrovirus, avian nephritis virus, and avian rotavirus. Avian. Dis. 2007, 51, 681–684. [Google Scholar] [CrossRef]
- Cagirgan, A.A.; Yazici, Z. Development of a multiplex RT-PCR assay for the routine detection of seven RNA viruses in Apis mellifera. J. Virol. Methods 2020, 281, 113858. [Google Scholar] [CrossRef]
- Zhao, Z.P.; Yang, Z.; Lin, W.D.; Wang, W.Y.; Yang, J.; Jin, W.J.; Qin, A.J. The rate of co-infection for piglet diarrhea viruses in China and the genetic characterization of porcine epidemic diarrhea virus and porcine kobuvirus. Acta Virol. 2016, 60, 55–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Q.; Wang, L.; Zheng, Y.; Zhang, J.; Guo, B.; Yoon, K.-J.; Gauger, P.C.; Harmon, K.M.; Main, R.G.; Li, G. Metagenomic analysis of the RNA fraction of the fecal virome indicates high diversity in pigs infected by porcine endemic diarrhea virus in the United States. Virol. J. 2018, 15, 95. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Theuns, S.; Vanmechelen, B.; Bernaert, Q.; Deboutte, W.; Vandenhole, M.; Beller, L.; Matthijnssens, J.; Maes, P.; Nauwynck, H.J. Nanopore sequencing as a revolutionary diagnostic tool for porcine viral enteric disease complexes identifies porcine kobuvirus as an important enteric virus. Sci. Rep. 2018, 8, 9830. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Shen, Q.; Zhang, W.; Zhao, T.; Li, Y.; Jiang, J.; Yu, X.; Guo, Z.; Cui, L.; Hua, X. Genomic organization and recombination analysis of a porcine sapovirus identified from a piglet with diarrhea in China. Virol. J. 2017, 14, 57. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, Z.K.; Li, J.Y.; Pan, H. Seroprevalence and molecular detection of porcine sapovirus in symptomatic suckling piglets in Guangdong Province, China. Trop. Anim. Health Prod. 2014, 46, 583–587. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.; Chen, L.; Zhao, J.; Wang, X.; Yang, X.; Yao, H.; Wang, C. Complete genome sequence of porcine encephalomyocarditis virus from an aardvark in China. Genome Announc. 2014, 2, e00017-14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yuan, W.; Song, Q.; Zhang, X.; Zhang, L.; Sun, J. Isolation and molecular analysis of porcine encephalomyocarditis virus strain BD2 from northern China. Infect. Genet. Evol. 2014, 21, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Matthijnssens, J.; Ciarlet, M.; McDonald, S.M.; Attoui, H.; Bányai, K.; Brister, J.R.; Buesa, J.; Esona, M.D.; Estes, M.K.; Gentsch, J.R. Uniformity of rotavirus strain nomenclature proposed by the Rotavirus Classification Working Group (RCWG). Arch. Virol. 2011, 156, 1397–1413. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xiao, C.-T.; Giménez-Lirola, L.G.; Gerber, P.F.; Jiang, Y.-H.; Halbur, P.G.; Opriessnig, T. Identification and characterization of novel porcine astroviruses (PAstVs) with high prevalence and frequent co-infection of individual pigs with multiple PAstV types. J. Gen. Virol. 2013, 94, 570–582. [Google Scholar] [CrossRef] [PubMed]
- Kumthip, K.; Khamrin, P.; Saikruang, W.; Kongkaew, A.; Vachirachewin, R.; Ushijima, H.; Maneekarn, N. Detection and genetic characterization of porcine astroviruses in piglets with and without diarrhea in Thailand. Arch. Virol. 2018, 163, 1823–1829. [Google Scholar] [CrossRef] [PubMed]
- Su, M.; Qi, S.; Yang, D.; Guo, D.; Yin, B.; Sun, D. Coinfection and Genetic Characterization of Porcine Astrovirus in Diarrheic Piglets in China from 2015 to 2018. Front. Vet. Sci. 2020, 7, 462. [Google Scholar] [CrossRef] [PubMed]
- Xiao, C.-T.; Luo, Z.; Lv, S.-L.; Opriessnig, T.; Li, R.-C.; Yu, X.-L. Identification and characterization of multiple porcine astrovirus genotypes in Hunan province, China. Arch. Virol. 2017, 162, 943–952. [Google Scholar] [CrossRef] [Green Version]
- Boniotti, M.B.; Papetti, A.; Lavazza, A.; Alborali, G.; Sozzi, E.; Chiapponi, C.; Faccini, S.; Bonilauri, P.; Cordioli, P.; Marthaler, D. Porcine Epidemic Diarrhea Virus and Discovery of a Recombinant Swine Enteric Coronavirus, Italy. Emerg. Infect. Dis. 2016, 22, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Akimkin, V.; Beer, M.; Blome, S.; Hanke, D.; Höper, D.; Jenckel, M.; Pohlmann, A. New chimeric porcine coronavirus in swine feces, Germany, 2012. Emerg. Infect. Dis. 2016, 22, 1314. [Google Scholar] [CrossRef] [Green Version]
| Primer | Sequence | Target Gene | Accession No. | Target Region (bp) | Amplicon Size (bp) |
|---|---|---|---|---|---|
| SaV-F | AGCCAGAAGTGTTCGTGATGG | ORF1 | MK965898.1 | 5124–5429 | 306 |
| SaV-R | GGACARGTGRAGYGTGTARGG | ||||
| EMCV-F | CCGTCAAGTCTTCCAACCAG | 3D | MH191297.1 | 6288–6725 | 438 |
| EMCV-R | GCGGCTTGAACCTTCTCTATC | ||||
| RVA-F | GCAAACGAAGTCTTCGACATGG | VP6 | MH308723.1 | 6–575 | 570 |
| RVA-R | GGCGTTAATCCACATAGTYCCCA | ||||
| AstV -F | TTGTGGAGCTTGACTGGACC | ORF1ab | MK613068.1 | 3341–4042 | 702 |
| AstV-R | CTGTGAGTCTTGCAGGCAGA |
| Age Group | Health Status | Number of Samples (n) | % (Number of Positive/n) | |||
|---|---|---|---|---|---|---|
| SaV | EMCV | PRVA | AstV | |||
| Piglet | Diarrheic | 31 | 71 (22/31) | 19.4 (6/31) | 19.4 (6/31) | 41.9 (13/31) |
| Non-Diarrheic | 41 | 7.3 (3/41) | 2.4 (1/41) | 2.4 (1/41) | 17.1 (7/41) | |
| Sub-total | 72 | 34.7 (25/72) | 9.7 (7/72) | 9.7 (7/72) | 27.8 (20/72) | |
| Weaner | Diarrheic | 12 | 75 (9/12) | 0.0 | 16.7 (2/12) | 50 (6/12) |
| Non-Diarrheic | 30 | 10 (3/30) | 13.3 (4/30) | 3.3 (1/30) | 16.7 (5/30) | |
| Sub-total | 42 | 28.6 (12/42) | 9.5 (4/42) | 7.1 (3/42) | 26.2 (11/42) | |
| Fattening pig | Diarrheic | 20 | 80 (16/20) | 15 (3/20) | 5 (1/20) | 30 (6/20) |
| Non-Diarrheic | 42 | 2.4 (1/42) | 0 | 0 | 7.1 (3/42) | |
| Sub-total | 62 | 27.4 (17/62) | 4.8 (3/62) | 1.6 (1/62) | 14.5 (9/62) | |
| Sow | Diarrheic | 19 | 78.9 (15/19) | 0 | 5.3 (1/19) | 26.3 (5/19) |
| Non-Diarrheic | 85 | 0 | 0 | 0 | 4.7 (4/85) | |
| Sub-total | 104 | 14.4 (15/104) | 0.0 | 1 (1/104) | 8.7 (9/104) | |
| 280 | 24.6 (69/280) | 5 (14/280) | 4.3 (12/280) | 17.5 (49/280) | ||
| Assay | Number of Positive Samples | |||
|---|---|---|---|---|
| SaV | EMCV | RVA | AstV | |
| sRT-PCR | 29 | 8 | 2 | 13 |
| mRT-PCR | 27 * | 8 | 2 | 13 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Werid, G.M.; Zhang, H.; Ibrahim, Y.M.; Pan, Y.; Zhang, L.; Xu, Y.; Zhang, W.; Wang, W.; Chen, H.; Fu, L.; et al. Development of a Multiplex RT-PCR Assay for Simultaneous Detection of Four Potential Zoonotic Swine RNA Viruses. Vet. Sci. 2022, 9, 176. https://doi.org/10.3390/vetsci9040176
Werid GM, Zhang H, Ibrahim YM, Pan Y, Zhang L, Xu Y, Zhang W, Wang W, Chen H, Fu L, et al. Development of a Multiplex RT-PCR Assay for Simultaneous Detection of Four Potential Zoonotic Swine RNA Viruses. Veterinary Sciences. 2022; 9(4):176. https://doi.org/10.3390/vetsci9040176
Chicago/Turabian StyleWerid, Gebremeskel Mamu, He Zhang, Yassein M. Ibrahim, Yu Pan, Lin Zhang, Yunfei Xu, Wenli Zhang, Wei Wang, Hongyan Chen, Lizhi Fu, and et al. 2022. "Development of a Multiplex RT-PCR Assay for Simultaneous Detection of Four Potential Zoonotic Swine RNA Viruses" Veterinary Sciences 9, no. 4: 176. https://doi.org/10.3390/vetsci9040176
APA StyleWerid, G. M., Zhang, H., Ibrahim, Y. M., Pan, Y., Zhang, L., Xu, Y., Zhang, W., Wang, W., Chen, H., Fu, L., & Wang, Y. (2022). Development of a Multiplex RT-PCR Assay for Simultaneous Detection of Four Potential Zoonotic Swine RNA Viruses. Veterinary Sciences, 9(4), 176. https://doi.org/10.3390/vetsci9040176








