Reduced FXR Target Gene Expression in Copper-Laden Livers of COMMD1-Deficient Dogs
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics, Animals, and Samples
2.2. Genotyping and Phenotyping
2.3. RNA Isolation, cDNA Synthesis, and q-RT-PCR
2.4. Isolation of Canine Biliary Duct Fragments and Liver Organoid Culture
2.5. Statistical Analysis
3. Results
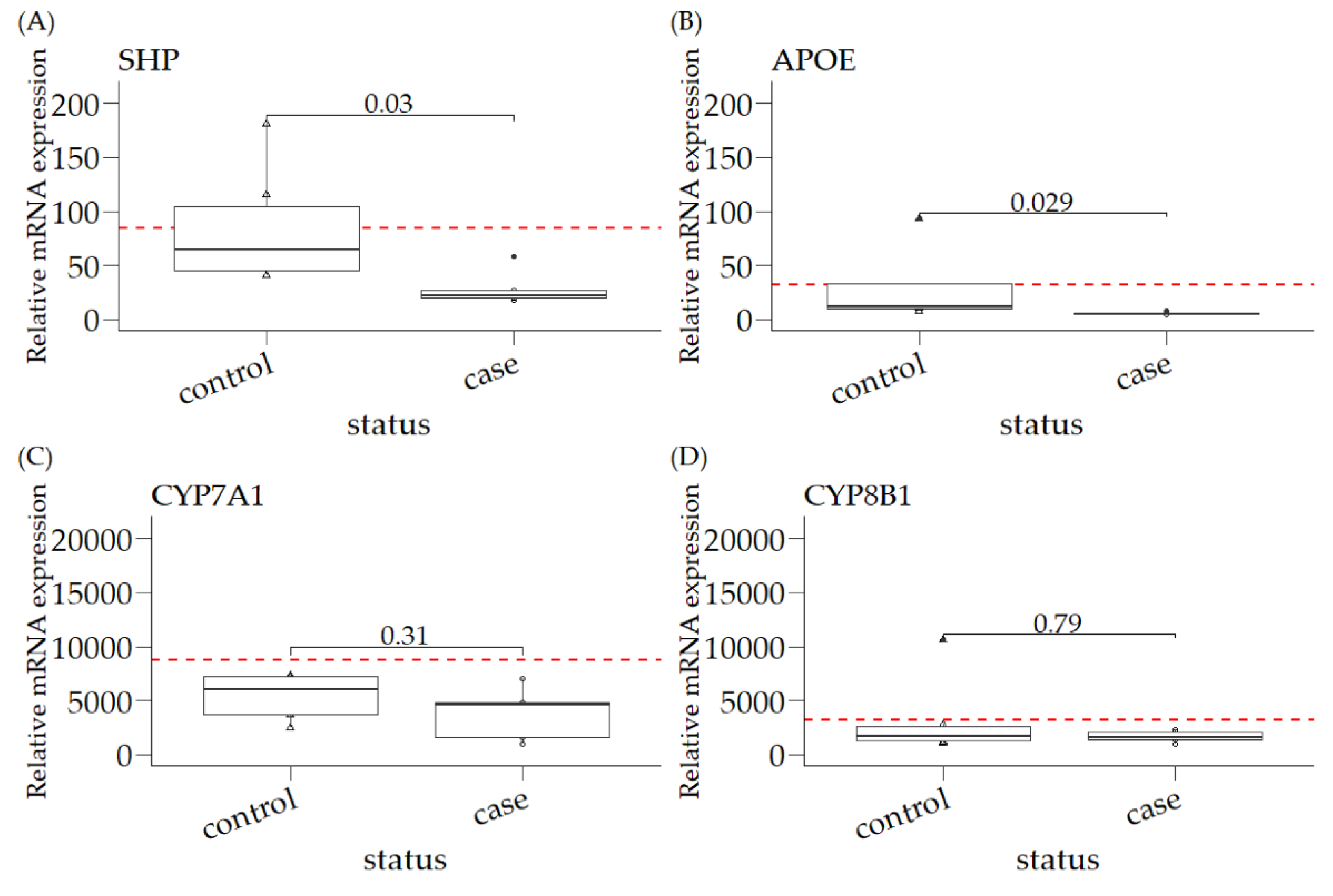
3.1. Relative mRNA Expression of SHP, APOE, CYP7A1, and CYP8B1 in Canine Liver Samples
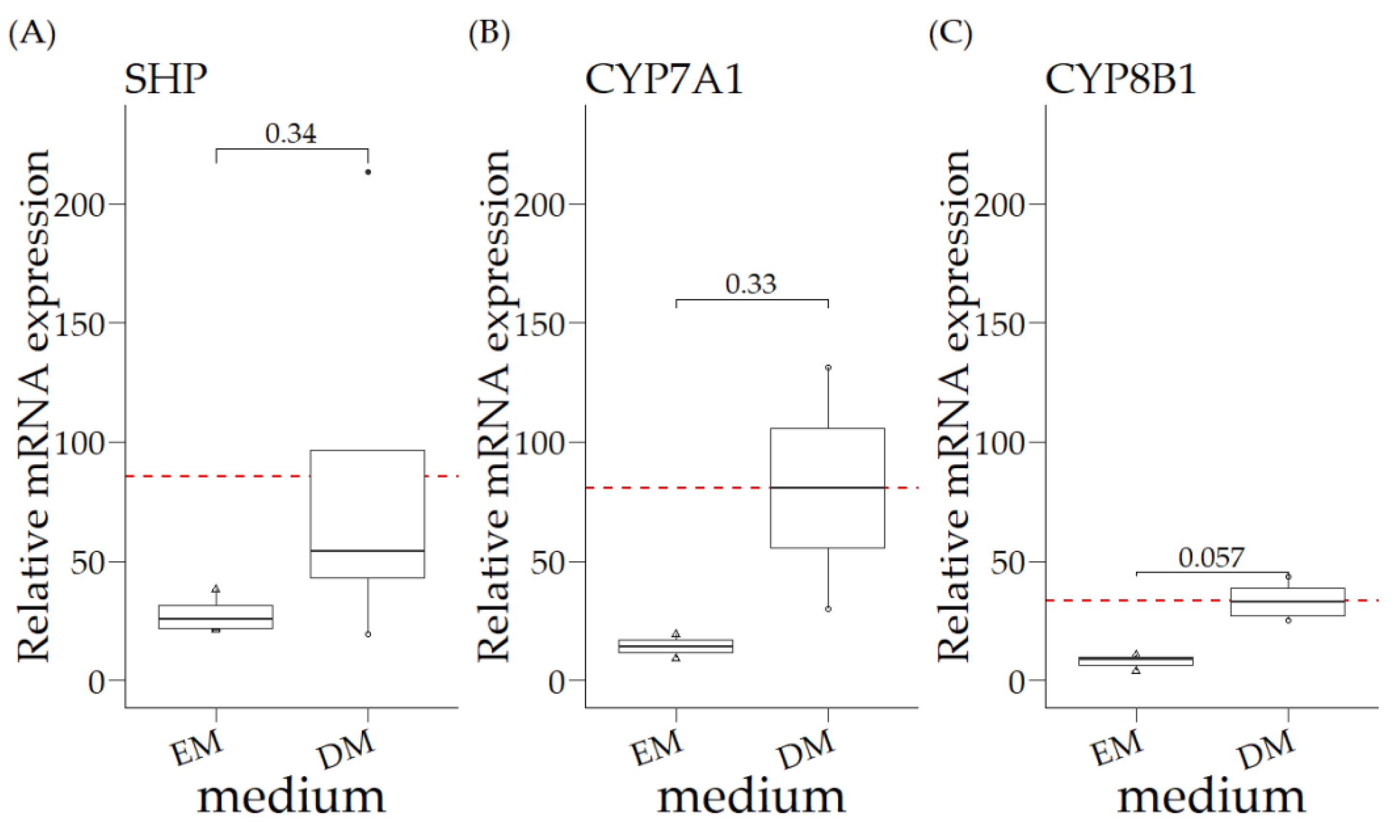
3.2. Relative mRNA Expression of COMMD1 +/+ Organoids in Expansion Medium (EM) and Differentiation Medium (DM)
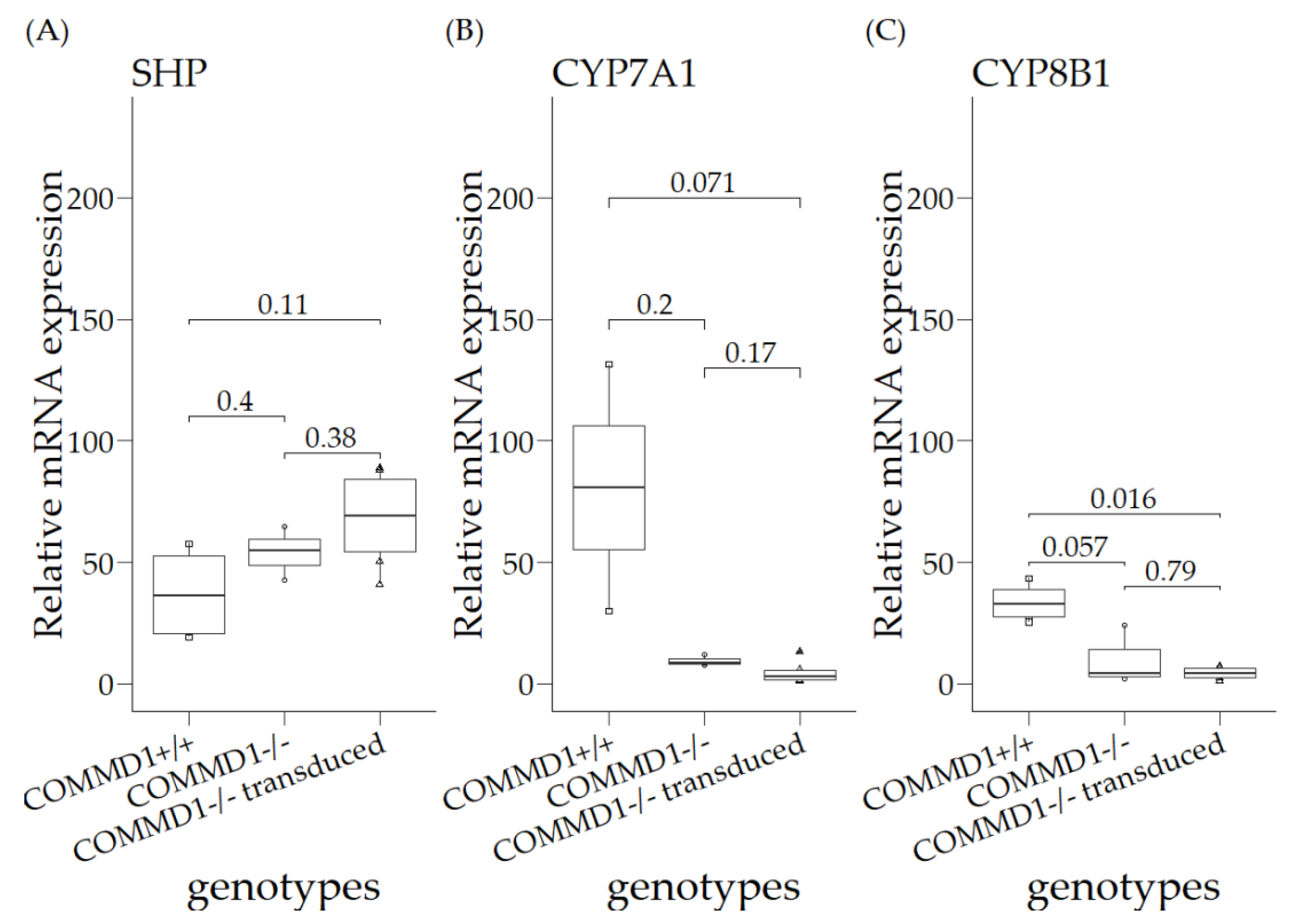
3.3. Relative mRNA Expression Levels in Organoids of COMMD1+/+, COMMD1−/−, and COMMD1−/−Transduced
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| WD | Wilson’s disease |
| ATP7B | ATPase copper transporting beta |
| NR | nuclear receptors |
| FXR | Farnesoid X nuclear receptor |
| LXR | liver X receptor |
| EM | Expansion medium |
| DM | differentiation medium |
| SHP | small heterodimer partner |
| ApoE | Apolipoprotein E |
| CYP7A1 | cholesterol 7 alpha-hydroxylase |
| CYP8B1 | sterol 12 alpha-hydroxylase |
| HPRT | hypoxanthine phosphoribosyltransferase |
| HMBS | hydroxymethylbilane synthase |
| SRPR | signal receptor particle receptor |
| BSEP | Bile Salt Export Pump |
| IBABP | Ileal Bile Acid Binding protein |
| RPS5 | ribosomal protein S5 |
References
- Weiss, K.H.; Stremmel, W. Clinical considerations for an effective medical therapy in Wilson’s disease. Ann. N. Y. Acad. Sci. 2014, 1315, 81–85. [Google Scholar] [CrossRef] [PubMed]
- Tanzi, R.E.; Petrukhin, K.; Chernov, I.; Pellequer, J.L.; Wasco, W.; Ross, B.; Romano, D.M.; Parano, E.; Pavone, L.; Brzustowicz, L.M. The Wilson disease gene is a copper transporting ATPase with homology to the Menkes disease gene. Nat. Genet. 1993, 5, 344–350. [Google Scholar] [CrossRef] [PubMed]
- Roth, E.; Manhart, N.; Wessner, B. Assessing the antioxidative status in critically ill patients. Curr. Opin. Clin. Nutr. Metab. Care 2004, 7, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Webb, C.; Twedt, D. Oxidative stress and liver disease. Vet. Clin. N. Am. Small Anim. Pract. 2008, 38, 125–135. [Google Scholar] [CrossRef] [PubMed]
- Fahrni, C.J. Synthetic fluorescent probes for monovalent copper. Curr. Opin. Chem. Biol. 2013, 17, 656–662. [Google Scholar] [CrossRef] [PubMed]
- Pham, A.N.; Xing, G.; Miller, C.J.; Waite, T.D. Fenton-like copper redox chemistry revisited: Hydrogen peroxide and superoxide mediation of copper-catalyzed oxidant production. J. Catal. 2013, 301, 54. [Google Scholar] [CrossRef]
- Wooton-Kee, C.R.; Jain, A.K.; Wagner, M.; Grusak, M.A.; Finegold, M.J.; Lutsenko, S.; Moore, D.D. Elevated copper impairs hepatic nuclear receptor function in Wilson’s disease. J. Clin. Investig. 2015, 125, 3449–3460. [Google Scholar] [CrossRef]
- Hamilton, J.P.; Koganti, L.; Muchenditsi, A.; Pendyala, V.S.; Huso, D.; Hankin, J.; Murphy, R.C.; Huster, D.; Merle, U.; Mangels, C.; et al. Activation of liver X receptor/retinoid X receptor pathway ameliorates liver disease in Atp7B(-/-) (Wilson disease) mice. Hepatology 2016, 63, 1828–1841. [Google Scholar] [CrossRef]
- van De Sluis, B.; Rothuizen, J.; Pearson, P.L.; van Oost, B.A.; Wijmenga, C. Identification of a new copper metabolism gene by positional cloning in a purebred dog population. Hum. Mol. Genet. 2002, 11, 165–173. [Google Scholar] [CrossRef]
- Fieten, H.; Gill, Y.; Martin, A.J.; Concilli, M.; Dirksen, K.; van Steenbeek, F.G.; Spee, B.; van den Ingh, T.S.; Martens, E.C.; Festa, P.; et al. The Menkes and Wilson disease genes counteract in copper toxicosis in Labrador retrievers: A new canine model for copper-metabolism disorders. Dis. Model Mech. 2016, 9, 25–38. [Google Scholar] [CrossRef]
- Wu, X.; Mandigers, P.J.J.; Watson, A.L.; van den Ingh, T.S.G.A.M.; Leegwater, P.A.J.; Fieten, H. Association of the canine ATP7A and ATP7B with hepatic copper accumulation in Dobermann dogs. J. Vet. Intern. Med. 2019, 33, 1646–1652. [Google Scholar] [CrossRef] [PubMed]
- Favier, R.P.; Spee, B.; Schotanus, B.A.; van den Ingh, T.S.; Fieten, H.; Brinkhof, B.; Viebahn, C.S.; Penning, L.C.; Rothuizen, J. COMMD1-deficient dogs accumulate copper in hepatocytes and provide a good model for chronic hepatitis and fibrosis. PLoS ONE 2012, 7, e42158. [Google Scholar] [CrossRef]
- Favier, R.P.; Spee, B.; Fieten, H.; van den Ingh, T.S.; Schotanus, B.A.; Brinkhof, B.; Rothuizen, J.; Penning, L.C. Aberrant expression of copper associated genes after copper accumulation in COMMD1-deficient dogs. J. Trace Elem. Med. Biol. 2015, 29, 347–353. [Google Scholar] [CrossRef] [PubMed]
- van Steenbeek, F.G.; Spee, B.; Penning, L.C.; Kummeling, A.; van Gils, I.H.; Grinwis, G.C.; Van Leenen, D.; Holstege, F.C.; Vos-Loohuis, M.; Rothuizen, J.; et al. Altered subcellular localization of heat shock protein 90 is associated with impaired expression of the aryl hydrocarbon receptor pathway in dogs. PLoS ONE 2013, 8, e57973. [Google Scholar] [CrossRef] [PubMed]
- Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 2003, 31, 3406–3415. [Google Scholar] [CrossRef] [PubMed]
- Spee, B.; Arends, B.; van den Ingh, T.S.; Penning, L.C.; Rothuizen, J. Copper metabolism and oxidative stress in chronic inflammatory and cholestatic liver diseases in dogs. J. Vet. Intern. Med. 2006, 20, 1085–1092. [Google Scholar] [CrossRef] [PubMed]
- Brinkhof, B.; Spee, B.; Rothuizen, J.; Penning, L.C. Development and evaluation of canine reference genes for accurate quantification of gene expression. Anal. Biochem. 2006, 356, 36–43. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, research0034-1. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Beaulieu, J.F.; Huggett, J.; Jaggi, R.; Kibenge, F.S.; Olsvik, P.A.; Penning, L.C.; Toegel, S. MIQE precis: Practical implementation of minimum standard guidelines for fluorescence-based quantitative real-time PCR experiments. BMC Mol. Biol. 2010, 11, 74. [Google Scholar] [CrossRef] [PubMed]
- Nantasanti, S.; Spee, B.; Kruitwagen, H.S.; Chen, C.; Geijsen, N.; Oosterhoff, L.A.; van Wolferen, M.E.; Pelaez, N.; Fieten, H.; Wubbolts, R.W.; et al. Disease Modeling and Gene Therapy of Copper Storage Disease in Canine Hepatic Organoids. Stem Cell Rep. 2015, 5, 895–907. [Google Scholar] [CrossRef] [PubMed]
- RStudio Team. RStudio: Integrated Development for R; RStudio, Inc.: Boston, MA, USA, 2015. [Google Scholar]
- Bijsmans, I.T.; Milona, A.; Ijssennagger, N.; Willemsen, E.C.; Ramos Pittol, J.M.; Jonker, J.W.; Lange, K.; Hooiveld, G.J.; van Mil, S.W. Characterization of stem cell-derived liver and intestinal organoids as a model system to study nuclear receptor biology. Biochim. Biophys. Acta 2017, 1863, 687–700. [Google Scholar] [CrossRef] [PubMed]
- Forman, B.M.; Goode, E.; Chen, J.; Oro, A.E.; Bradley, D.J.; Perlmann, T.; Noonan, D.J.; Burka, L.T.; McMorris, T.; Lamph, W.W.; et al. Identification of a nuclear receptor that is activated by farnesol metabolites. Cell 1995, 81, 687–693. [Google Scholar] [CrossRef]
- Wagner, M.; Zollner, G.; Trauner, M. Nuclear receptors in liver disease. Hepatology 2011, 53, 1023–1034. [Google Scholar] [CrossRef] [PubMed]
- Fieten, H.; Hooijer-Nouwens, B.D.; Biourge, V.C.; Leegwater, P.A.; Watson, A.L.; van den Ingh, T.S.; Rothuizen, J. Association of dietary copper and zinc levels with hepatic copper and zinc concentration in Labrador Retrievers. J. Vet. Intern. Med. 2012, 26, 1274–1280. [Google Scholar] [CrossRef] [PubMed]
- Fieten, H.; Penning, L.C.; Leegwater, P.A.; Rothuizen, J. New canine models of copper toxicosis: Diagnosis, treatment, and genetics. Ann. N. Y. Acad. Sci. 2014, 1314, 42–48. [Google Scholar] [CrossRef] [PubMed]
| Genes | Sequences (5’ to 3’) | Annealing Temperature | Amplicon Size in bp |
|---|---|---|---|
| SHP | FW: AACATTCTCCCGTTTGACCAC | 62 °C | 99 |
| RV: GTAGTTGGCGTTGATGTAGTCG | |||
| APOE | FW: CGCTTCTGGGATTACCTG | 58 °C | 124 |
| RV: CCTTCACCTCCTTCATGG | |||
| CYP7A1 | FW: TTTCAAATGATTAGGAGCCCTG | 66 °C | 115 |
| RV: TGATTCAGACAAATAGGACTGC | |||
| CYP8B1 | FW: CCAAGCATGGAGATGTGTTCAC | 66 °C | 162 |
| RV: CCCGAGTGGTATCCAAATACC | |||
| HPRT | FW: AGCTTGCTGGTGAAAAGGAC | 58 °C | 104 |
| RV: TTATAGTCAAGGGCATATCC | |||
| HMBS | FW: TCACCATCGGAGCCATCT | 61 °C | 112 |
| RV: GTTCCCACCACGCTCTTCT | |||
| SRPR | FW: GCTTCAGGATCTGGACTGC | 61 °C | 81 |
| RV: GTTCCCTTGGTAGCACTGG | |||
| RPS5 | FW: TCACTGGTGAGAACCCCCT | 63 °C | 141 |
| RV: CCTGATTCACACGGCGTAG |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, X.; Chien, H.; van Wolferen, M.E.; Kruitwagen, H.S.; Oosterhoff, L.A.; Penning, L.C. Reduced FXR Target Gene Expression in Copper-Laden Livers of COMMD1-Deficient Dogs. Vet. Sci. 2019, 6, 78. https://doi.org/10.3390/vetsci6040078
Wu X, Chien H, van Wolferen ME, Kruitwagen HS, Oosterhoff LA, Penning LC. Reduced FXR Target Gene Expression in Copper-Laden Livers of COMMD1-Deficient Dogs. Veterinary Sciences. 2019; 6(4):78. https://doi.org/10.3390/vetsci6040078
Chicago/Turabian StyleWu, Xiaoyan, Hsiaotzu Chien, Monique E. van Wolferen, Hedwig S. Kruitwagen, Loes A. Oosterhoff, and Louis C. Penning. 2019. "Reduced FXR Target Gene Expression in Copper-Laden Livers of COMMD1-Deficient Dogs" Veterinary Sciences 6, no. 4: 78. https://doi.org/10.3390/vetsci6040078
APA StyleWu, X., Chien, H., van Wolferen, M. E., Kruitwagen, H. S., Oosterhoff, L. A., & Penning, L. C. (2019). Reduced FXR Target Gene Expression in Copper-Laden Livers of COMMD1-Deficient Dogs. Veterinary Sciences, 6(4), 78. https://doi.org/10.3390/vetsci6040078





