Development and Evaluation of a Novel Taqman Real-Time PCR Assay for Rapid Detection of Mycoplasma bovis: Comparison of Assay Performance with a Conventional PCR Assay and Another Taqman Real-Time PCR Assay †
Abstract
:1. Introduction
2. Experimental Section
2.1. Clinical Specimens
2.2. DNA Extraction
2.3. TaqMan Real-Time PCR Assay
| Gene Target | Oligo | Sequence (5'→3') | Reference | Product Size |
|---|---|---|---|---|
| uvrC | Forward Primer | GAG AAT GCT TCA GTA TTT TGA CGG | This study | 170 bp |
| Reverse Primer | CAA AAG CAA AAT GTT AAA TTC AGG | This study | ||
| Probe | (6 FAM) CAT ATA TAA GTG AGA CTA ACT TAT T(MGB) | [18] | ||
| uvrC | F2024 | TCT AAT TTT TTC ATC ATC GCT AAT GC | [12] | 112 bp |
| R2135 | TCA GGC CTT TGC TAC AAT GAA C | |||
| M.bov | (FAM) AAC TGC ATC ATA TCA CAT ACT (MGB) | |||
| 16S rRNA | Forward Primer | CCT TTT AGA TTG GGA TAG CGG ATG | [19] | 360 bp |
| Reverse Primer | CCG TCA AGG TAG CAT CAT TTC CTA T |
2.4. Culture
2.5. Sensitivity and Specificity Analyses
| Number | Species | Strain | Real-Time PCR |
|---|---|---|---|
| 1 | Mycoplasma bovis | ATCC 25523 | Positive |
| 2 | Mycoplasma bovis | ATCC 27369 | Positive |
| 3 | Mycoplasma bovis | Field isolate | Positive |
| 4 | Mycoplasma alkalescens | ATCC 29103 | Negative |
| 5 | Mycoplasma canadense | ATCC 29418 | Negative |
| 6 | Mycoplasma arginini | ATCC 23838 | Negative |
| 7 | Mycoplasma bovigenitalium | ATCC 27748 | Negative |
| 8 | Staphylococcus aureus | Field isolate | Negative |
| 9 | Streptococcus agalactiae | Field isolate | Negative |
| 10 | Escherichia coli | Field isolate | Negative |
| 11 | Corynebacterium pseudotuberculosis | Field isolate | Negative |
| 12 | Streptococcus uberis | Field isolate | Negative |
| 13 | Coagulase Negative Staphylococcus | Field isolate | Negative |
| 14 | Pseudomonas aeruginosa | Field isolate | Negative |
| 15 | Klebsiella pneumonia | Field isolate | Negative |
| 16 | Proteus mirabilis | Field isolate | Negative |
| 17 | Serratia marcescens | Field isolate | Negative |
| 18 | Entereococcus faecalis | Field isolate | Negative |
| 19 | Arcanobacterium pyogenes | Field isolate | Negative |
| 20 | Candida spp | Field isolate | Negative |
| 21 | Histophilus somnus | Field isolate | Negative |
| 22 | Salmonella spp | Field isolate | Negative |
| 23 | Bacillus cereus | Field isolate | Negative |
| 24 | Pasteurella multocida | Field isolate | Negative |
| 25 | Mannheimia haemolytica | Field isolate | Negative |
3. Results and Discussion
3.1. Limit of Detection
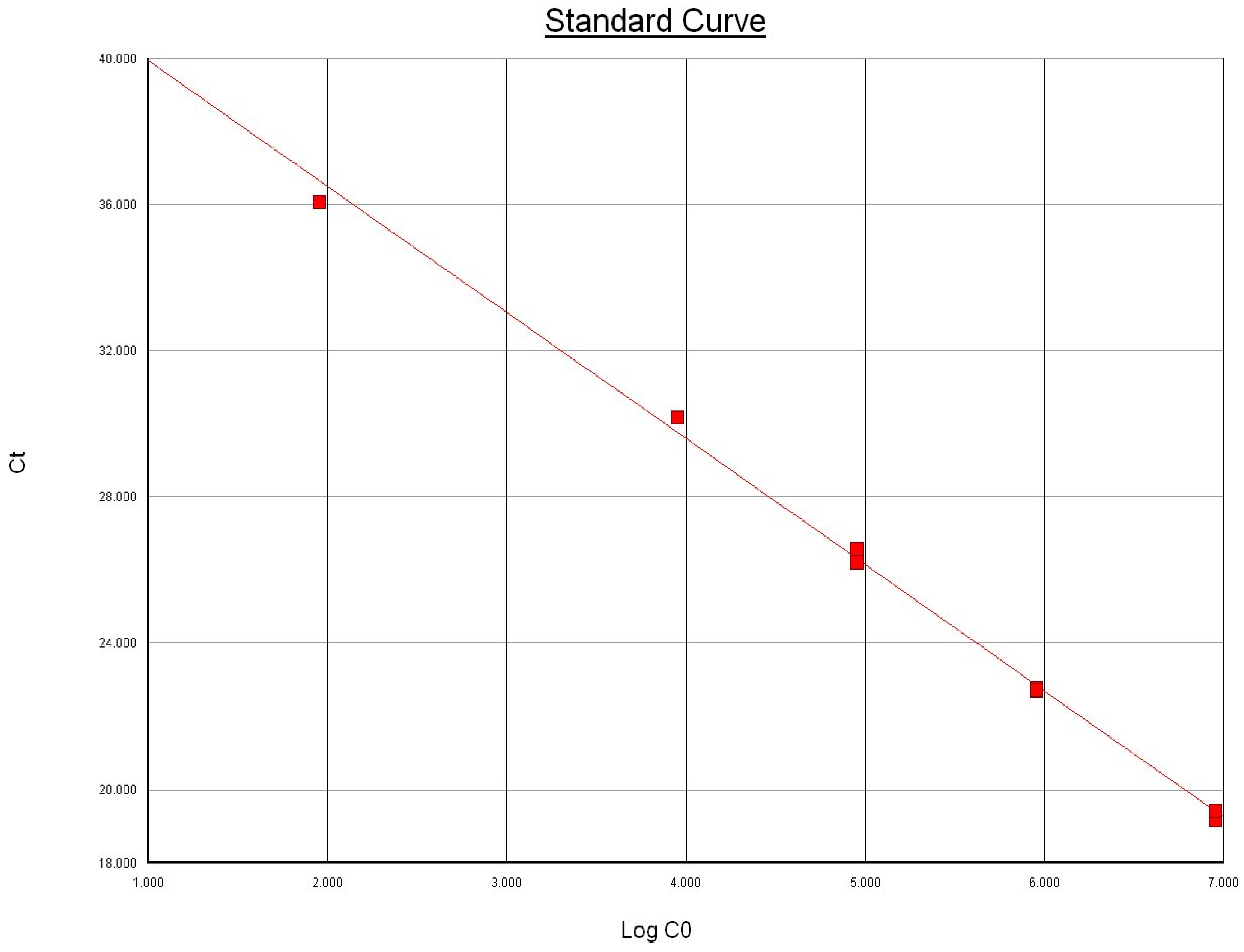
3.2. Assay Specificity
3.3. Assay Performance with Clinical Specimens
| Specimen | Our Taqman PCR Assay | Conventional PCR | Clothier et al. Taqman PCR Assay | |||
|---|---|---|---|---|---|---|
| Positive | Negative | Positive | Negative | Positive | Negative | |
| Lung (n = 45) | 42 | 3 | 42 | 3 | 42 | 3 |
| Milk (n = 78) | 15 | 63 | 15 | 63 | 15 | 63 |
| Nasal swab (n = 78) | 22 | 56 | 22 | 56 | 22 | 56 |
| Joint fluid (n = 5) | 2 | 3 | 2 | 3 | 2 | 3 |
| Semen (n = 8) | 1 | 7 | 1 | 7 | 1 | 7 |
| Total (n = 214) | 82 | 132 | 82 | 132 | 82 | 132 |
3.4. Performance Comparison of Culture and Taqman PCR Assay
| Specimen | Culture | Our Taqman PCR Assay | ||
|---|---|---|---|---|
| Positive | Negative | Positive | Negative | |
| Lung (n = 40) | 17 | 23 | 39 | 1 |
| Milk(n = 18) | 4 | 14 | 4 | 14 |
| Swabs(n = 10) | 4 | 6 | 6 | 4 |
| Joint Fluid (n = 3) | 0 | 3 | 1 | 2 |
| Total (n = 71) | 25 | 46 | 50 | 21 |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nicholas, R.A.; Ayling, R.D. Mycoplasma bovis: Disease, diagnosis and control. Res. Vet. Sci. 2003, 74, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Goll, F., Jr. Identification of mycoplasmas isolated from domestic animals. In Mycoplasmosis in Animals: Laboratory Diagnosis; Whitford, H.W., Rosenbusch, R.F., Lauerman, L.H., Eds.; Iowa State University Press: Ames, IA, USA, 1994; pp. 15–27. [Google Scholar]
- Rosenbusch, R.F. Biology and Taxonomy of Mycoplasmas. In Mycoplasmosis in Animals: Laboratory Diagnosis; Whitford, H.W., Rosenbusch, R.F., Lauerman, L.H., Eds.; Iowa State University Press: Ames, IA, USA, 1994; pp. 3–11. [Google Scholar]
- Pfutzner, H.; Sachse, K. Mycoplasma bovis as an agent of mastitis, pneumonia, arthritis and genital disorders. Revue Sci. Tech. 1994, 15, 1477–1494. [Google Scholar]
- Nicholas, R.A.J.; Baker, S.E. Recovery of Mycoplasmas from Animals. In Mycoplasma Protocols; Miles, R.J., Nicholas, R.A.J., Eds.; Humana Press: Totowa, NJ, USA, 1998; pp. 37–44. [Google Scholar]
- Jasper, D.E. Bovine Mycoplasma mastitis. Adv. Vet. Sci. Comp. Med. 1981, 25, 121–159. [Google Scholar] [PubMed]
- Caswell, J.L.; Archambault, M. Mycoplasma bovis pneumonia in cattle. Anim. Health Res. Rev. 2007, 8, 161–186. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Verma, A.K.; Rahal, A. Mycoplasma bovis, A Multi Disease Producing Pathogen: An Overview. Asian J. Anim. Vet. Adv. 2011, 6, 537–546. [Google Scholar] [CrossRef]
- Wise, K.S.; Calcutt, M.J.; Foecking, M.F.; Röske, K.; Madupu, R.; Methé, B.A. Complete genome sequence of Mycoplasma bovis type strain PG45 (ATCC 25523). Infect. Immun. 2011, 79, 982–983. [Google Scholar] [CrossRef] [PubMed]
- Subramaniam, S.; Bergonier, D.; Poumarat, F.; Capaul, S.; Schlatter, Y.; Nicolet, J.; Frey, J. Species identification of Mycoplasma bovis and Mycoplasma agalactiae based on the uvrC genes by PCR. Mol. Cell. Probes 1998, 12, 161–169. [Google Scholar] [CrossRef] [PubMed]
- Bashiruddin, J.B.; Frey, J.; Königsson, M.H.; Johansson, K.E.; Hotzel, H.; Diller, R.; deSantis, P.; Botelho, A.; Ayling, R.D.; Nicholas, R.A.; et al. Evaluation of PCR systems for the identification and differentiation of Mycoplasma agalactiae and Mycoplasma bovis: A collaborative trial. Vet. J. 2005, 169, 268–275. [Google Scholar]
- Clothier, K.A.; Jordan, D.M.; Thompson, C.J.; Kinyon, J.M.; Frana, T.S.; Strait, E.L. Mycoplasma bovis real-time polymerase chain reaction assay validation and diagnostic performance. J. Vet. Diagn. Invest. 2010, 22, 956–960. [Google Scholar] [CrossRef] [PubMed]
- Cai, H.Y.; Bell-Rogers, P.; Parker, L.; Prescott, J.F. Development of a real-time PCR for detection of Mycoplasma bovis in bovine milk and lung samples. J. Vet. Diagn. Invest. 2005, 17, 537–545. [Google Scholar] [CrossRef] [PubMed]
- McAuliffe, L.; Ellis, R.J.; Lawes, J.R.; Ayling, R.D.; Nicholas, R.A. 16S rDNA PCR and denaturing gradient gel electrophoresis; a single generic test for detecting and differentiating Mycoplasma species. J. Med. Microbiol. 2005, 54, 731–739. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.; Shi, L.; Hu, C.; Chen, X.; Qi, J.; Ba, X.; Peng, Q.; Chen, Y.; Chen, H.; Guo, A. Development of a loop-mediated isothermal amplification assay for sensitive and rapid detection of Mycoplasma bovis. Afr. J. Biotechnol. 2011, 10, 12333–12338. [Google Scholar]
- Thomas, A.; Dizier, I.; Linden, A.; Mainil, J.; Frey, J.; Vilei, E.M. Conservation of the uvrC gene sequence in Mycoplasma bovis and its use in routine PCR diagnosis. Vet. J. 2004, 168, 100–102. [Google Scholar] [CrossRef] [PubMed]
- Marenda, M.S.; Sagne, E.; Poumarat, F.; Citti, C. Suppression subtractive hybridization as a basis to assess Mycoplasma agalactiae and Mycoplasma bovis genomic diversity and species-specific sequences. Microbiology 2005, 151, 475–489. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.H.; Lee, J.W.; Wang, S.W.; Liu, L.Y.; Lee, M.F.; Chuang, S.T.; Shy, Y.M.; Chang, C.L.; Wu, M.C.; Chi, C.H. Development of a novel biochip for rapid multiplex detection of seven mastitis causing pathogens in bovine milk samples. J. Vet. Diagn. Investig. 2008, 20, 463–471. [Google Scholar] [CrossRef]
- Gonzalez, Y.R.C.; Baccunana, C.R.; Bolske, G.; Mattsson, J.G.; Molina, C.F.; Johansson, K.E. In vitro amplification of the 16S rRNA genes from Mycoplasma bovis and Mycoplasma agalactiae by PCR. Vet. Microbiol. 1995, 47, 183–190. [Google Scholar] [CrossRef] [PubMed]
- Moser, M.J.; Christensen, D.R.; Norwood, D.; Prudent, J.R. Multiplexed detection of anthrax-related toxin genes. J. Mol. Diagn. 2006, 8, 89–96. [Google Scholar] [CrossRef] [PubMed]
- González, R.N.; Sears, P.M.; Wilson, D.J. Diagnosis of Intramammary Infections Due to Mycoplasma bovis in Dairy Cattle. In Proceedings of the 3rd IDF International Mastitis Seminar, 1995; Volume S-2, pp. 23–27.
- Bell, C.J.; Blackurn, P.; Elliott, M.; Patterson, T.I.; Ellison, S.; Lahuerta-Marin, A.; Ball, H.J. Investigation of polymerase chain reaction assays to improve detection of bacterial involvement in bovine respiratory disease. J. Vet. Diagn. Investig. 2014, 26, 631–634. [Google Scholar] [CrossRef]
- Castillo-Alcala, F.; Bateman, K.G.; Cai, H.Y.; Schott, C.R.; Parker, L.; Clark, M.E.; McRaild, P.; McDowall, R.M.; Foster, R.A.; Archambault, M.; et al. Prevalence and genotype of Mycoplasma bovis in beef cattle after arrival at a feedlot. Am. J. Vet. Res. 2012, 73, 1932–1943. [Google Scholar]
- Justice-Allen, A.; Trujillo, J.; Goodell, G.; Wilson, D. Detection of multiple Mycoplasma species in bulk tank milk samples using real-time PCR and conventional culture and comparison of test sensitivities. J. Dairy Sci. 2011, 94, 3411–3419. [Google Scholar] [CrossRef] [PubMed]
- Hoorfar, J.; Malorny, B.; Abdulmawjood, A.; Cook, N.; Wagner, M.; Fach, P. Practical considerations in design of internal amplification controls for diagnostic PCR assays. J. Clin. Microbiol. 2004, 42, 1863–1868. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naikare, H.; Bruno, D.; Mahapatra, D.; Reinisch, A.; Raleigh, R.; Sprowls, R. Development and Evaluation of a Novel Taqman Real-Time PCR Assay for Rapid Detection of Mycoplasma bovis: Comparison of Assay Performance with a Conventional PCR Assay and Another Taqman Real-Time PCR Assay. Vet. Sci. 2015, 2, 32-42. https://doi.org/10.3390/vetsci2010032
Naikare H, Bruno D, Mahapatra D, Reinisch A, Raleigh R, Sprowls R. Development and Evaluation of a Novel Taqman Real-Time PCR Assay for Rapid Detection of Mycoplasma bovis: Comparison of Assay Performance with a Conventional PCR Assay and Another Taqman Real-Time PCR Assay. Veterinary Sciences. 2015; 2(1):32-42. https://doi.org/10.3390/vetsci2010032
Chicago/Turabian StyleNaikare, Hemant, Daniela Bruno, Debabrata Mahapatra, Alesia Reinisch, Russell Raleigh, and Robert Sprowls. 2015. "Development and Evaluation of a Novel Taqman Real-Time PCR Assay for Rapid Detection of Mycoplasma bovis: Comparison of Assay Performance with a Conventional PCR Assay and Another Taqman Real-Time PCR Assay" Veterinary Sciences 2, no. 1: 32-42. https://doi.org/10.3390/vetsci2010032
APA StyleNaikare, H., Bruno, D., Mahapatra, D., Reinisch, A., Raleigh, R., & Sprowls, R. (2015). Development and Evaluation of a Novel Taqman Real-Time PCR Assay for Rapid Detection of Mycoplasma bovis: Comparison of Assay Performance with a Conventional PCR Assay and Another Taqman Real-Time PCR Assay. Veterinary Sciences, 2(1), 32-42. https://doi.org/10.3390/vetsci2010032





