Abstract
Polymorphic yeast, Candida albicans, forms thick-walled structures called chlamydospores in order to survive under adverse conditions. We present proteomic profile changes occurring during chlamydospore formation. Chlamydospores were induced by inoculating C. albicans cells (grown for 48 h) on rice extract and semisolid agar containing tween 80 (1%), and were overlaid by a polyethene sheet to induce microaerophilic conditions at 30 °C. Proteins extracted from chlamydospores and hyphae (producing chlamydospores) were identified by LC-MS/MS analysis. Present datasets include proteomic data (Swath spectral libraries) of chlamydospores and yeast phase cells, as well as methodologies and tools used for the data generation. Further analysis is expected to provide an opportunity to understand modulations in metabolic processes, molecular architecture (i.e., cell wall, membrane, and cytoskeleton) and stress response pathways leading to chlamydospore formation and thus facilitating survival of C. albicans under adverse conditions.
Data Set: http://www.peptideatlas.org/PASS/PASS01061 (Username: PASS01061, Password: PF8546i)
Data Set License: There is no specific license.
1. Introduction
Morphophysiological plasticity enables Candida albicans to survive under a variety of extreme microenvironments [1,2,3,4,5,6,7,8,9,10,11]. It exists in the form of yeast, hyphae, pseudohyphae, chlamydospore, white/opaque cells, according to the microenvironment [1,2,3,4,5,6,7,8,9,10,11]. Cells in different morphological forms respond differently towards host defense mechanisms, antifungal agents, stress conditions [8]. It makes C. albicans one of the most successful opportunistic pathogens of humans [1,2]. Among these morphological forms, chlamydospore is one of the most interesting structures, and develops under unfavorable conditions such as oxygen limitation and embedded growth in matrix (Figure 1a) [5,6,7]. The significance and regulation of chlamydospore formation are not yet fully understood, though Böttcher et al. (2016) provides a detailed account of signaling pathways regulating chlamydospore formation in response to different environmental and nutritional factors [1,3,4]. Chlamydospores are a relatively unusual morphological form that facilitates survival of C. albicans under extreme micro environments by significantly lowering metabolism; however, they can germinate once conditions become favorable [3,4,6,8,9]. The thick walls could provide protection against the adverse microenvironment; however, not much study is available on the structure and composition of chlamydospore cell walls [9,10,11,12,13,14]. In the present study, a proteomic profile of chlamydospores of C. albicans generated using LC-MS/MS provides insights into the regulation of chlamydospore formation and survival strategies.
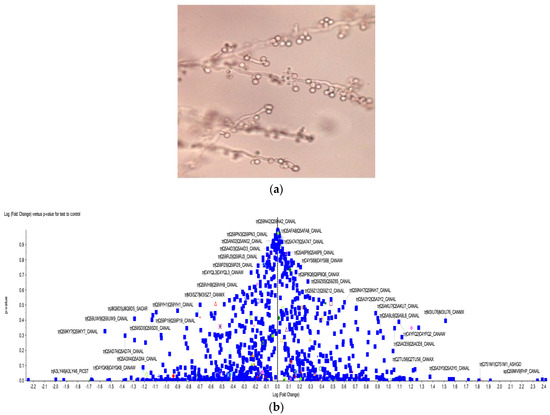
Figure 1.
(a) Light microscopy images of chlamydospores with hyphae (chlamydospore-producing) formation of C. albicans (ATCC 10231); (b) Scatter plot of differentially expressed proteins during Chlamydospore formation.
2. Results
Proteomic data of chlamydospore (test) and yeast phase cells (control) of C. albicans were generated through processed peptides using Micro LC-Triple TOF 5600 SWATH MS analysis. Specifically, SWATH MS mode was utilized for optimization of quadrupole settings in the instrument [15,16,17,18,19,20]. Samples were spiked from each treatment (chlamydospores and yeast phase cells) to produce the SWATH spectral library, and were analyzed via LC-MS/MS by information dependent acquisition (IDA) (Figure 1b, Supplementary Tables 1–17). Data was uploaded to PeptideAtlas and is available at the Official URL for this dataset: http://www.peptideatlas.org/PASS/PASS01061. Functional annotation (using David software, Candida genome database, Saccharomyces genome database, Kyoto Encyclopedia of Genes and Genomes and UniProt databases) revealed significant modulations in metabolism, cellular architecture, gene expression and stress response pathways required for survival of Candida albicans under adverse microenvironments [11,12].
3. Discussion
Candida albicans is one of the most important opportunistic pathogens in humans, worldwide [21]. Morphological plasticity (i.e., yeast, hyphae, pseudohyphae, chlamydospore, white and opaque cells) enables C. albicans to survive and grow in extreme niches by evading host immune responses [21,22]. Among these morphological forms, chlamydospores (dormant spores), though rarely observed in tissues [23], exhibit different responses towards the host immune system, and germinate and form hyphae once conditions becomes favorable through multiple, interlinked signaling pathways [1,2,3,8]. Our proteomic data will provide an opportunity to understand the regulation of morphogenesis (i.e., chlamydospores formation) and survival under adverse environmental conditions in addition to modulation of molecular architecture (i.e., cell wall, membrane, and cytoskeleton of chlamydospore).
4. Materials and Methods
4.1. Candida albicans Strain and Growth Conditions
Candida albicans (ATCC 10231), a quality control strain, was obtained from the Microbial Type Culture Collection (MTCC), Institute of Microbial Technology (IMTECH), Chandigarh, India and maintained on yeast extract peptone dextrose (YPD) agar slants at 4 °C. Yeast Extract Peptone Dextrose (YEPD) Broth and Rice extract agar were purchased from Hi-media Laboratories, Pvt. Ltd. (Mumbai, India). All the fine chemicals used in this study were purchased from Sigma-Aldrich Pvt. Ltd (Bangalore, India), Bangalore, and solvents used were purchased from Qualigens and SD Fine Chemicals Ltd. (Mumbai, India).
4.2. Test: Induction of Chlamydospore Formation
Candida albicans cells (grown for 48 h in YEPD broth at 28 °C) were grown on a nutrient-limiting rice extract agar (rice extract 1.3%) containing 1% tween 80 (for exerting surface stress). The inoculum was overlaid with circles of sterile polyethylene transparent sheets for microaerophilic conditions under embedded growth necessary for chlamydospore production. Plates were further incubated at 30 °C for 14 days [13,14].
4.3. Control Experiment
Candida albicans cells were grown on nutrient-limiting rice extract agar (rice extract 1.3%) plates containing 1% tween 80 (without microaerophilic conditions and embedded growth) at 30 °C for 14 days. Surface-grown cells were used as control.
4.4. Extraction of Proteins
Chlamydospores and hyphae (chlamydospore producing) from test and yeast phase cells (from control) were harvested from twelve-day old plates. Chlamydospores were lysed and proteins were extracted as per the protocol optimized by Haar (2007), and concentration was estimated by using the Bradford method [15,16].
4.5. Sample Preparation for LC-MS
Extracted proteins were subjected to trypsin digestion and peptides generated were washed using Zip tip C18 chromatography columns (Millipore; Billerica, MA, USA) and concentrated.
4.6. Liquid Chromatography and Mass Spectrometry
LC-MS/MS was carried out using a Triple-TOF 5600 (AB Sciex; Vaughan, ON, Canada) mass spectrometer coupled with a Micro LC 200 (Eksigent; Dublin, CA, USA) in high-sensitivity mode. Samples were spiked from each treatment to produce the SWATH spectral library and analyzed via LC-MS/MS by information-dependent acquisition (IDA) in duplicate [17,18,19,20]. In brief, a digested protein sample (4 µg) was injected into a Eskigent C18-RP HPLC column (100 × 0.3 mm, 3 µm, 120 Å) and then separated using a 95-min gradient of 3 to 35% ACN (acetonitrile) for IDA run and (1 µg) for SWATH run [17,18,19,20].
4.7. SWATH MS Analysis
Datasets were acquired (in triplicate) on Micro LC-Triple TOF 5600 SWATH MS. The instrument was specifically tuned in SWATH-MS mode to optimize the quadrupole settings for the selection of precursor ion selection windows 25 m/z wide. Using an isolation width of 25 m/z (containing 1 m/z for the window overlap), a set of 34 overlapping windows was constructed covering the precursor mass range of 400–1250 m/z. From 100 to 2000 m/z SWATH MS/MS spectra were collected. According to the calculation for a charge 2+ ions, for each window, the collision energy was optimized, centered upon the window with a spread of 15 eV. For all fragment-ion scans in high-sensitivity mode accumulation time (dwell time) of 100 ms was used. For 100 ms each SWATH-MS cycle a survey scan in high-resolution mode was also acquired resulting in a duty cycle of 3.4 s [17,18,19,20]. A spectral library was generated from IDA run using Protein pilot software version 4 by analyzing data against yeast database. Paragon algorithm is used in Protein pilot software. Approximately 1000 proteins were identified, with 5% FDR. The generated spectral library was used for SWATH analysis with 20 ppm error. SWATH runs were acquired in triplicate. SWATH runs of control and test were uploaded in SWATH processing window with 5 min RT window, 50 ppm mass error and 99% confidence. Processed SWATH file was exported to marker view software [17,18,19,20].
Exported files were saved as ion, peptide and protein files format. Protein file format was used for further analysis in Marker view. Data was normalized by Total sum normalization. Processed SWATH runs were grouped as control and test and compared using t-test [17,18,19,20]. Protein expression data was generated, and proteins with p-value < 0.05 and fold change ≥2 fold were considered as differentially expressed proteins in test with respect to control [11,12]. Uniprot ID of all differentially expressed proteins were retrieved and functionally annotated using David software [11,12].
4.8. Statistical Analysis
Statistical analysis was carried out using t-test. Probability p value less than 0.05 were considered to be significant and proteins with a number of matching peptides (≥2) and fold change (≥2) were considered for further analysis (samples were acquired in triplicate).
5. Conclusions
Proteomic profile of chlamydospores of a polymorphic fungus C. albicans were generated using LC-MS/MS analysis. Approximately 1000 proteins identified were characterized using different databases and tools. The dataset presented in this paper indicates significant modulation in proteins associated with morpho-physiological (chlamydospore formation) transition and stress responses. This data will be useful in understanding underlying mechanisms that facilitate survival of eukaryotic cells under extreme micro-environments.
Supplementary Materials
The following are available online at https://figshare.com/s/2c6887e058245d0b612c. DOI: 10.6084/m9.figshare.5286097.
Acknowledgments
Authors are thankful to the Honorable Vice Chancellor, Pandit Vidyasagar for constant support and encouragement. Mahesh Kulkarni and B. Santhakumari is thanked for extending LC-MS/MS facility. SPPU is thanked for financial assistance to R.P. under Biotechnology Department Research and Developmental Grant.
Author Contributions
Gajanan Zore and Sujata Ingle designed experiment, Sujata Ingle performed microbiological work and fixed samples for protein preparation. Rajendra Patil prepared the protein extract, Santosh Kodgire and Asha Shiradhone performed LC-MS/MS analysis and Gajanan Zore and Sujata Ingle wrote the MS.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Lim, C.Y.; Rosli, R.; Seow, H.F.; Chong, P.P. Candida and invasive candidiasis: Back to basics. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Ernst, J.F. Transcriptional factors in Candida albicans-environmental control of morphogenesis. Microbiology 2000, 146, 1763–1774. [Google Scholar] [CrossRef] [PubMed]
- Böttcher, B.; Pöllath, C.; Staib, P.; Hube, B.; Brunke, S. Candida species Rewired Hyphae Developmental Programs for Chlamydospore Formation. Front. Microbiol. 2016, 7, 1697. [Google Scholar] [CrossRef] [PubMed]
- Citiulo, F.; Moran, G.P.; Coleman, D.C.; Sullivan, D.J. Purification and ger mination of Candida albicans and Candida dubliniensis chlamydospores cultured in liquid media. FEMS Yeast Res. 2009, 9, 1051–1060. [Google Scholar] [CrossRef] [PubMed]
- Sonneborn, A.; Tebarth, B.; Ernst, J.F. Control of white-opaque phenotypic switching in Candida albicans by the Efg1p morphogenetic regulator. Infect. Immun. 1999, 67, 4655–4660. [Google Scholar] [PubMed]
- Nobile, C.J.; Bruno, V.M.; Richard, M.L.; Davis, D.A.; Mitchell, A.P. Genetic control of chlamydospore formation in Candida albicans. Microbiology 2003, 149, 3629–3637. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Shin, W.S.; Lee, K.H.; Kim, K.; Park, J.Y.; Koh, C.M. Rapid differentiation of Candida albicans from other Candida species using its unique germ tube formation at 39 °C. Yeast 2002, 19, 957–962. [Google Scholar] [CrossRef] [PubMed]
- Sosinska, G.J. Adaptations in the Wall Proteome of the Clinical Fungus Candida albicans in Response to Infection-Related Environmental Conditions. Ph.D. Thesis, University of Amsterdam, Amsterdam, The Netherlands, 2012. [Google Scholar]
- Staib, P.; Morschhäuser, J. Chlamydospore formation in Candida albicans and Candida dubliniensis–An enigmatic developmental programme. Mycoses 2006, 50, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Jansons, V.K.; Nickerson, W.J. Chemical composition of chlamydospores of Candida albicans. J. Bacteriol. 1970, 104, 922–932. [Google Scholar] [PubMed]
- Ingle, S. Metabolic Plasticity of Candida albicans during Yeast to Chlamydospore Transition. Master’s Thesis, SRTM University, Nanded, India, 2015. [Google Scholar]
- Ingle, S.; Kodgire, S.; Kazi, R.; Patil, R.; Bayitigeri, S.; Kulkarni, M.; Zore, G. Proteomic analysis of Candida albicans cells grown under chlamydospore inducing condition. J. Proteom. 2017. under review. [Google Scholar]
- Taschdjian, C.L. Routine identification of Candida albicans: Current methods and a new medium. Mycologia 1957, 49, 332–338. [Google Scholar] [CrossRef]
- Miller, S.E.; Spurlock, B.O.; Michaels, G.E. Electron microscopy of young Candida albicans chlamydospores. J. Bacteriol. 1974, 119, 992–999. [Google Scholar] [PubMed]
- Von Der Haar, T. Optimized protein extraction for quantitative proteomics of yeasts. PLoS ONE 2007, 2, e1078. [Google Scholar] [CrossRef] [PubMed]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Gillet, L.C.; Navarro, P.; Tate, S.; Röst, H.; Selevsek, N.; Reiter, L.; Aebersold, R. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: A new concept for consistent and accurate proteome analysis. Mol. Cell. Proteom. 2012, 11, O111–016717. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.S.; Li, H.C.; Lai, Y.M.; Lo, H.F.; Chen, L.F.O. Proteomics and transcriptomics of broccoli subjected to exogenously supplied and transgenic senescence-induced cytokinin for amelioration of postharvest yellowing. J. Proteom. 2013, 93, 133–144. [Google Scholar] [CrossRef] [PubMed]
- Collins, B.C.; Gillet, L.C.; Rosenberger, G.; Röst, H.L.; Vichalkovski, A.; Gstaiger, M.; Aebersold, R. Quantifying protein interaction dynamics by SWATH mass spectrometry: Application to the 14-3-3 system. Nat. Methods 2013, 10, 1246–1253. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Qian, W.J.; Mottaz, H.M.; Gritsenko, M.A.; Norbeck, A.D.; Moore, R.J.; Smith, R.D. Evaluation of multiprotein immunoaffinity subtraction for plasma proteomics and candidate biomarker discovery using mass spectrometry. Mol. Cell. Proteom. 2006, 5, 2167–2174. [Google Scholar] [CrossRef] [PubMed]
- Biswas, S.; Van Dijck, P.; Datta, A. Environmental sensing and signal transduction pathways regulating morphopathogenic deter minants of Candida albicans. Microbiol. Mol. Biol. Rev. 2007, 71, 348–376. [Google Scholar] [CrossRef] [PubMed]
- Mayer, F.L.; Wilson, D.; Jacobsen, I.D.; Miramón, P.; Slesiona, S.; Bohovych, I.M.; Hube, B. Small but crucial: The novel small heat shock protein Hsp21 mediates stress adaptation and virulence in Candida albicans. PLoS ONE 2012, 7, e38584. [Google Scholar] [CrossRef] [PubMed]
- Palige, K.; Linde, J.; Martin, R.; Böttcher, B.; Citiulo, F.; Sullivan, D.J.; Morschhäuser, J. Global transcriptome sequencing identifies chlamydospore specific markers in Candida albicans and Candida dubliniensis. PLoS ONE 2013, 8, e61940. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).