COVID-19 Biomarkers and Advanced Sensing Technologies for Point-of-Care (POC) Diagnosis
Abstract
1. Introduction
1.1. COVID-19 History and Statistics
1.2. Importance of COVID-19 Biomarkers in Virus Detection and Sensing
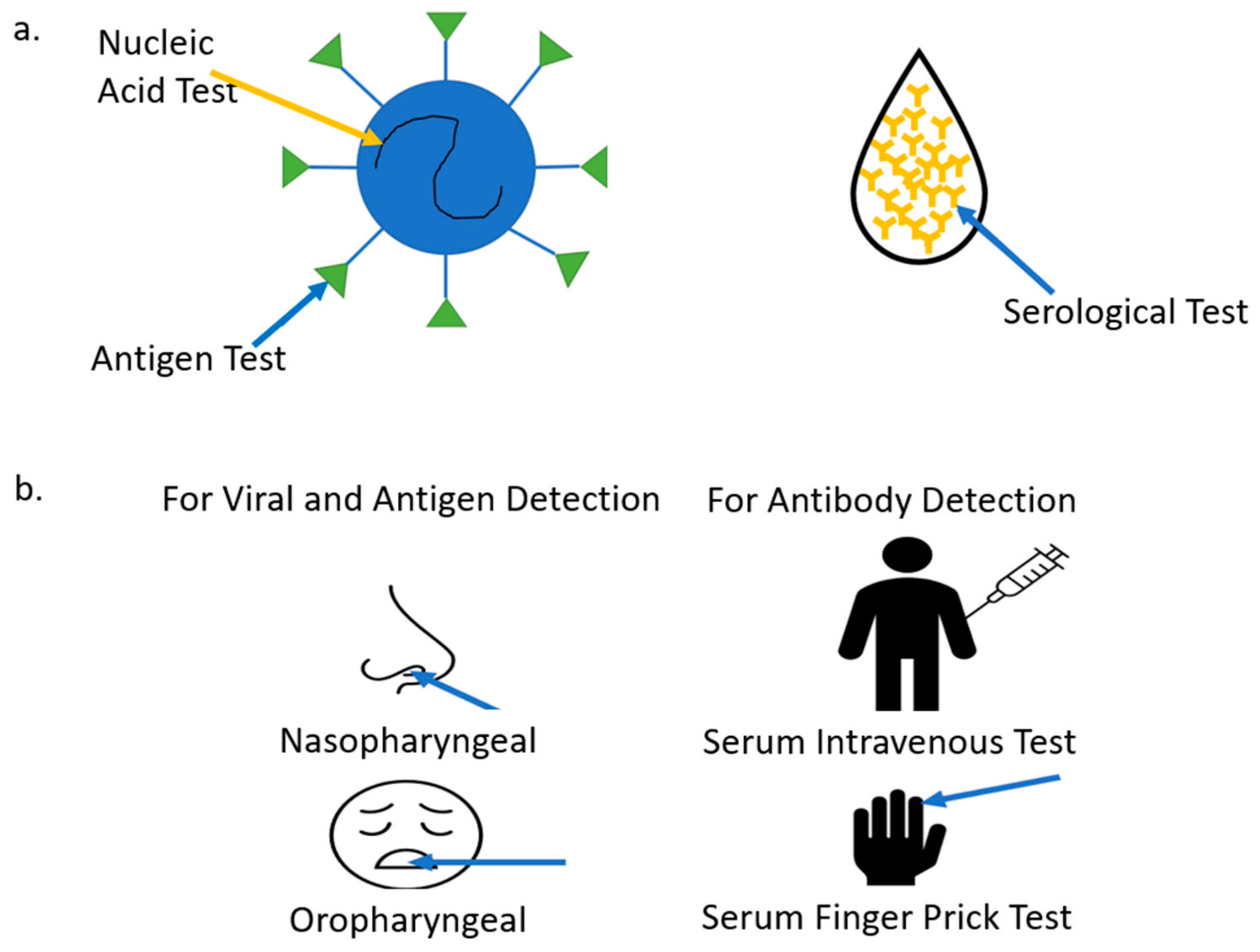
2. COVID-19 Biomarkers Classification
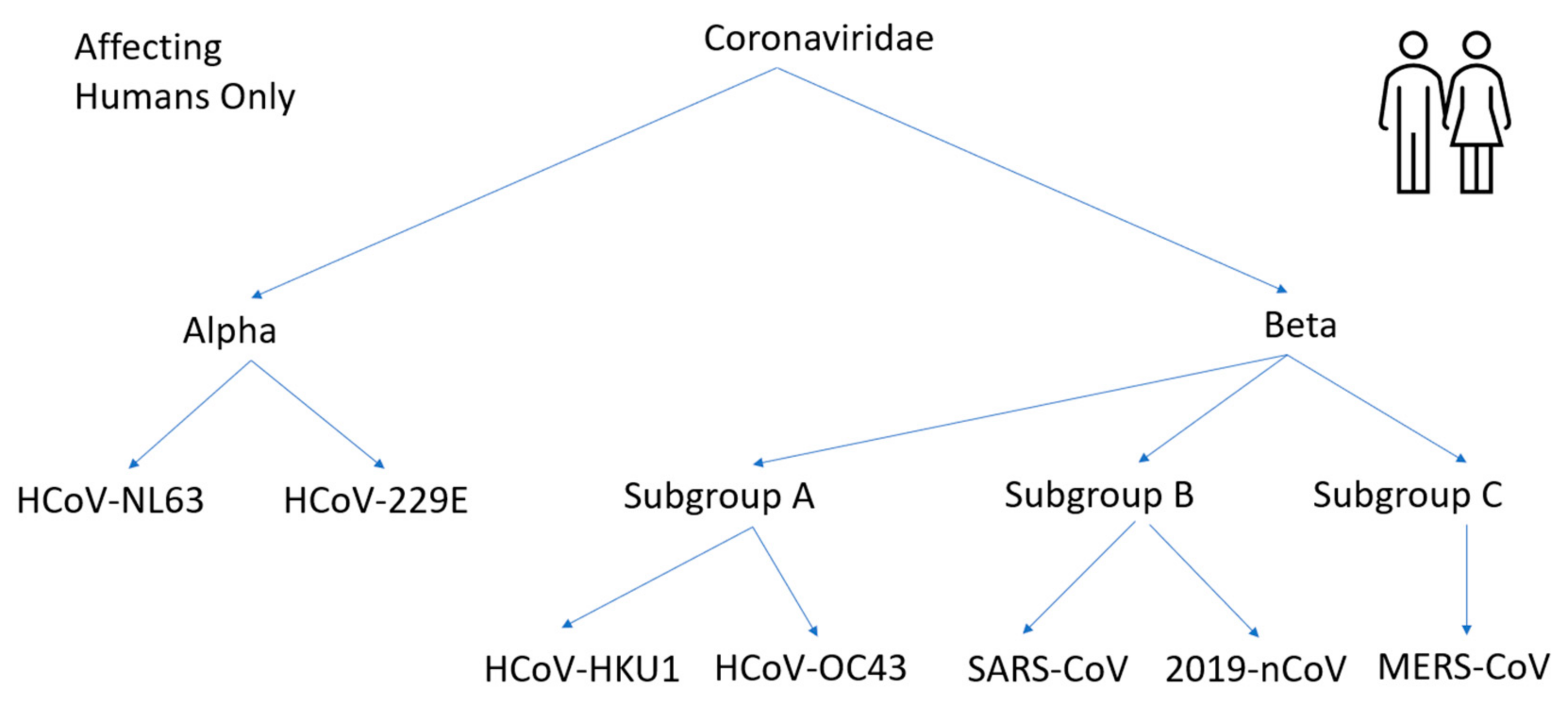
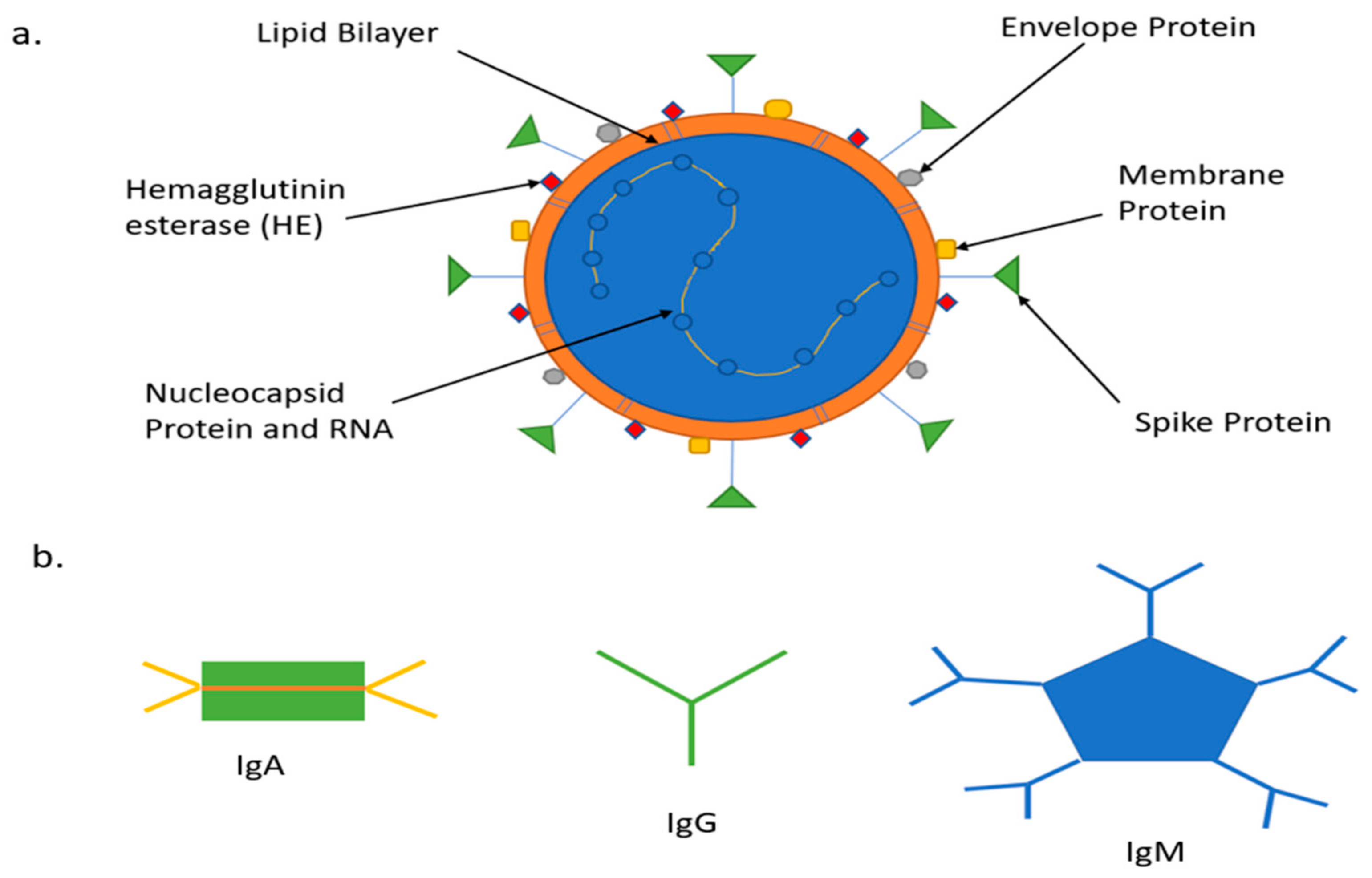
2.1. Genome Structure and Antigens
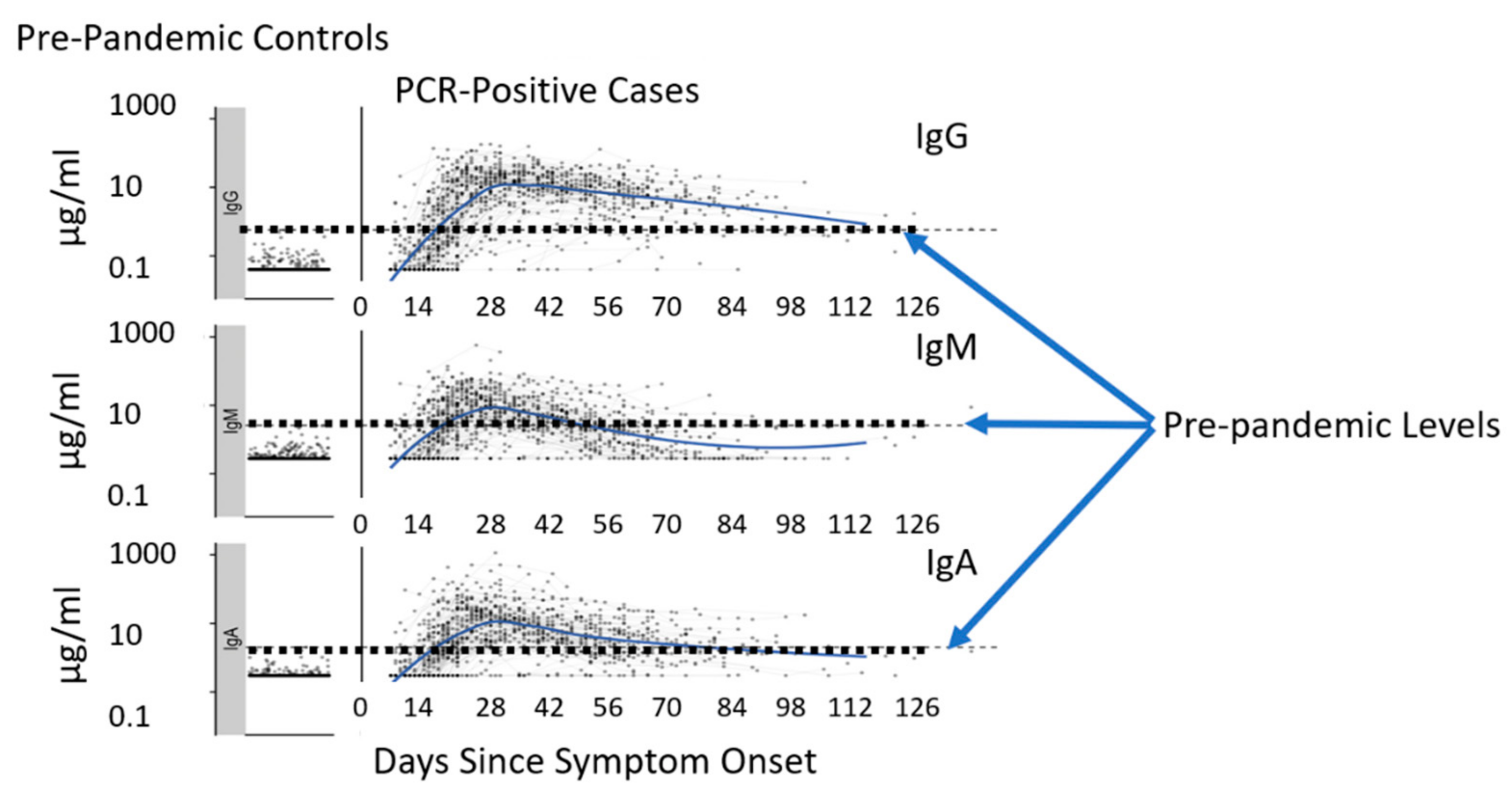
2.2. COVID-19 Antibodies
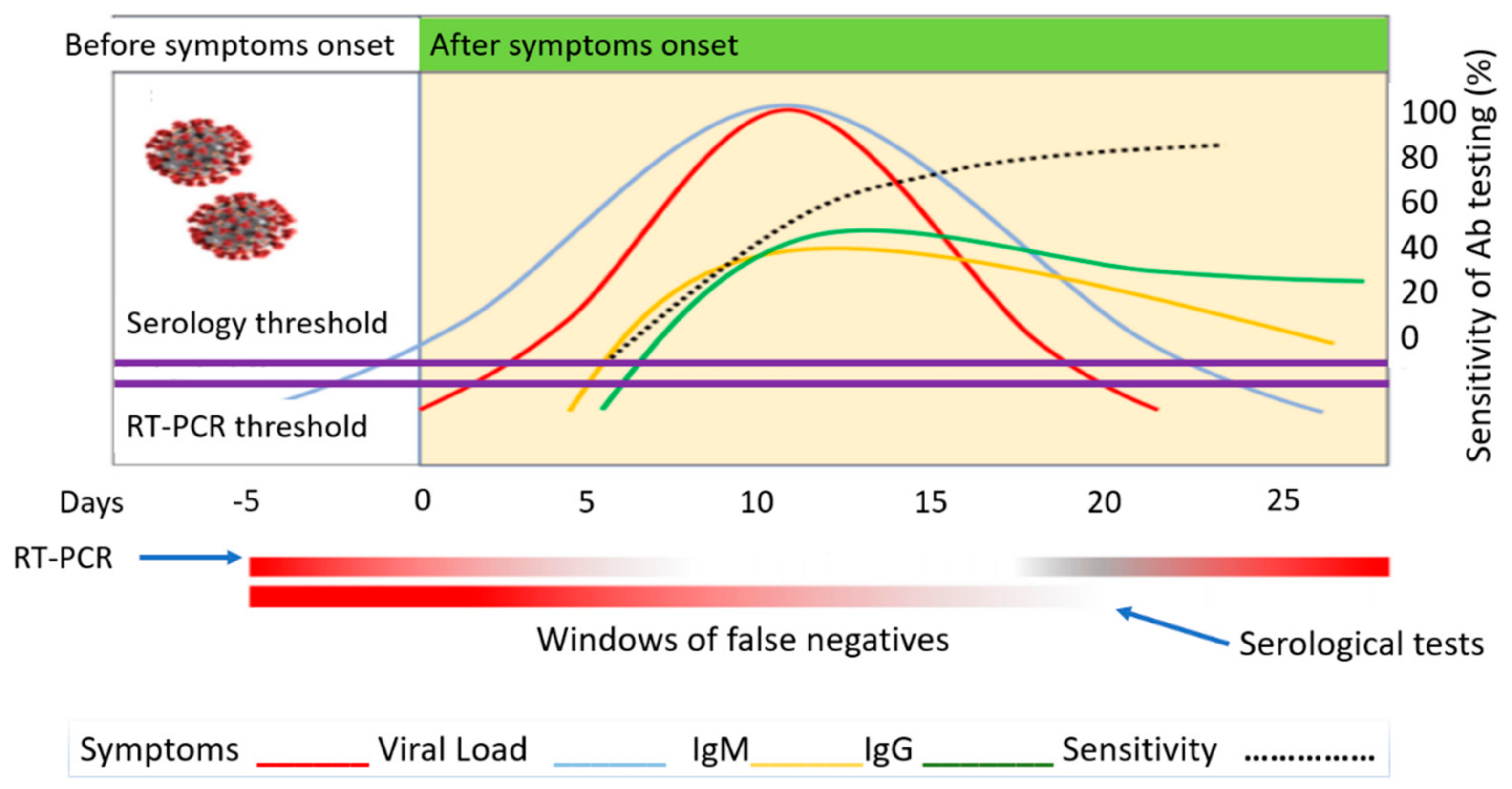
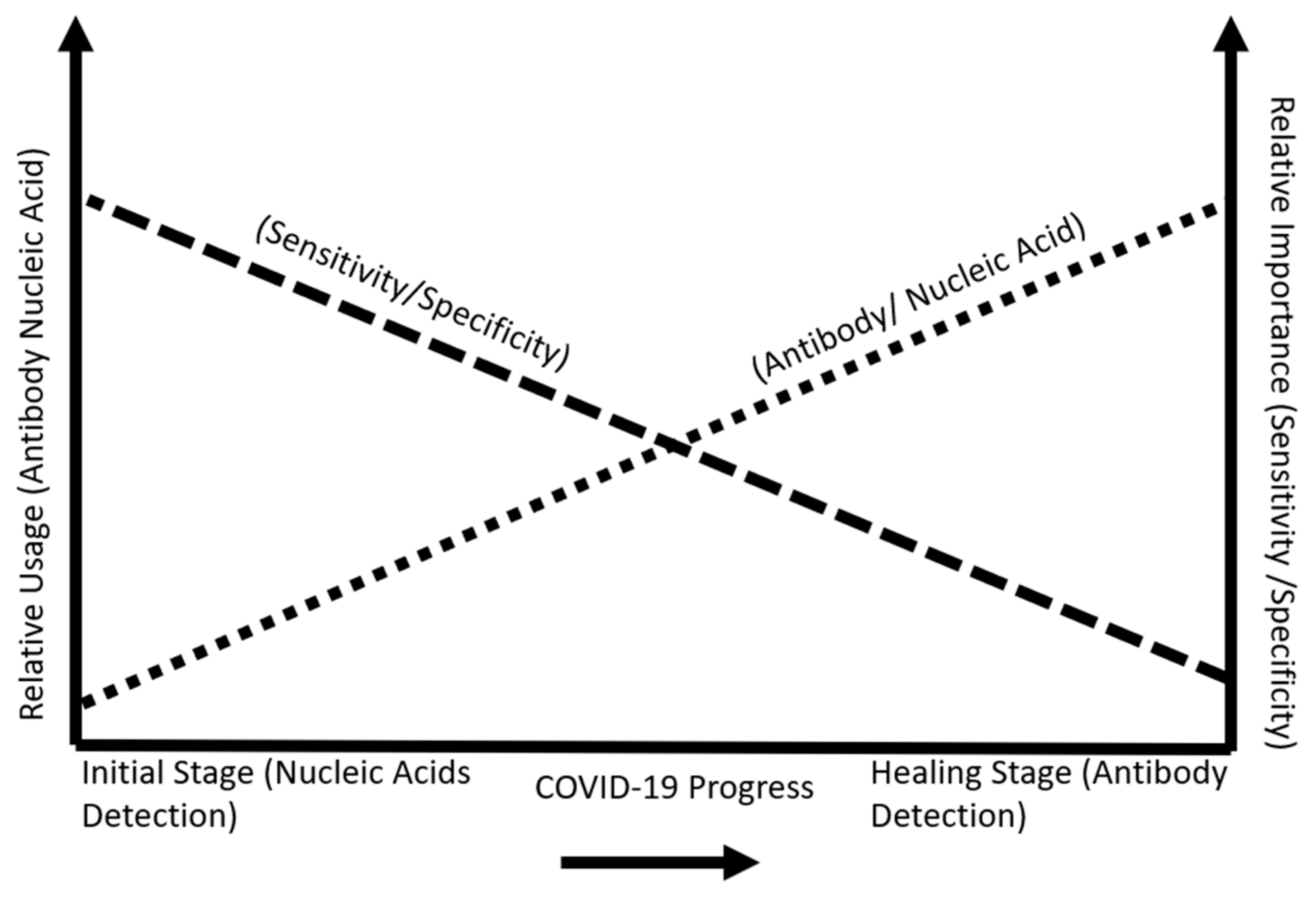
2.3. Biomarker Progression in the Human Body
3. Advanced Sensing Technologies and COVID-19 Biomarker Detection
3.1. Specimen and Sample Types
3.2. Currently Implemented Tests
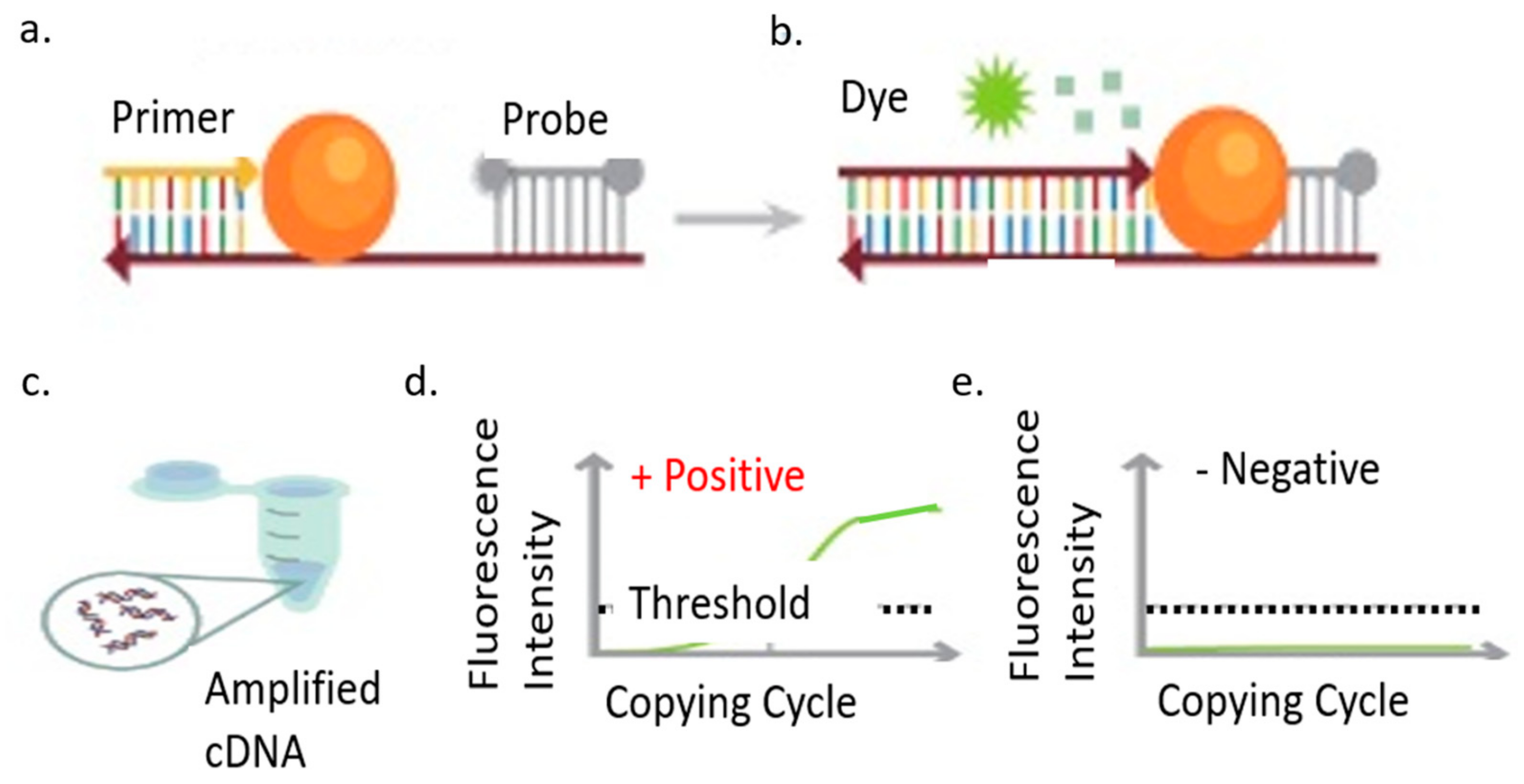
3.2.1. Reverse Transcription-Quantitative Polymerase Chain Reaction (RT-PCR)
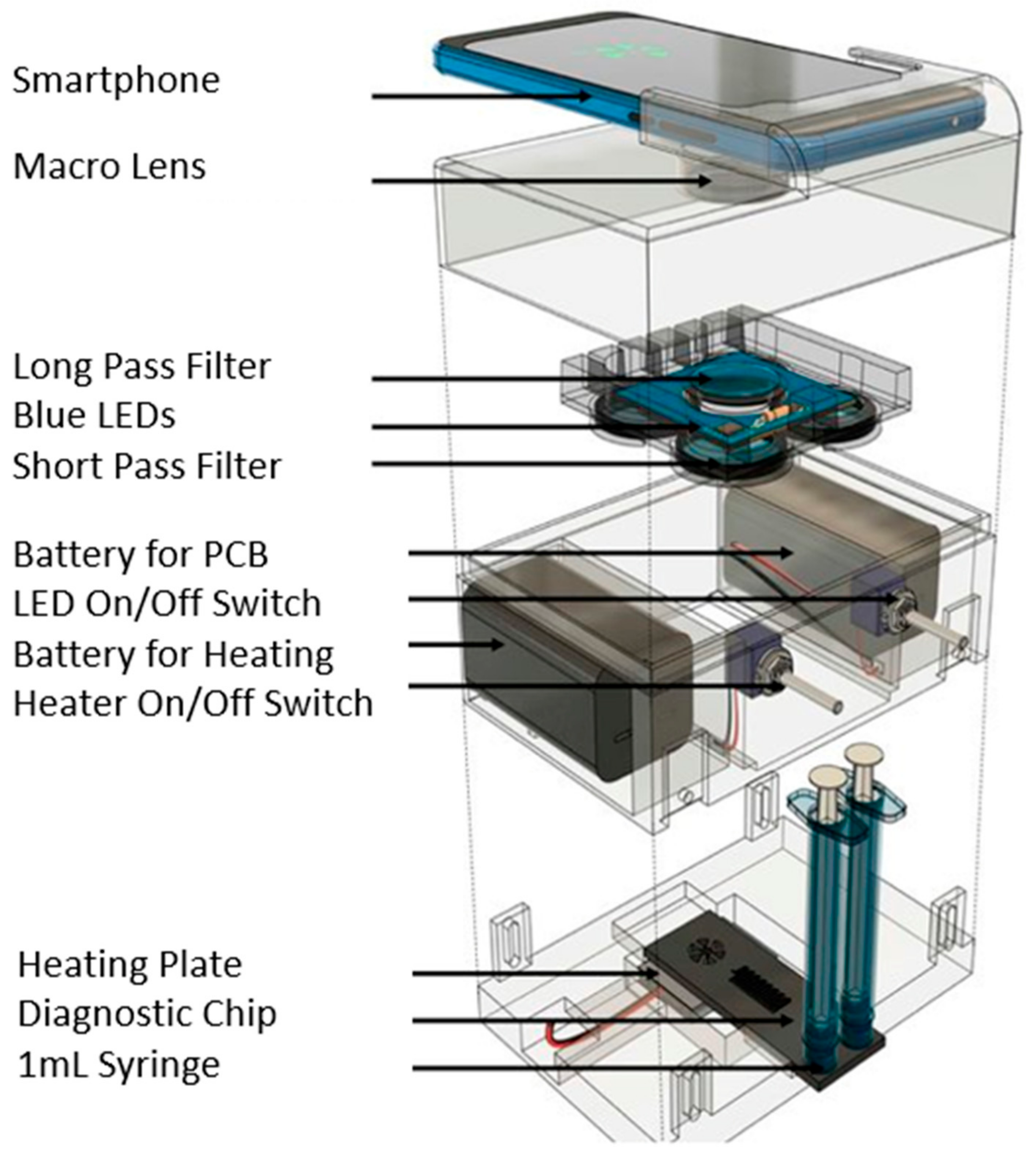
3.2.2. Reverse Transcription-Loop Mediated Isothermal Amplification (RT-LAMP)
3.3. Recent Advances in COVID-19 Sensing
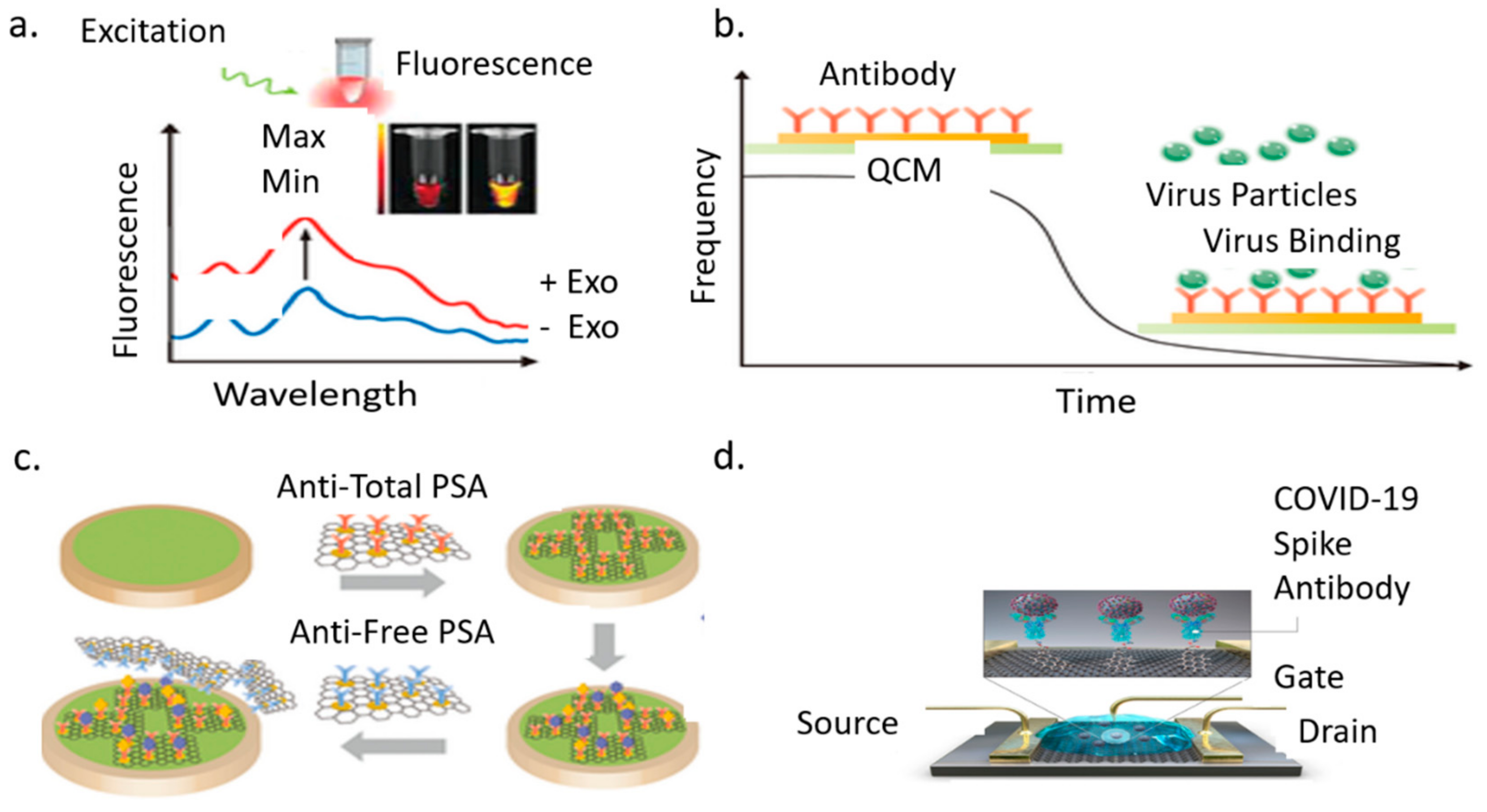
3.3.1. Optical Biosensor
3.3.2. Piezoelectric Biosensor
3.3.3. Electrochemical Biosensor
3.3.4. Graphene Field Effect Transistor
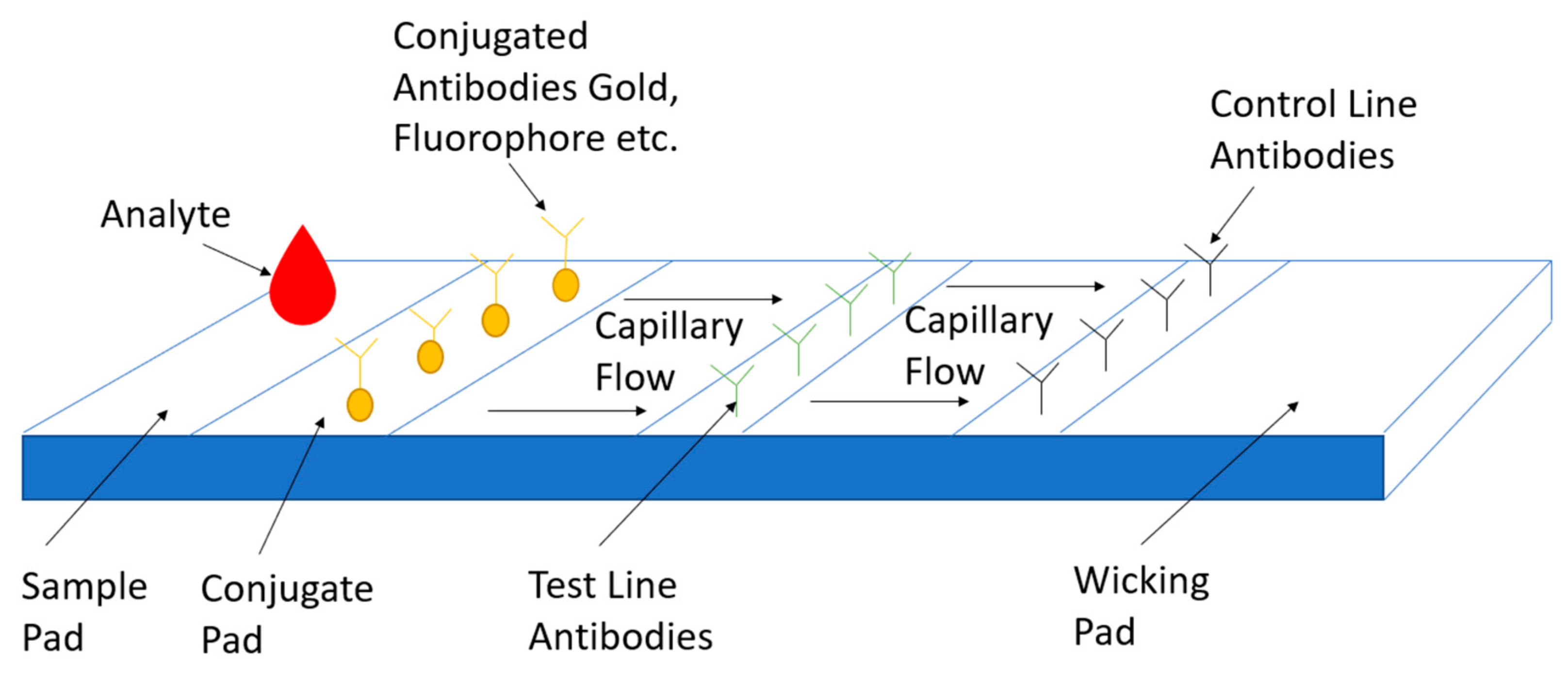
3.3.5. Lateral Flow Assay
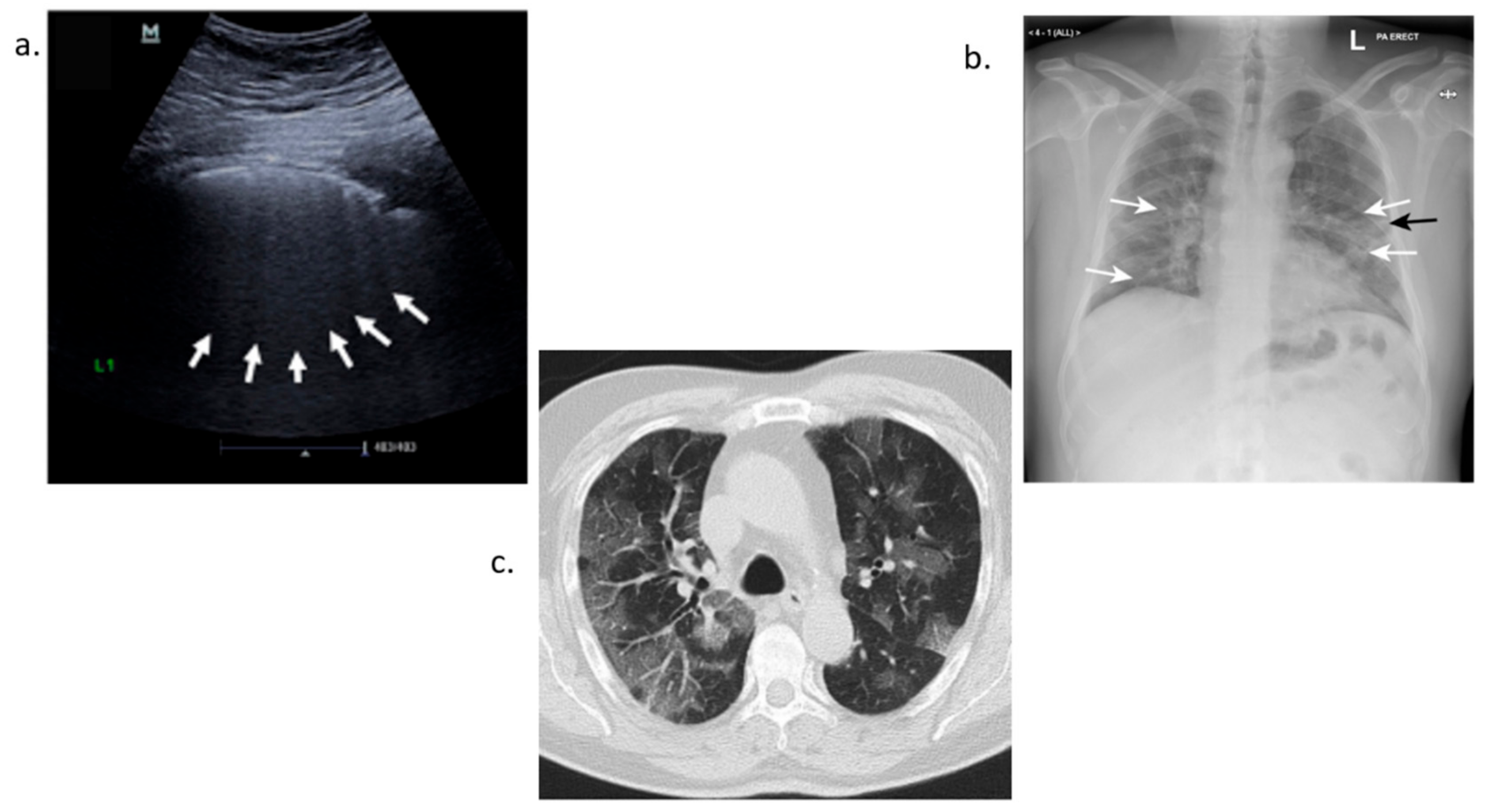
3.4. Imaging Detection of COVID-19
3.4.1. Ultrasound Detection
3.4.2. X-ray Detection
3.4.3. Computerized Tomography Scan-CT Scan Detection
4. Future Directions
4.1. Challenges Currently Facing Tests on the Market
4.2. Cross-Reactivity between Different Types of Coronaviruses
4.3. Multiple Biomarkers Used for COVID-19 Detection
5. Conclusions
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
Abbreviations
| MDPI | Multidisciplinary Digital Publishing Institute |
| DOAJ | Directory of open access journals |
| TLA | Three-letter acronym |
| LD | Linear dichroism |
| RT-PCR | Reverse Transcription-Polymerase Chain Reaction |
| FET | Field Effect Transistor |
| WHO | World Health Organization |
| SARS-CoV-2 | Severe Acute Respiratory Syndrome 2 |
| MERS-CoV | Middle East Respiratory Syndrome |
| 2019-nCoV | COVID-19 |
| ACE2 | Angiotensin Converting Enzyme |
| TMPRSS2 | Transmembrane serine protease 2 |
| DDP4 | Dipeptidyl peptidase-4 |
| EUA | Emergency Use Authorization |
| USD | United States Dollar |
| POC | Point-Of-Care |
| N | Nucleocapsid |
| S | Spike |
| Ig | Immunoglobulin |
| IL | Interleukin |
| ssRNA | single-strand Ribonucleic Acid |
| kbp | kilobase pairs |
| ORF | Open reading frame |
| NSP | Non-Structural Protein |
| RdRp | RNA dependent RNA Polymerase |
| NTD | Amino Group Terminal Domain |
| CTD | Carboxyl Group Terminal Domain |
| nM | nanoMolar |
| Abs | Antibodies |
| U/mL | Units per milliliter |
| IFN | Interferon |
| EGF | Epidermal Growth Factor |
| PDGF | Platelet Derived Growth Factor |
| VEGF | Vascular endothelial Growth Factor |
| CXCL | Chemokine |
| TNF | Tumor Necrosis Factor |
| ng/mL | nanograms per milliliter |
| pg/mL | picograms per milliliter |
| µg/mL | microgram per milliliter |
| fg/mL | femtogram per milliliter |
| DNA | Deoxyribonucleic acid |
| cDNA | Complimentary DNA |
| NP | Nucleocapsid Protein |
| SP | Spike Protein |
| NAAT | Nucleic Acid Amplification Technique |
| LAMP | Loop-mediated isothermal amplification |
| CRISPR | Clustered Regularly Interspaced Short Palindromic Repeats |
| DETECTR | DNA Endonuclease Targeted CRISPR Trans Reporter |
| TIR | Internal Reflection |
| RI | Refractive Index |
| RIU | Refractive Index Units |
| SPR | Surface Plasmon Resonance |
| LSPR | Localized Surface Plasmon Resonance |
| AuNI | Gold Nano Island |
| PPT | Plasmonic Photothermal |
| PQC | Piezoelectric Quartz Crystal |
| MB | Magnetic Beads |
| pfu/mL | plaque forming units |
| AuNPs | Gold Nanoparticles |
| fM | femtoMolar |
| FTO | Fluorine Doped Tin Oxide |
| SPCE | Screen Printed Carbon Electrodes |
| AC | Alternating Current |
| FET | Field Effect Transistor |
| PBASE | 1-pyrenebutanoic acid succinimidyl ester |
| LOD | Limit of Detection |
| LFA | Lateral Flow Assay |
| LED | Led Emitting Diode |
| LUS | Lung Ultrasound Score |
| Xray | Energetic High-Frequency Electromagnetic Radiation |
| CT | Computerized Tomography |
| nt | nucleotide |
References
- Liu, Y.-C.; Kuo, R.-L.; Shih, S.-R. COVID-19: The first documented coronavirus pandemic in history. Biomed. J. 2020, 43, 328–333. [Google Scholar] [CrossRef]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta BioMed. 2020, 91, 157–160. [Google Scholar]
- Dos Santos, W.G. Natural history of COVID-19 and current knowledge on treatment therapeutic options. Biomed. Pharmacother. 2020, 129, 110493. [Google Scholar] [CrossRef] [PubMed]
- WHO Novel Coronavirus (2019-nCoV) Situation Report 1. Available online: https://www.who.int/docs/default-source/coronaviruse/situationreports/20200121-sitrep-1-2019-ncov.pdf?sfvrsn=20a99c104 (accessed on 15 April 2020).
- COVID-19 Dashboard by the Center for Systems Science and Engineering at John Hopkins University (JHU). Available online: https://gisanddata.maps.arcgis.com/apps/opsdashboard/index.html/bda7594740fd40299423467b48e9ecf6 (accessed on 15 April 2020).
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus-Infected Pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Guo, H.; Zhou, P.; Shi, Z.-L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 2021, 19, 141–154. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Liu, X.; Pan, W.; Douglas, M.W.; Bao, S. Epidemiology of Coronavirus Disease in Gansu Province, China. Emerg. Infect. Dis. 2020, 26, 1257–1265. [Google Scholar] [CrossRef]
- Mansour, N. Coronavirus the Financial Crisis. J. Int. Acad. Case Stud. 2020, 26, 3. [Google Scholar] [CrossRef]
- Lam, T.T.Y.; Jia, N.; Zhang, Y.W.; Shum, M.H.; Jiang, J.F.; Zhu, H.C.; Tong, Y.G.; Shi, Y.X.; Ni, X.B.; Liao, Y.S.; et al. Identifying SARS-CoV-2-related coronaviruses in Malayan pangolins. Nature 2020, 583, 282–285. [Google Scholar] [CrossRef]
- Zhang, L.; Guo, H. Biomarkers of COVID-19 and technologies to combat SARS-CoV-2. Adv. Biomark. Sci. Technol. 2020, 2, 1–23. [Google Scholar] [CrossRef]
- La Marca, A.; Capuzzo, M.; Paglia, T.; Roli, L.; Trenti, T.; Nelson, S.M. Testing for SARS-CoV-2 (COVID-19): A systematic review and clinical guide to molecular and serological in-vitro diagnostic assays. Reprod. Biomed. Online 2020, 41, 483–499. [Google Scholar] [CrossRef]
- Pokhrel, P.; Hu, C.; Mao, H. Detecting the Coronavirus (COVID-19). ACS Sens. 2020, 5, 2283–2296. [Google Scholar] [CrossRef]
- Hodgens, A.; Gupta, V. Severe Acute Respiratory Syndrome; Stat Pearls Publishing: Tresure Island, FL, USA, 2021. Available online: https://www.ncbi.nlm.nih.gov/books/NBK558977/ (accessed on 20 April 2021).
- Li, F. Evidence for a Common Evolutionary Origin of Coronavirus Spike Protein Receptor-Binding Subunits. J. Virol. 2012, 86, 2856–2858. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.P.; Woo, P.C.Y.; Yip, C.C.Y.; Fan, R.Y.Y.; Huang, Y.; Wang, M.; Guo, R.; Lam, C.S.F.; Tsang, A.K.L.; Lai, K.K.Y.; et al. Isolation and Characterization of a Novel Betacoronavirus Subgroup A Coronavirus, Rabbit Coronavirus HKU14, from Domestic Rabbits. J. Virol. 2012, 86, 5481–5496. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, T.; Noman, M.; Almatroudi, A.; Shahid, M.; Khurshid, M.; Tariq, F.; ul Qamar, M.T.; Yu, R.; Li, B. A Novel Coronavirus 2019 Linked with Pneumonia in China: Current Status and Future Prospects. Preprints 2020, 2020020358. [Google Scholar] [CrossRef]
- De Assis, R.R.; Jain, A.; Nakajima, R.; Jasinskas, A.; Felgner, J.; Obiero, J.M.; Norris, P.J.; Stone, M.; Simmons, G.; Bagri, A.; et al. Analysis of SARS-CoV-2 antibodies in COVID-19 convalescent blood using a coronavirus antigen microarray. Nat. Commun. 2021, 12, 6. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef] [PubMed]
- Hu, T.; Liu, Y.; Zhao, M.; Zhuang, Q.; Xu, L.; He, Q. A comparison of COVID-19, SARS and MERS. PeerJ 2020, 8, e9725. [Google Scholar] [CrossRef]
- Zhu, Z.; Lian, X.; Su, X.; Wu, W.; Marraro, G.A.; Zeng, Y. From SARS and MERS to COVID-19: A brief summary and comparison of severe acute respiratory infections caused by three highly pathogenic human coronaviruses. Respir. Res. 2020, 21, 224. [Google Scholar] [CrossRef]
- Tong, T.R. Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV). Perspect. Med. Virol. 2006, 16, 43–95. [Google Scholar] [CrossRef]
- Federal Register/Volume 85, No. 63/Wednesday, 1 April 2020. Available online: https://www.govinfo.gov/content/pkg/FR-2020-04-01/pdf/2020-06746.pdf (accessed on 30 April 2021).
- Afzal, A. Molecular diagnostic technologies for COVID-19: Limitations and challenges. J. Adv. Res. 2020, 26, 149–159. [Google Scholar] [CrossRef]
- Sachdeva, S.; Davis, R.W.; Saha, A.K. Microfluidic Point-of-Care Testing: Commercial Landscape and Future Directions. Front. Bioeng. Biotechnol. 2021, 8, 602659. [Google Scholar] [CrossRef] [PubMed]
- Singh, H.; Zhuang, S.; Ingis, B.; Nunna, B.B.; Lee, E.S. Carbon-based catalysts for oxygen reduction reaction: A review on degradation mechanisms. Carbon 2019, 151, 160–174. [Google Scholar] [CrossRef]
- Nunna, B.B.; Mandal, D.; Lee, J.U.; Singh, H.; Zhuang, S.; Misra, D.; Bhuyian, N.U.; Lee, E.S. Detection of cancer antigens (CA-125) using gold nano particles on interdigitated electrode-based microfluidic biosensor. Nano Converg. 2019, 6, 1–12. [Google Scholar] [CrossRef]
- Singh, H.; Zhuang, S.; Nunna, B.B.; Lee, E.S. Thermal Stability and Potential Cycling Durability of Nitrogen-Doped Graphene Modified by Metal-Organic Framework for Oxygen Reduction Reactions. Catalysts 2018, 8, 607. [Google Scholar] [CrossRef]
- Zhuang, S.; Singh, H.; Nunna, B.B.; Mandal, D.; Boscoboinik, J.A.; Lee, E.S. Nitrogen-doped graphene-based catalyst with metal-reduced organic framework: Chemical analysis and structure control. Carbon 2018, 139, 933–944. [Google Scholar] [CrossRef]
- COVID-19-Serology-Testing Explained. Available online: https://asm.org/Articles/2020/May/COVID-19-Serology-Testing-Explained (accessed on 22 May 2021).
- Alanagreh, L.; Alzoughool, F.; Atoum, M. The Human Coronavirus Disease COVID-19: Its Origin, Characteristics, and Insights into Potential Drugs and Its Mechanisms. Pathogens 2020, 9, 331. [Google Scholar] [CrossRef]
- Khailany, R.A.; Safdar, M.; Ozaslan, M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 2020, 19, 100682. [Google Scholar] [CrossRef]
- Demogines, A.; Farzan, M.; Sawyer, S. Evidence for ACE2-Utilizing Coronaviruses (CoVs) Related to Severe Acute Respiratory Syndrome CoV in Bats. J. Virol. 2012, 86, 6350–6353. [Google Scholar] [CrossRef] [PubMed]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.-L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef]
- Shah, J.; Liu, S.; Potula, H.-H.; Bhargava, P.; Cruz, I.; Force, D.; Bazerbashi, A.; Ramasamy, R. IgG and IgM antibody formation to spike and nucleocapsid proteins in COVID-19 characterized by multiplex immunoblot assays. BMC Infect. Dis. 2021, 21, 325. [Google Scholar] [CrossRef] [PubMed]
- Coutard, B.; Valle, C.; De Lamballerie, X.; Canard, B.; Seidah, N.; Decroly, E. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 2020, 176, 104742. [Google Scholar] [CrossRef]
- Rabaan, A.A.; Al-Ahmed, S.H.; Haque, S.; Sah, R.; Tiwari, R.; Malik, Y.S.; Dhama, K.; Yatoo, M.I.; Bonilla-Aldana, D.K.; Rodriguez-Morales, A.J. SARS-CoV-2, SARS-CoV, and MERS-COV: A comparative overview. Infez. Med. 2020, 28, 174–184. [Google Scholar]
- Ingis, B.; Lee, E.S. 3D Printing for Whole Blood Filters Designed for Simple Integration with a Variety of Sensor Platforms. In Proceedings of the 2019 IEEE Healthcare Innovations and Point of Care Technologies, Bethesda, MA, USA, 20–22 November 2019; pp. 143–146. [Google Scholar] [CrossRef]
- Vidarsson, G.; Dekkers, G.; Rispens, T. IgG Subclasses and Allotypes: From Structure to Effector Functions. Front. Immunol. 2014, 5, 520. [Google Scholar] [CrossRef]
- Protective Antibodies Persist for Months in Survivors of Serious COVID-19 Infections. Available online: https://www.sciencedaily.com/releases/2020/10/201008121314.htm (accessed on 22 May 2021).
- Sterlin, D.; Mathian, A.; Miyara, M.; Mohr, A.; Anna, F.; Clear, L.; Quentric, P.; Fadlallah, J.; Devilliers, H.; Ghillani, P.; et al. IgA dominates the early neutralizing antibody response to SARS-CoV-2. Sci. Transl. Med. 2021, 13, eabd2223. [Google Scholar] [CrossRef] [PubMed]
- Goh, Y.S.; Chavatte, J.M.; Jieling, A.L.; Lee, B.; Hor, P.X.; Amrun, S.N.; Lee, C.Y.; Chee, R.S.; Wang, B.; Lee, C.Y.; et al. Sensitive detection of total anti-Spike antibodies and isotype switching in asymptomatic and symptomatic individuals with COVID-19. Cell Rep. Med. 2021, 2, 100193. [Google Scholar] [CrossRef]
- Iyer, A.S.; Jones, F.K.; Nodoushani, A.; Kelly, M.; Becker, M.; Slater, D.; Mills, R.; Teng, E.; Kamruzzaman, M.; Garcia-Beltran, W.F.; et al. Persistence and decay of human antibody responses to the receptor binding domain of SARS-CoV-2 spike protein in COVID-19 patients. Sci. Immunol. 2020, 5, eabe0367. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Pan, Y.; Li, Z.; Li, Y. The Utility of Specific Antibodies against SARS-CoV-2 in Laboratory Diagnosis. Front. Microbiol. 2021, 11, 603058. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Zheng, X.; Liu, B.; Wu, M.; Zhang, Z.; Zhang, G.; Su, X. Serum IgM against SARS-CoV-2 correlates with in-hospital mortality in severe/critical patients with COVID-19 in Wuhan, China. Aging 2020, 12, 12432–12440. [Google Scholar] [CrossRef] [PubMed]
- Rubin, E.J.; Longo, D.L.; Baden, L.R. Interleukin-6 Receptor Inhibition in COVID-19—Cooling the Inflammatory Soup. N. Engl. J. Med. 2021, 384, 1564–1565. [Google Scholar] [CrossRef] [PubMed]
- COVID-19 transmission-up in the air. COVID-19 transmission-up in the air. The Lancet Respiratory Medicine. Lancet 2020, 8, 1159. [Google Scholar] [CrossRef]
- Jia, H.P.; Look, D.C.; Shi, L.; Hickey, M.; Pewe, L.; Netland, J.; Farzan, M.; Wohlford-Lenane, C.; Perlman, S.; McCray, P.B. ACE2 Receptor Expression and Severe Acute Respiratory Syndrome Coronavirus Infection Depend on Differentiation of Human Airway Epithelia. J. Virol. 2005, 79, 14614–14621. [Google Scholar] [CrossRef] [PubMed]
- Trypsteen, W.; Van Cleemput, J.; Van Snippenberg, W.; Gerlo, S.; Vandekerckhove, L. On the whereabouts of SARS-CoV-2 in the human body: A systematic review. PLoS Pathog. 2020, 16, e1009037. [Google Scholar] [CrossRef] [PubMed]
- Parasher, A. COVID-19: Current understanding of its Pathophysiology, Clinical presentation and Treatment. Postgrad. Med. J. 2021, 97, 312–320. [Google Scholar] [CrossRef]
- Kermali, M.; Khalsa, R.K.; Pillai, K.; Ismail, Z.; Harky, A. The role of biomarkers in diagnosis of COVID-19—A systematic review. Life Sci. 2020, 254, 117788. [Google Scholar] [CrossRef]
- Alberts, B.; Johnson, A.; Lewis, J. Blood Vessels and Endothelial Cells. In Molecular Biology of the Cell, 4th ed.; Garland Science: New York, NY, USA, 2002. Available online: https://www.ncbi.nlm.nih.gov/books/NBK26848/ (accessed on 29 April 2021).
- Khalil, B.A.; Elemam, N.M.; Maghazachi, A.A. Chemokines and chemokine receptors during COVID-19 infection. Comput. Struct. Biotechnol. J. 2021, 19, 976–988. [Google Scholar] [CrossRef]
- Mallapaty, S. Mini organs reveal how the coronavirus ravages the body. Nat. Cell Biol. 2020, 583, 15–16. [Google Scholar] [CrossRef]
- Sungnak, W.; Network, H.L.B.; Huang, N.; Bécavin, C.; Berg, M.; Queen, R.; Litvinukova, M.; Talavera-López, C.; Maatz, H.; Reichart, D.; et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 2020, 26, 681–687. [Google Scholar] [CrossRef]
- Chen, L.Y.; Hoiland, R.L.; Stukas, S.; Wellington, C.L.; Sekhon, M.S. Confronting the controversy: Interleukin-6 and the COVID-19 cytokine storm syndrome. Eur. Respir. J. 2020, 56, 2003006. [Google Scholar] [CrossRef]
- Hamming, V.; Timens, W.; Bulthuis, M.L.C.; Lely, T.; Navis, G.J.; van Goor, H. Tissue distribution of ACE2 protein, the functional receptor for SARS coronavirus. A first step in understanding SARS pathogenesis. J. Pathol. 2004, 203, 631–637. [Google Scholar] [CrossRef]
- Alhalaili, B.; Popescu, I.; Kamoun, O.; Alzubi, F.; Alawadhia, S.; Vidu, R. Nanobiosensors for the Detection of Novel Coronavirus 2019-nCoV and Other Pandemic/Epidemic Respiratory Viruses: A Review. Sensors 2020, 20, 6591. [Google Scholar] [CrossRef]
- Marty, F.M.; Chen, K.; Verrill, K.A. How to Obtain a Nasopharyngeal Swab Specimen. N. Engl. J. Med. 2020, 382, e76. [Google Scholar] [CrossRef] [PubMed]
- Petruzzi, G.; De Virgilio, A.; Pichi, B.; Mazzola, F.; Zocchi, J.; Mercante, G.; Spriano, G.; Pellini, R. COVID-19: Nasal and oropharyngeal swab. Head Neck 2020, 42, 1303–1304. [Google Scholar] [CrossRef] [PubMed]
- Isho, B.; Abe, K.T.; Zuo, M.; Jamal, A.J.; Rathod, B.; Wang, J.H.; Li, Z.; Chao, G.; Rojas, O.L.; Bang, Y.M.; et al. Persistence of serum and saliva antibody responses to SARS-CoV-2 spike antigens in COVID-19 patients. Sci. Immunol. 2020, 5, abe5511. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Tan, L.; Wang, X.; Liu, W.; Lu, Y.; Cheng, L.; Sun, Z. Comparison of nasopharyngeal and oropharyngeal swabs for SARS-CoV-2 detection in 353 patients received tests with both specimens simultaneously. Int. J. Infect. Dis. 2020, 94, 107–109. [Google Scholar] [CrossRef] [PubMed]
- Padoan, A.; Sciacovelli, L.; Basso, D.; Negrini, D.; Zuin, S.; Cosma, C.; Faggian, D.; Matricardi, P.; Plebani, M. IgA-Ab response to spike glycoprotein of SARS-CoV-2 in patients with COVID-19: A longitudinal study. Clin. Chim. Acta 2020, 507, 164–166. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, S.; Nunna, B.B.; Lee, E.S. Metal organic framework-modified nitrogen-doped graphene oxygen reduction reaction catalyst synthesized by nanoscale high-energy wet ball-milling structural and electrochemical characterization. MRS Commun. 2018, 8, 40–48. [Google Scholar] [CrossRef]
- Nunna, B.B.; Lee, E.S. Point-of-Care (POC) Micro Biochip for Cancer Diagnostics. In TechConnect Briefs 2017: Biomaterials and Biomedical; Taylor Francis: Abingdon, UK, 2017; Volume 3, pp. 110–113. ISBN 978-0-9988782-0-1. [Google Scholar]
- Nunna, B.B.; Mandal, D.; Zhuang, S.; Lee, E.S. A standalone micro biochip to monitor the cancer progression by measuring cancer antigens as a point-of-care (POC) device for enhanced cancer management. In Proceedings of the 2017 IEEE Healthcare Innovations and Point of Care Technologies (HI-POCT), Bethesda, MA, USA, 6–8 November 2017; pp. 212–215. [Google Scholar]
- Lee, E.S.; Nunna, B.B. Biomarker Detection and Self-Separation of Serum during Capillary Flow. U.S. Patent US10481154B2, 19 November 2019. Available online: https://patents.google.com/patent/US10481154B2/en (accessed on 1 May 2021).
- Mesoraca, A.; Margiotti, K.; Viola, A.; Cima, A.; Sparacino, D.; Giorlandino, C. Evaluation of SARS-CoV-2 viral RNA in fecal samples. Virol. J. 2020, 17, 1–3. [Google Scholar] [CrossRef]
- Types of COVID-19 Tests. Available online: https://mammoth.bio/2020/06/17/types-of-covid-19-tests/ (accessed on 1 May 2021).
- WHO. Coronavirus Disease (COVID-19) Advice for the Public. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/advice-for-public (accessed on 22 May 2021).
- Bhalla, N.; Pan, Y.; Yang, Z.; Payam, A.F. Opportunities and Challenges for Biosensors and Nanoscale Analytical Tools for Pandemics: COVID-19. ACS Nano 2020, 14, 7783–7807. [Google Scholar] [CrossRef]
- Park, G.-S.; Ku, K.; Baek, S.-H.; Kim, S.-J.; Kim, S.I.; Kim, B.-T.; Maeng, J.-S. Development of Reverse Transcription Loop-Mediated Isothermal Amplification Assays Targeting Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). J. Mol. Diagn. 2020, 22, 729–735. [Google Scholar] [CrossRef]
- Shen, M.; Zhou, Y.; Ye, J.; Al-Maskri, A.A.A.; Kang, Y.; Zeng, S.; Cai, S. Recent advances and perspectives of nucleic acid detection for coronavirus. J. Pharm. Anal. 2020, 10, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Wang, Y.-C.; Shen, C.-F.; Cheng, C.-M. Point-of-Care RNA-Based Diagnostic Device for COVID-19. Diagnostics 2020, 10, 165. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Fan, C.; Li, M.; Nie, H.-L.; Wang, F.-B.; Wang, H.; Wang, R.; Xia, J.; Zheng, X.; Zuo, X.; et al. COVID-19: A Call for Physical Scientists and Engineers. ACS Nano 2020, 14, 3747–3754. [Google Scholar] [CrossRef] [PubMed]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.; Chan, W.C. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano 2020, 14, 3822. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Tong, J.; Liu, M.; Shen, Y.; Guo, D. Evaluation of coronavirus in tears and cunjunctival secretions of patients with SARS-CoV-2 infection. J. Med. Virol. 2020, 92, 584–594. [Google Scholar] [CrossRef]
- Alpdagtas, S.; Ilhan, E.; Uysal, E.; Sengor, M.; Ustundag, C.B.; Gunduz, O. Evaluation of current diagnostic methods for COVID-19. APL Bioeng. 2020, 4, 041506. [Google Scholar] [CrossRef]
- Sheridan, C. Fast, portable tests come online to curb coronavirus pandemic. Nat. Biotechnol. 2020, 38, 515–518. [Google Scholar] [CrossRef]
- Thompson, D.; Lei, Y. Mini Review: Recent Progress in RT-LAMP enabled COVID-19 detection. Sens. Actuators Rep. 2020, 2, 100017. [Google Scholar] [CrossRef]
- Emergency Use Authorization (EUA) Summary. Covid-19 RT-PCR Test. (Laboratory Corporation of America). Available online: https://www.fda.gov/media/136151/download (accessed on 29 April 2021).
- Tahamtan, A.; Ardebili, A. Real-time RT-PCR in COVID-19 detection: Issues affecting the results. Expert Rev. Mol. Diagn. 2020, 20, 453–454. [Google Scholar] [CrossRef]
- Shiao, Y.-H. A new reverse transcription-polymerase chain reaction method for accurate quantification. BMC Biotechnol. 2003, 3, 22. [Google Scholar] [CrossRef]
- Nunna, B.B.; Zhuang, S.; Javier, J.; Mandal, D.; Lee, E.S. Biomolecular Detection using Molecularly Imprinted Polymers (MIPs) at Point-of-Care (POC) Micro Biochip. In Proceedings of the 2016 IEEE-NIH 2016 Healthcare Innovation Point of Care Technologies Conference HIPOCT16, (PCHT16-0099), Cancun, Mexico, 9–11 November 2016. [Google Scholar]
- Nunna, B.B.; Zhuang, S.; Lee, E.S. Hemorheology in PDMS micro channel with varied surface roughness. In APS Division of Fluid Dynamics Meeting Abstracts; American Physical Society: Boston, MA, USA, 2015; p. KP1-116. [Google Scholar]
- Nunna, B.B.; Zhuang, S.; Lee, E.S. Flow control mechanism of capillary driven flow in microchannel using non-mechanical forces. In Proceedings of the Bulletin of the American Physical Society, APS—69th Annual Meeting of the APS Division of Fluid Dynamics—Session Index DFD16, Portland, OR, USA, 20–22 November 2016; pp. 1–2. [Google Scholar]
- Alekseenko, A.; Barrett, D.; Pareja-Sanchez, Y.; Howard, R.J.; Strandback, E.; Ampah-Korsah, H.; Rovšnik, U.; Zuniga-Veliz, S.; Klenov, A.; Malloo, J.; et al. Direct detection of SARS-CoV-2 using non-commercial RT-LAMP reagents on heat-inactivated samples. Sci. Rep. 2021, 11, 1820. [Google Scholar] [CrossRef]
- Silva, S.; Pardee, K.; Pena, L. Loop Mediated Isothermal Amplification (LAMP) for the Diagnosis of Zika Virus: A review. Viruses 2020, 17, 19. [Google Scholar] [CrossRef]
- Zhang, Y.; Odiwuor, N.; Xiong, J.; Sun, L.; Nyaruaba, R.O.; Wei, H.; Tanner, N.A. A Rapid Molecular Detection of SARS-CoV-2(COVID-19 Virus RNA using colorimetric lamp. medRxiv 2020. [Google Scholar] [CrossRef]
- Carter, L.J.; Garner, L.V.; Smoot, J.W.; Li, Y.; Zhou, Q.; Saveson, C.J.; Sasso, J.M.; Gregg, A.C.; Soares, D.J.; Beskid, T.R.; et al. Assay Techniques and Test Development for COVID-19 Diagnosis. ACS Cent. Sci. 2020, 6, 591–605. [Google Scholar] [CrossRef]
- SARS-CoV-2 Reference Panel Comparative Data. Available online: https://www.fda.gov/medical-devices/coronavirus-covid-19-and-medical-devices/sars-cov-2-reference-panel-comparative-data (accessed on 29 April 2021).
- Diego, J.G.-B.; Fernández-Soto, P.; Domínguez-Gil, M.; Belhassen-García, M.; Bellido, J.; Muro, A. A Simple, Affordable, Rapid, Stabilized, Colorimetric, Versatile RT-LAMP Assay to Detect SARS-CoV-2. Diagnostics 2021, 11, 438. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Abudayyeh, O.O.; Gootenberg, J.S. A Protocol for Detection of COVID-19 Using CRISPR Diagnostics. 2020. Available online: https://broad.io/sherlockprotocol (accessed on 29 April 2021).
- Ganguli, A.; Ornob, A.; Yu, H.; Damhorst, G.L.; Chen, W.; Sun, F.; Bhuiya, A.; Cunningham, B.T.; Bashir, R. Hands free smartphone based diagnostics for siumultaneous detection of Zika, Chikungunya and Dengue at point of care. Biomed. Microdev. 2017, 19, 73. [Google Scholar] [CrossRef] [PubMed]
- Mautner, L.; Baillie, C.-K.; Herold, H.M.; Volkwein, W.; Guertler, P.; Eberle, U.; Ackermann, N.; Sing, A.; Pavlovic, M.; Goerlich, O.; et al. Rapid point-of-care detection of SARS-CoV-2 using reverse transcription loop-mediated isothermal amplification (RT-LAMP). Virol. J. 2020, 17, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.K.; Du, P.; Lgnacio, C.; Mehta, S.; Newton, I.G.; Steinmetz, N.F. Virus Like Particles as Positive Controls for COVID-19 RT LMAP Diangostic Assays. Biomacromolecules 2021, 22, 1231–1243. [Google Scholar]
- Zhu, X.; Wang, X.; Han, L.; Chen, T.; Wang, L.; Li, H.; Li, S.; He, L.; Fu, X.; Chen, S.; et al. Multiplex reverse transcription loop-mediated isothermal amplification combined with nanoparticle-based lateral flow biosensor for the diagnosis of COVID-19. Biosens. Bioelectron. 2020, 166, 112437. [Google Scholar] [CrossRef]
- Vogels, C.B.F.; Brito, A.F.; Wyllie, A.L.; Fauver, J.R.; Ott, I.M.; Kalinich, C.C.; Petrone, M.E.; Casanovas-Massana, A.; Muenker, M.C.; Moore, A.J.; et al. Analytical Sensitivity and Efficiency Comparisons of SARS-CoV-2 RTQPCR PrimerProbe Sets. Nat. Microbiol. 2020, 5, 12991305. [Google Scholar] [CrossRef]
- Hasan, M.R.; Mirza, F.; Al-Hail, H.; Sundararaju, S.; Xaba, T.; Iqbal, M.; Alhussain, H.; Yassine, H.M.; Perez-Lopez, A.; Tang, P. Detection of SARS-CoV-2 RNA by Direct RT-QPCR on Nasopharyngeal Specimens without Extraction of Viral RNA. PLoS ONE 2020, 15, e0236564. [Google Scholar]
- Thi, V.L.D.; Herbst, K.; Boerner, K.; Meurer, M.; Kremer, L.P.; Kirrmaier, D.; Freistaedter, A.; Papagiannidis, D.; Galmozzi, C.; Stanifer, M.L.; et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl. Med. 2020, 12, eabc7075. [Google Scholar] [CrossRef]
- Yan, C.; Cui, J.; Huang, L.; Du, B.; Chen, L.; Xue, G.; Li, S.; Zhang, W.; Zhao, L.; Sun, Y.; et al. Rapid and Visual Detection of 2019 Novel Coronavirus (SARS-CoV-2) by A Reverse Transcription LoopMediated Isothermal Amplification Assay. Clin. Microbiol. Infect. 2020, 26, 773779. [Google Scholar] [CrossRef] [PubMed]
- The Basic Science Behind COVID-19 Molecular Testing Methods. Available online: https://acmgloballab.com/about-us/resources/COVID19-molecular-testing-methods (accessed on 22 May 2021).
- Ganguli, A.; Mostafa, A.; Berger, J.; Aydin, M.Y.; Sun, F.; de Ramirez, S.A.S.; Valera, E.; Cunningham, B.T.; King, W.P.; Bashir, R. Rapid isothermal amplification and portable detection system for SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 22727–22735. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, S.; Nunna, B.B.; Mandal, D.; Lee, E.S. A review of nitrogen-doped graphene catalysts for proton exchange membrane fuel cells-synthesis, characterization, and improvement. Nano-Struct. Nano-Objects 2018, 15, 140–152. [Google Scholar] [CrossRef]
- Lee, E.S.; Nunna, B.B.; Suh, S.K. Microfluidic Diagnostic Assembly. U.S. Patent US10898894B2, 26 January 2021. Available online: https://patents.google.com/patent/US10898894B2/en (accessed on 22 April 2021).
- Lee, E.S.; Nunna, B.B. Enhanced Sensitivity and Specificity for Point-of-Care (POC) Micro Biochip. U.S. Patent US20200182864A1, 11 June 2020. Available online: https://patents.google.com/patent/US20200182864A1/en (accessed on 22 April 2021).
- Lee, E.S.; Nunna, B.B. Microfluidic Biochip with Enhanced Sensitivity. U.S. Patent US11020740B2, 1 June 2021. Available online: https://patents.google.com/patent/US11020740B2/en (accessed on 22 April 2021).
- Bhalla, N.; Jolly, P.; Formisano, N.; Estrela, P. Introduction to biosensors. Essays Biochem. 2016, 60, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Mehrotra, P. Biosensors and their applications—A review. J. Oral Biol. Craniofacial Res. 2016, 6, 153–159. [Google Scholar] [CrossRef]
- Carpenter, A.C.; Paulsen, I.T.; Williams, T.C. Blueprints for Biosensors: Design, Limitations, and Applications. Genes 2018, 9, 375. [Google Scholar] [CrossRef] [PubMed]
- Parihar, A.; Ranjan, P.; Sanghi, S.; Srivastava, A.K.; Khan, R. Point of Care Biosensor Based Diagnosis of COVID-19 Holds Promise to Combat Current and Future Pandemics. ACS Appl. Bio Mater. 2020, 3, 7326–7343. [Google Scholar] [CrossRef]
- Fan, Z.; Geng, Z.; Fang, W.; Lu, X.; Su, Y.; Wang, S.; Chen, H. Smartphone Biosensor System with Multi Testing Unit Based on Localized Surface Plasmon Resonance Integrated with Microfluidics Chip. Sensors 2020, 20, 446. [Google Scholar] [CrossRef]
- Saylan, Y.; Erden, O.; Unal, S.; Denzli, A. An Alternative Medical Diagnosis Method: Biosensors for Virus Detection. Biosensors 2019, 9, 65. [Google Scholar] [CrossRef]
- Daaboul, G.G.; Lopez, C.A.; Yurt, A.; Goldberg, B.B.; Connor, J.; Ünlü, M.S. Label-Free Optical Biosensors for Virus Detection and Characterization. IEEE J. Sel. Top. Quantum Electron. 2011, 18, 1422–1433. [Google Scholar] [CrossRef]
- Ciminelli, C.; Dell’Olio, F.; Conteduca, D.; Armenise, M.N. Silicon photonic biosensors. IET Optoelectron. 2019, 13, 48–54. [Google Scholar] [CrossRef]
- Leonardi, A.A.; Lo Faro, M.J.; Petralia, S.; Fazio, B.; Musumeci, P.; Conoci, S.; Irrera, A.; Priolo, F. Ultrasensitive Lable and PCR free Genome Detection Based on Cooperative Hybridization of Silicon Nanowires Optical Biosensors. ACS Sens. 2018, 3, 1690–1697. [Google Scholar] [CrossRef]
- Oliverio, M.; Perotto, S.; Messira, G.C.; Lovato, L.; De Anglis, F. Chemical Functionalization of Plasmonic Surface Biosensors: A Tutorial REview on Issues, Strategies, and Costs. ACS Appl. Mater. Interfaces 2017, 35, 29394–29411. [Google Scholar] [CrossRef]
- Steglich, P.; Hulsemann, M.; Dietzel, B.; Mai, A. Optical Biosensors Based on Silicon on Insulater Ring Resonators: A Review. Molecules 2019, 24, 519. [Google Scholar] [CrossRef] [PubMed]
- Damborský, P.; Švitel, J.; Katrlík, J. Optical biosensors. Essays Biochem. 2016, 60, 91–100. [Google Scholar] [CrossRef] [PubMed]
- Qui, G.; Gai, Z.; Tao, Y.; Schmitt, J.; Kullak-Ublick, G.A.; Wang, J. Dual-Functional Plasmonic Photothermal Biosensors for Highly Accurate Severe Acute Respiratory Syndrome Coronavirus 2 Detection. ACS Nano 2020, 14, 5268–5277. [Google Scholar]
- Raman, R.; Patel, K.J.; Ranjan, K. COVID-19: Unmasking Emerging SARS-CoV-2 Variants, Vaccines and Therapeutic Strategies. Biomolecules 2021, 11, 993. [Google Scholar] [CrossRef]
- Mattoli, A.I.; Hassan, A.; Oliveira, O.N., Jr.; Crespilho, F.N. On the challenges for the diagnosis of SARS-CoV-2 Based on a Review on Current Methodologies. ACS Sens. 2020, 5, 3655–3677. [Google Scholar] [CrossRef]
- Pohanka, M. Overview of Piezoelectric Biosensors, Immunosensors and DNA Sensors and Their Applications. Materials 2018, 11, 448. [Google Scholar] [CrossRef]
- Chen, Y.; Qian, C.; Liu, C.; Shen, H.; Wang, Z.; Ping, J.; Wu, J.; Chen, H. Nucleic Acid amplification free biosensors for pathogen detection. Biosens. Bioelectron. 2020, 153, 112049. [Google Scholar] [CrossRef]
- Kuma, S.; Woldemariam, M.M. Structural, Electronic, Lattice Dynamic, and Elastic Properties of SnTiO3 and PbTiO3 Using Density Functional Theory. Adv. Condens. Matter Phys. 2019, 2019, 3176148. [Google Scholar] [CrossRef]
- Ravina, A.D.; Mohan, H.; Prasad, M.; Pundir, C.S. Detection methods for influenza A H1N1 virus with special reference to biosensors: A review. Biosci. Rep. 2020, 40, BSR20193852. [Google Scholar] [CrossRef]
- Pirich, C.L.; de Freitas, R.A.; Torresi, R.M.; Picheth, G.F.; Sierakowski, M.R. Piezoelectric immunochip coated with thin films of bacterial cellulose nanocrystal for dengue detection. Biosens. Bioelectron. 2017, 92, 47. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Zhao, Z.; Wang, B.; Qian, Z. The Design of a Frame-Like ZnO FBAR Sensor For Achieving Unifrom Mass Sensitivity Distribution. Sensors 2020, 20, 2408. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Guo, X.; Liu, R.; Guo, J.; Zhang, Y.; Zhang, W.; Sang, S. Real-time and label-free detection of VKORC1 genes based on a magneto elastic biosensor for warfarin therapy. J. Mater. Chem. B 2020, 8, 6271–6276. [Google Scholar] [CrossRef]
- Imran, S.; Ahmadi, S.; Kerman, K. Electrochemical Biosensors for the Detection of SARS-CoV-2 and Other Viruses. Micromachines 2021, 12, 174. [Google Scholar] [CrossRef] [PubMed]
- Antiochia, R. Developments in biosensors for CoV detection and future trends. Biosens. Bioelectron. 2021, 173, 112777. [Google Scholar] [CrossRef] [PubMed]
- Hampitak, P.; Jowitt, T.A.; Melendrez, D.; Fresquet, M.; Hamilton, P.; Iliut, M.; Nie, K.; Spencer, B.; Lennon, R.; Vijayaraghavan, A. A Point-of-Care Immunosensor Based on a Quartz Crystal Microbalance with Graphene Biointerface for Antibody Assay. ACS Sens. 2020, 5, 3520–3532. [Google Scholar] [CrossRef] [PubMed]
- Drobysh, M.; Ramanaviciene, A.; Viter, R.; Ramanavicius, A. Affinity Sensors for the Diagnosis of COVID-19. Micromachines 2021, 12, 390. [Google Scholar] [CrossRef]
- Nunna, B.B.; Zhuang, S.; Malave, I.; Lee, E.S. Ovarian Cancer Diagnosis using Micro Biochip. In Proceedings of the NIH-IEEE 2015 Strategic Conference on Healthcare Innovations and Point-of-Care Technologies for Precision Medicine, (PCHT15-0056), Bethesda, MD, USA, 9–10 November 2015. [Google Scholar]
- Chaibun, T.; Puenpa, J.; Ngamdee, T.; Boonapatcharoen, N.; Athamanolap, P.; O’Mullane, A.P.; Vongpunsawad, S.; Poovorawan, Y.; Lee, S.Y.; Lertanantawong, B. Rapid electrochemical detection of coronavirus SARS-CoV-2. Nat. Commun. 2021, 12, 802. [Google Scholar] [CrossRef]
- Zhuang, S.; Nunna, B.B.; Boscoboinik, J.A.; Lee, E.S. Nitrogen-doped graphene catalysts: High energy wet ball milling synthesis and characterizations of functional groups and particle size variation with time and speed. Int. J. Energy Res. 2017, 41, 2535–2554. [Google Scholar] [CrossRef]
- Tripathy, S.; Singh, S.G. Label-Free Electrochemical Detection of DNA Hybridization: A Method for COVID-19 Diagnosis. Trans. Indian Natl. Acad. Eng. 2020, 5, 205–209. [Google Scholar] [CrossRef]
- Morales-Narvaez, E.; Dincer, C. The Impact of Biosensing in a pandemic outbreak: COVID-19. Biosens. Bioelectron. 2020, 163, 112274. [Google Scholar] [CrossRef]
- Singh, H.; Zhuang, S.; Nunna, B.B.; Lee, E.S. Morphology and Chemical Structure of Modified Nitrogen-Doped Graphene for Highly Active Oxygen Reduction Reactions. In Proceedings of the 48th Power Source Conference, Denver, CO, USA, 11–14 June 2018. [Google Scholar]
- Mahshid, S.S.; Flynn, S.E.; Mahshid, S. The potential application of electrochemical biosensor in the COVID-19 pandemic: A perspective on the rapid diagnostics of SARS-CoV-2. Biosens. Bioelectron. 2021, 176, 112905. [Google Scholar] [CrossRef]
- Zhuang, S.; Nunna, B.B.; Lei, L.; Lee, E.S. Synthesis of Nitrogen-doped Graphene Catalyst by Wet Ball Milling for Electrochemical Systems. In Proceedings of the 251st ACS National Meeting Exposition, San Diego, CA, USA, 13–17 March 2016. [Google Scholar]
- Mahmoudi, G.; Babashkina, M.; Maniukiewicz, W.; Afkhami, F.; Nunna, B.; Zubkov, F.; Ptaszek, A.; Szczepanik, D.; Mitoraj, M.; Safin, D. Solvent-Induced Formation of Novel Ni(II) Complexes Derived from Bis-Thiosemicarbazone Ligand: An Insight from Experimental and Theoretical Investigations. Int. J. Mol. Sci. 2021, 22, 5337. [Google Scholar] [CrossRef]
- Szunerits, S.; Pagneux, Q.; Swaidan, A.; Mishyn, V.; Roussel, A.; Cambillau, C.; Devos, D.; Engelmann, I.; Alidjinou, E.K.; Happy, H.; et al. The role of the surface ligand on the performance of electrochemical SARS-CoV-2 antigen biosensors. Anal. Bioanal. Chem. 2021, 1–11. [Google Scholar] [CrossRef]
- Rasmi, Y.; Li, X.; Khan, J.; Ozer, T.; Choi, J.R. Emerging point-of-care biosensors for rapid diagnosis of COVID-19: Current progress, challenges, and future prospects. Anal. Bioanal. Chem. 2021, 413, 4137–4159. [Google Scholar] [CrossRef] [PubMed]
- Jayamohan, H.; Lambert, C.J.; Sant, H.J.; Jafek, A.; Patel, D.; Feng, H.; Beeman, M.; Mahmood, T.; Nze, U.; Gale, B.K. SARS-CoV-2 pandemic: A review of molecular diagnostic tools including sample collection and commercial response with associated advantages and limitations. Anal. Bioanal. Chem. 2021, 413, 49–71. [Google Scholar] [CrossRef] [PubMed]
- Giri, B.; Pandey, S.; Shrestha, R.; Pokharel, K.; Ligler, F.S.; Neupane, B.B. Review of analytical performance of COVID-19 detection methods. Anal. Bioanal. Chem. 2021, 413, 35–48. [Google Scholar] [CrossRef] [PubMed]
- Lei, Y.-M.; Xiao, M.-M.; Li, Y.-T.; Xu, L.; Zhang, H.; Zhang, Z.-Y.; Zhang, G.-J. Detection of heart failure-related biomarker in whole blood with graphene field effect transistor biosensor. Biosens. Bioelectron. 2017, 91, 1–7. [Google Scholar] [CrossRef]
- Orooji, Y.; Sohrabi, H.; Hemmat, N.; Oroojalian, F.; Baradaran, B.; Mokhtarzadeh, A.; Mohaghegh, M.; Karimi-Maleh, H. An Overview on SARS-CoV-2 (COVID-19) and Other Human Coronaviruses and Their Detection Capability via Amplification Assay, Chemical Sensing, Biosensing, Immunosensing, and Clinical Assays. Nano-Micro Lett. 2021, 13, 18. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Mao, H.; Wu, C.; Tang, L.; Wu, Z.; Sun, H.; Zhang, H.; Zhou, H.; Jia, C.; Jin, Q.; et al. Label-free graphene biosensor targeting cancer molecules based on non-covalent modification. Biosens. Bioelectron. 2017, 87, 701–707. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Lee, G.-H.; Kim, S.Y.; Kwon, S.Y.; Kim, H.-R.; Park, S. From Diagnosis to Treatment: Recent Advances in Patient-Friendly Biosensors and Implantable Devices. ACS Nano 2021, 15, 1960–2004. [Google Scholar] [CrossRef] [PubMed]
- Shetti, N.P.; Mishra, A.; Bukkitgar, S.D.; Basu, S.; Narang, J.; Reddy, K.R.; Aminabhavi, T.M. Conventional and Nanotechnology-Based Sensing Methods for SARS Coronavirus (2019-nCoV). ACS Appl. Bio Mater. 2021, 4, 1178–1190. [Google Scholar] [CrossRef] [PubMed]
- Yousefi, H.; Mahmud, A.; Chang, D.; Das, J.; Gomis, S.; Chen, J.B.; Wang, H.; Been, T.; Yip, L.; Coomes, E.; et al. Detection of SARS-CoV-2 Viral Particles Using Direct, Reagent-Free Electrochemical Sensing. J. Am. Chem. Soc. 2021, 143, 1722–1727. [Google Scholar] [CrossRef]
- Eissa, S.; Zourob, M. Development of a Low-Cost Cotton-Tipped Electrochemical Immunosensor for the Detection of SARS-CoV-2. Anal. Chem. 2021, 93, 1826–1833. [Google Scholar] [CrossRef]
- Zhuang, S.; Lei, L.; Nunna, B.; Lee, E.S. New Nitrogen-Doped Graphene/MOF-modified catalyst for Fuel Cell Systems. ECS Trans. 2016, 72, 149–154. [Google Scholar] [CrossRef]
- Sharafeldin, M.; Davis, J.J. Point of Care Sensors for Infectious Pathogens. Anal. Chem. 2021, 93, 184–197. [Google Scholar] [CrossRef]
- Sengupta, J.; Hussain, C. Graphene based field effect transistor biosensors for the rapid detection and analysis of viruses: A perspective in view of COVID-19. Carbon Trends 2020, 2, 100011. [Google Scholar] [CrossRef]
- Huang, H.; Su, S.; Wu, N.; Wan, H.; Wan, S.; Bi, H.; Sun, L. Graphene-Based Sensors for Human Health Monitoring. Front. Chem. 2019, 7, 399. [Google Scholar] [CrossRef]
- Islam, A.E.; Martineau, R.; Crasto, C.M.; Kim, H.; Rao, R.S.; Maruyama, B.; Kim, S.S.; Drummy, L.F. Graphene-Based Electrolyte-Gated Field-Effect Transistors for Potentiometrically Sensing Neuropeptide Y in Physiologically Relevant Environments. ACS Appl. Nano Mater. 2020, 3, 5088–5097. [Google Scholar] [CrossRef]
- Narita, F.; Wang, Z.; Kurita, H.; Li, Z.; Shi, Y.; Jia, Y.; Soutis, C. A review of Piezoelectric and Magnetostrictive Biosensor Materials for Detection of COVID-19 and Other Viruses. Adv. Mater. 2021, 33, 2065448. [Google Scholar] [CrossRef] [PubMed]
- Cumulative Cases. Available online: https://coronavirus.jhu.edu/data/cumulative-cases (accessed on 5 January 2021).
- Seo, G.; Lee, G.; Kim, M.J.; Baek, S.-H.; Choi, M.; Ku, K.B.; Lee, C.-S.; Jun, S.; Park, D.; Kim, H.G.; et al. Rapid Detection of COVID-19 Causative Virus (SARS-CoV-2) in Human Nasopharyngeal Swab Specimens Using Field-Effect Transistor-Based Biosensor. ACS Nano 2020, 14, 5135–5142. [Google Scholar] [CrossRef] [PubMed]
- Andryukov, B.G. Six decades of lateral flow immunoassay: From determining metabolic markers to diagnosing COVID-19. AIMS Microbiol. 2020, 6, 280–304. [Google Scholar] [CrossRef]
- Anfossi, L.; Di Nardo, F.; Cavalera, S.; Giovannoli, C.; Baggiani, C. Multiplex Lateral Flow Immunoassay: An Overview of Strategies towards High-throughput Point-of-Need Testing. Biosensors 2018, 9, 2. [Google Scholar] [CrossRef]
- Urusov, A.E.; Zherdev, A.V.; Dzantiev, B.B. Towards Lateral Flow Quantitative Assays: Detection Approaches. Biosensors 2019, 9, 89. [Google Scholar] [CrossRef] [PubMed]
- You, M.; Lin, M.; Gong, Y.; Wang, S.; Li, A.; Ji, L.; Zhao, H.; Ling, K.; Wen, T.; Huang, Y.; et al. Household Fluorescent Lateral Flow Strip Platform for Sensitive and Quantitative Prognosis of Heart Failure Using Dual-Color Upconversion Nanoparticles. ACS Nano 2017, 11, 6261–6270. [Google Scholar] [CrossRef] [PubMed]
- Foysal, K.H.; Seo, S.E.; Kim, M.J.; Kwon, O.S.; Chong, J.W. Analyte Quantity Detection from Lateral Flow Assay Using a Smartphone. Sensors 2019, 19, 4812. [Google Scholar] [CrossRef] [PubMed]
- Candel, F.J.; Viñuela-Prieto, J.M.; González Del Castillo, J.; Garcia, P.B.; Saavedra, M.F.; Piriz, A.H.; Virumbrales, D.J.; Lebrato, J.C.; de Casasola, G.G.; Prieto, R.G.; et al. Utility of lateral flow tests in SARS-CoV-2 infection monitorization. Rev. Esp. Quimioter. 2020, 33, 258–266. [Google Scholar] [CrossRef]
- Crozier, A.; Rajan, S.; Buchan, I.; McKee, M. Put to the test: Use of rapid testing technologies for COVID-19. BMJ 2021, 372, n208. [Google Scholar] [CrossRef] [PubMed]
- Lateral Flow Immunoassay. Available online: https://med.unr.edu/ddl/technology/lateral-flow-immunoassay (accessed on 29 April 2021).
- Buda, N.; Grau, E.S.; Cylwik, J.; Welnicki, M. Lung Ultrasound in the diagnosis of COVID-19 infection—A case series and review of the literature. Adv. Med. Sci. 2020, 2, 318–385. [Google Scholar] [CrossRef]
- Bernheim, A.; Mei, X.; Huang, M.; Yang, Y.; Fayad, Z.A.; Zhang, N.; Diao, K.; Lin, B.; Zhu, X.; Li, K.; et al. Chest CT Findings in Coronavirus Disease-19 (COVID-19): Relationship to Duration of Infection. Radiology 2020, 295, 200463. [Google Scholar] [CrossRef] [PubMed]
- Mayo, P.H.; Copetti, R.; Feller-Kopman, D.; Mathis, G.; Maury, E.; Mongodi, S. Thoracic ultrasonography: A narrative review. Intensive Care Med. 2019, 45, 1200–1211. [Google Scholar] [CrossRef]
- Jackson, K.; Butler, R.; Aujayeb, A. Lung ultrasound in the COVID-19 pandemic. Postgrad. Med. J. 2021, 97, 34–39. [Google Scholar] [CrossRef] [PubMed]
- de Alencar, J.C.G.; Marchini, J.F.M.; Marino, L.O.; Ribeiro, S.C.D.C.; Bueno, C.G.; da Cunha, V.P.; Neto, F.L.; Neto, R.A.B.; Souza, H.P. Lung ultrasound score predicts outcomes in COVID-19 patients admitted to the emergency department. Ann. Intensiv. Care 2021, 11, 1–8. [Google Scholar] [CrossRef]
- Ji, L.; Cao, C.; Gao, Y.; Zhang, W.; Xie, Y.; Duan, Y.; Kong, S.; You, M.; Ma, R.; Jiang, L.; et al. Prognostic value of bedside lung ultrasound score in patients with COVID-19. Crit. Care 2020, 24, 1–12. [Google Scholar] [CrossRef]
- Alam, N.A.; Ahsan, M.; Based, A.; Haider, J.; Kowalski, M. COVID-19 Detection from Chest X-Ray Images Using Feature Fusion and Deep Learning. Sensors 2021, 21, 1480. [Google Scholar] [CrossRef]
- Jacobi, A.; Chung, M.; Bernheim, A.; Eber, C. Portable chest X-ray in coronavirus disease-19 (COVID-19): A pictorial review. Clin. Imaging 2020, 64, 35–42. [Google Scholar] [CrossRef]
- Chest X-rays. Available online: https://www.mayoclinic.org/tests-procedures/chest-x-rays/about/pac-20393494 (accessed on 22 May 2021).
- Ng, M.-Y.; Lee, E.Y.P.; Yang, J.; Yang, F.; Li, X.; Wang, H.; Lui, M.M.-S.; Lo, C.S.-Y.; Leung, B.; Khong, P.-L.; et al. Imaging Profile of the COVID-19 Infection: Radiologic Findings and Literature Review. Radiol. Cardiothorac. Imaging 2020, 2, e200034. [Google Scholar] [CrossRef]
- Available online: https://www.dartmouth.edu/anatomy/HAE/Thorax/chestwall/radiology/CXRintro.html (accessed on 29 April 2021).
- Wang, L.; Lin, Z.Q.; Wong, A. COVID-Net: A tailored deep convolutional neural network design for detection of COVID-19 cases from chest X-ray images. Sci. Rep. 2020, 10, 19549. [Google Scholar] [CrossRef] [PubMed]
- Gietema, H.A.; Zelis, N.; Nobel, J.M.; Lambriks, L.J.G.; Van Alphen, L.B.; Lashof, A.M.L.O.; Wildberger, J.E.; Nelissen, I.C.; Stassen, P.M. CT in relation to RT-PCR in diagnosing COVID-19 in The Netherlands: A prospective study. PLoS ONE 2020, 15, e0235844. [Google Scholar] [CrossRef]
- Zhu, Y.; Liu, Y.L.; Li, Z.P.; Kuang, J.Y.; Li, X.M.; Yang, Y.Y.; Feng, S.T. Clinical and CT imaging features of 2019 novel coronavirus disease (COVID-19). J. Infect. 2020. [Google Scholar] [CrossRef]
- Kevadiya, B.D.; Machhi, J.; Herskovitz, J.; Oleynikov, M.D.; Blomberg, W.R.; Bajwa, N.; Soni, D.; Das, S.; Hasan, M.; Patel, M.; et al. Diagnostics for SARS-CoV-2 infections. Nat. Mater. 2021, 20, 593–605. [Google Scholar] [CrossRef]
- Chua, F.; Armstrong-James, D.; Desai, S.R.; Barnett, J.; Kouranos, V.; Kon, O.M.; Jose, R.; Vancheeswaran, R.; Loebinger, M.R.; Wong, J.; et al. The role of CT in case ascertainment and management of COVID-19 pneumonia in the UK: Insights from high-incidence regions. Lancet Respir. Med. 2020, 8, 438–440. [Google Scholar] [CrossRef]
- Zhao, W.; Zhong, Z.; Xie, X.; Yu, Q.; Liu, J. Relation Between Chest CT Findings and Clinical Conditions of Coronavirus Disease (COVID-19) Pneumonia: A Multicenter Study. AJR Am. J. Roentgenol. 2020, 214, 1072–1077. [Google Scholar] [CrossRef]
- Rahaman, M.M.; Li, C.; Yao, Y.; Kulwa, F.; Rahman, M.A.; Wang, Q.; Qi, S.; Kong, F.; Zhu, X.; Zhao, X. Identification of COVID-19 samples from chest X-ray images using deep learning: A comparison of transfer approaches. J. X-ray Sci. Technol. 2020, 28, 821–839. [Google Scholar] [CrossRef]
- Xie, X.; Zhong, Z.; Zhao, W.; Zheng, C.; Wang, F.; Liu, J. Chest CT for Typical Coronavirus Disease 2019 (COVID-19) Pneumonia: Relationship to Negative RT-PCR Testing. Radiology 2020, 296, E41–E45. [Google Scholar] [CrossRef]
- Zhao, L.; Yu, K.; Zhao, Q.; Tian, R.; Xie, H.; Xie, L.; Deng, P.; Xie, G.; Bao, A.; Du, J. Lung Ultrasound score in evaluating the severity of coronavirus disease 2019 (COVID-19) Pneumonia. Ultrasound Med. Biol. 2020, 46, 2938–2944. [Google Scholar] [CrossRef] [PubMed]
- Cleverley, J.; Piper, J.; Jones, M.M. The role of Chest Radiography in confimring COVID-19 Pneumonia. BMJ 2020, 28, 821–839. [Google Scholar] [CrossRef]
- Kwee, C.T.; Kwee, M.R. Chest CT in COVID-19 What the Radiologist needs to know. Radiographics 2020, 40, 7. [Google Scholar] [CrossRef]
- Nunna, B.B.; Mandal, D.; Lee, J.U.; Zhuang, S.; Lee, E.S. Sensitivity Study of Cancer Antigens (CA-125) Detection Using Interdigitated Electrodes under Microfluidic Flow Condition. BioNano Sci. 2019, 9, 203–214. [Google Scholar] [CrossRef]
- Mandal, D.; Nunna, B.B.; Zhuang, S.; Lee, E.S. Carbon Nanotubes Based Biosensor for Detection of Cancer Antigens (CA-125) Under Shear Flow Condition. Nano-Struct. Nano-Objects 2017, 15, 180–185. [Google Scholar] [CrossRef]
- Nunna, B.B.; Mandal, D.; Zhuang, S.; Lee, E.S. Innovative Point-of-Care (POC) Micro Biochip for Early Stage Ovarian Cancer Diagnostics. Sens. Transducers J. 2017, 214, 12–20. [Google Scholar]
- Okba, N.; Muller, M.A.; Li, W.; Wang, C.; Kessel, C.H.; Corman, V.M.; Lamers, M.M.; Sikkema, R.S.; de Bruin, E.; Chandler, F.D.; et al. Severe Acute Respiratory Syndrome Coronavirus-2 Specific Antibody Responses in Coronavirus Disease Patients. Emerg. Infect. Dis. 2020, 26, 1478–1488. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Feng, Y.; Mo, X.; Zheng, P.; Wang, Q.; Li, P.; Peng, P.; Liu, X.; Chen, Z.; Huang, H.; et al. Kinetics of SARS-CoV-2 specific IgM and IgG responses in COVID-19 patients. Emerg. Microbes Infect. 2020, 9, 940–948. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Zhang, Z.; Chen, W.; Cai, Z.; Ge, X.; Zhu, H.; Jiang, T.; Tan, W.; Peng, Y. Predicting the receptor-binding domain usage of the coronavirus based on kmer frequency on spike protein. Infect. Genet. Evol. 2018, 61, 183–184. [Google Scholar] [CrossRef]
| Biomarkers | Normal Patient | Infected Patient |
|---|---|---|
| Serum Ferritin | 15.0–150.0 (ng/mL) | 452.9–1451.6 (ng/mL) |
| C reactive protein | 0.0–1.0 (ng/mL) | 20.9–103.2 (ng/mL) |
| Interleukin 2R | 223.0–710.0 (U/mL) | 528.5–1136.3 (U/mL) |
| Cytokines (IL-6) | 0.0–7.0 (pg/mL) | 7.9 (pg/mL) |
| D-Dimer | 0–0.243 (µg/mL) | 0.5 (µg/mL) |
| Serum Amyloid A (SAA) | 0–10 (mg/mL) | 108.4 (mg/mL) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Etienne, E.E.; Nunna, B.B.; Talukder, N.; Wang, Y.; Lee, E.S. COVID-19 Biomarkers and Advanced Sensing Technologies for Point-of-Care (POC) Diagnosis. Bioengineering 2021, 8, 98. https://doi.org/10.3390/bioengineering8070098
Etienne EE, Nunna BB, Talukder N, Wang Y, Lee ES. COVID-19 Biomarkers and Advanced Sensing Technologies for Point-of-Care (POC) Diagnosis. Bioengineering. 2021; 8(7):98. https://doi.org/10.3390/bioengineering8070098
Chicago/Turabian StyleEtienne, Ernst Emmanuel, Bharath Babu Nunna, Niladri Talukder, Yudong Wang, and Eon Soo Lee. 2021. "COVID-19 Biomarkers and Advanced Sensing Technologies for Point-of-Care (POC) Diagnosis" Bioengineering 8, no. 7: 98. https://doi.org/10.3390/bioengineering8070098
APA StyleEtienne, E. E., Nunna, B. B., Talukder, N., Wang, Y., & Lee, E. S. (2021). COVID-19 Biomarkers and Advanced Sensing Technologies for Point-of-Care (POC) Diagnosis. Bioengineering, 8(7), 98. https://doi.org/10.3390/bioengineering8070098







