Breeding Chlorophyll-Deficient Mutants of Chlorella vulgaris to Enhance Consumer Acceptance
Abstract
1. Introduction
2. Materials and Methods
2.1. Culture Conditions, Mutagenesis and Mutant Selection
2.2. UV Radiation
2.3. Screening and Strain Selection
2.4. Cultivation
2.5. Dry Matter Determination
2.6. Light Microscopy
2.7. Amino Acid Profile
2.8. Total Lipid
2.9. Pigment Analysis
2.10. Cryo Focus Ion Beam Scanning Electron Microscopy for Cryo Volume Electron Microscopy (cvEM)
2.11. Quantification of Lipid Volume to Cell Volume
2.12. Statistical Analysis
3. Results
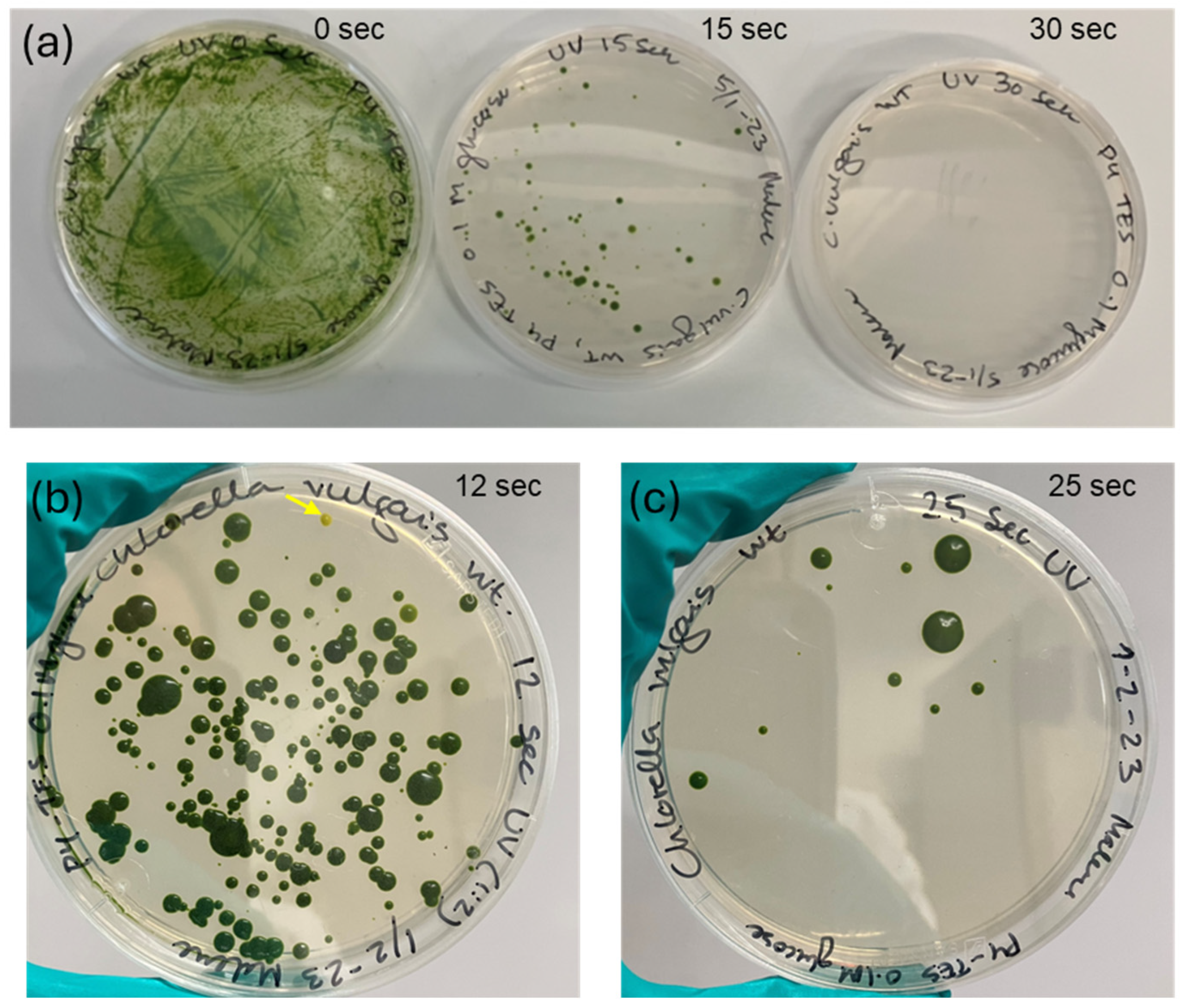
3.1. UV-Mutagenesis
3.2. Selection of Pigment-Deficient Mutants
3.3. Cultivations and Biomass Productivity
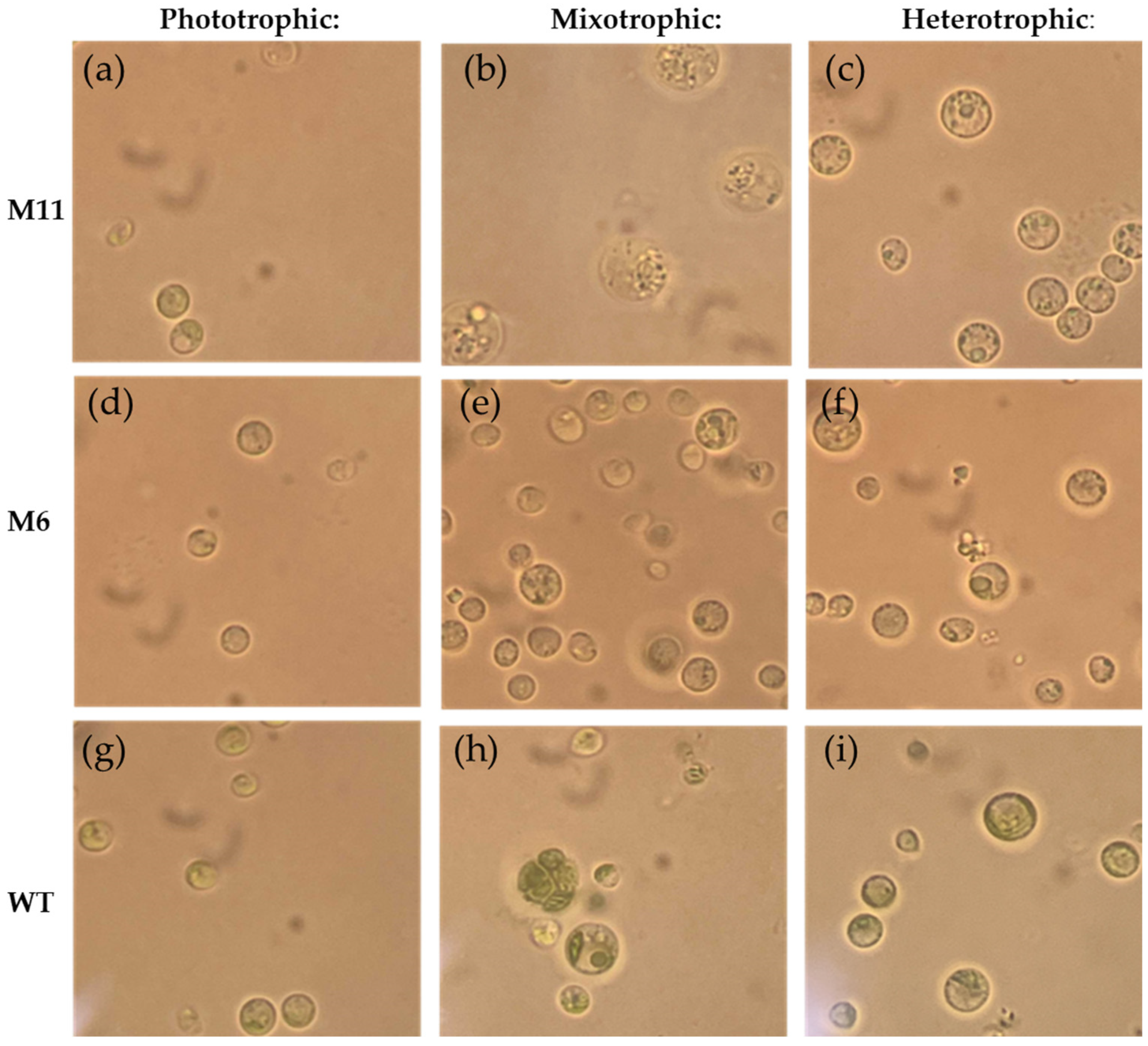
3.4. Visual Inspection Using Light Microscopy and Cell Size Measurement
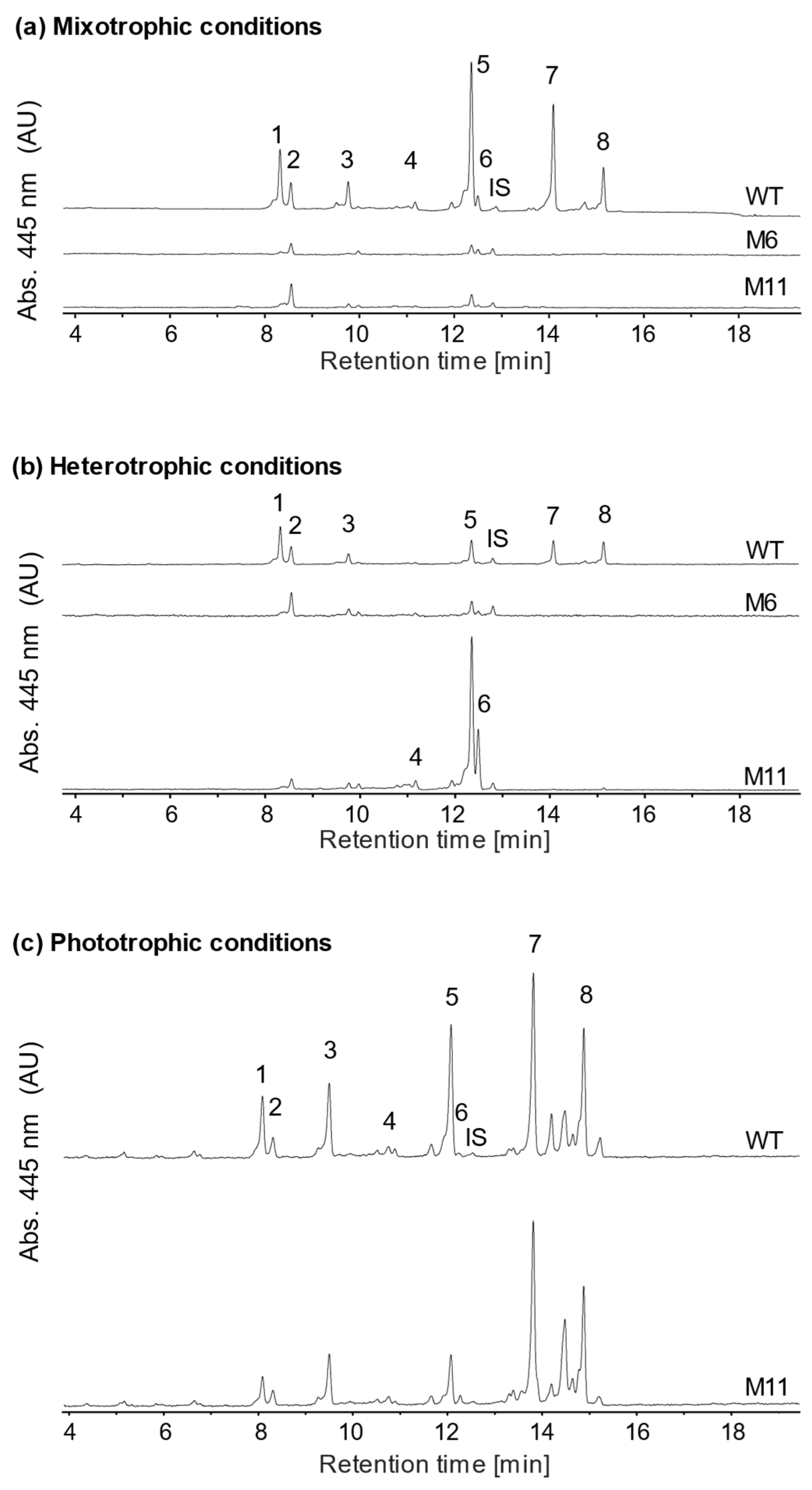
3.5. Analysis of Pigment Composition
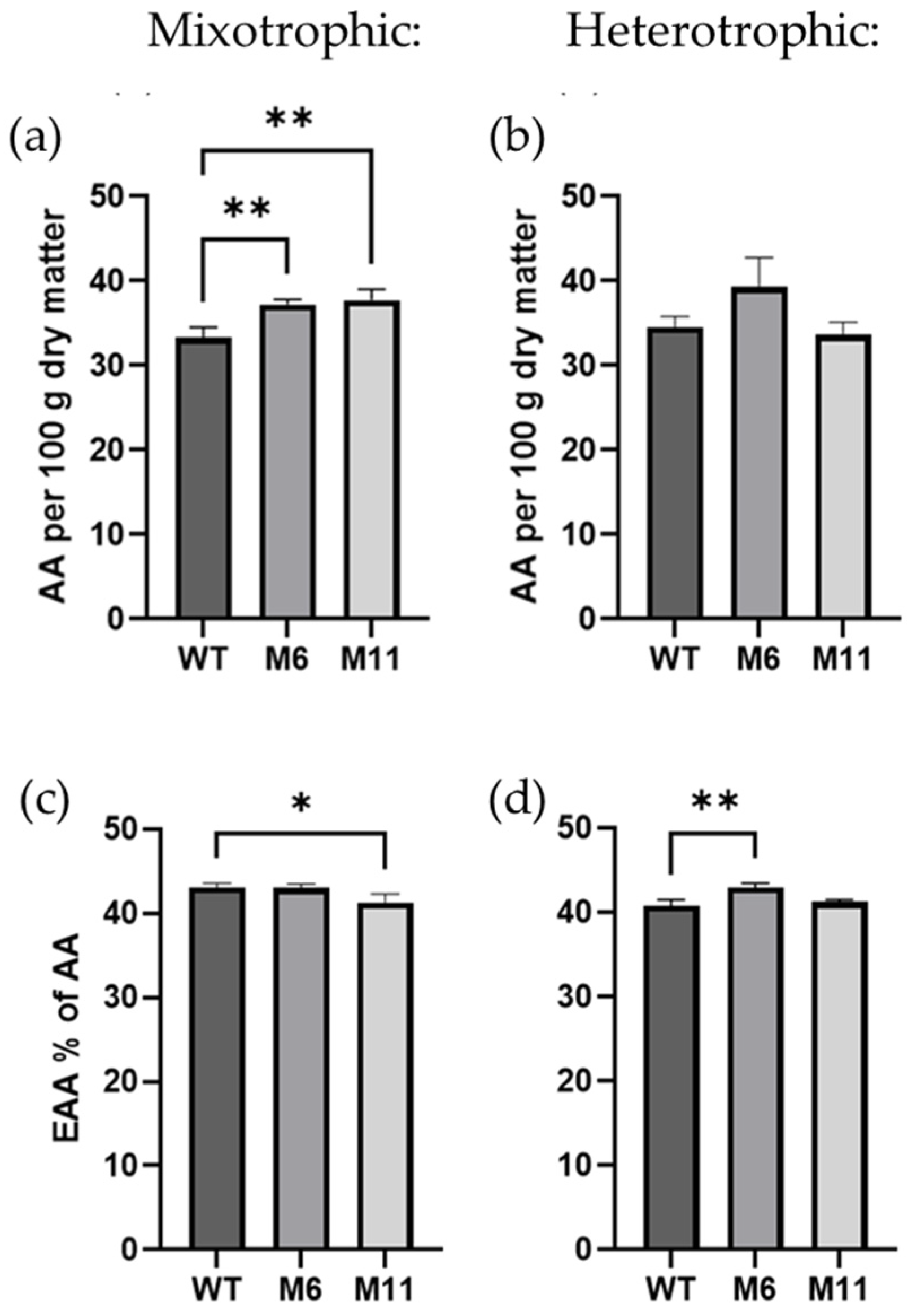
3.6. Amino Acid Content and Composition
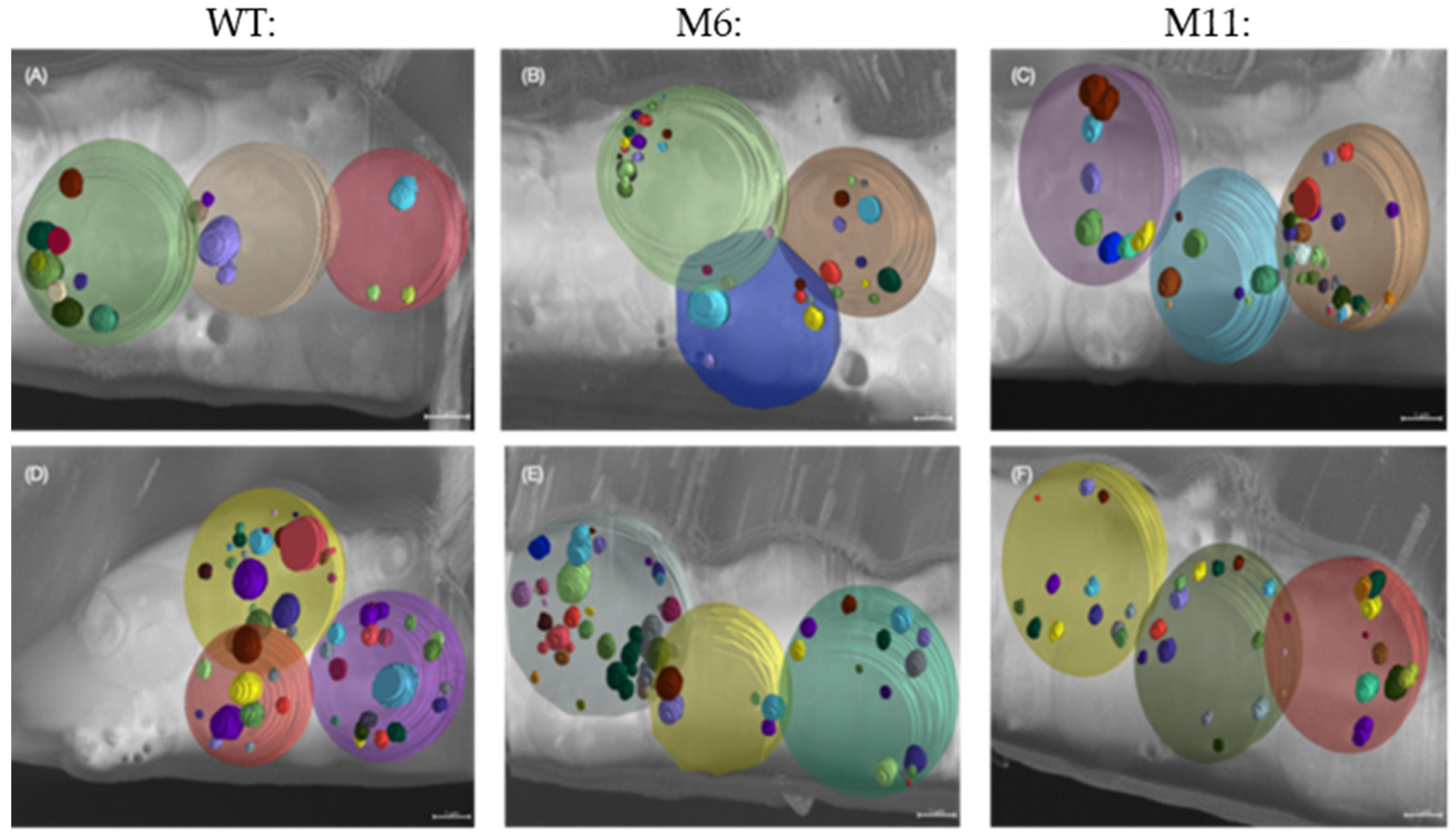
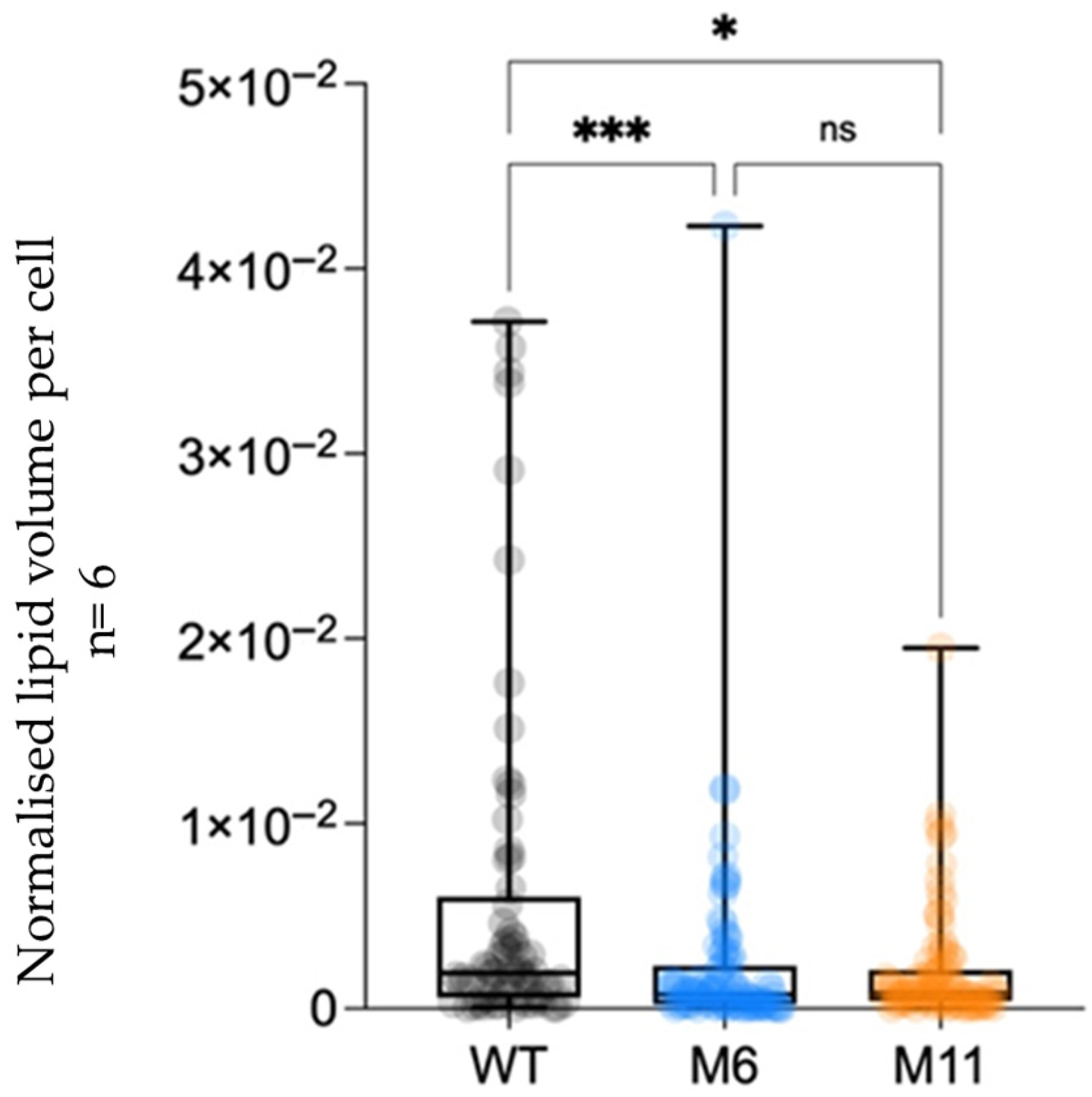
3.7. Analysis of Cellular Structures Using Cryo-Focused Ion Beam Scanning Microscopy (Cryo FIB-SEM)
4. Discussion
4.1. Ultrastructure of the C. vulgaris Cells
4.2. Pigment Content and Composition
4.3. Biomass Productivity
4.4. Biomass Composition
4.5. Limitations, Robustness, and Scalability
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AA | Amino Acids |
| AMST | Alignment to Median Smoothed Template |
| ANOVA | Analysis of Variance |
| CCAP | Culture Collection of Algae and Protozoa |
| cvEM | Cryo Volume Electron Microscopy |
| Cryo FIB-SEM | Cryo Focused Ion Beam Scanning Electron Microscopy |
| EAA | Essential Amino Acids |
| EMS | Ethyl Methane Sulphonate |
| ESI | Electrospray Ionization |
| HPLC | High Performance Liquid Chromatography |
| LC-MS | Liquid Chromatography–Mass Spectrometry |
| OD750 | Optical Density at 750 nm |
| PVDF | Polyvinylidene Fluoride |
| ROS | Reactive Oxygen Species |
| UHPLC | Ultra High Performance Liquid Chromatography |
| UVC | Ultraviolet C |
| CVDE | Violaxanthin De-epoxidase |
| WT | Wild Type |
| ZE | Zeaxanthin Epoxidase |
Appendix A. Recipe for the Modified P4-TES Medium Applied in This Study
References
- Canelli, G.; Tarnutzer, C.; Carpine, R.; Neutsch, L.; Bolten, C.J.; Dionisi, F.; Mathys, A. Biochemical and Nutritional Evaluation of Chlorella and Auxenochlorella Biomasses Relevant for Food Application. Front. Nutr. 2020, 7, 565996. [Google Scholar] [CrossRef]
- Draaisma, R.B.; Wijffels, R.H.; Slegers, P.; Brentner, L.B.; Roy, A.; Barbosa, M.J. Food Commodities from Microalgae. Curr. Opin. Biotechnol. 2013, 24, 169–177. [Google Scholar] [CrossRef]
- Becker, E.W. Microalgae for Human and Animal Nutrition. In Handbook of Microalgal Culture: Applied Phycology and Biotechnology; John Wiley & Sons: Hoboken, NJ, USA, 2013; pp. 461–503. [Google Scholar]
- Barbosa, M.J.; Janssen, M.; Südfeld, C.; D’Adamo, S.; Wijffels, R.H. Hypes, Hopes, and the Way Forward for Microalgal Biotechnology. Trends Biotechnol. 2023, 41, 452–471. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.D.; Vasconcelos, V. Legal Aspects of Microalgae in the European Food Sector. Foods 2023, 13, 124. [Google Scholar] [CrossRef] [PubMed]
- Abu-Ghosh, S.; Dubinsky, Z.; Verdelho, V.; Iluz, D. Unconventional High-Value Products from Microalgae: A Review. Bioresour. Technol. 2021, 329, 124895. [Google Scholar] [CrossRef] [PubMed]
- Weinrich, R.; Elshiewy, O. A Cross-Country Analysis of How Food-Related Lifestyles Impact Consumers’ Attitudes towards Microalgae Consumption. Algal Res. 2023, 70, 102999. [Google Scholar] [CrossRef]
- Olsen, M.L.; Olsen, K.; Jensen, P.E. Consumer Acceptance of Microalgae as a Novel Food—Where are we now? And how to get further. Physiol. Plant. 2024, 176, e14337. [Google Scholar] [CrossRef]
- Falcão, R.L.; Pinheiro, V.; Ribeiro, C.; Sousa, I.; Raymundo, A.; Nunes, M.C. Nutritional Improvement of Fresh Cheese with Microalga Chlorella vulgaris: Impact on Composition, Structure and Sensory Acceptance. Food Technol. Biotechnol. 2023, 61, 259–270. [Google Scholar] [CrossRef]
- Lafarga, T. Effect of Microalgal Biomass Incorporation into Foods: Nutritional and sensorial attributes of the end products. Algal Res. 2019, 41, 101566. [Google Scholar] [CrossRef]
- Fernández, F.G.A.; Reis, A.; Wijffels, R.H.; Barbosa, M.; Verdelho, V.; Llamas, B. The Role of Microalgae in the Bioeconomy. New Biotechnol. 2021, 61, 99–107. [Google Scholar] [CrossRef]
- Shin, W.-S.; Lee, B.; Jeong, B.; Chang, Y.K.; Kwon, J.-H. Truncated Light-Harvesting Chlorophyll Antenna Size in Chlorella vulgaris Improves Biomass Productivity. J. Appl. Phycol. 2016, 28, 3193–3202. [Google Scholar] [CrossRef]
- Abreu, A.P.; Morais, R.C.; Teixeira, J.A.; Nunes, J. A Comparison Between Microalgal Autotrophic Growth and Metabolite Accumulation with Heterotrophic, Mixotrophic and Photoheterotrophic Cultivation Modes. Renew. Sustain. Energy Rev. 2022, 159, 112247. [Google Scholar] [CrossRef]
- Ruiz, J.; Wijffels, R.H.; Dominguez, M.; Barbosa, M.J. Heterotrophic vs Autotrophic Production of Microalgae: Bringing Some Light into the Everlasting Cost Controversy. Algal Res. 2022, 64, 102698. [Google Scholar] [CrossRef]
- Milrad, Y.; Mosebach, L.; Buchert, F. Regulation of Microalgal Photosynthetic Electron Transfer. Plants 2024, 13, 2103. [Google Scholar] [CrossRef]
- Willows, R.D. Biosynthesis of Chlorophyll and Bilins in Algae. In Photosynthesis in Algae: Biochemical and Physiological Mechanisms; Springer: Berlin/Heidelberg, Germany, 2020; pp. 83–103. [Google Scholar]
- Xu, X.; Li, J.; Ma, Y.; Zhang, L.; Wang, Y.; Chen, Z. Effect of Chlorophyll on Lipid Oxidation of Rapeseed Oil. Eur. J. Lipid Sci. Technol. 2018, 120, 1800078. [Google Scholar] [CrossRef]
- Matusiak, M.; Radecka, H.; Siger, A.; Nogala-Kalucka, M. The effect of chlorophyll dyes on the oxidation of photosensitized rapeseed oil. Zywność Nauka Technol. Jakość 2021, 28, 126–372. [Google Scholar] [CrossRef]
- Zhang, Q.; Li, W.; Zhou, X.; Chen, H.; He, Q.; Li, F.; Wang, B. Comparative evaluation of sensory and instrumental flavor profiles of four edible microalgae: Spirulina platensis, Chlorella pyrenoidosa, Chlamydomonas reinhardtii, and Haematococcus pluvialis. Algal Res. 2024, 76, 103628. [Google Scholar] [CrossRef]
- Liu, H.; Yang, Z.; Chen, Y.; Feng, J.; Wei, Q.; Zhou, X.; Wen, Y. Metabolome and ROAV analysis reveal the effective flavor improvement in the golden biomass of chlorophyll-deficient mutant of Auxenochlorella pyrenoidosa. Food Biosci. 2024, 57, 103891. [Google Scholar] [CrossRef]
- Liu, Y.; Chen, X.; Wei, D.; Xing, X. Breeding a novel chlorophyll-deficient mutant of Auxenochlorella pyrenoidosa for high-quality protein production by atmospheric room temperature plasma mutagenesis. Bioresour. Technol. 2023, 384, 129907. [Google Scholar] [CrossRef]
- Trovão, M.; Schüler, L.M.; Machado, A.; Bombo, G.; Navalho, S.; Barros, A.; Pereira, H.; Silva, J.; Freitas, F.; Varela, J. Random Mutagenesis as a Promising Tool for Microalgal Strain Improvement towards Industrial Production. Mar. Drugs 2022, 20, 440. [Google Scholar] [CrossRef]
- Ruiz, J.; Olivieri, G.; de Vree, J.; Bosma, R.; Willems, P.; Reith, J.H.; Eppink, M.H.M.; Kleinegris, D.M.M.; Wijffels, H.; Barbosa, M.J. Towards Industrial Products from Microalgae. Energy Environ. Sci. 2016, 9, 3036–3043. [Google Scholar] [CrossRef]
- Schüler, L.; Greque de Morais, E.; Trovão, M.; Machado, A.; Carvalho, B.; Carneiro, M.; Maia, I.; Soares, M.; Duarte, P.; Barros, A.; et al. Isolation and Characterization of Novel Chlorella vulgaris Mutants with Low Chlorophyll and Improved Protein Contents for Food Applications. Front. Bioeng. Biotechnol. 2020, 8, 469. [Google Scholar] [CrossRef] [PubMed]
- Jian, J.; Torres-Montero, P.; Bai, M.; Lu, Y.; Simonsen, H.T. Genome Analysis of Chlorella vulgaris (CCAP 211/12) mutants provided insight into the molecular basis of chlorophyll deficiency. Algal Res. 2024, 78, 103426. [Google Scholar] [CrossRef]
- Grahl, S.; Strack, M.; Mensching, A.; Mörlein, D. Alternative Protein Sources in Western Diets: Food product development and consumer acceptance of spirulina-filled pasta. Food Qual. Prefer. 2020, 84, 103933. [Google Scholar] [CrossRef]
- Siegrist, M.; Hartmann, C. Consumer Acceptance of Novel Food Technologies. Nat. Food 2020, 1, 343–350. [Google Scholar] [CrossRef]
- Rozin, P.; Vollmecke, T.A. Food Likes and Dislikes. Annu. Rev. Nutr. 1986, 6, 433–456. [Google Scholar] [CrossRef]
- Cvetkovska, M.; Orgnero, S.; Hüner, N.P.A.; Smith, D.R. The Enigmatic Loss of Light-Independent Chlorophyll Biosynthesis from an Antarctic green alga in a light-limited environment. New Phytol. 2019, 222, 651–656. [Google Scholar] [CrossRef]
- Wu, Y.; Liao, W.; Dawuda, M.M.; Hu, L.; Yu, J. 5-Aminolevulinic acid (ALA) biosynthetic and metabolic pathways and its role in higher plants: A reviews. Plant Growth Regul. 2019, 87, 357–374. [Google Scholar] [CrossRef]
- Bearth, A.; Siegrist, M. “As long as it is not irradiated”—Influencing factors of US consumers’ acceptance of food irradiation. Food Qual. Prefer. 2019, 71, 141–148. [Google Scholar] [CrossRef]
- Marette, S.; Disdier, A.-C.; Beghin, J.C. A comparison of EU and US consumers’ willingness to pay for gene-edited food: Evidence from apples. Appetite 2021, 159, 105064. [Google Scholar] [CrossRef]
- Gong, Q.; Wang, J.; Zhao, W.; Liu, J.; Yang, S.; Gao, Y.; Mou, H.; Zhu, C.; Sun, H. Development of Chlorophyll-Deficient Chlorella pyrenoidosa Mutant using atmospheric and room temperature plasma mutagenesis. Bioresource Technol. 2025, 422, 132245. [Google Scholar] [CrossRef]
- Trovão, M.; Barros, A.; Machado, A.; Reis, A. Heterotrophic cultivation of Chlorella vulgaris yellow mutant on sidestreams: Medium formulation and process scale-up. J. Environ. Chem. Eng. 2025, 13, 115361. [Google Scholar] [CrossRef]
- Liu, S.; Zhao, Y.; Liu, L.; Ao, X.; Ma, L.; Wu, M.; Ma, F. Improving Cell Growth and Lipid Accumulation in Green Microalgae Chlorella sp. via UV Irradiation. Appl. Biochem. Biotechnol. 2015, 175, 3507–3518. [Google Scholar] [CrossRef] [PubMed]
- Borlaug, N.E. Contributions of Conventional Plant Breeding to Food Production. Science 1983, 219, 689–693. [Google Scholar] [CrossRef]
- Bradshaw, J.E. Plant Breeding: Past, Present and Future. Euphytica 2017, 213, 60. [Google Scholar] [CrossRef]
- Raus, M. Decontamination of Chlorella sp. Culture Using Antibiotics and antifungal cocktail treatment. J. Eng. Appl. Sci. 2016, 11, 104–109. [Google Scholar]
- Lippi, L.; Bähr, L.; Wüstenberg, A.; Wilde, A.; Steuer, R. Exploring the potential of high-density cultivation of cyanobacteria for the production of cyanophycin. Algal Res. 2018, 31, 363–366. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Gundersen, E.; Jacobsen, C. Cultivation of Nannochloropsis oceanica using food grade industrial side streams: Effect on growth and relative abundance of selected amino acids and unsaturated fatty acids. J. Biotechnol. 2025, 405, 249–253. [Google Scholar] [CrossRef]
- Bligh, E.G.; Dyer, W.J. A Rapid Method of Total Lipid Extraction and Purification. Can. J. Biochem. Physiol. 1959, 37, 911–917. [Google Scholar] [CrossRef]
- Ljubic, A.; Holdt, S.L.; Jakobsen, J.; Bysted, A.; Jacobsen, C. Fatty Acids, Carotenoids, and Tocopherols from Microalgae: Targeting the accumulation by manipulating the light during growth. J. Appl. Phycol. 2021, 33, 2783–2793. [Google Scholar] [CrossRef]
- Bijttebier, S.; D’Hondt, E.; Noten, B.; Hermans, N.; Apers, S.; Voorspoels, S. Ultra high performance liquid chromatography versus high performance liquid chromatography: Stationary phase selectivity for generic carotenoid screening. J. Chromatogr. A 2014, 1332, 46–56. [Google Scholar] [CrossRef] [PubMed]
- Kapishnikov, S.; Kobylynska, M.; Fleck, R.A.; Olsen, M.F.L.; Jensen, P.E.; Fahy, K.; Sheridan, P.; Fyans, W.; O’Reilly, F.; McEnroe, T. Sample Preparation for Correlative Light, Soft X-ray tomography, and cryo FIB-SEM imaging of biological cells. BIO Web Conf. 2024, 129, 19010. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An Open-Source Platform for Biological-Image Analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Hennies, J.; Lleti, J.M.S.; Schieber, N.L.; Templin, R.M.; Steyer, A.M.; Schwab, Y. AMST: Alignment to Median Smoothed Template for Focused Ion Beam Scanning Electron Microscopy Image Stacks. Sci. Rep. 2020, 10, 2004. [Google Scholar] [CrossRef]
- Lowe, D.G. Distinctive Image Features from Scale-Invariant Keypoints. Int. J. Comput. Vis. 2004, 60, 91–110. [Google Scholar] [CrossRef]
- Berg, S.; Kutra, D.; Kroeger, T.; Straehle, C.N.; Kausler, B.X.; Haubold, C.; Schiegg, M.; Ales, J.; Beier, T.; Rudy, M.; et al. Ilastik: Interactive Machine Learning for (Bio)image Analysis. Nat. Methods 2019, 16, 1226–1232. [Google Scholar] [CrossRef]
- Lafarga, T.; Acién-Fernández, F.G.; Castellari, M.; Villaró, S.; Bobo, G.; Aguiló-Aguayo, I. Effect of microalgae incorporation on the physicochemical, nutritional, and sensorial properties of an innovative broccoli soup. LWT 2019, 111, 167–174. [Google Scholar] [CrossRef]
- Meyer, M.T.; Whittaker, C.; Griffiths, H. The Algal Pyrenoid: Key unanswered questions. J. Exp. Bot. 2017, 68, 3739–3749. [Google Scholar] [CrossRef]
- He, S.; Crans, V.L.; Jonikas, M.C. The Pyrenoid: The Eukaryotic CO2-Concentrating Organelle. Plant Cell 2023, 35, 3236–3259. [Google Scholar] [CrossRef]
- Dall’Osto, L.; Bressan, M.; Bassi, R. Biogenesis of Light Harvesting Proteins. Biochim. Biophys. Acta Bioenerg. 2015, 1847, 861–871. [Google Scholar] [CrossRef] [PubMed]
- Perin, G.; Bellan, A.; Michelberger, T.; Lyska, D.; Wakao, S.; Niyogi, K.K.; Morosinotto, T. Modulation of xanthophyll cycle impacts biomass productivity in the marine microalga Nannochloropsis. Proc. Natl. Acad. Sci. USA 2023, 120, e2214119120. [Google Scholar] [CrossRef] [PubMed]
- Latowski, D.; Grzyb, J.; Strzałka, K. The xanthophyll cycle—Molecular mechanism and physiological significance. Acta Physiol. Plant. 2004, 26, 197–212. [Google Scholar] [CrossRef]
- Fernández-Marín, B.; Roach, T.; Verhoeven, A.; García-Plazaola, J.I. Shedding light on the dark side of xanthophyll cycles. New Phytol. 2021, 230, 1336–1344. [Google Scholar] [CrossRef]
- Hu, J.; Nagarajan, D.; Zhang, Q.; Chang, J.-S.; Lee, D.-J. Heterotrophic cultivation of microalgae for pigment production: A review. Biotechnol. Adv. 2018, 36, 54–67. [Google Scholar] [CrossRef]
- Stadnichuk, I.N.; Rakhimberdieva, M.G.; Bolychevtseva, Y.V.; Yurina, N.P.; Karapetyan, N.V.; Selyakh, I.O. Inhibition by glucose of chlorophyll a and phycocyanobilin biosynthesis in the unicellular red alga Galdieria partita at the stage of coproporphyrinogen III formation. Plant Sci. 1998, 136, 11–23. [Google Scholar] [CrossRef]
- Beale, S.I. Enzymes of Chlorophyll Biosynthesis. Photosynth. Res. 1999, 60, 43–73. [Google Scholar] [CrossRef]
- Azizi, S.; Bayat, B.; Tayebati, H.; Hashemi, A.; Pajoum Shariati, F. Nitrate and phosphate removal from treated wastewater by Chlorella vulgaris under various light regimes within membrane flat plate photobioreactor. Environ. Prog. Sustain. Energy 2021, 40, e13519. [Google Scholar] [CrossRef]
- Møller, I.M.; Jensen, P.E.; Hansson, A. Oxidative Modifications to Cellular Components in Plants. Annu. Rev. Plant Biol. 2007, 58, 459–481. [Google Scholar] [CrossRef]
- Olsen, M.F.L.; Pedersen, J.S.; Thomsen, S.T.; Martens, H.J.; Petersen, A.; Jensen, P.E. Outdoor cultivation of a novel isolate of the microalgae Scenedesmus sp. and the evaluation of its potential as a novel protein crop. Physiol. Plant. 2021, 173, 483–494. [Google Scholar] [CrossRef]
- Lourenço, S.O.; Barbarino, E.; Lavín, P.L.; Lanfer Marquez, U.M.; Aidar, E. Distribution of intracellular nitrogen in marine microalgae: Calculation of new nitrogen-to-protein conversion factors. Eur. J. Phycol. 2004, 39, 17–32. [Google Scholar] [CrossRef]
- Young, V.R.; El-Khoury, A.E. Human Amino Acid Requirements: A Re-Evaluation. Food Nutr. Bull. 1996, 17, 1–15. [Google Scholar] [CrossRef]
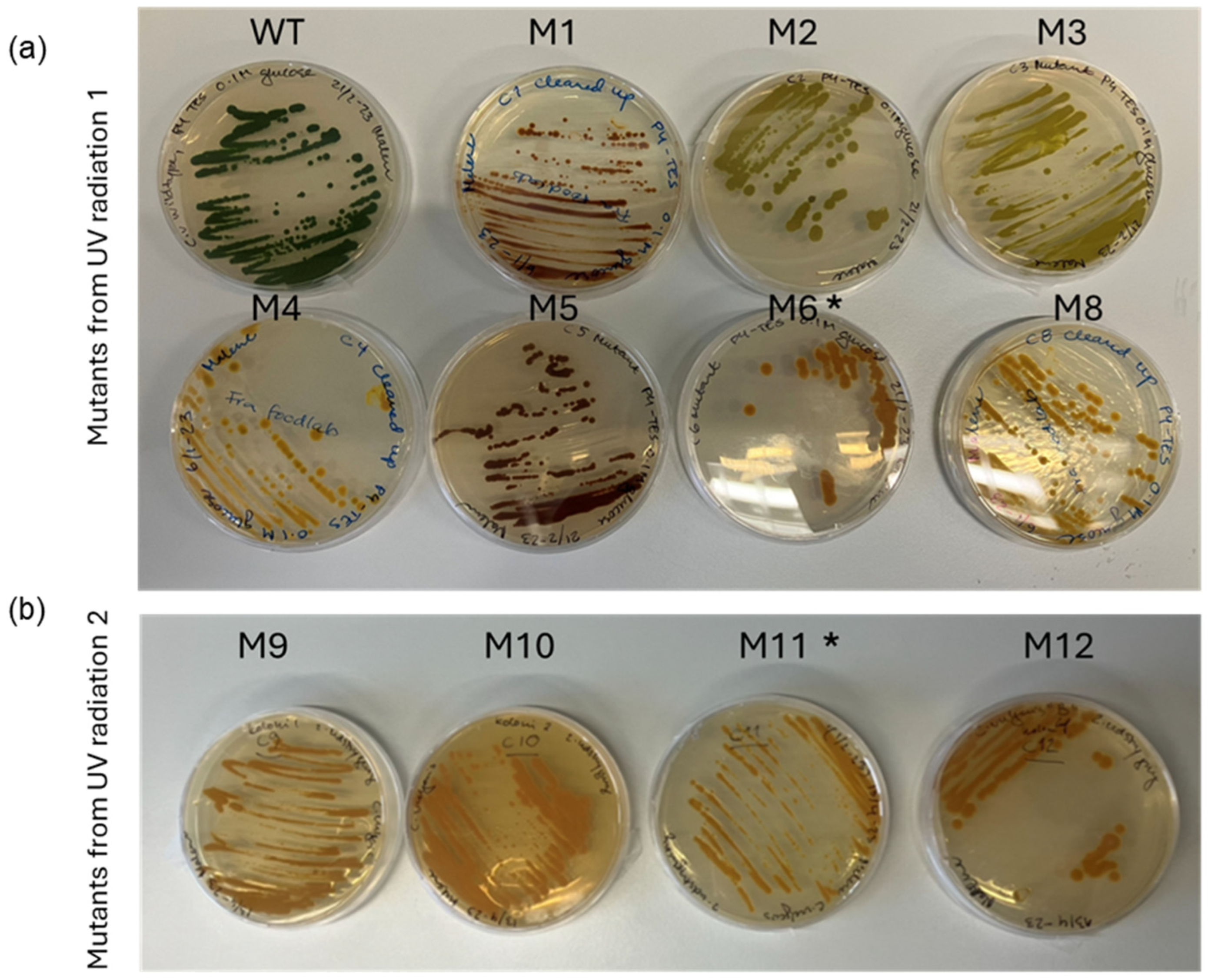



| Mutant | Mutagenesis | Colour, Heterotrophic |
|---|---|---|
| M1 | 1st trial | Orange/red |
| M2 | 1st trial | Lime green |
| M3 | 1st trial | Lime green |
| M4 | 1st trial | Yellow |
| M5 | 1st trial | Orange/red |
| M6 | 1st trial | Light orange |
| M7 | 1st trial | Almost white, very poor growth |
| M8 | 1st trial | Yellow |
| M9 | 2nd trial | Light green |
| M10 | 2nd trial | Orange, reverted to green |
| M11 | 2nd trial | Orange |
| M12 | 2nd trial | Orange |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Olsen, M.L.; Poveda-Huertes, D.; Ozcelik, D.; Gundersen, E.; Thøfner, J.F.B.; Kobylynska, M.; Marcotti, S.; Fleck, R.A.; McGrouther, D.; Andersen-Ranberg, J.; et al. Breeding Chlorophyll-Deficient Mutants of Chlorella vulgaris to Enhance Consumer Acceptance. Bioengineering 2026, 13, 318. https://doi.org/10.3390/bioengineering13030318
Olsen ML, Poveda-Huertes D, Ozcelik D, Gundersen E, Thøfner JFB, Kobylynska M, Marcotti S, Fleck RA, McGrouther D, Andersen-Ranberg J, et al. Breeding Chlorophyll-Deficient Mutants of Chlorella vulgaris to Enhance Consumer Acceptance. Bioengineering. 2026; 13(3):318. https://doi.org/10.3390/bioengineering13030318
Chicago/Turabian StyleOlsen, Malene Lihme, Daniel Poveda-Huertes, Duygu Ozcelik, Emil Gundersen, Jens Frederik Bang Thøfner, Maryna Kobylynska, Stefania Marcotti, Roland A. Fleck, Damien McGrouther, Johan Andersen-Ranberg, and et al. 2026. "Breeding Chlorophyll-Deficient Mutants of Chlorella vulgaris to Enhance Consumer Acceptance" Bioengineering 13, no. 3: 318. https://doi.org/10.3390/bioengineering13030318
APA StyleOlsen, M. L., Poveda-Huertes, D., Ozcelik, D., Gundersen, E., Thøfner, J. F. B., Kobylynska, M., Marcotti, S., Fleck, R. A., McGrouther, D., Andersen-Ranberg, J., Jacobsen, C., & Jensen, P. E. (2026). Breeding Chlorophyll-Deficient Mutants of Chlorella vulgaris to Enhance Consumer Acceptance. Bioengineering, 13(3), 318. https://doi.org/10.3390/bioengineering13030318











