Towards Feasible Home ECG Monitoring: AI-Driven Detection of Clinically Critical Arrhythmias Using Single-Lead Signals
Abstract
1. Introduction
1.1. Deep-Learning-Based ECG Classification
1.2. The Objective and Core Innovations of This Study
- We developed a lightweight architecture featuring a highly efficient Transformer model comprising approximately 0.7 M parameters (with a reduced model dimension d_model = 256). This architecture automates single-lead ECG analysis, bypassing the need for manual feature extraction processes, such as R-R interval calculation.
- The model achieves precise frequency distinction by accurately classifying cardiac rhythms that share similar P-QRS-T waveforms but operate at distinctly different heart rates, including normal sinus rhythm, ST, and SB.
- For advanced pathology identification, the framework effectively differentiates highly confusable conditions that fall within similar heart-rate ranges but possess fundamentally different pathological characteristics, most notably distinguishing between ST, SVT, and VT.
- To ensure the system is ready for wearable integration, the model was trained extensively on Lead I time-series data. Its low computational cost renders it exceptionally well-suited for real-world deployment in single-lead ECG patches and smartwatches.
2. Materials and Methods
2.1. Dataset Description
2.1.1. Database Specifications
2.1.2. ECG Selection and Labeling
2.1.3. Patient-Wise Data Partitioning
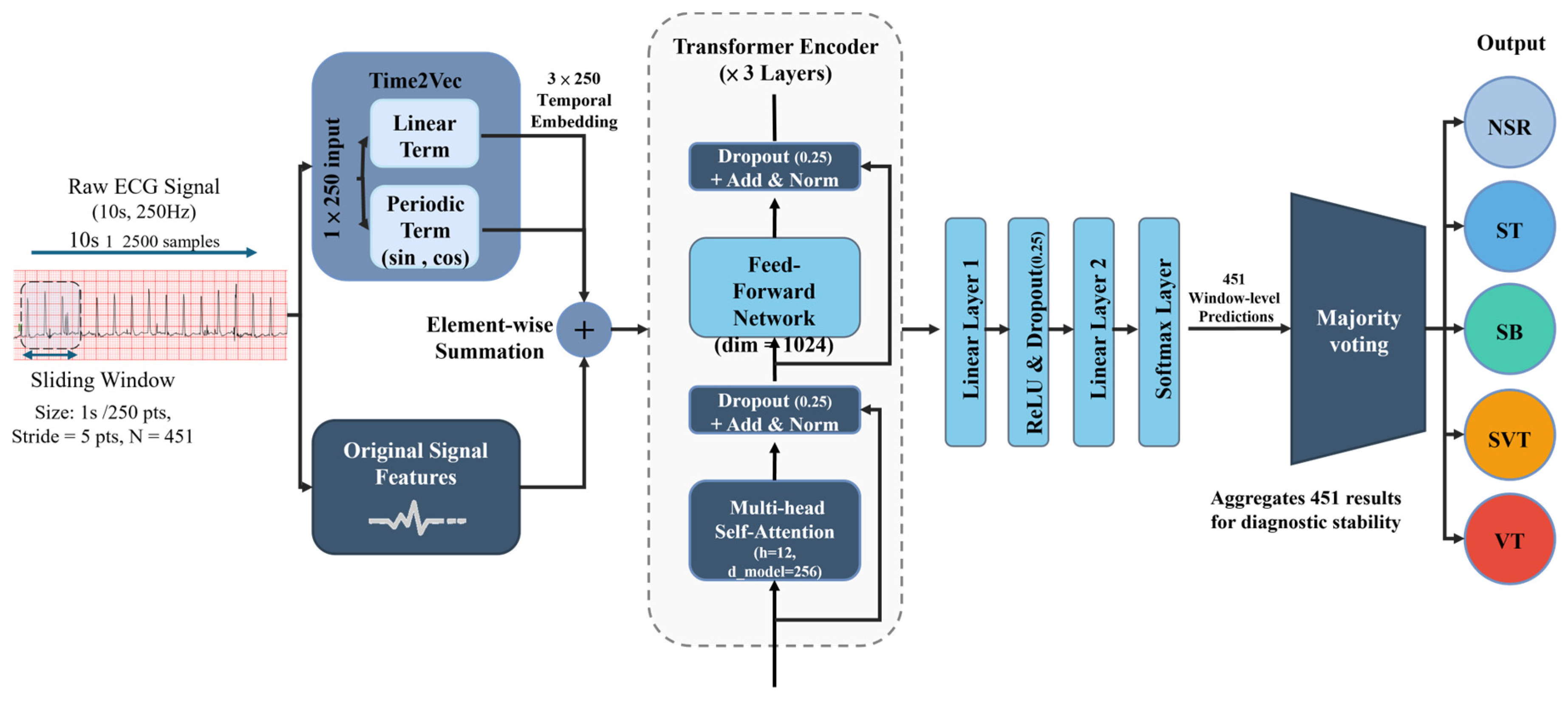
2.2. Model Architecture
2.2.1. Temporal Feature Embedding via Time2Vec
2.2.2. Transformer Encoder
2.2.3. Arrhythmia Classification Output Layer
2.2.4. Decision Mechanism: Majority Voting
2.3. Data Preprocessing
2.3.1. Signal Normalization
2.3.2. Input Layer Specifications
2.4. Model Performance Evaluation
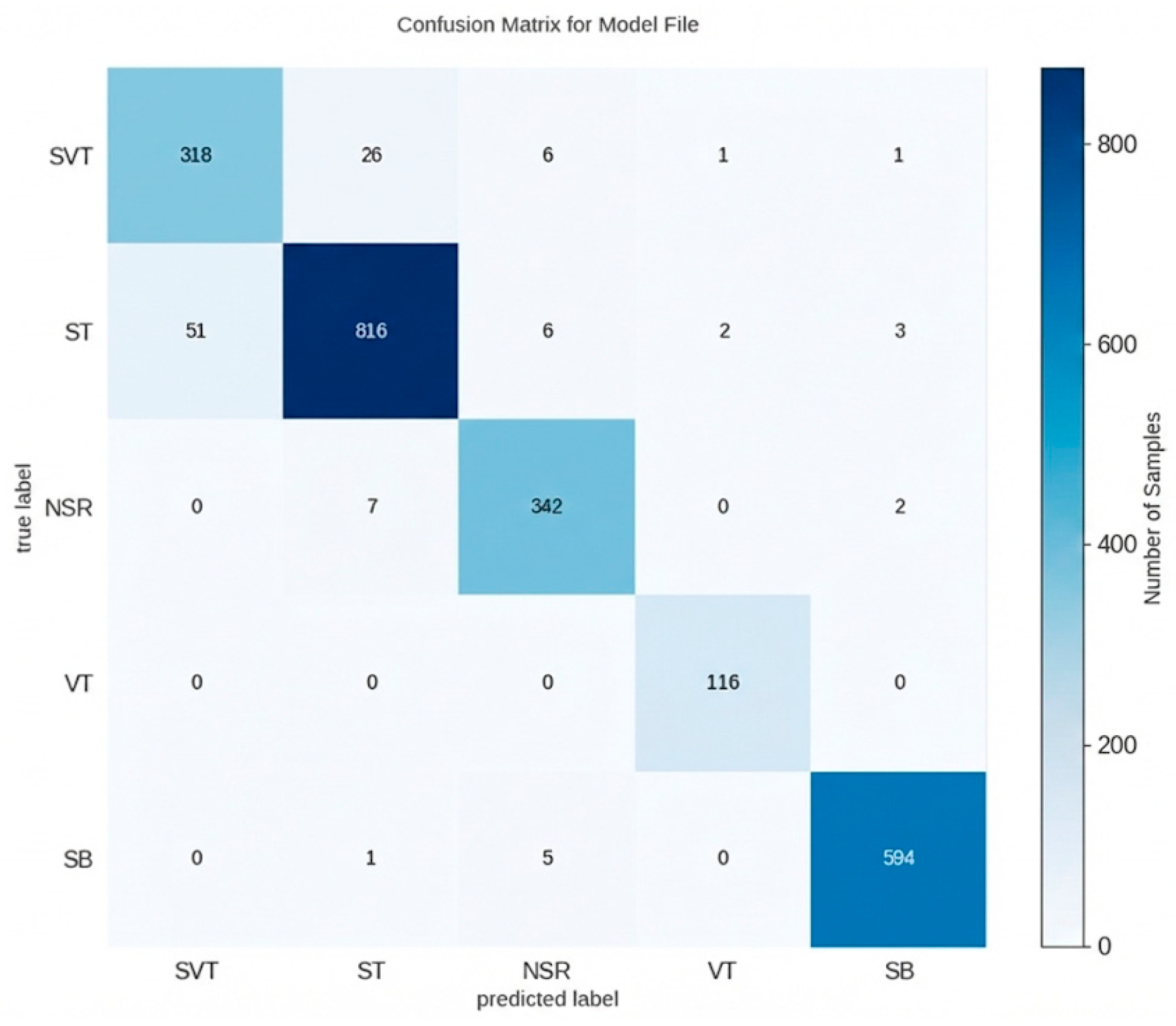
3. Results
4. Discussion
4.1. The Importance of Rhythm-Level Analysis in Clinical Diagnosis
4.2. Methodological Advantages over Traditional Analysis
4.3. Data Integration Strategy and Representativeness
4.4. Pathological Mechanisms and Interpretability Limitations
4.5. Computational Efficiency and Hardware Feasibility
4.6. Limitations and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AAMI | Association for the Advancement of Medical Instrumentation |
| AFib | Atrial Fibrillation |
| AF | Atrial Flutter |
| AI | Artificial Intelligence |
| AT | Atrial Tachycardia |
| AUC | Area Under the Curve |
| AVB | Atrioventricular Block |
| APC | Atrial Premature Contraction |
| BEiTs | BERT pre-training of image transformers |
| CD | Conduction Disturbance |
| CLSM | Central-towards LSTM Supportive Model |
| CNN | Convolutional Neural Network |
| CPSC | China Physiological Signal Challenge |
| DeiTs | Data-efficient image Transformers |
| DL | Deep Learning |
| DNN | Deep Neural Network |
| ECG | Electrocardiogram |
| FFNN | Feed-Forward Neural Network |
| FN | False Negative |
| FP | False Positive |
| Grad-CAM | Gradient-weighted Class Activation Mapping |
| GRU | Gated Recurrent Unit |
| HYP | Hypertrophy |
| IAVB | First-Degree Atrioventricular Block |
| LBBB | Left Bundle Branch Block |
| LSTM | Long Short-Term Memory |
| LVH | Left Ventricular Hypertrophy |
| MI | Myocardial Infarction |
| NSR | Normal Sinus Rhythm |
| PAC | Premature Atrial Contraction |
| PMI | Previous Myocardial Infarction |
| PVC | Premature Ventricular Contraction |
| QRS | QRS complex |
| QT | QT interval |
| RBBB | Right Bundle Branch Block |
| ROC | Receiver Operating Characteristic |
| SA | Sinus Arrhythmia |
| SB | Sinus Bradycardia |
| SR | Sinus Rhythm |
| ST | Sinus Tachycardia |
| STD | ST-segment Depression |
| STE | ST-segment Elevation |
| STTCs | ST-segment and T Wave Changes |
| SVT | Supraventricular Tachycardia |
| TN | True Negative |
| TP | True Positive |
| VFDB | Malignant Ventricular Ectopy Database |
| ViT | Vision Transformers |
| VPC | Ventricular Premature Contraction |
| VT | Ventricular Tachycardia |
References
- Sridhar, A.R.; Cheung, J.W.; Lampert, R.; Silva, J.N.; Gopinathannair, R.; Sotomonte, J.C.; Lakkireddy, D. State of the art of mobile health technologies use in clinical arrhythmia care. Commun. Med. 2024, 4, 218. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, A. Artificial intelligence enabled mobile health technologies in arrhythmias—An opinion article on recent findings. Front. Cardiovasc. Med. 2025, 12, 1548554. [Google Scholar] [CrossRef]
- Showrav, T.T.; Lincoln, S.I.; Hasan, M.K. EXGnet: A single-lead explainable-AI guided multiresolution network with train-only quantitative features for trustworthy ECG arrhythmia classification. arXiv 2025, arXiv:2506.12404. [Google Scholar] [CrossRef]
- Ezz, M. Deep learning-driven single-lead ECG classification: A rapid approach for comprehensive cardiac diagnostics. Diagnostics 2025, 15, 384. [Google Scholar] [CrossRef] [PubMed]
- Gliner, V.; Levy, I.; Tsutsui, K.; Acha, M.R.; Schliamser, J.; Schuster, A.; Yaniv, Y. Clinically meaningful interpretability of an AI model for ECG classification. npj Digit. Med. 2025, 8, 109. [Google Scholar] [CrossRef] [PubMed]
- Alghieth, M. DeepECG-Net: A hybrid transformer-based deep learning model for real-time ECG anomaly detection. Sci. Rep. 2025, 15, 20714. [Google Scholar] [CrossRef]
- Ikram, S.; Ikram, A.; Singh, H.; Ali Awan, M.D.; Naveed, S.; De la Torre Díez, I.; Candelaria Chio Montero, T. Transformer-based ECG classification for early detection of cardiac arrhythmias. Front. Med. 2025, 12, 1600855. [Google Scholar] [CrossRef]
- Yisimitila, T.; Wang, C.; Hou, M.; Maimaitiniyazi, M.; Aili, Z.; Liu, T.; Nijiati, M. Bridging clinical knowledge and AI: An interpretable transformer framework for ECG diagnosis. npj Digit. Med. 2025, 8, 120. [Google Scholar] [CrossRef]
- Li, Y.; Qian, R.; Li, K. Inter-patient arrhythmia classification with improved deep residual convolutional neural network. Comput. Methods Programs Biomed. 2022, 214, 106582. [Google Scholar] [CrossRef]
- Luo, K.; Li, J.; Wang, Z.; Celi, L.A. Patient-specific deep architectural model for ECG classification. J. Healthc. Eng. 2017, 2017, 4108720. [Google Scholar] [CrossRef]
- Alamr, A.; Artoli, A. Unsupervised transformer-based anomaly detection in ECG signals. Algorithms 2023, 16, 152. [Google Scholar] [CrossRef]
- Hu, R.; Chen, J.; Zhou, L. A transformer-based deep neural network for arrhythmia detection using continuous ECG signals. Comput. Biol. Med. 2022, 144, 105325. [Google Scholar] [CrossRef] [PubMed]
- Varghese, A.; Kamal, S.; Kurian, J. Transformer-based temporal sequence learners for arrhythmia classification. Med. Biol. Eng. Comput. 2023, 61, 1993–2000. [Google Scholar] [CrossRef] [PubMed]
- Yan, G.; Liang, S.; Zhang, Y.; Liu, F. Fusing transformer model with temporal features for ECG heartbeat classification. In Proceedings of the 2019 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), San Diego, CA, USA, 18–21 November 2019; pp. 898–905. [Google Scholar] [CrossRef]
- Cai, J.; Zhou, G.; Dong, M.; Wei, X.; Zhang, Y. Real-time arrhythmia classification algorithm using time-domain ECG feature based on FFNN and CNN. Math. Probl. Eng. 2021, 2021, 6648432. [Google Scholar] [CrossRef]
- Guo, L.; Sim, G.; Matuszewski, B. Inter-patient ECG classification with convolutional and recurrent neural networks. Biocybern. Biomed. Eng. 2019, 39, 568–579. [Google Scholar] [CrossRef]
- He, J.; Rong, J.; Sun, L.; Wang, H.; Zhang, Y.; Tian, J. An advanced two-step DNN-based framework for arrhythmia detection. In Advances in Knowledge Discovery and Data Mining, PAKDD 2020, Lecture Notes in Computer Science, Proceedings of the Pacific-Asia Conference on Knowledge Discovery and Data Mining, Singapore, 11–14 May 2020; Springer: Cham, Switzerland, 2020; Volume 12085, pp. 422–434. [Google Scholar] [CrossRef]
- Jangra, M.; Dhull, S.K.; Singh, K.K. O-WCNN: An optimized integration of spatial and spectral feature map for arrhythmia classification. Complex Intell. Syst. 2021, 7, 1–14. [Google Scholar] [CrossRef]
- Mousavi, S.; Afghah, F. Inter- and intra-patient ECG heartbeat classification for arrhythmia detection: A sequence to sequence deep learning approach. In Proceedings of the ICASSP 2019 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP), Brighton, UK, 12–17 May 2019; pp. 1308–1312. [Google Scholar] [CrossRef]
- Rizqyawan, M.I.; Siradj, Y.; Amri, M.F.; Pratama, R.P. Re-implementation of convolutional neural network for arrhythmia detection. Int. J. Adv. Sci. Eng. Inf. Technol. 2022, 12, 1319–1326. [Google Scholar] [CrossRef]
- Wang, T.; Lu, C.; Sun, Y.; Yang, M.; Liu, C. Automatic ECG classification using continuous wavelet transform and convolutional neural network. Entropy 2021, 23, 119. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, A.; Liang, D.; Chen, X.; Gao, M. Interpatient ECG heartbeat classification with an adversarial convolutional neural network. J. Healthc. Eng. 2021, 2021, 5522766. [Google Scholar] [CrossRef]
- Zubair, M.; Yoon, C. Cost-sensitive learning for anomaly detection in imbalanced ECG data using convolutional neural networks. Sensors 2022, 22, 4075. [Google Scholar] [CrossRef]
- Che, C.; Zhang, P.; Min, Z. Constrained transformer network for ECG signal processing and arrhythmia classification. BMC Med. Inform. Decis. Mak. 2021, 21, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Hao, C.Z.; Nugroho, H. A spatio-temporal approach with transformer network for heart disease classification with 12-lead electrocardiogram signals. In Lecture Notes in Electrical Engineering, Proceedings of the 6th International Conference on Electrical, Control and Computer Engineering, Pahang, Malaysia, 23 August 2021; Springer: Singapore, 2022; pp. 673–684. [Google Scholar] [CrossRef]
- Natarajan, A.; Chang, Y.; Mariani, S.; Rahman, A.; Boverman, G.; Vij, S.; Rubin, J. A wide and deep transformer neural network for 12-lead ECG classification. In Proceedings of the 2020 Computing in Cardiology, Rimini, Italy, 13–16 September 2020; pp. 1–4. [Google Scholar] [CrossRef]
- Apostol, A.; Nutu, M. Arrhythmia classification from 12-lead ECG signals using convolutional and transformer-based deep learning models. arXiv 2025, arXiv:2502.17887. [Google Scholar] [CrossRef]
- Ojha, J.; Haugerud, H.; Yazidi, A.; Lind, P.G. Exploring interpretable ai methods for ECG data classification. In Proceedings of the 5th ACM Workshop on Intelligent Cross-Data Analysis and Retrieval, Phuket, Thailand, 10 June 2024; pp. 11–18. [Google Scholar] [CrossRef]
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Mietus, J.E.; Moody, G.B.; Peng, C.-K.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a new research resource for complex physiologic signals. Circulation 2000, 101, e215–e220. [Google Scholar] [CrossRef]
- Perez Alday, E.A.; Gu, A.; Shah, A.J.; Robichaux, C.; Wong, A.K.; Liu, C.; Liu, F.; Rad, A.B.; Elola, A.; Seyedi, S.; et al. Classification of 12-lead ECGs: The PhysioNet/Computing in Cardiology Challenge 2020. Physiol. Meas. 2020, 41, 124003. [Google Scholar] [CrossRef]
- Greenwald, S.D. Development and Analysis of a Ventricular Fibrillation Detector. Master’s Thesis, Massachusetts Institute of Technology, Cambridge, MA, USA, 1986. [Google Scholar]
- Irani, H.; Metsis, V. Positional encoding in transformer-based time series models: A survey. arXiv 2025, arXiv:2502.12370. [Google Scholar] [CrossRef]
- Thwal, C.M.; Tun, Y.L.; Kim, K.; Park, S.B.; Hong, C.S. Transformers with Attentive Federated Aggregation for Time Series Stock Forecasting. In Proceedings of the 2023 International Conference on Information Networking (ICOIN), Bangkok, Thailand, 11–13 January 2023; pp. 499–504. [Google Scholar] [CrossRef]
- Yang, M.; Li, X.; Liu, Y. Sequence to point learning based on an attention neural network for nonintrusive load decomposition. Electronics 2021, 10, 1657. [Google Scholar] [CrossRef]
- Kazemi, S.M.; Goel, R.; Eghbali, S.; Ramanan, J.; Sahota, J.; Thakur, S.; Wu, S.; Smyth, C.; Poupart, P.; Brubaker, M. Time2Vec: Learning a vector representation of time. arXiv 2019, arXiv:1907.05321. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. In Advances in Neural Information Processing Systems; Guyon, I., Luxburg, U.V., Bengio, S., Wallach, H., Fergus, R., Vishwanathan, S., Garnett, R., Eds.; Curran Associates, Inc.: Long Beach, CA, USA, 2017; Volume 30. [Google Scholar] [CrossRef]
- Kavyashree, P.S.; El-Sharkawy, M. Compressed mobilenet v3: A light weight variant for resource-constrained platforms. In Proceedings of the 2021 IEEE 11th Annual Computing and Communication Workshop and Conference (CCWC), Online, 27–30 January 2021; pp. 0104–0107. [Google Scholar] [CrossRef]

| Data Source | Input Features | Network Architecture | Classes | Performance | References |
|---|---|---|---|---|---|
| MIT-BIH Arrhythmia Database | single QRS beat | PCA-based Transformer Encoder | 5 (Normal, APC, VPC, Fusion beat and Others) | Acc of all = 97.10% F1 = 95.00% | Ikram et al., 2025 [7] |
| MIT-BIH Arrhythmia Database & PhysioNet/Computing in Cardiology (CiC) Challenge 2017 | 10 s ECG sequences | DeepECG-Net (Hybrid CNN–Transformer) | 2 (Normal vs. Abnormal/Anomaly detection) | Acc of all = 98.30% Se = 96.50% Pre = 97.10% F1 = 96.50% | Alghieth 2025 [6] |
| PTB-XL, PTB Diagnostic, China 12-Lead, Georgia 12-Lead and St. Petersburg INCART | 2D grayscale ECG images (converted from 12-lead signals) | 2D Pre-trained Vision Transformers (ViT, BEiT, DeiT) | 5 (AF, IAVB, SB, NSR, ST) | ViT pretrained: Acc = 84.60% F1 = 0.84 BeiT_pretrained: Acc = 81.89% F1 = 0.79 DeiT pretrained: Acc = 84.60% F1 = 0.81 | Apostol and Nutu 2025 [27] |
| PhysioNet/Computing in Cardiology (CiC) Challenge 2021 (Chapman and Ningbo datasets) | 10 s ECG sequences | EXGnet (CNN + Multiresolution block + XAI guidance) | Chapman: 5 (SR, SB, ST, AF + AFib, SVT + AT) Ningbo: 5 (SR, SB, ST, AF + AFib, SA) | Chapman: Acc = 98.76% Se = 97.85% Spe = 99.70% F1 = 97.91% Ningbo: Acc = 96.93% Se = 95.40% Spe = 99.21% F1 = 95.53% | Showrav et al., 2025 [3] |
| A publicly available ECG Images Dataset (Mendeley Data) comprising 928 clinical ECG recordings from cardiac patients | 2D images of segmented single-lead ECG signals (Lead V4 was identified as the optimal diagnostic lead) | A comparative evaluation of deep CNN architectures, including VGG16, MobileNetV2, InceptionV3, DenseNet201, and NASNetLarge | 4 (Normal, Abnormal/Arrhythmia, MI and PMI) | VGG16: F1 = 98.11%, Prediction time = 4.2 ms MobileNetV2: F1 = 97.24%, Prediction time = 3.2 ms | Ezz 2025 [4] |
| MIT-BIH Arrhythmia Database & Chapman-Shaoxing dataset | 2D oscillographic representations (converted from 1D ECG signals via the OSC module) and auxiliary clinical features (including heart rate, QRS duration, ST segment changes, and QTc interval). | MDOT (Momentum Distillation Oscillographic Transformer) | MIT-BIH: 8 Chapman: 12 | MIT-BIH: Acc = 99.53% F1 = 97.26% Chapman: Acc = 99.03% F1 = 96.38% | Yisimitila et al., 2025 [8] |
| NYU dataset (clinical ECGs) and a dataset of scanned or photographed ECG images | Digitized 12-lead ECG images, including scanned ECG images and images obtained via mobile camera in clinical settings | ECG-AIO, an ensemble of deep learning models (specifically ResNet-18-based architectures) | 8 (AFib, AF, SB, ST, LBBB, LVH, PVC and RBBB) | Specific numerical metrics not reported; achieved high correlation with gold standard interpretations of 3 electrophysiologists. | Gliner et al., 2025 [5] |
| PhysioNet/Computing in Cardiology (CiC) Challenge 2020 | 10 s ECG sequences | ST-CNN-5: a deep Convolutional Neural Network (CNN) architecture focused on interpretable AI (XAI) using Grad-CAM | 5 (Normal, MI, STTC, CD and HYP) | Acc of all = 89.10% Pre of all = 79.80% Senof all = 69.30% Spe of all = 93.40% | Ojha et al., 2024 [28] |
| Arrhythmia Class | Estimated No. of Patients | No. of 10 s ECG Recordings | Training Dataset | Validation Dataset | Test Dataset |
|---|---|---|---|---|---|
| NSR | 751 | 751 | 300 | 100 | 351 |
| ST | 1278 | 1278 | 300 | 100 | 878 |
| SB | 1000 | 1000 | 300 | 100 | 600 |
| SVT (including AT) | 952 | 952 | 300 | 100 | 352 |
| VT | 16 | 516 | 300 | 100 | 116 |
| Total | 3997 | 4297 | 1500 | 500 | 2297 |
| Classification Performances of SVT, ST, NSR, VT and SB. | ||||||
|---|---|---|---|---|---|---|
| Accuracy | Sensitivity | Specificity | Precision | F1-Score | AUC | |
| SVT | 96.3% | 90.3% | 97.4% | 86.2% | 88.2% | 0.978 |
| ST | 95.8% | 92.9% | 97.6% | 96.0% | 94.4% | 0.987 |
| NSR | 98.9% | 97.4% | 99.1% | 95.3% | 96.3% | 0.997 |
| VT | 99.9% | 100.0% | 99.9% | 97.5% | 98.7% | 1.000 |
| SB | 99.5% | 99.0% | 99.6% | 99.0% | 99.0% | 0.999 |
| Total | 95.2% | - | - | - | - | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Hsu, C.-H.; Hsieh, J.-C.; Su, P.-Y.; Yang, C.-C. Towards Feasible Home ECG Monitoring: AI-Driven Detection of Clinically Critical Arrhythmias Using Single-Lead Signals. Bioengineering 2026, 13, 317. https://doi.org/10.3390/bioengineering13030317
Hsu C-H, Hsieh J-C, Su P-Y, Yang C-C. Towards Feasible Home ECG Monitoring: AI-Driven Detection of Clinically Critical Arrhythmias Using Single-Lead Signals. Bioengineering. 2026; 13(3):317. https://doi.org/10.3390/bioengineering13030317
Chicago/Turabian StyleHsu, Chia-Hsien, Jui-Chien Hsieh, Po-Yuan Su, and Chung-Chi Yang. 2026. "Towards Feasible Home ECG Monitoring: AI-Driven Detection of Clinically Critical Arrhythmias Using Single-Lead Signals" Bioengineering 13, no. 3: 317. https://doi.org/10.3390/bioengineering13030317
APA StyleHsu, C.-H., Hsieh, J.-C., Su, P.-Y., & Yang, C.-C. (2026). Towards Feasible Home ECG Monitoring: AI-Driven Detection of Clinically Critical Arrhythmias Using Single-Lead Signals. Bioengineering, 13(3), 317. https://doi.org/10.3390/bioengineering13030317







