4D Flow MRI at 0.6 T—Self-Gating Versus Camera-Based Respiratory Binning
Abstract
1. Introduction
2. Methods
2.1. Signal Considerations
2.2. Self-Gating
2.3. Camera Gating
2.4. Data Acquisition and Reconstruction
2.5. Data Analysis
2.5.1. Respiratory Analysis
2.5.2. 2D Analysis
2.5.3. 3D Analysis
3. Results
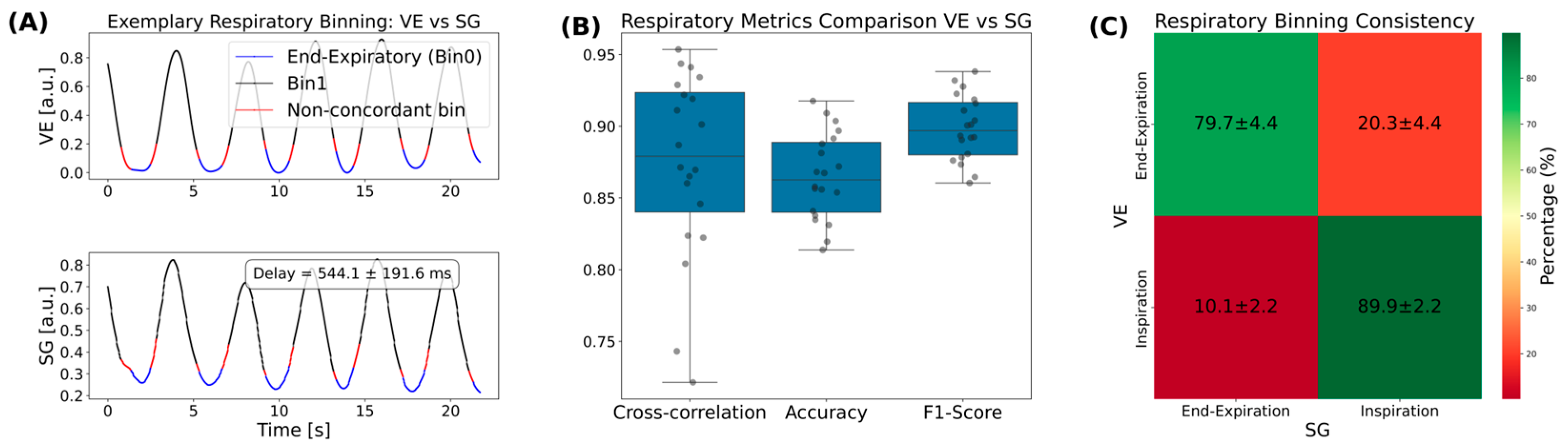
3.1. Respiratory Gating Analysis
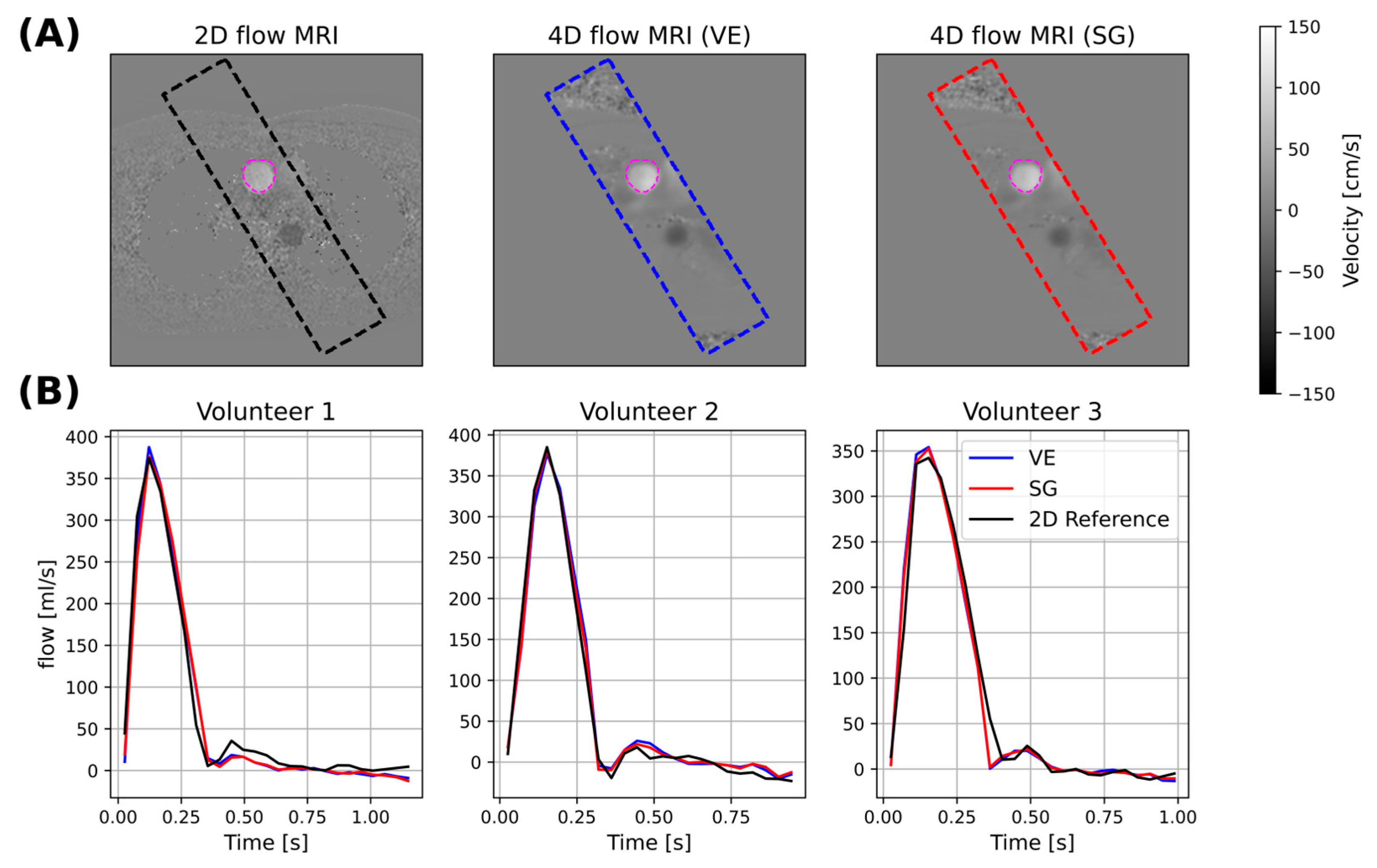
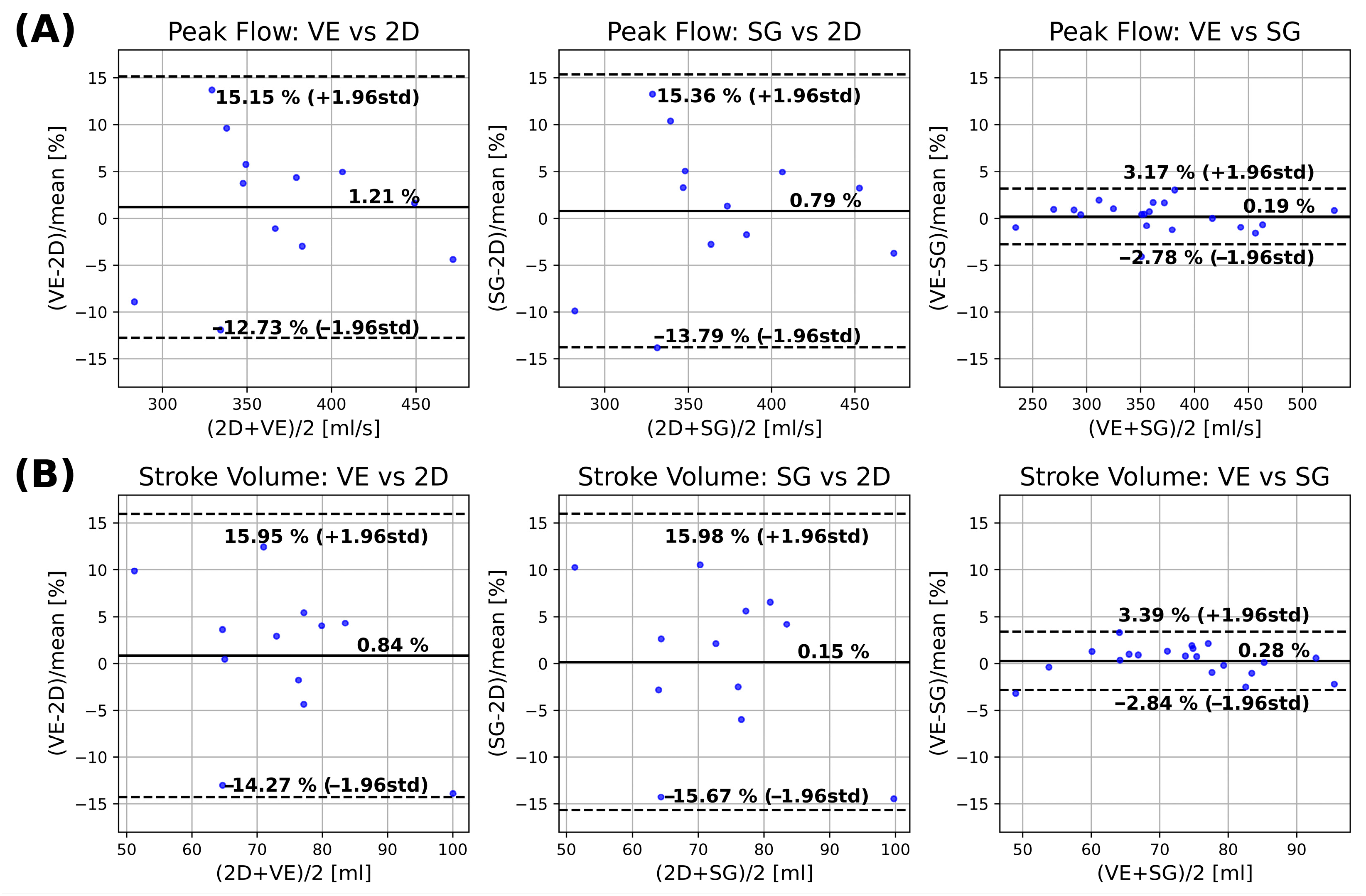
3.2. 2D Analysis
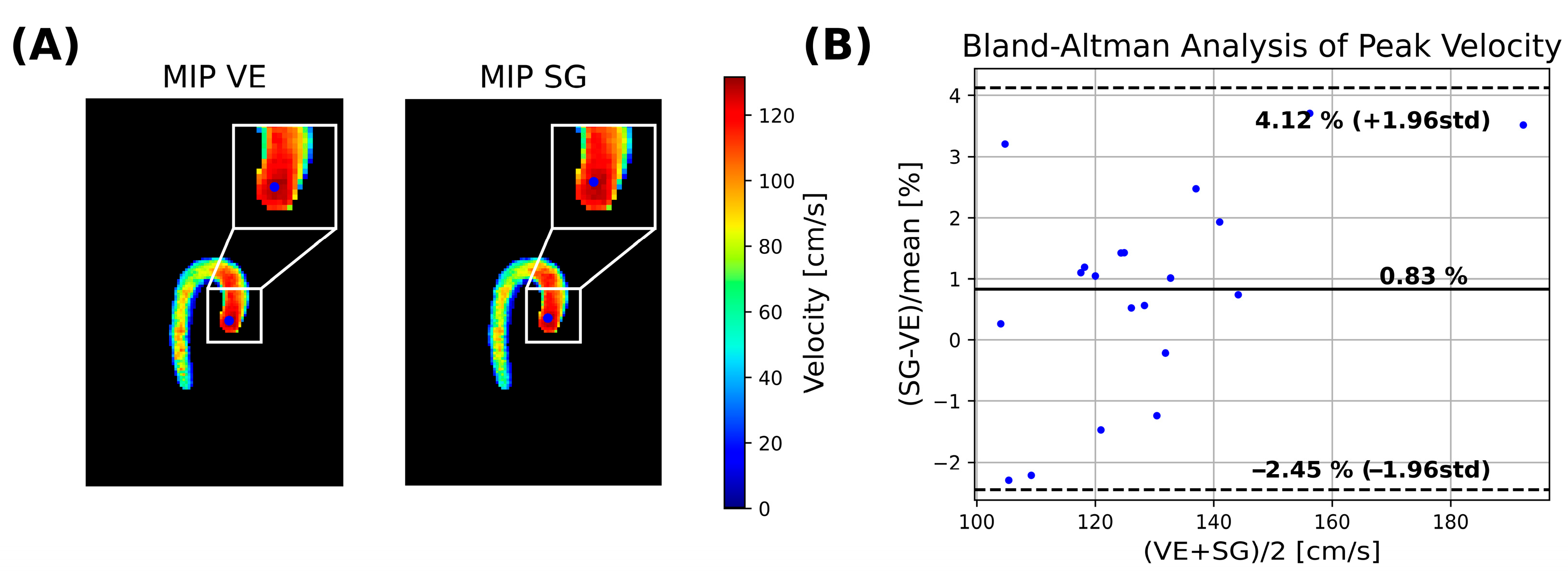
3.3. 3D Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| 1D | One-dimensional |
| 3D | Three-dimensional |
| 4D | Four-dimensional |
| 5D | Five-dimensional |
| RF | radiofrequency |
| MRI | Magnetic resonance imaging |
| VE | VitalEye |
| SG | Self-gating |
| EC | Eddy currents |
| PCA | Principal component analysis |
| SNR | Signal-to-noise ratio |
| CNR | Contrast-to-noise ratio |
| BMI | Body mass index |
| TR | Repetition time |
| TE | Echo time |
| SI | Superior-inferior |
| SOS | Sum-of-square |
| ECG | Electrocardiogram |
| FH | Feet-head |
| AP | Antero-posterior |
| RL | Right-left |
| VNR | Velocity-to-noise ratio |
| LoA | Limits of agreement |
| VDRad | Variable-Density and Radial view ordering |
| mDirrErr | Directional error |
| RMSE | Root-mean-square error |
| CMR | Cardiovascular magnetic resonance |
| bSSFP | Balanced steady-state free precession |
| PC | Phase-contrast |
References
- Markl, M.; Frydrychowicz, A.; Kozerke, S.; Hope, M.; Wieben, O. 4D Flow MRI. J. Magn. Reson. Imaging 2012, 36, 1015–1036. [Google Scholar] [CrossRef]
- Dyverfeldt, P.; Bissell, M.; Barker, A.J.; Bolger, A.F.; Carlhäll, C.-J.; Ebbers, T.; Francios, C.J.; Frydrychowicz, A.; Geiger, J.; Giese, D.; et al. 4D Flow Cardiovascular Magnetic Resonance Consensus Statement. J. Cardiovasc. Magn. Reson. 2015, 17, 72. [Google Scholar] [CrossRef]
- Dyverfeldt, P.; Ebbers, T. Comparison of Respiratory Motion Suppression Techniques for 4D Flow MRI. Magn. Reson. Med. 2017, 78, 1877–1882. [Google Scholar] [CrossRef]
- Santelli, C.; Nezafat, R.; Goddu, B.; Manning, W.J.; Smink, J.; Kozerke, S.; Peters, D.C. Respiratory Bellows Revisited for Motion Compensation: Preliminary Experience for Cardiovascular MR. Magn. Reson. Med. 2011, 65, 1097–1102. [Google Scholar] [CrossRef] [PubMed]
- Ehman, R.; McNamara, M.; Pallack, M.; Hricak, H.; Higgins, C. Magnetic Resonance Imaging with Respiratory Gating: Techniques and Advantages. Am. J. Roentgenol. 1984, 143, 1175–1182. [Google Scholar] [CrossRef] [PubMed]
- Bissell, M.M.; Raimondi, F.; Ait Ali, L.; Allen, B.D.; Barker, A.J.; Bolger, A.; Burris, N.; Carhäll, C.-J.; Collins, J.D.; Ebbers, T.; et al. 4D Flow Cardiovascular Magnetic Resonance Consensus Statement: 2023 Update. J. Cardiovasc. Magn. Reson. 2023, 25, 40. [Google Scholar] [CrossRef] [PubMed]
- Ehman, R.L.; Felmlee, J.P. Adaptive Technique for High-Definition MR Imaging of Moving Structures. Radiology 1989, 173, 255–263. [Google Scholar] [CrossRef]
- Danias, P.G.; McConnell, M.V.; Khasgiwala, V.C.; Chuang, M.L.; Edelman, R.R.; Manning, W.J. Prospective Navigator Correction of Image Position for Coronary MR Angiography. Radiology 1997, 203, 733–736. [Google Scholar] [CrossRef]
- Wang, Y.; Rossman, P.J.; Grimm, R.C.; Riederer, S.J.; Ehman, R.L. Navigator-Echo-Based Real-Time Respiratory Gating and Triggering for Reduction of Respiration Effects in Three-Dimensional Coronary MR Angiography. Radiology 1996, 198, 55–60. [Google Scholar] [CrossRef]
- Falcão, M.B.L.; Di Sopra, L.; Ma, L.; Bacher, M.; Yerly, J.; Speier, P.; Rutz, T.; Prša, M.; Markl, M.; Stuber, M.; et al. Pilot Tone Navigation for Respiratory and Cardiac Motion-Resolved Free-Running 5D Flow MRI. Magn. Reson. Med. 2022, 87, 718–732. [Google Scholar] [CrossRef]
- Walheim, J.; Dillinger, H.; Kozerke, S. Multipoint 5D Flow Cardiovascular Magnetic Resonance—Accelerated Cardiac- and Respiratory-Motion Resolved Mapping of Mean and Turbulent Velocities. J. Cardiovasc. Magn. Reson. 2019, 21, 42. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, L.; Piccirelli, M.; Vishnevskiy, V.; Kozerke, S. FlowMRI-Net: A Generalizable Self-Supervised 4D Flow MRI Reconstruction Network. J. Cardiovasc. Magn. Reson. 2025, 27, 101913. [Google Scholar] [CrossRef]
- Ma, L.E.; Yerly, J.; Piccini, D.; Di Sopra, L.; Roy, C.W.; Carr, J.C.; Rigsby, C.K.; Kim, D.; Stuber, M.; Markl, M. 5D Flow MRI: A Fully Self-Gated, Free-Running Framework for Cardiac and Respiratory Motion–Resolved 3D Hemodynamics. Radiol. Cardiothorac. Imaging 2020, 2, e200219. [Google Scholar] [CrossRef]
- Walheim, J.; Dillinger, H.; Gotschy, A.; Kozerke, S. 5D Flow Tensor MRI to Efficiently Map Reynolds Stresses of Aortic Blood Flow In-Vivo. Sci. Rep. 2019, 9, 18794. [Google Scholar] [CrossRef]
- Di Sopra, L.; Piccini, D.; Coppo, S.; Stuber, M.; Yerly, J. An Automated Approach to Fully Self-Gated Free-Running Cardiac and Respiratory Motion-Resolved 5D Whole-Heart MRI. Magn. Reson. Med. 2019, 82, 2118–2132. [Google Scholar] [CrossRef]
- Feng, L.; Coppo, S.; Piccini, D.; Yerly, J.; Lim, R.P.; Masci, P.G.; Stuber, M.; Sodickson, D.K.; Otazo, R. 5D Whole-Heart Sparse MRI. Magn. Reson. Med. 2018, 79, 826–838. [Google Scholar] [CrossRef]
- Madore, B.; Hess, A.T.; van Niekerk, A.M.J.; Hoinkiss, D.C.; Hucker, P.; Zaitsev, M.; Afacan, O.; Günther, M. External Hardware and Sensors, for Improved MRI. J. Magn. Reson. Imaging 2023, 57, 690–705. [Google Scholar] [CrossRef]
- Maclaren, J.; Aksoy, M.; Bammer, R. Contact-Free Physiological Monitoring Using a Markerless Optical System. Magn. Reson. Med. 2015, 74, 571–577. [Google Scholar] [CrossRef]
- Gottwald, L.M.; Blanken, C.P.S.; Tourais, J.; Smink, J.; Planken, R.N.; Boekholdt, S.M.; Meijboom, L.J.; Coolen, B.F.; Strijkers, G.J.; Nederveen, A.J.; et al. Retrospective Camera-Based Respiratory Gating in Clinical Whole-Heart 4D Flow MRI. J. Magn. Reson. Imaging 2021, 54, 440–451. [Google Scholar] [CrossRef] [PubMed]
- Solomon, E.; Rigie, D.S.; Vahle, T.; Paška, J.; Bollenbeck, J.; Sodickson, D.K.; Boada, F.E.; Block, K.T.; Chandarana, H. Free-Breathing Radial Imaging Using a Pilot-Tone Radiofrequency Transmitter for Detection of Respiratory Motion. Magn. Reson. Med. 2021, 85, 2672–2685. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Liu, Y.; Simonetti, O.P.; Tong, M.; Jin, N.; Bacher, M.; Speier, P.; Ahmad, R. Cardiac and Respiratory Motion Extraction for MRI Using Pilot Tone–A Patient Study. Int. J. Cardiovasc. Imaging 2024, 40, 93–105. [Google Scholar] [CrossRef]
- Vahle, T.; Bacher, M.; Rigie, D.; Fenchel, M.; Speier, P.; Bollenbeck, J.; Schäfers, K.P.; Kiefer, B.; Boada, F.E. Respiratory Motion Detection and Correction for MR Using the Pilot Tone: Applications for MR and Simultaneous PET/MR Examinations. Investig. Radiol. 2020, 55, 153. [Google Scholar] [CrossRef] [PubMed]
- Tasdelen, B.; Yagiz, E.; Cinbis, B.R.; Tian, Y.; Nayak, K.S. Contactless Cardiac Gating at 0.55 T Using High-Amplitude Pilot Tone with Interference Cancellation (HAPTIC). Magn. Reson. Med. 2025, 94, 1182–1190. [Google Scholar] [CrossRef]
- Liang, X.; Bi, Z.; Yang, C.; Sheng, R.; Xia, X.; Zhang, Z.; Dai, Y.; Zeng, M. Free-Breathing Liver Magnetic Resonance Imaging with Respiratory Frequency-Modulated Continuous-Wave Radar-Trigger Technique: A Preliminary Study. Front. Oncol. 2022, 12, 918173. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Ryu, K.; Li, Z.; Oscanoa, J.; Wu, Y.; Robb, F.; Vasanawala, S.; Pauly, J.; Scott, G. MRI Retrospective Respiratory Gating and Cardiac Sensing by CW Doppler Radar: A Feasibility Study. IEEE Trans. Biomed. Eng. 2025, 72, 112–122. [Google Scholar] [CrossRef]
- Wang, H.; Li, Y.; Xia, X.; Hu, L.; Zhao, J.; Chen, Q. Non-Contact Respiratory Triggering for Clinical MRI Using Frequency Modulated Continuous Wave Radar. In Proceedings of the Medical Imaging 2021: Physics of Medical Imaging; SPIE: Bellingham, WA, USA, 2021; Volume 11595, pp. 268–275. [Google Scholar]
- McGrath, C.; Bieri, O.; Kozerke, S.; Bauman, G. Self-Gated Cine Phase-Contrast Balanced SSFP Flow Quantification at 0.55 T. Magn. Reson. Med. 2024, 91, 174–189. [Google Scholar] [CrossRef]
- Paul, J.; Divkovic, E.; Wundrak, S.; Bernhardt, P.; Rottbauer, W.; Neumann, H.; Rasche, V. High-Resolution Respiratory Self-Gated Golden Angle Cardiac MRI: Comparison of Self-Gating Methods in Combination with k-t SPARSE SENSE. Magn. Reson. Med. 2015, 73, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Piccini, D.; Feng, L.; Bonanno, G.; Coppo, S.; Yerly, J.; Lim, R.P.; Schwitter, J.; Sodickson, D.K.; Otazo, R.; Stuber, M. Four-Dimensional Respiratory Motion-Resolved Whole Heart Coronary MR Angiography. Magn. Reson. Med. 2017, 77, 1473–1484. [Google Scholar] [CrossRef]
- Pang, J.; Sharif, B.; Fan, Z.; Bi, X.; Arsanjani, R.; Berman, D.S.; Li, D. ECG and Navigator-Free Four-Dimensional Whole-Heart Coronary MRA for Simultaneous Visualization of Cardiac Anatomy and Function. Magn. Reson. Med. 2014, 72, 1208–1217. [Google Scholar] [CrossRef]
- Weiss, E.K.; Baraboo, J.; Rigsby, C.K.; Robinson, J.D.; Ma, L.; Falcão, M.B.L.; Roy, C.W.; Stuber, M.; Markl, M. Respiratory-Resolved Five-Dimensional Flow Cardiovascular Magnetic Resonance: In-Vivo Validation and Respiratory-Dependent Flow Changes in Healthy Volunteers and Patients with Congenital Heart Disease. J. Cardiovasc. Magn. Reson. 2024, 26, 101077. [Google Scholar] [CrossRef]
- Holtackers, R.J.; Stuber, M. Free-Running Cardiac and Respiratory Motion-Resolved Imaging: A Paradigm Shift for Managing Motion in Cardiac MRI? Diagnostics 2024, 14, 1946. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Riederer, S.J.; Ehman, R.L. Respiratory Motion of the Heart: Kinematics and the Implications for the Spatial Resolution in Coronary Imaging. Magn. Reson. Med. 1995, 33, 713–719. [Google Scholar] [CrossRef]
- Roy, C.W.; Milani, B.; Yerly, J.; Si-Mohamed, S.; Romanin, L.; Bustin, A.; Tenisch, E.; Rutz, T.; Prsa, M.; Stuber, M. Intra-Bin Correction and Inter-Bin Compensation of Respiratory Motion in Free-Running Five-Dimensional Whole-Heart Magnetic Resonance Imaging. J. Cardiovasc. Magn. Reson. 2024, 26, 101037. [Google Scholar] [CrossRef]
- Deng, Z.; Pang, J.; Yang, W.; Yue, Y.; Sharif, B.; Tuli, R.; Li, D.; Fraass, B.; Fan, Z. Four-Dimensional MRI Using Three-Dimensional Radial Sampling with Respiratory Self-Gating to Characterize Temporal Phase-Resolved Respiratory Motion in the Abdomen. Magn. Reson. Med. 2016, 75, 1574–1585. [Google Scholar] [CrossRef]
- Pruitt, A.; Rich, A.; Liu, Y.; Jin, N.; Potter, L.; Tong, M.; Rajpal, S.; Simonetti, O.; Ahmad, R. Fully Self-Gated Whole-Heart 4D Flow Imaging from a 5-Minute Scan. Magn. Reson. Med. 2021, 85, 1222–1236. [Google Scholar] [CrossRef] [PubMed]
- Denecken, E.; Sotelo, J.; Arrieta, C.; Andia, M.E.; Uribe, S. Impact of Respiratory Gating on Hemodynamic Parameters from 4D Flow MRI. Appl. Sci. 2022, 12, 2943. [Google Scholar] [CrossRef]
- Bastkowski, R.; Weiss, K.; Maintz, D.; Giese, D. Self-Gated Golden-Angle Spiral 4D Flow MRI. Magn. Reson. Med. 2018, 80, 904–913. [Google Scholar] [CrossRef]
- Campbell-Washburn, A.E.; Ramasawmy, R.; Restivo, M.C.; Bhattacharya, I.; Basar, B.; Herzka, D.A.; Hansen, M.S.; Rogers, T.; Bandettini, W.P.; McGuirt, D.R.; et al. Opportunities in Interventional and Diagnostic Imaging by Using High-Performance Low-Field-Strength MRI. Radiology 2019, 293, 384–393. [Google Scholar] [CrossRef]
- Guenthner, C.; Peereboom, S.M.; Dillinger, H.; McGrath, C.; Albannay, M.M.; Vishnevskiy, V.; Fuetterer, M.; Luechinger, R.; Jenneskens, T.; Sturzenegger, U.; et al. Ramping down a Clinical 3 T Scanner: A Journey into MRI and MRS at 0.75 T. Magn. Reson. Mater. Phys. Biol. Med. 2023, 36, 355–373. [Google Scholar] [CrossRef]
- Bandettini, W.P.; Shanbhag, S.M.; Mancini, C.; McGuirt, D.R.; Kellman, P.; Xue, H.; Henry, J.L.; Lowery, M.; Thein, S.L.; Chen, M.Y.; et al. A Comparison of Cine CMR Imaging at 0.55 T and 1.5 T. J. Cardiovasc. Magn. Reson. 2020, 22, 37. [Google Scholar] [CrossRef] [PubMed]
- Varghese, J.; Jin, N.; Giese, D.; Chen, C.; Liu, Y.; Pan, Y.; Nair, N.; Shalaan, M.T.; Khan, M.; Tong, M.S.; et al. Building a Comprehensive Cardiovascular Magnetic Resonance Exam on a Commercial 0.55 T System: A Pictorial Essay on Potential Applications. Front. Cardiovasc. Med. 2023, 10, 1120982. [Google Scholar] [CrossRef] [PubMed]
- Segeroth, M.; Winkel, D.J.; Vosshenrich, J.; Breit, H.-C.; Giese, D.; Haaf, P.; Zellweger, M.J.; Bremerich, J.; Santini, F.; Pradella, M. Cardiac Cine MRI Using a Commercially Available 0.55-T Scanner. Radiol. Cardiothorac. Imaging 2024, 6, e230331. [Google Scholar] [CrossRef]
- Sieber, X.; Binzel, K.; Varghese, J.; Liu, Y.; Yerly, J.; Roy, C.W.; Antiochos, P.; Prsa, M.; van Heeswijk, R.B.; Simonetti, O.P.; et al. Measuring Biventricular Function and Left Atrial Volume in a Single Five-Dimensional Whole-Heart Cardiovascular Magnetic Resonance Scan at 0.55 T. J. Cardiovasc. Magn. Reson. 2025, 27, 101906. [Google Scholar] [CrossRef]
- Xiang, J.; Ramasawmy, R.; Seemann, F.; Peters, D.C.; Campbell-Washburn, A.E. Balanced Steady-State Free Precession Phase Contrast at 0.55 T Applied to Aortic Flow. J. Cardiovasc. Magn. Reson. 2024, 26, 101098. [Google Scholar] [CrossRef]
- Kruger, M.G.; Springer, R.P.W.; Kersten, G.M.; Bril, R.J. Contact-Less Vital Sign Monitoring Using a COTS Resource-Constrained Multi-Core System. In Proceedings of the IECON 2019—45th Annual Conference of the IEEE Industrial Electronics Society, Lisbon, Portugal, 14–17 October 2019; Volume 1, pp. 3057–3062. [Google Scholar]
- Rocque, M. Fully Automated Contactless Respiration Monitoring Using a Camera. In Proceedings of the 2016 IEEE International Conference on Consumer Electronics (ICCE), Las Vegas, NV, USA, 7–11 January 2016; pp. 478–479. [Google Scholar]
- Stelter, J.; Weiss, K.; Steinhelfer, L.; Meineke, J.; Zhang, W.; Kainz, B.; Braren, R.F.; Karampinos, D.C. Abdominal Simultaneous 3D Water T1 and T2 Mapping Using a Free-Breathing Cartesian Acquisition with Spiral Profile Ordering. Magn. Reson. Med. 2026, 95, 268–285. [Google Scholar] [CrossRef]
- Krishnamoorthy, G.; Tourais, J.; Smink, J.; Breeuwer, M.; Kouwenhoven, M. Free-Breathing 2D Radial Cine MRI with Respiratory Auto-Calibrated Motion Correction (RAMCO). Magn. Reson. Med. 2023, 89, 977–989. [Google Scholar] [CrossRef]
- Wu, C.; Krishnamoorthy, G.; Yu, V.; Subashi, E.; Rimner, A.; Otazo, R. 4D Lung MRI with High-Isotropic-Resolution Using Half-Spoke (UTE) and Full-Spoke 3D Radial Acquisition and Temporal Compressed Sensing Reconstruction. Phys. Med. Biol. 2023, 68, 035017. [Google Scholar] [CrossRef]
- Murray, V.; Wu, C.; Otazo, R. High-Definition Motion-Resolved MRI Using 3D Radial Kooshball Acquisition and Deep Learning Spatial-Temporal 4D Reconstruction. Phys. Med. Biol. 2025, 70, 125012. [Google Scholar] [CrossRef] [PubMed]
- Harder, F.; Lohöfer, F.K.; Kaissis, G.A.; Zoellner, C.; Kamal, O.; Katemann, C.; Hock, A.; Senegas, J.; Peeters, J.M.; Rummeny, E.J.; et al. Camera-Based Respiratory Triggering Improves the Image Quality of 3D Magnetic Resonance Cholangiopancreatography. Eur. J. Radiol. 2019, 120, 108675. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Xu, J.; Wu, Q.; Wang, X.; Ren, J.; Wang, X.; Xue, H.; Jin, Z. Application of Compressed Sensing 3D MR Cholangiopancreatography (CS-MRCP) with Contact-Free Physiological Monitoring (CFPM) for Pancreaticobiliary Disorders. Acad. Radiol. 2021, 28, S148–S156. [Google Scholar] [CrossRef]
- (ISMRM 2020) Retrospective Camera-Based Respiratory Binning for 4D Flow MRI—A Comparison with Liver Navigator and Self-Gating. Available online: https://archive.ismrm.org/2020/1330.html (accessed on 7 January 2026).
- Markl, M.; Leupold, J. Gradient Echo Imaging. J. Magn. Reson. Imaging 2012, 35, 1274–1289. [Google Scholar] [CrossRef]
- Zhang, T.; Cheng, J.Y.; Chen, Y.; Nishimura, D.G.; Pauly, J.M.; Vasanawala, S.S. Robust Self-Navigated Body MRI Using Dense Coil Arrays. Magn. Reson. Med. 2016, 76, 197–205. [Google Scholar] [CrossRef]
- Jacobs, L.; Piccirelli, M.; Vishnevskiy, V.; Kozerke, S. FlowMRI-Net Dataset, for Aortic and Cerebrovascular 4D Flow MRI; ETH Zurich: Zurich, Switzerland, 2025. [Google Scholar] [CrossRef]
- Bernstein, M.A.; Zhou, X.J.; Polzin, J.A.; King, K.F.; Ganin, A.; Pelc, N.J.; Glover, G.H. Concomitant Gradient Terms in Phase Contrast MR: Analysis and Correction. Magn. Reson. Med. 1998, 39, 300–308. [Google Scholar] [CrossRef] [PubMed]
- Busch, J.; Giese, D.; Kozerke, S. Image-Based Background Phase Error Correction in 4D Flow MRI Revisited. J. Magn. Reson. Imaging 2017, 46, 1516–1525. [Google Scholar] [CrossRef] [PubMed]
- Loecher, M.; Schrauben, E.; Johnson, K.M.; Wieben, O. Phase Unwrapping in 4D MR Flow with a 4D Single-Step Laplacian Algorithm. J. Magn. Reson. Imaging 2016, 43, 833–842. [Google Scholar] [CrossRef] [PubMed]
- (ISMRM 2025) Joint Magnitude and Phase Neural Network Segmentation for Cine 2D Flow MRI. Available online: https://archive.ismrm.org/2025/3505.html (accessed on 25 November 2025).
- (ISMRM 2025) Synthetically Trained Convolutional Neural Network Vessel Segmentation for 4D Flow MRI. Available online: https://archive.ismrm.org/2025/3506_0XsfdUQkw.html (accessed on 25 November 2025).
- Bernstein, M.A.; King, K.F.; Zhou, X.J. Chapter 9—Motion-Sensitizing Gradients. In Handbook of MRI Pulse Sequences; Bernstein, M.A., King, K.F., Zhou, X.J., Eds.; Academic Press: Burlington, MA, USA, 2004; pp. 274–291. [Google Scholar]





| Characteristic | Value |
|---|---|
| Number of Volunteers | 20 |
| Age [years] | 32.8 ± 12.6 |
| Heart Rate [bpm] | 59.5 ± 8.3 |
| Weight [kg] | 67.2 ± 7.3 |
| Height [cm] | 174.4 ± 5.1 |
| Females (%) | 50 |
| Parameters | 2D Flow | 4D Flow |
|---|---|---|
| ECG gating | Retrospective | Retrospective |
| Respiratory compensation | Breath-hold | SG or VE Gating |
| FOV [mm3] | 236–260 × 260–280 × 10 | 360 × 240–288 × 60 |
| Voxel size [mm3] | 1.75 × 1.75 × 10.0 | 2.5 × 2.5 × 2.5 |
| Undersampling factor R | 2 | ~4–4.5 |
| TR [ms] | 5.9 | 5.5–5.9 |
| TE [ms] | 3.5 | 3.3–3.5 |
| Temporal resolution [ms] | 41–42 | 42–46 |
| Flip angle [°] | 15 | 15 |
| Venc [cm/s] (FH-AP-RL) | 150–100–100 | 150–100–100 |
| Bandwidth [Hz] | 335 | 478 |
| Scan duration | ~20 s | ~9.6 min |
| Metrics | Mean ± Std. |
|---|---|
| RMSE [cm/s] | 3.9 ± 1.02 |
| mDirrErr [a.u.] | 0.01 ± 0.007 |
| Slope [a.u.] | 1.0 ± 0.008 |
| Intercept [cm/s] | 0.19 ± 0.28 |
| R2 [a.u.] | 0.99 ± 0.009 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Emery, S.; Jacobs, L.; Malich, J.; Wolkerstorfer, G.; Dong, Y.; Ercan, E.; Smink, J.; Nagtegaal, M.; Kozerke, S. 4D Flow MRI at 0.6 T—Self-Gating Versus Camera-Based Respiratory Binning. Bioengineering 2026, 13, 282. https://doi.org/10.3390/bioengineering13030282
Emery S, Jacobs L, Malich J, Wolkerstorfer G, Dong Y, Ercan E, Smink J, Nagtegaal M, Kozerke S. 4D Flow MRI at 0.6 T—Self-Gating Versus Camera-Based Respiratory Binning. Bioengineering. 2026; 13(3):282. https://doi.org/10.3390/bioengineering13030282
Chicago/Turabian StyleEmery, Sébastien, Luuk Jacobs, Jacob Malich, Gloria Wolkerstorfer, Yiming Dong, Ece Ercan, Jouke Smink, Martijn Nagtegaal, and Sebastian Kozerke. 2026. "4D Flow MRI at 0.6 T—Self-Gating Versus Camera-Based Respiratory Binning" Bioengineering 13, no. 3: 282. https://doi.org/10.3390/bioengineering13030282
APA StyleEmery, S., Jacobs, L., Malich, J., Wolkerstorfer, G., Dong, Y., Ercan, E., Smink, J., Nagtegaal, M., & Kozerke, S. (2026). 4D Flow MRI at 0.6 T—Self-Gating Versus Camera-Based Respiratory Binning. Bioengineering, 13(3), 282. https://doi.org/10.3390/bioengineering13030282







