Simulation-Driven Annotation-Free Deep Learning for Automated Detection and Segmentation of Airway Mucus Plugs on Non-Contrast CT Images
Abstract
1. Introduction
2. Methods and Materials
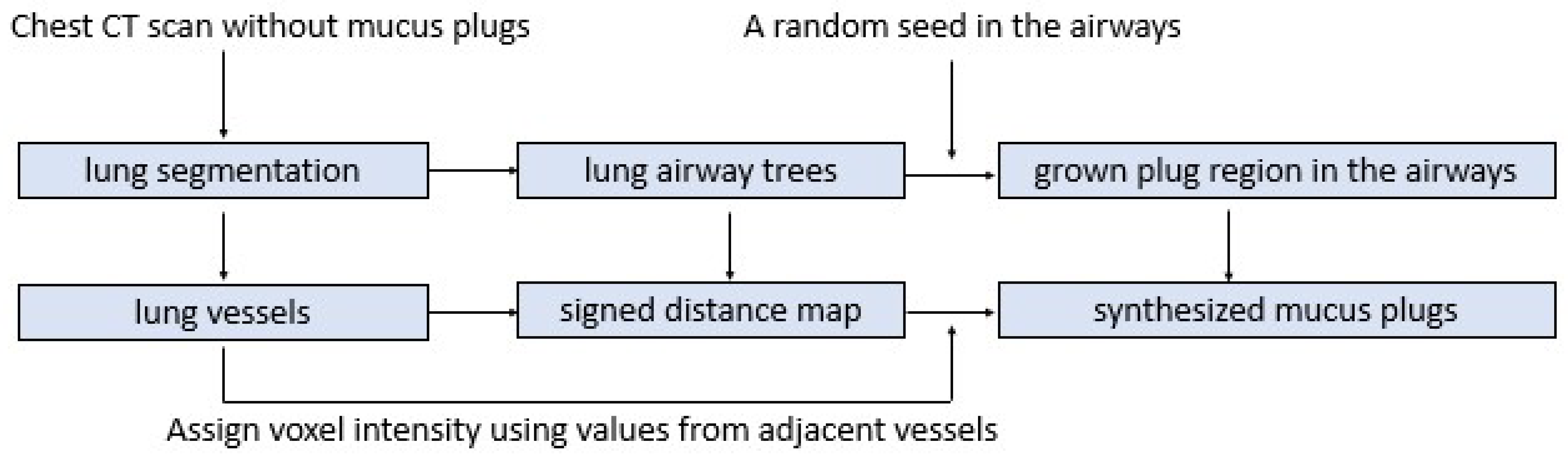
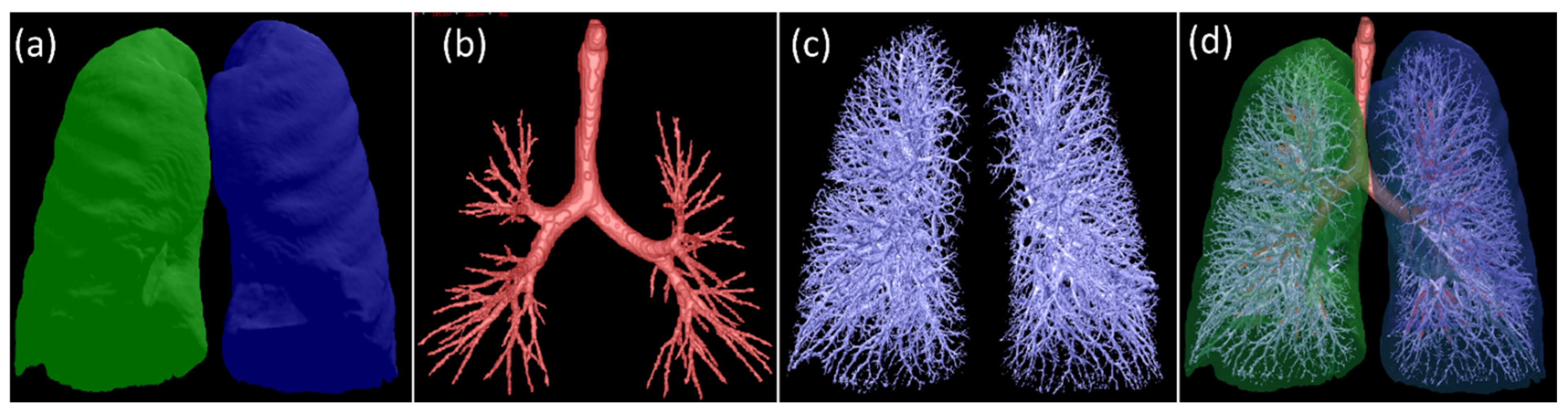
2.1. Workflow Overview
2.2. Chest CT Data and Expert Annotation
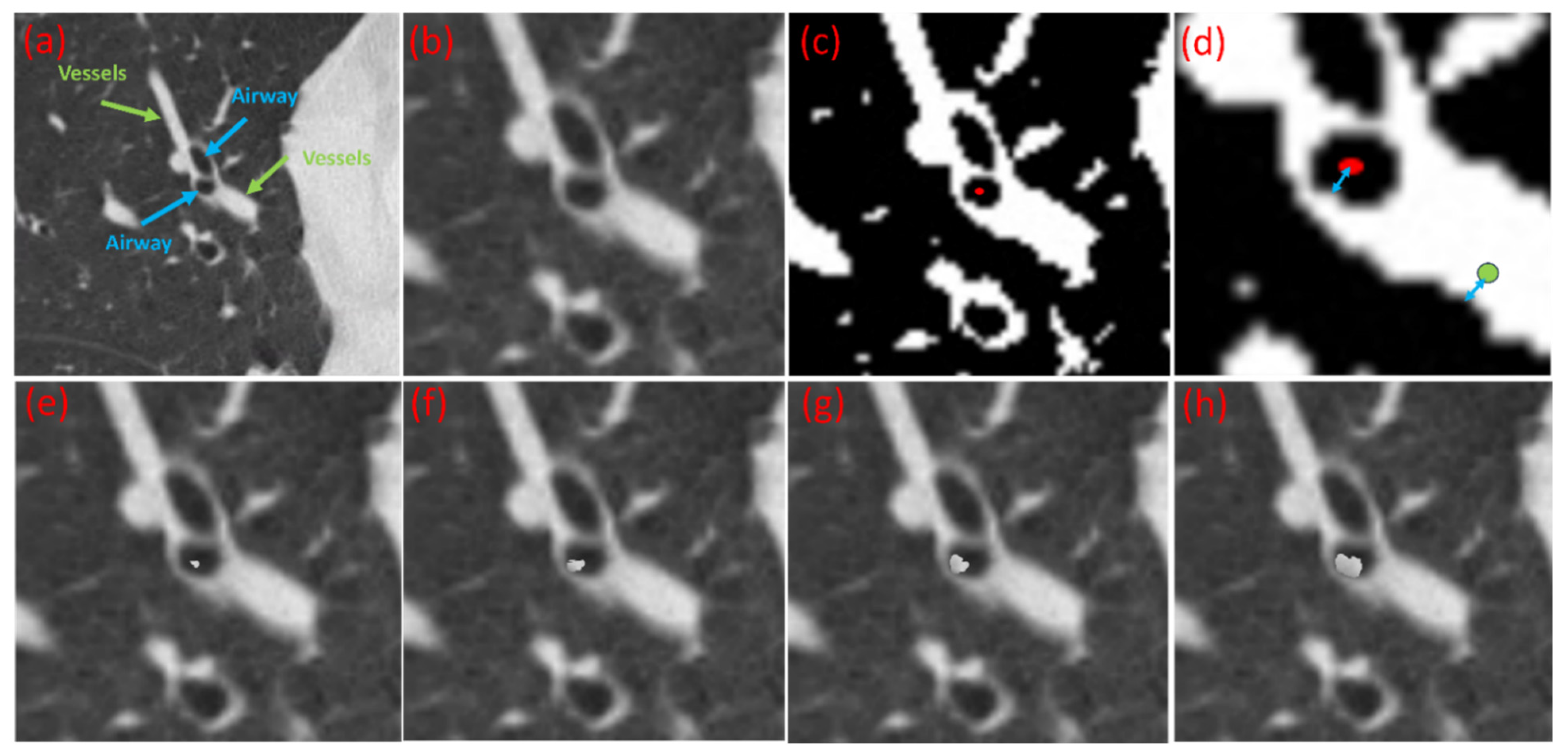
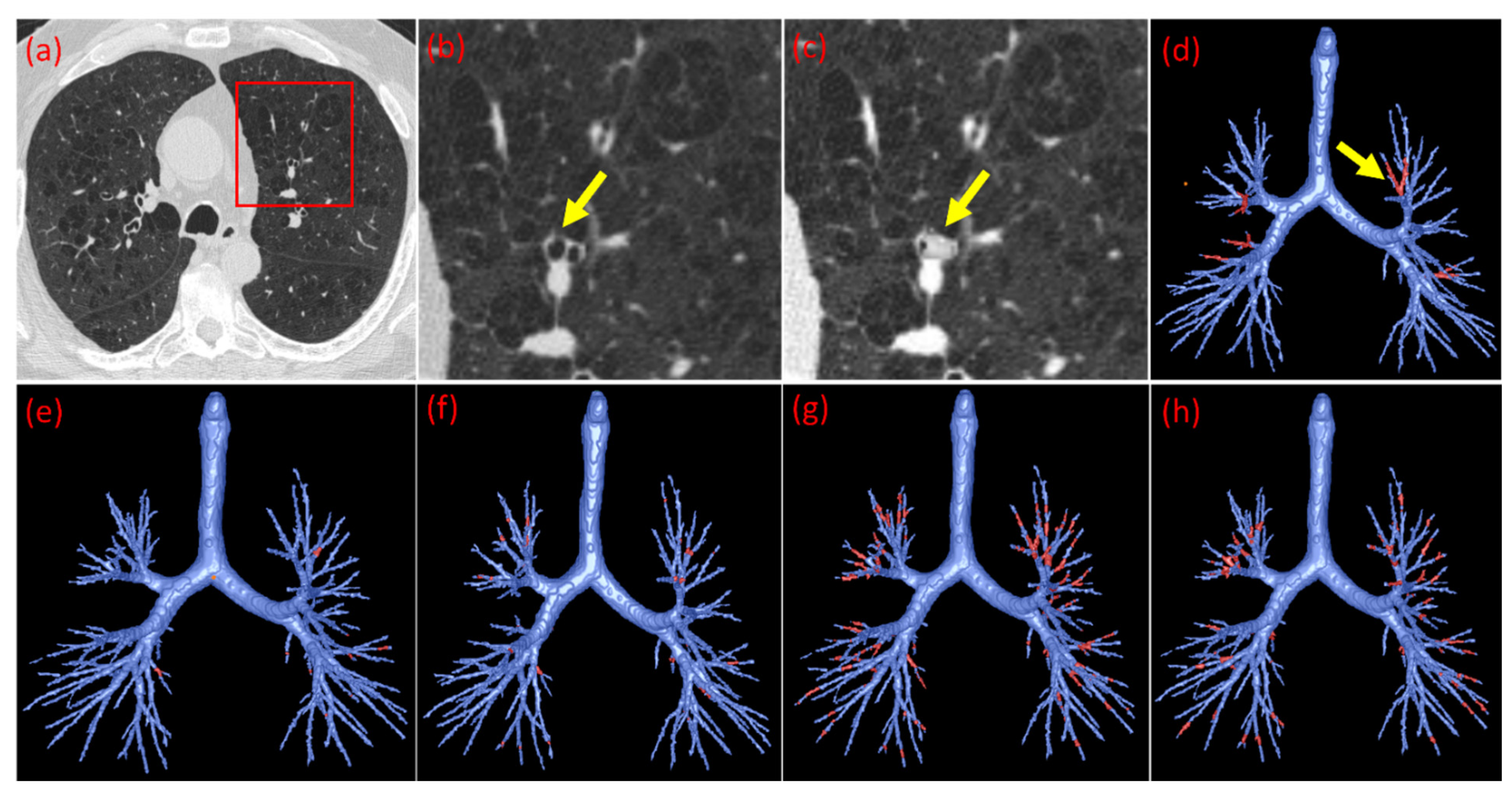
2.3. Simulating Mucus Plugs in Chest CT Scans
2.4. CNN-Based Mucus Plug Detection and Segmentation
2.5. Performance Evaluation
3. Results
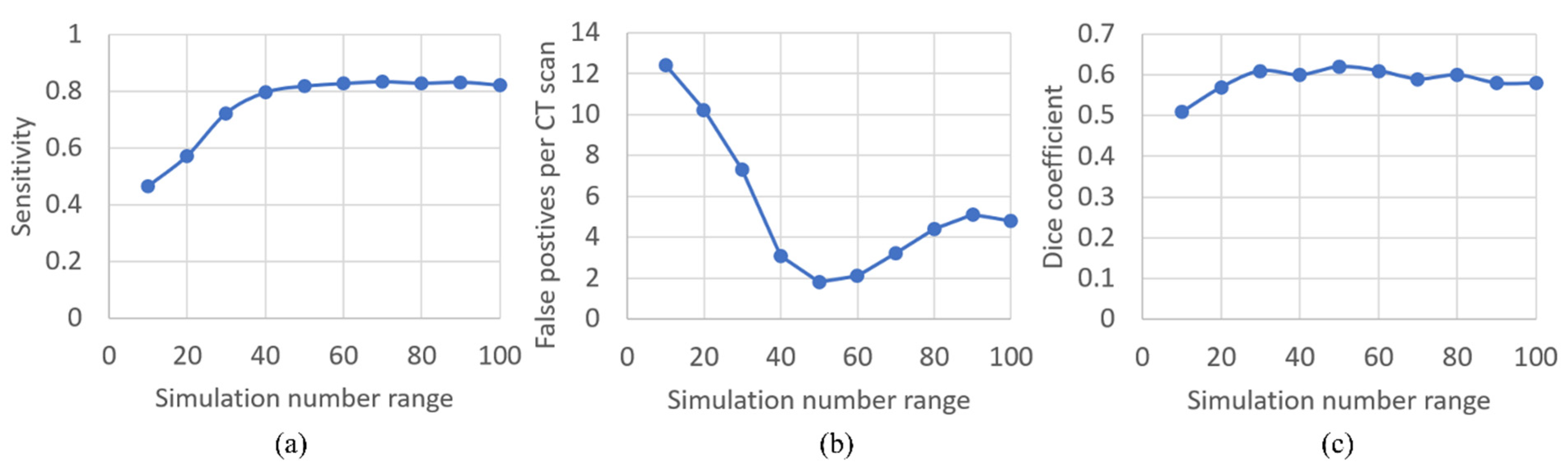
3.1. Optimization of Synthetic Plug Augmentation Count
3.2. Summary of the Synthetic and Real Mucus Plugs in the Study Cohort
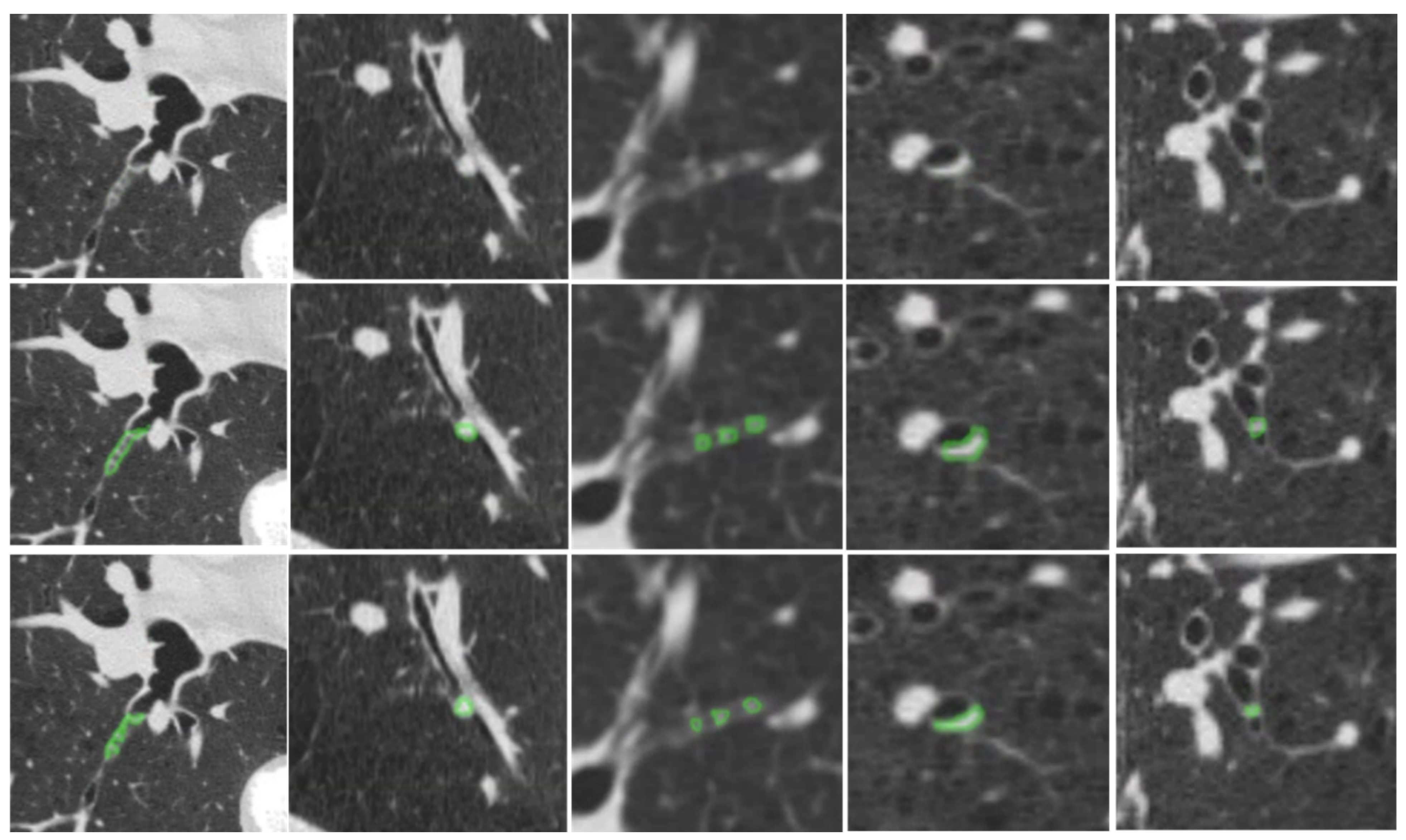
3.3. Performance Comparison: S-Model vs. M-Model
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Dunican, E.M.; Elicker, B.M.; Gierada, D.S.; Nagle, S.K.; Schiebler, M.L.; Newell, J.D.; Raymond, W.W.; Lachowicz-Scroggins, M.E.; Di Maio, S.; Hoffman, E.A.; et al. Mucus plugs in patients with asthma linked to eosinophilia and airflow obstruction. J. Clin. Investig. 2018, 128, 997–1009. [Google Scholar] [CrossRef]
- Mummy, D.G.; Dunican, E.M.; Carey, K.J.; Evans, M.D.; Elicker, B.M.; Newell, J.D., Jr.; Gierada, D.S.; Nagle, S.K.; Schiebler, M.L.; Sorkness, R.L.; et al. Mucus Plugs in Asthma at CT Associated with Regional Ventilation Defects at 3He MRI. Radiology 2022, 303, 184–190. [Google Scholar] [CrossRef]
- Huang, B.K.; Elicker, B.M.; Henry, T.S.; Kallianos, K.G.; Hahn, L.D.; Tang, M.; Heng, F.; McCulloch, C.E.; Bhakta, N.R.; Majumdar, S.; et al. Persistent mucus plugs in proximal airways are consequential for airflow limitation in asthma. JCI Insight 2024, 9, e174124. [Google Scholar] [CrossRef]
- Georas, S.N. All plugged up—Noninvasive mucus score to assess airway dysfunction in asthma. J. Clin. Investig. 2018, 128, 906–909. [Google Scholar] [CrossRef]
- Aikawa, T.; Shimura, S.; Sasaki, H.; Ebina, M.; Takishima, T. Marked goblet cell hyperplasia with mucus accumulation in the airways of patients who died of severe acute asthma attack. Chest 1992, 101, 916–921. [Google Scholar] [CrossRef]
- Rosenman, K.D.; Hanna, E.A.; Lyon-Callo, S.K.; Wasilevich, E.A. Investigating asthma deaths among children and young adults: Michigan Asthma Mortality Review. Public Health Rep. 2007, 122, 373–381. [Google Scholar] [CrossRef]
- Kuyper, L.M.; Pare, P.D.; Hogg, J.C.; Lambert, R.K.; Ionescu, D.; Woods, R.; Bai, T.R. Characterization of airway plugging in fatal asthma. Am. J. Med. 2003, 115, 6–11. [Google Scholar] [CrossRef]
- Diaz, A.A.; Orejas, J.L.; Grumley, S.; Nath, H.P.; Wang, W.; Dolliver, W.R.; Yen, A.; Kligerman, S.J.; Jacobs, K.; Manapragada, P.P.; et al. Airway-Occluding Mucus Plugs and Mortality in Patients with Chronic Obstructive Pulmonary Disease. JAMA 2023, 329, 1832–1839. [Google Scholar] [CrossRef] [PubMed]
- Okajima, Y.; Come, C.E.; Nardelli, P.; Sonavane, S.K.; Yen, A.; Nath, H.P.; Terry, N.; Grumley, S.A.; Ahmed, A.; Kligerman, S.; et al. Luminal Plugging on Chest CT Scan: Association with Lung Function, Quality of Life, and COPD Clinical Phenotypes. Chest 2020, 158, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Howell, I.; Howell, A.; Pavord, I.D. Type 2 inflammation and biological therapies in asthma: Targeted medicine taking flight. J. Exp. Med. 2023, 220, e20221212. [Google Scholar] [CrossRef] [PubMed]
- Sakai, N.; Koya, T.; Murai, Y.; Tsubokawa, F.; Tanaka, K.; Naramoto, S.; Aoki, A.; Shima, K.; Kimura, Y.; Watanabe, S.; et al. Effect of Benralizumab on Mucus Plugs in Severe Eosinophilic Asthma. Int. Arch. Allergy Immunol. 2023, 184, 783–791. [Google Scholar] [CrossRef]
- Chassagnon, G.; Burgel, P.R. Mucus Plugs in Medium-sized Airways: A Novel Imaging Biomarker for Phenotyping Chronic Obstructive Pulmonary Disease. Am. J. Respir. Crit. Care Med. 2021, 203, 932–934. [Google Scholar] [CrossRef]
- Nordenmark, L.H.; Hellqvist, Å.; Emson, C.; Diver, S.; Porsbjerg, C.; Griffiths, J.M.; Newell, J.D.; Peterson, S.; Pawlikowska, B.; Parnes, J.R.; et al. Tezepelumab and Mucus Plugs in Patients with Moderate-to-Severe Asthma. NEJM Evid. 2023, 2, EVIDoa2300135. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Chen, Z.; Lei, Q.; Ma, D.; Chen, M.; Liu, J. Chest CT assess the impact of omalizumab treatment on airway remodeling in refractory asthma. Pulm. Pharmacol. Ther. 2024, 87, 102329. [Google Scholar] [CrossRef] [PubMed]
- Audousset, C.; Swaleh, S.; Olivenstein, R.; Vameghestahbanati, M.; Kirby, M.; Semionov, A.; Smith, B.M.; Martin, J.G. Mucus plugs in the airways of asthmatic subjects and smoking status. Respir. Res. 2024, 25, 52. [Google Scholar] [CrossRef] [PubMed]
- Heng, F.; Henry, T.; Huang, B.; Dunican, E.; Elicker, B.; Fahy, J. Detailed annotation of airway mucus plugs to inform deep learning-based methods for automated mucus scoring. Eur. Respir. J. 2020, 56, 854. [Google Scholar] [CrossRef]
- van der Veer, T.; Andrinopoulou, E.R.; Braunstahl, G.J.; Charbonnier, J.P.; Kim, V.; Latisenko, R.; Lynch, D.A.; Tiddens, H. Association between automatic AI-based quantification of airway-occlusive mucus plugs and all-cause mortality in patients with COPD. Thorax 2025, 80, 105–108. [Google Scholar] [CrossRef]
- Moya-Saez, E.; Pena-Nogales, O.; Luis-Garcia, R.; Alberola-Lopez, C. A deep learning approach for synthetic MRI based on two routine sequences and training with synthetic data. Comput. Methods Programs Biomed. 2021, 210, 106371. [Google Scholar] [CrossRef]
- Nishio, M.; Fujimoto, K.; Matsuo, H.; Muramatsu, C.; Sakamoto, R.; Fujita, H. Lung Cancer Segmentation with Transfer Learning: Usefulness of a Pretrained Model Constructed from an Artificial Dataset Generated Using a Generative Adversarial Network. Front. Artif. Intell. 2021, 4, 694815. [Google Scholar] [CrossRef]
- Hill, D.B.; Button, B.; Rubinstein, M.; Boucher, R.C. Physiology and pathophysiology of human airway mucus. Physiol. Rev. 2022, 102, 1757–1836. [Google Scholar] [CrossRef]
- Pu, J.; Sechrist, J.; Meng, X.; Leader, J.K.; Sciurba, F.C. A pilot study: Quantify lung volume and emphysema extent directly from two-dimensional scout images. Med. Phys. 2021, 48, 4316–4325. [Google Scholar] [CrossRef] [PubMed]
- Pu, J.; Gezer, N.S.; Ren, S.; Alpaydin, A.O.; Avci, E.R.; Risbano, M.G.; Rivera-Lebron, B.; Chan, S.Y.; Leader, J.K. Automated detection and segmentation of pulmonary embolisms on computed tomography pulmonary angiography (CTPA) using deep learning but without manual outlining. Med. Image Anal. 2023, 89, 102882. [Google Scholar] [CrossRef]
- Pu, J.; Leader, J.K.; Sechrist, J.; Beeche, C.A.; Singh, J.P.; Ocak, I.K.; Risbano, M.G. Automated identification of pulmonary arteries and veins depicted in non-contrast chest CT scans. Med. Image Anal. 2022, 77, 102367. [Google Scholar] [CrossRef] [PubMed]
- Pu, J.; Zheng, B.; Leader, J.K.; Fuhrman, C.; Knollmann, F.; Klym, A.; Gur, D. Pulmonary lobe segmentation in CT examinations using implicit surface fitting. IEEE Trans. Med. Imaging 2009, 28, 1986–1996. [Google Scholar] [CrossRef]
- Pu, J.; Roos, J.; Yi, C.A.; Napel, S.; Rubin, G.D.; Paik, D.S. Adaptive border marching algorithm: Automatic lung segmentation on chest CT images. Comput. Med. Imaging Graph. 2008, 32, 452–462. [Google Scholar] [CrossRef] [PubMed]
- Pu, J.; Fuhrman, C.; Good, W.F.; Sciurba, F.C.; Gur, D. A differential geometric approach to automated segmentation of human airway tree. IEEE Trans. Med. Imaging 2011, 30, 266–278. [Google Scholar] [CrossRef]
- Sethian, J.A. A fast marching level set method for monotonically advancing fronts. Proc. Natl. Acad. Sci. USA 1996, 93, 1591–1595. [Google Scholar] [CrossRef]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 2021, 18, 203–211. [Google Scholar] [CrossRef]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. arXiv 2015, arXiv:1505.04597. [Google Scholar] [CrossRef]
- Muller, D.; Soto-Rey, I.; Kramer, F. Towards a guideline for evaluation metrics in medical image segmentation. BMC Res. Notes 2022, 15, 210. [Google Scholar] [CrossRef]
- Thompson, A.C.; Jammal, A.A.; Berchuck, S.I.; Mariottoni, E.B.; Medeiros, F.A. Assessment of a Segmentation-Free Deep Learning Algorithm for Diagnosing Glaucoma from Optical Coherence Tomography Scans. JAMA Ophthalmol. 2020, 138, 333–339. [Google Scholar] [CrossRef] [PubMed]




| All Subjects (n = 200) | Male (n = 120) | Female (n = 80) | |
|---|---|---|---|
| Age, Mean (Range) | 66.9 (46–82) | 67.4 (50–82) | 66.1 (46–79) |
| Race, n (%) | |||
| White | 189 (94.5) | 114 (95.0) | 75 (93.8) |
| African American | 10 (5.0) | 5 (4.2) | 5 (6.2) |
| Other races | 1 (0.5) | 1 (0.8) | 0 (0) |
| Weight, kg, mean (SD) | 81.4 (16.4) | 88.1 (15.5) | 71.8 (12.3) |
| Height, cm, mean (SD) | 170.0 (9.5) | 175.7 (6.7) | 161.9 (6.4) |
| COPD Severity | |||
| Normal | 76 | 42 | 34 |
| Mild | 35 | 20 | 15 |
| Moderate | 59 | 38 | 21 |
| Severe | 20 | 14 | 6 |
| Very severe | 10 | 6 | 4 |
| Mucus plug status, n (%) | |||
| Yes | 98 (49.0) | 62 (51.7) | 36 (45.0) |
| No | 83 (41.5) | 48 (40.0) | 35 (43.8) |
| Uncertain | 19 (9.5) | 10 (8.3) | 9 (11.2) |
| Real Mucus (in 98 CT Scans) | Synthetic Mucus (in 83 CT Scans) | |
|---|---|---|
| Count: total | 1643 | 4150 |
| mean (SD), range, per CT scan | 17.1 (26.5), 1–146 | 50 (0), 50 |
| Volume (mm3): mean (SD), range | 49.35 (220.0), 1.47–2136.28 | 46.86 (45.8), 1.35–530.2 |
| Length (mm): mean (SD), range | 6.06 (6.89), 1.14–29.75 | 8.90 (4.62), 1.35–27.76 |
| Distribution based on length: | ||
| (0, 3) | 770 (46.9%) | 768 (18.5%) |
| [3, 6) | 459 (27.9%) | 2652 (63.9%) |
| [6, 15) | 280 (17.0%) | 402 (9.7%) |
| [15, ∞) | 134 (8.2%) | 328 (7.9%) |
| Mucus Size (Length: mm) | Metrics | S-Model | M-Model |
|---|---|---|---|
| (0, 3) | Dice coefficient | 0.475 ± 0.145 | 0.482 ± 0.132 |
| Sensitivity | 0.789 (608/770; 95% CI: 0.762–0.814) | 0.794 (611/770; 95% CI: 0.768–0.819) | |
| False positives per scan | 1.34 (131/98) | 1.21 (119/98) | |
| [3, 6) | Dice coefficient | 0.520 ± 0.099 | 0.494 ± 0.113 |
| Sensitivity | 0.843 (387/459; 95% CI: 0.808–0.873) | 0.725 (333/459; 95% CI: 0.686–0.761) | |
| False positives per scan | 0.378 (37/98) | 1.04 (102/98) | |
| [6, 15) | Dice coefficient | 0.672 ± 0.082 | 0.610 ± 0.109 |
| Sensitivity | 0.907 (254/280; 95% CI: 0.868–0.938) | 0.736 (206/280; 95% CI: 0.678–0.788) | |
| False positives per scan | 0.204 (20/98) | 0.643 (63/98) | |
| [15, ∞) | Dice coefficient | 0.777 ± 0.080 | 0.694 ± 0.118 |
| Sensitivity | 0.940 (126/134; 95% CI: 0.884–0.973) | 0.694 (93/134; 95% CI: 0.606–0.771) | |
| False positives per scan | 0 (0/98) | 0.786 (77/98) | |
| All | Dice coefficient | 0.631 ± 0.088 | 0.557 ± 0.119 |
| Sensitivity | 0.837 (1375/1643; 95% CI: 0.818–0.854) | 0.757 (1243/1643; 95% CI: 0.737–0.776) | |
| False positives per scan | 1.91 (188/98) | 3.68 (361/98) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Pu, L.; Gezer, N.S.; Yu, T.; Kirshenboim, Z.; Duman, E.; Dhupar, R.; Meng, X. Simulation-Driven Annotation-Free Deep Learning for Automated Detection and Segmentation of Airway Mucus Plugs on Non-Contrast CT Images. Bioengineering 2026, 13, 153. https://doi.org/10.3390/bioengineering13020153
Pu L, Gezer NS, Yu T, Kirshenboim Z, Duman E, Dhupar R, Meng X. Simulation-Driven Annotation-Free Deep Learning for Automated Detection and Segmentation of Airway Mucus Plugs on Non-Contrast CT Images. Bioengineering. 2026; 13(2):153. https://doi.org/10.3390/bioengineering13020153
Chicago/Turabian StylePu, Lucy, Naciye Sinem Gezer, Tong Yu, Zehavit Kirshenboim, Emrah Duman, Rajeev Dhupar, and Xin Meng. 2026. "Simulation-Driven Annotation-Free Deep Learning for Automated Detection and Segmentation of Airway Mucus Plugs on Non-Contrast CT Images" Bioengineering 13, no. 2: 153. https://doi.org/10.3390/bioengineering13020153
APA StylePu, L., Gezer, N. S., Yu, T., Kirshenboim, Z., Duman, E., Dhupar, R., & Meng, X. (2026). Simulation-Driven Annotation-Free Deep Learning for Automated Detection and Segmentation of Airway Mucus Plugs on Non-Contrast CT Images. Bioengineering, 13(2), 153. https://doi.org/10.3390/bioengineering13020153







