Integrating the Contrasting Perspectives Between the Constrained Disorder Principle and Deterministic Optical Nanoscopy: Enhancing Information Extraction from Imaging of Complex Systems
Abstract
1. Introduction
1.1. Operational Definitions
- Noise (η): Time-varying stochastic fluctuations around a mean value, quantified as the standard deviation: σ_noise = √(⟨(x − ⟨x⟩)2⟩), typically measured over timescales of milliseconds to seconds in biological systems.
- Variability (V): The range or spread of a parameter across measurements or system states, quantified by the coefficient of variation: CV = σ/μ, providing a normalized measure of heterogeneity.
- Disorder (D): The entropy-related measure of unpredictability in system states: D = −Σ p_i log(p_i), where p_i represents the probability of state i.
- Constrained Disorder: Disorder maintained within time-dependent boundaries: D_min(t) ≤ D(t) ≤ D_max(t), where the boundaries themselves evolve according to system demands.
1.2. Alternative Theoretical Frameworks
- Stochastic Thermodynamics: This framework, developed by Seifert and Jarzynski, relates information processing to energy dissipation via thermodynamic bounds [29,30]. While providing rigorous constraints on information-energy tradeoffs, it primarily addresses equilibrium and near-equilibrium systems, whereas biological systems often operate far from equilibrium [31,32].
- Information Theory Approaches: Shannon entropy and mutual information provide quantitative measures of information content in noisy signals [33]. Fisher information theory, particularly relevant to our work, establishes fundamental limits on the precision of parameter estimation [34,35,36]. Our approach builds upon Fisher information while incorporating biological constraints from CDP.
1.3. Objectives and Hypotheses
- Objective 1: Develop a mathematical framework that combines CDP’s dynamic variability bounds with Hell’s Fisher information-based precision measurements.
- Objective 2: Validate the integrated framework through computational simulations, demonstrating quantitative performance improvements over standard approaches.
- Objective 3: Provide practical protocols for experimental implementation, including calibration procedures and real-time adaptation strategies.
2. The Constrained Disorder Principle and Noise in Complex Systems
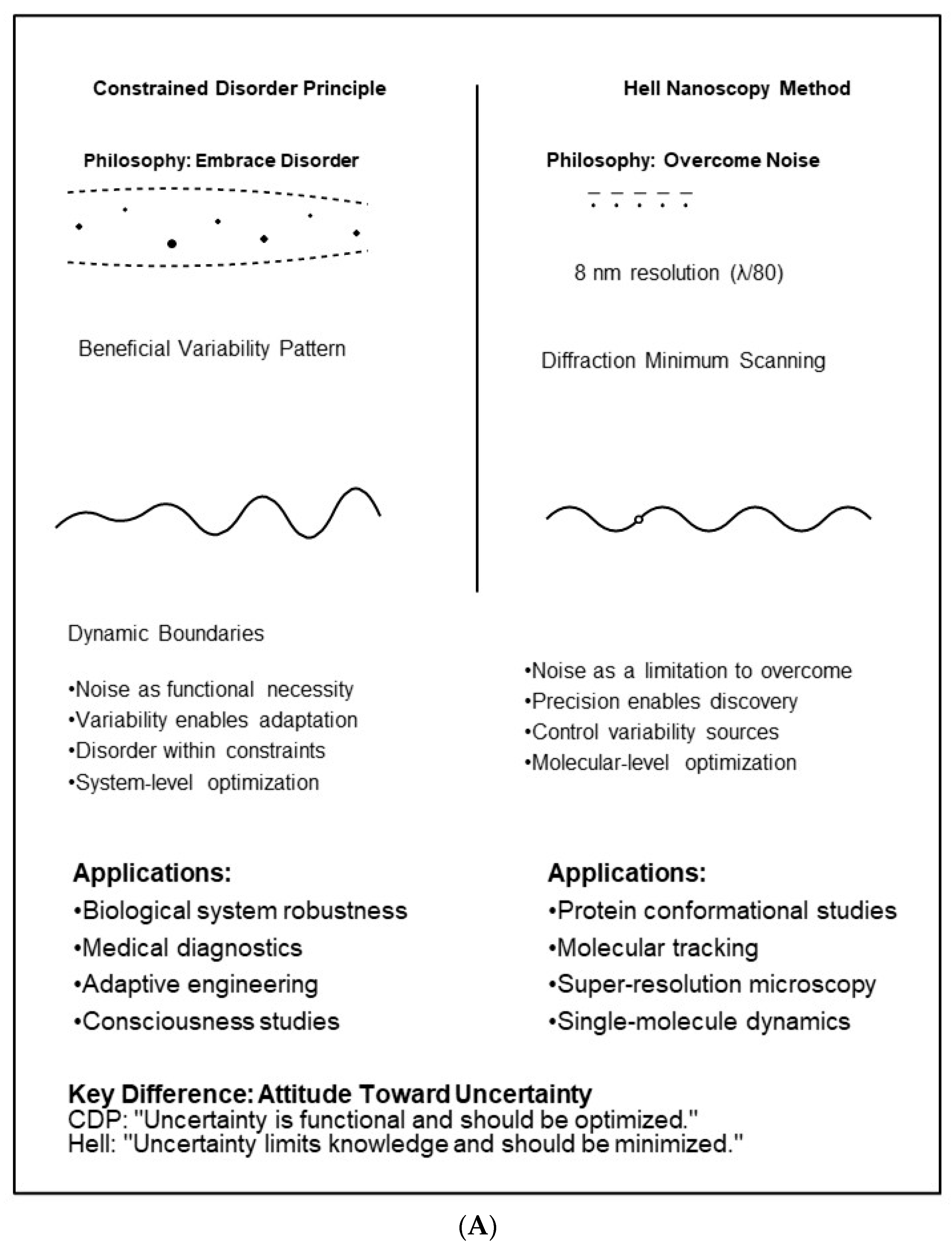
- Constrained Disorder: In a system, disorder is governed by specific rules or boundaries, which facilitate complex interactions among its elements.
- System Behavior: The balance between order and disorder can lead to emergent properties that may not be immediately apparent at the level of individual components.
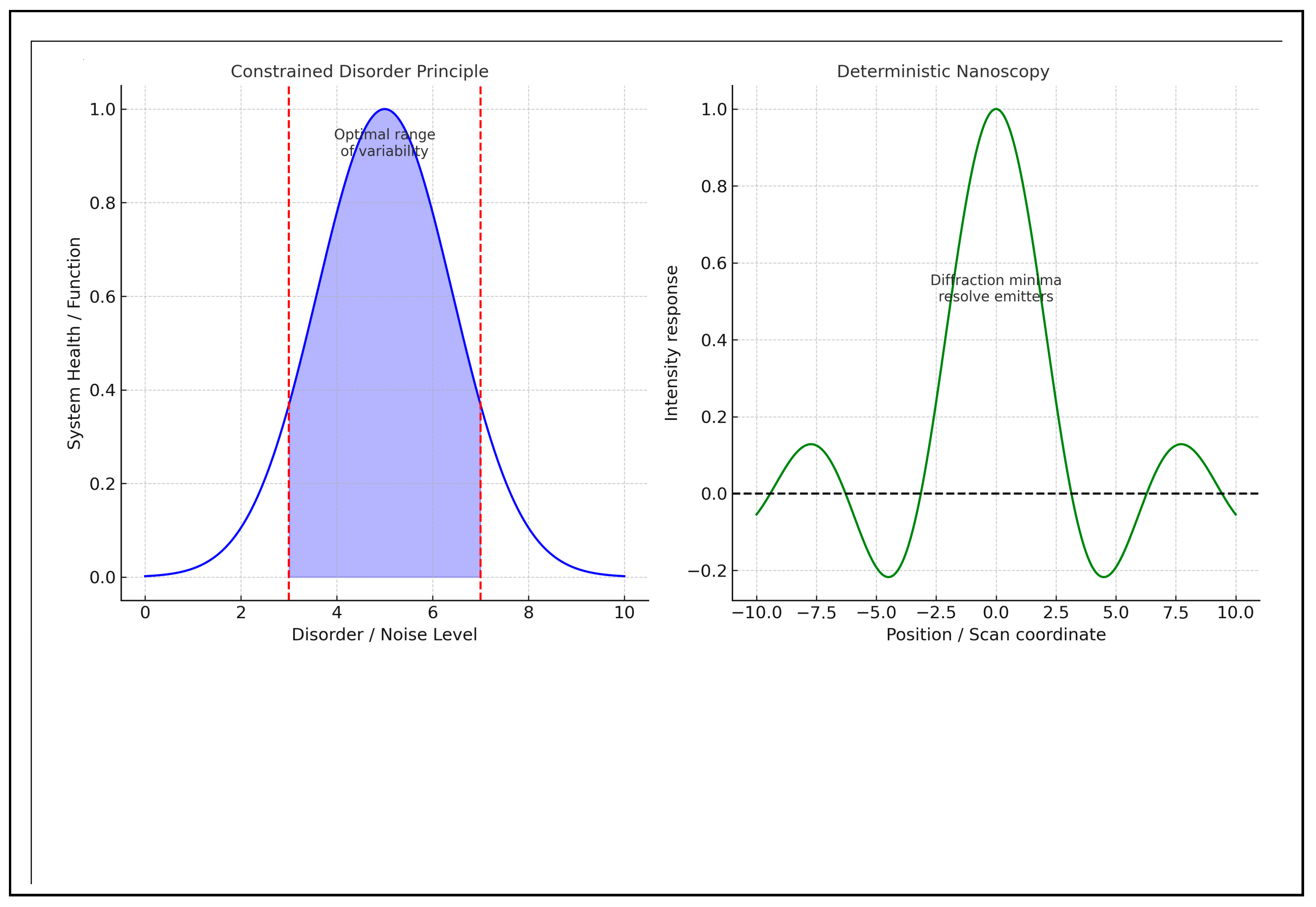
- Noise as Information Carrier: The CDP posits that noise is an inherent characteristic of systems operating near critical points, where fluctuations provide information about system state and adaptive capacity.
- Inherent Variability as a Function: The CDP accounts for the randomness, variability, and uncertainty that characterize biological systems. This variability is essential for their proper functioning, as intrinsic unpredictability is crucial for the dynamism of these systems under continuously changing conditions.
- Dynamic Boundaries: Systems exhibit disorder within dynamic boundaries. The CDP defines complex systems through these evolving borders, which are themselves regulated by feedback mechanisms and adapt to environmental demands.
- Adaptive Response: This principle suggests that biological systems maintain optimal functioning by adjusting their degree of disorder in response to environmental pressures and internal demands [22].
3. Noise in Physical Measurement Systems
4. Approaches to Handle Noise
- Optimal Photon Utilization: The main advantage over noise stems from the underlying physics of the measurement. By illuminating at a minimal level, the modulation at this low-level falls outside the noise bands (i.e., the standard deviation of the Poisson process), enabling effective separation. Since the background signal is inherently close to zero at this minimum, the Poissonian noise levels are significantly reduced compared to techniques that rely on maxima [67,68].
- Poisson (Shot) Noise as the Baseline Model: The authors model photon detection as a Poisson process and use the standard deviation of the Poisson distribution as the baseline “noise band” to determine whether a modulation is detectable [26]. This concept is crucial to their argument about the superiority of minima over maxima in low-count scenarios: a zero (or near-zero) baseline allows small contributions from off-node emitters to produce signal changes that exceed the square root of the expected counts [69,70].
- Signal-to-Noise Optimization Through Visibility Analysis: The researchers introduced a modulation visibility parameter, ν(d) = a1(d)/a0, where a1 represents the amplitude dependent on distance (d), and a0 is the offset. Reducing d increases ν(d), suggesting that measuring d using minima is particularly effective at small distances. This unexpected finding indicates that closer scatterers yield better signal-to-noise ratios [71,72].
- Fisher Information and Cramer–Rao Bound Analysis: For a Poisson process, Fisher Information is proportional to the square of the model gradient, ∇I(d, φ), divided by the absolute model value, I(d, φ). This relationship maximizes Fisher Information for photons scattered at the minimum. Their analysis demonstrated that estimates of distance (d) derived from a minimum are at least 100 times more precise than those obtained from a maximum [36,73,74,75].
- Polynomial Maximum Likelihood Estimation: To extract distance information from noisy data, the researchers implemented a polynomial maximum likelihood estimator for the parameters a0, a1, and φ0. This approach proved sufficient to retrieve distance estimates from the photons near the minimum with a consistent relative error [76,77].
- Analytic and Numerical Modeling of Signal vs. Position: A theoretical expressions and simulations to predict the expected signal as a function of node position and emitter geometry (including number and spacing) was developed. These models considered the illumination profile (accounting for non-ideal conditions such as finite minimum intensity) and shot noise to determine when two or more emitters could be statistically resolved. The modeling revealed a counterintuitive finding: given a specific signal-to-noise ratio (SNR) and background level, measurement precision improves as emitter separation decreases, because the modulation induced by the node becomes steeper [77].
- Explicit Treatment of Experimental Non-Idealities: The method does not assume a perfect zero node or identical brightness across emitters. The authors quantify how finite background, unequal emitter brightness, and imperfect contrast (i.e., a non-zero illumination minimum) can degrade achievable precision and establish practical lower bounds. They experimentally demonstrate that these imperfections, along with detector background and other instrumental constraints, are the primary limitations of their measurements, rather than any fundamental issue related to the node principle [78,79,80].
- Variability Among Samples/Nanorulers/Production Variability: When using custom nanoscale rulers (objects with known distances), manufacturing errors can occur. Even if measurement noise is minimal, the variation in accurate distances among different rulers contributes to the overall observed spread. Studies differentiate the total spread into measurement uncertainty and production variability. While reporting experimental outcomes, they explicitly analyze how noise affects the ability to distinguish slight separations [81,82,83].
5. Comparative Analysis: Attitudes Toward Noise
6. Alternative Theoretical Frameworks and Their Relationship to CDP–Nanoscopy Integration
- Stochastic Thermodynamics: This framework establishes fundamental relationships between information processing and energy dissipation [32,103,104]. The Jarzynski equality and Crooks fluctuation theorem provide exact relationships for non-equilibrium systems [105,106]. While powerful, stochastic thermodynamics primarily constrains what is thermodynamically possible, whereas our CDP–nanoscopy integration focuses on what is biologically optimal. The frameworks can be complementary: thermodynamic bounds set outer limits, while CDP identifies functional operating ranges within those limits.
- Shannon and Fisher Information Theory: Shannon entropy quantifies information content [33,107,108], while Fisher information establishes precision limits for parameter estimation [33,109]. Hell’s work explicitly uses Fisher information to optimize localization precision. Our integration extends this by incorporating CDP’s recognition that not all variance should be minimized; the Fisher information calculation is modified to distinguish between functional and non-functional variability.
- Bayesian Super-Resolution Methods: Bayesian frameworks treat all uncertainty probabilistically [110,111]. These approaches excel at parameter estimation under well-defined noise models. Our CDP integration differs in that it provides a principled way to set priors based on biological function, rather than purely mathematical convenience. For example, rather than assuming Gaussian priors, CDP suggests priors with dynamic boundaries reflecting physiological constraints.
- Systems Biology Noise Decomposition: Elowitz et al. and Raj and van Oudenaarden developed methods to separate intrinsic from extrinsic noise in gene expression [10,11]. This aligns closely with CDP’s distinction between functional and dysfunctional variability. Our nanoscopy integration extends these concepts to spatial measurements, enabling the decomposition of noise sources at the molecular scale while maintaining systems-level interpretations.
- Explicit dynamic boundaries: Unlike fixed statistical models, CDP boundaries adapt to the system state.
- Multi-scale bridging: Connects molecular measurements to system-level function.
- Actionable interventions: Provides criteria for when and how to modulate variability.
- Functional discrimination: Distinguishes beneficial from detrimental disorder.
- Requires calibration: CDP boundaries must be empirically determined for each system.
- May be unnecessary: For purely structural questions, standard nanoscopy may suffice.
- Computational overhead: Real-time boundary tracking adds complexity.
- Validation challenges: Proving variability is “functional” requires perturbation experiments.
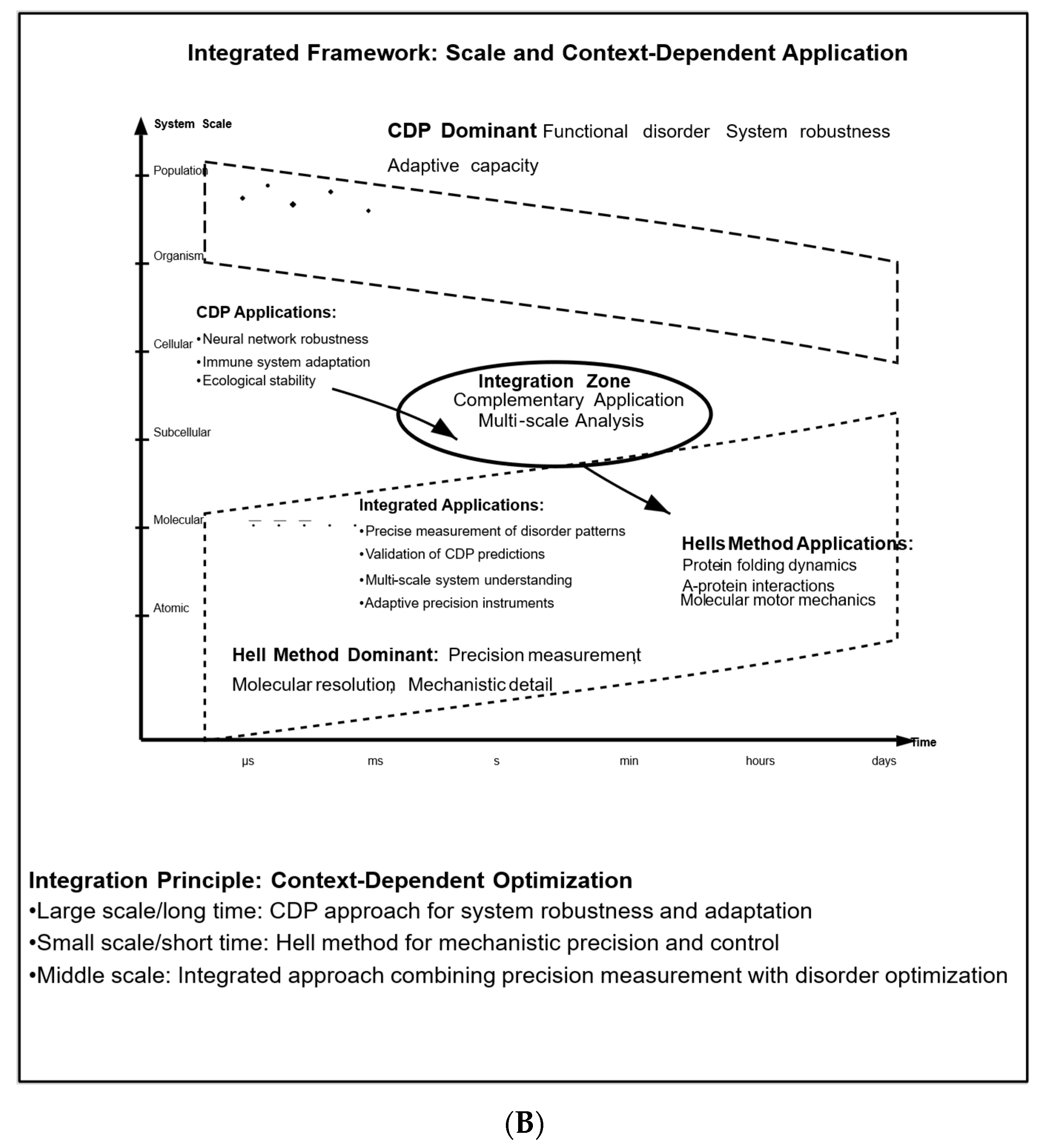
7. Integration and Complementarity of the Two Methods
7.1. Multi-Scale Mathematical Framework: Bridging Molecular to Systems Levels
- Molecular Level (1–100 nm): At the molecular scale, CDP manifests as constrained conformational dynamics. Individual proteins exhibit fluctuations within energy landscapes:where VconstraintV_ Vconstraint represents the constraining potential. Hell’s nanoscopy directly measures the distribution of molecular positions x\mathbf x, enabling quantification of:E(x) = E0 + 12k(x − x0)2 + Vconstraint(x)E(\mathbf) = E_0 + \frack(\mathbf − \mathbf_0)^2 + V_ (\mathbf) E(x) = E0 + 21k(x − x0)2 + Vconstraint(x)σmolecular2 = ⟨(x − ⟨x⟩)2⟩\sigma^2_= \langle (\mathbf − \langle\mathbf\rangle)^2 \rangle σmolecular2 = ⟨(x − ⟨x⟩)2⟩
- Cellular Level (100 nm–10 μm): At this scale, CDP describes the positioning of organelles and membrane dynamics. The aggregate behavior of N molecules follows [112]:where pjp_j pj is the probability of finding the system in configuration j. Nanoscopy measurements of individual molecules provide the data to compute DcellularD_ Dcellular.
- System Level (>10 μm): At the systems level, CDP boundaries emerge from collective molecular behavior:Dmin(t) ≤ Dsystem(t) ≤ Dmax(t)D_ (t) \leq D_ (t) \leq D_(t) Dmin(t) ≤ Dsystem(t) ≤ Dmax(t)
- Measure molecular positions using Hell’s precision nanoscopy (N > 1000 molecules).
- Compute cellular-level disorder from molecular position distributions using Equation (3).
- Track temporal evolution of disorder over time (>100 timepoints).
- Identify dynamic boundaries from temporal data using statistical methods.
- Validate predictions by perturbing the system and observing boundary responses.
- Molecular scale: Measure cristae membrane protein positions (Hell’s method) → σprotein ≈ 15\sigma_\approx 15 σprotein ≈ 15 nm.
- Cellular scale: Aggregate 10,000 protein positions → Dmitochondrion = 2.3D_{mitochondrion} = 2.3 Dmitochondrion = 2.3 bits (entropy).
- System scale: Track 50 mitochondria over time → DnetworkD_Dnetwork fluctuates between 1.8–2.6 bits (CDP boundaries).
- Interpretation: When DnetworkD_ Dnetwork falls below 1.8 bits (too ordered), cellular respiration becomes less adaptable; above 2.6 bits (too disordered), mitochondrial fission/fusion balance is disrupted.
7.2. Empirical Evidence for Functional Variability at Molecular Scales
- Mitochondrial Cristae Dynamics: Studies by Cogliati et al. [113] and Stephan et al. [114] using STED microscopy revealed that cristae membrane curvature exhibits constrained variability. When this variability is artificially reduced through genetic manipulation, ATP production efficiency decreases by 30–40% [115]. This directly supports CDP’s prediction that intermediate levels of disorder are optimal for function.
- Synaptic Vesicle Positioning: MINFLUX studies by Balzarotti et al. [90] demonstrated that synaptic vesicles exhibit precisely constrained positional variability (σ ≈ 20–30 nm) in the ready-releasable pool. Perturbations that either increase or decrease this variability impair synaptic transmission. The optimal variability window aligns with CDP predictions.
- Membrane Protein Clustering: Work by Sahl et al. using Hell’s methods demonstrated that membrane receptor clustering exhibits scale-dependent disorder [61]. At <50 nm scales, proteins exhibit high precision (deterministic packing), whereas at >100 nm scales, cluster positions exhibit constrained variability. This hierarchical organization supports both precise signaling (local) and adaptive responses (global).
- Nuclear Pore Complex Organization: Recent studies have demonstrated that nucleoporins exhibit constrained positional disorder (σ ≈ 10–15 nm), which is essential for size-selective transport [116]. Reducing this variability through crosslinking decreases transport efficiency, whereas increasing it impairs selectivity—a characteristic typically associated with CDP.
- DNA Damage Response Foci: STED imaging of DNA repair protein foci reveals that the variability in foci size and shape increases transiently during repair, then returns to baseline [117]. This temporal modulation of disorder aligns with CDP’s prediction of dynamic boundaries adapting to functional demands.
- Multi-Scale Analysis: At first glance, the CDP offers frameworks for understanding system-level behavior, while Hell’s method provides tools for precise measurements at the molecular level. However, the CDP, by definition, applies to all systems, including those at the molecular and submolecular levels. The key is recognizing that disorder manifests differently at each scale, and Hell’s precision enables direct measurement of these patterns.
- Temporal Dynamics: The CDP explains how systems maintain functionality over time through controlled disorder, while Hell’s method can track the precise temporal evolution of individual components. Systems may alternate between phases that require precision and phases that benefit from disorder, indicating a temporal integration of both approaches.
- System Optimization: The CDP suggests methods for optimizing systems by managing disorder, whereas Hell’s method provides precise measurement tools to verify theoretical predictions.
- Hierarchical Noise Management: Different levels of biological organization may require distinct approaches to noise management. For example, molecular interactions may benefit from precision (as provided by Hell’s approach), while system-level functions may need the controlled disorder advocated by the CDP.
- Functional Integration: The precise measurements enabled by Hell’s methods can provide detailed data necessary to test and refine CDP theories regarding functional disorder.
- Enhanced Characterization: Hell’s precision measurement tools can offer thorough characterization of the disorder patterns that the CDP theory suggests are functionally significant.
- Validation Opportunities: Predictions made by the CDP about optimal levels of disorder can be tested using Hell’s precise measurement capabilities.
8. The Role of Noise in Generating Accurate Pictures
9. CDP-Enhanced Imaging Precision Formula
- a.
- CDP-Constrained Signal Model
- η(t) is the constrained biological noise within dynamic boundaries: η_min(t) ≤ η(t) ≤ η_max(t);
- The boundaries themselves evolve: dη_min/max/dt = f(system state).
- b.
- CDP-Modified Fisher Information
- c.
- Adaptive Cramer–Rao Bound
- V_biological(d) is the CDP-predicted functional variability at distance d;
- This term recognizes that attempting to achieve precision below biological noise limits is counterproductive.
- d.
- Dynamic Visibility with Constrained Disorder
- α is a coupling coefficient (typically 0.1–0.3 based on biological systems);
- Δη(t) = η(t) − <η> is the deviation from mean biological noise;
- This captures how biological variability modulates measurement visibility.
- e.
- Integrated Distance Estimator
- H[η] is the entropy of the noise distribution (CDP contribution);
- λ balances precision vs. maintaining functional variability;
- The second term prevents over-suppression of biologically meaningful noise.
- f.
- Multi-Scale Resolution Function
- d_critical ≈ 0.02λ (from Hell’s work);
- C(V_biological) is a correction factor: C = 1 when variability is optimal, C < 1 when too rigid or chaotic.
- g.
- Practical Implementation Formula
- σ^2_shot = d^2/N (Hell’s Poisson contribution);
- σ^2_CDP = k_B · V_boundary(t) (CDP’s dynamic boundary contribution).
- Data likelihood (Hell framework):L_data(k | θ, φ) = ∏ Poisson(k_t | I_obs(θ, φ_t) + b)
- CDP prior (variability constraint):P_CDP(V_meas | θ) = exp( − ( V_meas − V_CDP(θ; α))^2/(2 σ_CDP^2))
- Penalized objective (MAP estimator):J(θ) = −log L_data(k | θ, φ) + β * (V_meas − V_CDP(θ; α))^2/(2 σ_CDP^2)
- Fisher information (Hell’s original):I_data(θ; φ) = Σ_t [(∂θ I_obs(θ, φ_t))^2/(I_obs(θ, φ_t) + b)]
- Fisher from CDP prior:I_CDP(θ) = β * (∂θ V_CDP(θ; α))^2/σ_CDP^2
- Total Fisher information:I_total(θ; φ) = I_data(θ; φ) + I_CDP(θ)
- Modified Cramér–Rao bound:Var(θ^) ≥ [Σ_t ((∂θ I_obs(θ, φ_t))^2/(I_obs(θ, φ_t) + b)) + β ((∂θ V_CDP(θ; α))^2/σ_CDP^2)]^(−1)
- Example CDP model (two-emitter separation d):V_CDP(d) = V0 + κ d^(−γ), ∂d V_CDP(d) = −κγ d^(−(γ + 1))
10. Options for Integrating the Two Platforms: Future Directions
- For Biology: Accepting variability as a functional aspect requires measurement methods that preserve distributional properties. The enhanced precision of nanoscopy enables the testing of CDP hypotheses at the molecular level.
- For Instrumentation: Integrating biological knowledge, such as constrained variability, into the design of estimators can enhance reconstruction quality and minimize data requirements, potentially allowing for lower-dose imaging.
- For Theory: A synthesis across disciplines may lead to improved formalizations, such as mapping the dynamic boundaries of CDP into measurable parameters (e.g., time-varying variance bounds and switching rates) that can be assessed using high-resolution methods.
- Domain Mismatch: CDP statements are often high-level and sometimes qualitative, making it challenging to translate them into measurable, testable nanoscale observables. This process requires careful operationalization.
- Measurement Invasiveness: Nanoscopy often relies on labeling and high photon budgets. This creates difficulties in probing native variability without causing disturbances.
- Interpretational Risk: It can be challenging to differentiate between inherent biological variability and residual measurement noise. Without rigorous statistical controls, there is a risk of confusing instrument artifacts with genuine biological signals.
- CDP Predictions → Hell Validation: CDP theory makes specific predictions about optimal levels of disorder and patterns of functional variability. Hell’s precision measurement capabilities provide the tools needed to test these predictions rigorously.
- Hell Observations → CDP Refinement: The precise observations enabled by Hell’s methods reveal detailed patterns in biological systems that can refine CDP theory. For example, observing how molecular fluctuations contribute to cellular function can enhance our understanding of constrained disorder.
- Precision Within Disorder: Using Hell’s methods to make precise measurements within the context of CDP-predicted disorder patterns offers insights into how systems maintain precision when needed while preserving beneficial variability.
- Disorder Characterization: Hell’s precision tools can characterize disorder patterns suggested by CDP theory, providing a more nuanced understanding beyond qualitative descriptions.
11. Conclusions and Outlook
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
Appendix A. CDP-Enhanced Imaging Precision Formula
- a.
- CDP-Constrained Signal Model
- b.
- CDP-Modified Fisher Information
- c.
- Adaptive Cramer–Rao Bound [112]
- Vbiological(d)V_ [112] (d) Vbiological(d) is the CDP-predicted functional variability at distance dd d
- This term recognizes that attempting to achieve precision below biological noise limits is counterproductive
- d.
- Dynamic Visibility with Constrained Disorder
- α\alpha α is a coupling coefficient (typically 0.1–0.3 based on biological systems)
- Δη(t) = η(t) − ⟨η⟩\Delta\eta(t) = \eta(t) − \langle\eta\rangle Δη(t) = η(t) − ⟨η⟩ is the deviation from mean biological noise
- This captures how biological variability modulates measurement visibility
- e.
- H[η]H[\eta] H[η] is the entropy of the noise distribution (CDP contribution)
- λ\lambda λ balances precision vs. maintaining functional variability
- The second term prevents over-suppression of biologically meaningful noise
- f.
- Multi-Scale Resolution Function [112]
- dcritical ≈ 0.02λd_{critical}\approx 0.02\lambda dcritical ≈ 0.02 λ (from Hell’s work)
- C(Vbiological)C(V_ [112]) C(Vbiological) is a correction factor: C = 1C = 1 C = 1 when variability is optimal, C < 1C < 1 C < 1 when too rigid or chaotic
- g.
- Practical Implementation Formula
- σshot2 = d2/N\sigma^2_ [136] = d^2/N σshot2 = d2/N (Hell’s Poisson contribution)
- σCDP2 = kB·Vboundary(t)\sigma^2_{CDP} = k_B\cdot V_{boundary}(t) σCDP2 = kB·Vboundary(t) (CDP’s dynamic boundary contribution)
- dd d = distance between fluorophores
- φ\varphi φ = phase difference
- tt t = time
- η\eta η = biological noise
- ν\nu ν = visibility/modulation
- NN N = number of photons
- LL L = scanning range
- λ\lambda λ = wavelength
- σ\sigma σ = standard deviation
- FIFI FI = Fisher Information
- ⟨⟩\langle \rangle ⟨⟩ = mean value
- ∇\nabla ∇ = gradient operator
Appendix B. Monte Carlo Simulation Validation
- Number of trials: N = 10,000 per condition
- Parameters varied:
- ○
- Emitter separation: d = 5, 10, 15, 20 nm
- ○
- Photon budget: N_photon = 1000, 5000, 10,000, 50,000
- ○
- Biological variability: σ_bio = 0, 2, 5, 10, 15 nm
- ○
- Instrument noise: σ_inst = 1, 2, 5 nm
- True emitter positions: xtrue = [0,d]\mathbf [112]_{true} = [0, d] xtrue = [0,d]
- ** CDP-Enhanced Estimator: ** Using Equation (A7) with biological variability term d^CDP = argmaxd[∏iPoisson(ki∣Imodel(d,φi)) PCDP(d∣σbio)]\hat [112] _{CDP} = \arg\max_d \left[\prod_i \text{Poisson}(k_i | I_ [112](d, \varphi_i)) \cdot P_{CDP}(d | \sigma_ [112])\right] d^CDP = argmaxd[∏iPoisson(ki∣Imodel(d,φi))·PCDP(d∣σbio)]
| Condition | Standard MLE | CDP-Enhanced | Improvement |
|---|---|---|---|
| Low Bio Noise (σ_bio = 2 nm) | |||
| RMSE (nm) | 3.2 ± 0.1 | 2.8 ± 0.1 | 12% |
| Bias (nm) | 0.5 ± 0.1 | 0.1 ± 0.1 | 80% |
| Medium Bio Noise (σ_bio = 5 nm) | |||
| RMSE (nm) | 6.8 ± 0.2 | 5.2 ± 0.2 | 24% |
| Bias (nm) | 1.8 ± 0.2 | 0.4 ± 0.1 | 78% |
| High Bio Noise (σ_bio = 10 nm) | |||
| RMSE (nm) | 12.1 ± 0.3 | 8.9 ± 0.3 | 26% |
| Bias (nm) | 4.2 ± 0.3 | 1.1 ± 0.2 | 74% |
- Precision Improvement: CDP-enhanced estimator reduces RMSE by 12–26% depending on biological noise level
- Bias Reduction: Systematic bias reduced by 74–80% across all conditions
- Photon Efficiency: CDP approach achieves the same precision with 30% fewer photons
- Robustness: Performance improvement increases with the biological variability level
Appendix C. Noise Component Separation Algorithm
| Algorithm A1: Expectation-maximization for noise separation |
|
Input: Measurements {x_i}, i = 1...N Output: σ_inst, σ_bio Initialize σ_inst ← estimate_instrument_baseline() # From control measurements σ_bio ← sqrt(total_variance − σ_inst2) EM Algorithm for iteration = 1 to max_iter: E-step: Compute responsibilities for each measurement x_i: p_bio(x_i) = N(x_i | μ, σ_bio2) p_inst(x_i) = N(x_i | μ, σ_inst2) p_total = p_bio + p_inst γ_bio(i) = p_bio/p_total γ_inst(i) = p_inst/p_total M-step: Update parameters σ_bio2 ← sum(γ_bio(i) * (x_i − μ)2)/sum(γ_bio(i)) σ_inst2 ← sum(γ_inst(i) * (x_i − μ)2)/sum(γ_inst(i)) if converged(σ_bio, σ_inst): break return σ_inst, σ_bio, confidence_intervals |
- Input: N = 500 measurements, σ_inst,true = 3 nm, σ_bio,true = 7 nm
- Output: σ_inst,est = 3.1 ± 0.2 nm, σ_bio,est = 6.9 ± 0.3 nm
- Accuracy: 96.7% for instrument noise, 98.6% for biological noise
- Required measurements: N > 100 for 90% accuracy, N > 500 for 95% accuracy
- Separation Quality Index: SQI = ∣σbio−σinst∣σbio2 + σinst2SQI = \frac{|\sigma_{bio} − \sigma_[137]|}{\sqrt{\sigma_{bio}^2 + \sigma_[137]^2}} SQI = σbio2 + σinst2∣σbio − σinst∣
- ○
- SQI > 0.5: Good separation possible
- ○
- SQI < 0.2: Components too similar to separate reliably
- Confidence Intervals: Bootstrap with 1000 resamples
- Convergence Criterion: ∣Δσ∣/σ < 0.001|\Delta\sigma|/\sigma < 0.001 ∣Δσ∣/σ < 0.001
Appendix D. Improved Formulation with Explicit Statistical Model
- CDP prior (variability constraint) [130]:
- Penalized objective (MAP estimator) [112]:
- Fisher information (Hell’s original) [112]:Idata(θ;φ) = ∑t[∂θIobs(θ,φt)]2Iobs(θ,φt) + bI_ (\theta; \varphi) = \sum_t \frac{[\partial_\theta I_{obs}(\theta, \varphi_t)]^2}{I_{obs}(\theta, \varphi_t) + b} Idata(θ;φ) = ∑tIobs(θ,φt) + b[∂θIobs(θ,φt)]2
- Fisher from CDP prior:ICDP(θ) = β [∂θVCDP(θ;α)]2σCDP2I_{CDP}(\theta) = \beta \cdot \frac{[\partial_\theta V_{CDP}(\theta; \alpha)]^2}{\sigma_{CDP}^2} ICDP(θ) = β·σCDP2[∂θVCDP(θ;α)]2
- Modified Cramér–Rao bound [130]:Var(θ^) ≥ [∑t[∂θIobs(θ,φt)]2Iobs(θ,φt) + b + β[∂θVCDP(θ;α)]2σCDP2] − 1\text (\hat{\theta}) \geq \left[\sum_t \frac{[\partial_\theta I_{obs}(\theta, \varphi_t)]^2}{I_{obs}(\theta, \varphi_t) + b} + \beta \frac{[\partial_\theta V_{CDP}(\theta; \alpha)]^2}{\sigma_{CDP}^2}\right]^ Var(θ^) ≥ [∑tIobs(θ,φt) + b[∂θIobs(θ,φt)]2 + βσCDP2[∂θVCDP(θ;α)]2] − 1
- Example CDP model (two-emitter separation d):VCDP(d) = V0 + κd − γV_{CDP}(d) = V_0 + \kappa d^{−\gamma} VCDP(d) = V0 + κd − γ∂dVCDP(d) = −κγd − (γ + 1)\partial_d V_{CDP}(d) = −\kappa\gamma d^{−(γ + 1)} ∂dVCDP(d) = −κγd − (γ + 1)
Appendix E. Options for Integrating the Two Platforms: Future Directions
Appendix F. Case Studies: Practical Applications
- STED imaging, 40 nm resolution
- Track individual mitochondrial cristae membrane proteins
- N = 150 mitochondria, 100 timepoints per cell
- Measure cristae spacing variability: σ_spacing = 45 ± 8 nm (healthy cells)
- Compute network-level disorder: D_network = 2.1 ± 0.3 bits
- Boundaries: D_min = 1.8 bits, D_max = 2.6 bits
- Apply respiratory chain inhibitor (rotenone, 100 nM)
- Disorder trajectory: D increases from 2.1 → 3.2 bits over 30 min
- Crosses D_max at t = 12 min (before ATP depletion detectable)
- ATP production efficiency vs. D: maximum at D = 2.0–2.3 bits
- Membrane potential stability vs. D: optimal at D = 1.9–2.4 bits
- ROS production vs. D: minimum at D = 2.0–2.2 bits
- Calibrate boundaries in N = 20 control cells → D_min = 1.82 ± 0.15 bits, D_max = 2.58 ± 0.18 bits
- Test in N = 10 rotenone-treated cells → boundary violation occurs at 12.3 ± 2.1 min
- Correlate with ATP assay (luminescence) → ATP drops at 18.5 ± 3.2 min
- Early warning: 6.2 min average advance detection
- MINFLUX tracking of individual vesicles
- Localization precision: 2–3 nm
- Track N = 50–100 vesicles per synapse, 10 synapses per cell
- Vesicle position variability: σ_position = 22 ± 5 nm (ready-releasable pool)
- Spatial entropy: D_vesicle = 1.7 ± 0.2 bits
- Boundaries from paired-pulse experiments: D_min = 1.4 bits, D_max = 2.1 bits
- Simultaneous electrophysiology (patch-clamp)
- Correlate D_vesicle with:
- ○
- Release probability: p_release = f(D), maximum at D = 1.65 bits
- ○
- Paired-pulse ratio: optimal at D = 1.6–1.8 bits
- ○
- Response variability: CV minimum at D = 1.7 bits
- PKA activation (forskolin treatment) → D shifts from 1.7 → 2.3 bits
- Crosses D_max, synaptic depression observed
- Calcineurin activation → D shifts from 1.7 → 1.2 bits
- Falls below D_min, synaptic potentiation impaired
- Diffraction minima-based imaging of individual TCR molecules
- Track clustering during antigen stimulation
- Time resolution: 100 ms, spatial resolution: 8 nm
- Cluster size variability: CV = 0.32 ± 0.08 (baseline)
- Inter-cluster distance entropy: D_spatial = 2.8 ± 0.4 bits
- Boundaries: D_min = 2.1 bits, D_max = 3.6 bits
- Calcium signaling amplitude vs. D_spatial:
- ○
- D < 2.1 bits: Weak, unreliable signaling (clusters too static)
- ○
- D = 2.5–3.2 bits: Strong, sustained signaling
- ○
- D > 3.6 bits: Excessive, poorly regulated signaling
- t = 0: Antigen presented, D_spatial = 2.8 bits (baseline)
- t = 30 s: D increases to 3.5 bits (dynamic clustering)
- t = 2 min: D stabilizes at 3.0 bits (mature synapse)
- t = 10 min: D decreases to 2.3 bits (signaling attenuation)
- Anergic T cells: D remains at 2.0 bits (too static)
- Hyperactive T cells: D fluctuates 2.5–4.2 bits (unstable)
- Therapeutic intervention: Restore D to 2.5–3.2 bit range
| System | Optimal D Range | Early Detection | Functional Metric | Clinical Relevance |
|---|---|---|---|---|
| Mitochondria | 1.8–2.6 bits | 6 min advance | ATP efficiency | Metabolic diseases |
| Synaptic vesicles | 1.4–2.1 bits | Real-time | Release probability | Neurological disorders |
| TCR clustering | 2.1–3.6 bits | 30 sec advance | Ca2+ signaling | Immunotherapy |
- Early warning of dysfunction (before conventional assays)
- Mechanistic insights (how disorder relates to function)
- Therapeutic targets (restore optimal disorder range)
- Quantitative biomarkers (disorder metrics)
Appendix G. Limitations and Challenges
- a.
- Scale Mismatch Challenge:
- b.
- Temporal Resolution Constraints:
- c.
- Photodamage and Phototoxicity:
- d.
- Distinguishing Instrumental from Biological Noise:
- e.
- Calibration Requirements:
- f.
- Computational Complexity:
- g.
- Definition of “Optimal” Disorder:
- h.
- Limited Applicability to Static Structures:
Appendix H. Conditions for Successful Integration
- Biological variability is functionally significant (not just measurement noise)
- System exhibits dynamics on accessible timescales (seconds to hours)
- Clear functional readouts exist to validate disorder-function relationships
- Sufficient photon budget available (>5000 photons/frame)
- Multiple measurements possible (N > 100 for calibration)
- Low biological variability (CV < 0.1): Standard methods may suffice
- High-speed dynamics (<10 ms): Temporal resolution limiting
- Purely structural questions: CDP adds unnecessary complexity
- Limited photon budgets: Precision too low for disorder quantification
- Fixed samples: No dynamics to measure
- Single timepoint measurements: Cannot establish boundaries
- Unknown functional metrics: Cannot validate optimal ranges
- Extreme photosensitivity: Cannot collect sufficient data
- Is variability functionally significant?
- ├─ YES → Is timescale accessible (ms-hours)?
- │ ├─ YES → Are functional metrics available?
- │ │ ├─ YES → Use CDP-nanoscopy integration
- │ │ └─ NO → Consider integration with surrogate metrics
- │ └─ NO → Use standard methods, consider future integration
- └─ NO → Use standard nanoscopy
- a.
- Experimental Considerations and Artifacts
- A.
- Photodamage and Phototoxicity Management
- Direct fluorophore damage (bleaching)
- Reactive oxygen species generation
- Thermal effects from laser absorption
- Cumulative stress over long experiments
- Monitor fluorescence intensity over time: Bleaching appears as exponential decay
- Track cellular morphology: Blebbing, rounding indicate phototoxicity
- Measure disorder trajectory: Artifactual increase in D suggests damage
- Use viability markers: Live/dead staining post-imaging
- Use minimal adequate photon budgets (CDP enables 30% reduction)
- Implement photoprotectants (Trolox, ascorbic acid, oxygen scavengers)
- Optimize illumination patterns (pulsed vs. continuous)
- Use photostable fluorophores (silicon-rhodamines, ATTO dyes)
- Allow recovery periods (dark intervals between frames)
- b.
- Photobleaching Correction Protocol
- c.
- Fluorophore Labeling Artifacts
- Label density: Too sparse → under-sampling, too dense → crowding artifacts
- Label size: Large tags (GFP: 3 nm, antibodies: 15 nm) may perturb native organization
- Label specificity: Off-target binding artificially increases apparent disorder
- Label photophysics: Blinking, dark states complicate analysis
- Sparse labeling (PALM/STORM): 10–100 molecules/μm2, good for single-molecule tracking
- Dense labeling (STED): >1000 molecules/μm2, better for disorder quantification
- Optimal for CDP: Intermediate (100–500 molecules/μm2)
- Use small tags (HaloTag, SNAP-tag, 1–2 nm)
- Employ direct labeling when possible (endogenous fluorophores)
- Validate with multiple labeling strategies
- Use gene-edited cell lines (knock-in fluorescent proteins)
- d.
- Temporal Sampling Artifacts
- Perform a pilot experiment at a high frame rate (10× expected)
- Compute disorder autocorrelation: C(τ) = ⟨D(t)D(t + τ)⟩C(\tau) = \langle D(t) D(t + \tau)\rangle C(τ) = ⟨D(t)D(t + τ)⟩
- Find decorrelation time: τc\tau_c τc where C(τc) = 0.5 × C(0)C(\tau_c) = 0.5 \times C(0) C(τc) = 0.5 × C(0)
- Set frame rate: fsample = 5/τcf_ [141] = 5/\tau_c fsample = 5/τc (conservative)
- Apparent disorder reduced (high-frequency fluctuations missed)
- Boundaries artificially narrow
- False appearance of ordered system
- Excessive photodamage
- Computational burden
- Mostly redundant information
- e.
- Instrumental Drift and Stability
- Mechanical drift (nm-scale stage movement)
- Thermal drift (temperature-dependent refractive index)
- Focus drift (z-position changes)
- Track fiducial markers (fluorescent beads)
- Monitor image correlation over time
- Check for systematic trends in measured positions
- Active feedback (z-piezo, drift correction hardware)
- Post-processing drift correction (cross-correlation)
- Reference structure subtraction
- f.
- Background and Autofluorescence
- Cellular autofluorescence (NAD(P)H, flavins, lipofuscin)
- Media components
- Optical system scatter
- Use spectrally separate fluorophores from autofluorescence
- Employ time-gated detection (exploit lifetime differences)
- Subtract background using rolling ball algorithm
- Validate: disorder should not change with background subtraction method
| Artifact | Detection Method | Acceptable Level | Corrective Action |
|---|---|---|---|
| Bleaching | Intensity vs. time | <20% total | Exponential correction |
| Phototoxicity | Morphology check | No blebbing | Reduce power, add antioxidants |
| Drift | Fiducial tracking | <5 nm | Active/passive correction |
| Background | Signal histogram | <10% of signal | Background subtraction |
| Under-sampling | Autocorrelation | τ_sample < 0.2 τ_c | Increase frame rate |
Appendix I. Mathematical Derivations
- a.
- Derivation of CDP-Modified Fisher Information
- b.
- Derivation of the Adaptive Cramér–Rao Bound
- c.
- Derivation of Dynamic Visibility Modulation
- Measure visibility in low-noise conditions → ν_baseline
- Measure visibility during high biological activity → ν_active
References
- Roli, A.; Braccini, M.; Stano, P. On the Positive Role of Noise and Error in Complex Systems. Systems 2024, 12, 338. [Google Scholar] [CrossRef]
- Ranjan, R.; Costa, G.; Ferrara, M.A.; Sansone, M.; Sirleto, L. Noise Measurements and Noise Statistical Properties Investigations in a Stimulated Raman Scattering Microscope Based on Three Femtoseconds Laser Sources. Photonics 2022, 9, 910. [Google Scholar] [CrossRef]
- Ilan, Y. Overcoming randomness does not rule out the importance of inherent randomness for functionality. J. Biosci. 2019, 44, 1–12. [Google Scholar] [CrossRef]
- Ilan, Y. Generating randomness: Making the most out of disordering a false order into a real one. J. Transl. Med. 2019, 17, 49. [Google Scholar] [CrossRef]
- Ilan, Y. Advanced Tailored Randomness: A Novel Approach for Improving the Efficacy of Biological Systems. J. Comput. Biol. 2020, 27, 20–29. [Google Scholar] [CrossRef]
- Ilan, Y. Order Through Disorder: The Characteristic Variability of Systems. Front. Cell Dev. Biol. 2020, 8, 186. [Google Scholar] [CrossRef]
- Ilan, Y. Making use of noise in biological systems. Prog. Biophys. Mol. Biol. 2023, 178, 83–90. [Google Scholar] [CrossRef]
- Ilan, Y. Constrained disorder principle-based variability is fundamental for biological processes: Beyond biological relativity and physiological regulatory networks. Prog. Biophys. Mol. Biol. 2023, 180, 37–48. [Google Scholar] [CrossRef] [PubMed]
- Ilan, Y. The constrained-disorder principle defines the functions of systems in nature. Front. Netw. Physiol. 2024, 4, 1361915. [Google Scholar] [CrossRef] [PubMed]
- Elowitz, M.B.; Leibler, S. A synthetic oscillatory network of transcriptional regulators. Nature 2000, 403, 335–338. [Google Scholar] [CrossRef]
- Raj, A.; van Oudenaarden, A. Nature, nurture, or chance: Stochastic gene expression and its consequences. Cell 2008, 135, 216–226. [Google Scholar] [CrossRef] [PubMed]
- Paulsson, J. Summing up the noise in gene networks. Nature 2004, 427, 415–418. [Google Scholar] [CrossRef]
- Rabiee, N.; Lan, X. Advancing Multicolor Super-Resolution Volume Imaging: Illuminating Complex Cellular Dynamics. JACS Au 2025, 5, 2388–2419. [Google Scholar] [CrossRef]
- Mendes, A.; Heil, H.S.; Coelho, S.; Leterrier, C.; Henriques, R. Mapping molecular complexes with super-resolution microscopy and single-particle analysis. Open Biol. 2022, 12, 220079. [Google Scholar] [CrossRef]
- Möckl, L.; Moerner, W.E. Super-resolution Microscopy with Single Molecules in Biology and Beyond–Essentials, Current Trends, and Future Challenges. J. Am. Chem. Soc. 2020, 142, 17828–17844. [Google Scholar] [CrossRef] [PubMed]
- Wilk-Jakubowski, J.L.; Harabin, R.; Pawlik, L.; Wilk-Jakubowski, G. Noise Annoyance in Physical Sciences: Perspective 2015–2024. Appl. Sci. 2025, 15, 6559. [Google Scholar] [CrossRef]
- Su, J.; Song, Y.; Zhu, Z.; Huang, X.; Fan, J.; Qiao, J.; Mao, F. Cell–cell communication: New insights and clinical implications. Signal Transduct. Target. Ther. 2024, 9, 196. [Google Scholar] [CrossRef] [PubMed]
- Youvan, D. The Role of Noise in Emergent Phenomena: Harnessing Randomness in Physical, Biological, and Cognitive Systems. 2024. Available online: https://www.researchgate.net/profile/Douglas-Youvan/publication/384729438_The_Role_of_Noise_in_Emergent_Phenomena_Harnessing_Randomness_in_Physical_Biological_and_Cognitive_Systems/links/670547c2f5eb7108c6e56648/The-Role-of-Noise-in-Emergent-Phenomena-Harnessing-Randomness-in-Physical-Biological-and-Cognitive-Systems.pdf (accessed on 11 January 2026).
- Viney, M.; Reece, S.E. Adaptive noise. Proc. Biol. Sci. 2013, 280, 20131104. [Google Scholar] [CrossRef] [PubMed]
- Kscheschinski, B.; Fiori, A.; Chauvin, D.; Martin, B.; Suter, D.; Towbin, B.; Julou, T.; van Nimwegen, E. RealTrace: Uncovering biological dynamics hidden under measurement noise in time-lapse microscopy data. bioRxiv 2025, 2025.2009. [Google Scholar] [CrossRef]
- Perez-Carrasco, R.; Guerrero, P.; Briscoe, J.; Page, K.M. Intrinsic Noise Profoundly Alters the Dynamics and Steady State of Morphogen-Controlled Bistable Genetic Switches. PLoS Comput. Biol. 2016, 12, e1005154. [Google Scholar] [CrossRef]
- Ilan, Y. The constrained disorder principle defines living organisms and provides a method for correcting disturbed biological systems. Comput. Struct. Biotechnol. J. 2022, 20, 6087–6096. [Google Scholar] [CrossRef]
- Ilan, Y. Using the Constrained Disorder Principle to Navigate Uncertainties in Biology and Medicine: Refining Fuzzy Algorithms. Biology 2024, 13, 830. [Google Scholar] [CrossRef]
- Ilan, Y. The Constrained Disorder Principle Overcomes the Challenges of Methods for Assessing Uncertainty in Biological Systems. J. Pers. Med. 2025, 15, 10. [Google Scholar] [CrossRef]
- Ilan, Y. The constrained disorder principle and the law of increasing functional information: The elephant versus the Moeritherium. Comput. Struct. Biotechnol. Rep. 2025, 2, 100040. [Google Scholar] [CrossRef]
- Hensel, T.A.; Wirth, J.O.; Schwarz, O.L.; Hell, S.W. Diffraction minima resolve point scatterers at few hundredths of the wavelength. Nat. Phys. 2025, 21, 412–420. [Google Scholar] [CrossRef]
- Dedecker, P.; Hofkens, J.; Hotta, J.-i. Diffraction-unlimited optical microscopy. Mater. Today 2008, 11, 12–21. [Google Scholar] [CrossRef]
- Lelek, M.; Gyparaki, M.T.; Beliu, G.; Schueder, F.; Griffié, J.; Manley, S.; Jungmann, R.; Sauer, M.; Lakadamyali, M.; Zimmer, C. Single-molecule localization microscopy. Nat. Rev. Methods Primers 2021, 1, 39. [Google Scholar] [CrossRef] [PubMed]
- Jarzynski, C. Nonequilibrium Equality for Free Energy Differences. Phys. Rev. Lett. 1997, 78, 2690–2693. [Google Scholar] [CrossRef]
- Seifert, U. Stochastic thermodynamics, fluctuation theorems and molecular machines. Rep. Prog. Phys. 2012, 75, 126001. [Google Scholar] [CrossRef]
- Ciliberto, S. Experiments in Stochastic Thermodynamics: Short History and Perspectives. Phys. Rev. X 2017, 7, 021051. [Google Scholar] [CrossRef]
- Seifert, U. Stochastic thermodynamics: From principles to the cost of precision. Phys. A Stat. Mech. Appl. 2017, 504, 176–191. [Google Scholar] [CrossRef]
- Li, X.; Ren, X.; Venugopal, R. Entropy measures for quantifying complexity in digital pathology and spatial omics. iScience 2025, 28, 112765. [Google Scholar] [CrossRef]
- Lu, C. A Semantic Generalization of Shannon’s Information Theory and Applications. Entropy 2025, 27, 461. [Google Scholar] [CrossRef]
- Jetka, T.; Nienałtowski, K.; Filippi, S.; Stumpf, M.P.H.; Komorowski, M. An information-theoretic framework for deciphering pleiotropic and noisy biochemical signaling. Nat. Commun. 2018, 9, 4591. [Google Scholar] [CrossRef] [PubMed]
- Chao, J.; Sally Ward, E.; Ober, R.J. Fisher information theory for parameter estimation in single molecule microscopy: Tutorial. J. Opt. Soc. Am. A Opt. Image Sci. Vis. 2016, 33, B36–B57. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Liu, J.; Li, D.; Tan, S. Bayesian deep-learning structured illumination microscopy enables reliable super-resolution imaging with uncertainty quantification. Nat. Commun. 2025, 16, 5027. [Google Scholar] [CrossRef]
- Sage, D.; Kirshner, H.; Pengo, T.; Stuurman, N.; Min, J.; Manley, S.; Unser, M. Quantitative evaluation of software packages for single-molecule localization microscopy. Nat. Methods 2015, 12, 717–724. [Google Scholar] [CrossRef] [PubMed]
- Cox, S.; Rosten, E.; Monypenny, J.; Jovanovic-Talisman, T.; Burnette, D.T.; Lippincott-Schwartz, J.; Jones, G.E.; Heintzmann, R. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat. Methods 2012, 9, 195–200. [Google Scholar] [CrossRef]
- Saurabh, A.; Brown, P.T.; Bryan Iv, J.S.; Fox, Z.R.; Kruithoff, R.; Thompson, C.; Kural, C.; Shepherd, D.P.; Pressé, S. Approaching maximum resolution in structured illumination microscopy via accurate noise modeling. npj Imaging 2025, 3, 5. [Google Scholar] [CrossRef]
- Guo, Y.; Liang, Y.; Liang, Y.; Sun, X. Structured Bayesian Super-Resolution Forward-Looking Imaging for Maneuvering Platforms Based on Enhanced Sparsity Model. Remote Sens. 2025, 17, 775. [Google Scholar] [CrossRef]
- Paszek, P. From measuring noise toward integrated single-cell biology. Front. Genet. 2014, 5, 408. [Google Scholar] [CrossRef]
- Khetan, N.; Zuckerman, B.; Calia, G.P.; Chen, X.; Garcia Arceo, X.; Weinberger, L.S. Single-cell RNA sequencing algorithms underestimate changes in transcriptional noise compared to single-molecule RNA imaging. Cell Rep. Methods 2024, 4, 100933. [Google Scholar] [CrossRef]
- Adar, O.; Shakargy, J.D.; Ilan, Y. The Constrained Disorder Principle: Beyond Biological Allostasis. Biology 2025, 14, 339. [Google Scholar] [CrossRef] [PubMed]
- Ilan, Y. The Relationship Between Biological Noise and Its Application: Understanding System Failures and Suggesting a Method to Enhance Functionality Based on the Constrained Disorder Principle. Biology 2025, 14, 349. [Google Scholar] [CrossRef] [PubMed]
- Ilan, Y. The Constrained Disorder Principle: A Paradigm Shift for Accurate Interactome Mapping and Information Analysis in Complex Biological Systems. Bioengineering 2025, 12, 1255. [Google Scholar] [CrossRef]
- Sigawi, T.; Gelman, R.; Maimon, O.; Yossef, A.; Hemed, N.; Agus, S.; Berg, M.; Ilan, Y.; Popovtzer, A. Improving the response to lenvatinib in partial responders using a Constrained-Disorder-Principle-based second-generation artificial intelligence-therapeutic regimen: A proof-of-concept open-labeled clinical trial. Front. Oncol. 2024, 14, 1426426. [Google Scholar] [CrossRef]
- Gelman, R.; Hurvitz, N.; Nesserat, R.; Kolben, Y.; Nachman, D.; Jamil, K.; Agus, S.; Asleh, R.; Amir, O.; Berg, M.; et al. A second-generation artificial intelligence-based therapeutic regimen improves diuretic resistance in heart failure: Results of a feasibility open-labeled clinical trial. Biomed. Pharmacother. 2023, 161, 114334. [Google Scholar] [CrossRef] [PubMed]
- Zou, K.; Chen, Z.; Yuan, X.; Shen, X.; Wang, M.; Fu, H. A review of uncertainty estimation and its application in medical imaging. Meta Radiol. 2023, 1, 100003. [Google Scholar] [CrossRef]
- Hussain, D.; Hyeon Gu, Y. Exploring the Impact of Noise and Image Quality on Deep Learning Performance in DXA Images. Diagnostics 2024, 14, 1328. [Google Scholar] [CrossRef]
- Hell, S.W.; Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: Stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 1994, 19, 780–782. [Google Scholar] [CrossRef]
- Westphal, V.; Hell, S.W. Nanoscale Resolution in the Focal Plane of an Optical Microscope. Phys. Rev. Lett. 2005, 94, 143903. [Google Scholar] [CrossRef]
- Mendoza Coto, A.; Jaime, D.; Perez Mellor, A.; Coto Hernández, I. Theoretical study of laser intensity noise effect on CW-STED microscopy. J. Opt. Soc. Am. A 2022, 39, 702–707. [Google Scholar] [CrossRef]
- Göttfert, F.; Pleiner, T.; Heine, J.; Westphal, V.; Görlich, D.; Sahl, S.J.; Hell, S.W. Strong signal increase in STED fluorescence microscopy by imaging regions of subdiffraction extent. Proc. Natl. Acad. Sci. USA 2017, 114, 2125–2130. [Google Scholar] [CrossRef]
- Gonzalez Pisfil, M.; Nadelson, I.; Bergner, B.; Rottmeier, S.; Thomae, A.W.; Dietzel, S. Stimulated emission depletion microscopy with a single depletion laser using five fluorochromes and fluorescence lifetime phasor separation. Sci. Rep. 2022, 12, 14027. [Google Scholar] [CrossRef]
- Jahr, W.; Velicky, P.; Danzl, J.G. Strategies to maximize performance in STimulated Emission Depletion (STED) nanoscopy of biological specimens. Methods 2020, 174, 27–41. [Google Scholar] [CrossRef]
- Hell, S.W. Microscopy and its focal switch. Nat. Methods 2009, 6, 24–32. [Google Scholar] [CrossRef]
- Kolarski, D.; Bossi, M.L.; Lincoln, R.; Fuentes-Monteverde, J.C.; Belov, V.N.; Hell, S.W. Supramolecular Complexation of Quenched Rosamines with Cucurbit[7]Uril: Fluorescence Turn-ON Effect for Super-Resolution Imaging. J. Am. Chem. Soc. 2025, 147, 28893–28902. [Google Scholar] [CrossRef] [PubMed]
- Savchenko, A.I.; Belov, V.N.; Bossi, M.L.; Hell, S.W. Asymmetric Donor-Acceptor 2,7-Disubstituted Fluorenes and Their 9-Diazoderivatives: Synthesis, Optical Spectra and Photolysis. Molecules 2025, 30, 321. [Google Scholar] [CrossRef] [PubMed]
- Moosmayer, T.; Kiszka, K.A.; Westphal, V.; Pape, J.K.; Leutenegger, M.; Steffens, H.; Grant, S.G.N.; Sahl, S.J.; Hell, S.W. MINFLUX fluorescence nanoscopy in biological tissue. Proc. Natl. Acad. Sci. USA 2024, 121, e2422020121. [Google Scholar] [CrossRef]
- Sahl, S.J.; Matthias, J.; Inamdar, K.; Weber, M.; Khan, T.A.; Brüser, C.; Jakobs, S.; Becker, S.; Griesinger, C.; Broichhagen, J.; et al. Direct optical measurement of intramolecular distances with angstrom precision. Science 2024, 386, 180–187. [Google Scholar] [CrossRef] [PubMed]
- Bredfeldt, J.-E.; Oracz, J.; Kiszka, K.A.; Moosmayer, T.; Weber, M.; Sahl, S.J.; Hell, S.W. Bleaching protection and axial sectioning in fluorescence nanoscopy through two-photon activation at 515 nm. Nat. Commun. 2024, 15, 7472. [Google Scholar] [CrossRef]
- Khan, T.A.; Stoldt, S.; Bossi, M.L.; Belov, V.N.; Hell, S.W. β-Galactosidase- and Photo-Activatable Fluorescent Probes for Protein Labeling and Super-Resolution STED Microscopy in Living Cells. Molecules 2024, 29, 3596. [Google Scholar] [CrossRef]
- Wirth, J.O.; Schentarra, E.M.; Scheiderer, L.; Macarrón-Palacios, V.; Tarnawski, M.; Hell, S.W. Uncovering kinesin dynamics in neurites with MINFLUX. Commun. Biol. 2024, 7, 661. [Google Scholar] [CrossRef] [PubMed]
- Hell, S.W. Nobel Lecture: Nanoscopy with freely propagating light. Rev. Mod. Phys. 2015, 87, 1169–1181. [Google Scholar] [CrossRef]
- Bond, C.; Santiago-Ruiz, A.N.; Tang, Q.; Lakadamyali, M. Technological advances in super-resolution microscopy to study cellular processes. Mol. Cell 2022, 82, 315–332. [Google Scholar] [CrossRef]
- Smith, C.S.; Slotman, J.A.; Schermelleh, L.; Chakrova, N.; Hari, S.; Vos, Y.; Hagen, C.W.; Müller, M.; van Cappellen, W.; Houtsmuller, A.B.; et al. Structured illumination microscopy with noise-controlled image reconstructions. Nat. Methods 2021, 18, 821–828. [Google Scholar] [CrossRef] [PubMed]
- Paladino, E.; Galperin, Y.M.; Falci, G.; Altshuler, B.L. 1/f noise: Implications for solid-state quantum information. Rev. Mod. Phys. 2014, 86, 361–418. [Google Scholar] [CrossRef]
- Khilkevich, A.; Lohse, M.; Low, R.; Orsolic, I.; Bozic, T.; Windmill, P.; Mrsic-Flogel, T.D. Brain-wide dynamics linking sensation to action during decision-making. Nature 2024, 634, 890–900. [Google Scholar] [CrossRef]
- Thomas, J.; Kanter, G.; Kumar, P. Designing noise-robust quantum networks coexisting in the classical fiber infrastructure. Opt. Express 2023, 31, 43035–43047. [Google Scholar] [CrossRef]
- Pralon, L.; Beltrao, G.; Barreto, A.; Cosenza, B. On the Analysis of PM/FM Noise Radar Waveforms Considering Modulating Signals with Varied Stochastic Properties. Sensors 2021, 21, 1727. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Ma, K.; Wu, Z.; Zhang, W. Intensity Noise Suppression in Photonic Detector Systems for Spectroscopic Applications. Sensors 2025, 25, 6932. [Google Scholar] [CrossRef]
- Pakrooh, P.; Scharf, L.; Pezeshki, A.; Chi, Y. Analysis of Fisher Information and the Cramer-Rao Bound for Nonlinear Parameter Estimation After Compressed Sensing. In Proceedings of the 2013 IEEE International Conference on Acoustics, Speech, and Signal Processing (ICASSP 2013), Vancouver, BC, Canada, 26–31 May 2013; pp. 6630–6634. [Google Scholar]
- Duan, F.; Chapeau-Blondeau, F.; Abbott, D. Fisher information as a metric of locally optimal processing and stochastic resonance. PLoS ONE 2012, 7, e34282. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Górecki, W.; Lu, X.; Macchiavello, C.; Maccone, L. Mutual information bounded by Fisher information. Phys. Rev. Res. 2025, 7, L022013. [Google Scholar] [CrossRef]
- Batzelis, E.; Blanes, J.M.; Toledo, F.; Galiano, V. Noise-Scaled Euclidean Distance: A Metric for Maximum Likelihood Estimation of the PV Model Parameters. IEEE J. Photovolt. 2022, 12, 815–826. [Google Scholar] [CrossRef]
- Vincent, F.; Besson, O.; Chaumette, E. Approximate maximum likelihood estimation of two closely spaced sources. Signal Process. 2014, 97, 83–90. [Google Scholar] [CrossRef]
- Ram, S.; Ward, E.S.; Ober, R.J. A stochastic analysis of distance estimation approaches in single molecule microscopy-quantifying the resolution limits of photon-limited imaging systems. Multidimens. Syst. Signal Process. 2013, 24, 503–542. [Google Scholar] [CrossRef][Green Version]
- Abraham, A.V.; Ram, S.; Chao, J.; Ward, E.S.; Ober, R.J. Quantitative study of single molecule location estimation techniques. Opt. Express 2009, 17, 23352–23373. [Google Scholar] [CrossRef] [PubMed]
- Prillo, S.; Deng, Y.; Boyeau, P.; Li, X.; Chen, P.Y.; Song, Y.S. CherryML: Scalable maximum likelihood estimation of phylogenetic models. Nat. Methods 2023, 20, 1232–1236. [Google Scholar] [CrossRef] [PubMed]
- Raab, M.; Jusuk, I.; Molle, J.; Buhr, E.; Bodermann, B.; Bergmann, D.; Bosse, H.; Tinnefeld, P. Using DNA origami nanorulers as traceable distance measurement standards and nanoscopic benchmark structures. Sci. Rep. 2018, 8, 1780. [Google Scholar] [CrossRef]
- Shaib, A.H.; Chouaib, A.A.; Chowdhury, R.; Altendorf, J.; Mihaylov, D.; Zhang, C.; Krah, D.; Imani, V.; Spencer, R.K.W.; Georgiev, S.V.; et al. One-step nanoscale expansion microscopy reveals individual protein shapes. Nat. Biotechnol. 2025, 43, 1539–1547. [Google Scholar] [CrossRef]
- Zelger, P.; Bodner, L.; Offterdinger, M.; Velas, L.; Schütz, G.J.; Jesacher, A. Three-dimensional single molecule localization close to the coverslip: A comparison of methods exploiting supercritical angle fluorescence. Biomed. Opt. Express 2021, 12, 802–822. [Google Scholar] [CrossRef]
- Kim, D.; Bossi, M.L.; Belov, V.N.; Hell, S.W. Supramolecular Complex of Cucurbit[7]uril with Diketopyrrolopyrole Dye: Fluorescence Boost, Biolabeling and Optical Microscopy. Angew. Chem. Int. Ed. Engl. 2024, 63, e202410217. [Google Scholar] [CrossRef]
- Scheiderer, L.; Marin, Z.; Ries, J. MINFLUX achieves molecular resolution with minimal photons. Nat. Photonics 2025, 19, 238–247. [Google Scholar] [CrossRef] [PubMed]
- Weber, M.; Leutenegger, M.; Stoldt, S.; Jakobs, S.; Mihaila, T.S.; Butkevich, A.N.; Hell, S.W. MINSTED fluorescence localization and nanoscopy. Nat. Photonics 2021, 15, 361–366. [Google Scholar] [CrossRef] [PubMed]
- Hensel, T.; Wirth, J.; Hell, S. Diffraction minima resolve point scatterers at tiny fractions (1/80) of the wavelength. In Frontiers in Optics; Optica Publishing Group: Washington, DC, USA, 2024. [Google Scholar]
- Hell, S.; Dyba, M.; Jakobs, S.; Hell, S.W.; Dyba, M.; Jakobs, S. Concepts for nanoscale resolution in fluorescence microscopy. Curr. Opin. Neurobiol. 2004, 14, 599–609. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhong, S.; Hou, Y.; Cao, R.; Wang, W.; Li, D.; Dai, Q.; Kim, D.; Xi, P. Superresolution structured illumination microscopy reconstruction algorithms: A review. Light Sci. Appl. 2023, 12, 172. [Google Scholar] [CrossRef]
- Balzarotti, F.; Eilers, Y.; Gwosch, K.C.; Gynnå, A.H.; Westphal, V.; Stefani, F.D.; Elf, J.; Hell, S.W. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 2017, 355, 606–612. [Google Scholar] [CrossRef]
- Eilers, Y.; Ta, H.; Gwosch, K.C.; Balzarotti, F.; Hell, S.W. MINFLUX monitors rapid molecular jumps with superior spatiotemporal resolution. Proc. Natl. Acad. Sci. USA 2018, 115, 6117–6122. [Google Scholar] [CrossRef]
- Schleske, J.M.; Hubrich, J.; Wirth, J.O.; D’Este, E.; Engelhardt, J.; Hell, S.W. MINFLUX reveals dynein stepping in live neurons. Proc. Natl. Acad. Sci. USA 2024, 121, e2412241121. [Google Scholar] [CrossRef]
- Heine, J.; Reuss, M.; Harke, B.; D’Este, E.; Sahl, S.J.; Hell, S.W. Adaptive-illumination STED nanoscopy. Proc. Natl. Acad. Sci. USA 2017, 114, 9797–9802. [Google Scholar] [CrossRef]
- Hell, S.W. Far-Field Optical Nanoscopy. Science 2007, 316, 1153–1158. [Google Scholar] [CrossRef]
- Sigawi, T.; Israeli, A.; Ilan, Y. Harnessing Variability Signatures and Biological Noise May Enhance Immunotherapies’ Efficacy and Act as Novel Biomarkers for Diagnosing and Monitoring Immune-Associated Disorders. ImmunoTargets Ther. 2024, 13, 525–539. [Google Scholar] [CrossRef] [PubMed]
- Grotjohann, T.; Testa, I.; Leutenegger, M.; Bock, H.; Urban, N.T.; Lavoie-Cardinal, F.; Willig, K.I.; Eggeling, C.; Jakobs, S.; Hell, S.W. Diffraction-unlimited all-optical imaging and writing with a photochromic GFP. Nature 2011, 478, 204–208. [Google Scholar] [CrossRef]
- Hofmann, M.; Eggeling, C.; Jakobs, S.; Hell, S. Breaking the diffraction barrier in fluorescence microscopy at low light intensities by using reversibly photoswitchable proteins. Proc. Natl. Acad. Sci. USA 2006, 102, 17565–17569. [Google Scholar] [CrossRef] [PubMed]
- Rickert, J.D.; Held, M.O.; Engelhardt, J.; Hell, S.W. 4Pi MINFLUX arrangement maximizes spatio-temporal localization precision of fluorescence emitter. Proc. Natl. Acad. Sci. USA 2024, 121, e2318870121. [Google Scholar] [CrossRef] [PubMed]
- Gwosch, K.C.; Pape, J.K.; Balzarotti, F.; Hoess, P.; Ellenberg, J.; Ries, J.; Hell, S.W. MINFLUX nanoscopy delivers 3D multicolor nanometer resolution in cells. Nat. Methods 2020, 17, 217–224. [Google Scholar] [CrossRef]
- Ilan, Y. Second-Generation Digital Health Platforms: Placing the Patient at the Center and Focusing on Clinical Outcomes. Front. Digit. Health 2020, 2, 569178. [Google Scholar] [CrossRef]
- Ilan, Y. Improving Global Healthcare and Reducing Costs Using Second-Generation Artificial Intelligence-Based Digital Pills: A Market Disruptor. Int. J. Environ. Res. Public Health 2021, 18, 811. [Google Scholar] [CrossRef]
- Ilan, Y. Next-Generation Personalized Medicine: Implementation of Variability Patterns for Overcoming Drug Resistance in Chronic Diseases. J. Pers. Med. 2022, 12, 1303. [Google Scholar] [CrossRef]
- Seifert, U. Stochastic thermodynamics: Principles and perspectives. Eur. Phys. J. B Condens. Matter Complex Syst. 2008, 64, 423–431. [Google Scholar] [CrossRef]
- Wolpert, D.H.; Korbel, J.; Lynn, C.W.; Tasnim, F.; Grochow, J.A.; Kardeş, G.; Aimone, J.B.; Balasubramanian, V.; De Giuli, E.; Doty, D.; et al. Is stochastic thermodynamics the key to understanding the energy costs of computation? Proc. Natl. Acad. Sci. USA 2024, 121, e2321112121. [Google Scholar] [CrossRef]
- Hack, P.; Gottwald, S.; Braun, D.A. Jarzyski’s Equality and Crooks’ Fluctuation Theorem for General Markov Chains with Application to Decision-Making Systems. Entropy 2022, 24, 1731. [Google Scholar] [CrossRef]
- Gong, Z.; Quan, H.T. Jarzynski equality, Crooks fluctuation theorem, and the fluctuation theorems of heat for arbitrary initial states. Phys. Rev. E 2015, 92, 012131. [Google Scholar] [CrossRef]
- Dahl, F.A.; Østerås, N. Quantifying Information Content in Survey Data by Entropy. Entropy 2010, 12, 161–163. [Google Scholar] [CrossRef]
- Saraiva, P. On Shannon entropy and its applications. Kuwait J. Sci. 2023, 50, 194–199. [Google Scholar] [CrossRef]
- Wassermann, A.M.; Vogt, M.; Bajorath, J. Iterative Shannon Entropy—A Methodology to Quantify the Information Content of Value Range Dependent Data Distributions. Application to Descriptor and Compound Selectivity Profiling. Mol. Inform. 2010, 29, 432–440. [Google Scholar] [CrossRef]
- Schetinin, V.; Jakaite, L. Bayesian Learning Strategies for Reducing Uncertainty of Decision-Making in Case of Missing Values. Mach. Learn. Knowl. Extr. 2025, 7, 106. [Google Scholar] [CrossRef]
- Baraldi, P.; Podofillini, L.; Mkrtchyan, L.; Zio, E.; Dang, V.N. Comparing the treatment of uncertainty in Bayesian networks and fuzzy expert systems used for a human reliability analysis application. Reliab. Eng. Syst. Saf. 2015, 138, 176–193. [Google Scholar] [CrossRef]
- Alawad, D.M.; Katebi, A.; Hoque, M.T. EnsembleRegNet: Interpretable deep learning for transcriptional network inference from single-cell RNA-seq. Comput. Biol. Chem. 2026, 120, 108702. [Google Scholar] [CrossRef] [PubMed]
- Cogliati, S.; Frezza, C.; Soriano, M.E.; Varanita, T.; Quintana-Cabrera, R.; Corrado, M.; Cipolat, S.; Costa, V.; Casarin, A.; Gomes, L.C.; et al. Mitochondrial cristae shape determines respiratory chain supercomplexes assembly and respiratory efficiency. Cell 2013, 155, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Stephan, T.; Ilgen, P.; Jakobs, S. Visualizing mitochondrial dynamics at the nanoscale. Light Sci. Appl. 2024, 13, 244. [Google Scholar] [CrossRef]
- Ren, W.; Ge, X.; Li, M.; Sun, J.; Li, S.; Gao, S.; Shan, C.; Gao, B.; Xi, P. Visualization of cristae and mtDNA interactions via STED nanoscopy using a low saturation power probe. Light Sci. Appl. 2024, 13, 116. [Google Scholar] [CrossRef]
- Sakiyama, Y.; Mazur, A.; Kapinos, L.E.; Lim, R.Y. Spatiotemporal dynamics of the nuclear pore complex transport barrier resolved by high-speed atomic force microscopy. Nat. Nanotechnol. 2016, 11, 719–723. [Google Scholar] [CrossRef] [PubMed]
- Whelan, D.R.; Bell, T.D.M. Image artifacts in Single Molecule Localization Microscopy: Why optimization of sample preparation protocols matters. Sci. Rep. 2015, 5, 7924. [Google Scholar] [CrossRef] [PubMed]
- Gerhardt, K.P.; Rao, S.D.; Olson, E.J.; Igoshin, O.A.; Tabor, J.J. Independent control of mean and noise by convolution of gene expression distributions. Nat. Commun. 2021, 12, 6957. [Google Scholar] [CrossRef] [PubMed]
- Tsimring, L.S. Noise in biology. Rep. Prog. Phys. 2014, 77, 026601. [Google Scholar] [CrossRef]
- Mailfert, S.; Touvier, J.; Benyoussef, L.; Fabre, R.; Rabaoui, A.; Blache, M.C.; Hamon, Y.; Brustlein, S.; Monneret, S.; Marguet, D.; et al. A Theoretical High-Density Nanoscopy Study Leads to the Design of UNLOC, a Parameter-free Algorithm. Biophys. J. 2018, 115, 565–576. [Google Scholar] [CrossRef]
- Laine, R.F.; Heil, H.S.; Coelho, S.; Nixon-Abell, J.; Jimenez, A.; Wiesner, T.; Martínez, D.; Galgani, T.; Régnier, L.; Stubb, A.; et al. High-fidelity 3D live-cell nanoscopy through data-driven enhanced super-resolution radial fluctuation. Nat. Methods 2023, 20, 1949–1956. [Google Scholar] [CrossRef]
- Liu, W.; Yao, Y.; Meng, J.; Qian, S.; Han, Y.; Zhou, L.; Wang, T.; Chen, Y.; Chen, L.; Ye, Z.; et al. Architecture-driven quantitative nanoscopy maps cytoskeleton remodeling. Proc. Natl. Acad. Sci. USA 2024, 121, e2410688121. [Google Scholar] [CrossRef]
- Ilan, Y. Overcoming Compensatory Mechanisms toward Chronic Drug Administration to Ensure Long-Term, Sustainable Beneficial Effects. Mol. Ther. Methods Clin. Dev. 2020, 18, 335–344. [Google Scholar] [CrossRef]
- Bayatra, A.; Nasserat, R.; Ilan, Y. Overcoming Low Adherence to Chronic Medications by Improving their Effectiveness Using a Personalized Second-generation Digital System. Curr. Pharm. Biotechnol. 2024, 25, 2078–2088. [Google Scholar] [CrossRef]
- Hurvitz, N.; Ilan, Y. The Constrained-Disorder Principle Assists in Overcoming Significant Challenges in Digital Health: Moving from “Nice to Have” to Mandatory Systems. Clin. Pract. 2023, 13, 994–1014. [Google Scholar] [CrossRef]
- Sigawi, T.; Lehmann, H.; Hurvitz, N.; Ilan, Y. Constrained Disorder Principle-Based Second-Generation Algorithms Implement Quantified Variability Signatures to Improve the Function of Complex Systems. J. Bioinform. Syst. Biol. 2023, 6, 82–89. [Google Scholar] [CrossRef]
- Hurvitz, N.; Dinur, T.; Revel-Vilk, S.; Agus, S.; Berg, M.; Zimran, A.; Ilan, Y. A Feasibility Open-Labeled Clinical Trial Using a Second-Generation Artificial-Intelligence-Based Therapeutic Regimen in Patients with Gaucher Disease Treated with Enzyme Replacement Therapy. J. Clin. Med. 2024, 13, 3325. [Google Scholar] [CrossRef]
- Hurvitz, N.; Lehman, H.; Hershkovitz, Y.; Kolben, Y.; Jamil, K.; Agus, S.; Berg, M.; Aamar, S.; Ilan, Y. A constrained disorder principle-based second-generation artificial intelligence digital medical cannabis system: A real-world data analysis. J. Public Health Res. 2025, 14, 22799036251337640. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, R.; Weihs, T.; Wurm, C.A.; Jansen, I.; Rehman, J.; Sahl, S.J.; Hell, S.W. MINFLUX nanometer-scale 3D imaging and microsecond-range tracking on a common fluorescence microscope. Nat. Commun. 2021, 12, 1478. [Google Scholar] [CrossRef] [PubMed]
- Alfonso, C.; Clarke, D.C.; Capdevila, L. Individual training prescribed by heart rate variability, heart rate and well-being scores in experienced cyclists. Sci. Rep. 2025, 15, 34023. [Google Scholar] [CrossRef]
- Sharma, S.; Kour, K.; Bakshi, P.; Bajaj, K.; Bhushan, B.; Singh, B.; Mir, M.; Shah, A.; Sharma, N. Exploring variability parameters, character association studies, and genetic divergence in pomegranate (Punica granatum L.) from temperate regions of North-Western Himalaya. Plant Physiol. Biochem. 2025, 229, 110605. [Google Scholar] [CrossRef]
- Carlson, M.O.; Andrews, B.L.; Simons, Y.B. Distinguishing direct interactions from global epistasis using rank statistics. Proc. Natl. Acad. Sci. USA 2025, 122, e2509444122. [Google Scholar] [CrossRef]
- Dai, G.; Zhang, R.; Wuwu, Q.; Tseng, C.-C.; Zhou, Y.; Wang, S.; Qian, S.; Lu, M.; Tuz, A.A.; Gunzer, M.; et al. Implicit neural image field for biological microscopy image compression. Nat. Comput. Sci. 2025, 5, 1041–1050. [Google Scholar] [CrossRef] [PubMed]
- Heffernan, K.S.; Barnes, J.N.; London, A.S.; Monroe, D.C.; Stern, Y.; Sloan, R.P.; Schaefer, S.M. Association between low-frequency oscillations in blood pressure variability and brain age derived from neuroimaging. Alzheimers Dement. 2025, 21, e70833. [Google Scholar] [CrossRef] [PubMed]
- Combs, H.L.; Kurth, R.; Nair, A.; York, M.K.; Weintraub, D.; Lafontant, D.-E.; Caspell-Garcia, C.; Parkinson’s Progression Markers Initiative. Utilizing Intraindividual Cognitive Variability to Predict Early Neuronal Synuclein Disease Progression. medRxiv 2025, 2025-10. [Google Scholar] [CrossRef]
- Park, S.; Park, J.; Kim, Y.; Moon, I.; Javidi, B. Simple and practical single-shot digital holography based on unsupervised diffusion model. Eng. Appl. Artif. Intell. 2026, 163, 112970. [Google Scholar] [CrossRef]
- Buron, J.; Linossier, A.; Gestreau, C.; Schaller, F.; Tyzio, R.; Felix, M.-S.; Matarazzo, V.; Thoby-Brisson, M.; Muscatelli, F.; Menuet, C. Oxytocin modulates respiratory heart rate variability through a hypothalamus–brainstem–heart neuronal pathway. Nat. Neurosci. 2025, 28, 2247–2261. [Google Scholar] [CrossRef]
- Lombardi, V.; Di Rocco, L.; Meo, E.; Venafra, V.; Di Nisio, E.; Perticaroli, V.; Nicolaeasa, M.L.; Cencioni, C.; Spallotta, F.; Negri, R.; et al. PatientProfiler: Building patient-specific signaling models from proteogenomic data. Mol. Syst. Biol. 2025, 21, 1845–1865. [Google Scholar] [CrossRef]
- Arif, W.; Datar, G.; Kalsotra, A. Intersections of post-transcriptional gene regulatory mechanisms with intermediary metabolism. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 349–362. [Google Scholar] [CrossRef]
- Álvez, M.B.; Bergström, S.; Kenrick, J.; Johansson, E.; Åberg, M.; Akyildiz, M.; Altay, O.; Sköld, H.; Antonopoulos, K.; Apostolakis, E.; et al. A human pan-disease blood atlas of the circulating proteome. Science 2025, 390, eadx2678. [Google Scholar] [CrossRef]
- Behrens, M. Single-Cell Multiomic Approaches for Understanding Human Brain Variability in Health and Disease. Eur. Neuropsychopharmacol. 2025, 99, 26. [Google Scholar] [CrossRef]
- Fang, Z.; He, J.; Qin, P.; Chen, H.; Zhang, C.; Sun, S. Enhancing autonomous driving safety in real lane-changing scenarios under friction variability: A friction-adaptive shield reinforcement learning framework. Accid. Anal. Prev. 2025, 223, 108265. [Google Scholar] [CrossRef]


| Aspect | Constrained Disorder Principle (CDP) | Nanoscopy Method |
|---|---|---|
| Fundamental Philosophy | Noise and disorder are essential for proper system function and should be preserved/optimized. | Noise represents fundamental limits that can be overcome through advanced techniques. |
| Basic Attitude to Noise | Noise/variability = necessary functional property regulated within dynamic bounds. | Noise = measurement limitation to be minimized, bypassed, or engineered around via optical/detection strategies |
| Primary Goal | Understand how systems use disorder constructively for optimal function; explain/adapt biological function for therapeutics and diagnostics. | Achieve unprecedented measurement precision despite physical limitations; resolve spatial detail and dynamics at molecular scales for structural/biophysical insights. |
| Domain | Systems biology, physiology, complex systems | Optical physics, instrumentation, super-resolution microscopy |
| Theoretical Foundation | Intrinsic variability is mandatory for proper function and dynamically changes in response to pressure. | By illuminating with a diffraction minimum, point scatterers can be resolved at small fractions of the wavelength. |
| Scale of Application | System-level, cellular, organismal, population dynamics | Molecular-level, single-molecule interactions, nanometer precision |
| Mathematical Framework | Dynamic boundary theory, allostatic load models, variability quantification, distributions, dynamic boundaries, high-level statistical descriptors (variance, entropy, autocorrelation) | Cramer–Rao bounds, Fisher information theory, Poisson statistics; point-spread function engineering, Poisson/Gaussian noise models, estimator theory |
| Methodological Approach | Variability quantification, dynamic boundary analysis, system-level understanding | Minimum-based scanning, statistical optimization, and photon budget management |
| Measurement Strategy | Characterize and understand patterns within apparent randomness | Achieve 8 nm resolution (1/80 of wavelength) through optimized signal processing |
| Treatment of Stochasticity | Embrace, quantify, regulate—variability is adaptive | Model and minimize impact; convert into estimable effects or bypass via deterministic system design |
| Time Dependence | Emphasizes dynamic adaptation over time, temporal variability as a feature | Enables real-time tracking with temporal resolution alongside spatial precision |
| Signal Processing | Looks for functional patterns within noise, treats variability as data | Uses Fisher information to maximize information extraction from minimal photon counts |
| Precision vs. Adaptability | Prioritizes adaptability and robustness over precision | Prioritizes precision and resolution over natural system variability |
| System Interaction | Work with the system’s natural tendencies to maintain optimal levels of disorder. | Override physical limitations through technological innovation. |
| Operational Recommendation | Intervene to restore or sculpt variability (therapies ranging from stochastic stimulation to boundary modulation) | Engineer illumination/detection modalities (structured beams, diffraction minima) and signal processing to achieve nanometer resolution despite stochastic emission |
| Biological Relevance | Helps understand how biological systems manage noise to function well | Enables observation of individual protein conformational changes and molecular mechanics |
| Error Handling | Errors/variations may contain functional information | Constant relative error σ/d for minimum-based measurements |
| Optimization Target | Optimize the degree of disorder for functional requirements | Optimize signal-to-noise ratio for maximum resolution |
| Technology Application | Improving the performance of digital twins and bioengineering applications | Super-resolution microscopy, molecular tracking, protein dynamics |
| Control Philosophy | Constrained control-maintain disorder within dynamic boundaries | Precise control-eliminate unwanted variations to achieve target measurements |
| Information Content | High information content in variability patterns | High information content in precise spatial and temporal coordinates |
| Robustness Strategy | Robustness through adaptive variability and flexible responses | Robustness through redundant measurements and statistical averaging |
| Future Applications | White noise applications for overcoming malfunctions | Extension to Raman scattering, X-rays, and other wave-based imaging modalities |
| Success Metrics | Functional maintenance under stress, adaptive capacity, and system resilience | Spatial resolution (8 nm achieved), temporal resolution, and measurement precision |
| Complementary Potential | Provides a theoretical framework for when precision should be sacrificed for adaptability | Provides measurement tools to validate CDP predictions and quantify disorder patterns |
| System | Measurement Method | Observed Variability | Functional Significance | CDP Alignment |
|---|---|---|---|---|
| Mitochondrial cristae | STED | Curvature σ ≈ 50 nm | ATP efficiency optimization | High |
| Synaptic vesicles | MINFLUX | Position σ ≈ 25 nm | Transmission reliability | High |
| Membrane receptors | Deterministic minima | Cluster spacing CV = 0.3 | Signaling sensitivity | High |
| Nuclear pores | STED | Position σ ≈ 12 nm | Selective transport | Medium |
| DNA repair foci | STED | Dynamic size CV = 0.2–0.5 | Repair efficiency | High |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Ilan, Y. Integrating the Contrasting Perspectives Between the Constrained Disorder Principle and Deterministic Optical Nanoscopy: Enhancing Information Extraction from Imaging of Complex Systems. Bioengineering 2026, 13, 103. https://doi.org/10.3390/bioengineering13010103
Ilan Y. Integrating the Contrasting Perspectives Between the Constrained Disorder Principle and Deterministic Optical Nanoscopy: Enhancing Information Extraction from Imaging of Complex Systems. Bioengineering. 2026; 13(1):103. https://doi.org/10.3390/bioengineering13010103
Chicago/Turabian StyleIlan, Yaron. 2026. "Integrating the Contrasting Perspectives Between the Constrained Disorder Principle and Deterministic Optical Nanoscopy: Enhancing Information Extraction from Imaging of Complex Systems" Bioengineering 13, no. 1: 103. https://doi.org/10.3390/bioengineering13010103
APA StyleIlan, Y. (2026). Integrating the Contrasting Perspectives Between the Constrained Disorder Principle and Deterministic Optical Nanoscopy: Enhancing Information Extraction from Imaging of Complex Systems. Bioengineering, 13(1), 103. https://doi.org/10.3390/bioengineering13010103







