MedSegNet10: A Publicly Accessible Network Repository for Split Federated Medical Image Segmentation
Abstract
1. Introduction
2. Related Works
2.1. Split Federated Learning (SplitFed)
2.2. Publicly Available Federated Repositories
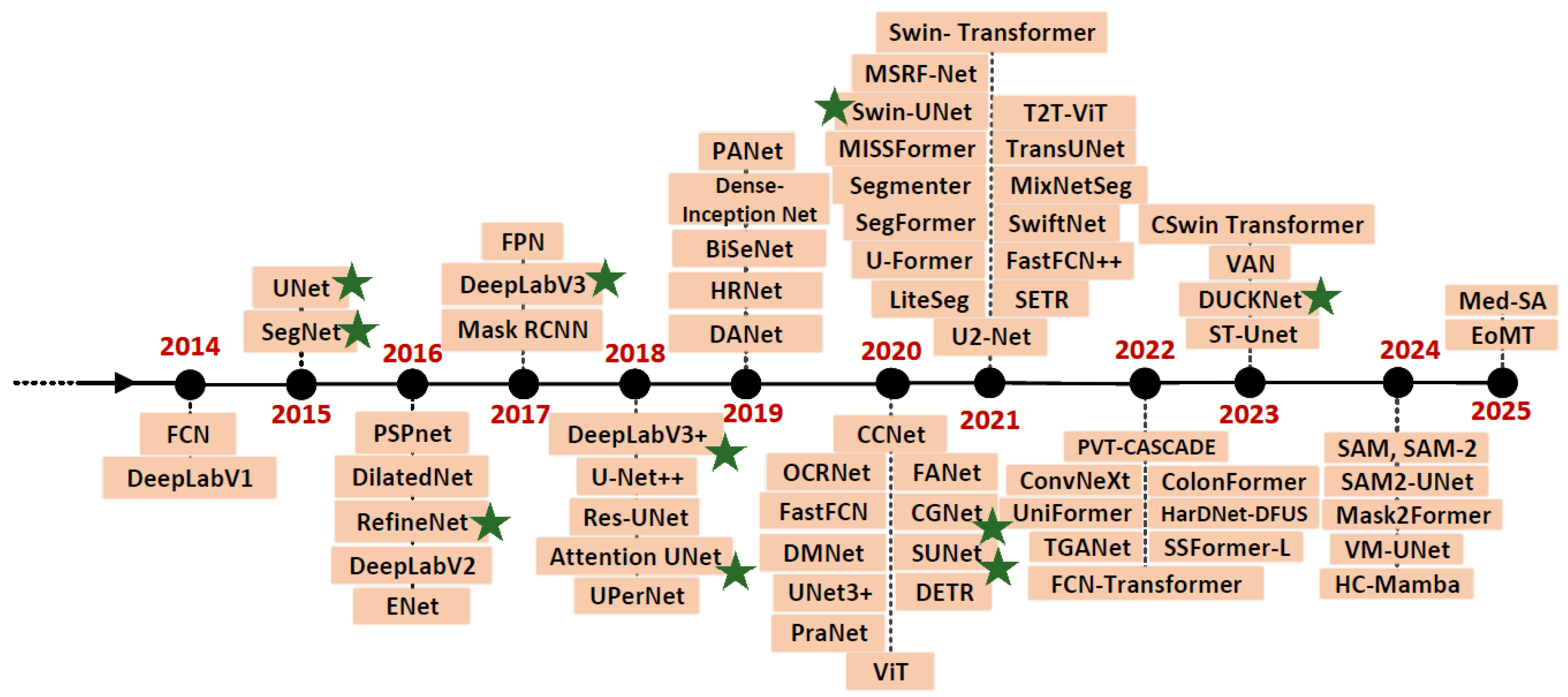
2.3. Image Segmentation Models
- FCN [51]: Fully Connected Network.
- UNet [10]: UNetwork.
- SegNet [11]: Segmentation Network.
- PSPnet [52]: Pyramid Scene Parsing Network.
- ENet [53]: Efficient Neural Network.
- RefineNet: Refining Segmentation-based Network.
- Attention UNet: Attention-based UNet.
- FPN [56]: Feature Pyramid Network.
- Mask RCNN [57]: Mask Region-based Convolutional Neural Network.
- PANet [58]: Path Aggregation Network.
- BiseNet [59]: Bilateral Segmentation Network.
- HRNet [60]: High-Resolution Network.
- OCRNet [61]: Object-Contextual Representations for Semantic Segmentation.
- DANet [62]: Dual Attention Network.
- CCNet [63]: Criss-Cross Attention Network.
- SETR [64]: Spatially Enhanced Transformer.
- UPerNet [65]: Unified Perceptual Parsing Network.
- FastFCN [66]: Fast Fully Convolutional Network.
- SUNet [16]: Strong UNet.
- FANet [67]: Feature Aggregation Network.
- DMNet [68]: Dense Multi-scale Network.
- CGNet [15]: Context-Guided Network.
- DETR [69]: DEtection Transformer.
- PraNet [70]: Parallel Reverse Attention Network.
- ViT [71]: Vision Transformer.
- Swin-UNet: Swin Transformer-based UNet.
- MSRF-Net [72]: Multi-Scale Residual Fusion Network.
- T2T-ViT [73]: Token-to-Token Vision Transformer.
- VAN [74]: Visual Attention Network.
- CSwin Transformer [75]: Cross-Stage win transformer.
- DUCK-Net [17]: Deep Understanding Convolutional Kernel Network.
- ST-UNet [76]: Spatiotemporal UNet.
- SAM [77]: Segment Anything.
- VM-UNet [78]: Vision Mamba UNet.
- HC-Mamba [79]: Hybrid-convolution version of Vision Mamba.
- EoMT [80]: Encoder-only Mask Transformer.
- Med-SA [81]: Medical SAM Adapter.
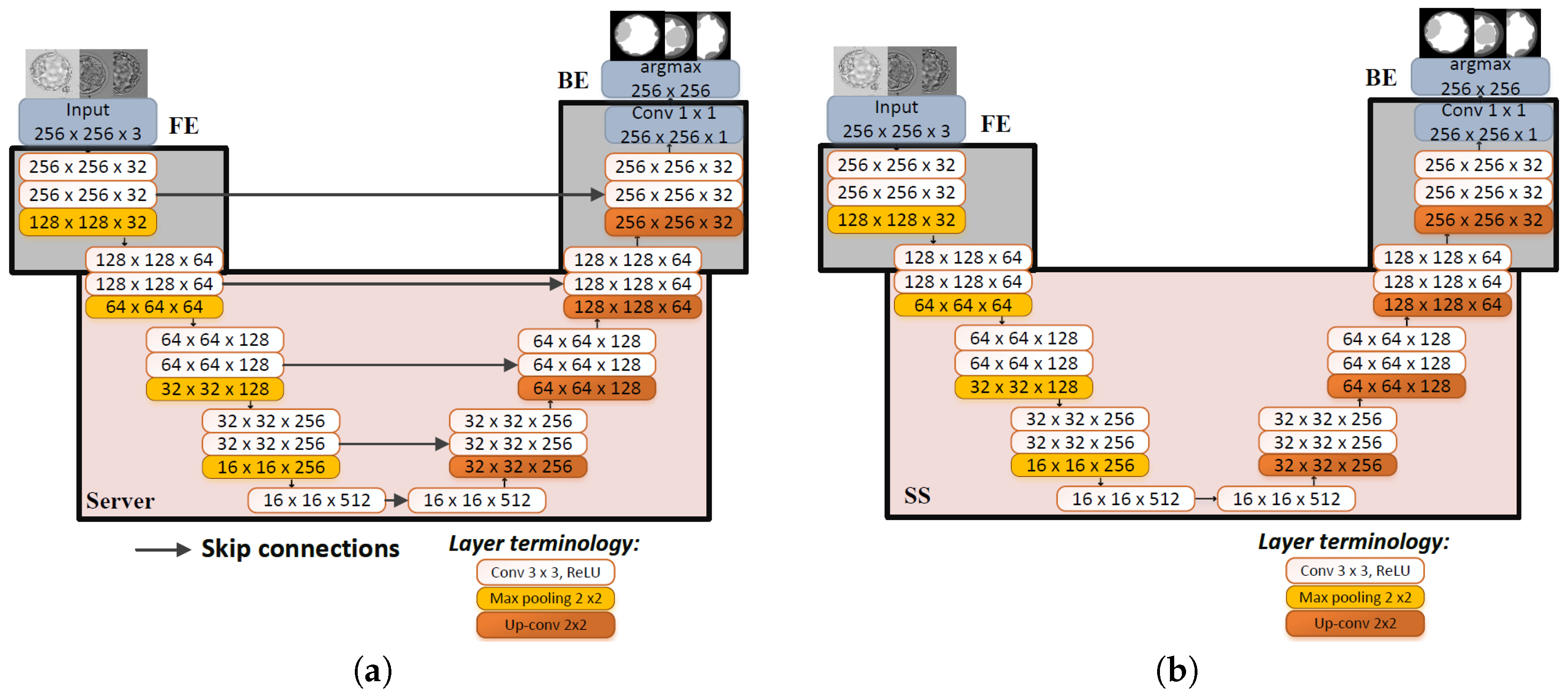
2.3.1. UNet
2.3.2. SegNet
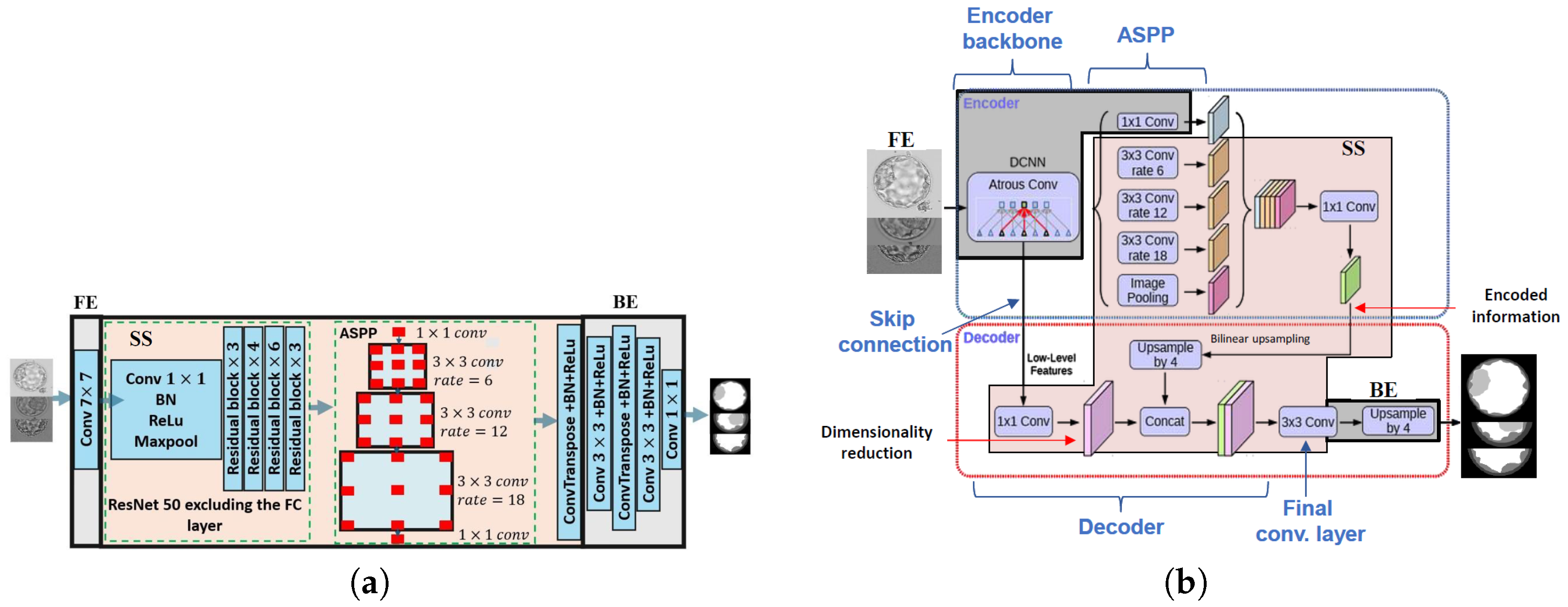
2.3.3. DeepLab
2.3.4. RefineNet
2.3.5. SUNet
2.3.6. CGNet
2.3.7. DUCK-Net
2.3.8. Attention UNet
2.3.9. Swin-UNet
2.4. Decision on Split Points Selection
- Task-specific concerns: The choice of split points is often guided by the nature of the machine learning task. For instance, in natural language processing, splits should occur at layers that capture semantic features, whereas in computer vision, splits should be at layers that capture high-level visual features.
- Communication constraints: Split points should be strategically selected to minimize the overall computational load and communication costs associated with information transfer. This involves choosing points where computations are most intensive or sensitive, thus reducing overall latency and communication overhead.
- Model architecture: Split points are selected at layers representing high-level features to enable clients to effectively learn task-specific details, ensuring that the model architecture supports the desired learning outcomes. Moreover, the edge blocks maintain the same dimensions, which is necessary for backpropagating gradients in the backward pass. Each sub-model generates its own gradients, making consistent dimensionality crucial.
- Privacy and security concerns: To maintain data privacy, splits must be designed so that sensitive data remains on the client side. This approach involves creating two distinct model parts on the client side, with the front end handling sensitive data and the back end managing sensitive GTs.
- Computational capabilities of clients: Split points are chosen to ensure that clients perform minimal computations, allowing those with limited resources to participate in the SplitFed training process without facing computational constraints.
3. Experiments
3.1. Experimental Setup
- -
- Blastocyst dataset [22]: includes 801 Blastocyst RGB images along with their GTs created for a multi-class embryo segmentation task. Each image is segmented into five classes: zona pellucida (ZP), trophectoderm (TE), blastocoel (BL), inner cell mass (ICM), and background (BG).
- -
- HAM10K dataset [20]: The Human Against Machine dataset contains 10,015 dermatoscopic RGB images along with the corresponding binary GT masks, representing seven different types of skin lesions, including melanoma and benign conditions. Each image is segmented into two classes: skin lesion and background.
- -
- KVASIR-SEG dataset [21]: contains 1000 annotated endoscopic RGB images of polyps from colonoscopy procedures, each paired with a binary GT segmentation mask. Each image is segmented into two classes: abnormal condition (such as a lesion, polyp, or ulcer) and background.
3.2. Experimental Results
3.2.1. Quantitative Results
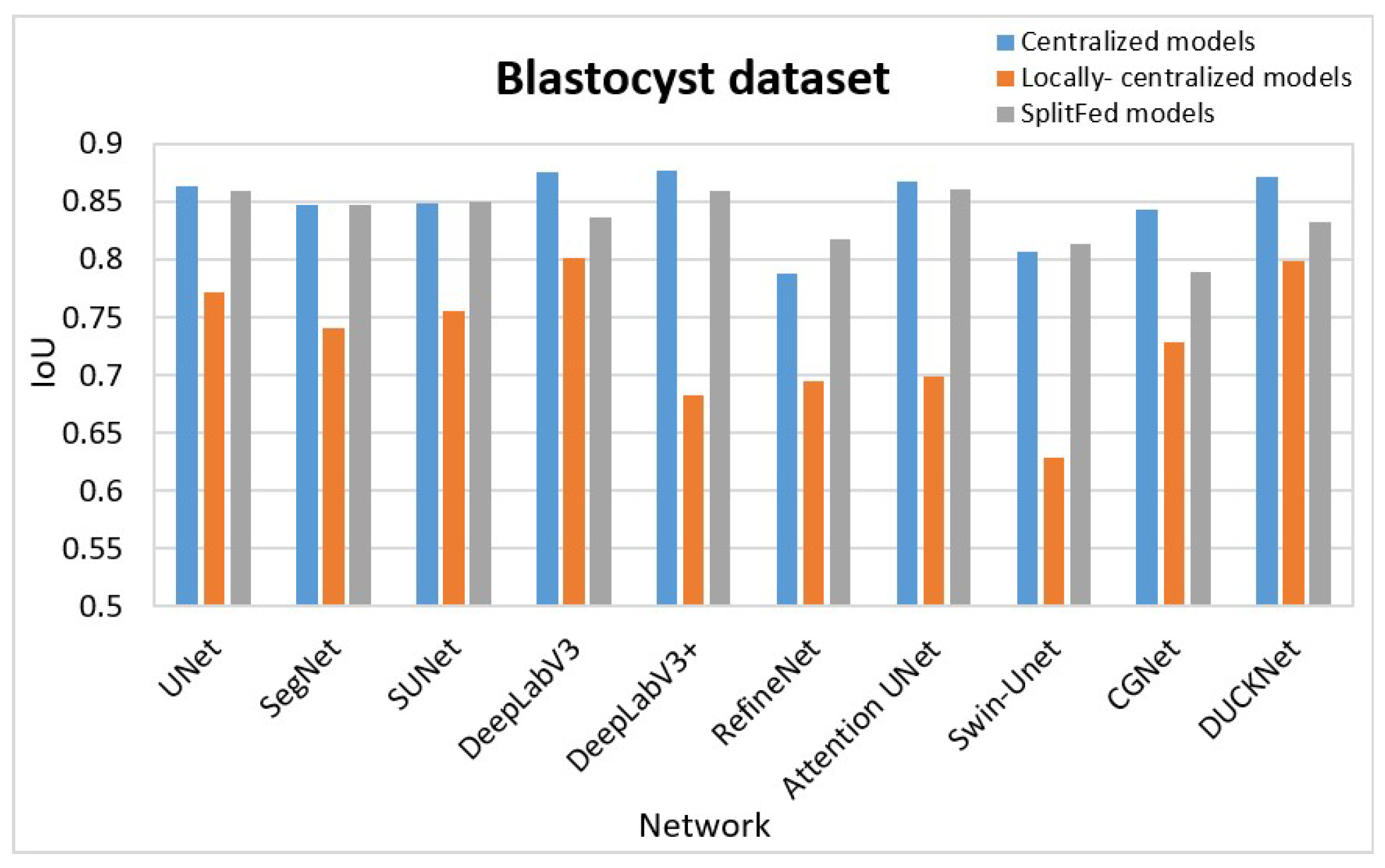
- Centralized learning on full data: We initially trained each network without splitting for image segmentation. We utilized the entire data from the three datasets separately. The average IoUs for all data samples in each set for the centralized models are displayed in the C column of Table 1.
- Centralized learning locally at each client: Secondly, we trained each client’s local data in a client-specific, centralized manner to ensure a fair comparison. In this step, each client trained the networks without data splitting. We recorded the IoUs for each client and computed the average, which is presented in the L column for each segmentation task in Table 1.
- SplitFed learning: Thirdly, we trained the SplitFed networks in collaboration with all clients. The IoUs of the SplitFed models are recorded in the S column for each segmentation task in Table 1.
3.2.2. Qualitative Results
3.3. Evaluation
3.3.1. Testing Performance Comparison
3.3.2. Comparison of Computational Complexity
3.3.3. Performance Comparison with Other Existing Methods
4. Limitations & Future Works
- First, our evaluation is limited to three distinct and commonly studied publicly available image types. Although we considered both multi-class (Blastocyst) and binary (HAM10K and KVASIR-SEG) segmentation datasets with varying sample sizes and styles to broaden the scope of generalization, these datasets may still not fully reflect the diversity of imaging characteristics, modalities, and annotation practices encountered in large-scale clinical deployments. Consequently, the reported results should not be regarded as definitive evidence of cross-domain generalizability. Future work will involve evaluating MedSegNet10 across a wider range of imaging modalities (e.g., CT, MRI, and multimodal acquisitions), annotation practices, and datasets to more comprehensively assess its robustness.
- Second, the datasets used in this study are modest in size. This limitation reflects the broader reality of medical imaging research- large open-source datasets are rare, expert-annotated data are costly, many image types are difficult to obtain, and privacy regulations restrict access. Consequently, it is naturally infeasible to evaluate decentralized frameworks on large-scale public datasets simply because such resources do not exist for many medical imaging tasks. MedSegNet10 should therefore be viewed as a foundational resource rather than a demonstration of large-scale scalability. Future work will involve expanding MedSegNet10 using larger institutional datasets and multi-centre cohorts, enabling more rigorous evaluation under realistic data volumes, acquisition variability, and deployment conditions.
- Third, the current experiments intentionally adopt IID data partitions to establish a controlled baseline and support benchmark reproducibility. This design choice does not reflect real clinical environments, where hospitals often exhibit strongly non-IID data distributions due to differing demographics, imaging devices, and annotation protocols. Non-IID robustness is a central challenge in federated learning, and evaluating MedSegNet10 under a range of realistic non-IID scenarios is an important direction for future work.
- Finally, recent capability-oriented reviews in smart healthcare [105] highlight how AI contributes to integrated monitoring [105,106], remote diagnostics [107], decentralized decision support [108], and data-driven hospital systems [109]. SplitFed architectures align naturally with these developments because they enable collaboration across institutions while preserving data privacy. Incorporating SplitFed networks from MedSegNet10 into smart-healthcare frameworks could facilitate interoperable, privacy-preserving segmentation tools that operate across hospitals or global imaging networks. This integration represents a promising avenue for extending MedSegNet10 beyond standalone model training towards deployment in real clinical infrastructures.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Alzahrani, Y.; Boufama, B. Biomedical image segmentation: A survey. SN Comput. Sci. 2021, 2, 310. [Google Scholar] [CrossRef]
- Narayan, V.; Faiz, M.; Mall, P.K.; Srivastava, S. A Comprehensive Review of Various Approach for Medical Image Segmentation and Disease Prediction. Wirel. Pers. Commun. 2023, 132, 1819–1848. [Google Scholar] [CrossRef]
- McMahan, B.; Moore, E.; Ramage, D.; Hampson, S.; y Arcas, B.A. Communication-Efficient Learning of Deep Networks from Decentralized Data. In Proceedings of the AISTATS, Fort Lauderdale, FL, USA, 20–22 April 2017; pp. 1273–1282. [Google Scholar]
- Gupta, O.; Raskar, R. Distributed learning of deep neural network over multiple agents. J. Netw. Comput. 2018, 116, 1–8. [Google Scholar] [CrossRef]
- Thapa, C.; Arachchige, P.C.M.; Camtepe, S.; Sun, L. SplitFed: When Federated Learning Meets Split Learning. In Proceedings of the AAAI, Virtual, 22 February–1 March 2022; Volume 36, pp. 8485–8493. [Google Scholar]
- Abbas, Q.; Jeong, W.; Lee, S.W. Explainable AI in clinical decision support systems: A meta-analysis of methods, applications, and usability challenges. Healthcare 2025, 13, 2154. [Google Scholar] [CrossRef] [PubMed]
- Barragan-Montero, A.; Bibal, A.; Dastarac, M.H.; Draguet, C.; Valdes, G.; Nguyen, D.; Willems, S.; Vandewinckele, L.; Holmström, M.; Löfman, F.; et al. Towards a safe and efficient clinical implementation of machine learning in radiation oncology by exploring model interpretability, explainability and data-model dependency. Phys. Med. Biol. 2022, 67, 11TR01. [Google Scholar] [CrossRef]
- Guo, Y.; Liu, Y.; Georgiou, T.; Lew, M.S. A review of semantic segmentation using deep neural networks. Int. J. Multimed. Inf. Retr. 2018, 7, 87–93. [Google Scholar] [CrossRef]
- Strudel, R.; Garcia, R.; Laptev, I.; Schmid, C. Segmenter: Transformer for semantic segmentation. In Proceedings of the CVPR, Virtual, 19–25 June 2021; pp. 7262–7272. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional networks for biomedical image segmentation. In Proceedings of the MICCAI, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar]
- Badrinarayanan, V.; Kendall, A.; Cipolla, R. Segnet: A deep convolutional encoder–decoder architecture for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 2481–2495. [Google Scholar] [CrossRef]
- Chen, L.C.; Papandreou, G.; Schroff, F.; Adam, H. Rethinking atrous convolution for semantic image segmentation. arXiv 2017, arXiv:1706.05587. [Google Scholar] [CrossRef]
- Chen, L.C.; Zhu, Y.; Papandreou, G.; Schroff, F.; Adam, H. Encoder-decoder with atrous separable convolution for semantic image segmentation. In Proceedings of the ECCV, Munich, Germany, 8–14 September 2018; pp. 801–818. [Google Scholar]
- Lin, G.; Milan, A.; Shen, C.; Reid, I. Refinenet: Multi-path refinement networks for high-resolution semantic segmentation. In Proceedings of the CVPR, Honolulu, HI, USA, 21–26 July 2017; pp. 1925–1934. [Google Scholar]
- Wu, T.; Tang, S.; Zhang, R.; Cao, J.; Zhang, Y. CGNet: A light-weight context guided network for semantic segmentation. IEEE Trans. Image Process. 2020, 30, 1169–1179. [Google Scholar] [CrossRef]
- Yi, L.; Zhang, J.; Zhang, R.; Shi, J.; Wang, G.; Liu, X. SU-Net: An efficient encoder–decoder model of federated learning for brain tumor segmentation. In Proceedings of the ICANN, Bratislava, Slovakia, 15–18 September 2020; pp. 761–773. [Google Scholar]
- Dumitru, R.G.; Peteleaza, D.; Craciun, C. Using DUCK-Net for polyp image segmentation. Sci. Rep. 2023, 13, 9803. [Google Scholar] [CrossRef]
- Oktay, O.; Schlemper, J.; Folgoc, L.L.; Lee, M.; Heinrich, M.; Misawa, K.; Mori, K.; McDonagh, S.; Hammerla, N.Y.; Kainz, B.; et al. Attention u-net: Learning where to look for the pancreas. arXiv 2018, arXiv:1804.03999. [Google Scholar] [CrossRef]
- Cao, H.; Wang, Y.; Chen, J.; Jiang, D.; Zhang, X.; Tian, Q.; Wang, M. Swin-unet: Unet-like pure transformer for medical image segmentation. In Proceedings of the ECCV, Tel Aviv, Israel, 23–27 October 2022; pp. 205–218. [Google Scholar]
- Tschandl, P.; Rosendahl, C.; Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 2018, 5, 1–9. [Google Scholar] [CrossRef]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Halvorsen, P.; de Lange, T.; Johansen, D.; Johansen, H.D. Kvasir-seg: A segmented polyp dataset. In Proceedings of the MMM, Daejeon, South Korea, 5–8 January 2020; pp. 451–462. [Google Scholar]
- Lockhart, L.; Saeedi, P.; Au, J.; Havelock, J. Multi-Label Classification for Automatic Human Blastocyst Grading with Severely Imbalanced Data. In Proceedings of the MMSP, Kuala Lumpur, Malaysia, 27–29 September 2019; pp. 1–6. [Google Scholar]
- Zhang, M.; Qu, L.; Singh, P.; Kalpathy-Cramer, J.; Rubin, D.L. Splitavg: A heterogeneity-aware federated deep learning method for medical imaging. IEEE J. Biomed. Health Inform. 2022, 26, 4635–4644. [Google Scholar] [CrossRef]
- Yang, Z.; Chen, Y.; Huangfu, H.; Ran, M.; Wang, H.; Li, X.; Zhang, Y. Dynamic corrected split federated learning with homomorphic encryption for u-shaped medical image networks. IEEE J. Biomed. Health Inform. 2023, 27, 5946–5957. [Google Scholar] [CrossRef]
- Kafshgari, Z.H.; Shiranthika, C.; Saeedi, P.; Bajić, I.V. Quality-adaptive split-federated learning for segmenting medical images with inaccurate annotations. In Proceedings of the IEEE ISBI, Cartagena de Indias, Colombia, 18–21 April 2023; pp. 1–5. [Google Scholar]
- Kafshgari, Z.H.; Bajić, I.V.; Saeedi, P. Smart split-federated learning over noisy channels for embryo image segmentation. In Proceedings of the IEEE ICASSP, Rhodes Island, Greece, 4–10 June 2023; pp. 1–5. [Google Scholar]
- Baek, S.; Park, J.; Vepakomma, P.; Raskar, R.; Bennis, M.; Kim, S.L. Visual transformer meets cutmix for improved accuracy, communication efficiency, and data privacy in split learning. arXiv 2022, arXiv:2207.00234. [Google Scholar] [CrossRef]
- Qu, L.; Zhou, Y.; Liang, P.P.; Xia, Y.; Wang, F.; Adeli, E.; Fei-Fei, L.; Rubin, D. Rethinking architecture design for tackling data heterogeneity in federated learning. In Proceedings of the CVPR, New Orleans, LA, USA, 19–24 June 2022; pp. 10061–10071. [Google Scholar]
- Oh, S.; Park, J.; Baek, S.; Nam, H.; Vepakomma, P.; Raskar, R.; Bennis, M.; Kim, S.L. Differentially private cutmix for split learning with vision transformer. arXiv 2022, arXiv:2210.15986. [Google Scholar] [CrossRef]
- Park, S.; Kim, G.; Kim, J.; Kim, B.; Ye, J.C. Federated split vision transformer for COVID-19 CXR diagnosis using task-agnostic training. arXiv 2021, arXiv:2111.01338. [Google Scholar] [CrossRef]
- Park, S.; Ye, J.C. Multi-Task Distributed Learning Using Vision Transformer With Random Patch Permutation. IEEE Trans. Med. Imaging 2023, 42, 2091–2105. [Google Scholar] [CrossRef] [PubMed]
- Almalik, F.; Alkhunaizi, N.; Almakky, I.; Nandakumar, K. FeSViBS: Federated Split Learning of Vision Transformer with Block Sampling. arXiv 2023, arXiv:2306.14638. [Google Scholar]
- Shiranthika, C.; Saeedi, P.; Bajić, I.V. Decentralized Learning in Healthcare: A Review of Emerging Techniques. IEEE Access 2023, 11, 54188–54209. [Google Scholar] [CrossRef]
- Carter, K.W.; Francis, R.W.; Carter, K.; Francis, R.; Bresnahan, M.; Gissler, M.; Grønborg, T.; Gross, R.; Gunnes, N.; Hammond, G.; et al. ViPAR: A software platform for the Virtual Pooling and Analysis of Research Data. Int. J. Epidemiol. 2016, 45, 408–416. [Google Scholar] [CrossRef] [PubMed]
- Gazula, H.; Kelly, R.; Romero, J.; Verner, E.; Baker, B.T.; Silva, R.F.; Imtiaz, H.; Saha, D.K.; Raja, R.; Turner, J.A.; et al. COINSTAC: Collaborative informatics and neuroimaging suite toolkit for anonymous computation. JOSS 2020, 5, 2166. [Google Scholar] [CrossRef]
- Marcus, D.S.; Olsen, T.R.; Ramaratnam, M.; Buckner, R.L. The Extensible Neuroimaging Archive Toolkit: An informatics platform for managing, exploring, and sharing neuroimaging data. Neuroinformatics 2007, 5, 11–33. [Google Scholar] [CrossRef]
- Tensorflow Federated. Available online: https://github.com/tensorflow/federated/ (accessed on 8 May 2025).
- Google Federated Research. Available online: https://github.com/google-research/federated/ (accessed on 8 May 2025).
- LEAF. Available online: https://github.com/TalwalkarLab/leaf/ (accessed on 8 May 2025).
- Fedmint. Available online: https://github.com/fedimint/fedimint/ (accessed on 8 May 2025).
- Xie, Y.; Wang, Z.; Gao, D.; Chen, D.; Yao, L.; Kuang, W.; Li, Y.; Ding, B.; Zhou, J. FederatedScope: A Flexible Federated Learning Platform for Heterogeneity. Proc. VLDB Endow. 2023, 16, 1059–1072. [Google Scholar] [CrossRef]
- Zeng, D.; Liang, S.; Hu, X.; Wang, H.; Xu, Z. FedLab: A flexible federated learning framework. arXiv 2021, arXiv:2107.11621. [Google Scholar]
- FATE. Available online: https://github.com/FederatedAI/FATE/ (accessed on 8 May 2025).
- Wu, N.; Yu, L.; Yang, X.; Cheng, K.T.; Yan, Z. FedIIC: Towards Robust Federated Learning for Class-Imbalanced Medical Image Classification. In Proceedings of the MICCAI, Vancouver, Canada, 8–12, October, 2023. [Google Scholar]
- Quicksql. Available online: https://github.com/Qihoo360/Quicksql/ (accessed on 8 May 2025).
- Graphql-mesh. Available online: https://github.com/ardatan/graphql-mesh/ (accessed on 8 May 2025).
- MedAugment. Available online: https://github.com/NUS-Tim/MedAugment/ (accessed on 8 May 2025).
- OpenMined’s PyGrid. Available online: https://github.com/OpenMined/PyGrid-deprecated---see-PySyft-/ (accessed on 8 May 2025).
- FedCT. Available online: https://github.com/evison/FedCT/ (accessed on 8 May 2025).
- Zhuang, W.; Xu, J.; Chen, C.; Li, J.; Lyu, L. COALA: A Practical and Vision-Centric Federated Learning Platform. In Proceedings of the ICML, Vienna, Austria, 21–27 July 2024. [Google Scholar]
- Long, J.; Shelhamer, E.; Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the CVPR, Boston, MA, USA, 7–12 June 2015; pp. 3431–3440. [Google Scholar]
- Zhao, H.; Shi, J.; Qi, X.; Wang, X.; Jia, J. Pyramid scene parsing network. In Proceedings of the CVPR, Honolulu, HI, USA, 21–26 July 2017; pp. 2881–2890. [Google Scholar]
- Paszke, A.; Chaurasia, A.; Kim, S.; Culurciello, E. Enet: A deep neural network architecture for real-time semantic segmentation. arXiv 2016, arXiv:1606.02147. [Google Scholar] [CrossRef]
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. Semantic image segmentation with deep convolutional nets and fully connected crfs. arXiv 2014, arXiv:1412.7062. [Google Scholar]
- Chen, L.C.; Papandreou, G.; Kokkinos, I.; Murphy, K.; Yuille, A.L. Deeplab: Semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected crfs. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 40, 834–848. [Google Scholar] [CrossRef]
- Lin, T.Y.; Dollár, P.; Girshick, R.; He, K.; Hariharan, B.; Belongie, S. Feature pyramid networks for object detection. In Proceedings of the CVPR, Honolulu, HI, USA, 21–26 July 2017; pp. 2117–2125. [Google Scholar]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R. Mask r-cnn. In Proceedings of the CVPR, Honolulu, HI, USA, 21–26 July 2017; pp. 2961–2969. [Google Scholar]
- Wang, K.; Liew, J.H.; Zou, Y.; Zhou, D.; Feng, J. PANet: Few-shot image semantic segmentation with prototype alignment. In Proceedings of the ICCV, Seoul, South Korea, 27 October–2 November 2019; pp. 9197–9206. [Google Scholar]
- Yu, C.; Wang, J.; Peng, C.; Gao, C.; Yu, G.; Sang, N. BiSeNet: Bilateral segmentation network for real-time semantic segmentation. In Proceedings of the ECCV, Munich, Germany, 8–14 September 2018; pp. 325–341. [Google Scholar]
- Sun, K.; Xiao, B.; Liu, D.; Wang, J. Deep High-Resolution Representation Learning for Human Pose Estimation. In Proceedings of the CVPR, Long Beach, CA, USA, 16–20 June 2019; pp. 5686–5696. [Google Scholar] [CrossRef]
- Yuan, Y.; Chen, X.; Wang, J. Object-contextual representations for semantic segmentation. In Proceedings of the ECCV, Glasgow, UK, 23–28 August 2020; pp. 173–190. [Google Scholar]
- Fu, J.; Liu, J.; Tian, H.; Li, Y.; Bao, Y.; Fang, Z.; Lu, H. Dual attention network for scene segmentation. In Proceedings of the CVPR, Long Beach, CA, 16–20 June 2019; pp. 3146–3154. [Google Scholar]
- Huang, Z.; Wang, X.; Huang, L.; Huang, C.; Wei, Y.; Liu, W. CCNet: Criss-cross attention for semantic segmentation. In Proceedings of the CVPR, Long Beach, CA, 16–20 June 2019; pp. 603–612. [Google Scholar]
- Heo, B.; Yun, S.; Han, D.; Chun, S.; Choe, J.; Oh, S.J. Rethinking spatial dimensions of vision transformers. In Proceedings of the ICCV, Virtual, 11–17 October 2021; pp. 11936–11945. [Google Scholar]
- Xiao, T.; Liu, Y.; Zhou, B.; Jiang, Y.; Sun, J. Unified perceptual parsing for scene understanding. In Proceedings of the ECCV, Munich, Germany, 8–14 September 2018; pp. 418–434. [Google Scholar]
- Wu, H.; Zhang, J.; Huang, K.; Liang, K.; Yu, Y. FastFCN: Rethinking dilated convolution in the backbone for semantic segmentation. arXiv 2019, arXiv:1903.11816. [Google Scholar] [CrossRef]
- Singha, T.; Pham, D.S.; Krishna, A. FANet: Feature aggregation network for semantic segmentation. In Proceedings of the DICTA, Melbourne, Australia, 29 November–2 December 2020; pp. 1–8. [Google Scholar]
- Fang, K.; Li, W.J. DMNet: Difference minimization network for semi-supervised segmentation in medical images. In Proceedings of the MICCAI, Lima, Peru, 4–8 October 2020; pp. 532–541. [Google Scholar]
- Carion, N.; Massa, F.; Synnaeve, G.; Usunier, N.; Kirillov, A.; Zagoruyko, S. End-to-end object detection with transformers. In Proceedings of the ECCV, Virtual, 23–28 August 2020; pp. 213–229. [Google Scholar]
- Fan, D.P.; Ji, G.P.; Zhou, T.; Chen, G.; Fu, H.; Shen, J.; Shao, L. PraNet: Parallel reverse attention network for polyp segmentation. In Proceedings of the MICCAI, Lima, Peru, 4–8 October 2020; pp. 263–273. [Google Scholar]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An image is worth 16 × 16 words: Transformers for image recognition at scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Srivastava, A.; Jha, D.; Chanda, S.; Pal, U.; Johansen, H.D.; Johansen, D.; Riegler, M.A.; Ali, S.; Halvorsen, P. MSRF-Net: A multi-scale residual fusion network for biomedical image segmentation. IEEE J. Biomed. Inform. 2021, 26, 2252–2263. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Chen, Y.; Wang, T.; Yu, W.; Shi, Y.; Jiang, Z.H.; Tay, F.E.; Feng, J.; Yan, S. Tokens-to-token ViT: Training vision transformers from scratch on imagenet. In Proceedings of the ICCV, Virtual, 11–17 October 2021; pp. 558–567. [Google Scholar]
- Guo, M.H.; Lu, C.Z.; Liu, Z.N.; Cheng, M.M.; Hu, S.M. Visual attention network. Comput. Vis. Media 2023, 9, 733–752. [Google Scholar] [CrossRef]
- Yang, H.; Yang, D. CSwin-PNet: A CNN-Swin Transformer combined pyramid network for breast lesion segmentation in ultrasound images. Expert Syst. Appl. 2023, 213, 119024. [Google Scholar] [CrossRef]
- Zhang, J.; Qin, Q.; Ye, Q.; Ruan, T. ST-UNet: Swin transformer boosted U-net with cross-layer feature enhancement for medical image segmentation. Comput. Biol. Med. 2023, 153, 106516. [Google Scholar] [CrossRef]
- Kirillov, A.; Mintun, E.; Ravi, N.; Mao, H.; Rolland, C.; Gustafson, L.; Xiao, T.; Whitehead, S.; Berg, A.C.; Lo, W.Y.; et al. Segment anything. In Proceedings of the CVPR, Vancouver, British Columbia, Canada, 18–22 June 2023; pp. 4015–4026. [Google Scholar]
- Ruan, J.; Li, J.; Xiang, S. VM-Unet: Vision mamba unet for medical image segmentation. arXiv 2024, arXiv:2402.02491. [Google Scholar] [CrossRef]
- Xu, J. HC-Mamba: Vision mamba with hybrid convolutional techniques for medical image segmentation. arXiv 2024, arXiv:2405.05007. [Google Scholar] [CrossRef]
- Kerssies, T.; Cavagnero, N.; Hermans, A.; Norouzi, N.; Averta, G.; Leibe, B.; Dubbelman, G.; de Geus, D. Your VIT is secretly an image segmentation model. In Proceedings of the CVPR, Nashville, TN, USA, 11–15 June 2025; pp. 25303–25313. [Google Scholar]
- Wu, J.; Wang, Z.; Hong, M.; Ji, W.; Fu, H.; Xu, Y.; Xu, M.; Jin, Y. Medical sam adapter: Adapting segment anything model for medical image segmentation. Med. Image Anal. 2025, 102, 103547. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the CVPR, Las Vegas, NV, 26 June–1 July 2016; pp. 770–778. [Google Scholar] [CrossRef]
- Chollet, F. Xception: Deep learning with depthwise separable convolutions. In Proceedings of the CVPR, Honolulu, HI, USA, 21–26 July 2017; pp. 1251–1258. [Google Scholar]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the inception architecture for computer vision. In Proceedings of the CVPR, Las Vegas, NV, 26 June–1 July 2016; pp. 2818–2826. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the CVPR, Honolulu, HI, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, Ł.; Polosukhin, I. Attention is all you need. In Proceedings of the 31st Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017; Volume 30. [Google Scholar]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin transformer: Hierarchical vision transformer using shifted windows. In Proceedings of the ICCV, Virtual, 11–17 October 2021; pp. 10012–10022. [Google Scholar]
- Shiranthika, C.; Kafshgari, Z.H.; Saeedi, P.; Bajić, I.V. SplitFed resilience to packet loss: Where to split, that is the question. In Proceedings of the MICCAI DeCaF, Vancouver, BC, Canada, 8–12 October 2023. [Google Scholar]
- Sudre, C.H.; Li, W.; Vercauteren, T.; Ourselin, S.; Jorge Cardoso, M. Generalised dice overlap as a deep learning loss function for highly unbalanced segmentations. In Proceedings of the MICCAI DLMIA, Quebec City, QC, Canada, 10–14 September 2017; pp. 240–248. [Google Scholar]
- Cox, M.A.A.; Cox, T.F. Multidimensional Scaling. In Handbook of Data Visualization; Springer: Berlin/Heidelberg, Germany, 2008; pp. 315–347. [Google Scholar]
- FLOPs Counting Tool for Neural Networks in Pytorch Framework. Available online: https://pypi.org/project/ptflops/ (accessed on 19 April 2025).
- Rad, R.M.; Saeedi, P.; Au, J.; Havelock, J. BLAST-NET: Semantic segmentation of human blastocyst components via cascaded atrous pyramid and dense progressive upsampling. In Proceedings of the ICIP, Taipei, Taiwan, 22-25 September 2019; pp. 1865–1869. [Google Scholar]
- Bdair, T.; Navab, N.; Albarqouni, S. FedPerl: Semi-supervised peer learning for skin lesion classification. In Proceedings of the MICCAI, Virtual, 27 September–1 October 2021; pp. 336–346. [Google Scholar]
- Wicaksana, J.; Yan, Z.; Zhang, D.; Huang, X.; Wu, H.; Yang, X.; Cheng, K.T. FedMix: Mixed supervised federated learning for medical image segmentation. IEEE Trans. Med. Imaging 2022, 42, 1955–1968. [Google Scholar] [CrossRef] [PubMed]
- Ruan, J.; Xiang, S.; Xie, M.; Liu, T.; Fu, Y. MALUNet: A multi-attention and light-weight unet for skin lesion segmentation. In Proceedings of the BIBM, Las Vegas, NV, USA and Changsha, Hunan, China, 6–9 December 2022; pp. 1150–1156. [Google Scholar]
- Yang, T.; Xu, J.; Zhu, M.; An, S.; Gong, M.; Zhu, H. FedZaCt: Federated learning with Z average and cross-teaching on image segmentation. Electronics 2022, 11, 3262. [Google Scholar] [CrossRef]
- Chen, Z.; Yang, C.; Zhu, M.; Peng, Z.; Yuan, Y. Personalized retrogress-resilient federated learning toward imbalanced medical data. IEEE Trans. Med. Imaging 2022, 41, 3663–3674. [Google Scholar] [CrossRef]
- Liu, F.; Yang, F. Medical image segmentation based on federated distillation optimization learning on non-IID data. In Proceedings of the IEEE ICIC, Zhengzhou, China, 10–13 August 2023; pp. 347–358. [Google Scholar]
- Sanderson, E.; Matuszewski, B.J. FCN-transformer feature fusion for polyp segmentation. In Proceedings of the MIUA, Cambridge, UK, 27–29 July 2022; pp. 892–907. [Google Scholar]
- Subedi, R.; Gaire, R.R.; Ali, S.; Nguyen, A.; Stoyanov, D.; Bhattarai, B. A Client-Server deep federated learning for cross-domain surgical image segmentation. arXiv 2023, arXiv:2306.08720. [Google Scholar]
- Tomar, N.K.; Jha, D.; Bagci, U. DilatedSegnet: A deep dilated segmentation network for polyp segmentation. In Proceedings of the MMM, Qui Nhon, Vietnam, 5–8 April 2023; pp. 334–344. [Google Scholar]
- Duc, N.T.; Oanh, N.T.; Thuy, N.T.; Triet, T.M.; Dinh, V.S. ColonFormer: An efficient transformer based method for colon polyp segmentation. IEEE Access 2022, 10, 80575–80586. [Google Scholar] [CrossRef]
- Wang, J.; Huang, Q.; Tang, F.; Meng, J.; Su, J.; Song, S. Stepwise feature fusion: Local guides global. In Proceedings of the MICCAI, Singapore, 18–22 September 2022; pp. 110–120. [Google Scholar]
- Zhu, M.; Chen, Z.; Yuan, Y. FedDM: Federated weakly supervised segmentation via annotation calibration and gradient de-conflicting. IEEE Trans. Med. Imaging 2023, 42, 1632–1643. [Google Scholar] [CrossRef] [PubMed]
- Nasr, M.; Islam, M.M.; Shehata, S.; Karray, F.; Quintana, Y. Smart healthcare in the age of AI: Recent advances, challenges, and future prospects. IEEE Access 2021, 9, 145248–145270. [Google Scholar] [CrossRef]
- Rath, K.C.; Khang, A.; Rath, S.K.; Satapathy, N.; Satapathy, S.K.; Kar, S. Artificial intelligence (AI)-enabled technology in medicine-advancing holistic healthcare monitoring and control systems. In Computer Vision and AI-Integrated IoT Technologies in the Medical Ecosystem; CRC Press: Boca Raton, FL, USA, 2024; pp. 87–108. [Google Scholar]
- Mansour, R.F.; El Amraoui, A.; Nouaouri, I.; Díaz, V.G.; Gupta, D.; Kumar, S. Artificial intelligence and internet of things enabled disease diagnosis model for smart healthcare systems. IEEE Access 2021, 9, 45137–45146. [Google Scholar] [CrossRef]
- Moreira, M.W.; Rodrigues, J.J.; Korotaev, V.; Al-Muhtadi, J.; Kumar, N. A comprehensive review on smart decision support systems for health care. IEEE Syst. J. 2019, 13, 3536–3545. [Google Scholar] [CrossRef]
- Katal, N. Ai-driven healthcare services and infrastructure in smart cities. In Smart Cities; CRC Press: Boca Raton, FL, USA, 2024; pp. 150–170. [Google Scholar]






| Model | Blastocyst Dataset | HAM10K Dataset | KVASIR-SEG Dataset | ||||||
|---|---|---|---|---|---|---|---|---|---|
| C | L | S | C | L | S | C | L | S | |
| UNet | 0.8643 | 0.7726 | 0.8593 | 0.8672 | 0.8320 | 0.8640 | 0.8271 | 0.6946 | 0.8042 |
| SegNet | 0.8475 | 0.7416 | 0.8475 | 0.8426 | 0.7773 | 0.8620 | 0.7337 | 0.5713 | 0.7669 |
| SUNet | 0.8487 | 0.7566 | 0.8504 | 0.8679 | 0.8241 | 0.8539 | 0.7280 | 0.6006 | 0.7233 |
| DeepLabV3 | 0.8768 | 0.8016 | 0.8369 | 0.8699 | 0.7715 | 0.8696 | 0.8438 | 0.7715 | 0.8262 |
| DeepLabV3+ | 0.8774 | 0.6834 | 0.8591 | 0.8715 | 0.8311 | 0.8262 | 0.8264 | 0.6965 | 0.8278 |
| RefineNet | 0.7881 | 0.6948 | 0.8181 | 0.8584 | 0.8161 | 0.8403 | 0.7083 | 0.6669 | 0.7652 |
| Attention UNet | 0.8673 | 0.6990 | 0.8605 | 0.8654 | 0.8241 | 0.8699 | 0.8236 | 0.6991 | 0.7961 |
| Swin-UNet | 0.8074 | 0.6283 | 0.8142 | 0.8492 | 0.7768 | 0.8478 | 0.7871 | 0.5483 | 0.6642 |
| CGNet | 0.8433 | 0.7287 | 0.7891 | 0.8728 | 0.8382 | 0.8490 | 0.8354 | 0.6868 | 0.8110 |
| DUCK-Net | 0.8725 | 0.7994 | 0.8321 | 0.8652 | 0.8389 | 0.8600 | 0.8824 | 0.7778 | 0.7800 |
| Average over models | 0.8493 | 0.7306 | 0.8367 | 0.8630 | 0.8130 | 0.8543 | 0.7996 | 0.6713 | 0.7765 |
| Sample | Ground Truth | UNet | SegNet | SUNet | DeepLab V3 | DeepLab V3+ | RefineNet | Attention UNet | Swin-UNet | CGNet | DUCK-Net |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Blastocyst Dataset | |||||||||||
 Blast_PCRM_R14-0411a.BMP |  |  |  |  |  |  |  |  |  |  |  |
| HAM10K Dataset | |||||||||||
 ISIC_0024308.jpg |  |  |  |  |  |  |  |  |  |  |  |
| KVASIR-SEG Dataset | |||||||||||
 cju7bgnvb1sf808717qa799ir.jpg |  |  |  |  |  |  |  |  |  |  |  |
| Model | Centralized Models | Locally Centralized Models | SplitFed Models |
|---|---|---|---|
| Blastocyst dataset | DeepLabV3+, DeepLabV3, DUCK-Net | DeepLabV3, DUCK-Net, CGNet | Attention UNet, DeepLabV3+, SUNet |
| HAM10K dataset | CGNet, DeepLabV3+, DeepLabV3 | DUCK-Net, CGNet, DeepLabV3+ | Attention UNet, DeepLabV3, UNet |
| KVASIR-SEG dataset | DUCK-Net, DeepLabV3, CGNet | DUCK-Net, DeepLabV3, DeepLabV3+ | DeepLabV3+, DeepLabV3, CGNet |
| Model | Trainable Parameters (TPs) | FLOPs | TP as a %. of UNet | FLOPs as a %. of UNet |
|---|---|---|---|---|
| UNet | 7.76M | 10.52 GMAC | 1% | 1% |
| SegNet | 9.44M | 7.04 GMAC | 1.22% | 0.67% |
| SUNet | 14.1M | 24 GMAC | 1.82% | 2.28% |
| DeepLabV3 | 28.32M | 12.83 GMAC | 3.64% | 1.21% |
| DeepLabV3+ | 54.70M | 15.85 GMAC | 7.05% | 1.50% |
| RefineNet | 118M | 50.24 GMAC | 15.20% | 4.77% |
| Attention UNet | 34.87M | 51.03 GMAC | 4.50% | 4.85% |
| Swin-UNet | 41.38M | 8.67 GMAC | 5.33% | 0.82% |
| CGNet | 0.30M | 541.69 MMAC | 0.039% | 0.05% |
| DUCK-Net | 22.67M | 12.55 GMAC | 2.92% | 1.19% |
| Model | Layer Proportions | Trainable Parameters (TP) | FLOPs | ||||||
|---|---|---|---|---|---|---|---|---|---|
| FE | SS | BE | FE | SS | BE | FE | SS | BE | |
| UNet | 2 (8.0%) | 21 (84.0%) | 2 (8.0%) | 0.002M (0.03%) | 7.75M (99.85%) | 0.009M (0.12%) | 123.73 MMAC | 13.09 GMAC | 614.5 MMAC |
| SegNet | 2 (6.25%) | 28 (87.50%) | 2 (6.25%) | 0.001M (0.01%) | 9.43M (99.90%) | 0.009M (0.10%) | 0.057 GMAC | 8.46 GMAC | 0.61 GMAC |
| SUNet | 4 (6.78%) | 52 (88.14%) | 3 (5.08%) | 0.006M (0.47%) | 13.91M (98.72%) | 0.11M (0.81%) | 0.415 GMAC | 38.817 GMAC | 1.030 GMAC |
| DeepLabV3 | 1 (1.61%) | 59 (95.16%) | 2 (3.23%) | 0.009M (0.03%) | 28.24M (99.69%) | 0.074M (0.26%) | 0.31 GMAC | 27.98 GMAC | 0.77 GMAC |
| DeepLabV3+ | 1 (1.30%) | 73 (94.81%) | 3 (3.89%) | 0.009M (0.02%) | 54.403M (99.41%) | 0.32M (0.57%) | 0.19 GMAC | 31.85 GMAC | 0.49 GMAC |
| RefineNet | 1 (0.71%) | 138 (98.57%) | 1 (0.71%) | 0.009M (0.01%) | 117.85M (99.95%) | 0.012M (0.01%) | 0.31 GMAC | 59.84 GMAC | 0.08 GMAC |
| Attention UNet | 1 (2.63%) | 35 (92.11%) | 2 (5.26%) | 0.35M (0.01%) | 34.82M (99.88%) | 0.009M (0.12%) | 0.042 GMAC | 13.10 GMAC | 0.61 GMAC |
| Swin-UNet | 2 (6.9%) | 24 (82.8%) | 3 (10.3%) | 0.29M (0.7%) | 39.23M (94.8%) | 1.86M (4.5%) | 0.08 GMAC | 8.18 GMAC | 0.42 GMAC |
| CGNet | 3 (11.11%) | 13 (48.15%) | 11 (40.74%) | 0.02M (6.69%) | 0.06M (20.13%) | 0.22M (73.18%) | 0.32 GMAC | 0.58 GMAC | 0.66 GMAC |
| DuckNet | 2 (6.67%) | 26 (86.67%) | 2 (6.67%) | 0.11M (0.50%) | 21.08M (93.00%) | 1.47M (6.50%) | 0.11 GMAC | 11.55 GMAC | 0.89 GMAC |
| SoTA Research | Centralized Models IoU | Federated Models IoU | SplitFed Models IoU |
|---|---|---|---|
| Blastocyst Dataset | |||
| Our Previous research | 0.798 (BLAST-NET [92]), 0.817 [88] | 0.810 [88] | 0.825 [88] |
| HAM10K Dataset | |||
| FedPerl (Efficient-Net) [93] | 0.769 | 0.747 | NA |
| FedMix (UNet) [94] | NA | 0.819 ± 1.7 | NA |
| MALUNet [95] | 0.802 | NA | NA |
| FedZaCt (UNet) [96] | 0.855 | 0.856 | NA |
| FedZaCt (DeepLabV3+) [96] | 0.861 | 0.863 | NA |
| Chen et al. [97] | NA | 0.892 | NA |
| FedDTM (UNet) [98] | NA | 0.7994 | NA |
| KVASIR-SEG Dataset | |||
| DUCK-Net [17] | 0.9502 | NA | NA |
| FCN-Transformer [99] | 0.9220 | NA | NA |
| MSRF-Net [72] | 0.8508 | NA | NA |
| PraNet [70] | 0.7286 | NA | NA |
| HRNetV2 [60] | 0.8530 | NA | NA |
| Subedi et al. [100] | 0.81 | 0.823 | NA |
| DilatedSegNet [101] | 0.8957 | NA | NA |
| DeepLabV3+ [101] | 0.8837 | NA | NA |
| Colonformer [102] | 0.877 | 0.876 | NA |
| SSFormer-S [103] | 0.8743 | NA | NA |
| SSFormer-L [103] | 0.8905 | NA | NA |
| FedDM [104] | 0.5275 ± 0.0002 | 0.6877 ± 0.0308 | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Shiranthika, C.; Kafshgari, Z.H.; Hadizadeh, H.; Saeedi, P. MedSegNet10: A Publicly Accessible Network Repository for Split Federated Medical Image Segmentation. Bioengineering 2026, 13, 104. https://doi.org/10.3390/bioengineering13010104
Shiranthika C, Kafshgari ZH, Hadizadeh H, Saeedi P. MedSegNet10: A Publicly Accessible Network Repository for Split Federated Medical Image Segmentation. Bioengineering. 2026; 13(1):104. https://doi.org/10.3390/bioengineering13010104
Chicago/Turabian StyleShiranthika, Chamani, Zahra Hafezi Kafshgari, Hadi Hadizadeh, and Parvaneh Saeedi. 2026. "MedSegNet10: A Publicly Accessible Network Repository for Split Federated Medical Image Segmentation" Bioengineering 13, no. 1: 104. https://doi.org/10.3390/bioengineering13010104
APA StyleShiranthika, C., Kafshgari, Z. H., Hadizadeh, H., & Saeedi, P. (2026). MedSegNet10: A Publicly Accessible Network Repository for Split Federated Medical Image Segmentation. Bioengineering, 13(1), 104. https://doi.org/10.3390/bioengineering13010104







