Oxidative Damage of Mussels Living in Seawater Enriched with Trace Metals, from the Viewpoint of Proteins Expression and Modification
Abstract
1. Introduction
2. Materials and Methods
2.1. Exposure of Mussels to Trace Metals
2.2. Biomarkers’ Analyses
2.2.1. Metal Concentration in Mussels
2.2.2. Oxidative Damage Indicators
2.2.3. Anti-Oxidant Barrier
2.3. Biochemical Preparations
2.4. DNPH Dot Blot Assay for theQuantification of Carbonylated Proteins
2.5. WesternBlot Detection of Specific Proteins
2.6. Statistical Analysis
3. Results
3.1. Stress Biomarkers
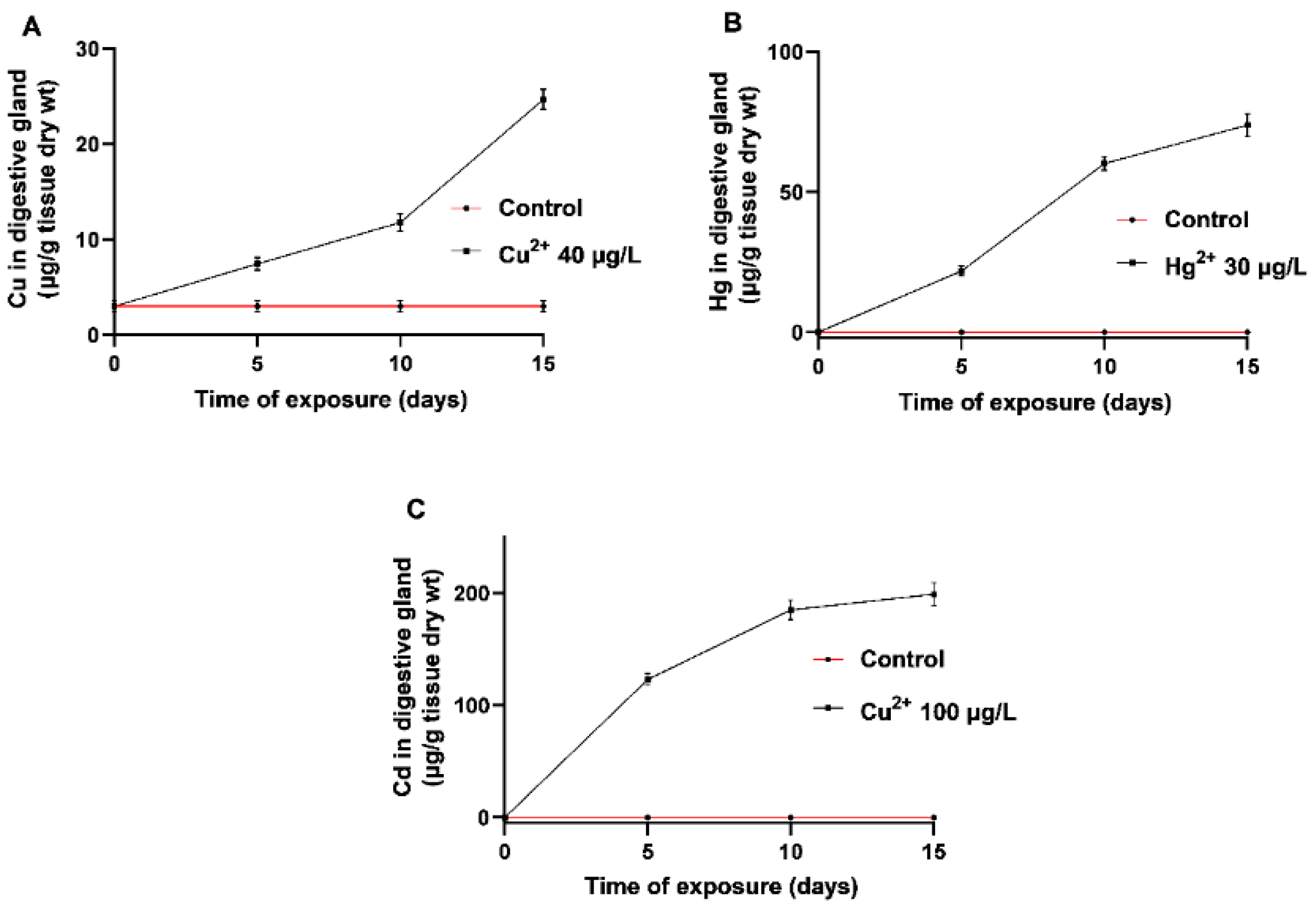
3.1.1. Metal Bioaccumulation
3.1.2. Oxidative Damage Indicators
3.1.3. Anti-OxidantBarrier
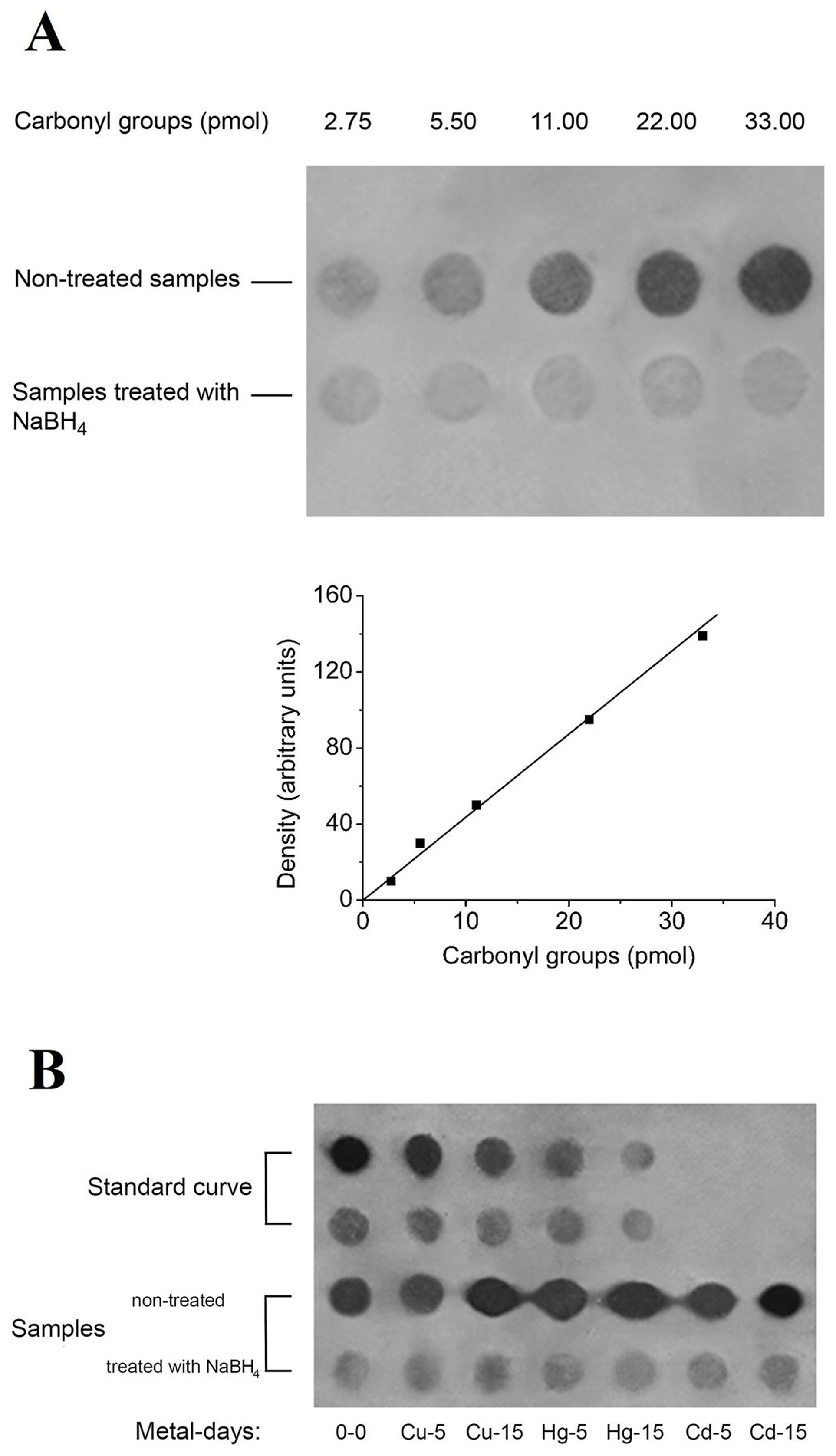
3.2. Dot Blot Analysis of Carbonylated Proteins
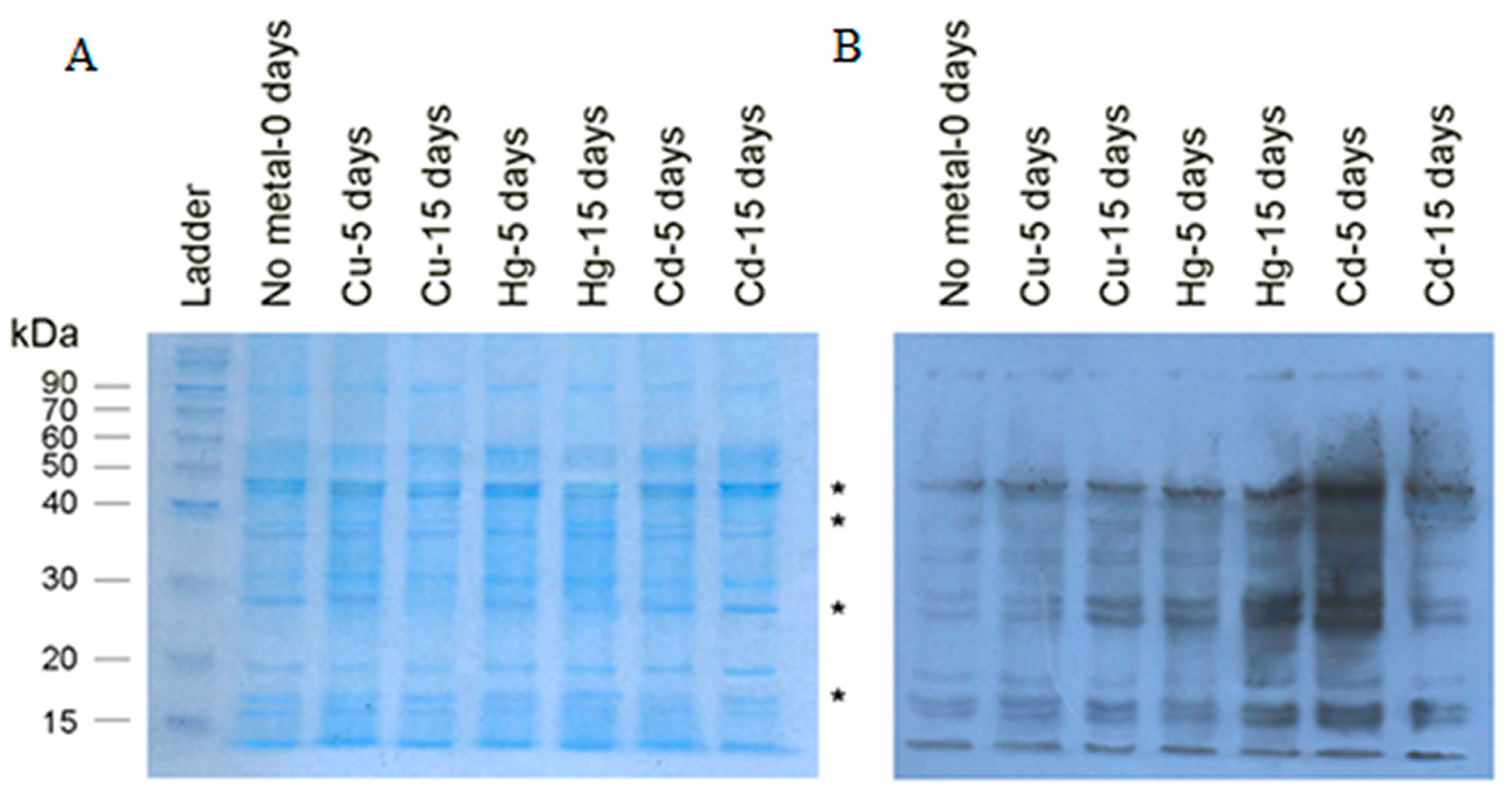
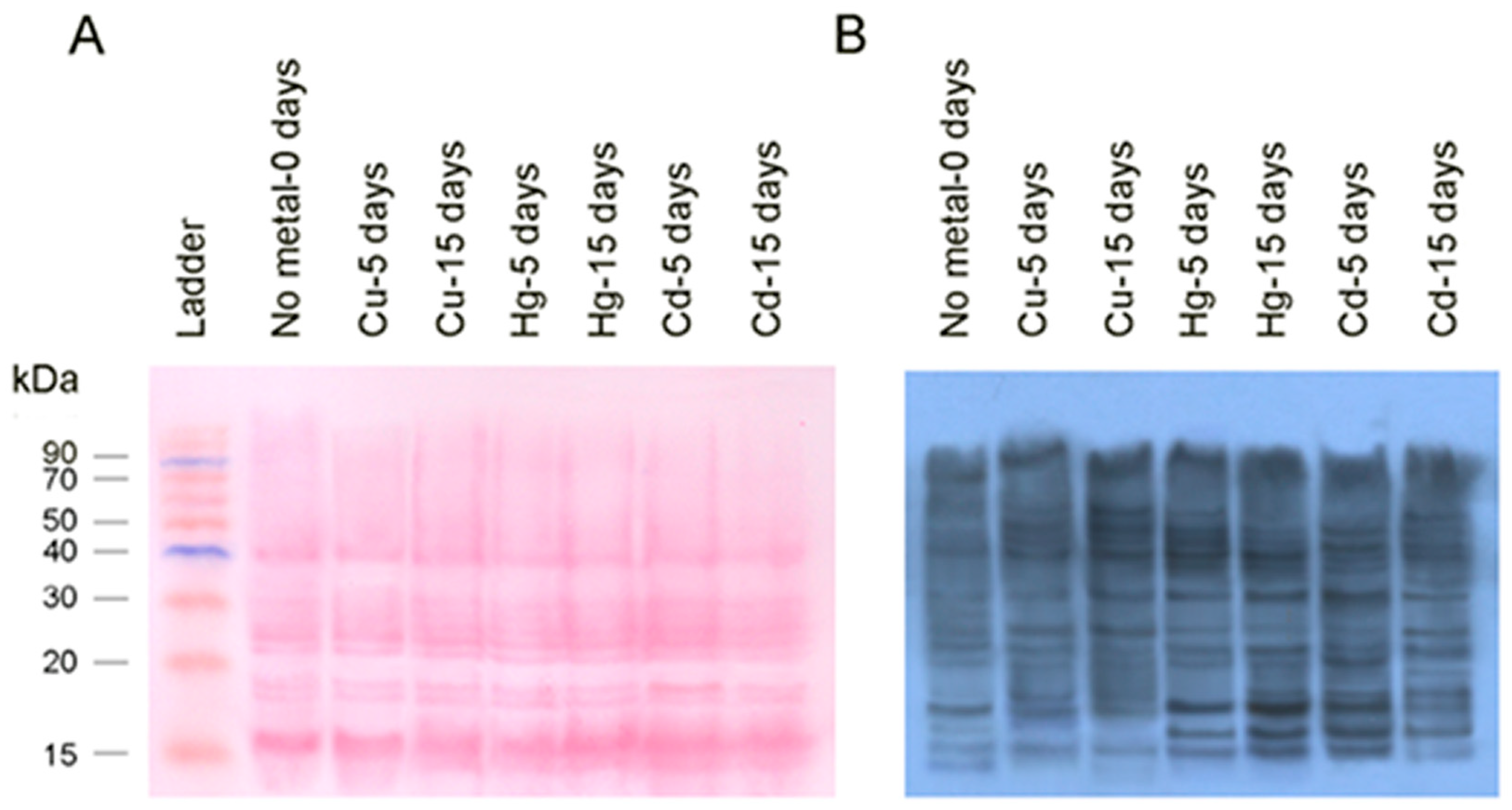
3.3. WesternBlot Analysis
- A.
- Carbonylated FWR
- B. Carbonylated Ribosomal Proteins
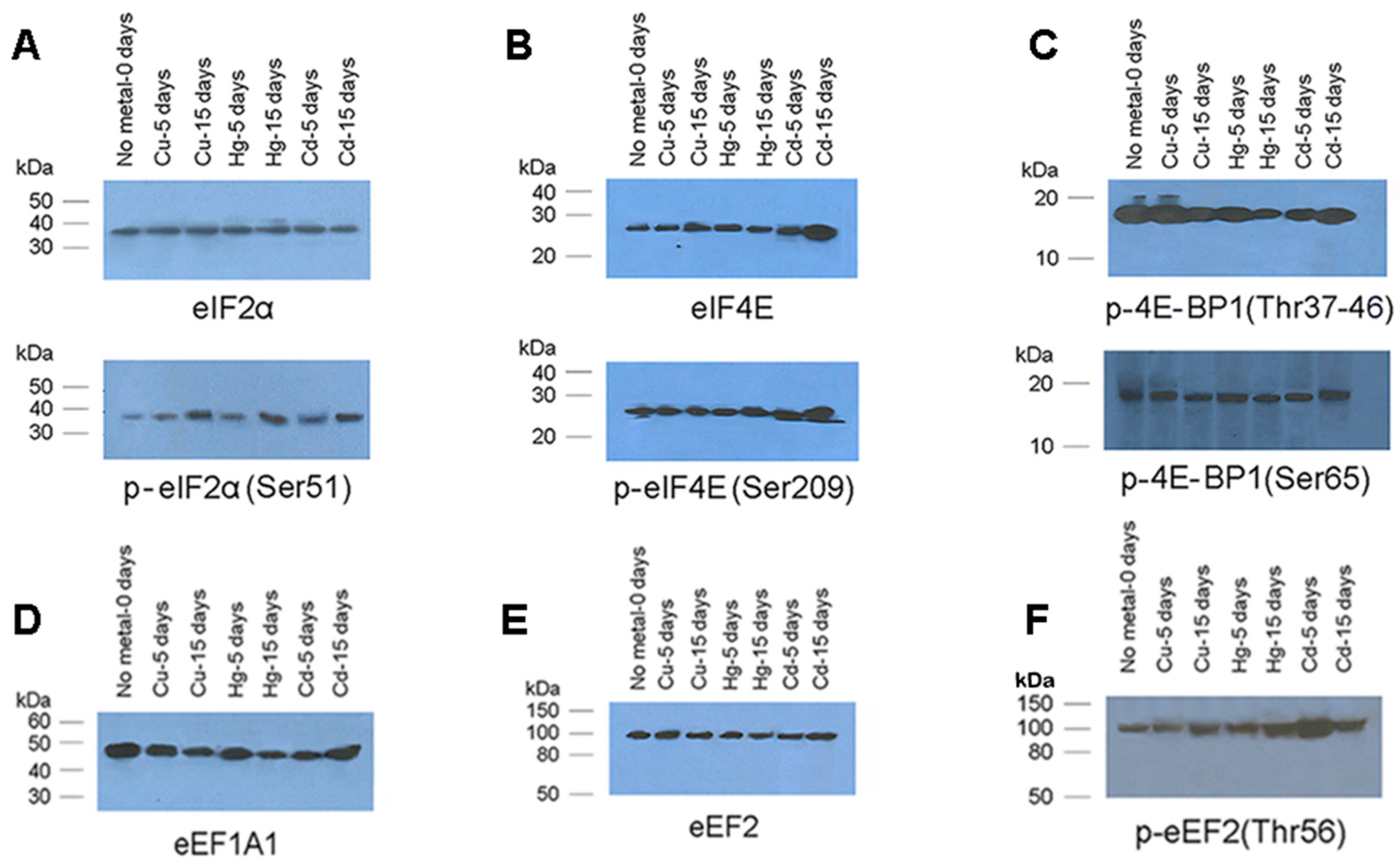
3.4. Translation Factors Expression
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Beyer, J.; Green, N.W.; Brooks, S.; Allan, I.J.; Ruus, A.; Gomes, T.; Bråte, I.; Schøyen, M. Blue mussels (Mytilus edulis spp.) as sentinel organisms in coastal pollution monitoring: A review. Mar. Environ. Res. 2017, 130, 338–365. [Google Scholar] [CrossRef]
- Sforzini, S.; Oliveri, C.; Orrù, A.; Chessa, G.; Pacchioni, B.; Millino, C.; Jha, A.N.; Viarengo, A.; Banni, M. Application of a new targeted low density microarray and conventional biomarkers to evaluate the health status of marine mussels: A field study in Sardinian coast, Italy. Sci. Total Environ. 2018, 628–629, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Franco, R.; Sánchez-Olea, R.; Reyes-Reyes, E.M.; Panayiotidis, M.I. Environmental toxicity, oxidative stress and apoptosis: Menage atrois. Mutat. Res. 2009, 674, 3–22. [Google Scholar] [CrossRef]
- Nzengue, Y.; Candéias, S.M.; Sauvaigo, S.; Douki, T.; Favier, A.; Rachidi, W.; Guiraud, P. The toxicity redox mechanisms of cadmium alone or together with copper and zinc homeostasis alteration: Its redox biomarkers. J. Trace Elem. Med. Biol. 2011, 25, 171–180. [Google Scholar] [CrossRef] [PubMed]
- Letelier, M.E.; Lepe, A.M.; Faúndez, M.; Salazar, J.; Marín, R.; Aracena, P.; Speisky, H. Possible mechanisms underlying copper-induced damage in biological membranes leading to cellular toxicity. Chem. Biol. Interact. 2005, 151, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Gomes, T.; Chora, S.; Pereira, C.G.; Cardoso, C.; Bebianno, M.J. Proteomic response of mussels Mytilus galloprovincialis exposed to CuONPs and Cu2+: An exploratory biomarker discovery. Aquat. Toxicol. 2014, 155, 327–336. [Google Scholar] [CrossRef]
- Calvo, J.; Jung, H.; Meloni, G. Copper metallothioneins. IUBMB Life 2017, 69, 236–245. [Google Scholar] [CrossRef]
- Dondero, F.; Piacentini, L.; Banni, M.; Rebelo, M.; Burlando, B.; Viarengo, A. Quantitative PCR analysis of two molluscan metallothionein genes unveils differential expression and regulation. Gene 2005, 345, 259–270. [Google Scholar] [CrossRef]
- Domouhtsidou, G.P.; Dailianis, S.; Kaloyanni, M.; Dimitriadis, V.K. Lysosomal membrane stability and metallothionein content in Mytilus galloprovincialis (Lam.), as biomarkers. Combination with trace metal concentrations. Mar. Pollut. Bull. 2004, 48, 572–586. [Google Scholar] [CrossRef]
- Kournoutou, G.G.; Giannopoulou, P.C.; Sazakli, E.; Leotsinidis, M.; Kalpaxis, D.L. Oxidative damage of 18S and 5S ribosomal RNA in digestive gland of mussels exposed to trace metals. Aquat. Toxicol. 2017, 192, 136–147. [Google Scholar] [CrossRef]
- Pytharopoulou, S.; Kournoutou, G.G.; Leotsinidis, M.; Georgiou, C.D.; Kalpaxis, D.L. Cadmium versus copper toxicity: Insights from an integrated dissection of protein synthesis pathway in the digestive glands of mussel Mytilus galloprovincialis. J. Hazard. Mater. 2013, 260, 263–271. [Google Scholar] [CrossRef] [PubMed]
- Zorita, I.; Ortiz-Zarragoitia, M.; Soto, M.; Cajaraville, M.P. Biomarkers in mussels from a copper site gradient (Visnes, Norway): An integrated biochemical, histochemical and histological study. Aquat. Toxicol. 2006, 78 (Suppl. 1), S109–S116. [Google Scholar] [CrossRef]
- Bolognesi, C.; Landini, E.; Roggieri, P.; Fabbri, R.; Viarengo, A. Genotoxicity biomarkers in the assessment of heavy metal effects in mussels: Experimental studies. Environ. Mol. Mutagen. 1999, 33, 287–292. [Google Scholar] [CrossRef]
- Pytharopoulou, S.; Grintzalis, K.; Sazakli, E.; Leotsinidis, M.; Georgiou, C.D.; Kalpaxis, D.L. Translational responses and oxidative stress of mussels experimentally exposed to Hg, Cu and Cd: One pattern does not fit at all. Aquat. Toxicol. 2011, 105, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Pytharopoulou, S.; Kournoutou, G.G.; Leotsinidis, M.; Georgiou, C.D.; Kalpaxis, D.L. Dysfunctions of the translational machinery in digestive glands of mussels exposed to mercury ions. Aquat. Toxicol. 2013, 134, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Tamás, M.J.; Fauvet, B.; Christen, P.; Goloubinoff, P. Misfolding and aggregation of nascent proteins: A novel mode of toxic cadmium action in vivo. Curr. Genet. 2018, 64, 177–181. [Google Scholar] [CrossRef]
- Gardarin, A.; Chédin, S.; Lagniel, G.; Aude, J.C.; Godat, E.; Catty, P.; Labarre, J. Endoplasmic reticulum is a major target of cadmium toxicity in yeast. Mol. Microbiol. 2010, 76, 1034–1048. [Google Scholar] [CrossRef] [PubMed]
- Jin, Y.H.; Clark, A.B.; Slebos, R.J.; Al-Refai, H.; Taylor, J.A.; Kunkel, T.A.; Resnick, M.A.; Gordenin, D.A. Cadmium is a mutagen that acts by inhibiting mismatch repair. Nat. Genet. 2003, 34, 326–329. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Ji, C.; Wu, H.; Tan, Q.; Wang, W.X. A comparative proteomic study on the effects of metal pollution in oysters Crassostrea hongkongensis. Mar. Pollut. Bull. 2016, 112, 436–442. [Google Scholar] [CrossRef]
- Polle, A.; Rennenberg, H. Significance of antioxidants in plant adaptation to environmental stress. In Plant Adaptation to Environmental Stress; Mansfield, T., Fowden, L., Stoddard, F., Eds.; Chapman & Hall: London, UK, 1993; pp. 263–273. [Google Scholar]
- Branca, J.J.; Fiorillo, C.; Carrino, D.; Paternostro, F.; Taddei, N.; Gulisano, M.; Pacini, A.; Becatti, M. Cadmium-Induced Oxidative Stress: Focus on the Central Nervous System. Antioxidants 2020, 9, 492. [Google Scholar] [CrossRef]
- Bernotiene, R.; Ivanoviene, L.; Sadauskiene, I.; Liekis, A.; Ivanov, L. The effects of cadmium chloride and sodium selenite on protein synthesis in mouse liver. Environ. Toxicol. Pharmacol. 2013, 36, 1261–1265. [Google Scholar] [CrossRef]
- Papaconstantinou, A.D.; Brown, K.M.; Noren, B.T.; McAlister, T.; Fisher, B.R.; Goering, P.L. Mercury, cadmium, and arsenite enhance heat shock protein synthesis in chick embryos prior to embryotoxicity. Birth Defects Res. B Dev. Reprod. Toxicol. 2003, 68, 456–464. [Google Scholar] [CrossRef]
- Smith, R.W.; Blaney, S.C.; Dowling, K.; Sturm, A.; Jönsson, M.; Houlihan, D.F. Protein synthesis costs could account for the tissue-specific effects of sub-lethal copper on protein synthesis in rainbow trout (Oncorhynchus mykiss). Aquat. Toxicol. 2001, 53, 265–277. [Google Scholar] [CrossRef]
- Franzellitti, S.; Fabbri, E. Differential HSP70 gene expression in the Mediterranean mussel exposed to various stressors. Biochem. Biophys. Res. Commun. 2005, 336, 1157–1163. [Google Scholar] [CrossRef] [PubMed]
- Géret, F.; Jouan, A.; Turpin, V.; Bebianno, M.J.; Cosson, R.P. Influence of metal exposure on metallothionein synthesis and lipid peroxidation in two bivalve mollusks: The oyster (Crassostrea gigas) and the mussel (Mytilus edulis). Aquat. Living Resour. 2002, 15, 61–66. [Google Scholar] [CrossRef]
- Liu, B.; Qian, S.B. Translational reprogramming in cellular stress response. WIREs RNA 2014, 5, 301–305. [Google Scholar] [CrossRef] [PubMed]
- Achard-Joris, M.; Gonzalez, P.; Marie, V.; Baudrimont, M.; Bourdineaud, J.P. cDNA cloning and gene expression of ribosomal S9 protein gene in the mollusk Corbicula fluminea: A new potential biomarker of metal contamination up-regulated by cadmium and repressed by zinc. Environ. Toxicol. Chem. 2006, 25, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, R.; Lourenço, J.; Pereira, P.; Carvalho, S.; Mendo, S. Effects of metal contamination on the gene expression profile of two benthic species: Cerastoderma edule and Ruditapes philippinarum. Mar. Pollut. Bull. 2017, 125, 157–165. [Google Scholar] [CrossRef]
- Rossi, F.; Palombella, S.; Pirrone, C.; Mancini, G.; Bernardini, G.; Gornati, R. Evaluation of tissue morphology and gene expression as biomarkers of pollution in mussel Mytilus galloprovincialis caging experiment. Aquat. Toxicol. 2016, 181, 57–66. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Varotto, L.; Domeneghetti, S.; Rosani, U.; Manfrin, C.; Cajaraville, M.P.; Raccanelli, S.; Pallavicini, A.; Venier, P. DNA damage and transcriptional changes in the gills of Mytilus galloprovincialis exposed to nanomolar doses of combined metal salts (Cd, Cu, Hg). PLoS ONE 2013, 8, e54602. [Google Scholar] [CrossRef]
- Xu, M.; Jiang, L.; Shen, K.N.; Wu, C.; He, G.; Hsiao, C.D. Transcriptome response to copper heavy metal stress in hard-shelled mussel (Mytilus coruscus). Genom. Data 2016, 7, 152–154. [Google Scholar] [CrossRef] [PubMed]
- Shcherbik, N.; Pestov, D.G. The Impact of Oxidative Stress on Ribosomes: From Injury to Regulation. Cells 2019, 8, 1379. [Google Scholar] [CrossRef] [PubMed]
- Storey, K.B.; Storey, J.M. Metabolic rate depression in animals: Transcriptional and translational controls. Biol. Rev. Camb. Philos. Soc. 2004, 79, 207–233. [Google Scholar] [CrossRef]
- Torrent, M.; Chalancon, G.; de Groot, N.S.; Wuster, A.; Babu, M.M. Cells alter their tRNA abundance to selectively regulate protein synthesis during stress conditions. Sci. Signal. 2018, 11, eaat6409. [Google Scholar] [CrossRef]
- Avery, S.V. Molecular targets of oxidative stress. Biochem. J. 2011, 434, 201–210. [Google Scholar] [CrossRef]
- Honda, K.; Smith, M.A.; Zhu, X.; Baus, D.; Merrick, W.C.; Tartakoff, A.M.; Harrier, T.; Harris, P.L.; Siedlack, S.L.; Fujioka, H.; et al. Ribosomal RNA in Alzheimer disease is oxidized by bound redox-active iron. J. Biol. Chem. 2005, 280, 20978–20986. [Google Scholar] [CrossRef]
- Liu, M.; Gong, X.; Alluri, R.K.; Wu, J.; Sablo, T.; Li, Z. Characterization of RNA damage under oxidative stress in Escherichia coli. Biol. Chem. 2012, 393, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Mirzaei, H.; Regnier, F. Protein-RNA cross-linking in the ribosomes of yeast under oxidative stress. J. Proteome Res. 2006, 5, 3249–3259. [Google Scholar] [CrossRef] [PubMed]
- Celi, P.; Gabai, G. Oxidant/antioxidant balance in animal nutrition and health: The role of protein oxidation. Front. Vet. Sci. 2015, 2, 48. [Google Scholar] [CrossRef]
- Fedorova, M.; Bollineni, R.C.; Hoffmann, R. Protein carbonylation as a major hallmark of oxidative damage: Update of analytical strategies. Mass Spectrom. Rev. 2014, 33, 79–97. [Google Scholar] [CrossRef]
- Moore, M.N. Cytochemical demonstration of latency of lysosomal hydrolases in digestive cells of the common mussel, Mytilus edulis, and changes induced by thermal stress. Cell Tissue Res. 1976, 175, 279–287. [Google Scholar] [CrossRef]
- Bolognesi, C.; Fenech, M. Mussel micronucleus cytome assay. Nat. Protoc. 2012, 7, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Zamora, R.; Alaiz, M.; Hidalgo, F.J. Feed-back inhibition of oxidative stress by oxidized lipid/amino acid reaction products. Biochemistry 1997, 36, 15765–15771. [Google Scholar] [CrossRef] [PubMed]
- Grintzalis, K.; Georgiou, C.D.; Schneider, Y.J. An accurate and sensitive Coomassie Brilliant Blue G-250-based assay for protein determination. Anal. Biochem. 2015, 480, 28–30. [Google Scholar] [CrossRef]
- Georgiou, C.D.; Papapostolou, I.; Patsoukis, N.; Tsegenidis, T.; Sideris, T. An ultrasensitive fluorescent assay for the in vivo quantification of superoxide radical in organisms. Anal. Biochem. 2005, 347, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Viarengo, A.; Ponzano, E.; Dondero, F.; Fabbri, R. A simple spectro-photometric method for metallothionein evaluation in marine organisms: An application to Mediterranean and Antarctic molluscs. Mar. Environ. Res. 1997, 44, 69–84. [Google Scholar] [CrossRef]
- Pan, J.; Huang, X.; Li, Y.; Li, M.; Yao, N.; Zhou, Z.; Li, X. Zinc protects against cadmium-induced toxicity by regulating oxidative stress, ions homeostasis and protein synthesis. Chemosphere 2017, 188, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Koontz, L. TCA precipitation. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2014; Volume 541, pp. 3–10. [Google Scholar]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Barritault, D.; Expert-Bezançon, A.; Guérin, M.F.; Hayes, D. The use of acetone precipitation in the isolation of ribosomal proteins. FEBS J. 1976, 63, 131–135. [Google Scholar] [CrossRef]
- Alamdari, D.H.; Kostidou, E.; Paletas, K.; Sarigianni, M.; Konstas, A.G.; Karapiperidou, A.; Koliakos, G. High sensitivity enzyme-linked immunosorbent assay (ELISA) method for measuring protein carbonyl in samples with low amounts of protein. Free Rad. Biol. Med. 2005, 39, 1362–1367. [Google Scholar] [CrossRef]
- Luzhna, L.; Kathiria, P.; Kovalchuk, O. Micronuclei in genotoxicity assessment: From genetics to epigenetics and beyond. Front. Genet. 2013, 4, 131. [Google Scholar] [CrossRef]
- Dalle-Donne, I.; Rossi, R.; Colombo, R.; Giustarini, D.; Milzani, A. Biomarkers of oxidative damage in human disease. Clin. Chem. 2006, 52, 601–623. [Google Scholar] [CrossRef] [PubMed]
- Dalle-Donne, I.; Rossi, R.; Giustarini, D.; Milzani, A.; Colombo, R. Protein carbonyl groups as biomarkers of oxidative stress. Clin. Chim. Acta 2003, 329, 23–38. [Google Scholar] [CrossRef]
- Genuth, N.R.; Barna, M. The Discovery of Ribosome Heterogeneity and Its Implications for Gene Regulation and Organismal Life. Mol. Cell 2018, 71, 364–374. [Google Scholar] [CrossRef] [PubMed]
- Katz, M.J.; Gándara, L.; Ezcurra, A.D.L.; Wappner, P. Hydroxylation and translational adaptation to stress: Some answers lie beyond the STOP codon. Cell. Mol. Life Sci. 2016, 73, 1881–1893. [Google Scholar] [CrossRef] [PubMed]
- Proud, C.G. eIF2 and the control of cell physiology. Semin. Cell Dev. Biol. 2005, 16, 3–12. [Google Scholar] [CrossRef]
- van Kolck, M.; Huijbregts, M.A.J.; Veltman, K.; Hendriks, A.J. Estimating bioconcentration factors, lethal concentrations and critical body residues of metals in the mollusks Pernaviridis and Mytilus edulis using ion characteristics. Environ. Toxicol. Chem. 2008, 27, 272–276. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Xu, L.; Yu, D.; Ji, C. Differential metabolic responses in three life stages of mussels Mytilus galloprovincialis exposed to cadmium. Ecotoxicology 2016, 26, 74–80. [Google Scholar] [CrossRef]
- Dheri, G.S.; Brar, M.S.; Malhi, S.S. Heavy-metal concentration of sewage contaminated water and its impact on underground water, soil, and crop plants in alluvial soils of northwestern India. Commun. Soil Sci. Plant. Anal. 2007, 38, 1353–1370. [Google Scholar] [CrossRef]
- Şeker, S.; Kutlu, B. Determination of Copper (Cu) Levels for Rivers in Tunceli, Turkey. World Environ. 2014, 4, 168–171. [Google Scholar]
- Chandurvelan, R.; Marsden, I.D.; Gaw, S.; Glover, C.N. Biochemical biomarker responses of green-lipped mussel, Perna canaliculus, to acute and subchronic waterborne cadmium toxicity. Aquat. Toxicol. 2013, 140, 303–313. [Google Scholar] [CrossRef]
- Ebrahimi, M.; Taherianfard, M. Concentration of four heavy metals (cadmium, lead, mercury, and arsenic) in organs of two cyprinid fish (Cyprinus carpio and Capoeta sp.) from the Kor River (Iran). Environ. Monit. Assess. 2010, 168, 575–585. [Google Scholar] [CrossRef]
- Pakos-Zebrucka, K.; Koryga, I.; Mnich, K.; Ljujic, M.; Samali, A.; Gorman, A.M. The integrated stress response. EMBO Rep. 2016, 17, 1374–1395. [Google Scholar] [CrossRef]
- Truitt, M.L.; Conn, C.S.; Shi, Z.; Pang, X.; Tokuyasu, T.; Coady, A.M.; Seo, Y.; Barna, M.; Ruggero, D. Differential requirements for eIF4E dose in normal development and cancer. Cell 2015, 162, 59–71. [Google Scholar] [CrossRef] [PubMed]
- Gao, B.; Roux, P.P. Translational control by oncogenic signaling pathways. Biochim. Biophys. Acta 2015, 1849, 753–765. [Google Scholar] [CrossRef] [PubMed]
- Sekiyama, N.; Arthanari, H.; Papadopoulos, E.; Rodriguez-Mias, R.A.; Wagner, G.; Léger-Abraham, M. Molecular mechanism of the dual activity of 4EGI-1: Dissociating eIF4G from eIF4E but stabilizing the binding of unphosphorylated 4E-BP1. Proc. Natl. Acad. Sci. USA 2015, 112, E4036–E4045. [Google Scholar] [CrossRef] [PubMed]
- Gingras, A.C.; Raught, B.; Gygi, S.P.; Niedzwiecka, A.; Miron, M.; Burley, S.K.; Polakiewicz, R.D.; Wyslouch-Cieszynska, A.; Aebersold, R.; Sonenberg, N. Hierarchical phosphorylation of the translation inhibitor 4E-BP1. Genes Dev. 2001, 15, 2852–2864. [Google Scholar] [PubMed]
- Lewis, S.S.; Keller, S.J. Identification of copper-responsive genes in an early life stage of the fathead minnow Pimephales promelas. Ecotoxicology 2009, 18, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Xia, Y.; Chen, C.; Zhuang, K.; Song, Y.; Shen, Z. Analysis of Copper-Binding Proteins in Rice Radicles Exposed to Excess Copper and Hydrogen Peroxide Stress. Front. Plant. Sci. 2016, 7, 1216. [Google Scholar] [CrossRef]
- Dumont-Miscopein, A.; Lavergne, J.P.; Guillot, D.; Sontag, B.; Reboud, J.P. Interaction of phosphorylated elongation factor EF-2 with nucleotides and ribosomes. FEBS Lett. 1994, 356, 283–286. [Google Scholar] [CrossRef]
| Time of Exposure (Days) | ||||
|---|---|---|---|---|
| Parameter | Metal | 0 | 5 | 15 |
| Oxidative damage | ||||
| MN frequency (ppt *) | Cu Hg Cd | 2.6 ± 0.5 2.8 ± 0.8 2.6 ± 0.5 | 5.6 ± 0.5 b 6.0 ± 0.7 b 3.4 ± 0.9 | 10.0 ± 1.0 b 12.0 ±1.6 b 4.0 ± 1.2 b |
| SR (pmol/mg protein) | Cu Hg Cd | 2.6 ± 0.2 2.5 ± 0.1 2.6 ± 0.2 | 1.5 ± 0.2 b 2.7 ± 0.1 b 8.0 ± 1.0 b | 0.4 ± 0.1 b 2.6 ± 0.2 22.4 ± 2.1 b |
| LP (min) | Cu Hg Cd | 28.0 ± 2.0 30.0 ± 3.2 27.0 ± 1.9 | 16.0 ± 1.6 b 18.0 ± 1.6 b 9.0 ± 1.4 b | 9.0 ± 1.6 b 11.0 ± 1.0 b 6.5 ± 0.5 b |
| MDA (nmol/mg protein) | Cu Hg Cd | 2.0 ± 0.3 2.1 ± 0.3 2.0 ± 0.2 | 2.2 ± 0.2 2.4 ± 0.1 2.6 ± 0.2 b | 2.8 ± 0.3 b 2.6 ± 0.2 b 6.2 ± 0.4 b |
| Anti-oxidant barrier | ||||
| SOD (units/mg protein) | Cu Hg Cd | 0.8 ± 0.1 0.7 ± 0.1 0.8 ± 0.1 | 0.9 ± 0.1 1.8 ± 0.2 b 1.6 ± 0.2 b | 0.2 ± 0.1 b 0.6 ± 0.1 0.6 ± 0.1 b |
| MTs (μg/g tissue w.w) | Cu Hg Cd | 52.0 ± 5.0 50.0 ± 4.5 50.0 ± 4.5 | 56.0 ± 5.3 50.0 ± 4.5 210.0 ± 21.6 b | 53.0 ± 3.9 85.0 ± 7.4 b 354.0 ± 30.5 b |
| GSH (nmol/g tissue w.w.) | Cu Hg Cd | 740.0 ± 67.3 780.0 ± 60.3 720.0 ± 69.9 | 511.0 ± 44.6 b 565.0 ± 60.0 b 369.0 ± 40.1 b | 421.0 ± 45.3 b 275.0 ± 38.1 b 852.0 ± 98.2 b |
| GSSG (nmol/g tissue w.w.) | Cu Hg Cd | 117.0 ± 16.9 122.0 ± 13.7 112.0 ± 14.4 | 104.5 ± 10.2 110.5 ± 10.3 76.0 ± 6.9 b | 102.0 ± 10.0 95.5 ± 8.0 b 23.0 ± 2.9 b |
| GSH/GSSG ratio | Cu Hg Cd | 6.32 ± 1.1 6.39 ± 0.9 6.42 ± 1.0 | 4.89 ± 0.6 b 5.11 ± 0.7 4.85 ± 0.7 b | 4.13 ± 0.6 b 2.88 ± 0.5 b 37.04 ± 6.3 b |
| Time of Exposure (Days) | ||||
|---|---|---|---|---|
| Cellular Component | Metal | 0 | 5 | 15 |
| Total cytosolic proteins | Cu Hg Cd | 7.8 ± 1.2 7.5 ± 0.8 7.2 ± 0.8 | 22.0 ± 2.4 a 44.7 ± 4.3 a 52.6 ± 6.6 a | 48.5 ± 4.2 a 53.6 ± 6.2 a 28.4 ± 4.1 a |
| Proteins FWR fraction | Cu Hg Cd | 7.5 ± 0.6 6.8 ± 0.6 6.9 ± 0.7 | 10.5 ± 1.5 a 13.9 ± 1.4 a 18.4 ± 1.7 a | 17.3 ± 2.2 a 18.6 ± 2.4 a 12.2 ± 2.3 a |
| Proteins in 80S ribosome | Cu Hg Cd | 2.8 ± 0.2 3.1 ± 0.3 2.8 ± 0.3 | 3.4 ± 0.3 a 5.8 ± 0.4 a 6.2 ± 0.6 a | 7.4 ± 0.7 a 8.1± 0.8 a 4.0 ± 0.4 a |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kournoutou, G.G.; Giannopoulou, P.C.; Sazakli, E.; Leotsinidis, M.; Kalpaxis, D.L.; Dinos, G.P. Oxidative Damage of Mussels Living in Seawater Enriched with Trace Metals, from the Viewpoint of Proteins Expression and Modification. Toxics 2020, 8, 89. https://doi.org/10.3390/toxics8040089
Kournoutou GG, Giannopoulou PC, Sazakli E, Leotsinidis M, Kalpaxis DL, Dinos GP. Oxidative Damage of Mussels Living in Seawater Enriched with Trace Metals, from the Viewpoint of Proteins Expression and Modification. Toxics. 2020; 8(4):89. https://doi.org/10.3390/toxics8040089
Chicago/Turabian StyleKournoutou, Georgia G., Panagiota C. Giannopoulou, Eleni Sazakli, Michalis Leotsinidis, Dimitrios L. Kalpaxis, and George P. Dinos. 2020. "Oxidative Damage of Mussels Living in Seawater Enriched with Trace Metals, from the Viewpoint of Proteins Expression and Modification" Toxics 8, no. 4: 89. https://doi.org/10.3390/toxics8040089
APA StyleKournoutou, G. G., Giannopoulou, P. C., Sazakli, E., Leotsinidis, M., Kalpaxis, D. L., & Dinos, G. P. (2020). Oxidative Damage of Mussels Living in Seawater Enriched with Trace Metals, from the Viewpoint of Proteins Expression and Modification. Toxics, 8(4), 89. https://doi.org/10.3390/toxics8040089





