Epigenetic Modifications, and Alterations in Cell Cycle and Apoptosis Pathway in A549 Lung Carcinoma Cell Line upon Exposure to Perfluoroalkyl Substances
Abstract
1. Introduction
2. Materials and Methods
2.1. Dosing Solutions
2.2. Cell Culture
2.3. Cell Viability Assay
2.4. Isolation and Quantification of RNA and cDNA Synthesis
2.5. Quantitative Real-Time Polymerase Chain Reaction (qRT-PCR)
2.6. Intracellular Concentration Determination by UPLC-MS Analysis
2.7. Hyperspectral Dark-Field Microscopy (HS-DFM)
2.8. Statistical Analysis
3. Results
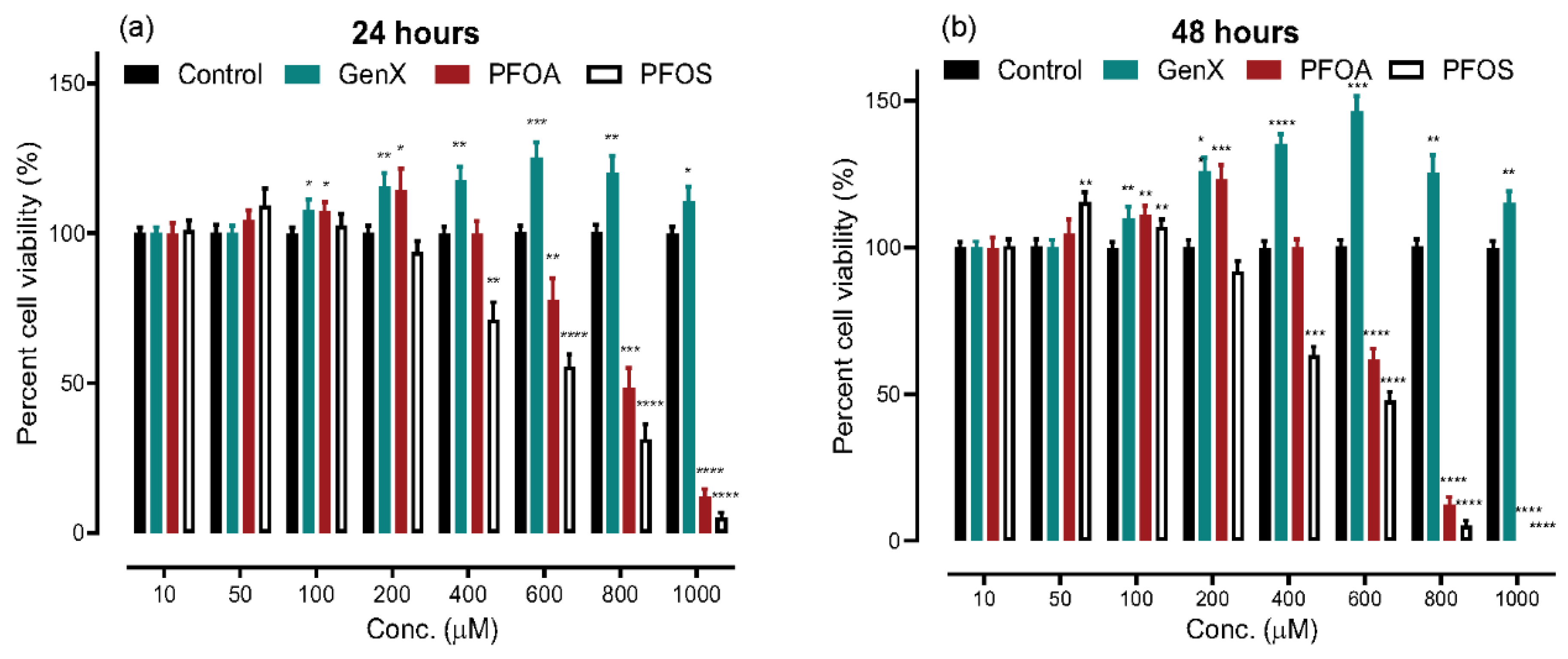
3.1. Cytotoxicity Estimation
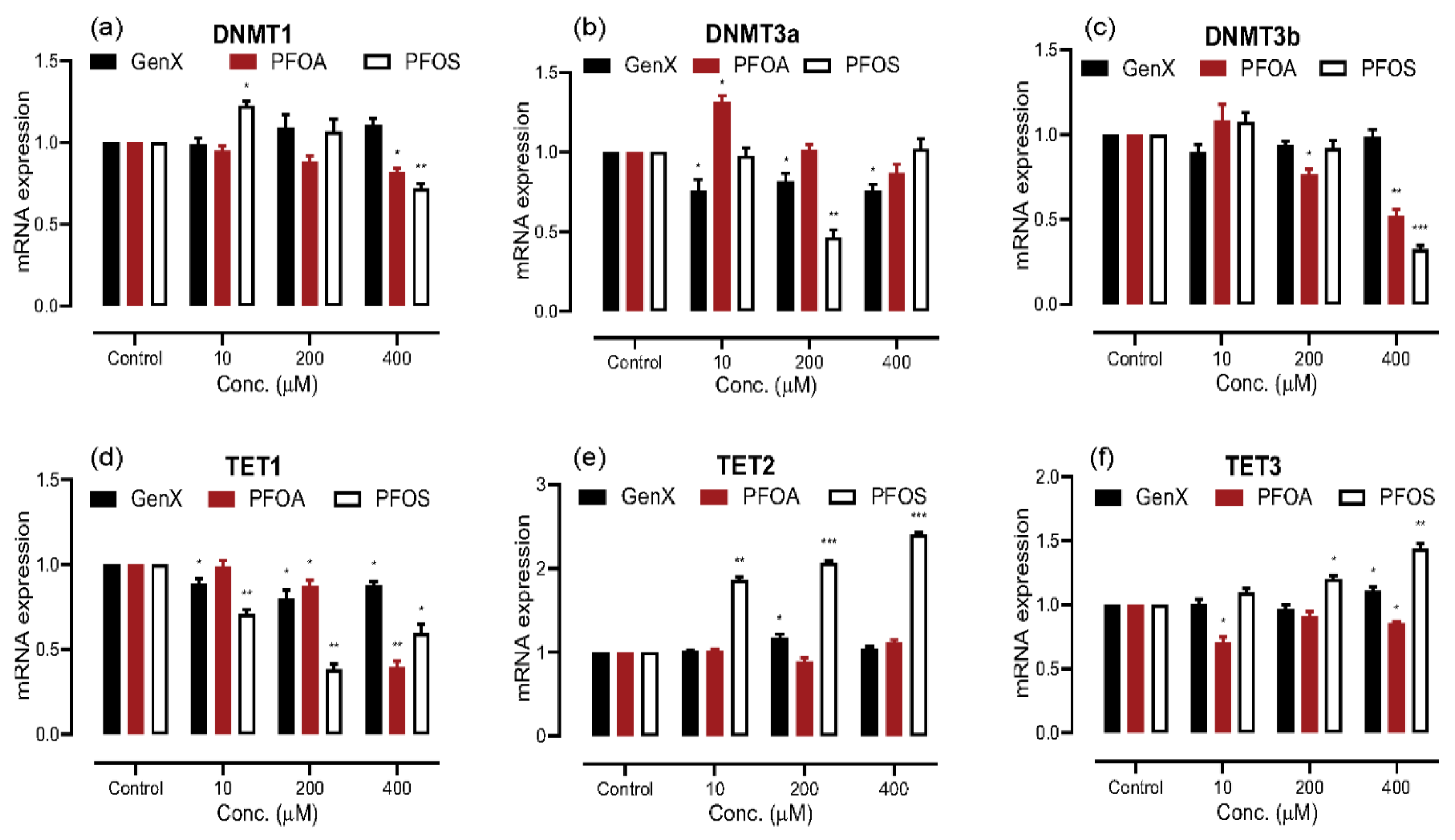
3.2. Exposure to PFAS Alters the mRNA Expression of DNA Methylation Regulators
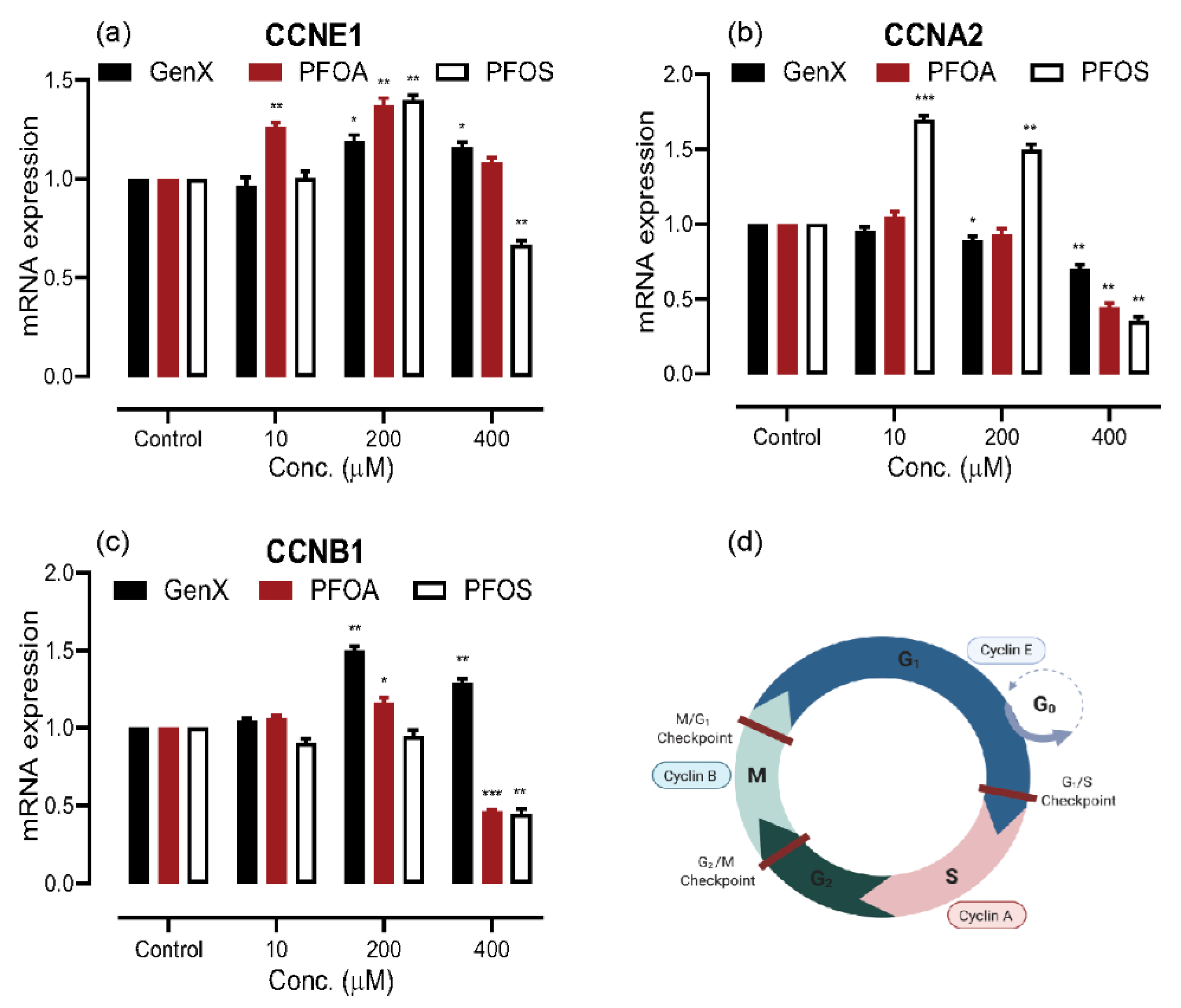
3.3. PFAS Exposure Causes Dysregulation in the Cell Cycle Proliferation Genes
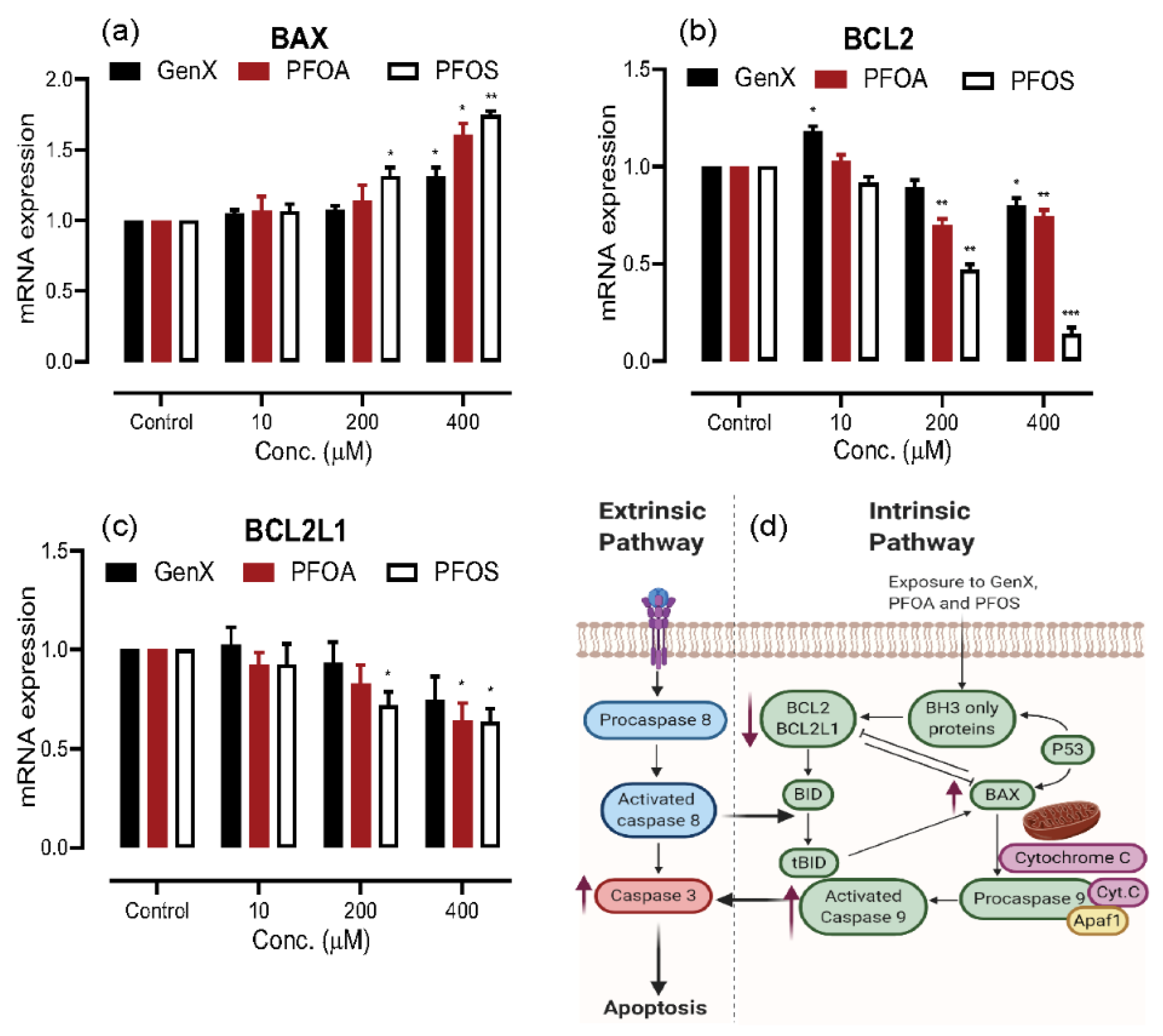
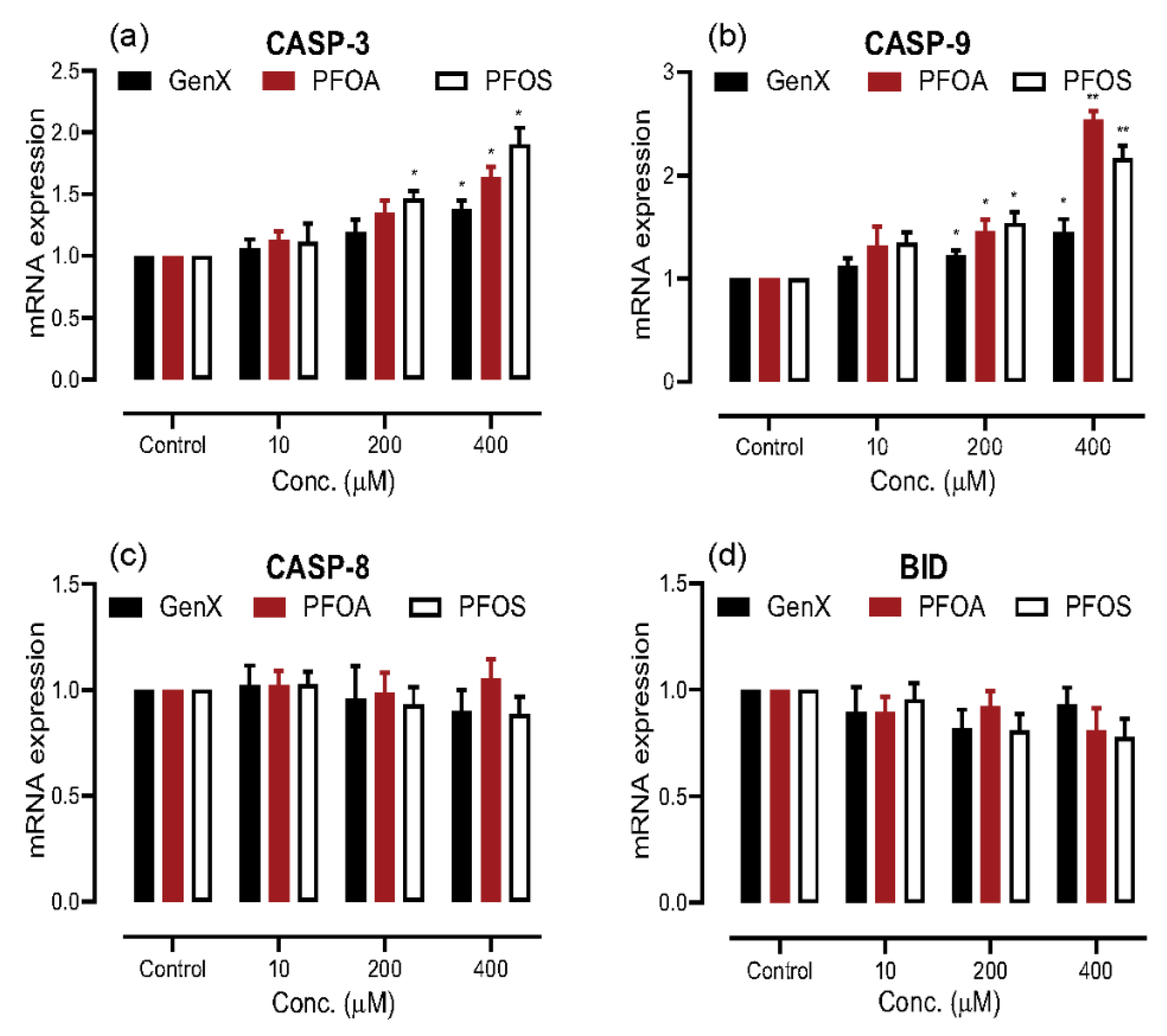
3.4. Induction of Apoptosis on PFAS Exposure
3.5. PFAS Induced Apoptosis through the Intrinsic Pathway
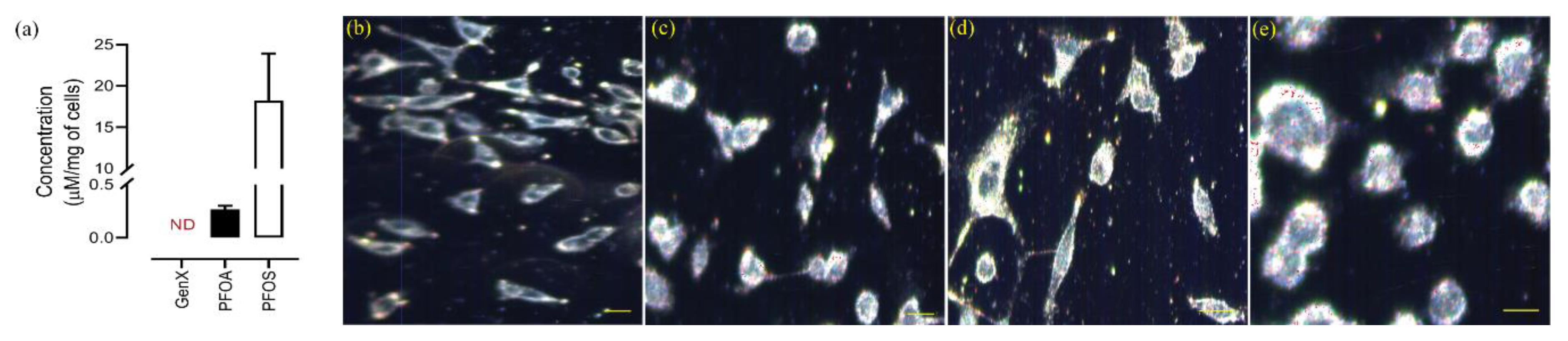
3.6. Intracellular Concentration Analysis
3.7. Cellular Accumulation of PFAS evaluated by Hyperspectral Dark-Field Microscopic (HS-DFM) Imaging
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Kissa, E. Fluorinated Surfactants and Repellents; CRC Press: Boca Raton, FL, USA, 2001; Volume 97. [Google Scholar]
- Schultz, M.M.; Barofsky, D.F.; Field, J.A. Fluorinated Alkyl Surfactants. Environ. Eng. Sci. 2003, 20, 487–501. [Google Scholar] [CrossRef]
- Walters, A.; Santillo, D. Uses of perfluorinated substances. Greenpeace Res. Lab. 2006. [Google Scholar]
- EPA. Perfluorooctanoic Acid (PFOA) and Fluorinated Telomers; U.S. Environmental Protection Agency: Washington, DC, USA, 2008.
- Prevedouros, K.; Cousins, I.T.; Buck, R.C.; Korzeniowski, S.H. Sources, fate and transport of perfluorocarboxylates. Environ. Sci. Technol. 2006, 40, 32–44. [Google Scholar] [PubMed]
- Öberg, T.; Liu, T. Extension of a prediction model to estimate vapor pressures of perfluorinated compounds (PFCs). Chemom. Intell. Lab. Syst. 2011, 107, 59–64. [Google Scholar] [CrossRef]
- Davis, K.L.; Aucoin, M.D.; Larsen, B.S.; Kaiser, M.A.; Hartten, A.S. Transport of ammonium perfluorooctanoate in environmental media near a fluoropolymer manufacturing facility. Chemosphere 2007, 67, 2011–2019. [Google Scholar] [CrossRef]
- ATSDR. Toxicological Profile for Perfluoroalkyls; Draft Public Comment; Agency for Toxic Substances and Disease Registry: Atlanta, Georgia, 2018.
- De Vos, M.G.; Huijbregts, M.A.J.; Heuvel-Greve, M.J.V.D.; Vethaak, A.D.; Van De Vijver, K.I.; Leonards, P.; Van Leeuwen, S.P.; De Voogt, P.; Hendriks, A.J. Accumulation of perfluorooctane sulfonate (PFOS) in the food chain of the Western Scheldt estuary: Comparing field measurements with kinetic modeling. Chemosphere 2008, 70, 1766–1773. [Google Scholar] [CrossRef]
- Furdui, V.I.; Stock, N.L.; Ellis, D.A.; Butt, C.M.; Whittle, D.M.; Crozier, P.W.; Reiner, E.J.; Muir, D.C.G.; Mabury, S.A. Spatial Distribution of Perfluoroalkyl Contaminants in Lake Trout from the Great Lakes. Environ. Sci. Technol. 2007, 41, 1554–1559. [Google Scholar] [CrossRef]
- Martin, J.W.; Whittle, D.M.; Muir, D.C.G.; Mabury, S.A. Perfluoroalkyl Contaminants in a Food Web from Lake Ontario. Environ. Sci. Technol. 2004, 38, 5379–5385. [Google Scholar] [CrossRef]
- Maestri, L.; Negri, S.; Ferrari, M.; Ghittori, S.; Fabris, F.; Danesino, P.; Imbriani, M. Determination of perfluorooctanoic acid and perfluorooctanesulfonate in human tissues by liquid chromatography/single quadrupole mass spectrometry. Rapid Commun. Mass Spectrom. 2006, 20, 2728–2734. [Google Scholar] [CrossRef]
- Garcia, D.S.; Sjödin, M.; Hellstrandh, M.; Norinder, U.; Nikiforova, V.; Lindberg, J.; Wincent, E.; Bergman, Å.; Cotgreave, I.; Kos, V.M. Cellular accumulation and lipid binding of perfluorinated alkylated substances (PFASs)—A comparison with lysosomotropic drugs. Chem. Interact. 2018, 281, 1–10. [Google Scholar] [CrossRef]
- EPA. Long-chain perfluoroalkyl carboxylate and perfluoroalkyl sulfonate chemical substances: Significan new use rule. Fed. Regist. 2015, 80, 2885–2898. [Google Scholar]
- Chen, Y.-M.; Guo, L.-H. Fluorescence study on site-specific binding of perfluoroalkyl acids to human serum albumin. Arch. Toxicol. 2008, 83, 255–261. [Google Scholar] [CrossRef] [PubMed]
- Sheng, N.; Li, J.; Liu, H.; Zhang, A.; Dai, J. Interaction of perfluoroalkyl acids with human liver fatty acid-binding protein. Arch. Toxicol. 2014, 90, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Calafat, A.M.; Wong, L.-Y.; Kuklenyik, Z.; Reidy, J.A.; Needham, L.L. Polyfluoroalkyl Chemicals in the U.S. Population: Data from the National Health and Nutrition Examination Survey (NHANES) 2003–2004 and Comparisons with NHANES 1999–2000. Environ. Health Perspect. 2007, 115, 1596–1602. [Google Scholar] [CrossRef]
- De Silva, A.O.; Mabury, S.A. Isomer Distribution of Perfluorocarboxylates in Human Blood: Potential Correlation to Source. Environ. Sci. Technol. 2006, 40, 2903–2909. [Google Scholar] [CrossRef]
- Kuklenyik, Z.; Reich, J.A.; Tully, J.S.; Needham, L.L.; Calafat, A.M. Automated Solid-Phase Extraction and Measurement of Perfluorinated Organic Acids and Amides in Human Serum and Milk. Environ. Sci. Technol. 2004, 38, 3698–3704. [Google Scholar] [CrossRef]
- Olsen, G.W.; Burris, J.M.; Ehresman, D.J.; Froehlich, J.W.; Seacat, A.M.; Butenhoff, J.L.; Zobel, L.R. Half-Life of Serum Elimination of Perfluorooctanesulfonate, Perfluorohexanesulfonate, and Perfluorooctanoate in Retired Fluorochemical Production Workers. Environ. Health Perspect. 2007, 115, 1298–1305. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, H.; Cui, Q.; Sheng, N.; Yeung, L.W.Y.; Guo, Y.; Sun, Y.; Dai, J. First Report on the Occurrence and Bioaccumulation of Hexafluoropropylene Oxide Trimer Acid: An Emerging Concern. Environ. Sci. Technol. 2017, 51, 9553–9560. [Google Scholar] [CrossRef]
- Morello-Frosch, R.; Cushing, L.J.; Jesdale, B.M.; Schwartz, J.M.; Guo, W.; Guo, T.; Wang, M.; Harwani, S.; Petropoulou, S.-S.E.; Duong, W.; et al. Environmental Chemicals in an Urban Population of Pregnant Women and Their Newborns from San Francisco. Environ. Sci. Technol. 2016, 50, 12464–12472. [Google Scholar] [CrossRef]
- Apelberg, B.J.; Witter, F.R.; Herbstman, J.B.; Calafat, A.M.; Halden, R.U.; Needham, L.L.; Goldman, L.R. Cord Serum Concentrations of Perfluorooctane Sulfonate (PFOS) and Perfluorooctanoate (PFOA) in Relation to Weight and Size at Birth. Environ. Health Perspect. 2007, 115, 1670–1676. [Google Scholar] [CrossRef]
- Fei, C.; McLaughlin, J.K.; Tarone, R.E.; Olsen, J. Perfluorinated Chemicals and Fetal Growth: A Study within the Danish National Birth Cohort. Environ. Health Perspect. 2007, 115, 1677–1682. [Google Scholar] [CrossRef] [PubMed]
- Inoue, K.; Okada, F.; Ito, R.; Kato, S.; Sasaki, S.; Nakajima, S.; Uno, A.; Saijo, Y.; Sata, F.; Yoshimura, Y.; et al. Perfluorooctane Sulfonate (PFOS) and Related Perfluorinated Compounds in Human Maternal and Cord Blood Samples: Assessment of PFOS Exposure in a Susceptible Population during Pregnancy. Environ. Health Perspect. 2004, 112, 1204–1207. [Google Scholar] [CrossRef] [PubMed]
- Midasch, O.; Drexler, H.; Hart, N.; Beckmann, M.W.; Angerer, J. Transplacental exposure of neonates to perfluorooctanesulfonate and perfluorooctanoate: A pilot study. Int. Arch. Occup. Environ. Health 2007, 80, 643–648. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Kannan, K.; Wong, C.M.; Arcaro, K.F.; Butenhoff, J.L. Perfluorinated Compounds in Human Milk from Massachusetts, U.S.A. Environ. Sci. Technol. 2008, 42, 3096–3101. [Google Scholar] [CrossRef]
- Kärrman, A.; Ericson, I.; Van Bavel, B.; Darnerud, P.O.; Aune, M.; Glynn, A.; Lignell, S.; Lindström, G. Exposure of Perfluorinated Chemicals through Lactation: Levels of Matched Human Milk and Serum and a Temporal Trend, 1996–2004, in Sweden. Environ. Health Perspect. 2007, 115, 226–230. [Google Scholar] [CrossRef]
- Kato, K.; Calafat, A.M.; Wong, L.-Y.; Wanigatunga, A.A.; Caudill, S.P.; Needham, L.L. Polyfluoroalkyl Compounds in Pooled Sera from Children Participating in the National Health and Nutrition Examination Survey 2001−2002. Environ. Sci. Technol. 2009, 43, 2641–2647. [Google Scholar] [CrossRef]
- Trudel, D.; Horowitz, L.; Wormuth, M.; Scheringer, M.; Cousins, I.T.; Hungerbühler, K. Estimating Consumer Exposure to PFOS and PFOA. Risk Anal. 2008, 28, 251–269. [Google Scholar] [CrossRef]
- Seacat, A.M.; Thomford, P.J.; Hansen, K.J.; Olsen, G.W.; Case, M.T.; Butenhoff, J.L. Subchronic Toxicity Studies on Perfluorooctanesulfonate Potassium Salt in Cynomolgus Monkeys. Toxicol. Sci. 2002, 68, 249–264. [Google Scholar] [CrossRef]
- Butenhoff, J.L.; Kennedy, G.L.; Hinderliter, P.M.; Lieder, P.H.; Noker, P.E.; Thomford, P.J.; Jung, R.; Hansen, K.J.; Gorman, G.S. Pharmacokinetics of Perfluorooctanoate in Cynomolgus Monkeys. Toxicol. Sci. 2004, 82, 394–406. [Google Scholar] [CrossRef]
- Chang, S.-C.; Noker, P.E.; Gorman, G.S.; Gibson, S.J.; Hart, J.A.; Ehresman, D.J.; Butenhoff, J.L. Comparative pharmacokinetics of perfluorooctanesulfonate (PFOS) in rats, mice, and monkeys. Reprod. Toxicol. 2012, 33, 428–440. [Google Scholar] [CrossRef]
- Kim, S.-J.; Heo, S.-H.; Lee, N.-S.; Hwang, I.G.; Lee, Y.-B.; Cho, H.-Y. Gender differences in pharmacokinetics and tissue distribution of 3 perfluoroalkyl and polyfluoroalkyl substances in rats. Food Chem. Toxicol. 2016, 97, 243–255. [Google Scholar] [CrossRef] [PubMed]
- Fujii, Y.; Niisoe, T.; Harada, K.H.; Uemoto, S.; Ogura, Y.; Takenaka, K.; Koizumi, A. Toxicokinetics of perfluoroalkyl carboxylic acids with different carbon chain lengths in mice and humans. J. Occup. Health 2015, 57, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Gannon, S.A.; Fasano, W.J.; Mawn, M.P.; Nabb, D.L.; Buck, R.C.; Buxton, L.W.; Jepson, G.W.; Frame, S.R. Absorption, distribution, metabolism, excretion, and kinetics of 2,3,3,3-tetrafluoro-2-(heptafluoropropoxy)propanoic acid ammonium salt following a single dose in rat, mouse, and cynomolgus monkey. Toxicology 2016, 340, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Conley, J.M.; Lambright, C.S.; Evans, N.; Strynar, M.J.; Mccord, J.; McIntyre, B.S.; Travlos, G.S.; Cardon, M.C.; Medlock-Kakaley, E.; Hartig, P.C.; et al. Adverse Maternal, Fetal, and Postnatal Effects of Hexafluoropropylene Oxide Dimer Acid (GenX) from Oral Gestational Exposure in Sprague-Dawley Rats. Environ. Health Perspect. 2019, 127, 037008. [Google Scholar] [CrossRef]
- Lau, C.; Butenhoff, J.L.; Rogers, J.M. The developmental toxicity of perfluoroalkyl acids and their derivatives. Toxicol. Appl. Pharmacol. 2004, 198, 231–241. [Google Scholar] [CrossRef]
- Yahia, D.; El-Nasser, M.A.; Abedel-Latif, M.; Tsukuba, C.; Yoshida, M.; Sato, I.; Tsuda, S. Effects of perfluorooctanoic acid (PFOA) exposure to pregnant mice on reproduction. J. Toxicol. Sci. 2010, 35, 527–533. [Google Scholar] [CrossRef][Green Version]
- Bijland, S.; Rensen, P.C.; Pieterman, E.J.; Maas, A.C.; Van Der Hoorn, J.W.; Van Erk, M.J.; Havekes, L.M.; Van Dijk, K.W.; Chang, S.-C.; Ehresman, D.J.; et al. Perfluoroalkyl Sulfonates Cause Alkyl Chain Length–Dependent Hepatic Steatosis and Hypolipidemia Mainly by Impairing Lipoprotein Production in APOE*3-Leiden CETP Mice. Toxicol. Sci. 2011, 123, 290–303. [Google Scholar] [CrossRef]
- Kennedy, G.L.; Butenhoff, J.L.; Olsen, G.W.; O’Connor, J.C.; Seacat, A.M.; Perkins, R.G.; Biegel, L.B.; Murphy, S.R.; Farrar, D.G. The Toxicology of Perfluorooctanoate. Crit. Rev. Toxicol. 2004, 34, 351–384. [Google Scholar] [CrossRef]
- Wang, L.; Wang, Y.; Liang, Y.; Li, J.; Liu, Y.; Zhang, J.; Zhang, A.; Fu, J.; Jiang, G. Specific Accumulation of Lipid Droplets in Hepatocyte Nuclei of PFOA-exposed BALB/c Mice. Sci. Rep. 2013, 3, srep02174. [Google Scholar] [CrossRef]
- Xing, J.; Wang, G.; Zhao, J.; Wang, E.; Yin, B.; Fang, D.; Zhao, J.; Zhang, H.; Chen, Y.Q.; Chen, W. Toxicity assessment of perfluorooctane sulfonate using acute and subchronic male C57BL/6J mouse models. Environ. Pollut. 2016, 210, 388–396. [Google Scholar] [CrossRef]
- Weiss, J.M.; Andersson, P.L.; Lamoree, M.H.; Leonards, P.E.G.; Van Leeuwen, S.P.J.; Hamers, T. Competitive Binding of Poly- and Perfluorinated Compounds to the Thyroid Hormone Transport Protein Transthyretin. Toxicol. Sci. 2009, 109, 206–216. [Google Scholar] [CrossRef] [PubMed]
- Corsini, E.; Luebke, R.W.; Germolec, D.R.; DeWitt, J.C. Perfluorinated compounds: Emerging POPs with potential immunotoxicity. Toxicol. Lett. 2014, 230, 263–270. [Google Scholar] [CrossRef] [PubMed]
- DeWitt, J.C.; Peden-Adams, M.M.; Keller, J.M.; Germolec, D.R. Immunotoxicity of Perfluorinated Compounds: Recent Developments. Toxicol. Pathol. 2012, 40, 300–311. [Google Scholar] [CrossRef] [PubMed]
- Guruge, K.S.; Hikono, H.; Shimada, N.; Murakami, K.; Hasegawa, J.; Yeung, L.W.; Yamanaka, N.; Yamashita, N. Effect of perfluorooctane sulfonate (PFOS) on influenza A virus-induced mortality in female B6C3F1 mice. J. Toxicol. Sci. 2009, 34, 687–691. [Google Scholar] [CrossRef]
- Rushing, B.R.; Hu, Q.; Franklin, J.N.; McMahen, R.L.; Dagnino, S.; Higgins, C.P.; Strynar, M.J.; DeWitt, J.C. Evaluation of the immunomodulatory effects of 2,3,3,3-tetrafluoro-2-(heptafluoropropoxy)-propanoate in C57BL/6 mice. Toxicol. Sci. 2017, 156, 179–189. [Google Scholar] [CrossRef]
- Rae, J.C.; Craig, L.; Slone, T.W.; Frame, S.R.; Buxton, L.; Kennedy, G.L. Evaluation of chronic toxicity and carcinogenicity of ammonium 2,3,3,3-tetrafluoro-2-(heptafluoropropoxy)-propanoate in Sprague–Dawley rats. Toxicol. Rep. 2015, 2, 939–949. [Google Scholar] [CrossRef]
- Barry, V.; Winquist, A.; Steenland, K. Perfluorooctanoic Acid (PFOA) Exposures and Incident Cancers among Adults Living Near a Chemical Plant. Environ. Health Perspect. 2013, 121, 1313–1318. [Google Scholar] [CrossRef]
- Raleigh, K.K.; Alexander, B.H.; Olsen, G.W.; Ramachandran, G.; Morey, S.Z.; Church, T.R.; Logan, P.W.; Scott, L.L.F.; Allen, E.M. Mortality and cancer incidence in ammonium perfluorooctanoate production workers. Occup. Environ. Med. 2014, 71, 500–506. [Google Scholar] [CrossRef]
- Steenland, K.; Zhao, L.; Winquist, A. A cohort incidence study of workers exposed to perfluorooctanoic acid (PFOA). Occup. Environ. Med. 2015, 72, 373–380. [Google Scholar] [CrossRef]
- Wen, Y.; Mirji, N.; Irudayaraj, J. Epigenetic toxicity of PFOA and GenX in HepG2 cells and their role in lipid metabolism. Toxicol. In Vitro 2020, 65, 104797. [Google Scholar] [CrossRef]
- Rashid, F.; Ramakrishnan, A.; Fields, C.; Irudayaraj, J. Acute PFOA exposure promotes epigenomic alterations in mouse kidney tissues. Toxicol. Rep. 2020, 7, 125–132. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Chen, J.; Li, J.; Arif, W.; Kalsotra, A.; Irudayaraj, J. Effect of PFOA on DNA Methylation and Alternative Splicing in Mouse Liver. Toxicol. Lett. 2020, 329, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Irudayaraj, J. Perfluorooctanoic acid (PFOA) exposure inhibits DNA methyltransferase activities and alters constitutive heterochromatin organization. Food Chem. Toxicol. 2020, 141, 111358. [Google Scholar] [CrossRef] [PubMed]
- Holliday, R.; Pugh, J. DNA modification mechanisms and gene activity during development. Science 1975, 187, 226–232. [Google Scholar] [CrossRef] [PubMed]
- Smith, Z.D.; Meissner, A. DNA methylation: Roles in mammalian development. Nat. Rev. Genet. 2013, 14, 204–220. [Google Scholar] [CrossRef]
- Okano, M.; Bell, D.W.; Haber, D.A.; Li, E. DNA Methyltransferases Dnmt3a and Dnmt3b Are Essential for De Novo Methylation and Mammalian Development. Cell 1999, 99, 247–257. [Google Scholar] [CrossRef]
- Bergman, Y.; Cedar, H. DNA methylation dynamics in health and disease. Nat. Struct. Mol. Biol. 2013, 20, 274–281. [Google Scholar] [CrossRef]
- Xu, G.-L.; Bestor, T.H.; Bourc’His, D.; Hsieh, C.-L.; Tommerup, N.; Bugge, M.; Hulten, M.; Qu, X.; Russo, J.J.; Viegas-Péquignot, E. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nat. Cell Biol. 1999, 402, 187–191. [Google Scholar] [CrossRef]
- Portela, A.; Esteller, M. Epigenetic modifications and human disease. Nat. Biotechnol. 2010, 28, 1057–1068. [Google Scholar] [CrossRef]
- Du, J.; Johnson, L.M.; Jacobsen, S.E.; Patel, D.J. DNA methylation pathways and their crosstalk with histone methylation. Nat. Rev. Mol. Cell Biol. 2015, 16, 519–532. [Google Scholar] [CrossRef]
- Robertson, K.D.; Uzvolgyi, E.; Liang, G.; Talmadge, C.; Sumegi, J.; Gonzales, F.A.; Jones, P.A. The human DNA methyltransferases (DNMTs) 1, 3a and 3b: Coordinate mRNA expression in normal tissues and overexpression in tumors. Nucleic Acids Res. 1999, 27, 2291–2298. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, K.D.; Helin, K. Role of TET enzymes in DNA methylation, development, and cancer. Genes Dev. 2016, 30, 733–750. [Google Scholar] [CrossRef] [PubMed]
- Lyko, F. The DNA methyltransferase family: A versatile toolkit for epigenetic regulation. Nat. Rev. Genet. 2018, 19, 81–92. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.C.; Cullen, S.P.; Martin, S.J. Apoptosis: Controlled demolition at the cellular level. Nat. Rev. Mol. Cell Biol. 2008, 9, 231–241. [Google Scholar] [CrossRef]
- Mao, Z.; Xia, W.; Wang, J.; Chen, T.; Zeng, Q.; Xu, B.; Li, W.; Chen, X.; Xu, S. Perfluorooctane sulfonate induces apoptosis in lung cancer A549 cells through reactive oxygen species-mediated mitochondrion-dependent pathway. J. Appl. Toxicol. 2012, 33, 1268–1276. [Google Scholar] [CrossRef]
- Liu, C.; Yu, K.; Shi, X.; Wang, J.; Lam, P.K.; Wu, R.S.; Zhou, B. Induction of oxidative stress and apoptosis by PFOS and PFOA in primary cultured hepatocytes of freshwater tilapia (Oreochromis niloticus). Aquat. Toxicol. 2007, 82, 135–143. [Google Scholar] [CrossRef]
- Evan, G.I.; Vousden, K.H. Proliferation, cell cycle and apoptosis in cancer. Nat. Cell Biol. 2001, 411, 342–348. [Google Scholar] [CrossRef]
- Stepanic, V.; Kostrun, S.; Malnar, I.; Hlevnjak, M.; Butkovic, K.; Caleta, I.; Dukši, M.; Kragol, G.; Makaruha-Stegic, O.; Mikac, L. Modeling cellular pharmacokinetics of 14-and 15-membered macrolides with physicochemical properties. J. Med. Chem. 2011, 54, 719–733. [Google Scholar] [CrossRef]
- Wang, X.; Cui, Y.; Irudayaraj, J. Single-Cell Quantification of Cytosine Modifications by Hyperspectral Dark-Field Imaging. ACS Nano 2015, 9, 11924–11932. [Google Scholar] [CrossRef]
- Cui, Y.; Wang, X.; Ren, W.; Liu, J.; Irudayaraj, J. Optical Clearing Delivers Ultrasensitive Hyperspectral Dark-Field Imaging for Single-Cell Evaluation. ACS Nano 2016, 10, 3132–3143. [Google Scholar] [CrossRef]
- Trosko, J.E.; Chang, C.-C.; Upham, B.; Wilson, M. Epigenetic toxicology as toxicant-induced changes in intracellular signalling leading to altered gap junctional intercellular communication. Toxicol. Lett. 1998, 102, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Reamon-Buettner, S.M.; Borlak, J. A new paradigm in toxicology and teratology: Altering gene activity in the absence of DNA sequence variation. Reprod. Toxicol. 2007, 24, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Szyf, M. The Implications of DNA Methylation for Toxicology: Toward Toxicomethylomics, the Toxicology of DNA Methylation. Toxicol. Sci. 2011, 120, 235–255. [Google Scholar] [CrossRef] [PubMed]
- Rashid, F.; Ahmad, S.; Irudayaraj, J.M.K. Effect of Perfluorooctanoic Acid on the Epigenetic and Tight Junction Genes of the Mouse Intestine. Toxics 2020, 8, 64. [Google Scholar] [CrossRef]
- Cohen, S.M.; Ellwein, L.B. Cell proliferation in carcinogenesis. Science 1990, 249, 1007–1011. [Google Scholar] [CrossRef]
- Riedl, S.J.; Shi, Y. Molecular mechanisms of caspase regulation during apoptosis. Nat. Rev. Mol. Cell Biol. 2004, 5, 897–907. [Google Scholar] [CrossRef]
- Cui, Y.; Liu, W.; Xie, W.; Yu, W.; Wang, C.; Chen, H. Investigation of the Effects of Perfluorooctanoic Acid (PFOA) and Perfluorooctane Sulfonate (PFOS) on Apoptosis and Cell Cycle in a Zebrafish (Danio rerio) Liver Cell Line. Int. J. Environ. Res. Public Health 2015, 12, 15673–15682. [Google Scholar] [CrossRef]
- Takata, T.; Tanaka, F.; Yamada, T.; Yanagihara, K.; Otake, Y.; Kawano, Y.; Nakagawa, T.; Miyahara, R.; Oyanagi, H.; Inui, K.; et al. Clinical significance of caspase-3 expression in pathologic-stage I, nonsmall-cell lung cancer. Int. J. Cancer 2001, 96, 54–60. [Google Scholar] [CrossRef]
- Roth, M.; Obaidat, A.; Hagenbuch, B. OATPs, OATs and OCTs: The organic anion and cation transporters of the SLCO and SLC22A gene superfamilies. Br. J. Pharmacol. 2012, 165, 1260–1287. [Google Scholar] [CrossRef]
- Aubert, L.; Motais, R. Molecular features of organic anion permeablity in ox red blood cell. J. Physiol. 1975, 246, 159–179. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jabeen, M.; Fayyaz, M.; Irudayaraj, J. Epigenetic Modifications, and Alterations in Cell Cycle and Apoptosis Pathway in A549 Lung Carcinoma Cell Line upon Exposure to Perfluoroalkyl Substances. Toxics 2020, 8, 112. https://doi.org/10.3390/toxics8040112
Jabeen M, Fayyaz M, Irudayaraj J. Epigenetic Modifications, and Alterations in Cell Cycle and Apoptosis Pathway in A549 Lung Carcinoma Cell Line upon Exposure to Perfluoroalkyl Substances. Toxics. 2020; 8(4):112. https://doi.org/10.3390/toxics8040112
Chicago/Turabian StyleJabeen, Musarrat, Muhammad Fayyaz, and Joseph Irudayaraj. 2020. "Epigenetic Modifications, and Alterations in Cell Cycle and Apoptosis Pathway in A549 Lung Carcinoma Cell Line upon Exposure to Perfluoroalkyl Substances" Toxics 8, no. 4: 112. https://doi.org/10.3390/toxics8040112
APA StyleJabeen, M., Fayyaz, M., & Irudayaraj, J. (2020). Epigenetic Modifications, and Alterations in Cell Cycle and Apoptosis Pathway in A549 Lung Carcinoma Cell Line upon Exposure to Perfluoroalkyl Substances. Toxics, 8(4), 112. https://doi.org/10.3390/toxics8040112






