Metabolism of Lactobacillus sakei Chr82 in the Presence of Different Amounts of Fermentable Sugars
Abstract
1. Introduction
2. Materials and Methods
2.1. Microorganism Used
2.2. Growth Media
2.3. Growth Modeling and pH Measurement
2.4. Microbiological Analysis
2.5. Organic Acids Content
2.6. Quantification of Amino Acids
2.7. Flow Cytometric Analysis
2.8. Untargeted Metabolomics Analysis by 1H-NMR
3. Results and Discussion
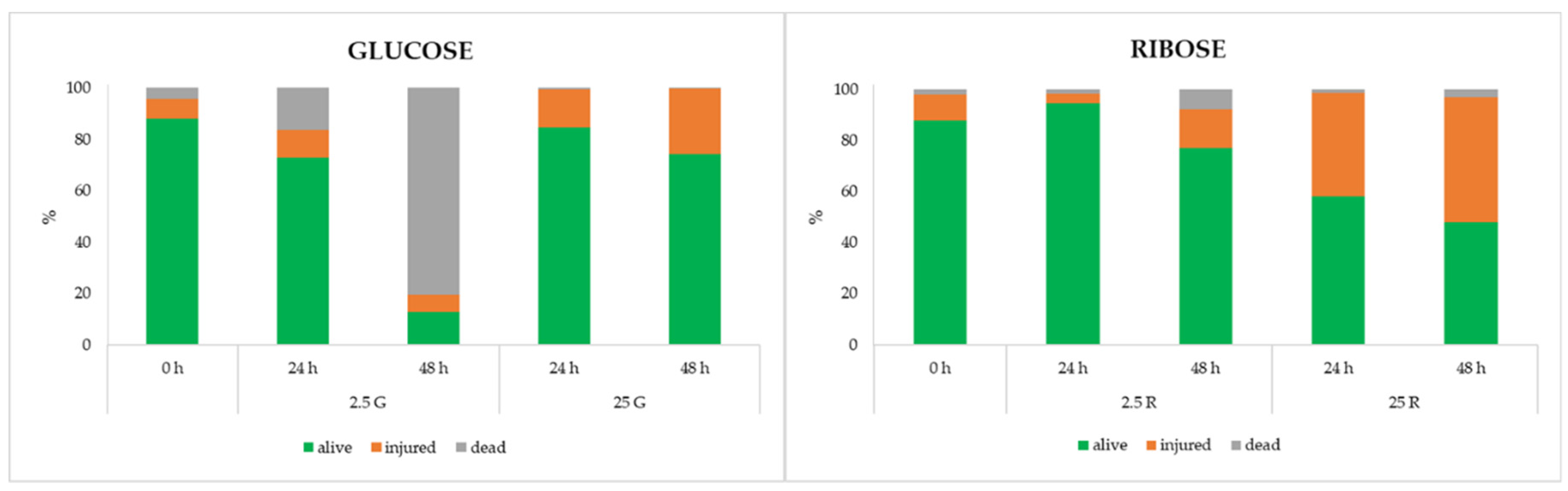
3.1. Determination of Growth Curves and Microbiological Analysis
3.2. Organic Acid Content and pH
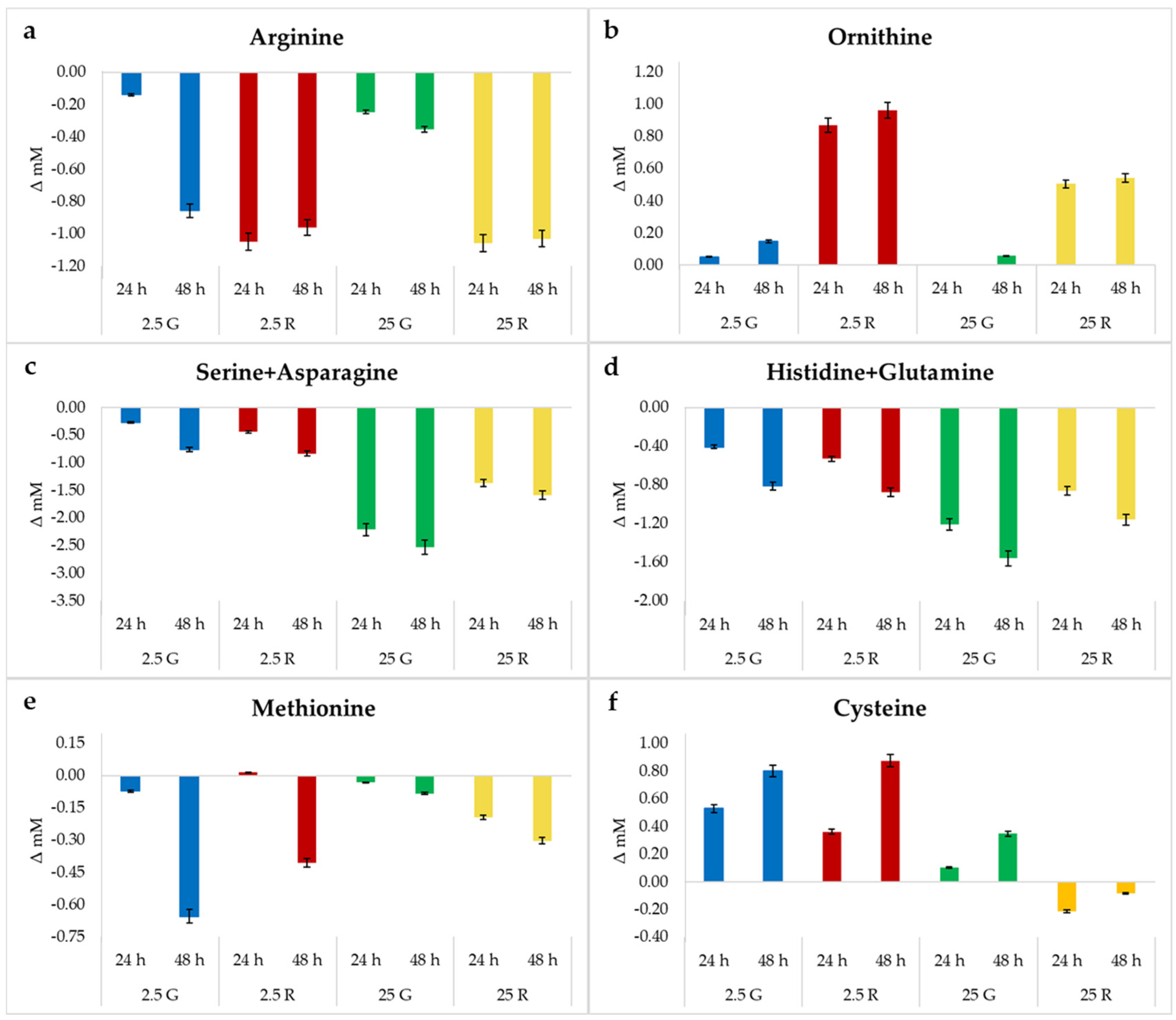
3.3. Amino Acids Quantification
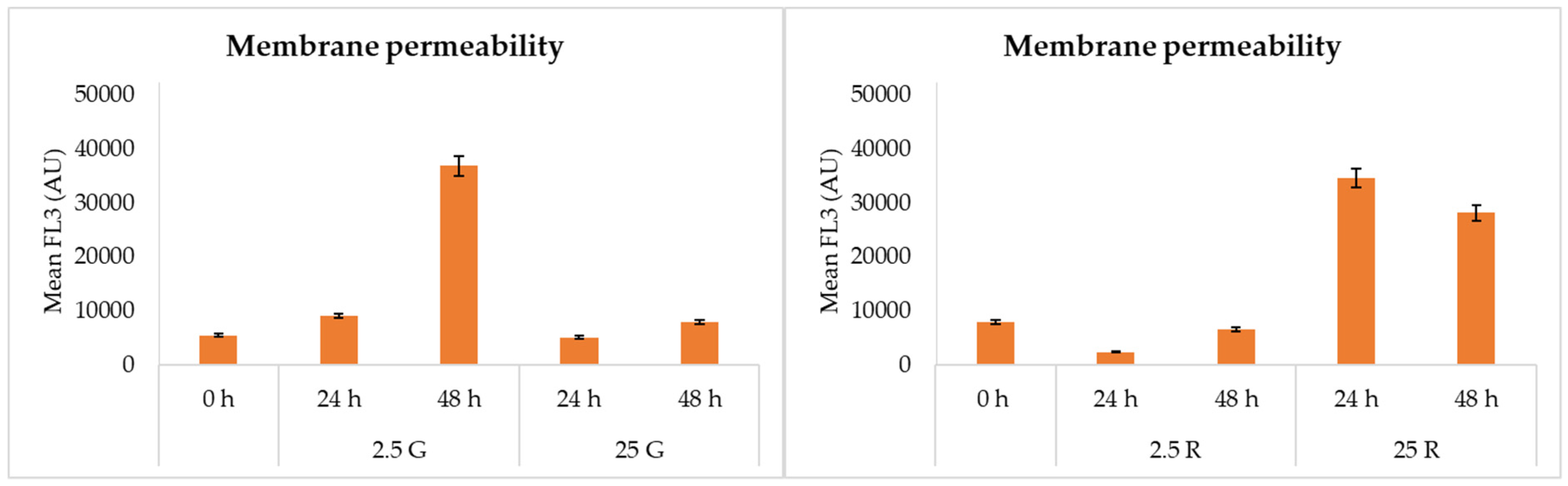
3.4. Flow Cytometric Analysis
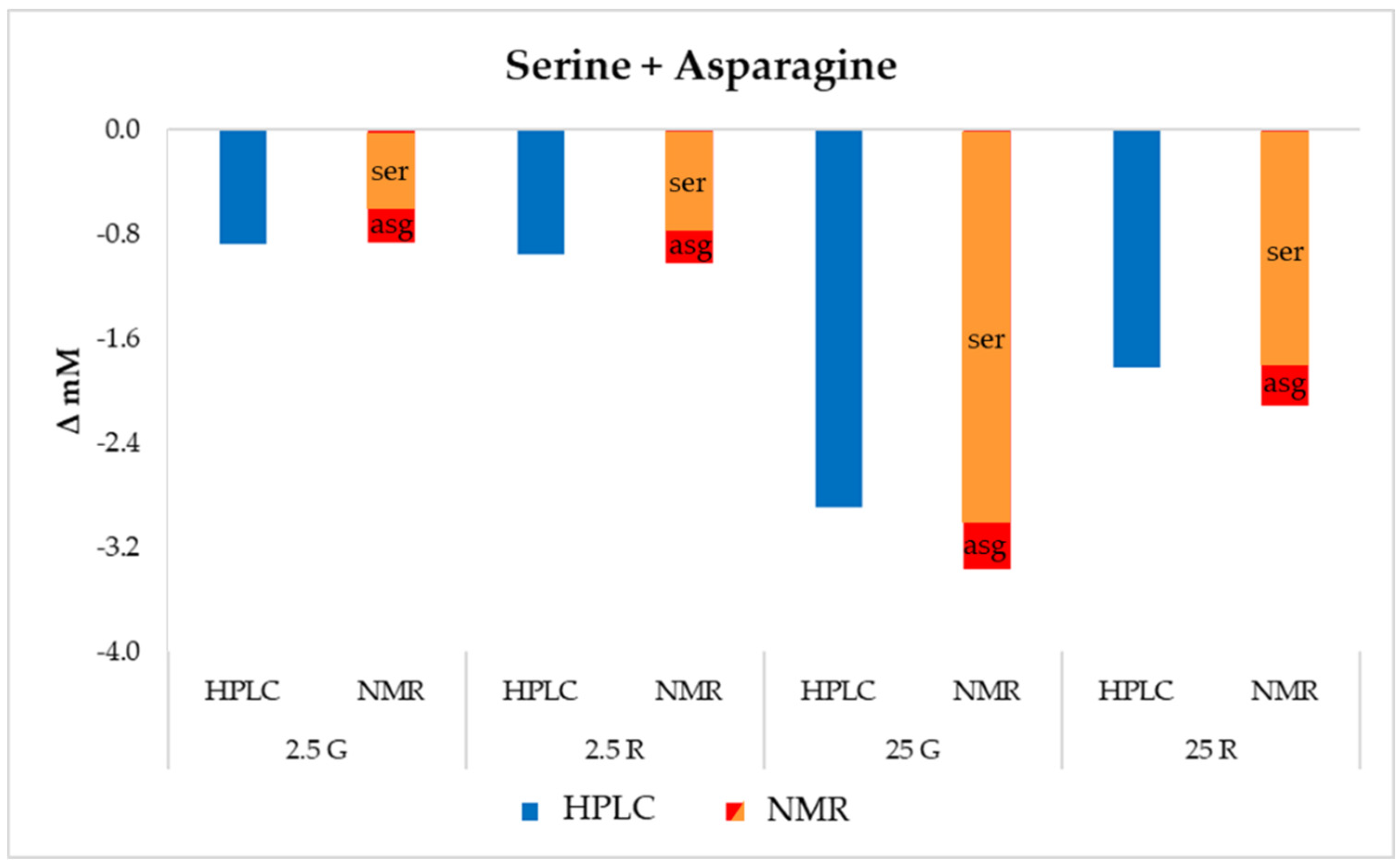
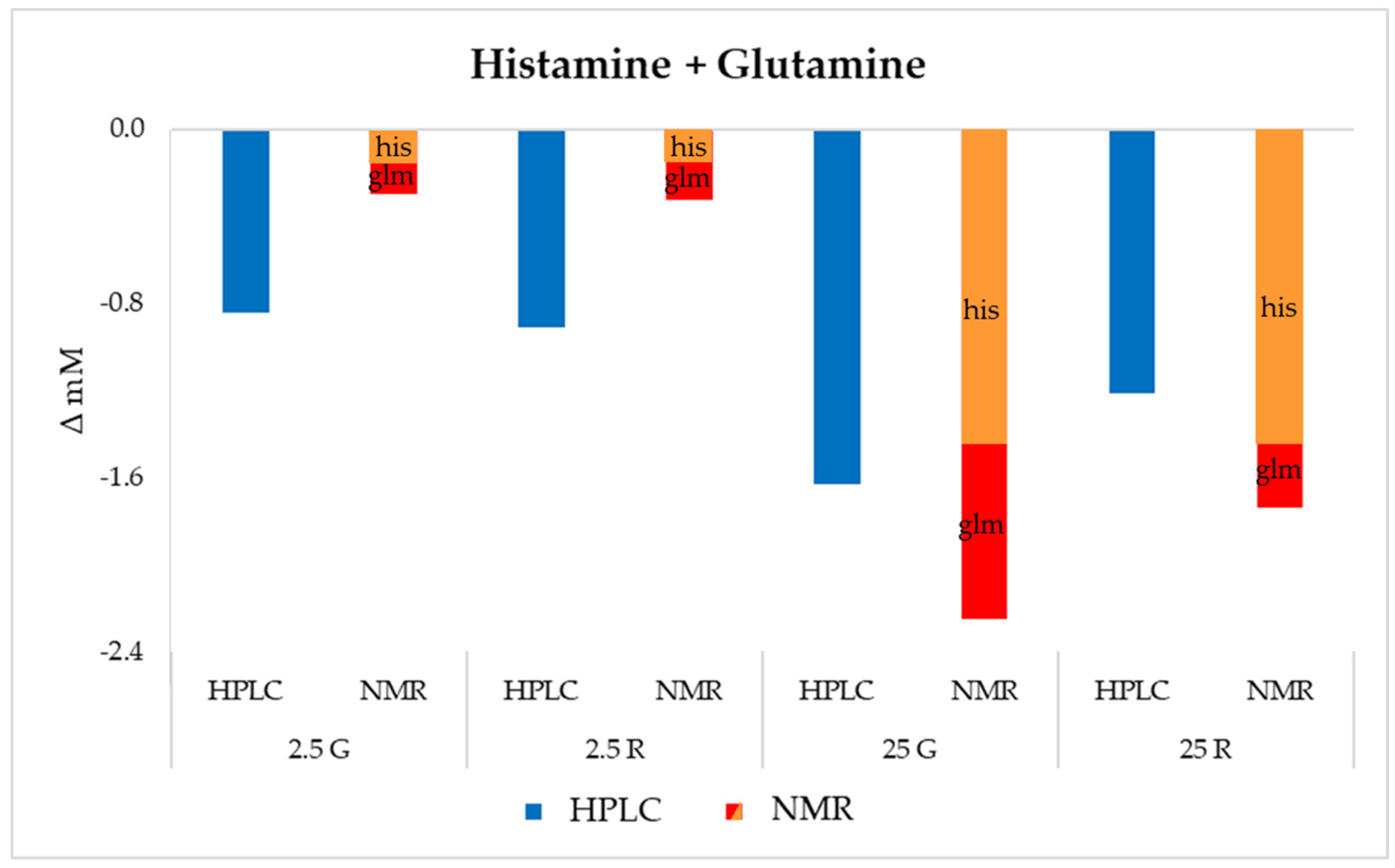
3.5. Untargeted Metabolomics Analysis by 1H-NMR
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Cocconcelli, P.S.; Fontana, C. Starter cultures for meat fermentation. In Handbook of Meat Processing; Toldrá, F., Ed.; Blackwell Publishing: Ames, IA, USA, 2010; pp. 199–218. ISBN 9780813820897. [Google Scholar]
- Chaillou, S.; Lucquin, I.; Najjari, A.; Zagorec, M.; Champomier-Vergès, M.C. Population genetics of Lactobacillus sakei reveals three lineages with distinct evolutionary histories. PLoS ONE 2013, 8, e73253. [Google Scholar] [CrossRef]
- McLeod, A.; Nyquistb, O.N.; Snipenb, L.; Naterstada, K.; Axelsson, L. Diversity of Lactobacillus sakei strains investigated by phenotypic and genotypic methods. Syst. Appl. Microbiol. 2008, 31, 393–403. [Google Scholar] [CrossRef]
- Rimaux, T.; Vrancken, G.; Vuylsteke, B.; De Vuyst, L.; Leroy, F. The pentose moiety of adenosine and inosine is an important energy source for the fermented-meat starter culture Lactobacillus sakei CTC 494. Appl. Environ. Microbiol. 2011, 77, 6539–6550. [Google Scholar] [CrossRef]
- McLeod, A.; Mosleth, E.F.; Rud, I.; Branco dos Santos, F.; Snipen, L.; Liland, K.H.; Axelsson, L. Effects of glucose availability in Lactobacillus sakei; metabolic change and regulation of the proteome and transcriptome. PLoS ONE 2017, 12, e0187542. [Google Scholar] [CrossRef] [PubMed]
- Montanari, C.; Barbieri, F.; Magnani, M.; Grazia, L.; Gardini, F.; Tabanelli, G. Phenotypic diversity of Lactobacillus sakei strains. Front. Microbiol. 2018, 9, 2003. [Google Scholar] [CrossRef] [PubMed]
- Chaillou, S.; Daty, M.; Baraige, F.; Dudez, A.M.; Anglade, P.; Jones, R.; Alpert, C.A.; Champomier-Vergès, M.C.; Zagorec, M. Intra-species genomic diversity and natural population structure of the meat borne lactic acid bacterium Lactobacillus sakei. Appl. Environ. Microbiol. 2009, 75, 970–980. [Google Scholar] [CrossRef]
- Chaillou, S.; Champomier-Vergès, M.C.; Cornet, M.; Crutz-Le Coq, A.M.; Dudez, A.M.; Martin, V.; Beaufi, S.; Darbon-Rongère, E.; Bossy, R.; Loux, R.; et al. The complete genome sequence of the meat-borne lactic acid bacterium Lactobacillus sakei 23K. Nat. Biotechnol. 2005, 23, 1527–1533. [Google Scholar] [CrossRef]
- McLeod, A.; Zagorec, M.; Champomier-Vergès, M.C.; Naterstad, K.; Axelsson, L. Primary metabolism in Lactobacillus sakei food isolates by proteomic analysis. BMC Microbiol. 2010, 10, 120. [Google Scholar] [CrossRef]
- McLeod, A.; Snipen, L.; Naterstad, K.; Axelsson, L. Global transcriptome response in Lactobacillus sakei during growth on ribose. BMC Microbiol. 2011, 11, 145. [Google Scholar] [CrossRef]
- Fiehn, O. Metabolomics – the link between genotypes and phenotypes. Plant Mol. Biol. 2002, 48, 155–171. [Google Scholar] [CrossRef]
- Laghi, L.; Versari, A.; Marcolini, E.; Parpinello, G.P. Metabonomic investigation by 1H-NMR to discriminate between red wines from organic and biodynamic grapes. Food Nutr. Sci. 2014, 5, 52–59. [Google Scholar] [CrossRef]
- Lauret, R.; Morel-Deville, F.; Berthier, F.; Champomier-Vergès, M.C.; Postma, P.; Ehrlich, S.D.; Zagorec, M. Carbohydrate utilization in Lactobacillus sake. Appl. Environ. Microbiol. 1996, 62, 1922–1927. [Google Scholar] [CrossRef]
- Zwietering, M.H.; Jongenburger, I.; Rombouts, F.M.; van’t Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 1990, 56, 1875–1881. [Google Scholar] [CrossRef]
- Arioli, S.; Montanari, C.; Magnani, M.; Tabanelli, G.; Patrignani, F.; Lanciotti, R.; Mora, D.; Gardini, F. Modelling of Listeria monocytogenes Scott A after a mild heat treatment in the presence of thymol and carvacrol: Effects on culturability and viability. J. Food Eng. 2019, 240, 73–82. [Google Scholar] [CrossRef]
- Tabanelli, G.; Montanari, C.; Arioli, S.; Magnani, M.; Patrignani, F.; Lanciotti, R.; Mora, D.; Gardini, F. Physiological response of Saccharomyces cerevisiae to citral combined with thermal treatment. LWT Food Sci. Technol. 2019, 101, 827–834. [Google Scholar] [CrossRef]
- Zhu, C.; Li, C.; Wang, Y.; Laghi, L. Characterization of yak common biofluids metabolome by means of proton nuclear magnetic resonance spectroscopy. Metabolites 2019, 9, 41. [Google Scholar] [CrossRef]
- Fadda, S.; Anglade, P.; Baraige, F.; Zagorec, M.; Talon, R.; Vignolo, G.; Champomier-Vergès, M.C. Adaptive response of Lactobacillus sakei 23K during growth in the presence of meat extracts: A proteomic approach. Int. J. Food Microbiol. 2010, 142, 36–43. [Google Scholar] [CrossRef]
- Axelsson, L. Lactic acid bacteria: Classification and physiology. In Lactic Acid Bacteria: Microbiology and Functional Aspects, 3rd ed.; Salminen, S., von Wright, A., Ouwehand, A., Eds.; Marcel Dekker: New York, NY, USA, 2004; pp. 1–66. ISBN 0-8247-5332-1. [Google Scholar]
- Rimaux, T.; Riviére, A.; Illeghems, K.; Weckx, S.; De Vuyst, L.; Leroy, F. Expression of the arginine deiminase pathway genes in Lactobacillus sakei is strain-dependent and is affected by environmental pH. Appl. Environ. Microbiol. 2012, 78, 4874–4883. [Google Scholar] [CrossRef]
- Gänzle, M.G. Lactic metabolism revisited: Metabolism of lactic acid bacteria in food fermentations and food spoilage. Curr. Opin. Food Sci. 2015, 2, 106–117. [Google Scholar] [CrossRef]
- Irmler, S.; Bavan, T.; Oberli, A.; Roetschi, A.; Badertscher, R.; Guggenbühl, B.; Berthoud, H. Catabolism of serine by Pediococcus acidilactici and Pediococcus pentosaceus. Appl. Environ. Microbiol. 2013, 79, 1309–1315. [Google Scholar] [CrossRef][Green Version]
- Skeie, S.; Kieronczyk, A.; Næss, R.M.; Østlie, H. Lactobacillus adjuncts in cheese: Their influence on the degradation of citrate and serine during ripening of a washed curd cheese. Int. Dairy J. 2008, 18, 158–168. [Google Scholar] [CrossRef]
- Biagioli, M.; Laghi, L.; Carino, A.; Cipriani, S.; Distrutti, E.; Marchianò, S.; Parolin, C.; Scarpelli, P.; Vitali, B.; Fiorucci, S. Metabolic variability of a multispecies probiotic preparation impacts on the anti-inflammatory activity. Front. Pharmacol. 2017, 8, 505. [Google Scholar] [CrossRef]
- Parolin, C.; Marangoni, A.; Laghi, L.; Foschi, C.; Ñahui Palomino, R.A.; Calonghi, N.; Cevenini, R.; Vitali, B. Isolation of vaginal Lactobacilli and characterization of anti-Candida activity. PLoS ONE 2015, 10, e0131220. [Google Scholar] [CrossRef]
- Picone, G.; Laghi, L.; Gardini, F.; Lanciotti, R.; Siroli, L.; Capozzi, F. Evaluation of the effect of carvacrol on the Escherichia coli 555 metabolome by using 1H-NMR spectroscopy. Food Chem. 2013, 141, 4367–4374. [Google Scholar] [CrossRef]
- Liu, S.Q.; Holland, R.; McJarrow, P.; Crow, V.L. Serine metabolism in Lactobacillus plantarum. Int. J. Food Microbiol. 2003, 89, 265–273. [Google Scholar] [CrossRef]


| Compound | Concentration (g/L) |
|---|---|
| Macro components | |
| Sodium acetate | 2.0 |
| K2HPO4 | 1.75 |
| MnSO4 4H2O | 0.012 |
| MgSO4 7H2O | 0.2 |
| Tween 80 | 1 mL |
| Vitamins | |
| Thiamine HCl | 0.0011 |
| Folic acid | 0.0002 |
| Riboflavin | 0.001 |
| Calcium pantothenate | 0.001 |
| Nicotinic acid | 0.001 |
| Pyridoxal-5-phosphate | 0.0005 |
| p-amino benzoic acid | 0.0004 |
| Nucleotides | |
| Adenine | 0.005 |
| Guanine | 0.01 |
| Uracil | 0.01 |
| Sample | Time | Cell Counts | Growth Parameters (Gompertz Equation) | |||
|---|---|---|---|---|---|---|
| (Log CFU/mL) | k | A | µmax | λ | ||
| 0 h | 6.94 (±0.21) | |||||
| 2.5 G | 24 h | 8.15 (±0.23) | 0.044 | 0.264 | 0.155 | 11.457 |
| 48 h | 6.67 (±0.11) | |||||
| 2.5 R | 24 h | 8.41 (±0.16) | 0.046 | 0.282 | 0.120 | 11.570 |
| 48 h | 7.62 (±0.13) | |||||
| 25 G | 24 h | 8.13 (±0.21) | 0.045 | 1.446 | 0.280 | 10.839 |
| 48 h | 6.98 (±0.13) | |||||
| 25 R | 24 h | 8.10 (±0.11) | 0.074 | 1.151 | 0.275 | 11.257 |
| 48 h | 6.89 (±0.14) | |||||
| Sample | Time | L-Lactic Acid (mM) | D-Lactic Acid (mM) | Acetic Acid (mM) | pH |
|---|---|---|---|---|---|
| 2.5 G | 24 h | 4.25 (±0.09) | - * | 0.52 (±0.20) | 5.78 (±0.29) |
| 48 h | 3.88 (±0.14) | - | 0.62 (±0.15) | 5.83 (±0.16) | |
| 2.5 R | 24 h | 1.66 (±0.07) | - | 2.85 (±0.11) | 6.17 (±0.30) |
| 48 h | 1.39 (±0.04) | - | 3.15 (±0.09) | 6.21 (±0.27) | |
| 25 G | 24 h | 39.60 (±1.98) | 0.41 (±0.09) | 2.58 (±0.03) | 4.15 (±0.17) |
| 48 h | 43.81 (±2.05) | 0.39 (±0.10) | 2.77 (±0.28) | 4.00 (±0.22) | |
| 25 R | 24 h | 15.61 (±0.90) | 1.43 (±0.20) | 21.00 (±0.43) | 4.73 (±0.21) |
| 48 h | 16.02 (±0.63) | 1.09 (±0.14) | 22.70 (±0.67) | 4.37 (±0.24) |
| Asp | Ser + Asg | Glu | Gly | His + Glm | Arg | Thr | Ala | Pro | Cys | Tyr | Val | Met | Lys | Ile | Leu | Phe | ||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Initial concentration | 1.50 | 3.42 | 1.36 | 2.66 | 2.66 | 1.15 | 1.68 | 2.24 | 1.74 | 1.65 | 1.10 | 1.71 | 1.34 | 1.37 | 1.52 | 1.52 | 1.21 | |
| Sample | Time | |||||||||||||||||
| 2.5 G | 24 h | 1.39 (−7.49) | 3.15 (−7.94) | 1.27 (−6.53) | 2.91 (9.53) | 2.25 (−15.27) | 1.01 (−12.00) | 1.79 (6.57) | 2.12 (−1.09) | 1.92 (10.49) | 2.18 (32.02) | 1.11 (0.85) | 1.84 (7.68) | 1.27 (−5.45) | 1.36 (−0.45) | 1.50 (−1.54) | 1.39 (−8.51) | 1.28 (6.02) |
| 48 h | 1.27 (−15.53) | 2.65 (−22.41) | 1.07 (−21.36) | 2.50 (−5.97) | 1.85 (−30.44) | 0.29 (−74.57) | 1.80 (7.29) | 2.03 (−9.22) | 1.80 (3.20) | 2.45 (48.56) | 1.04 (−5.83) | 1.66 (−2.90) | 0.69 (−48.88) | 1.44 (4.96) | 1.53 (0.68) | 1.49 (−2.22) | 1.41 (16.51) | |
| 2.5 R | 24 h | 1.32 (−12.26) | 2.98 (−13.00) | 1.18 (−13.50) | 2.81 (5.61) | 2.13 (−19.94) | 0.10 (−91.57) | 1.65 (−1.87) | 2.21 (−1.38) | 1.78 (2.09) | 2.01 (21.78) | 1.10 (0.12) | 1.69 (−0.96) | 1.36 (1.18) | 1.36 (−0.67) | 1.58 (3.63) | 1.43 (−6.03) | 1.26 (4.53) |
| 48 h | 1.44 (−11.04) | 2.58 (−24.42) | 1.08 (−20.47) | 2.48 (−6.82) | 1.79 (−32.85) | 0.19 (−83.76) | 1.71 (2.08) | 2.02 (−10.03) | 1.77 (1.75) | 2.53 (53.09) | 1.14 (3.93) | 1.62 (−5.05) | 0.94 (−30.22) | 1.41 (2.99) | 1.53 (0.39) | 1.50 (−1.58) | 1.34 (10.72) | |
| 25 G | 24 h | 1.30 (−4.00) | 1.21 (−64.72) | 1.73 (27.56) | 2.71 (1.78) | 1.45 (−45.33) | 0.90 (−21.38) | 1.56 (−6.93) | 1.92 (−14.09) | 1.69 (−3.07) | 1.75 (6.31) | 1.07 (−2.87) | 1.59 (−7.27) | 1.31 (−2.42) | 1.23 (−9.96) | 1.59 (4.93) | 1.39 (−8.25) | 1.42 (17.71) |
| 48 h | 1.41 (−13.27) | 0.89 (−73.95) | 1.63 (19.88) | 2.33 (−12.43) | 1.10 (−58.54) | 0.80 (−30.78) | 1.36 (−19.13) | 1.73 (−22.80) | 1.75 (0.81) | 2.00 (21.05) | 1.02 (−7.05) | 1.50 (−12.17) | 1.26 (−6.27) | 1.10 (−19.61) | 1.39 (−8.81) | 1.45 (−4.67) | 1.32 (9.38) | |
| 25 R | 24 h | 1.28 (−14.55) | 2.05 (−40.17) | 1.33 (−2.17) | 2.62 (−1.41) | 1.80 (−32.31) | 0.09 (−92.08) | 1.77 (5.53) | 2.02 (−9.78) | 1.75 (0.30) | 1.43 (−13.05) | 1.01 (−7.88) | 1.59 (−6.93) | 1.15 (−14.50) | 1.21 (−11.82) | 1.49 (−2.04) | 1.46 (−4.22) | 1.35 (11.82) |
| 48 h | 1.20 (−20.06) | 1.83 (−46.41) | 1.18 (−13.32) | 2.35 (−11.60) | 1.50 (−43.72) | 0.12 (−89.31) | 1.64 (−2.14) | 1.86 (−17.08) | 1.80 (3.60) | 1.57 (−5.09) | 1.08 (−2.27) | 1.52 (−11.15) | 1.04 (−22.76) | 1.09 (−20.45) | 1.34 (−11.72) | 1.37 (−9.93) | 1.32 (9.45) | |
| Ethanol | Acetoin | 2,3-Butanediol | Glucose | Ribose | |
|---|---|---|---|---|---|
| 2.5 G | 0.01 | 0.10 | 0.01 | - * | - |
| 2.5 R | 0.01 | 0.15 | 0.01 | - | - |
| 25 G | 0.03 | 0.14 | 0.06 | 0.14 | - |
| 25 R | 0.01 | 0.21 | 0.19 | - | 0.15 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barbieri, F.; Laghi, L.; Gardini, F.; Montanari, C.; Tabanelli, G. Metabolism of Lactobacillus sakei Chr82 in the Presence of Different Amounts of Fermentable Sugars. Foods 2020, 9, 720. https://doi.org/10.3390/foods9060720
Barbieri F, Laghi L, Gardini F, Montanari C, Tabanelli G. Metabolism of Lactobacillus sakei Chr82 in the Presence of Different Amounts of Fermentable Sugars. Foods. 2020; 9(6):720. https://doi.org/10.3390/foods9060720
Chicago/Turabian StyleBarbieri, Federica, Luca Laghi, Fausto Gardini, Chiara Montanari, and Giulia Tabanelli. 2020. "Metabolism of Lactobacillus sakei Chr82 in the Presence of Different Amounts of Fermentable Sugars" Foods 9, no. 6: 720. https://doi.org/10.3390/foods9060720
APA StyleBarbieri, F., Laghi, L., Gardini, F., Montanari, C., & Tabanelli, G. (2020). Metabolism of Lactobacillus sakei Chr82 in the Presence of Different Amounts of Fermentable Sugars. Foods, 9(6), 720. https://doi.org/10.3390/foods9060720






