Assessment of RNAlater® as a Potential Method to Preserve Bovine Muscle Proteins Compared with Dry Ice in a Proteomic Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Tissue Sampling
2.2. Warner–Bratzler Shear Force Measurement
2.3. Extraction of Muscle Proteins
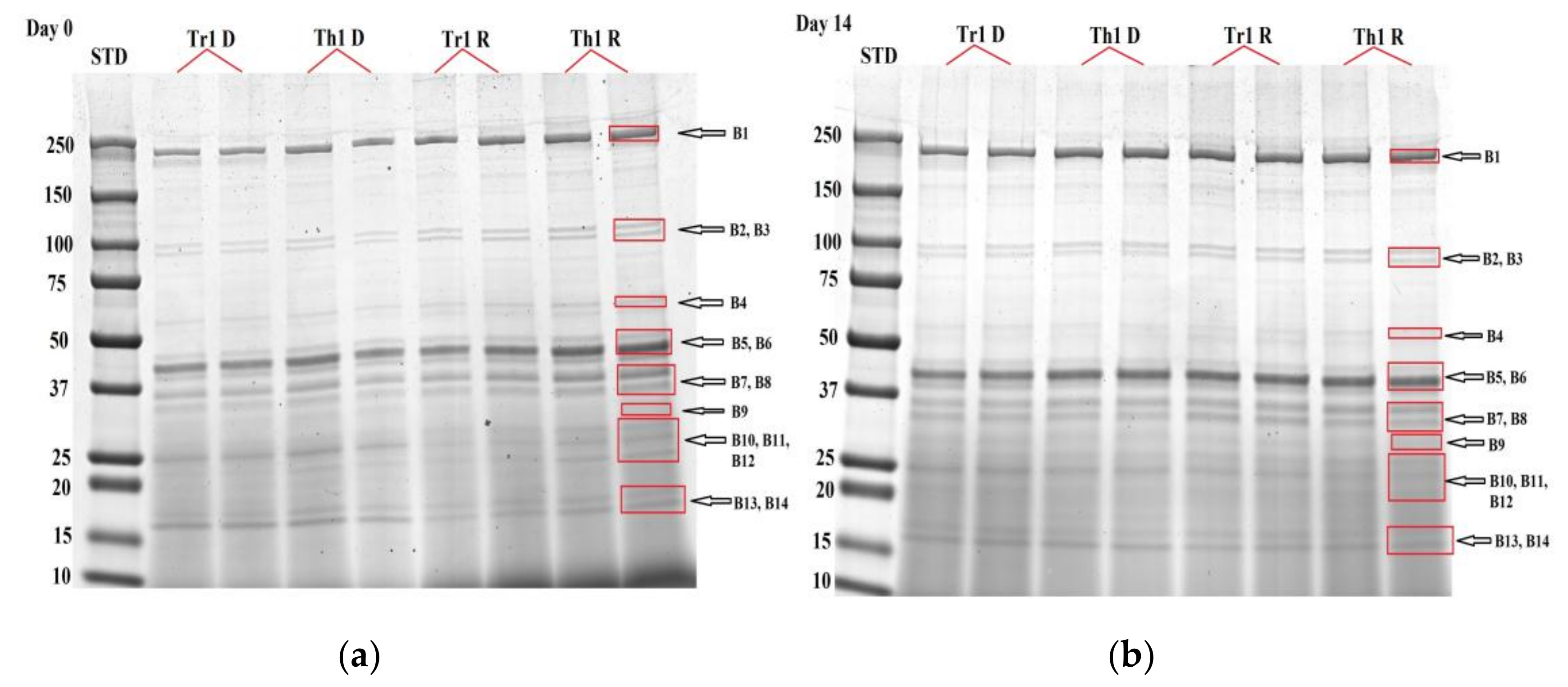
2.4. Proteomic Analysis
2.5. Image Analysis
2.6. Mass Spectrometry Analysis
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Saito, M.A.; Bulygin, V.V.; Moran, D.M.; Taylor, C.; Scholin, C. Examination of Microbial Proteome Preservation Techniques Applicable to Autonomous Environmental Sample Collection. Front. Microbiol. 2011, 2, 1–10. [Google Scholar] [CrossRef]
- Manoj, C.R.; Douglas, S.M.; Rebecca, A.R.; Kathleen, M.E. A Method for the Extraction of High-Quality RNA and Protein from Single Small Samples of Arteries and Veins Preserved in RNAlater. J. Pharmacol. Toxicol. Methods 2002, 47, 87–92. [Google Scholar]
- Bennike, T.B.; Kastaniegaard, K.; Padurariu, S.; Gaihede, M.; Birkelund, S.; Andersen, V.; Stensballe, A. Comparing the Proteome of Snap Frozen, RNAlater Preserved, and Formalin-Fixed Paraffin-Embedded Human Tissue Samples. EuPA Open Proteom. 2016, 10, 9–18. [Google Scholar] [CrossRef]
- Rader, J.S.; James, P.M.; Julia, G.; Petra, G.; Rebecca, A.B.; Loan, N.; Dan, L.C. A Unified Sample Preparation Protocol for Proteomic and Genomic Profiling of Cervical Swabs to Identify Biomarkers for Cervical Cancer Screening. Proteomics 2008, 2, 1658–1669. [Google Scholar] [CrossRef]
- Lenchik, N.I.; Desiderio, D.M.; Gerling, I.C. Two-Dimensional Gel Electrophoresis Characterization of the Mouse Leukocyte Proteome, Using a Tri-Reagent for Protein Extraction. Proteomics 2005, 5, 202–209. [Google Scholar] [CrossRef] [PubMed]
- Barclay, D.; Ruben, Z.; Andres, T.; Rajaie, N.; David, S.; Yoram, V. A Simple, Rapid, and Convenient LuminexTM-Compatible Method of Tissue Isolation. J. Clin. Lab. Anal. 2008, 22, 278–281. [Google Scholar] [CrossRef] [PubMed]
- Kruse, C.P.S.; Basu, P.; Luesse, D.R.; Wyatt, S.E. Transcriptome and Proteome Responses in RNAlater Preserved Tissue of Arabidopsis Thaliana. PLoS ONE 2017, 12, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Abbaraju, N.V.; Yang, C.; Bernard, B.R. Protein Recovery and Identification from the Gulf Killifish, Fundulus Grandis: Comparing Snap-Frozen and RNAlater® Preserved Tissues. Proteomics 2011, 11, 4257–4261. [Google Scholar] [CrossRef] [PubMed]
- Belew, J.B.; Brooks, J.C.; McKenna, D.R.; Savell, J.W. Warner–Bratzler Shear Evaluations of 40 Bovine Muscles. Meat Sci. 2003, 64, 507–512. [Google Scholar] [CrossRef]
- Bradford, M.M. A Rapid and Sensitive Method for the Quantitation of Microgram Quantities of Protein Utilizing the Principle of Protein-Dye Binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of Structural Proteins during the Assembly of the Head of Bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- Wilm, M.; Shevchenko, A.; Houthaeve, T.; Breit, S.; Schweigerer, L.; Fotsis, T.; Mann, M. Femtomole sequencing of proteins from polyacrylamide gels by nano-electrospray mass spectrometry. Nature 1996, 379, 466–469. [Google Scholar] [CrossRef] [PubMed]
- Buts, B.; Claeys, E.; Demyer, D. Relation between concentration of troponin-T, 30,000-Dalton, and titin on SDS–PAGE and tenderness of bull longissimus dorsi. In Proceedings of the 32nd European Meeting of Meat Research Workers, Ghent, Belgium, 24–29 August 1986. [Google Scholar]
- MacBride, M.A.; Parrish, F.C. 30,000-Dalton component of tender bovine longissimus muscle. J. Food Sci. 1977, 42, 1627–1629. [Google Scholar] [CrossRef]
- Ho, C.Y.; Stromer, M.H.; Robson, R.M. Identification of the 30 kDa Polypeptide in Post Mortem Skeletal Muscle as a Degradation Product of Troponin-T. Biochimie 1994, 76, 369–375. [Google Scholar] [CrossRef]
- Olson, D.G.; Parrish, F.C.; Dayton, W.R.; Goll, D.E. Effect of postmortem storage and calcium activated factor on myofibrillar proteins of bovine skeletal muscle. J. Food Sci. 1977, 42, 117–124. [Google Scholar] [CrossRef]
| Bands No. | Quality | Treatment | Time Point | Quality × Treatment | Quality × Time Point | Treat × Time Point |
|---|---|---|---|---|---|---|
| 1 | 0.93 | 0.90 | 0.17 | 0.52 | 0.50 | 0.94 |
| 2 | 0.58 | 0.25 | 0.42 | 0.57 | 0.67 | 0.60 |
| 3 | 0.51 | 0.51 | 0.52 | 0.22 | 0.61 | 0.24 |
| 4 | 0.91 | 0.78 | 0.93 | 0.69 | 0.77 | 0.71 |
| 5 | 0.34 | 0.86 | 0.15 | 0.08 Ϯ | 0.81 | 0.91 |
| 6 | 0.51 | 0.83 | 0.87 | 0.13 | 0.74 | 0.87 |
| 7 | 0.74 | 0.23 | 0.01 | 0.43 | 0.31 | 0.39 |
| 8 | 0.98 | 0.12 | 0.20 | 0.94 | 0.72 | 0.63 |
| 9 | 0.30 | 0.33 | <001 | 0.46 | 0.38 | 0.88 |
| 10 | 0.87 | 0.81 | 0.61 | 0.84 | 0.21 | 0.79 |
| 11 | 0.67 | 0.08 Ϯ | 0.69 | 0.87 | 0.71 | 0.77 |
| 12 | 0.83 | 0.61 | 0.86 | 0.91 | 0.62 | 0.54 |
| 13 | 0.64 | 0.60 | 0.45 | 0.45 | 0.68 | 0.68 |
| 14 | 0.63 | 0.21 | 0.66 | 0.78 | 0.96 | 0.88 |
| Dry Ice | RNAlater® | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Day 0 | Day 14 | e.s.e. | Tender | Tough | e.s.e. | Day 0 | Day 14 | e.s.e. | Tender | Tough | e.s.e. | |
| Band 7 | 0.076 ab | 0.052 a | 0.008 | 0.066 | 0.061 | 0.008 | 0.101 b | 0.056 ab | 0.014 | 0.072 | 0.085 | 0.014 |
| Band 9 | 0.005 a | 0.021 b | 0.002 | 0.012 | 0.015 | 0.002 | 0.003 a | 0.020 b | 0.002 | 0.011 | 0.012 | 0.002 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhu, Y.; Mullen, A.M.; Rai, D.K.; Kelly, A.L.; Sheehan, D.; Cafferky, J.; Hamill, R.M. Assessment of RNAlater® as a Potential Method to Preserve Bovine Muscle Proteins Compared with Dry Ice in a Proteomic Study. Foods 2019, 8, 60. https://doi.org/10.3390/foods8020060
Zhu Y, Mullen AM, Rai DK, Kelly AL, Sheehan D, Cafferky J, Hamill RM. Assessment of RNAlater® as a Potential Method to Preserve Bovine Muscle Proteins Compared with Dry Ice in a Proteomic Study. Foods. 2019; 8(2):60. https://doi.org/10.3390/foods8020060
Chicago/Turabian StyleZhu, Yao, Anne Maria Mullen, Dilip K. Rai, Alan L. Kelly, David Sheehan, Jamie Cafferky, and Ruth M. Hamill. 2019. "Assessment of RNAlater® as a Potential Method to Preserve Bovine Muscle Proteins Compared with Dry Ice in a Proteomic Study" Foods 8, no. 2: 60. https://doi.org/10.3390/foods8020060
APA StyleZhu, Y., Mullen, A. M., Rai, D. K., Kelly, A. L., Sheehan, D., Cafferky, J., & Hamill, R. M. (2019). Assessment of RNAlater® as a Potential Method to Preserve Bovine Muscle Proteins Compared with Dry Ice in a Proteomic Study. Foods, 8(2), 60. https://doi.org/10.3390/foods8020060







