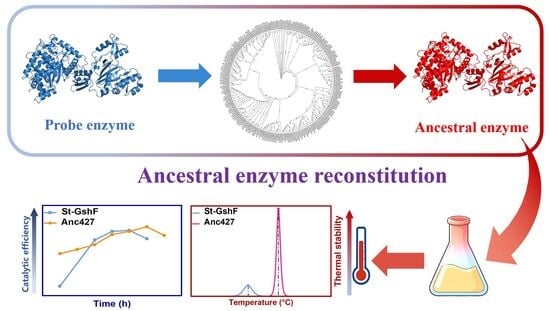
Ancestral Sequence Reconstruction for Novel Bifunctional Glutathione Synthetase with Enhanced Thermostability and Catalytic Efficiency
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals, Strains, and Plasmids
2.2. Ancestral Sequence Reconstruction
2.3. Protein Expression and Induction
2.4. Protein Purification
2.5. Enzyme Activity Assay
2.6. GSH Content Analysis
2.6.1. DTNB Spectrophotometric Assay
2.6.2. High-Performance Liquid Chromatography (HPLC) Analysis
2.7. Optimization of Enzymatic Reaction Conditions
2.8. Thermostability Analysis
2.9. Reaction with Elevated Concentration Substrate
2.10. Steady-State Kinetic Analysis
2.11. Molecular Dynamics (MD) Simulations
3. Results and Discussion
3.1. Ancestral Sequence Reconstruction for GshF
3.2. Heterologous Expression and Purification of the Ancestral Enzyme
3.3. Characterization of the Ancestral and Probe Enzymes
3.4. Thermostability of the Ancestral Enzyme and Probe Enzymes
3.5. High-Substrate Concentration Reaction
Structural Modeling and MD Simulations

4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| LB | Luria–Bertani |
| TB | Terrific Broth |
| GSH | Glutathione |
| PBS | Phosphate-buffered saline |
| Tris-HCl | Tris(hydroxymethyl)aminomethane hydrochloride |
| IPTG | Isopropyl β-d-1-thiogalactopyranoside |
| MMseqs2 | Many-against-Many sequence searching |
| PAML-X | Phylogenetic Analysis by Maximum Likelihood, X version |
| DTNB | Ellman’s Reagent Method |
| HPLC | High-Performance Liquid Chromatography |
| MD | Molecular dynamics |
| GshF | Bifunctional Glutathione Synthase |
| St-GshF | GshF from Streptococcus thermophilus |
| Anc427 | The node 427 ancestral enzyme |
| SDS-PAGE | Sodium dodecyl sulfate–polyacrylamide gel electrophoresis |
| Tm | Thermal denaturation temperature |
| T1/2 | Half-life |
| ASR | Ancestral sequence reconstruction |
| RMSD | Root-mean-square deviation |
| AlphaFold2 | A deep learning–based protein structure prediction algorithm developed by DeepMind |
| SAVES v6.1 | Structure Analysis and Verification Server, UCLA, USA |
| AKTA pure | Automated fast protein liquid chromatography system (Cytiva, Uppsala, Sweden) |
References
- Lapenna, D. Glutathione and glutathione-dependent enzymes: From biochemistry to gerontology and successful aging. Ageing Res. Rev. 2023, 92, 102066. [Google Scholar] [CrossRef]
- Gasmi, A.; Nasreen, A.; Lenchyk, L.; Lysiuk, R.; Peana, M.; Shapovalova, N.; Piscopo, S.; Komisarenko, M.; Shanaida, M.; Smetanina, K.; et al. An update on glutathione’s biosynthesis, metabolism, functions, and medicinal purposes. Curr. Med. Chem. 2024, 31, 4579–4601. [Google Scholar] [CrossRef]
- Al-Temimi, A.A.; Al-Mossawi, A.-E.-B.; Al-Hilifi, S.A.; Korma, S.A.; Esatbeyoglu, T.; Rocha, J.M.; Agarwal, V. Glutathione for food and health applications with emphasis on extraction, identification, and quantification methods: A review. Metabolites 2023, 13, 465. [Google Scholar] [CrossRef]
- Weschawalit, S.; Thongthip, S.; Phutrakool, P.; Asawanonda, P. Glutathione and its antiaging and ant melanogenic effects. Clin. Cosmet. Investig. Dermatol. 2017, 10, 147–153. [Google Scholar] [CrossRef]
- Forman, H.J.; Zhang, H.; Rinna, A. Glutathione: Overview of its protective roles, measurement, and biosynthesis. Mol. Asp. Med. 2009, 30, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Bitwell, C.; Indra, S.S.; Luke, C.; Kakoma, M.K. A review of modern and conventional extraction techniques and their applications for extracting phytochemicals from plants. Sci. Afr. 2023, 19, e01585. [Google Scholar] [CrossRef]
- Alqahtani, A.S.; Elbeltagi, S. Advancing chemistry sustainably: From synthesis to benefits and applications of green synthesis. J. Organomet. Chem. 2025, 1027, 123508. [Google Scholar] [CrossRef]
- Cui, X.; Wan, J.; Zhang, X.; Wu, H.; Li, Z.; Ye, Q. Efficient glutathione production in metabolically engineered Escherichia coli strains using constitutive promoters. J. Biotechnol. 2019, 289, 39–45. [Google Scholar] [CrossRef]
- Lin, H.; Wang, L.; Jiang, X.; Wang, J. Glutathione dynamics in subcellular compartments and implications for drug de-velopment. Curr. Opin. Chem. Biol. 2024, 81, 102505. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Su, M.; Liu, Q.; Li, Y.; Liang, Y.; Li, S.; Li, Y. Systems metabolic engineering of glutathione biosynthesis in saccharomyces cerevisiae: Pathway balancing coupled with enzyme screening for high-titer production. Eng. Microbiol. 2025, 5, 100243. [Google Scholar] [CrossRef]
- Cao, H.; Li, C.; Zhao, J.; Wang, F.; Tan, T.; Liu, L. Enzymatic production of glutathione coupling with an atp regeneration system based on polyphosphate kinase. Appl. Biochem. Biotechnol. 2018, 185, 385–395. [Google Scholar] [CrossRef] [PubMed]
- Zhu, W.; Sun, H.; Jiang, Q.; Zheng, R.; Wang, Q.; Zhang, Q.; Liu, L.; Cao, H. Enhancing the thermal stability of glutathione bifunctional synthase by b-factor strategy and un/folding free energy calculation. Catalysts 2022, 12, 1649. [Google Scholar] [CrossRef]
- Musatti, A.; Devesa, V.; Calatayud, M.; Vélez, D.; Manzoni, M.; Rollini, M. Glutathione-enriched baker’s yeast: Production, bioaccessibility and intestinal transport assays. J. Appl. Microbiol. 2014, 116, 304–313. [Google Scholar] [CrossRef]
- Jiang, Y.; Tao, R.; Shen, Z.; Sun, L.; Zhu, F.; Yang, S. Enzymatic production of glutathione by bifunctional γ-glutamyl cysteine synthetase/glutathione synthetase coupled with in vitro acetate kinase-based atp generation. Appl. Biochem. Biotechnol. 2016, 180, 1446–1455. [Google Scholar] [CrossRef] [PubMed]
- Ge, S.; Zhu, T.; Li, Y. Expression of bacterial gshf in Pichia pastoris for glutathione production. Appl. Environ. Microbiol. 2012, 78, 5435–5439. [Google Scholar] [CrossRef] [PubMed]
- Stout, J.; Vos, D.D.; Vergauwen, B.; Savvides, S.N. Glutathione biosynthesis in bacteria by bifunctional gshf is driven by a modular structure featuring a novel hybrid atp-grasp fold. J. Mol. Biol. 2012, 416, 486–494. [Google Scholar] [CrossRef]
- Cui, X.; Li, Z. High production of glutathione by in vitro enzymatic cascade after thermostability enhancement. AIChE J. 2021, 67, e17055. [Google Scholar] [CrossRef]
- Shi, J.; Yuan, B.; Yang, H.; Sun, Z. Recent advances on protein engineering for improved stability. BioDesign Res. 2025, 7, 100005. [Google Scholar] [CrossRef]
- Xiong, W.; Liu, B.; Shen, Y.; Jing, K.; Savage, T.R. Protein engineering design from directed evolution to de novo synthesis. Biochem. Eng. J. 2021, 174, 108096. [Google Scholar] [CrossRef]
- Drufva, E.E.; Hix, E.G.; Bailey, C.B. Site directed mutagenesis as a precision tool to enable synthetic biology with engineered modular polyketide synthases. Synth. Syst. Biotechnol. 2020, 5, 62–80. [Google Scholar] [CrossRef]
- Nirantar, S.R. Directed evolution methods for enzyme engineering. Molecules 2021, 26, 5599. [Google Scholar] [CrossRef]
- Fan, L.; Wang, H.; Gao, H.; Ding, Y.; Zhao, J.; Luo, H.; Tu, T.; Wu, N.; Yao, B.; Guan, F.; et al. Rational protein engineering using an omni-directional multipoint mutagenesis generation pipeline. iScience 2025, 28, 113273. [Google Scholar] [CrossRef]
- Liu, Q.L.; Liu, N.; Huang, H.W.; Su, C.; Xu, J.G.; Xu, G.Q.; Zhang, X.M.; Shi, J.S.; Xu, Z.H. Molecular modification and whole-cell catalytic optimization of bifunctional glutathione synthase. China Biotechnol. 2024, 44, 1–11. [Google Scholar] [CrossRef]
- Prakinee, K.; Phaisan, S.; Kongjaroon, S.; Chaiyen, P. Ancestral sequence reconstruction for designing biocatalysts and investigating their functional mechanisms. JACS Au 2024, 4, 4571–4591. [Google Scholar] [CrossRef] [PubMed]
- Spence, M.A.; Kaczmarski, J.A.; Saunders, J.W.; Jackson, C.J. Ancestral sequence reconstruction for protein engineers. Curr. Opin. Struct. Biol. 2021, 69, 131–141. [Google Scholar] [CrossRef]
- Niketan, S. Evolutionary insights into enzymes: Unraveling the molecular history of biological catalysts. J. Biochem. Res. 2023, 6, 103–106. [Google Scholar]
- Wang, S.; Gribskov, M. Finding homologous sequences using Uniprot. In Practical Bioinformatics: A Laboratory Manual; Springer Nature: Singapore, 2025; pp. 53–62. [Google Scholar]
- Wang, S.; Gribskov, M. Finding homologous sequences using BLAST. In Practical Bioinformatics: A Laboratory Manual; Springer Nature: Singapore, 2025; pp. 41–52. [Google Scholar]
- Menardo, F.; Loiseau, C.; Brites, D.; Coscolla, M.; Gygli, S.M.; Rutaihwa, L.K.; Trauner, A.; Beisel, C.; Borrell, S.; Gagneux, S. Treemmer: A tool to reduce large phylogenetic datasets with minimal loss of diversity. BMC Bioinform. 2018, 19, 164. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular evolutionary genetics analysis (mega) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Trifinopoulos, J.; Nguyen, L.-T.; von Haeseler, A.; Minh, B.Q. W-IQ-TREE: A fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 2016, 44, W232–W235. [Google Scholar] [CrossRef]
- Kulikov, N.; Derakhshandeh, F.; Mayer, C. Machine learning can be as good as maximum likelihood when recon-structing phylogenetic trees and determining the best evolutionary model on four taxon alignments. Mol. Phylogenet. Evol. 2024, 200, 108181. [Google Scholar] [CrossRef]
- Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 2007, 24, 1586–1591. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Sun, H.; Wei, L.; Hu, R.; Liu, H.; Lai, Y.; Li, X. Robust and sensitive colorimetric detection of glutathione with double-triggering MOF-Fe (DTNB). Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2025, 328, 125439. [Google Scholar] [CrossRef] [PubMed]
- Biterova, E.I.; Barycki, J.J. Mechanistic details of glutathione biosynthesis revealed by crystal structures of Saccharomyces cerevisiae glutamate cysteine ligase. J. Biol. Chem. 2009, 284, 32700–32708. [Google Scholar] [CrossRef]
- Robert, X.; Guillon, C.; Gouet, P. FoldScript: A web server for the efficient analysis of AI-generated 3D protein models. Nucleic Acids Res. 2025, 53, w277–w282. [Google Scholar] [CrossRef]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef]
- Gouet, P.; Robert, X.; Courcelle, E. ESPript/ENDscript: Extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 2003, 31, 3320–3323. [Google Scholar] [CrossRef]
- Gouet, P.; Courcelle, E. ENDscript: A workflow to display sequence and structure information. Bioinformatics 2002, 18, 767–768. [Google Scholar] [CrossRef] [PubMed]
- Gouet, P.; Courcelle, E.; Stuart, D.I.; Métoz, F. ESPript: Analysis of multiple sequence alignments in PostScript. Bioinformatics 1999, 15, 305–308. [Google Scholar] [CrossRef] [PubMed]
- Laskowski, R.A.; Rullmannn, J.A.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef]







Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Zhao, J.; Wang, B.; Di, J.; Zhou, J.; Dong, J.; Ni, Y.; Han, R. Ancestral Sequence Reconstruction for Novel Bifunctional Glutathione Synthetase with Enhanced Thermostability and Catalytic Efficiency. Foods 2026, 15, 309. https://doi.org/10.3390/foods15020309
Zhao J, Wang B, Di J, Zhou J, Dong J, Ni Y, Han R. Ancestral Sequence Reconstruction for Novel Bifunctional Glutathione Synthetase with Enhanced Thermostability and Catalytic Efficiency. Foods. 2026; 15(2):309. https://doi.org/10.3390/foods15020309
Chicago/Turabian StyleZhao, Jieru, Binhao Wang, Junhua Di, Jieyu Zhou, Jinjun Dong, Ye Ni, and Ruizhi Han. 2026. "Ancestral Sequence Reconstruction for Novel Bifunctional Glutathione Synthetase with Enhanced Thermostability and Catalytic Efficiency" Foods 15, no. 2: 309. https://doi.org/10.3390/foods15020309
APA StyleZhao, J., Wang, B., Di, J., Zhou, J., Dong, J., Ni, Y., & Han, R. (2026). Ancestral Sequence Reconstruction for Novel Bifunctional Glutathione Synthetase with Enhanced Thermostability and Catalytic Efficiency. Foods, 15(2), 309. https://doi.org/10.3390/foods15020309